Good Vibrations: Structural Remodeling of Maturing Yeast Pre-40S Ribosomal Particles Followed by Cryo-Electron Microscopy
Abstract
:1. Introduction
2. Results
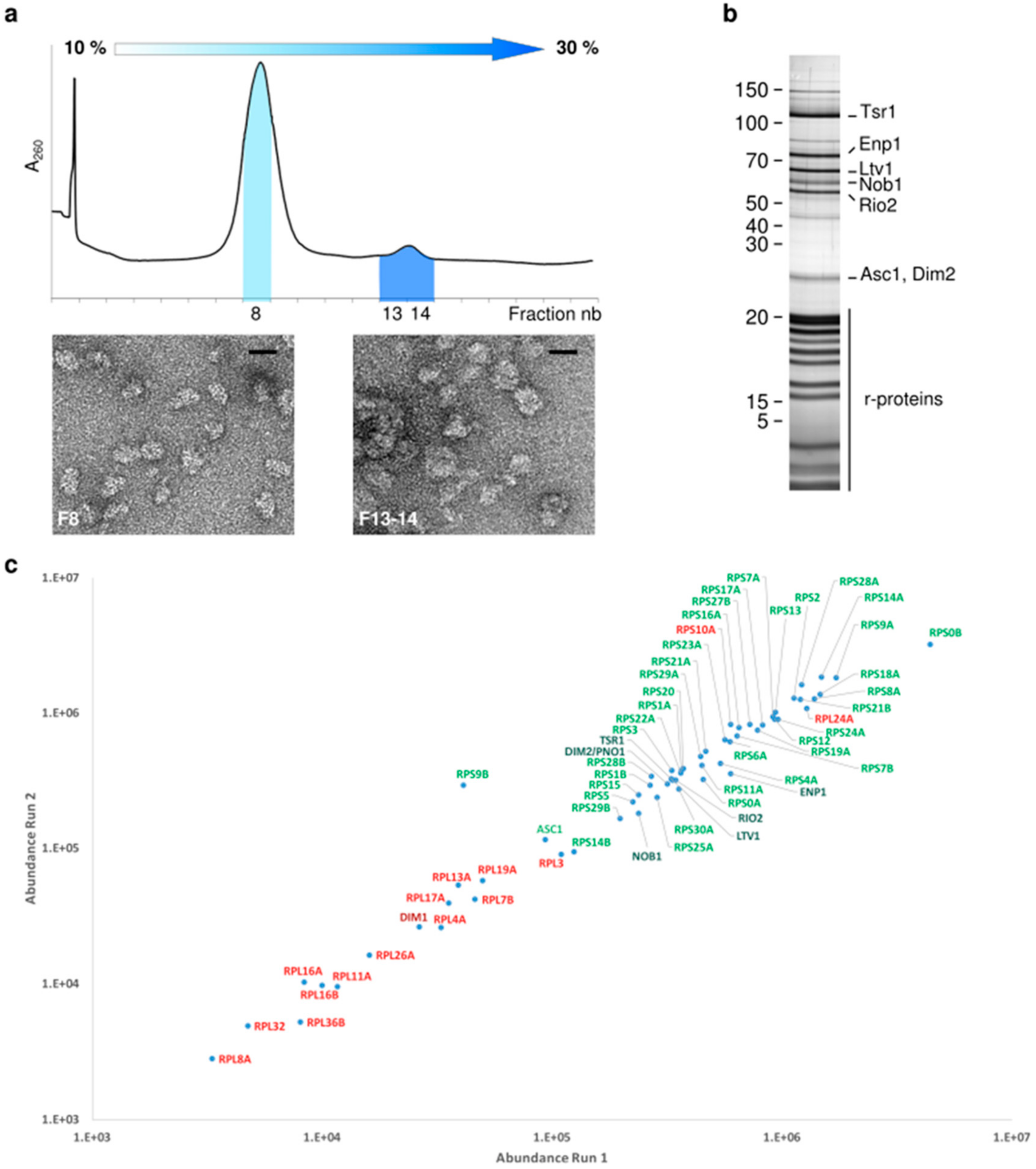
2.1. Proteomic Characterization of Tsr1-purified pre-40S Particles
2.2. Cryo-EM Analysis of Tsr1-FPZ pre-40S Particles
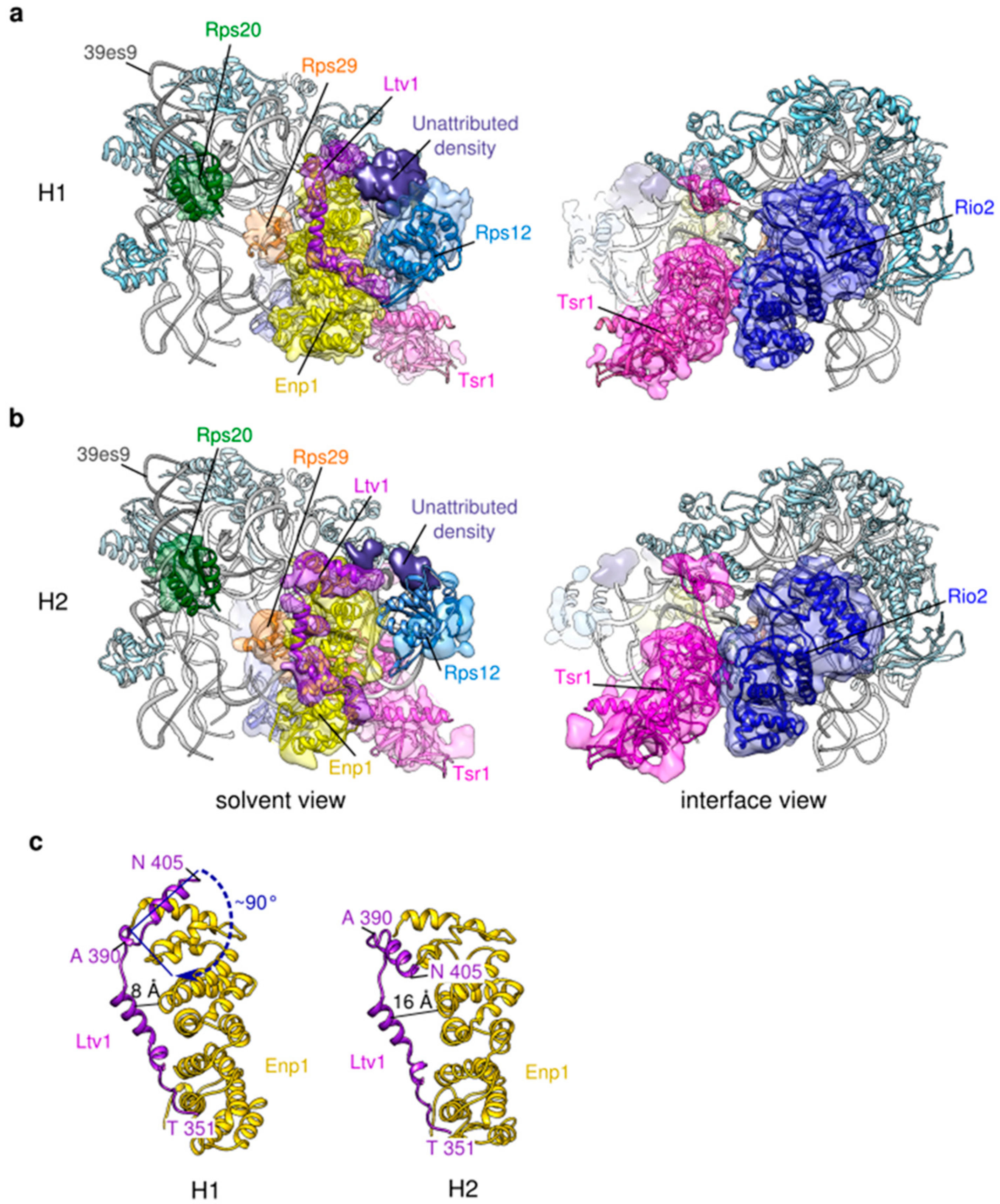
2.3. Structural Heterogeneity of the Head Domain
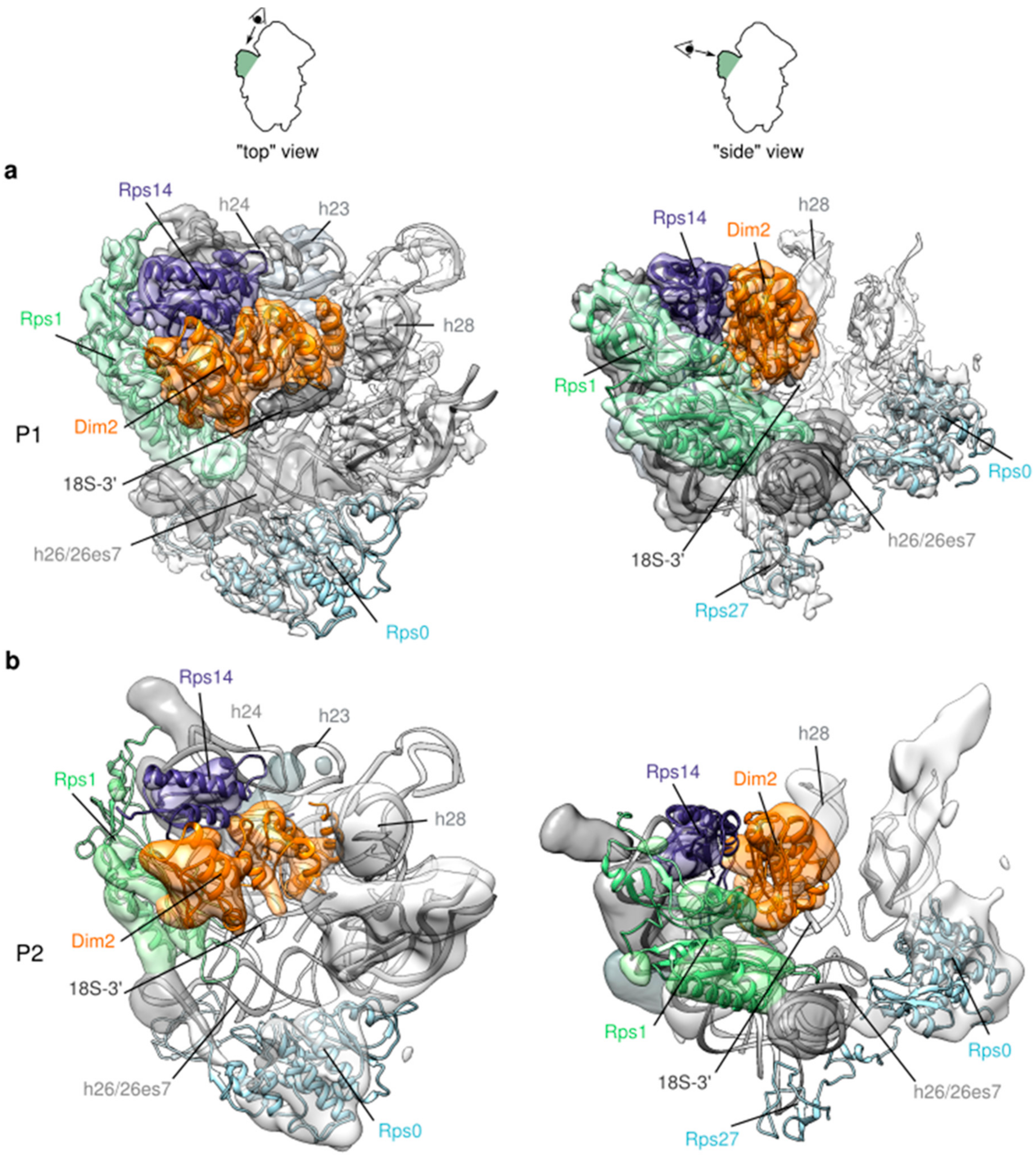
2.4. A “Wriggling” of the Platform Domain
3. Discussion
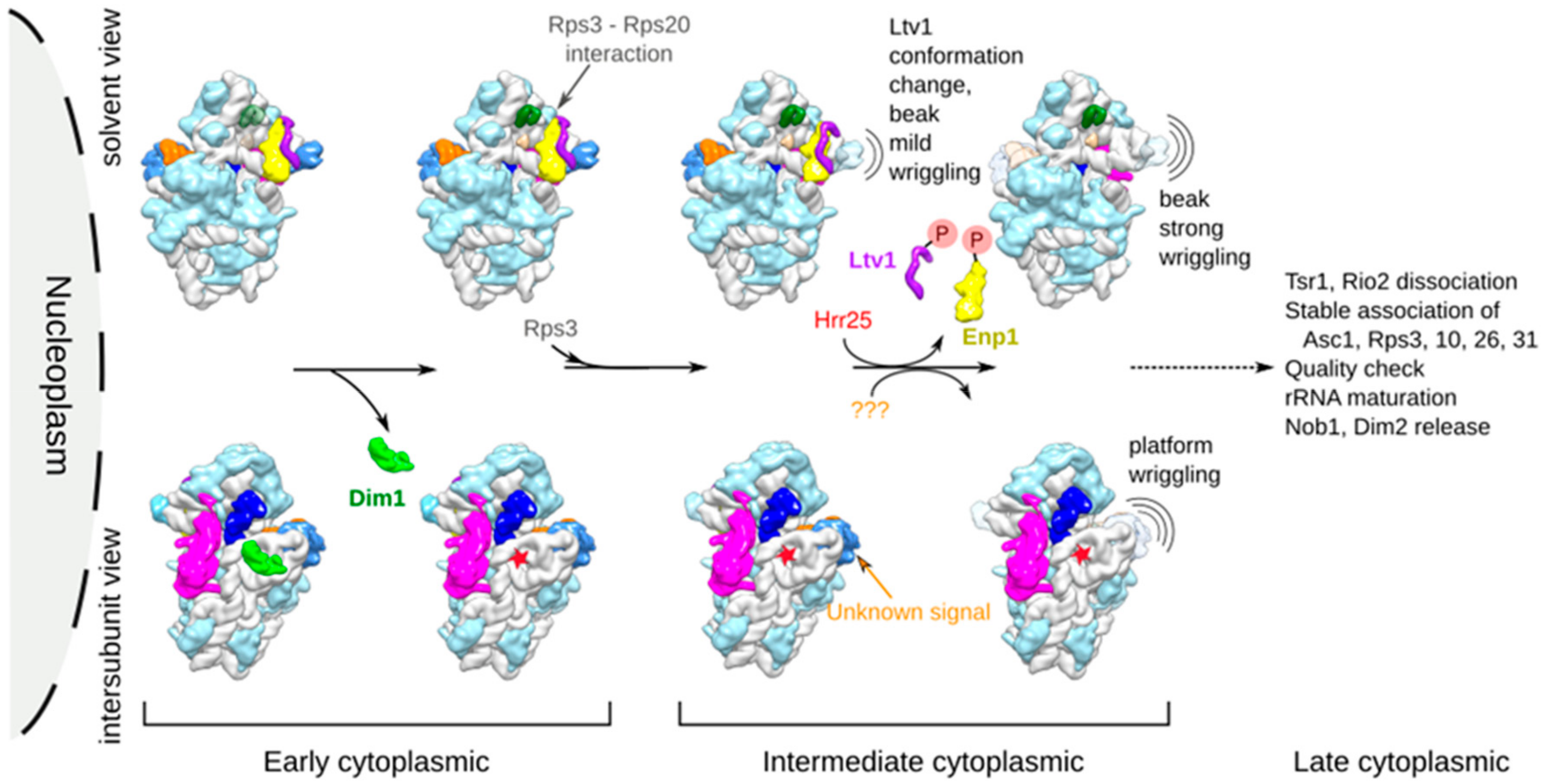
3.1. Cryo-EM Structures Suggest that pre-40S Particles Transit through a Vibrating State
3.2. Dim1 Activity and Release from pre-40S Particles Might Correspond to Early Cytoplasmic Events
3.3. Ltv1 Conformational Change and Beak Wriggling Precede Ltv1 Phosphorylation by Hrr25
4. Material and Methods
4.1. Purification of Cytoplasmic pre-40S Particles
4.2. Protein Digestion and NanoLC-MS/MS Analysis
4.3. Bioinformatic MS Data Analysis
4.4. Grid Preparation and cryo-EM Images Acquisition
4.5. Single Particle Analysis
4.6. Cryo-EM Maps Interpretation
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Data Availability
References
- Venema, J.; Tollervey, D. Ribosome Synthesis in Saccharomyces Cerevisiae. Annu. Rev. Genet. 1999, 33, 261–311. [Google Scholar] [CrossRef] [PubMed]
- Dragon, F.; Gallagher, J.E.G.; Compagnone-Post, P.A.; Mitchell, B.M.; Porwancher, K.A.; Wehner, K.A.; Wormsley, S.; Settlage, R.E.; Shabanowitz, J.; Osheim, Y.; et al. A Large Nucleolar U3 Ribonucleoprotein Required for 18S Ribosomal RNA Biogenesis. Nature 2002, 417, 967–970. [Google Scholar] [CrossRef]
- Schäfer, T.; Strauss, D.; Petfalski, E.; Tollervey, D.; Hurt, E. The Path from Nucleolar 90S to Cytoplasmic 40S Pre-Ribosomes. EMBO J. 2003, 22, 1370–1380. [Google Scholar] [CrossRef] [PubMed]
- Zemp, I.; Kutay, U. Nuclear Export and Cytoplasmic Maturation of Ribosomal Subunits. FEBS Lett. 2007, 581, 2783–2793. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thomson, E.; Ferreira-Cerca, S.; Hurt, E. Eukaryotic Ribosome Biogenesis at a Glance. J. Cell Sci. 2013, 126, 4815–4821. [Google Scholar] [CrossRef] [Green Version]
- Henras, A.K.; Plisson-Chastang, C.; O’Donohue, M.-F.; Chakraborty, A.; Gleizes, P.-E. An Overview of Pre-Ribosomal RNA Processing in Eukaryotes. Wiley Interdiscip. Rev. RNA 2015, 6, 225–242. [Google Scholar] [CrossRef]
- Cerezo, E.; Plisson-Chastang, C.; Henras, A.K.; Lebaron, S.; Gleizes, P.-E.; O’Donohue, M.-F.; Romeo, Y.; Henry, Y. Maturation of Pre-40S Particles in Yeast and Humans. Wiley Interdiscip. Rev. RNA 2019, 10, e1516. [Google Scholar] [CrossRef] [Green Version]
- Barandun, J.; Hunziker, M.; Klinge, S. Assembly and Structure of the SSU Processome—A Nucleolar Precursor of the Small Ribosomal Subunit. Curr. Opin. Struct. Biol. 2018, 49, 85–93. [Google Scholar] [CrossRef]
- Ferreira-Cerca, S.; Pöll, G.; Kühn, H.; Neueder, A.; Jakob, S.; Tschochner, H.; Milkereit, P. Analysis of the In Vivo Assembly Pathway of Eukaryotic 40S Ribosomal Proteins. Mol. Cell 2007, 28, 446–457. [Google Scholar] [CrossRef]
- Rouquette, J.; Choesmel, V.; Gleizes, P.-E. Nuclear Export and Cytoplasmic Processing of Precursors to the 40S Ribosomal Subunits in Mammalian Cells. EMBO J. 2005, 24, 2862–2872. [Google Scholar] [CrossRef] [Green Version]
- Schäfer, T.; Maco, B.; Petfalski, E.; Tollervey, D.; Böttcher, B.; Aebi, U.; Hurt, E. Hrr25-Dependent Phosphorylation State Regulates Organization of the Pre-40S Subunit. Nature 2006, 441, 651–655. [Google Scholar] [CrossRef] [PubMed]
- Granneman, S.; Petfalski, E.; Swiatkowska, A.; Tollervey, D. Cracking Pre-40S Ribosomal Subunit Structure by Systematic Analyses of RNA–Protein Cross-Linking. EMBO J. 2010, 29, 2026–2036. [Google Scholar] [CrossRef]
- Turowski, T.W.; Lebaron, S.; Zhang, E.; Peil, L.; Dudnakova, T.; Petfalski, E.; Granneman, S.; Rappsilber, J.; Tollervey, D. Rio1 Mediates ATP-Dependent Final Maturation of 40S Ribosomal Subunits. Nucleic Acids Res. 2014, 42, 12189–12199. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Strunk, B.S.; Loucks, C.R.; Su, M.; Vashisth, H.; Cheng, S.; Schilling, J.; Brooks, C.L.; Karbstein, K.; Skiniotis, G. Ribosome Assembly Factors Prevent Premature Translation Initiation by 40S Assembly Intermediates. Science 2011, 333, 1449–1453. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Johnson, M.C.; Ghalei, H.; Doxtader, K.A.; Karbstein, K.; Stroupe, M.E. Structural Heterogeneity in Pre-40S Ribosomes. Struct. Lond. Engl. 2017, 25, 329–340. [Google Scholar] [CrossRef] [Green Version]
- Heuer, A.; Thomson, E.; Schmidt, C.; Berninghausen, O.; Becker, T.; Hurt, E.; Beckmann, R. Cryo-EM Structure of a Late Pre-40S Ribosomal Subunit FromSaccharomyces Cerevisiae. eLife 2017, 6. [Google Scholar] [CrossRef]
- Scaiola, A.; Peña, C.; Weisser, M.; Böhringer, D.; Leibundgut, M.; Klingauf-Nerurkar, P.; Gerhardy, S.; Panse, V.G.; Ban, N. Structure of a Eukaryotic Cytoplasmic Pre-40S Ribosomal Subunit. EMBO J. 2018, 37, e98499. [Google Scholar] [CrossRef]
- Lafontaine, D.L.; Preiss, T.; Tollervey, D. Yeast 18S RRNA Dimethylase Dim1p: A Quality Control Mechanism in Ribosome Synthesis? Mol. Cell. Biol. 1998, 18, 2360–2370. [Google Scholar] [CrossRef] [Green Version]
- Mitterer, V.; Shayan, R.; Ferreira-Cerca, S.; Murat, G.; Enne, T.; Rinaldi, D.; Weigl, S.; Omanic, H.; Gleizes, P.-E.; Kressler, D.; et al. Conformational Proofreading of Distant 40S Ribosomal Subunit Maturation Events by a Long-Range Communication Mechanism. Nat. Commun. 2019, 10, 2754. [Google Scholar] [CrossRef] [Green Version]
- Pertschy, B.; Schneider, C.; Gnädig, M.; Schäfer, T.; Tollervey, D.; Hurt, E. RNA Helicase Prp43 and Its Co-Factor Pfa1 Promote 20 to 18 S RRNA Processing Catalyzed by the Endonuclease Nob1. J. Biol. Chem. 2009, 284, 35079–35091. [Google Scholar] [CrossRef] [Green Version]
- Larburu, N.; Montellese, C.; O’Donohue, M.-F.; Kutay, U.; Gleizes, P.-E.; Plisson-Chastang, C. Structure of a Human Pre-40S Particle Points to a Role for RACK1 in the Final Steps of 18S RRNA Processing. Nucleic Acids Res. 2016, 44, 8465–8478. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ameismeier, M.; Cheng, J.; Berninghausen, O.; Beckmann, R. Visualizing Late States of Human 40S Ribosomal Subunit Maturation. Nature 2018, 558, 249–253. [Google Scholar] [CrossRef] [PubMed]
- Ghalei, H.; Schaub, F.X.; Doherty, J.R.; Noguchi, Y.; Roush, W.R.; Cleveland, J.L.; Stroupe, M.E.; Karbstein, K. Hrr25/CK1 -Directed Release of Ltv1 from Pre-40S Ribosomes Is Necessary for Ribosome Assembly and Cell Growth. J. Cell Biol. 2015, 208, 745–759. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Holzer, S.; Ban, N.; Klinge, S. Crystal Structure of the Yeast Ribosomal Protein RpS3 in Complex with Its Chaperone Yar1. J. Mol. Biol. 2013, 425, 4154–4160. [Google Scholar] [CrossRef]
- Mitterer, V.; Murat, G.; Réty, S.; Blaud, M.; Delbos, L.; Stanborough, T.; Bergler, H.; Leulliot, N.; Kressler, D.; Pertschy, B. Sequential Domain Assembly of Ribosomal Protein S3 Drives 40S Subunit Maturation. Nat. Commun. 2016, 7, 10336. [Google Scholar] [CrossRef] [Green Version]
- Ferreira-Cerca, S.; Sagar, V.; Schäfer, T.; Diop, M.; Wesseling, A.-M.; Lu, H.; Chai, E.; Hurt, E.; LaRonde-LeBlanc, N. ATPase-Dependent Role of the Atypical Kinase Rio2 on the Evolving Pre-40S Ribosomal Subunit. Nat. Struct. Mol. Biol. 2012, 19, 1316–1323. [Google Scholar] [CrossRef] [Green Version]
- Ferreira-Cerca, S.; Kiburu, I.; Thomson, E.; LaRonde, N.; Hurt, E. Dominant Rio1 Kinase/ATPase Catalytic Mutant Induces Trapping of Late Pre-40S Biogenesis Factors in 80S-like Ribosomes. Nucleic Acids Res. 2014. [Google Scholar] [CrossRef] [Green Version]
- Hector, R.D.; Burlacu, E.; Aitken, S.; Le Bihan, T.; Tuijtel, M.; Zaplatina, A.; Cook, A.G.; Granneman, S. Snapshots of Pre-RRNA Structural Flexibility Reveal Eukaryotic 40S Assembly Dynamics at Nucleotide Resolution. Nucleic Acids Res. 2014, 42, 12138–12154. [Google Scholar] [CrossRef] [Green Version]
- Strunk, B.S.; Novak, M.N.; Young, C.L.; Karbstein, K. A Translation-like Cycle Is a Quality Control Checkpoint for Maturing 40S Ribosome Subunits. Cell 2012, 150, 111–121. [Google Scholar] [CrossRef] [Green Version]
- Lebaron, S.; Schneider, C.; van Nues, R.W.; Swiatkowska, A.; Walsh, D.; Böttcher, B.; Granneman, S.; Watkins, N.J.; Tollervey, D. Proofreading of Pre-40S Ribosome Maturation by a Translation Initiation Factor and 60S Subunits. Nat. Struct. Mol. Biol. 2012, 19, 744–753. [Google Scholar] [CrossRef] [Green Version]
- Mulder, A.M.; Yoshioka, C.; Beck, A.H.; Bunner, A.E.; Milligan, R.A.; Potter, C.S.; Carragher, B.; Williamson, J.R. Visualizing Ribosome Biogenesis: Parallel Assembly Pathways for the 30S Subunit. Science 2010, 330, 673–677. [Google Scholar] [CrossRef] [PubMed]
- Razi, A.; Guarné, A.; Ortega, J. The Cryo-EM Structure of YjeQ Bound to the 30S Subunit Suggests a Fidelity Checkpoint Function for This Protein in Ribosome Assembly. Proc. Natl. Acad. Sci. USA 2017, 114, E3396–E3403. [Google Scholar] [CrossRef] [Green Version]
- Léger-Silvestre, I.; Milkereit, P.; Ferreira-Cerca, S.; Saveanu, C.; Rousselle, J.-C.; Choesmel, V.; Guinefoleau, C.; Gas, N.; Gleizes, P.-E. The Ribosomal Protein Rps15p Is Required for Nuclear Exit of the 40S Subunit Precursors in Yeast. EMBO J. 2004, 23, 2336–2347. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ghalei, H.; Trepreau, J.; Collins, J.C.; Bhaskaran, H.; Strunk, B.S.; Karbstein, K. The ATPase Fap7 Tests the Ability to Carry Out Translocation-like Conformational Changes and Releases Dim1 during 40S Ribosome Maturation. Mol. Cell 2017, 67, 990–1000.e3. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Belhabich-Baumas, K.; Joret, C.; Jády, B.E.; Plisson-Chastang, C.; Shayan, R.; Klopp, C.; Henras, A.K.; Henry, Y.; Mougin, A. The Rio1p ATPase Hinders Premature Entry into Translation of Late Pre-40S Pre-Ribosomal Particles. Nucleic Acids Res. 2017, 45, 10824–10836. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McCaughan, U.M.; Jayachandran, U.; Shchepachev, V.; Chen, Z.A.; Rappsilber, J.; Tollervey, D.; Cook, A.G. Pre-40S Ribosome Biogenesis Factor Tsr1 Is an Inactive Structural Mimic of Translational GTPases. Nat. Commun. 2016, 7, 11789. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zemp, I.; Wandrey, F.; Rao, S.; Ashiono, C.; Wyler, E.; Montellese, C.; Kutay, U. CK1 and CK1 Are Components of Human 40S Subunit Precursors Required for Cytoplasmic 40S Maturation. J. Cell Sci. 2014, 127, 1242–1253. [Google Scholar] [CrossRef] [Green Version]
- Sloan, K.E.; Warda, A.S.; Sharma, S.; Entian, K.-D.; Lafontaine, D.L.J.; Bohnsack, M.T. Tuning the Ribosome: The Influence of RRNA Modification on Eukaryotic Ribosome Biogenesis and Function. RNA Biol. 2017, 14, 1138–1152. [Google Scholar] [CrossRef]
- Mohan, S.; Donohue, J.P.; Noller, H.F. Molecular Mechanics of 30S Subunit Head Rotation. Proc. Natl. Acad. Sci. USA 2014, 111, 13325–13330. [Google Scholar] [CrossRef] [Green Version]
- Vanrobays, E.; Gélugne, J.-P.; Caizergues-Ferrer, M.; Lafontaine, D.L.J. Dim2p, a KH-Domain Protein Required for Small Ribosomal Subunit Synthesis. RNA N. Y. N 2004, 10, 645–656. [Google Scholar] [CrossRef] [Green Version]
- Woolls, H.A.; Lamanna, A.C.; Karbstein, K. Roles of Dim2 in Ribosome Assembly. J. Biol. Chem. 2011, 286, 2578–2586. [Google Scholar] [CrossRef] [Green Version]
- Klinge, S.; Woolford, J.L. Ribosome Assembly Coming into Focus. Nat. Rev. Mol. Cell Biol. 2019, 20, 116–131. [Google Scholar] [CrossRef] [PubMed]
- Nakane, T.; Kimanius, D.; Lindahl, E.; Scheres, S.H. Characterisation of Molecular Motions in Cryo-EM Single-Particle Data by Multi-Body Refinement in RELION. eLife 2018, 7. [Google Scholar] [CrossRef] [PubMed]
- Nierhaus, K.H. The Assembly of Prokaryotic Ribosomes. Biochimie 1991, 73, 739–755. [Google Scholar] [CrossRef]
- O’Donohue, M.-F.; Choesmel, V.; Faubladier, M.; Fichant, G.; Gleizes, P.-E. Functional Dichotomy of Ribosomal Proteins during the Synthesis of Mammalian 40S Ribosomal Subunits. J. Cell Biol. 2010, 190, 853–866. [Google Scholar] [CrossRef] [Green Version]
- Campbell, M.G.; Karbstein, K. Protein-Protein Interactions within Late Pre-40S Ribosomes. PLoS OnNE 2011, 6, e16194. [Google Scholar] [CrossRef]
- Bouyssié, D.; Hesse, A.-M.; Mouton-Barbosa, E.; Rompais, M.; Macron, C.; Carapito, C.; de Peredo, A.G.; Couté, Y.; Dupierris, V.; Burel, A.; et al. Proline: An Efficient and User-Friendly Software Suite for Large-Scale Proteomics. Bioinforma. Oxf. Engl. 2020. [Google Scholar] [CrossRef] [Green Version]
- Zheng, S.Q.; Palovcak, E.; Armache, J.-P.; Verba, K.A.; Cheng, Y.; Agard, D.A. MotionCor2 - Anisotropic Correction of Beam-Induced Motion for Improved Cryo-Electron Microscopy. Nat. Methods 2017, 14, 331. [Google Scholar] [CrossRef] [Green Version]
- Rohou, A.; Grigorieff, N. CTFFIND4: Fast and Accurate Defocus Estimation from Electron Micrographs. J. Struct. Biol. 2015, 192, 216–221. [Google Scholar] [CrossRef]
- Scheres, S.H.W. RELION: Implementation of a Bayesian Approach to Cryo-EM Structure Determination. J. Struct. Biol. 2012, 180, 519–530. [Google Scholar] [CrossRef] [Green Version]
- Ben-Shem, A.; Garreau de Loubresse, N.; Melnikov, S.; Jenner, L.; Yusupova, G.; Yusupov, M. The Structure of the Eukaryotic Ribosome at 3.0 A Resolution. Science 2011, 334, 1524–1529. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Scheres, S.H.W. Processing of Structurally Heterogeneous Cryo-EM Data in RELION. Methods Enzymol. 2016, 579, 125–157. [Google Scholar] [CrossRef] [PubMed]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera--a Visualization System for Exploratory Research and Analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brown, A.; Long, F.; Nicholls, R.A.; Toots, J.; Emsley, P.; Murshudov, G. Tools for Macromolecular Model Building and Refinement into Electron Cryo-Microscopy Reconstructions. Acta Crystallogr. D Biol. Crystallogr. 2015, 71, 136–153. [Google Scholar] [CrossRef]
- Murshudov, G.N.; Skubák, P.; Lebedev, A.A.; Pannu, N.S.; Steiner, R.A.; Nicholls, R.A.; Winn, M.D.; Long, F.; Vagin, A.A. REFMAC5 for the Refinement of Macromolecular Crystal Structures. Acta Crystallogr. D Biol. Crystallogr. 2011, 67, 355–367. [Google Scholar] [CrossRef] [Green Version]
- Adams, P.D.; Afonine, P.V.; Bunkóczi, G.; Chen, V.B.; Davis, I.W.; Echols, N.; Headd, J.J.; Hung, L.-W.; Kapral, G.J.; Grosse-Kunstleve, R.W.; et al. PHENIX: A Comprehensive Python-Based System for Macromolecular Structure Solution. Acta Crystallogr. D Biol. Crystallogr. 2010, 66, 213–221. [Google Scholar] [CrossRef] [Green Version]
- Nicholls, R.A.; Fischer, M.; McNicholas, S.; Murshudov, G.N. Conformation-Independent Structural Comparison of Macromolecules with ProSMART. Acta Crystallogr. D Biol. Crystallogr. 2014, 70, 2487–2499. [Google Scholar] [CrossRef] [Green Version]
- Chen, V.B.; Arendall, W.B.; Headd, J.J.; Keedy, D.A.; Immormino, R.M.; Kapral, G.J.; Murray, L.W.; Richardson, J.S.; Richardson, D.C. MolProbity: All-Atom Structure Validation for Macromolecular Crystallography. Acta Crystallogr. D Biol. Crystallogr. 2010, 66, 12–21. [Google Scholar] [CrossRef] [Green Version]
- Perez-Riverol, Y.; Csordas, A.; Bai, J.; Bernal-Llinares, M.; Hewapathirana, S.; Kundu, D.J.; Inuganti, A.; Griss, J.; Mayer, G.; Eisenacher, M.; et al. The PRIDE Database and Related Tools and Resources in 2019: Improving Support for Quantification Data. Nucleic Acids Res. 2019, 47, D442–D450. [Google Scholar] [CrossRef]
Sample Availability: Yeast strains harboring a tagged version of Tsr1 are available from the authors for pre-40S particles purification. |

| Protein | Peptide Sequence | Phosphorylation Protein Position | % Phosphosite run 1 | % Phosphosite run 2 | % Phosphosite Average | |
|---|---|---|---|---|---|---|
| Tsr1 | DTLDEYAPEGTEDWSDYDEDFEYDGLTTAR | T354 or S358 | 100 | 100 | 0 | Non phosphorylated |
| 100 | Phosphorylated | |||||
| Ltv1 | GAMSDVSGFSMSSSAIAR | S336 or S339 | 7.8 | 5.2 | 93.5 | Non phosphorylated |
| 6.5 | Phosphorylated | |||||
| Ltv1 | VTNTLSSLKF | S460 | 0 | 2.5 | 98.7 | Non phosphorylated |
| 1.3 | Phosphorylated | |||||
| Enp1 | EKESQVEDMQDDEPLANEQNTSR | S172 | 0 | 4.3 | 97.9 | Non phosphorylated |
| 2.1 | Phosphorylated | |||||
| Enp1 | ESQVEDMQDDEPLANEQNTSRGNISSGLK | T189 or S195 or S196 | 0 | 1.2 | 99.4 | Non phosphorylated |
| 0.6 | Phosphorylated | |||||
| Enp1 | ILDDGSNGEDATR | S404 | 0.2 | 0.2 | 99.8 | Non phosphorylated |
| 0.2 | Phosphorylated | |||||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shayan, R.; Rinaldi, D.; Larburu, N.; Plassart, L.; Balor, S.; Bouyssié, D.; Lebaron, S.; Marcoux, J.; Gleizes, P.-E.; Plisson-Chastang, C. Good Vibrations: Structural Remodeling of Maturing Yeast Pre-40S Ribosomal Particles Followed by Cryo-Electron Microscopy. Molecules 2020, 25, 1125. https://doi.org/10.3390/molecules25051125
Shayan R, Rinaldi D, Larburu N, Plassart L, Balor S, Bouyssié D, Lebaron S, Marcoux J, Gleizes P-E, Plisson-Chastang C. Good Vibrations: Structural Remodeling of Maturing Yeast Pre-40S Ribosomal Particles Followed by Cryo-Electron Microscopy. Molecules. 2020; 25(5):1125. https://doi.org/10.3390/molecules25051125
Chicago/Turabian StyleShayan, Ramtin, Dana Rinaldi, Natacha Larburu, Laura Plassart, Stéphanie Balor, David Bouyssié, Simon Lebaron, Julien Marcoux, Pierre-Emmanuel Gleizes, and Célia Plisson-Chastang. 2020. "Good Vibrations: Structural Remodeling of Maturing Yeast Pre-40S Ribosomal Particles Followed by Cryo-Electron Microscopy" Molecules 25, no. 5: 1125. https://doi.org/10.3390/molecules25051125
APA StyleShayan, R., Rinaldi, D., Larburu, N., Plassart, L., Balor, S., Bouyssié, D., Lebaron, S., Marcoux, J., Gleizes, P.-E., & Plisson-Chastang, C. (2020). Good Vibrations: Structural Remodeling of Maturing Yeast Pre-40S Ribosomal Particles Followed by Cryo-Electron Microscopy. Molecules, 25(5), 1125. https://doi.org/10.3390/molecules25051125






