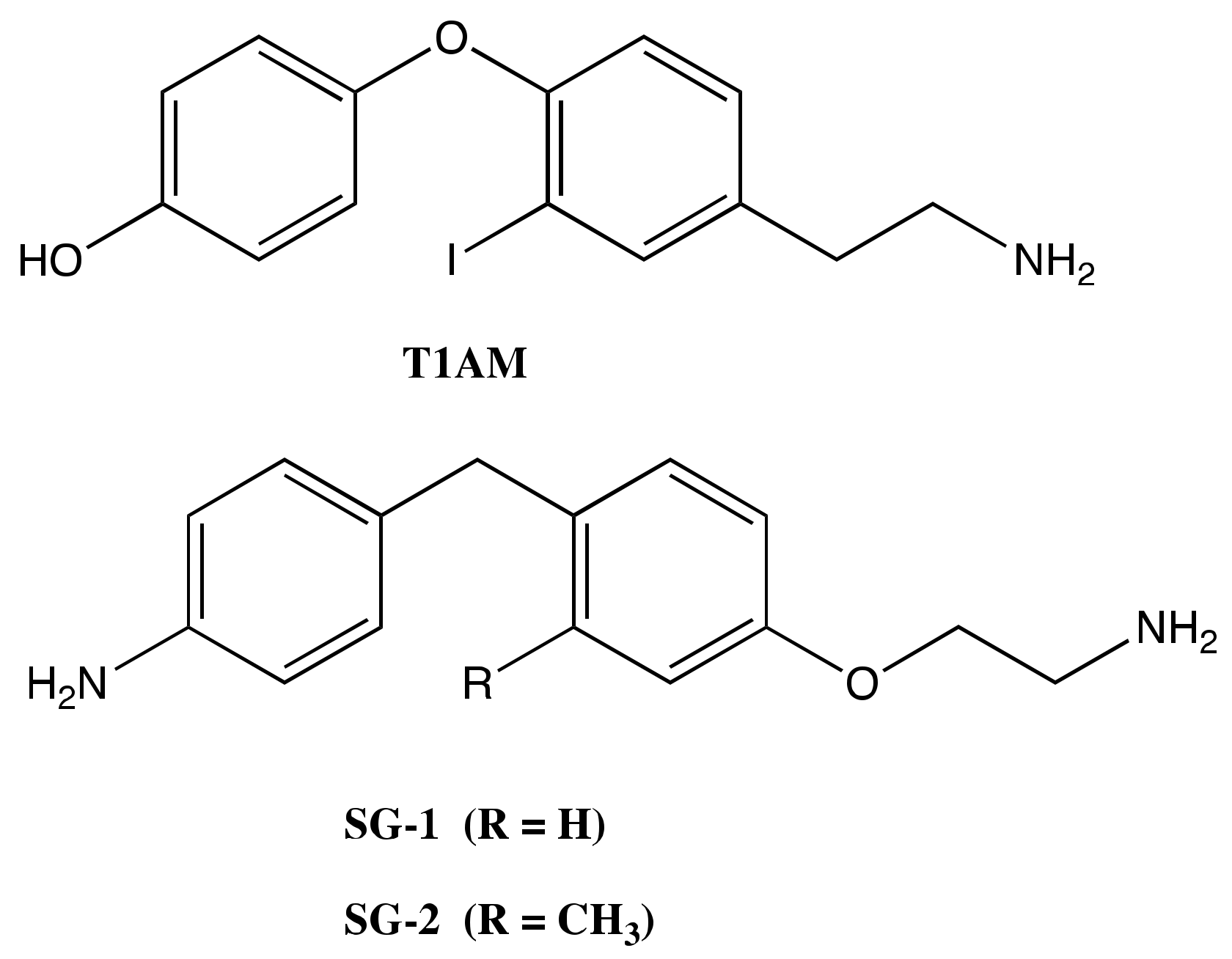
Endogenous 3-Iodothyronamine (T1AM) and Synthetic Thyronamine-Like Analog SG-2 Act as Novel Pleiotropic Neuroprotective Agents through the Modulation of SIRT6
Abstract
1. Introduction
2. Results
2.1. Effects of SG-1, SG-2 and T1AM on the Expression of Autophagy-Related Genes
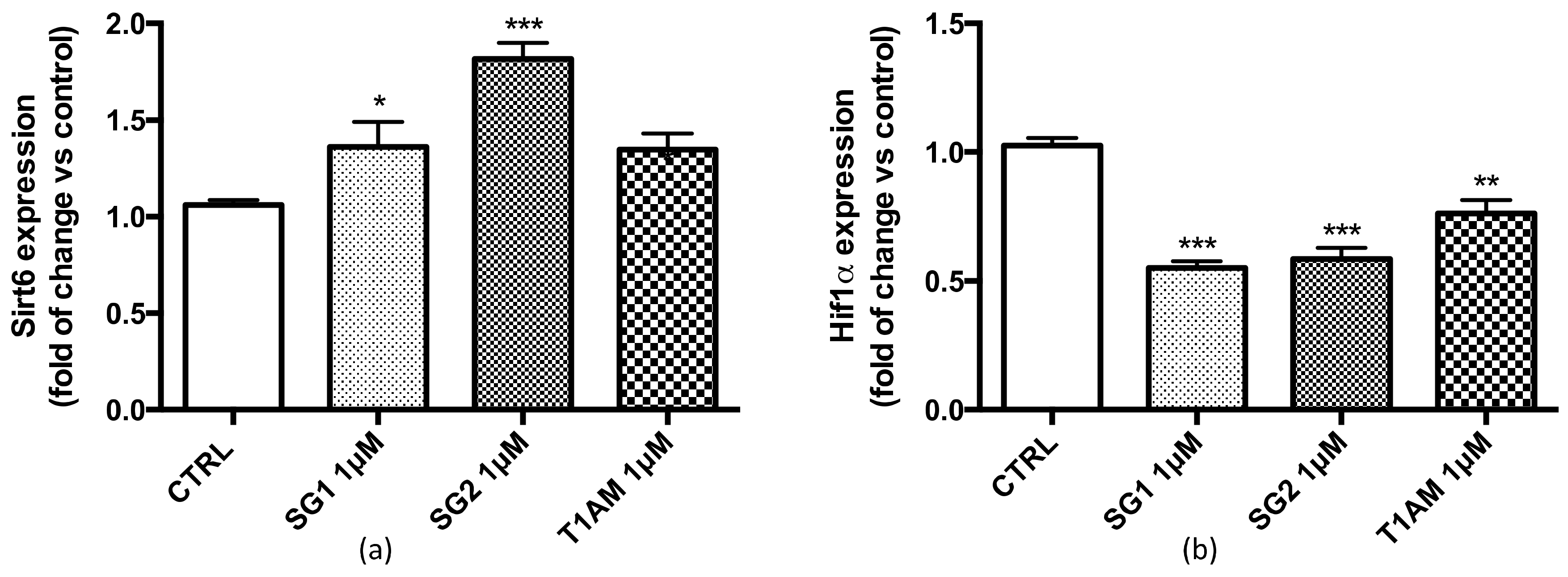
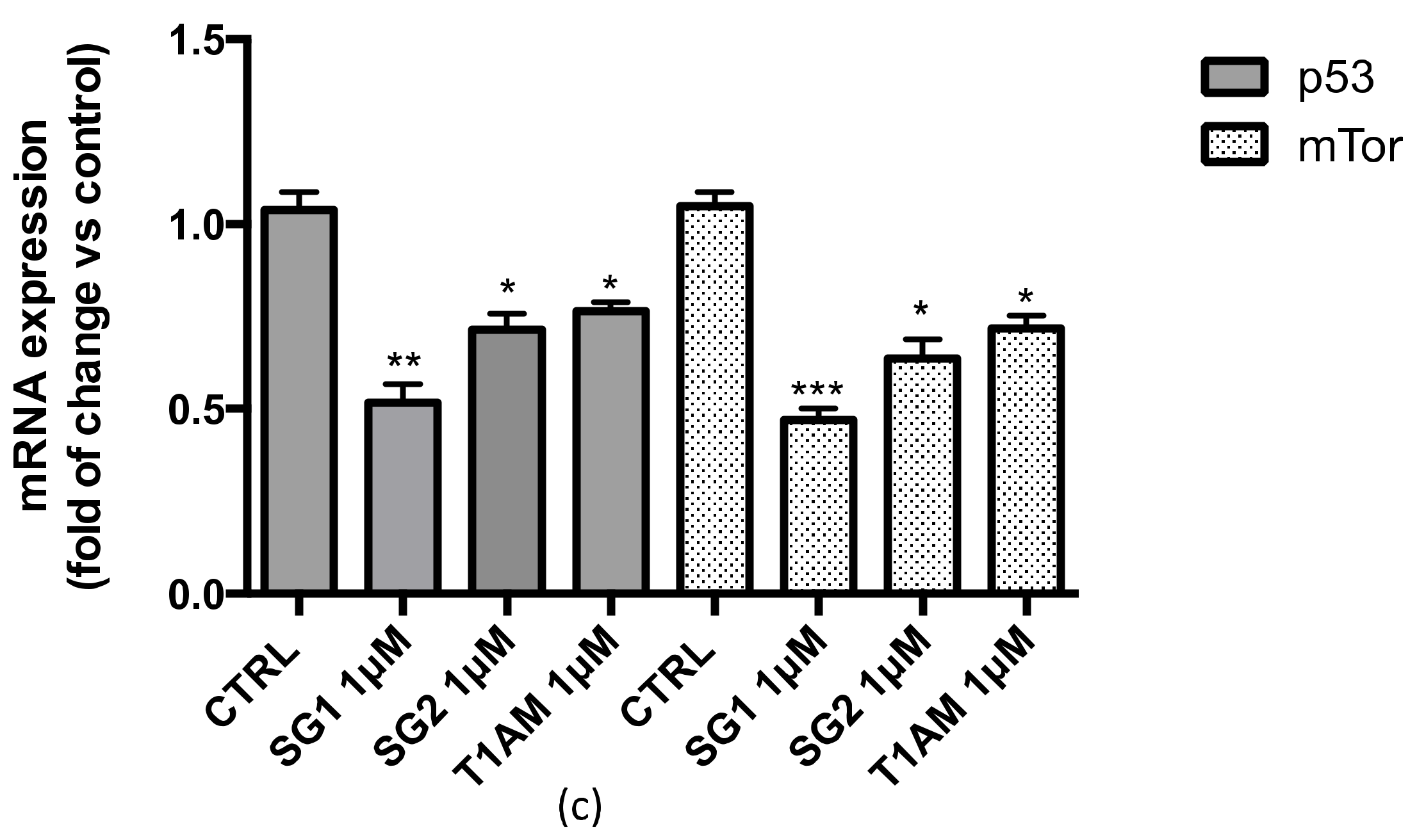
2.2. Effects of SG-1, SG-2 and T1AM on the Expression of SIRT6 and mTOR
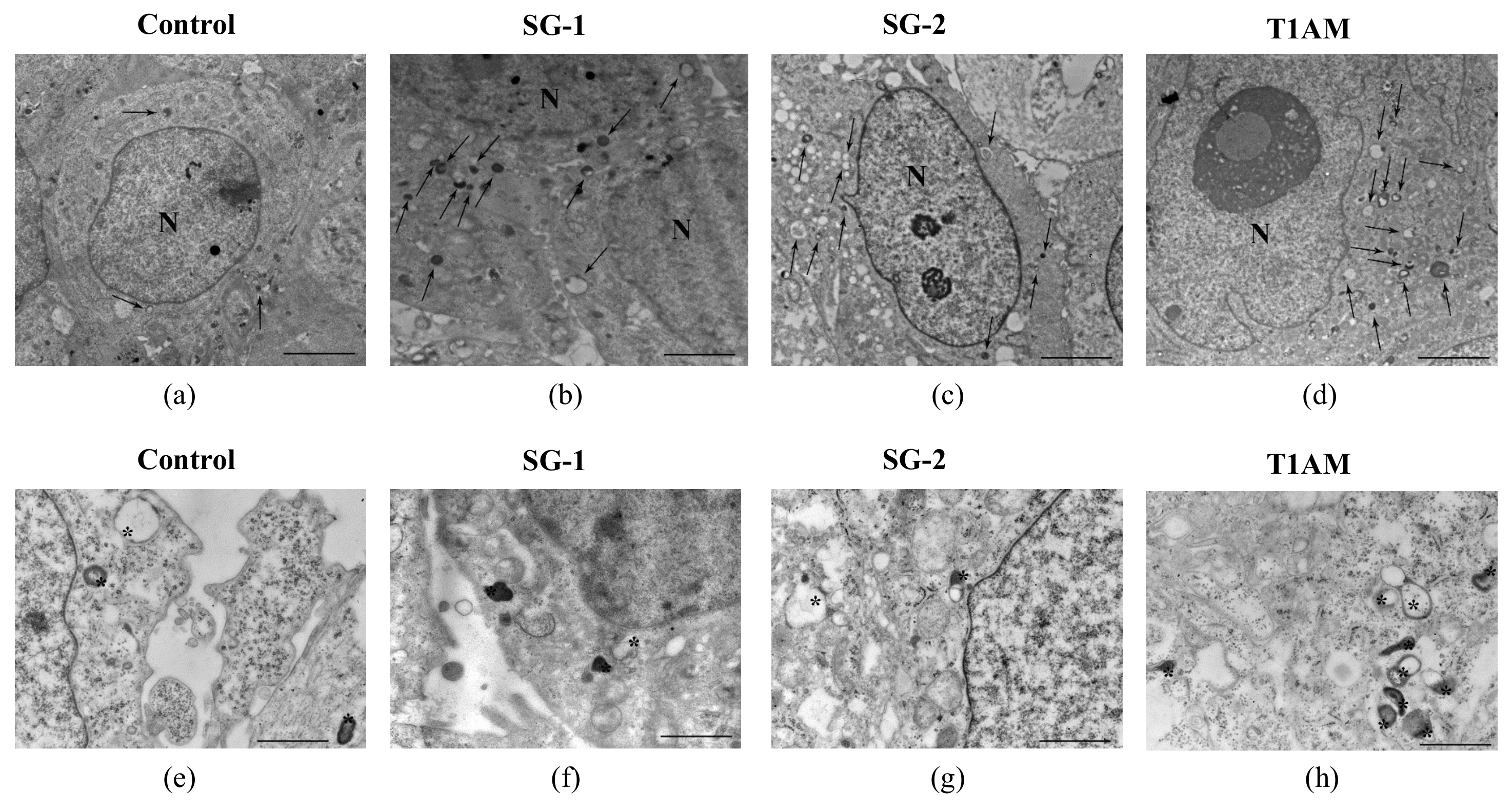
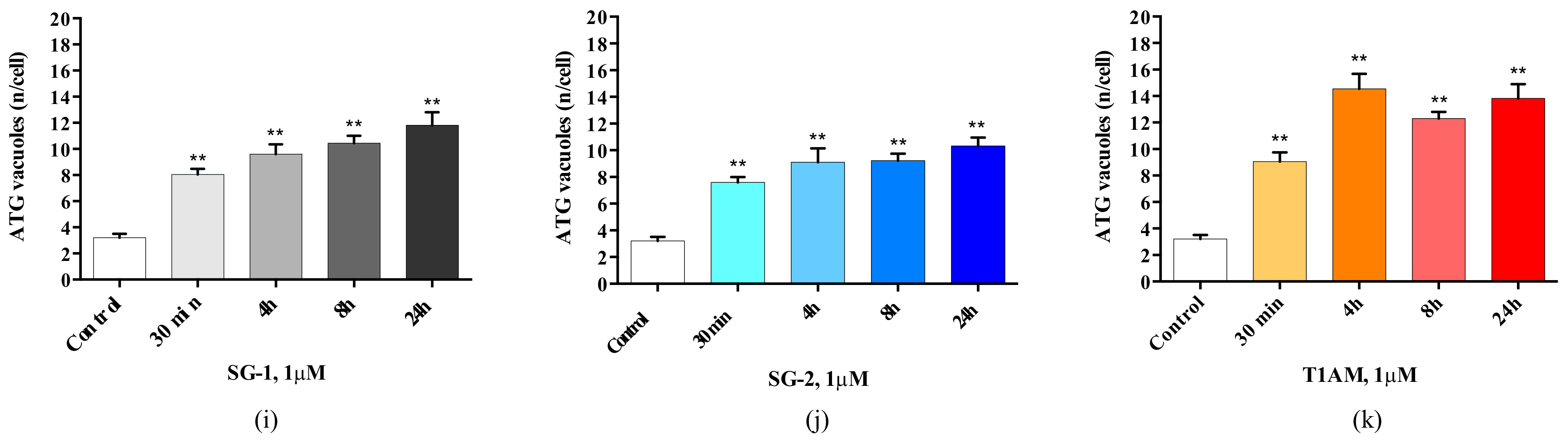
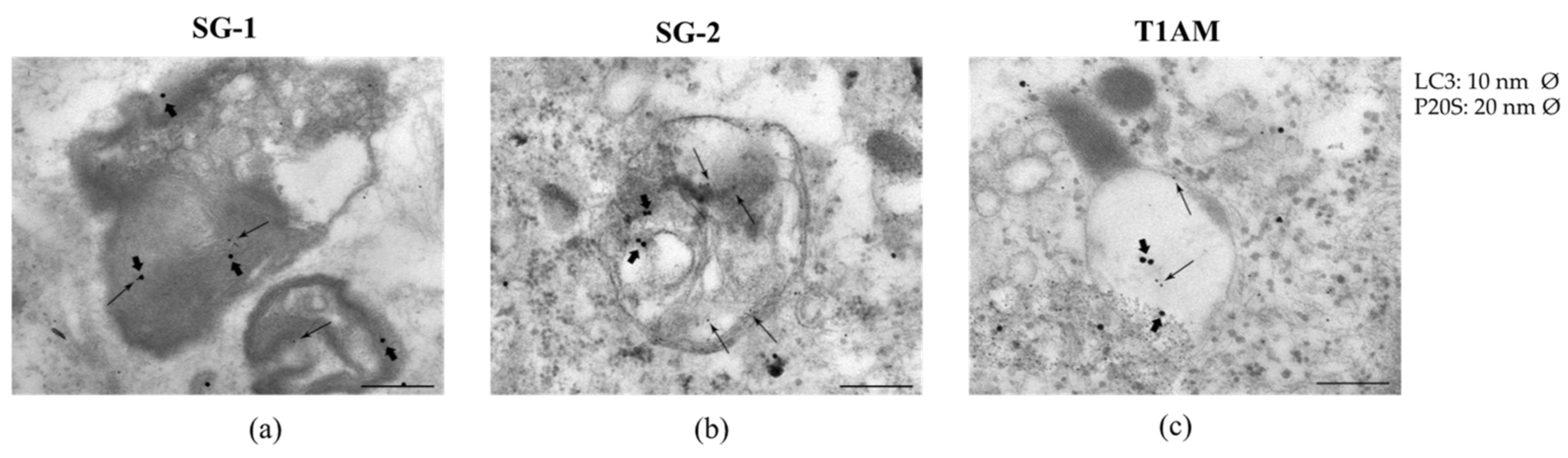
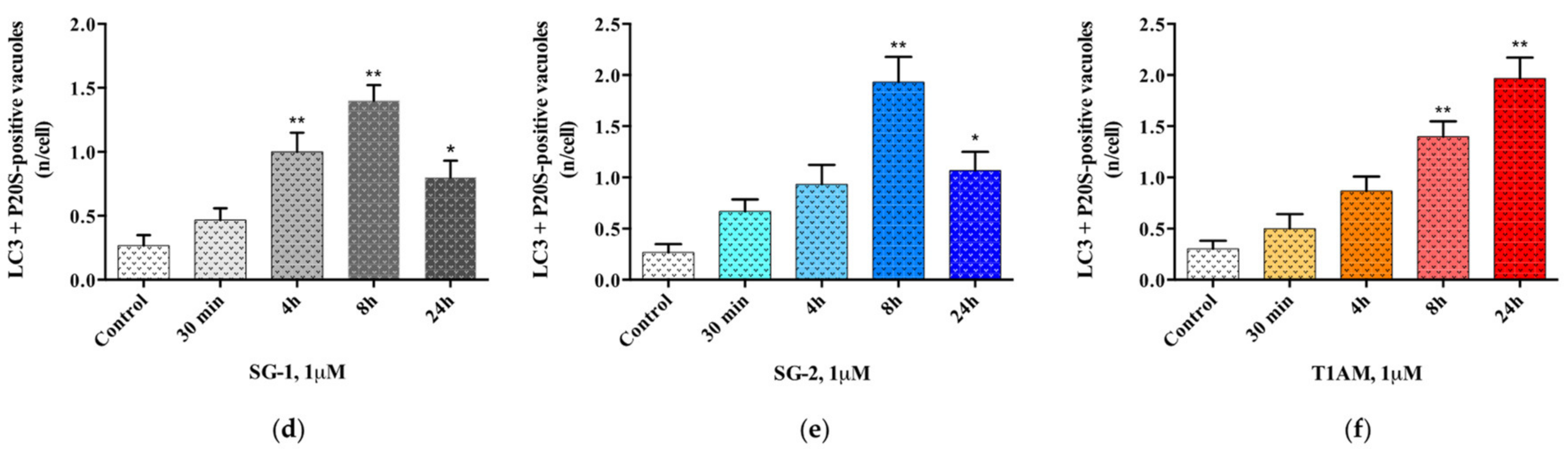
2.3. Effects of SG-1, SG-2 and T1AM on Cell Clearing Systems
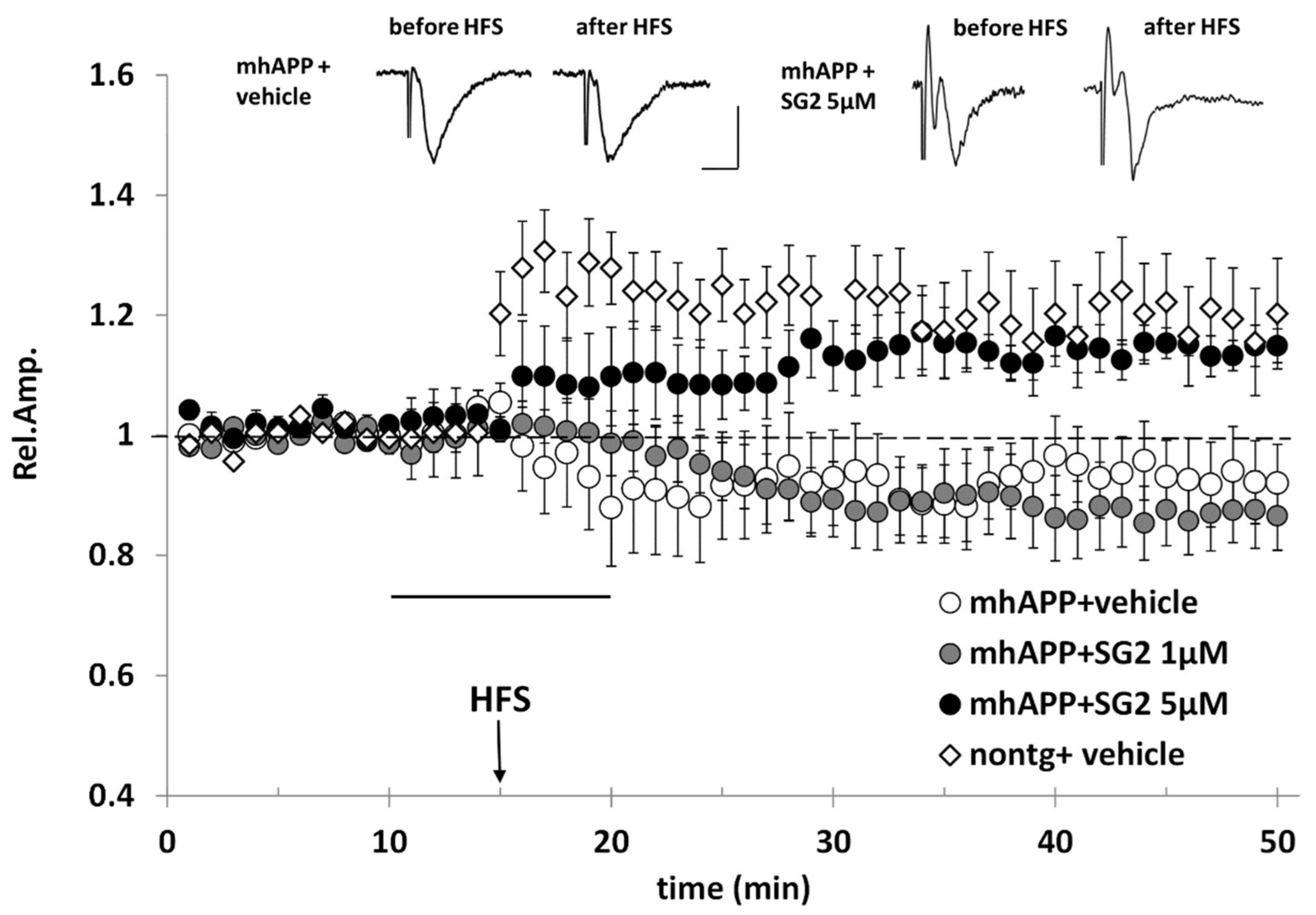
2.4. SG-2 Rescues LTP in hAPP-J20 Mouse Model of AD at the Level of the EC
3. Discussion
4. Materials and Methods
4.1. Chemicals
4.2. Cell Lines
4.3. Assessment of Autophagy-Like Vacuoles, and Count of LC3 and P20S Immunogold Particles
4.4. Gene Expression Analysis
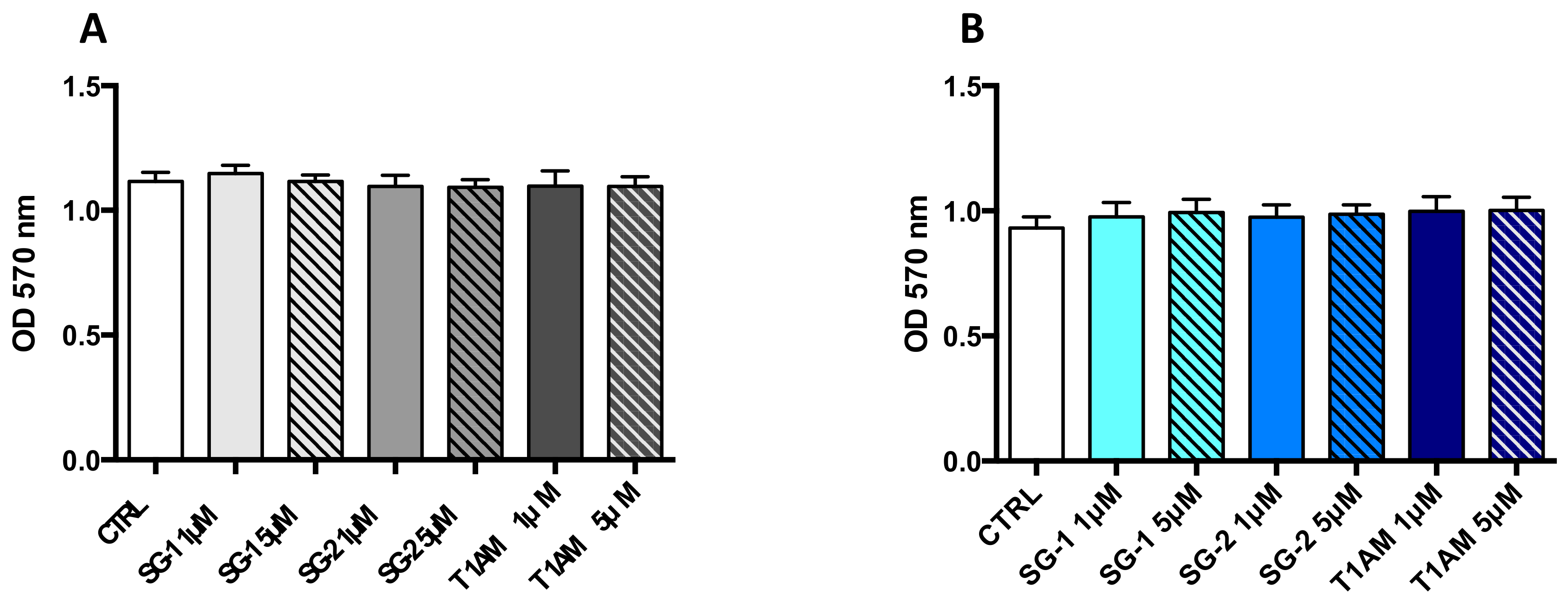
4.5. Western Blot Analysis and MTT Cell Viability Assay
4.6. Animals
4.7. In Vitro Electrophysiology
4.8. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Mazon, J.N.; de Mello, A.H.; Ferreira, G.K.; Rezin, G.T. The impact of obesity on neurodegenerative diseases. Life Sci. 2017, 182, 22–28. [Google Scholar] [CrossRef]
- Procaccini, C.; Santopaolo, M.; Faicchia, D.; Colamatteo, A.; Formisano, L.; de Candia, P.; Galgani, M.; De Rosa, V.; Matarese, G. Role of metabolism in neurodegenerative disorders. Metabolism 2016, 65, 1376–1390. [Google Scholar] [CrossRef]
- Cai, D. Neuroinflammation and neurodegeneration in overnutrition-induced diseases. Trends Endocrinol. Metab. 2013, 24, 40–47. [Google Scholar] [CrossRef]
- Lee, E.B.; Mattson, M.P. The Neuropathology of Obesity: Insights from Human Disease. Acta Neuropathol. 2014, 127, 3–28. [Google Scholar] [CrossRef]
- Perluigi, M.; Barone, E.; Di Domenico, F.; Butterfield, D.A. Aberrant protein phosphorylation in Alzheimer disease brain disturbs pro-survival and cell death pathways. Biochim. Biophys. Acta (BBA)—Mol. Basis Dis. 2016, 1862, 1871–1882. [Google Scholar] [CrossRef] [PubMed]
- Giblin, W.; Skinner, M.E.; Lombard, D.B. Sirtuins: Guardians of mammalian healthspan. Trends Genet. 2014, 30, 271–286. [Google Scholar] [CrossRef] [PubMed]
- Kugel, S.; Mostoslavsky, R. Chromatin and beyond: The multitasking roles for SIRT6. Trends Biochem. Sci. 2014, 39, 72–81. [Google Scholar] [CrossRef] [PubMed]
- Kaluski, S.; Portillo, M.; Besnard, A.; Stein, D.; Einav, M.; Zhong, L.; Ueberham, U.; Arendt, T.; Mostoslavsky, R.; Sahay, A.; et al. Neuroprotective Functions for the Histone Deacetylase SIRT6. Cell Rep. 2017, 18, 3052–3062. [Google Scholar] [CrossRef] [PubMed]
- Scanlan, T.S.; Suchland, K.L.; Hart, M.E.; Chiellini, G.; Huang, Y.; Kruzich, P.J.; Frascarelli, S.; Crossley, D.A.; Bunzow, J.R.; Ronca-Testoni, S.; et al. 3-Iodothyronamine is an endogenous and rapid-acting derivative of thyroid hormone. Nat. Med. 2004, 10, 638–642. [Google Scholar] [CrossRef] [PubMed]
- Scanlan, T.S. 3-Iodothyronamine (T1AM): A New Player on the Thyroid Endocrine Team? Endocrinology 2019, 150, 1108–1111. [Google Scholar] [CrossRef]
- Haviland, J.A.; Reiland, H.; Butz, D.E.; Tonelli, M.; Porter, W.P.; Zucchi, R.; Scanlan, T.S.; Chiellini, G.; Assadi-Porter, F.M. NMR-based metabolomics and breath studies show lipid and protein catabolism during low dose chronic T(1)AM treatment. Obesity 2013, 21, 2538–2544. [Google Scholar] [CrossRef] [PubMed]
- Assadi-Porter, F.M.; Reiland, H.; Sabatini, M.; Lorenzini, L.; Carnicelli, V.; Rogowski, M.; Selen Alpergin, E.S.; Tonelli, M.; Ghelardoni, S.; Saba, A.; et al. Metabolic Reprogramming by 3-Iodothyronamine (T1AM): A New Perspective to Reverse Obesity through Co-Regulation of Sirtuin 4 and 6 Expression. Int. J. Mol. Sci. 2018, 19, 1535. [Google Scholar] [CrossRef] [PubMed]
- Bellusci, L.; Laurino, A.; Sabatini, M.; Sestito, S.; Lenzi, P.; Raimondi, L.; Rapposelli, S.; Biagioni, F.; Fornai, F.; Salvetti, A.; et al. New Insights into the Potential Roles of 3-Iodothyronamine (T1AM) and Newly Developed Thyronamine-Like TAAR1 Agonists in Neuroprotection. Front. Pharmacol. 2017, 8, 905. [Google Scholar] [CrossRef] [PubMed]
- Cao, J.; Hou, J.; Ping, J.; Cai, D. Advances in developing novel therapeutic strategies for Alzheimer’s disease. Mol. Neurodegener. 2018, 13, 64. [Google Scholar] [CrossRef]
- Lenzi, P.; Lazzeri, G.; Biagioni, F.; Busceti, C.L.; Gambardella, S.; Salvetti, A.; Fornai, F. The Autophagoproteasome a Novel Cell Clearing Organelle in Baseline and Stimulated Conditions. Front. Neuroanat. 2016, 10, 78. [Google Scholar] [CrossRef]
- Blasiak, J.; Pawlowska, E.; Szczepanska, J.; Kaarniranta, K. Interplay between Autophagy and the Ubiquitin-Proteasome System and Its Role in the Pathogenesis of Age-Related Macular Degeneration. Int. J. Mol. Sci. 2019, 20, 210. [Google Scholar] [CrossRef]
- Accorroni, A.; Chiellini, G.; Origlia, N. Effects of Thyroid Hormones and their Metabolites on Learning and Memory in Normal and Pathological Conditions. Curr. Drug Metabalism 2017, 18, 225–236. [Google Scholar] [CrossRef]
- Accorroni, A.; Criscuolo, C.; Sabatini, M.; Donzelli, R.; Saba, A.; Origlia, N.; Zucchi, R. 3-Iodothyronamine and trace amine-associated receptor 1 are involved in the expression of long-term potentiation in mouse entorhinal cortex. Eur. Thyroid J. 2016, 5 (Suppl. 1), 57–176. [Google Scholar]
- Origlia, N.; Righi, M.; Capsoni, S.; Cattaneo, A.; Fang, F.; Stern, D.M.; Chen, J.X.; Schmidt, A.M.; Arancio, O.; Yan, S.D.; et al. Receptor for advanced glycation end product-dependent activation of p38 mitogen-activated protein kinase contributes to amyloid-beta-mediated cortical synaptic dysfunction. J. Neurosci. 2008, 28, 3521–3523. [Google Scholar] [CrossRef]
- Chiellini, G.; Bellusci, L.; Sabatini, M.; Zucchi, R. Thyronamines and Analogues—The Route from Rediscovery to Translational Research on Thyronergic Amines. Mol. Cell. Endocrinol. 2017, 458, 149–155. [Google Scholar] [CrossRef]
- Rogowski, M.; Bellusci, L.; Sabatini, M.; Rapposelli, S.; Rahman, S.M.; Chiellini, G.; Assadi-Porter, F.M. Lipolytic Effects of 3-Iodothyronamine (T1AM) and a Novel Thyronamine-Like Analog SG-2 through the AMPK Pathway. Int. J. Mol. Sci. 2019, 20, 4054. [Google Scholar] [CrossRef] [PubMed]
- Nixon, R.A. The role of autophagy in neurodegenerative disease. Nat. Med. 2013, 19, 983–997. [Google Scholar] [CrossRef] [PubMed]
- Wei, Y.; Liu, M.; Li, X.; Liu, J.; Li, H. Origin of the Autophagosome Membrane in Mammals. Biomed Res. Int. 2018, 1012789. [Google Scholar] [CrossRef] [PubMed]
- Condello, M.; Pellegrini, E.; Caraglia, M.; Meschini, S. Targeting Autophagy to Overcome Human Diseases. Int. J. Mol. Sci. 2019, 20, 725. [Google Scholar] [CrossRef]
- Shao, J.; Yang, X.; Liu, T.; Zhang, T.; Xie, Q.R.; Xia, W. Autophagy induction by SIRT6 is involved in oxidative stress-induced neuronal damage. Protein Cell 2016, 7, 281–290. [Google Scholar] [CrossRef]
- Iachettini, S.; Trisciuoglio, D.; Rotili, D.; Lucidi, A.; Salvati, E.; Zizza, P.; Di Leo, L.; Del Bufalo, D.; Ciriolo, M.R.; Leonetti, C.; et al. Pharmacological activation of SIRT6 triggers lethal autophagy in human cancer cells. Cell Death Discov. 2018, 9, 996. [Google Scholar] [CrossRef]
- Takasaka, N.; Araya, J.; Hara, H.; Ito, S.; Kobayashi, K.; Kurita, Y.; Wakui, H.; Yoshii, Y.; Yumino, Y.; Fujii, S.; et al. Autophagy induction by SIRT6 through attenuation of insulin-like growth factor signaling is involved in the regulation of human bronchial epithelial cell senescence. J. Immunol. 2014, 192, 958–968. [Google Scholar] [CrossRef]
- Zhong, L.; D’Urso, A.; Toiber, D.; Sebastian, C.; Henry, R.E.; Vadysirisack, D.D.; Guimaraes, A.; Marinelli, B.; Wikstrom, J.D.; Nir, T.; et al. The histone deacetylase Sirt6 regulates glucose homeostasis via Hif1alpha. Cell 2010, 140, 280–293. [Google Scholar] [CrossRef]
- Brown, C.J.; Lain, S.; Verma, C.S.; Fersht, A.R.; Lane, D.P. Awakening guardian angels: Drugging the p53 pathway. Nat. Rev. Cancer 2009, 9, 862–873. [Google Scholar] [CrossRef]
- Chang, J.R.; Ghafouri, M.; Mukerjee, R.; Bagashev, A.; Chabrashvili, T.; Sawaya, B.E. Role of p53 in neurodegenerative diseases. Neurodegener. Dis. 2012, 9, 68–80. [Google Scholar] [CrossRef]
- Tasdemir, E.; Maiuri, M.C.; Galluzzi, L.; Vitale, I.; Djavaheri-Mergny, M.; D’Amelio, M.; Criollo, A.; Morselli, E.; Zhu, C.; Harper, F.; et al. Regulation of autophagy by cytoplasmic p53. Nat. Cell Biol. 2008, 10, 676–687. [Google Scholar] [CrossRef] [PubMed]
- Magalingam, K.B.; Radhakrishnan, A.; Ping, N.S.; Haleagrahara, N. Current Concepts of Neurodegenerative Mechanisms in Alzheimer’s Disease. Biomed Res. Int. 2018, 3740461. [Google Scholar] [CrossRef] [PubMed]
- Mucke, L.; Masliah, E.; Yu, G.Q.; Mallory, M.; Rockenstein, E.M.; Tatsuno, G.; Hu, K.; Kholodenko, D.; Johnson-Wood, K.; McConlogue, L. High-level neuronal expression of abeta 1-42 in wild-type human amyloid protein precursor transgenic mice: Synaptotoxicity without plaque formation. J. Neurosci. 2000, 20, 4050–4058. [Google Scholar] [CrossRef] [PubMed]
- Criscuolo, C.; Fontebasso, V.; Middei, S.; Stazi, M.; Ammassari-Teule, M.; Yan, S.S.; Origlia, N. Entorhinal Cortex dysfunction can be rescued by inhibition of microglial RAGE in an Alzheimer’s disease mouse model. Sci. Rep. 2017, 7, 42370. [Google Scholar] [CrossRef]
- Accorroni, A.; Rutigliano, G.; Sabatini, M.; Frascarelli, S.; Borsò, M.; Novelli, E.; Bandini, L.; Ghelardoni, S.; Saba, A.; Zucchi, R.; et al. Exogenous 3-iodothyronamine rescues the entorhinal cortex from β-amyloid toxicity. Thyroid 2019. [Google Scholar] [CrossRef]
- Feng, J.; Yan, P.F.; Zhao, H.Y.; Zhang, F.C.; Zhao, W.H.; Feng, M. SIRT6 suppresses glioma cell growth via induction of apoptosis, inhibition of oxidative stress and suppression of JAK2/STAT3 signaling pathway activation. Oncol. Rep. 2016, 35, 1395–1402. [Google Scholar] [CrossRef]
- Rogowski, M.; Gollahon, L.; Chellini, G.; Assadi-Porter, F.M. Uptake of 3-iodothyronamine hormone analogs inhibits the growth and viability of cancer cells. FEBS Open Bio 2017, 7, 587–601. [Google Scholar] [CrossRef]
- Chiellini, G.; Nesi, G.; Digiacomo, M.; Malvasi, R.; Espinoza, S.; Sabatini, M.; Frascarelli, S.; Laurino, A.; Cichero, E.; Macchia, M.; et al. Design, Synthesis, and Evaluation of Thyronamine Analogues as Novel Potent Mouse Trace Amine Associated Receptor 1 (mTAAR1) Agonists. J. Med. Chem. 2015, 58, 5096–5107. [Google Scholar] [CrossRef]
- Klionsky, D.J. Developing a set of guidelines for your research field: A practical approach. Mol. Biol. Cell 2016, 27, 733–738. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]

| Reference Sequence (RefSeq) mRNA | Gene Symbol | Sense and Anti-Sense Sequences | Length of the Amplicon (bp) | |
|---|---|---|---|---|
| NM_001243084 | HIF1A | sense | TTGGCAGCAACGACACAG | 169 bp |
| anti-sense | GCAGGGTCAGCACTACTTC | |||
| NM_004958 | MTOR | sense | TGCCTTCACAGATACCCAGTA | 171 bp |
| anti-sense | AGACCTCACAGCCACAGA | |||
| NM_000546 | P53 | sense | TCAACAAGATGTTTTGCCAACTG | 118 bp |
| anti-sense | ATGTGCTGTGACTGCTTGTAGATG | |||
| NM_016539 | SIRT6 | sense | CTCCTCCGCTTCCTGGTC | 119 bp |
| anti-sense | TTACACTTGGCACATTCTTCC | |||
| NM_002046 | GAPDH | sense | CCCTTCATTGACCTCAACTACATG | 115bp |
| anti-sense | TGGGATTTCCATTGATGACAAGC | |||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bellusci, L.; Runfola, M.; Carnicelli, V.; Sestito, S.; Fulceri, F.; Santucci, F.; Lenzi, P.; Fornai, F.; Rapposelli, S.; Origlia, N.; et al. Endogenous 3-Iodothyronamine (T1AM) and Synthetic Thyronamine-Like Analog SG-2 Act as Novel Pleiotropic Neuroprotective Agents through the Modulation of SIRT6. Molecules 2020, 25, 1054. https://doi.org/10.3390/molecules25051054
Bellusci L, Runfola M, Carnicelli V, Sestito S, Fulceri F, Santucci F, Lenzi P, Fornai F, Rapposelli S, Origlia N, et al. Endogenous 3-Iodothyronamine (T1AM) and Synthetic Thyronamine-Like Analog SG-2 Act as Novel Pleiotropic Neuroprotective Agents through the Modulation of SIRT6. Molecules. 2020; 25(5):1054. https://doi.org/10.3390/molecules25051054
Chicago/Turabian StyleBellusci, Lorenza, Massimiliano Runfola, Vittoria Carnicelli, Simona Sestito, Federica Fulceri, Filippo Santucci, Paola Lenzi, Francesco Fornai, Simona Rapposelli, Nicola Origlia, and et al. 2020. "Endogenous 3-Iodothyronamine (T1AM) and Synthetic Thyronamine-Like Analog SG-2 Act as Novel Pleiotropic Neuroprotective Agents through the Modulation of SIRT6" Molecules 25, no. 5: 1054. https://doi.org/10.3390/molecules25051054
APA StyleBellusci, L., Runfola, M., Carnicelli, V., Sestito, S., Fulceri, F., Santucci, F., Lenzi, P., Fornai, F., Rapposelli, S., Origlia, N., Zucchi, R., & Chiellini, G. (2020). Endogenous 3-Iodothyronamine (T1AM) and Synthetic Thyronamine-Like Analog SG-2 Act as Novel Pleiotropic Neuroprotective Agents through the Modulation of SIRT6. Molecules, 25(5), 1054. https://doi.org/10.3390/molecules25051054












