The Generation of the Oxidant Agent of a Mononuclear Nonheme Fe(II) Biomimetic Complex by Oxidative Decarboxylation. A DFT Investigation
Abstract
1. Introduction
2. Results and Discussion
3. Conclusions
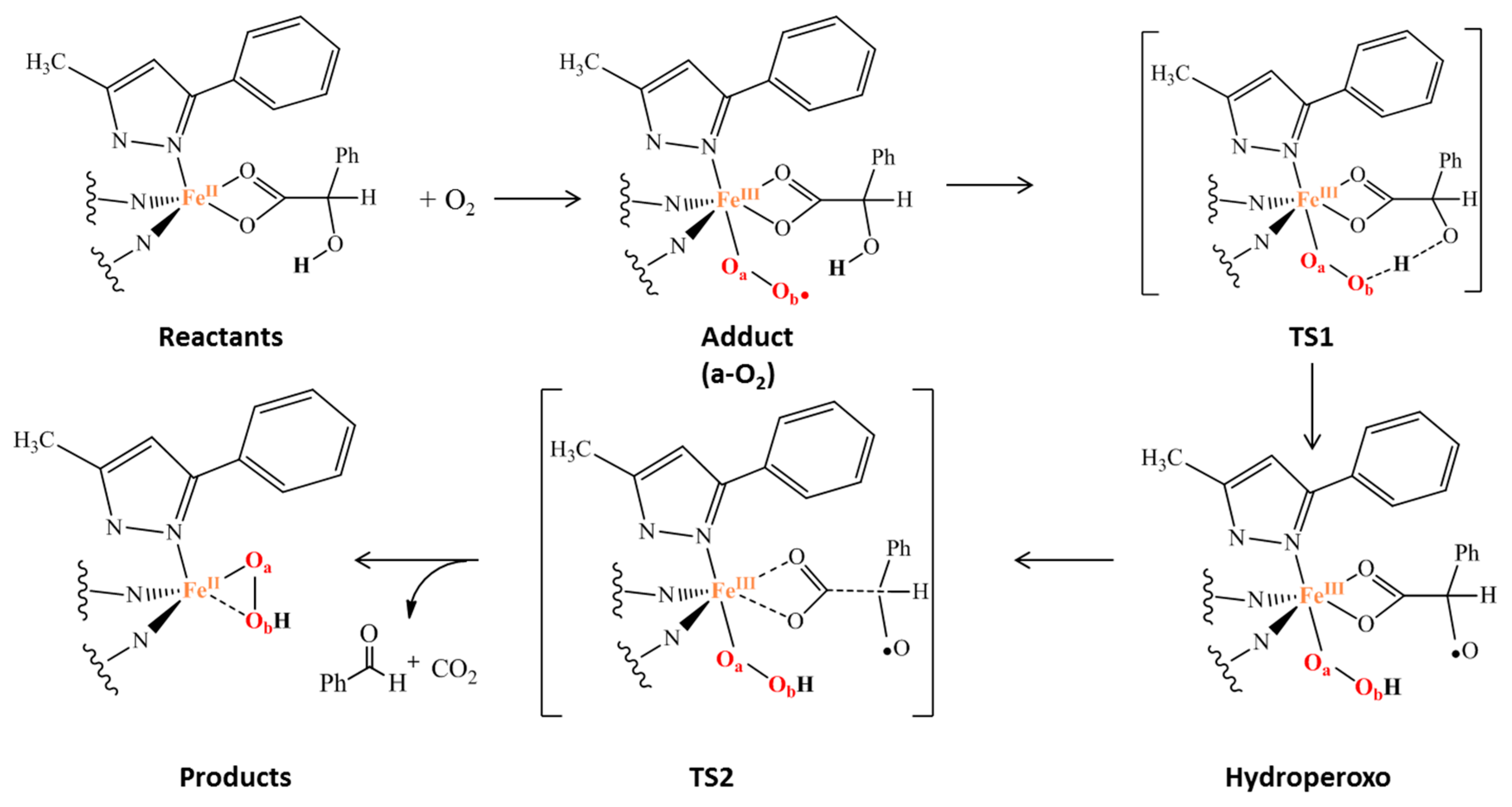
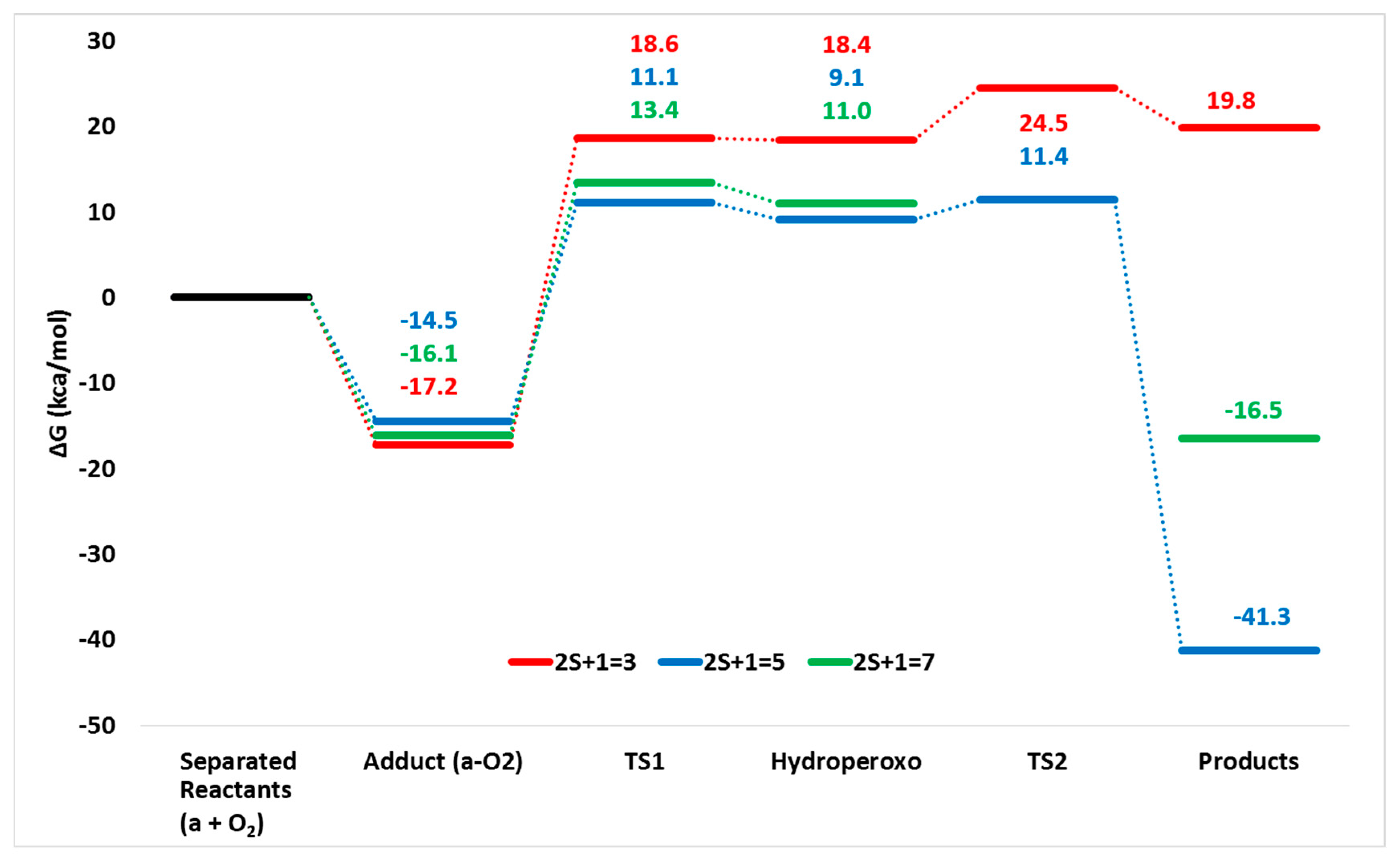
- The favored reaction path occurs in the quintet spin state as in the corresponding Rieske oxygenase enzymes;
- A spin cross takes place between the triplet and quintet electronic spin states after the adduct formation;
- The α-hydroxy group plays an important role in the oxidative decarboxylation;
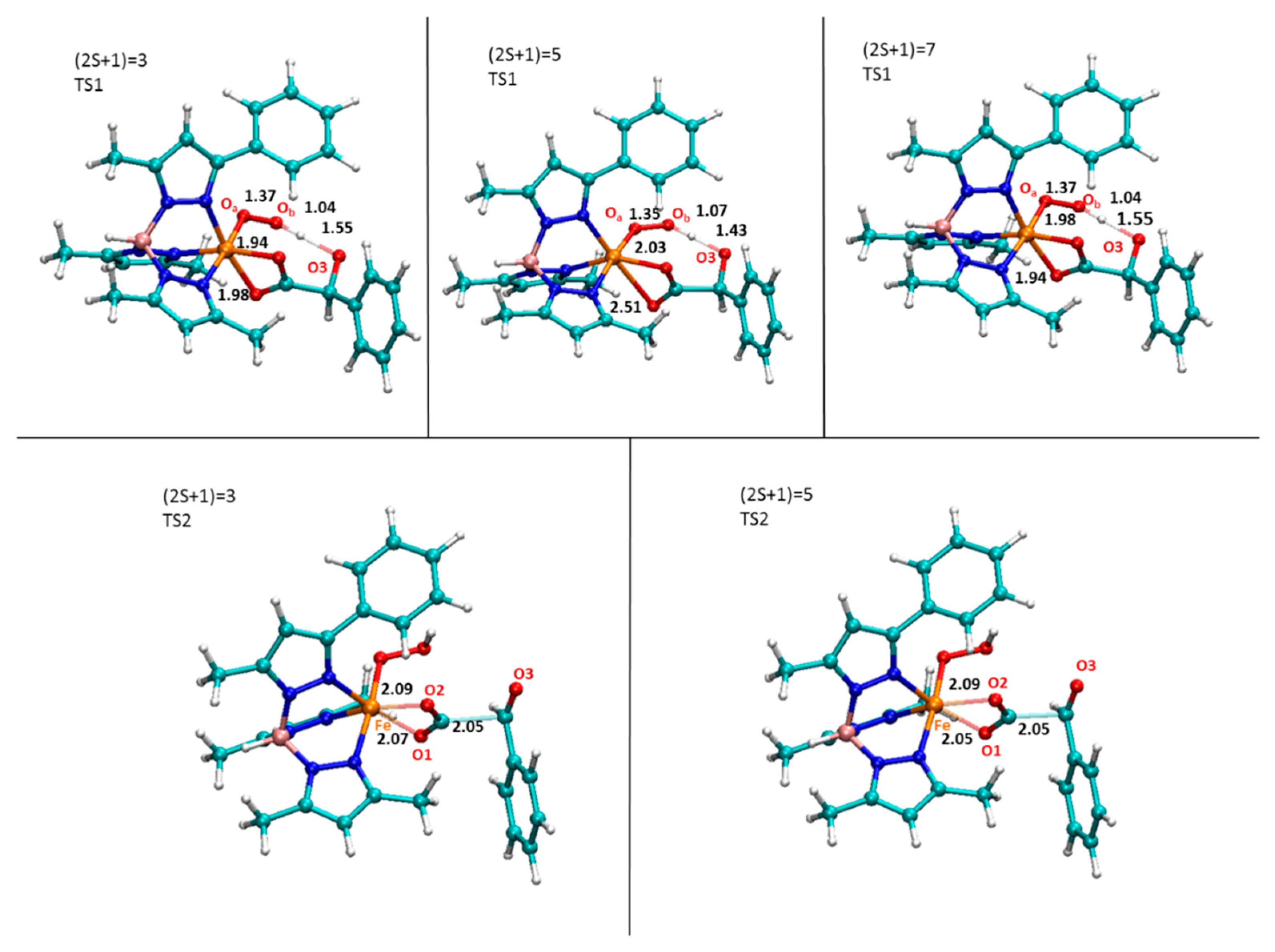
- The rate determining step is the proton transfer;
- The overall reaction results to be more exergonic in the quintet PES.
4. Computational Details
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
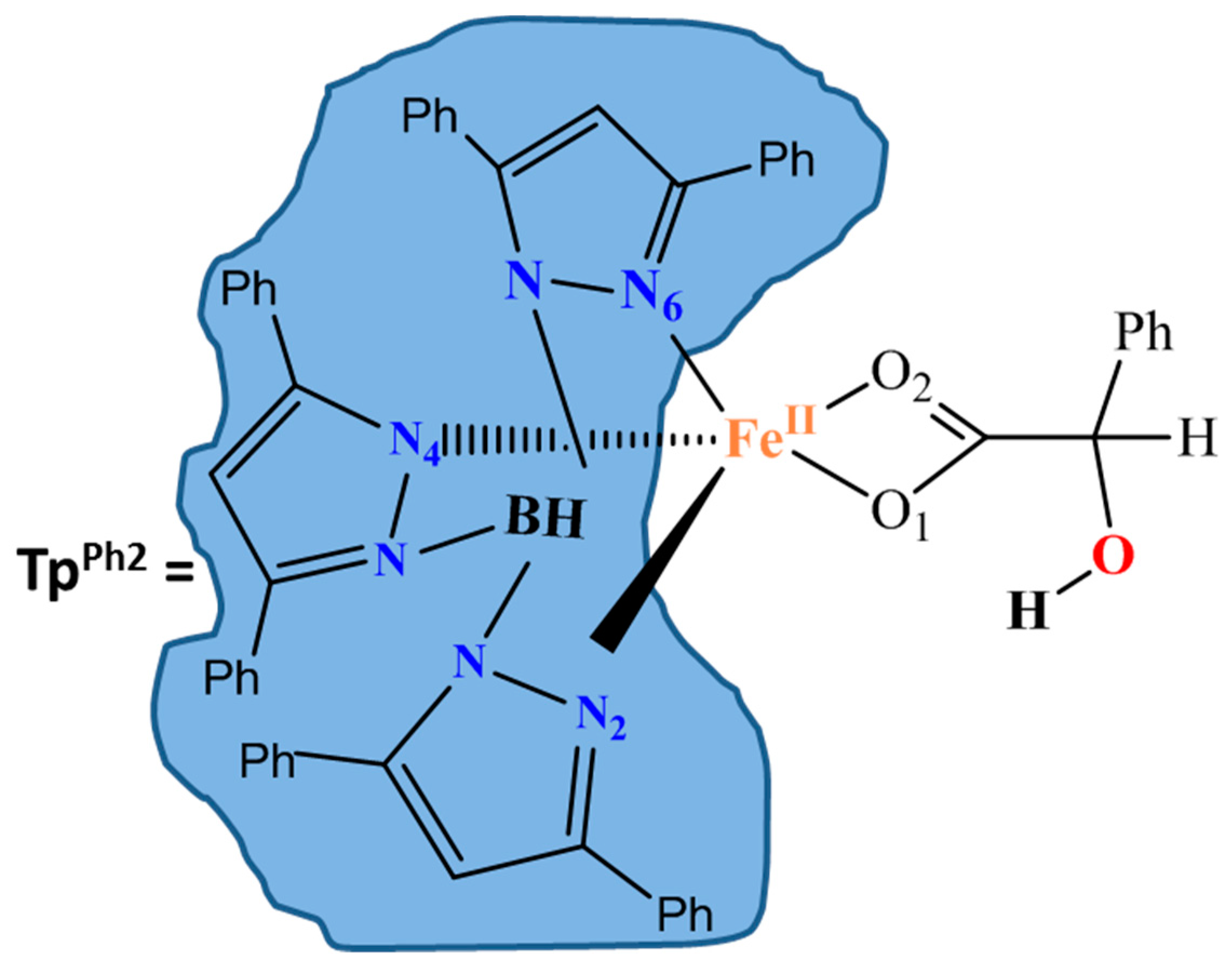
| TpPh2 | hydrotris (3,5-diphenylpyrazole-1-yl) borate |
| DFT | Density Functional Theory |
| PES | Potential Energy Surface |
| PCM | Polarizable Continuum Model |
| HOMO | High Occupied Molecular Orbital |
References
- Costas, M.; Mehn, M.P.; Jensen, M.P.; Que, L., Jr. Dioxygen activation at mononuclear nonheme iron active sites: Enzymes, models, and intermediates. Chem. Rev. 2004, 104, 939–986. [Google Scholar] [CrossRef] [PubMed]
- Talsi, E.P.; Bryliakov, K.P. Chemo- and stereoselective CH oxidations and epoxidations/cis-dihydroxylations with H2O2, catalyzed by non-heme iron and manganese complexes. Coord. Chem. Rev. 2012, 256, 1418–1434. [Google Scholar] [CrossRef]
- Paria, S.; Chatterjee, S.; Paine, T.K. Reactivity of an Iron− Oxygen Oxidant Generated upon Oxidative Decarboxylation of Biomimetic Iron(II) α-Hydroxy Acid Complexes. Inorg. Chem. 2014, 53, 2810–2821. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Costas, M.; Kim, J.; Tipton, A.K.; Que, L., Jr. Olefin Cis-Dihydroxylation versus Epoxidation by Non-Heme Iron Catalysts: Two Faces of an FeIII−OOH Coin. J. Am. Chem. Soc. 2002, 124, 3026–3035. [Google Scholar] [CrossRef] [PubMed]
- Thibon, A.; England, J.; Martinho, M.; Young, V.G., Jr.; Frisch, J.R.; Guillot, R.; Girerd, J.-J.; Münck, E.; Que, L., Jr.; Banse, F. Proton- and Reductant-Assisted Dioxygen Activation by a Nonheme Iron(II) Complex to Form an Oxoiron(IV) Intermediate. Angew. Chem. Int. Ed. Engl. 2008, 47, 7064–7067. [Google Scholar] [CrossRef] [PubMed]
- Hong, S.; Lee, Y.-M.; Shin, W.; Fukuzumi, S.; Nam, W. Dioxygen Activation by Mononuclear Nonheme Iron(II) Complexes Generates Iron−Oxygen Intermediates in the Presence of an NADH Analogue and Proton. J. Am. Chem. Soc. 2009, 131, 13910–13911. [Google Scholar] [CrossRef]
- Mukherjee, A.; Martinho, M.; Bominaar, E.L.; Münck, E.; Que, L., Jr. Shape-Selective Interception by Hydrocarbons of the O2- Derived Oxidant of a Biomimetic Nonheme Iron Complex. Angew. Chem. Int. Ed. Engl. 2009, 48, 1780–1783. [Google Scholar] [CrossRef]
- Martinho, M.; Blain, G.; Banse, F. Activation of dioxygen by a mononuclear non-heme iron complex: Characterization of a Fe(III)-(OOH) intermediate. Dalton Trans. 2010, 39, 1630–1634. [Google Scholar] [CrossRef]
- Lee, Y.-M.; Hong, S.; Morimoto, Y.; Shin, W.; Fukuzumi, S.; Nam, W. Dioxygen Activation by a Non-Heme Iron(II) Complex: Formation of an Iron(IV)−Oxo Complex via C−H Activation by a Putative Iron(III)−Superoxo Species. J. Am. Chem. Soc. 2010, 132, 10668–10670. [Google Scholar] [CrossRef]
- Mandon, D.; Jaafar, H.; Thibon, A. Exploring the Oxygen sensitivity of FeCl2 complexes with tris(2-pyridylmethyl)amine-type ligands: O2 coordination and a quest for superoxide. N. J. Chem. 2011, 35, 1986–2000. [Google Scholar] [CrossRef]
- Badiei, Y.M.; Siegler, M.A.; Goldberg, D.P. O2 Activation by Bis(imino)pyridine Iron(II)-Thiolate Complexes. J. Am. Chem. Soc. 2011, 133, 1274–1277. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Goldsmith, R. Observation of a ferric hydroperoxide complex during the non-heme iron catalysed oxidation of alkenes and alkanes by O2. Chem. Commun. 2012, 48, 10532–10534. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Van Heuvelen, K.M.; Meier, K.K.; Münck, E.; Que, L., Jr. Sc3+-Triggered Oxoiron(IV) Formation from O2 and its Non-Heme Iron(II) Precursor via a Sc3+−Peroxo−Fe3+ Intermediate. J. Am. Chem. Soc. 2013, 135, 10198–10201. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.O.; Sastri, C.V.; Seo, M.S.; Kim, J.; Nam, W. Dioxygen Activation and Catalytic Aerobic Oxidation by a Mononuclear Nonheme Iron(II) Complex. J. Am. Chem. Soc. 2005, 127, 4178–4179. [Google Scholar] [CrossRef]
- Bruijnincx, P.C.A.; van Koten, G.; Gebbink, R.J.M.K. Mononuclear non-heme iron enzymes with the 2-His-1-carboxylate facial triad: Recent developments in enzymology and modeling studies. Chem. Soc. Rev. 2008, 37, 2716–2744. [Google Scholar] [CrossRef]
- Shook, R.L.; Peterson, S.M.; Greaves, J.; Moore, C.; Rheingold, A.L.; Borovik, A.S. Catalytic Reduction of Dioxygen to Water with a Monomeric Manganese Complex at Room Temperature. J. Am. Chem. Soc. 2011, 133, 5810–5817. [Google Scholar] [CrossRef]
- Paine, T.K.; Que, L., Jr. Dioxygen Activation by Biomimetic Iron(II) Complexes of α-Keto Acids and α-Hydroxy Acids. Struct. Bond. 2014, 160, 39–56. [Google Scholar]
- Sallmann, M.; Limberg, C. Utilizing the Trispyrazolyl Borate Ligand for the Mimicking of O2-Activating Mononuclear Nonheme Iron Enzymes. Acc. Chem. Res. 2015, 48, 2734–2743. [Google Scholar] [CrossRef]
- Sahu, S.; Goldberg, D.P. Activation of Dioxygen by Iron and Manganese Complexes: A Heme and Nonheme Perspective. J. Am. Chem. Soc. 2016, 138, 11410–11428. [Google Scholar] [CrossRef]
- Funabiki, T. Functional model oxygenations by nonheme iron complexes. In Advances in Catalytic Activation of Dioxygen by Metal Complexes; Simandi, L.I., Ed.; Kluwer Academic Publishers: Dordrecht, The Netherlands, 2003; pp. 157–226. [Google Scholar]
- Chatterjee, S.; Paine, T.K. Olefin cis-Dihydroxylation and Aliphatic C-H Bond Oxygenation by a Dioxygen-Derived Electrophilic Iron-Oxygen Oxidant. Angew. Chem. Int. Ed. Engl. 2015, 54, 9338–9342. [Google Scholar] [CrossRef]
- Chatterjee, S.; Paine, T.K. Hydroxylation versus Halogenation of Aliphatic C-H Bonds by a Dioxygen-Derived Iron−Oxygen Oxidant: Functional Mimicking of Iron Halogenases. Angew. Chem. Int. Ed. Engl. 2016, 55, 7717–7722. [Google Scholar] [CrossRef] [PubMed]
- Paria, S.; Que, L., Jr.; Paine, T.K. Oxidative Decarboxylation of Benzilic Acid by a Biomimetic Iron(II) Complex: Evidence for an Iron(IV)−Oxo−Hydroxo Oxidant from O2. Angew. Chem. Int. Ed. Engl. 2011, 50, 11129–11132. [Google Scholar] [CrossRef] [PubMed]
- Costas, M.; Que, L., Jr. Ligand Topology Tuning of Iron-Catalyzed Hydrocarbon Oxidations. Angew. Chem. Int. Ed. Engl. 2002, 41, 2179–2181. [Google Scholar] [CrossRef]
- Oldenburg, P.D.; Feng, Y.; Pryjomska-Ray, I.; Ness, D.; Que, L., Jr. Olefin cis-dihydroxylation with bio-inspired iron catalysts. evidence for an Fe(II)/Fe(IV) catalytic cycle. J. Am. Chem. Soc. 2010, 132, 17713–17723. [Google Scholar]
- Borowski, T.; Bassan, A.; Siegbahn, P.E.M. A Hybrid Density Functional Study of O−O Bond Cleavage and Phenyl Ring Hydroxylation for a Biomimetic Non-Heme Iron Complex. Inorg. Chem. 2004, 43, 3277–3291. [Google Scholar] [CrossRef]
- Mehn, M.P.; Fujisawa, K.; Hegg, E.L.; Que, L., Jr. Oxygen Activation by Nonheme Iron(II) Complexes: α-Keto Carboxylate versus Carboxylate. J. Am. Chem. Soc. 2003, 125, 7828–7842. [Google Scholar] [CrossRef]
- Siegbahn, P.E.M.; Borowski, T. Modeling Enzymatic Reactions Involving Transition Metals. Acc. Chem. Res. 2006, 39, 729–738. [Google Scholar] [CrossRef]
- Borowski, T.; Bassan, A.; Siegbahn, P.E.M. Mechanism of Dioxygen Activation in 2-Oxoglutarate-Dependent Enzymes: A Hybrid DFT Study. Chem. Eur. J. 2004, 10, 1031–1041. [Google Scholar] [CrossRef]
- Frisch, M.J.; Trucks, G.W.; Schlegel, H.B.; Scuseria, G.E.; Robb, M.A.; Cheeseman, J.R.; Zakrzewski, V.G.; Montgomery, J.A., Jr.; Stratmann, R.E.; Burant, J.C.; et al. Gaussian 2009; Gaussian, Inc.: Wallingford, CT, USA, 2009. [Google Scholar]
- Zhao, Y.; Truhlar, D.G. A new local density functional for main-group thermochemistry, transition metal bonding, thermochemical kinetics, and noncovalent interactions. J. Chem. Phys. 2006, 125, 194101. [Google Scholar] [CrossRef]
- Valero, R.; Costa, R.; Moreira, I.D.P.R.; Truhlar, D.G.; Illas, F. Performance of the M06 family of exchange-correlation functionals for predicting magnetic coupling in organic and inorganic molecules. J. Chem. Phys. 2008, 128, 114103. [Google Scholar] [CrossRef]
- Marzouk, A.; Bolvin, H.; Reinhardt, P.; Manceron, L.; Perchard, J.P.; Tremblay, B.; Alikhani, M.E. A Combined Experimental and Theoretical Study of the Ti2 + N2O Reaction. J. Phys. Chem. A 2014, 118, 561–572. [Google Scholar] [CrossRef]
- Gusev, D.G. Assessing the Accuracy of M06-L Organometallic Thermochemistry. Organometallics 2013, 32, 4239–4243. [Google Scholar] [CrossRef]
- Marino, T.; Fortino, M.G.; Russo, N.; Toscano, M.; Alberto, M.E. Computational Mechanistic Insights on the NO Oxidation Reaction Catalyzed by Non-Heme Biomimetic Cr-N-Tetramethylated Cyclam Complexes. Int. J. Mol. Sci. 2019, 20, 3955. [Google Scholar] [CrossRef] [PubMed]
- Fortino, M.; Marino, T.; Russo, N. Theoretical Study of Silver-Ion-Mediated Base Pairs: The Case of C−Ag−C and C−Ag−A Systems. J. Phys. Chem. A 2015, 119, 5153–5157. [Google Scholar] [CrossRef] [PubMed]
- Boone, A.J.; Chang, C.H.; Greene, S.N.; Herz, T.; Richards, N.G.J. Modeling the spin-dependent properties of open-shell Fe(III)-containing systems: Towards a computational description of nitrile hydratase. Coord. Chem. Rev. 2003, 238–239, 291–314. [Google Scholar] [CrossRef]
- Barone, V.; Cossi, M. Quantum Calculation of Molecular Energies and Energy Gradients in Solution by a Conductor Solvent Model. J. Phys. Chem. A 1998, 102, 1995–2001. [Google Scholar] [CrossRef]
- Cossi, M.; Rega, N.; Scalmani, G.; Barone, V. Energies, Structures, and Electronic Properties of Molecules in Solution with the C-PCM Solvation Model. J. Comput. Chem. 2003, 24, 669–681. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of the compounds are not available from the authors. |
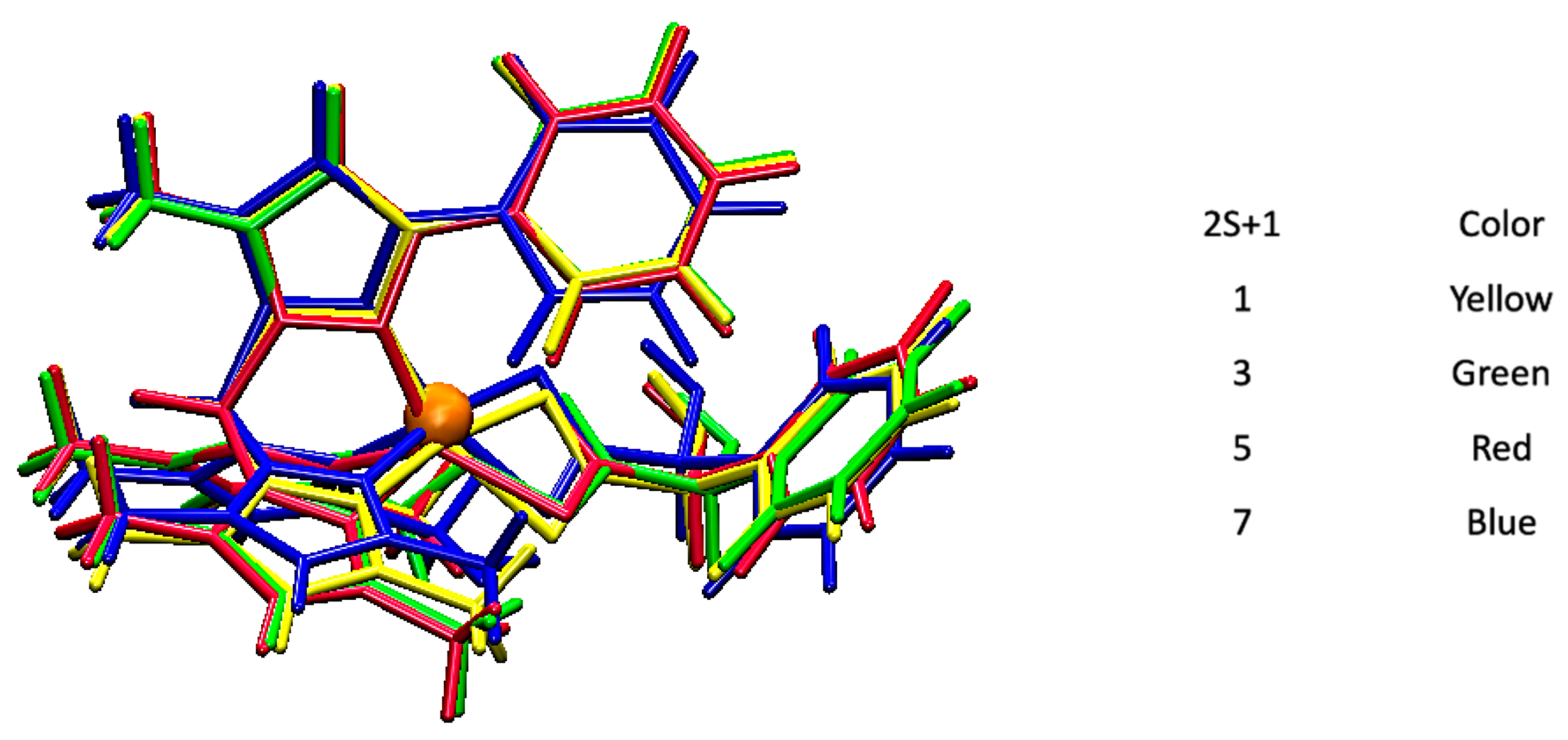
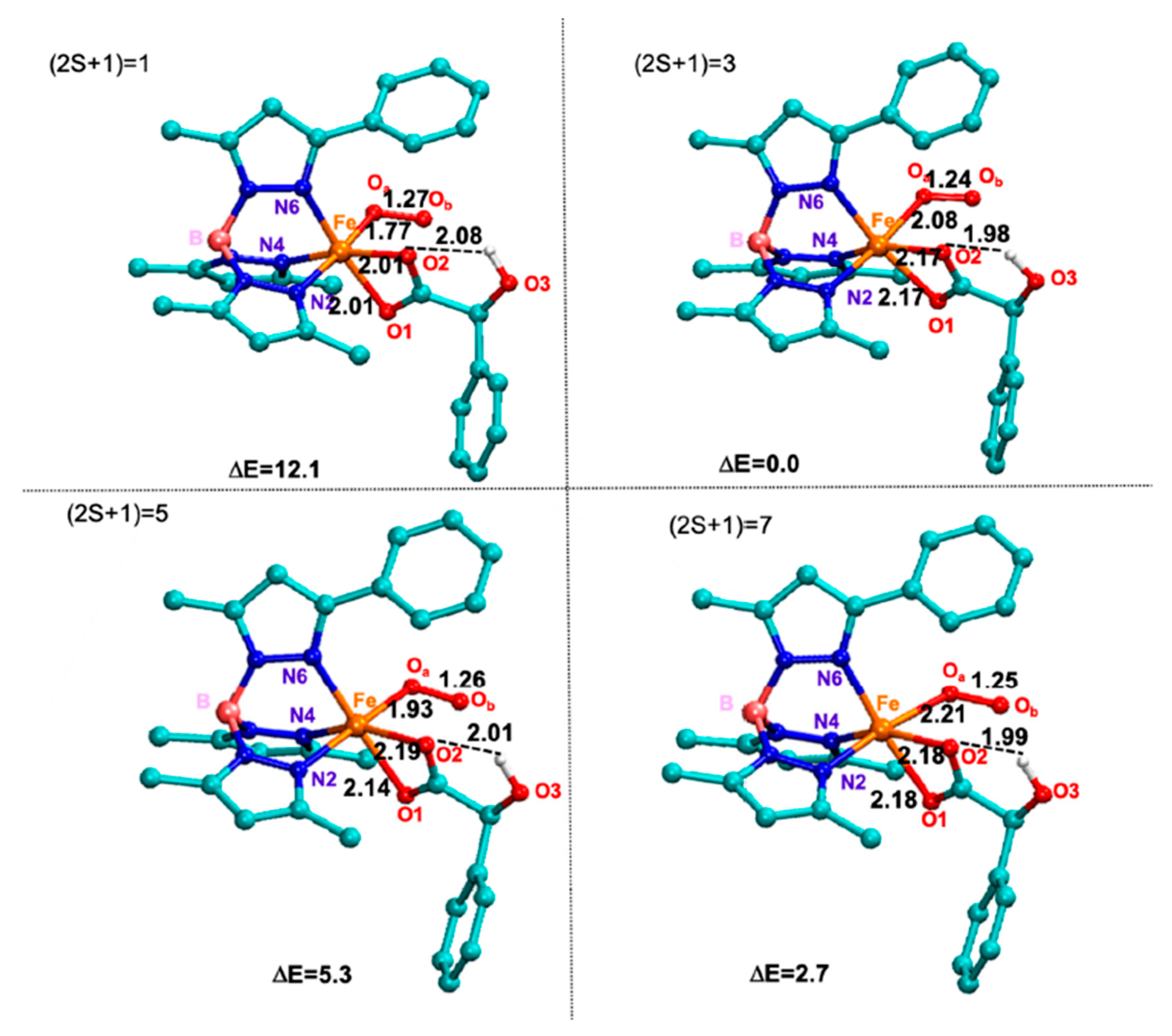
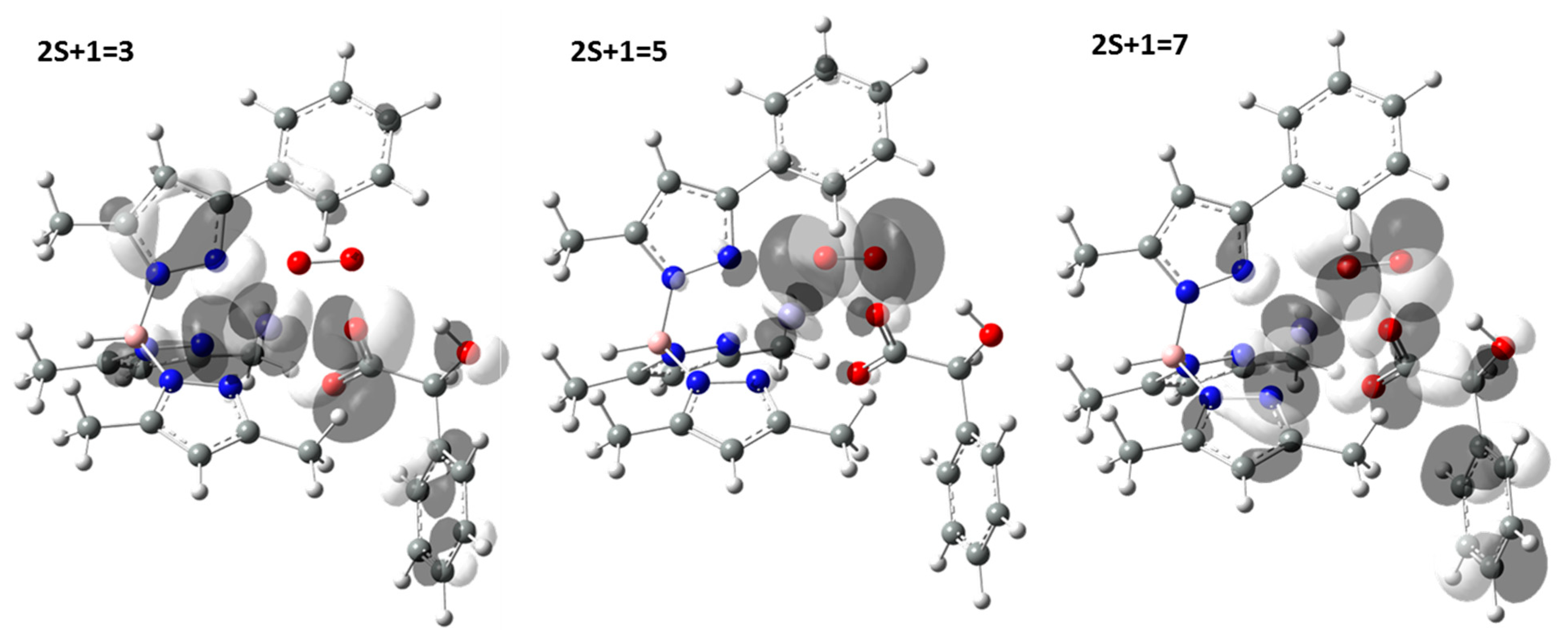
| Bond | 2S + 1 | X-Ray | |||
|---|---|---|---|---|---|
| 1 | 3 | 5 | 7 | ||
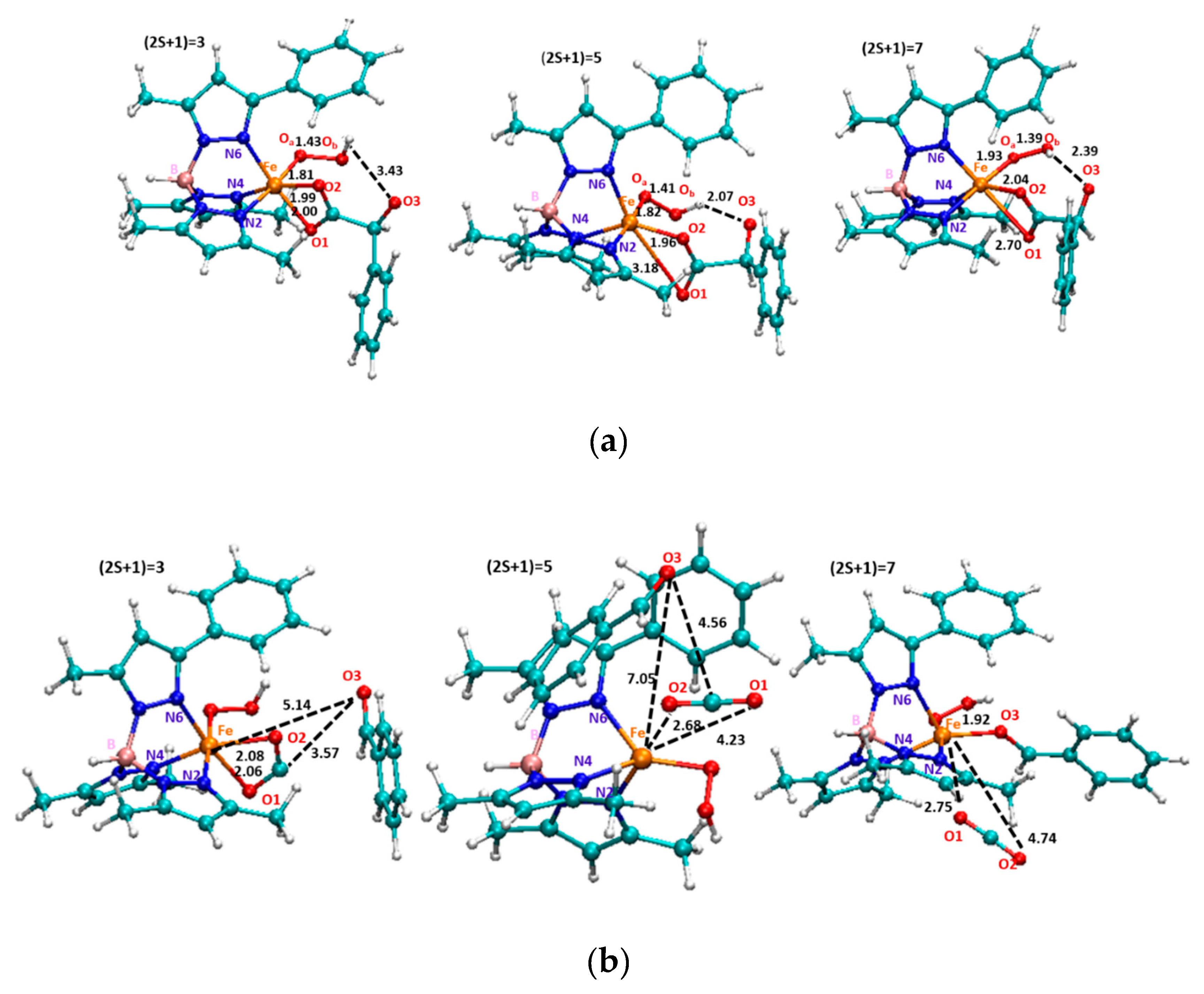
| Fe-O1 | 2.06 Å | 2.04 Å | 2.01 Å | 2.06 Å | 2.01 Å |
| Fe-O2 | 2.08 Å | 2.03 Å | 2.19 Å | 2.29 Å | 2.17 Å |
| Fe-N2 | 1.96 Å | 2.18 Å | 2.10 Å | 2.09 Å | 2.17 Å |
| Fe-N4 | 1.96 Å | 1.95 Å | 2.16 Å | 2.05 Å | 2.10 Å |
| Fe-N6 | 1.92 Å | 2.00 Å | 2.09 Å | 1.97 Å | 2.11 Å |
| DE | 17.2 | 13.1 | 0.0 | 65.1 | - |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Parise, A.; Muraca, M.C.; Russo, N.; Toscano, M.; Marino, T. The Generation of the Oxidant Agent of a Mononuclear Nonheme Fe(II) Biomimetic Complex by Oxidative Decarboxylation. A DFT Investigation. Molecules 2020, 25, 328. https://doi.org/10.3390/molecules25020328
Parise A, Muraca MC, Russo N, Toscano M, Marino T. The Generation of the Oxidant Agent of a Mononuclear Nonheme Fe(II) Biomimetic Complex by Oxidative Decarboxylation. A DFT Investigation. Molecules. 2020; 25(2):328. https://doi.org/10.3390/molecules25020328
Chicago/Turabian StyleParise, Angela, Maria Costanza Muraca, Nino Russo, Marirosa Toscano, and Tiziana Marino. 2020. "The Generation of the Oxidant Agent of a Mononuclear Nonheme Fe(II) Biomimetic Complex by Oxidative Decarboxylation. A DFT Investigation" Molecules 25, no. 2: 328. https://doi.org/10.3390/molecules25020328
APA StyleParise, A., Muraca, M. C., Russo, N., Toscano, M., & Marino, T. (2020). The Generation of the Oxidant Agent of a Mononuclear Nonheme Fe(II) Biomimetic Complex by Oxidative Decarboxylation. A DFT Investigation. Molecules, 25(2), 328. https://doi.org/10.3390/molecules25020328









