Metabolomics Combined with Multivariate Statistical Analysis for Screening of Chemical Markers between Gentiana scabra and Gentiana rigescens
Abstract
1. Introduction
2. Result and Discussion
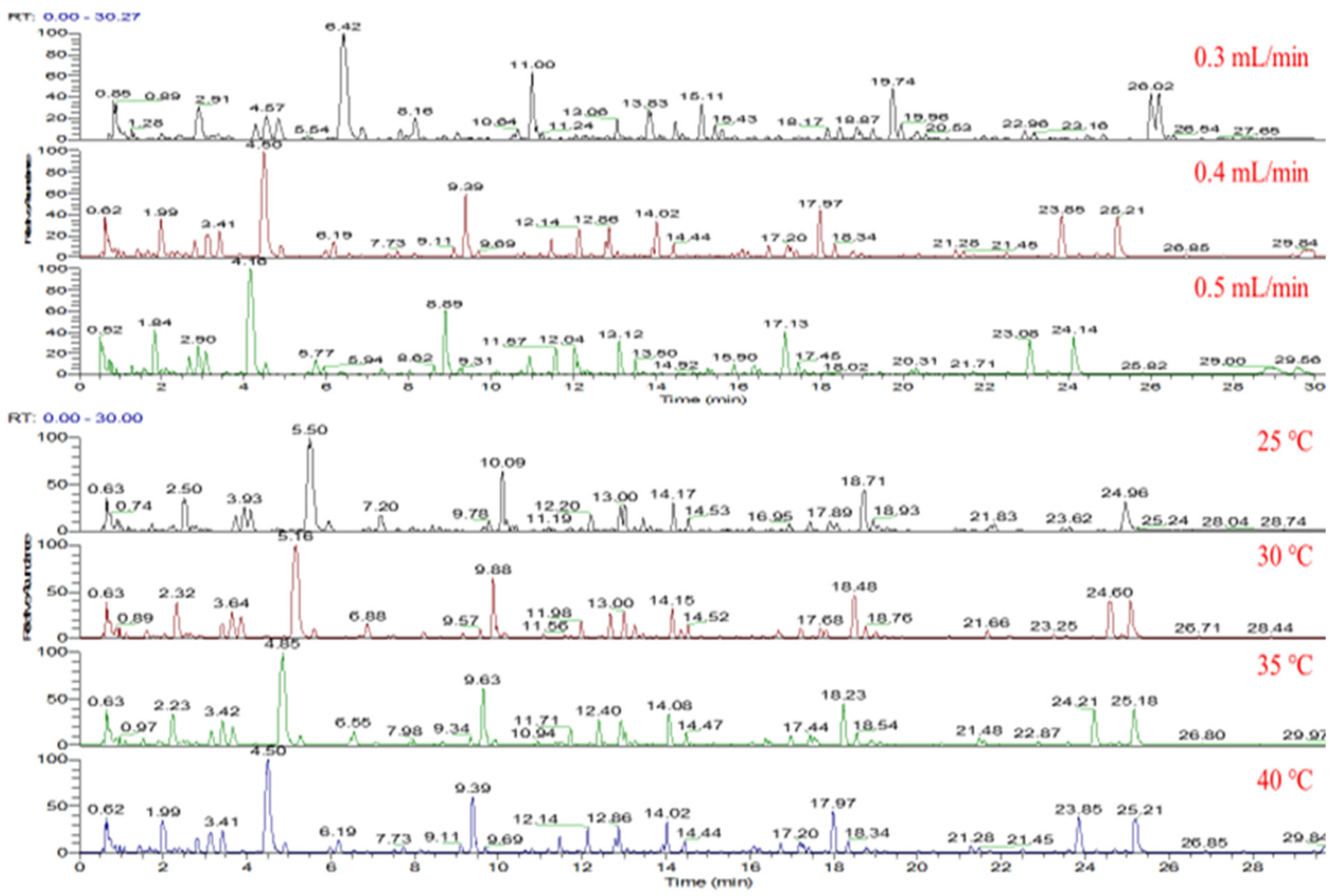
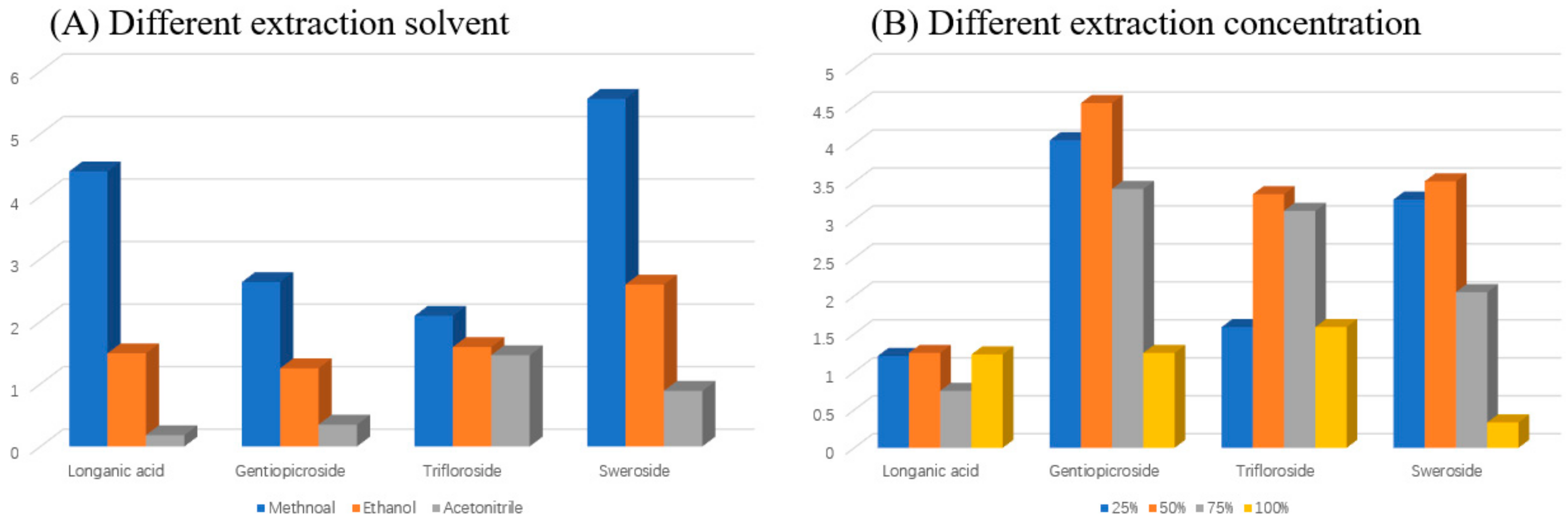
2.1. Optimization of Chromatographic Conditions and Sample Extraction
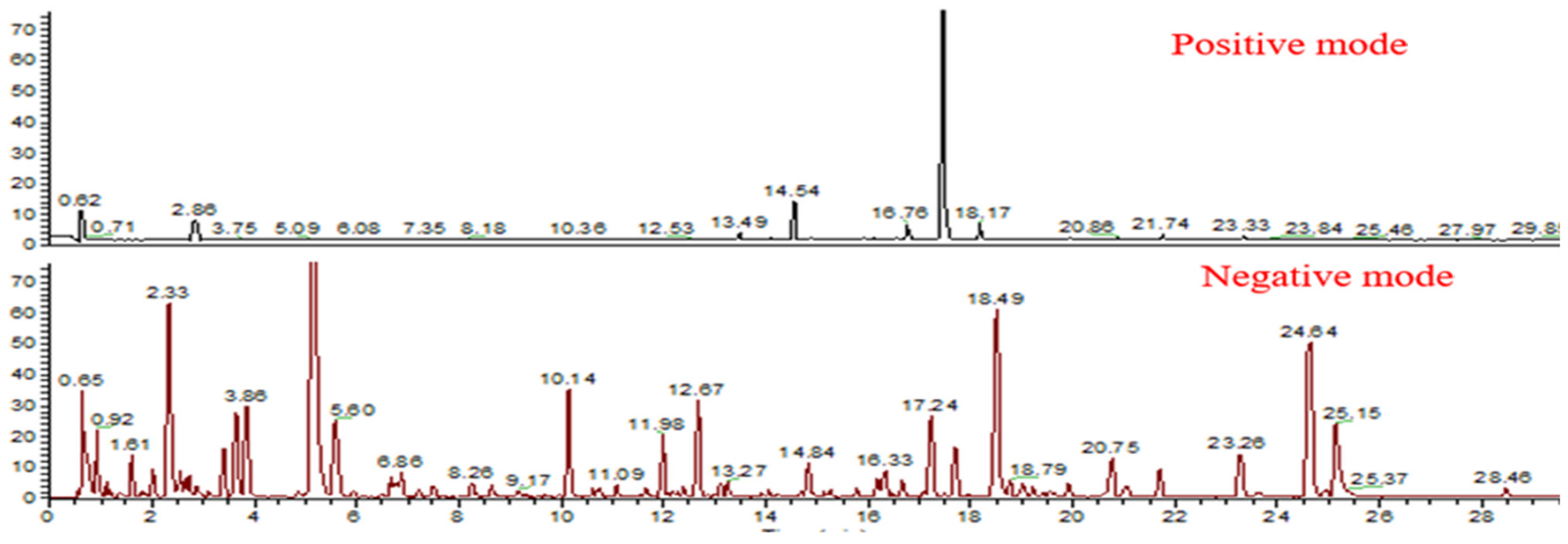
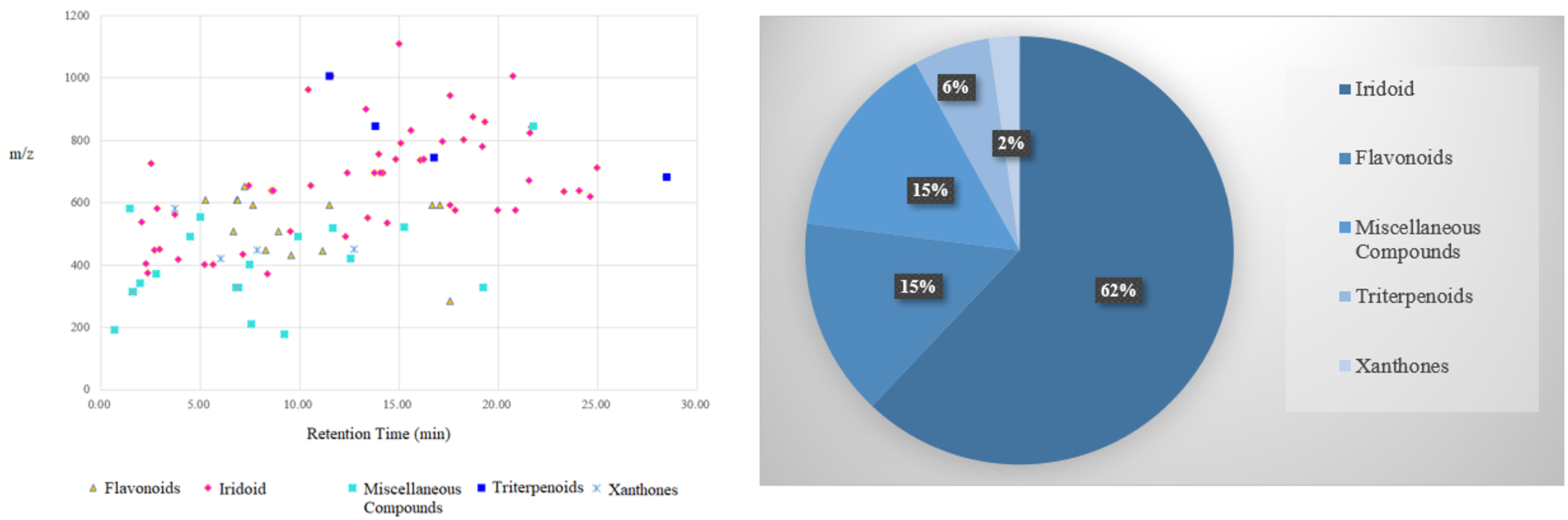
2.2. Characterization of Chemical Components by UPLC-LTQ-Orbitrap/MS
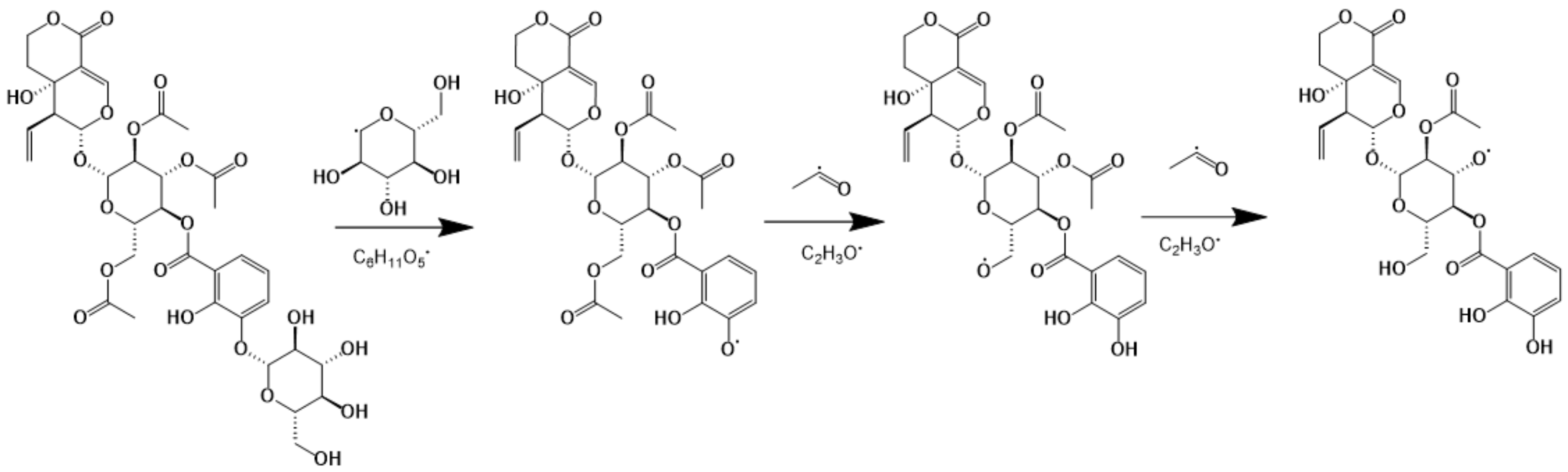
2.2.1. Deductive Fragmentation Pathway of Iridoids
2.2.2. Deductive Fragmentation Pathway of Flavonoids
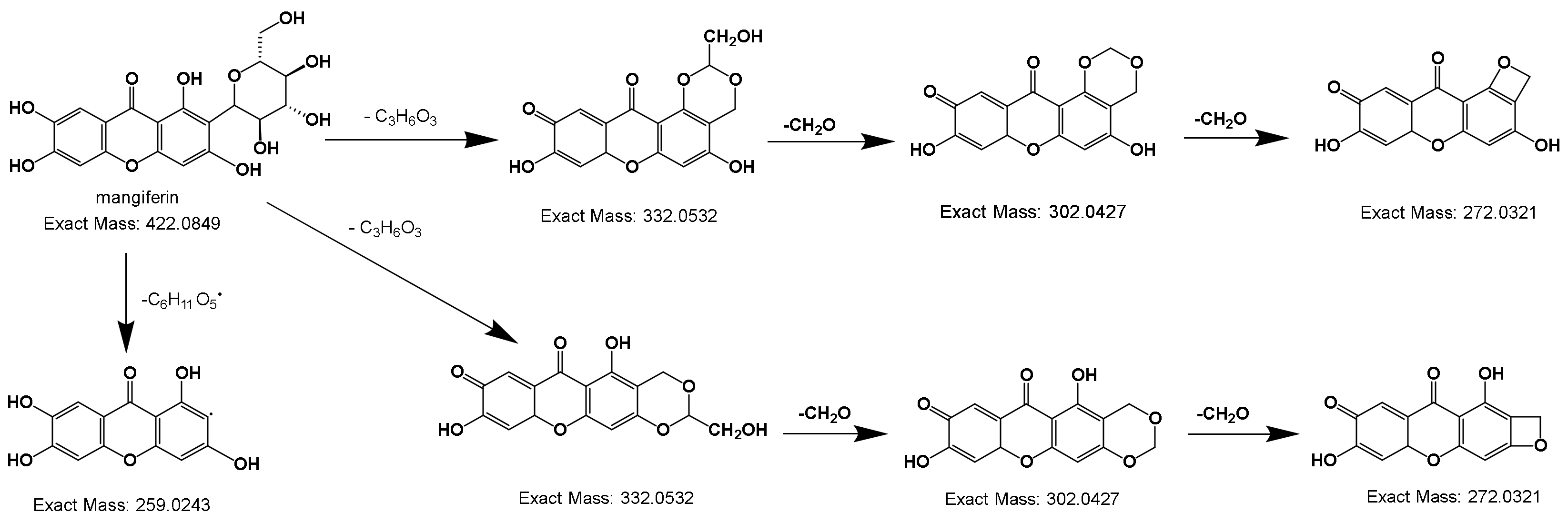
2.2.3. Deductive Fragmentation Pathway of Xanthones
2.3. Screening of Chemical Markers by Metabolomics and Chemometrics
2.3.1. Univariate Analysis for the Screening of Differential Metabolites
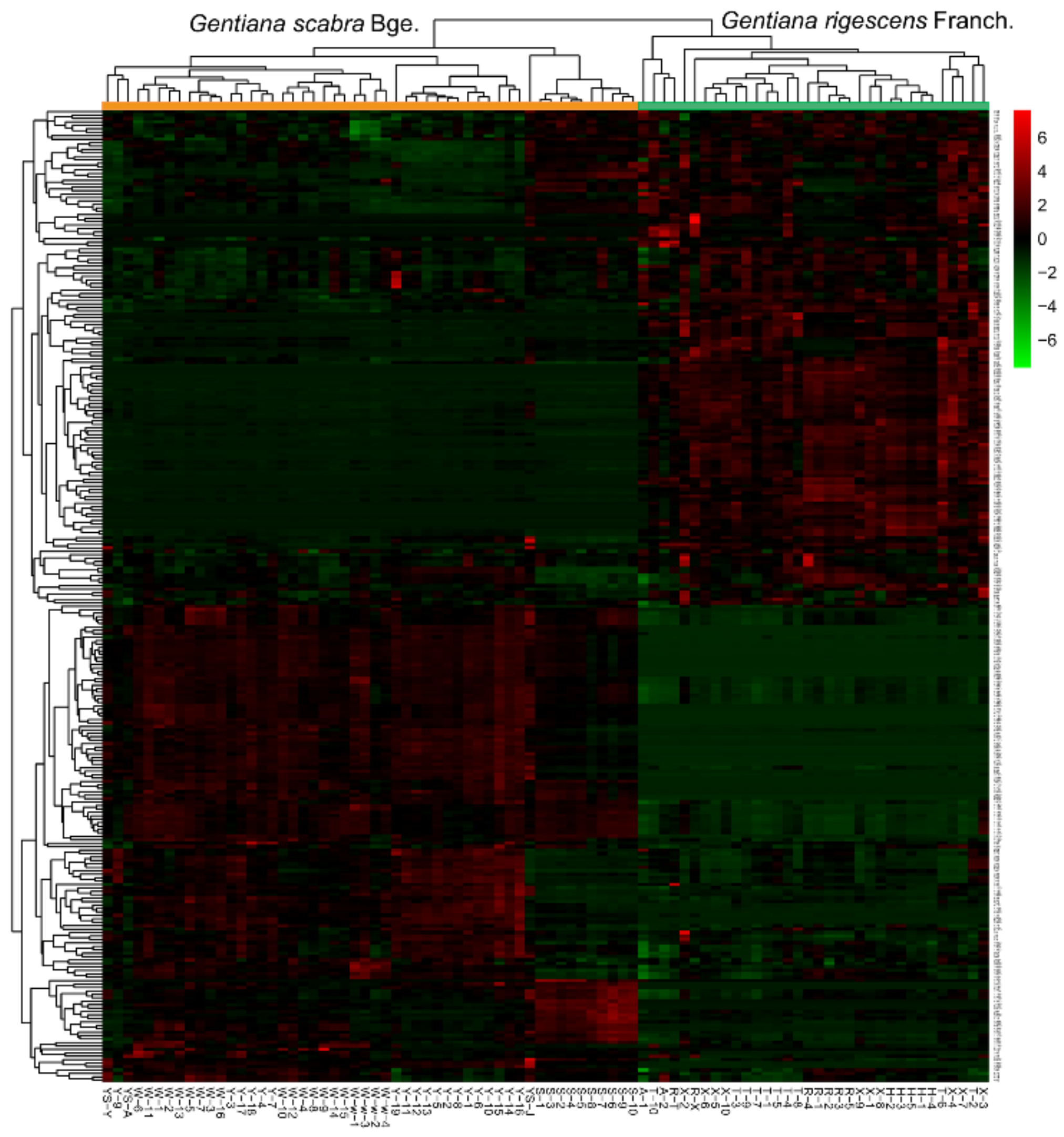
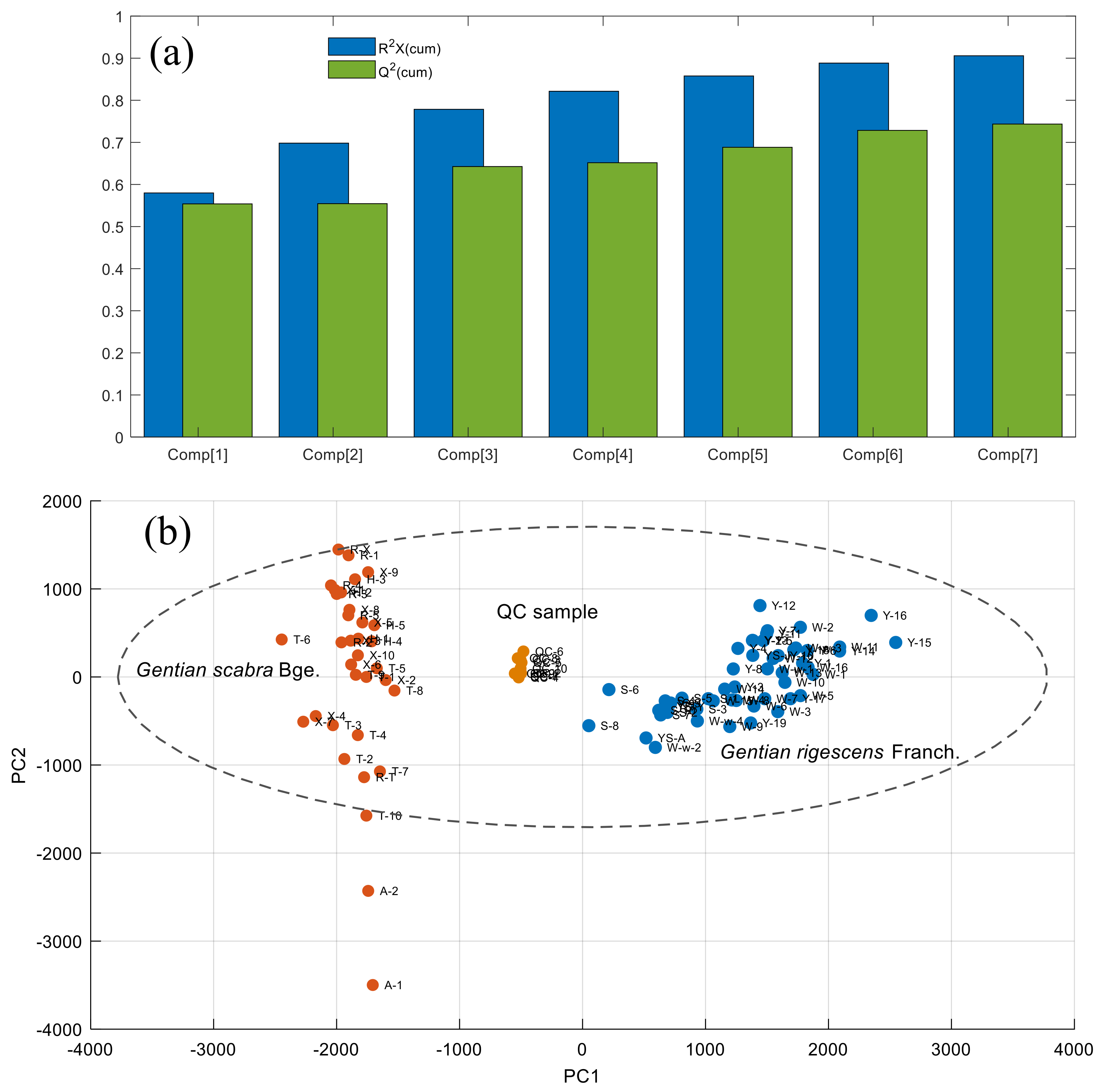
2.3.2. Data Visualization and Experimental Stability Evaluation
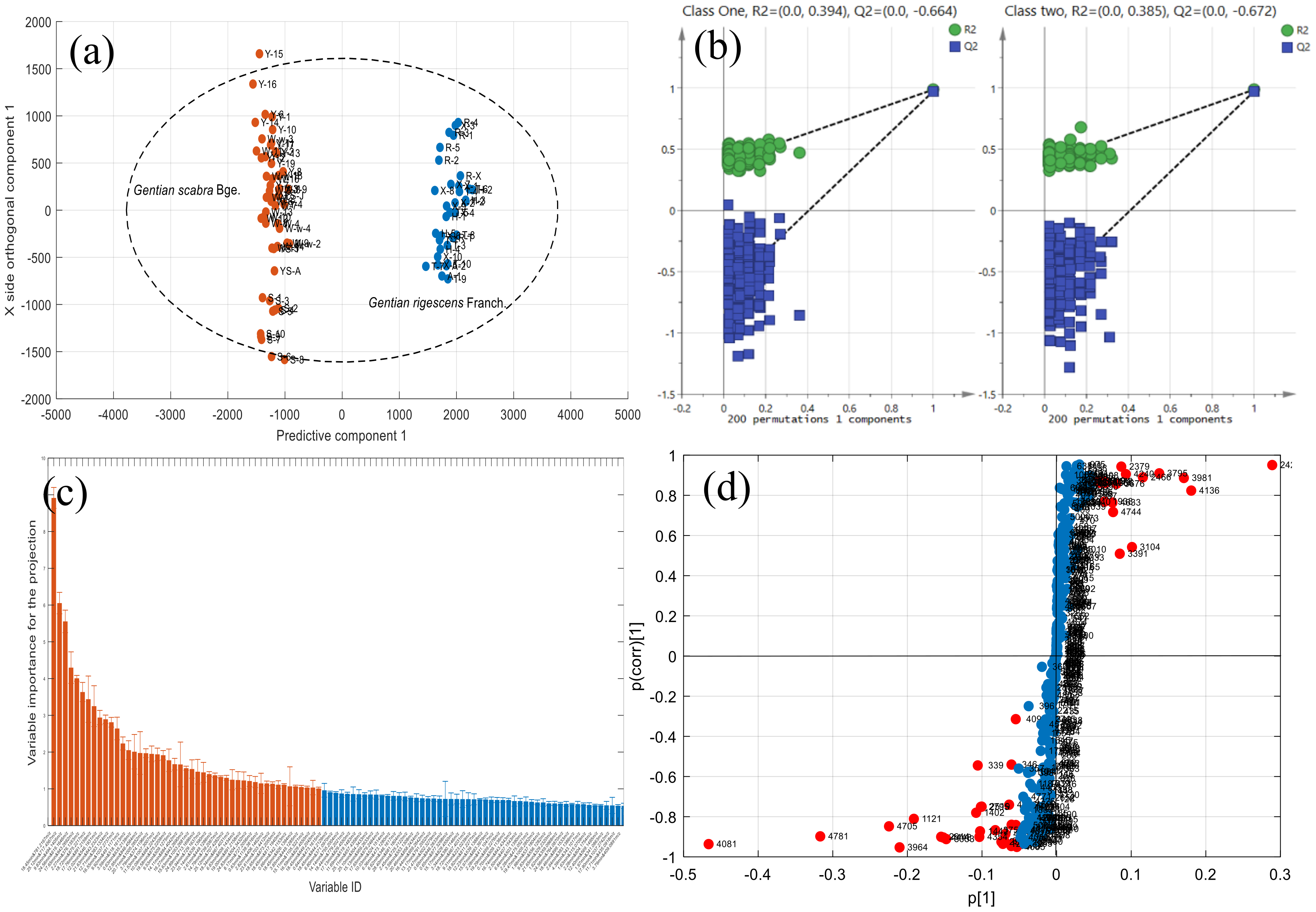
2.3.3. Screening of Differential Metabolic Characteristics
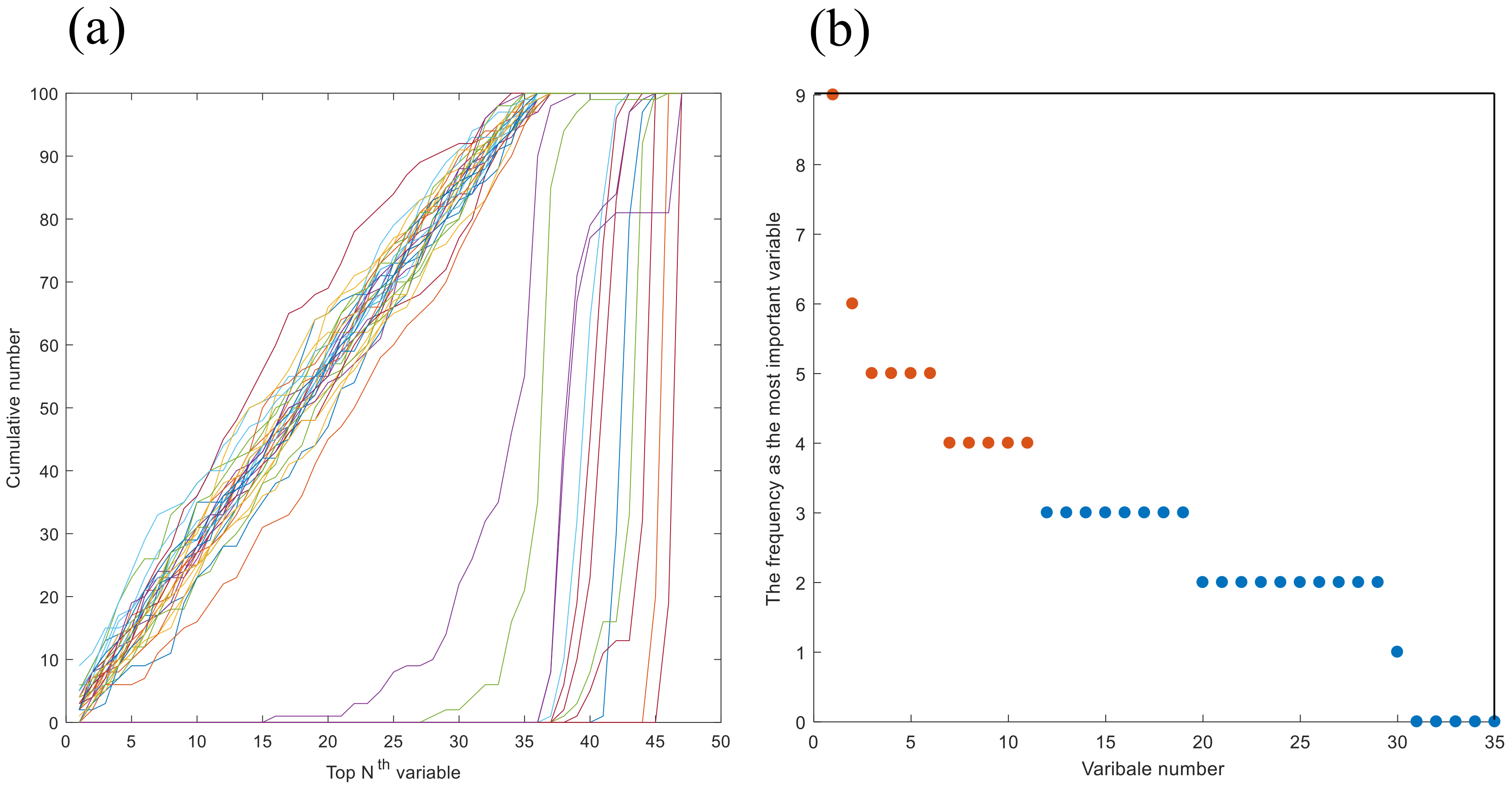
2.3.4. Identification of Chemical Markers
3. Experimental
3.1. Materials and Reagents
3.2. Sample and Standards Preparation
3.3. Instrumentation and Condition
3.4. Data Processing and Multivariate Analysis
3.5. Software
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviation
References
- Wang, Y.M.; Xu, M.; Wang, D.; Zhu, H.T.; Yang, C.R.; Zhang, Y.J. Review on “Long-Dan”, one of the traditional Chinese medicinal herbs recorded in Chinese pharmacopoeia. Nat. Prod. Bioprod. 2012, 2, 1–10. [Google Scholar] [CrossRef]
- State Pharmacopoeia Commission of the PRC. Pharamacopoeia of the People’s Republic of China; People’s Medical Publishing House: Beijing, China, 2015.
- Lian, L.H.; Wu, Y.L.; Bai, T.; Jiang, S.; Yang, Y.; Nan, J.X. Hepatoprotective effects of Gentiana manshurica Kitagawa. Acta Pharmacol. Sin. 2013, 34, 129. [Google Scholar]
- Yang, J.L.; Liu, L.L.; Shi, Y.P. Phytochemicals and biological activities of Gentiana species. Nat. Prod. Commun. 2010, 5, 1934578X1000500432. [Google Scholar] [CrossRef]
- Suh, H.W.; Lee, K.B.; Kim, K.S.; Yang, H.J.; Choi, E.K.; Shin, M.H.; Park, Y.S.; Na, Y.C.; Ahn, K.S.; Jang, Y.P.; et al. A bitter herbal medicine Gentiana scabra root extract stimulates glucagon-like peptide-1 secretion and regulates blood glucose in db/db mouse. J. Ethnopharmacol. 2015, 172, 219–226. [Google Scholar] [CrossRef] [PubMed]
- Yang, B.; Kim, S.; Kim, J.H.; Lim, C.; Kim, H.; Cho, S. Gentiana scabra Bge roots alleviates skin lesions of contact dermatitis in mice. J. Ethnopharmacol. 2019, 233, 141–147. [Google Scholar] [CrossRef]
- Wang, K.; Coyle, M.E.; Mansu, S.; Zhang, A.L.; Xue, C.C. Gentiana scabra Bge formula for herpes zoster: Biological actions of key herbs and systematic review of efficacy and safety. Phytother. Res. 2017, 31, 375–386. [Google Scholar] [CrossRef]
- Wang, Z.Y.; Wang, C.Y.; Su, T.T.; Zhang, J. Antioxidant and immunological activities of polysaccharides from Gentiana scabra Bge roots. Carbohydr. Polym. 2014, 112, 114–118. [Google Scholar] [CrossRef]
- Singh, A. Phytochemicals of Gentianaceae a review of pharmacological properties. Inter. J. Pharm. Sci. Nanotech. 2008, 1, 33–36. [Google Scholar]
- Pan, Y.; Zhao, Y.L.; Zhang, J.; Li, W.Y.; Wang, Y.Z. Phytochemistry and pharmacological activities of the genus Gentiana (Gentianaceae): Review. Phytother. Res. PTR 2016, 13, 107–150. [Google Scholar]
- Mirzaee, F.; Hosseini, A.; Jouybari, H.B.; Davoodi, A.; Azadbakht, M. Medicinal, biological and phytochemical properties of Gentiana species. J. Trad. Comple. Med. 2017, 7, 400–408. [Google Scholar] [CrossRef] [PubMed]
- Wang, A.Y.; Lian, L.H.; Jiang, Y.Z.; Wu, Y.L.; Nan, J.X. Gentiana manshurica Kitagawa prevents acetaminophen-induced acute hepatic injury in mice via inhibiting JNK/ERK MAPK pathway. World J. Gastroenter. 2010, 16, 384–391. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Zhou, W.; Kim, S.; Koo, J.E.; Kim, Y.; Koh, Y.S.; Shim, S.H.; Ma, J.Y.; Kim, Y.H. Three new secoiridoid glycosides from the rhizomes and roots of Gentiana scabra and their anti-inflammatory activities. Nat. Prod. Res. 2015, 29, 1920–1927. [Google Scholar] [CrossRef] [PubMed]
- He, Y.M.; Zhu, S.; Ge, Y.W.; Kazuma, K.; Zou, K.; Cai, S.Q.; Komatsu, K. The anti-inflammatory secoiridoid glycosides from Gentianae Scabrae Radix: The root and rhizome of Gentiana scabra. J. Nat. Med.-Tokyo 2015, 69, 303–312. [Google Scholar] [CrossRef] [PubMed]
- Choi, R.-Y.; Nam, S.-J.; Lee, H.-I.; Lee, J.; Leutou, A.S.; Ri Ham, J.; Lee, M.-K. Gentiopicroside isolated from Gentiana scabra Bge inhibits adipogenesis in 3T3-L1 cells and reduces body weight in diet-induced obese mice. Bioorg. Med. Chem. Let. 2019, 29, 1699–1704. [Google Scholar] [CrossRef]
- Cheng, Z.; Zhang, Y.; Song, H.; Zhou, H.; Zhong, F.; Hu, H.; Feng, Y. Extraction optimization, characterization and antioxidant activity of polysaccharide from Gentiana scabra Bge. Inter. J. Biol. Macromol. 2016, 93, 369–380. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Wang, Y.; Zhang, J.; Wang, Z. Optimization for the extraction of polysaccharides from Gentiana scabra Bge and their antioxidant in vitro and anti-tumor activity in vivo. J. Taiwan Inst. Chem. Eng. 2014, 45, 1126–1132. [Google Scholar] [CrossRef]
- Xu, Y.; Li, Y.; Maffucci, K.G.; Huang, L.; Zeng, R. Analytical methods of phytochemicals from the genus Gentiana. Molecules 2017, 22, 2080. [Google Scholar] [CrossRef]
- Zhang, W.; Jiang, H.; Yang, J.; Song, G.; Wen, D.; Liu, W.; Jin, M.; Wang, Q.; Du, Y.; Sun, Q.; et al. A high-throughput metabolomics approach for the comprehensive differentiation of four Pulsatilla Adans herbs combined with a nontargeted bidirectional screen for rapid identification of triterpenoid saponins. Anal. Bioanal. Chem. 2019, 411, 2071–2088. [Google Scholar] [CrossRef]
- Theodoridis, G.A.; Gika, H.G.; Want, E.J.; Wilson, I.D. Liquid chromatography-mass spectrometry based global metabolite profiling: A review. Anal. Chim. Acta 2012, 711, 7–16. [Google Scholar] [CrossRef]
- Ma, S.; Chowdhury, S.K. Data acquisition and data mining techniques for metabolite identification using LC coupled to high-resolution MS. Bioanalysis 2013, 5, 1285–1297. [Google Scholar] [CrossRef]
- Lv, X.; Sun, J.Z.; Xu, S.Z.; Cai, Q.; Liu, Y.Q. Rapid characterization and identification of chemical constituents in Gentiana radix before and after wine-processed by UHPLC-LTQ-Orbitrap MS(n). Molecules 2018, 23, 3222. [Google Scholar] [CrossRef] [PubMed]
- Saez, V.; Riquelme, S.; Baer, D.V.; Vallverdu-Queralt, A. Phenolic profile of Grape Canes: Novel compounds identified by LC-ESI-LTQ-Orbitrap-MS. Molecules 2019, 24, 3163. [Google Scholar] [CrossRef] [PubMed]
- Pan, H.; Yang, W.; Zhang, Y.; Yang, M.; Feng, R.; Wu, W.; Guo, D. An integrated strategy for the systematic characterization and discovery of new indole alkaloids from Uncaria rhynchophylla by UHPLC/DAD/LTQ-Orbitrap-MS. Anal. Bioanal. Chem. 2015, 407, 6057–6070. [Google Scholar] [CrossRef] [PubMed]
- Blazenovic, I.; Kind, T.; Ji, J.; Fiehn, O. Software tools and approaches for compound identification of LC-MS/MS data in metabolomics. Metabolites 2018, 8, 31. [Google Scholar] [CrossRef]
- Dinda, B. Pharmacology and Applications of Naturally Occurring Iridoids; Springer: Basel, Switzerland, 2019. [Google Scholar]
- Suying, L.; Fengrui, S.; Zhiqiang, L. Mass Spectrometry Analysis of Traditional Chinese Medicine; Science Press: Beijing, China, 2012. [Google Scholar]
- Qin, J.P.; Deng, J.G.; Feng, Y.Q.; Feng, X. Applications of organic mass spectrometry in structure identification of an impurity compound in prepared mangiferin extracted from Mangifera indica L. leaves. J. Chin. Mass Spectrom. Soc. 2008, 29, 218–225. [Google Scholar]
- Tugizimana, F.; Steenkamp, P.A.; Piater, L.A.; Dubery, I.A. A Conversation on data mining strategies in LC-MS untargeted metabolomics: Pre-processing and pre-treatment steps. Metabolites 2016, 6, 40. [Google Scholar] [CrossRef]
- Gromski, P.S.; Xu, Y.; Hollywood, K.A.; Turner, M.L.; Goodacre, R. The influence of scaling metabolomics data on model classification accuracy. Metabolomics 2014, 11, 684–695. [Google Scholar] [CrossRef]
- Pezzatti, J.; Boccard, J.; Codesido, S.; Gagnebin, Y.; Joshi, A.; Picard, D.; González-Ruiz, V.; Rudaz, S. Implementation of liquid chromatography–high resolution mass spectrometry methods for untargeted metabolomic analyses of biological samples: A tutorial. Anal. Chim. Acta 2020. [Google Scholar] [CrossRef]
Sample Availability: Not available. |

| No | Name | Formula | m/z | Adduction | Fragment Ion | RT | Scabra | Rigescens |
|---|---|---|---|---|---|---|---|---|
| Iridoid | ||||||||
| 1 | Secologanic acid | C16H22O10 | 373.11 | [M-H] | 179/108/204/282 | 8.40 | + | |
| 2 | Longanic acid * | C16H24O10 | 375.13 | [M-H] | 371/179 | 2.35 | + | |
| 3 | Gentiolactone | C10H12O5 | 211.10 | [M-H] | 167 | 7.50 | + | + |
| 4 | Gentiopicroside* | C16H20O9 | 401.11 | [M-H+HCOOH] | 179/355/149/119 | 5.22 | + | + |
| 5 | Sweroside * | C16H22O9 | 403.12 | [M-H+HCOOH] | 357/195/179/125 | 5.63 | + | + |
| 6 | 8-epikingside | C17H24O11 | 403.12 | [M-H] | 371/223/179 | 7.49 | + | |
| 7 | Caryptoside | C17H26O11 | 405.14 | [M-H+HCOOH] | 179 | 2.26 | + | + |
| 8 | Swertiamarin * | C16H22O10 | 419.12 | [M-H+HCOOH] | 179/355/211/119 | 3.85 | + | + |
| 9 | Loganin * | C17H26O10 | 435.22 | [M-H+HCOOH] | 389.22 | 7.17 | + | |
| 10 | Secoxyloganin | C17H24O11 | 449.13 | [M-H+HCOOH] | 179/241/359/403 | 2.66 | + | |
| 11 | Morroniside | C17H26O11 | 451.14 | [M-H+HCOOH] | 405/243/179/ | 2.94 | + | |
| 12 | 2′-O-(2,3-dihyrben)-gentiopicroside | C20H28O14 | 491.14 | [M-H] | 167/323/459 | 4.59 | + | + |
| 13 | 2′-O-(2,3-dihyben)-swertamairn | C23H26O13 | 509.13 | [M-H] | 153/297/315/367 | 10.53 | + | |
| 14 | 3′-O-(2,3-dihyben)-swertamairn | C23H26O13 | 509.22 | [M-H] | 153/517/411/321 | 6.11 | + | |
| 15 | Deglu-noneacetylate-rindoiside | C23H26O13 | 509.22 | [M-H] | 153/297/315/367 | 9.53 | + | |
| 16 | Rigenolide A | C25H28O12 | 519.15 | [M-H] | 307 | 11.20 | + | |
| 17 | 4-glu+D97RT | C26H34O11 | 521.20 | [M-H] | 359/329 | 9.44 | + | |
| 18 | Lacriciresinol | C26H34O11 | 521.20 | [M-H] | 473/355/375/415 | 15.26 | + | |
| 19 | 6′-O-d-glu swertiamarin | C25H28O13 | 535.14 | [M-H] | 409/491/153/339 | 14.39 | + | |
| 20 | 6′-O-d-glu-Loganic acid | C22H34O15 | 537.18 | [M-H] | 213 | 2.05 | + | + |
| 21 | Dideacetylate-deglu-rindoside | C25H28O14 | 551.14 | [M-H] | 491/409/509 | 12.34 | + | |
| 22 | Gentianaside | C22H36O13 | 553.13 | [M-H+HCOOH] | 507 | 5.03 | + | |
| 23 | 6-O-d-glu-gentiopicroside | C22H30O14 | 563.16 | [M-H+HCOOH] | 341/517 | 5.69 | + | + |
| 24 | Tortoside B | C28H38O13 | 581.15 | [M-H] | 401/357/313/269 | 1.47 | + | |
| 25 | Gentiabavaroside | C26H30O15 | 581.16 | [M-H] | 401/357/313/221 | 1.47 | + | |
| 26 | 6-O-d-glu-swertianmarin | C22H32O15 | 581.17 | [M-H+HCOOH] | 341/535/517/179 | 2.83 | + | |
| 27 | Deacetylate-deglu-rindoside | C27H30O15 | 593.15 | [M-H] | 451/531/551 | 19.54 | + | + |
| 28 | Deglu-trifloroside | C29H32O15 | 619.17 | [M-H] | 577 | 24.5 | + | + |
| 29 | Deglu-gelidoside | C29H32O16 | 635.16 | [M-H] | 551/451/593 | 23.19 | + | + |
| 30 | Gentiotrifloroside | C29H36O17 | 655.19 | [M-H] | 315/493/529 | 7.41 | + | + |
| 31 | Deglu-scabraside | C34H34O15 | 681.18 | [M-H] | 639/475/153 | 28.45 | + | + |
| 32 | Gentrigeoside A | C36H60O12 | 683.40 | [M-H] | 640/622 | 28.46 | + | |
| 33 | 2,3-deacetyl-trifloroside | C32H40O17 | 695.18 | [M-H] | 315 | 13.78 | + | |
| 34 | 6′-O-ace-3-O-glu-2-hy-sweroside | C31H38O18 | 697.17 | [M-H] | 315/535/571/315 | 14.25 | + | |
| 35 | Trideacetylate-trifloroside | C31H38O18 | 697.18 | [M-H+HCOOH] | 505/651/313/269 | 8.78 | + | |
| 36 | Scabran G3 | C28H40O19 | 725.21 | [M-H] | 341/383/503/679 | 2.55 | + | + |
| 37 | 2-deaceyl-trifloroside | C33H38O19 | 737.19 | [M-H] | 315/575/693 | 16.1 | + | |
| 38 | Dedihydroxybenzoate-Macrophylloside | C33H40O19 | 739.20 | [M-H] | 697/577/613/535 | 15.76 | + | + |
| 39 | Deacetylate-Trifloroside | C33H40O19 | 739.20 | [M-H] | 697/577/613/535 | 14.7 | + | + |
| 40 | Deacetylate-Rindoside | C33H40O20 | 755.20 | [M-H] | 593/315/713 | 13.92 | + | + |
| 41 | Trifloroside * | C35H42O20 | 781.22 | [M-H] | 619/739/577/315 | 18.44 | + | |
| 42 | Dideacetylate-Macophylloside | C36H40O20 | 791.24 | [M-H] | 521/629/315 | 15.11 | + | |
| 43 | Rindoside * | C35H42O21 | 797.21 | [M-H] | 315/493/635/755 | 17.19 | + | + |
| 44 | Deacetylate-scabraside | C38H42O19 | 801.22 | [M-H] | 639/597 | 18.41 | + | |
| 45 | Acetylate-trifloroside | C37H44O21 | 823.23 | [M-H] | 619/577/781 | 21.59 | + | |
| 46 | Deacetylatemacrophylloside A | C38H42O21 | 833.21 | [M-H] | 671/697/535/315 | 15.33 | + | + |
| 47 | Scabraside | C40H44O20 | 843.24 | [M-H] | 681/639/315/801 | 21.66 | + | + |
| 48 | Dideacetylate-4′-glu-trifloroside | C37H48O23 | 859.23 | [M-H] | 697 | 19.31 | + | + |
| 49 | Macrophylloside A | C40H44O22 | 875.22 | [M-H] | 739/577/535 | 18,74 | + | + |
| 50 | Deacetylate-4′-glu-trifloroside | C39H50O24 | 901.26 | [M-H] | 577/459/535/859 | 13.36 | + | |
| 51 | 4-O-β-d-trifloroside | C41H52O25 | 943.27 | [M-H] | 459/619/901/577 | 16.29 | + | + |
| 52 | Acetylate-4′-glu-scabraside | C46H54O25 | 1005.29 | [M-H] | 963/681/639/477 | 20.73 | + | |
| 53 | 4″-O-β-d-glucopyranosy1-6′-O-(4-O-β-d-glu-caffeoyl) linearroside | C46H56O25 | 1007.30 | [M-H] | 845/801/487/639 | 11.59 | + | |
| 54 | Benzoxy-4″-O-β-d-glucopyranosy1-6′-O-(4-O-β-d-glucopyranosylcaffeoyl)linearroside | C53H60O26 | 1111.33 | [M-H] | 845/487/639/801 | 15.03 | + | |
| Flavonoids | ||||||||
| 55 | Isovitexin | C21H20O10 | 431.10 | [M-H] | 9.7 | + | + | |
| 56 | Isoorientin(Homoorientin) * | C21H20O11 | 447.15 | [M-H] | 327/357/429 | 8.29 | + | + |
| 57 | Isoscoparin | C22H22O11 | 507.17 | [M-H+COOH] | 461 | 6.66 | + | |
| 58 | 2-glu-isovitexin | C27H30O15 | 593.15 | [M-H] | 551/451/531/409 | 19.54 | + | |
| 59 | Isosaponarin | C27H30O16 | 593.15 | [M-H] | 311/431/473/503 | 7.68 | + | |
| 60 | Saponarin | C27H30O17 | 593.15 | [M-H] | 367 | 11.51 | + | |
| 61 | 4′-glu-isoorientin | C27H30O16 | 609.14 | [M-H] | 447 | 5.27 | + | |
| 62 | Lutonarin | C27H30O16 | 609.14 | [M-H] | 447/519/489/327 | 6.87 | + | |
| 63 | Rutin | C27H30O16 | 609.14 | [M-H] | 447/519/489/327 | 6.9 | + | |
| 64 | Keampferol | C15H10O6 | 331.04 | [M-H+HCOOH] | 285/165 | 22.50 | + | |
| 65 | 7-glu-isopyrenine | C29H34O17 | 653.17 | [M-H] | 315 | 10.06 | + | + |
| 66 | Hyperoside | C21H20O12 | 509.22 | [M-H+HCOOH] | 463 | 8.95 | + | |
| 67 | Lonicerin | C27H30O15 | 593.19 | [M-H] | 551/451 | 17.06 | + | |
| Miscellaneous Compounds | ||||||||
| 68 | Caffeic acid | C9H8O4 | 179.03 | [M-H] | 135/109 | 9.25 | + | + |
| 69 | Ferulic acid | C10H10O4 | 193.02 | [M-H] | 149 | 3.77 | + | |
| 70 | Isoferulic acid | C10H10O4 | 193.02 | [M-H] | 149 | 4.01 | + | |
| 71 | Vanilloloside | C13H16O9 | 315.07 | [M-H] | 153 | 1.6 | + | + |
| 72 | Glu-2,3-dihydroxybenzoic acid | C14H20O8 | 315.11 | [M-H] | 187/297/253/145 | 22.81 | + | |
| 73 | Glu-2-hydro-3-methoben | C14H18O9 | 329.09 | [M-H] | 167 | 6.89 | + | + |
| 74 | Methyl-3-(β-d-glucopyranosyl)-2-hydroxybenzoate | C14H18O9 | 329.23 | [M-H] | 209/311 | 19.25 | + | |
| 75 | Glu-caffeic acid | C15H18O9 | 341.09 | [M-H] | 179/135/203/239 | 1.97 | + | |
| 76 | Syringin | C17H24O9 | 371.10 | [M-H] | 249 | 4.7 | + | |
| 77 | 3-[(6-O-Arabinopyranosyl)-β-d-glucopyranosyloxy]oct-1-en | C19H34O10 | 421.20 | [M-H] | 289/133 | 12.57 | + | |
| 78 | Methyl-3-[(6-O-β-d-glucopyranosyl)-β-d- glucopyranosyloxy]-2-hydroxybenzoate | C23H24O12 | 491.12 | [M-H] | 153/315/475 | 12.31 | + | |
| 79 | (+) Syringaresinol | C24H30O8 | 491.14 | [M-H+HCOOH] | 315/447/153 | 9.94 | + | |
| 80 | Lonicerin | C27H30O15 | 593.19 | [M-H] | 551/451/ | 16.68 | + | |
| Xanthones | ||||||||
| 81 | Mangiferin * | C19H18O11 | 421.08 | [M-H] | 403/331/301 | 5.98 | + | |
| 82 | Gentianabavaroside | C26H30O15 | 581.16 | [M-H] | 401/357/313/ | 3.71 | + | |
| Triterpenoids | ||||||||
| 83 | Gentrigeoside D | C36H60O13 | 745.40 | [M-H+HCOOH] | 699 | 11.99 | + | |
| 84 | Gentrigeoside C | C42H70O17 | 845.24 | [M-H] | 683/803 | 21.77 | + | |
| 85 | Glu-Gentigeoside C | C46H56O25 | 1007.31 | [M-H] | 845/801/639/487 | 11.52 | + | |
| 86 | Gentrigeoside A (Dammarane) | C36H60O12 | 683.40 | [M-H] | 640/622 | 28.46 | + | |
| 87 | Rha-Gentrigeoside | C46H54O27 | 1037.51 | [M-H+HCOOH] | 991 | 13.6 | + | |
| No. | Name | Formula | RT | m/z | Adduction | Fragment Ion |
|---|---|---|---|---|---|---|
| 1 | (+)Syringaresinol | C24H30O8 | 9.94 | 491.14 | [M-H+HCOOH] | 315/447/153 |
| 2 | Lutonarin | C27H30O16 | 6.87 | 609.14 | [M-H] | 447/519/489/327 |
| 3 | Trifloroside | C35H42O20 | 18.44 | 781.22 | [M-H] | 619/739/577/315 |
| 4 | 4-β-d-glu-trifloroside | C41H52O25 | 16.29 | 943.27 | [M-H] | 459/619/901/577 |
| 5 | 4″-O-β-d-glucopyranosy1-6′-O-(4-O-β-d-glucopyranosylcaffeoyl)linearroside | C46H56O25 | 11.59 | 1007.30 | [M-H] | 845/323/487/639/801 |
| 6 | Macrophylloside A | C40H44O22 | 18.74 | 875.22 | [M-H] | 739/577/535 |
| 7 | Scabraside | C40H44O20 | 21.66 | 843.24 | [M-H] | 681/639/315/801 |
| 8 | Unknown | 25.12 | 717.46 | |||
| 9 | Unknown | 15.11 | 1105.32 | |||
| 10 | Unknown | 19.00 | 891.22 | |||
| 11 | Unknown | 15.63 | 1053.27 |
| Species | location | Sample Number | Species | Species | Sample Number |
|---|---|---|---|---|---|
| Gentiana scabra | QY-wdz | 1–20 | Gentiana scabra | Ys-j | 52 |
| Gentiana scabra | QY--yem | 21–39 | Gentiana rigescens | Yx-hsc | 53–57 |
| Gentiana scabra | QY- | 40–49 | Gentiana rigescens | Yx-jfc | 58–62 |
| Gentiana scabra | Ys-y | 50 | Gentiana rigescens | Yx-xh | 63–72 |
| Gentiana scabra | Ys-a | 51 | Gentiana rigescens | Yx-th | 73–86 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, G.; Li, Y.; Wei, W.; Li, J.; Li, H.; Huang, Y.; Guo, D.-a. Metabolomics Combined with Multivariate Statistical Analysis for Screening of Chemical Markers between Gentiana scabra and Gentiana rigescens. Molecules 2020, 25, 1228. https://doi.org/10.3390/molecules25051228
Zhang G, Li Y, Wei W, Li J, Li H, Huang Y, Guo D-a. Metabolomics Combined with Multivariate Statistical Analysis for Screening of Chemical Markers between Gentiana scabra and Gentiana rigescens. Molecules. 2020; 25(5):1228. https://doi.org/10.3390/molecules25051228
Chicago/Turabian StyleZhang, Gaole, Yun Li, Wenlong Wei, Jiayuan Li, Haoju Li, Yong Huang, and De-an Guo. 2020. "Metabolomics Combined with Multivariate Statistical Analysis for Screening of Chemical Markers between Gentiana scabra and Gentiana rigescens" Molecules 25, no. 5: 1228. https://doi.org/10.3390/molecules25051228
APA StyleZhang, G., Li, Y., Wei, W., Li, J., Li, H., Huang, Y., & Guo, D.-a. (2020). Metabolomics Combined with Multivariate Statistical Analysis for Screening of Chemical Markers between Gentiana scabra and Gentiana rigescens. Molecules, 25(5), 1228. https://doi.org/10.3390/molecules25051228






