Impact of Cholesterol on the Stability of Monomeric and Dimeric Forms of the Translocator Protein TSPO: A Molecular Simulation Study
Abstract
1. Introduction
2. Results
2.1. TSPO Mouse Dimers
2.1.1. Models Refinement
2.1.2. Structural Properties of the Models
2.2. Mouse TSPO Monomers
2.2.1. Models Refinement
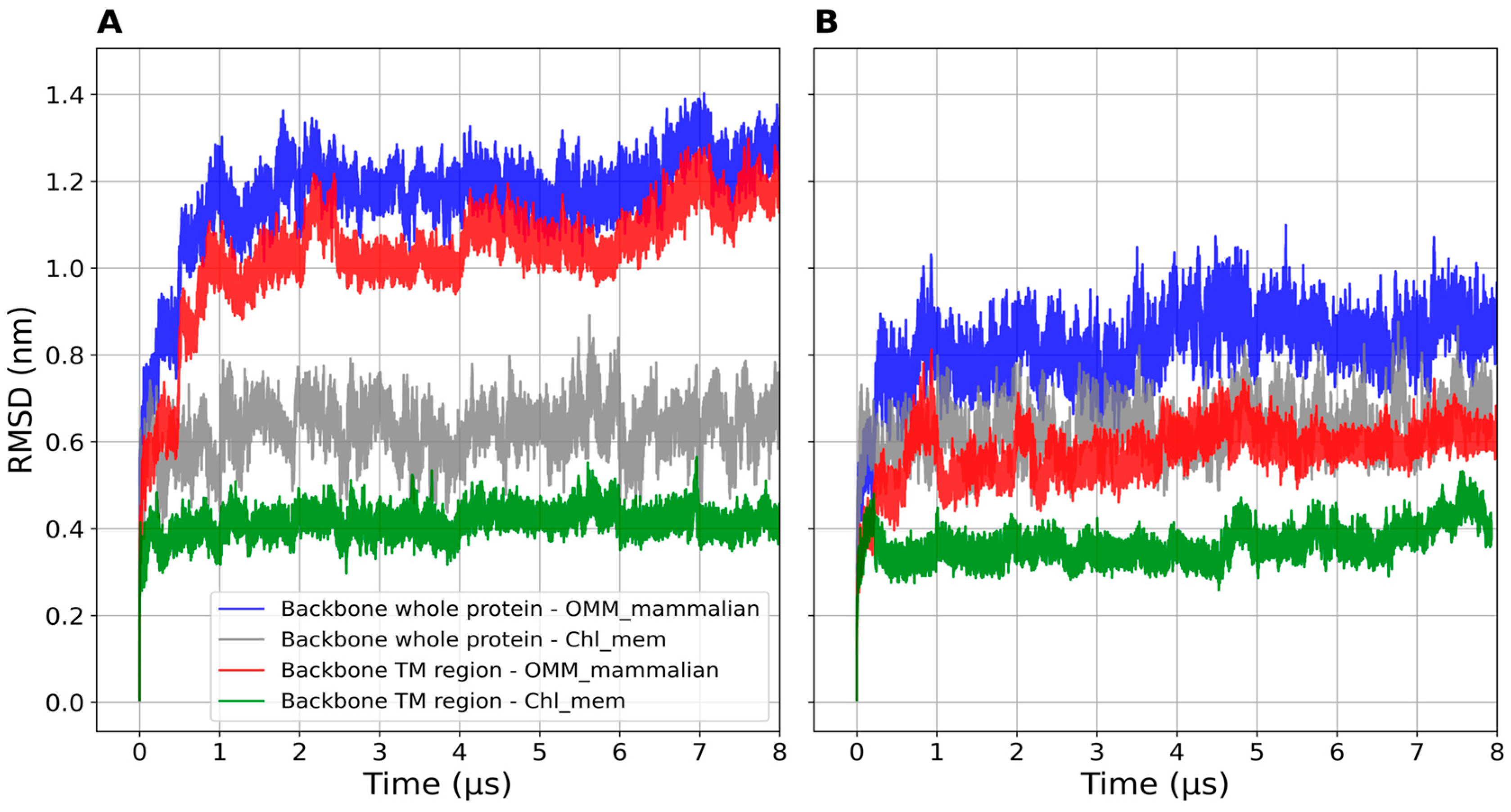
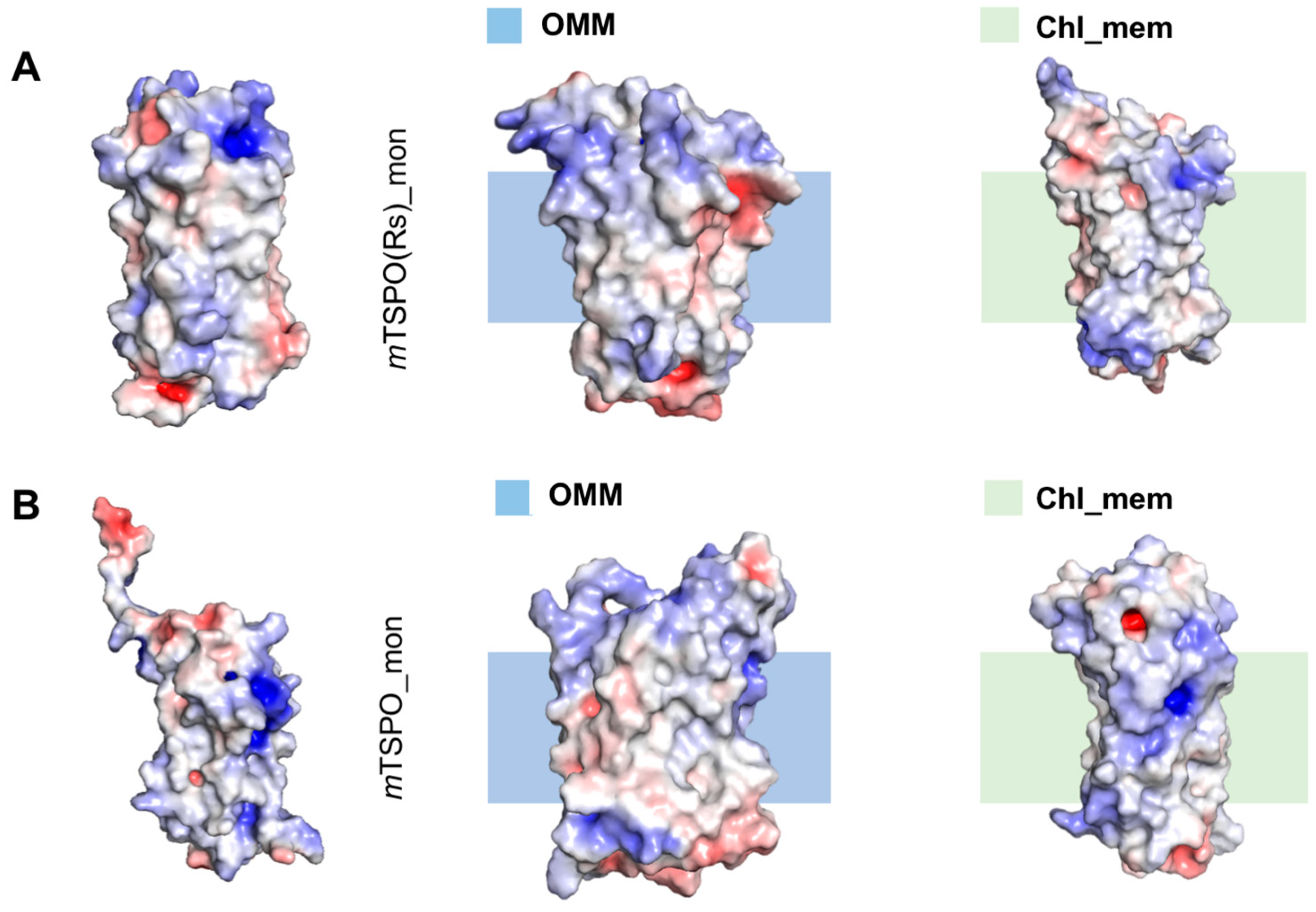
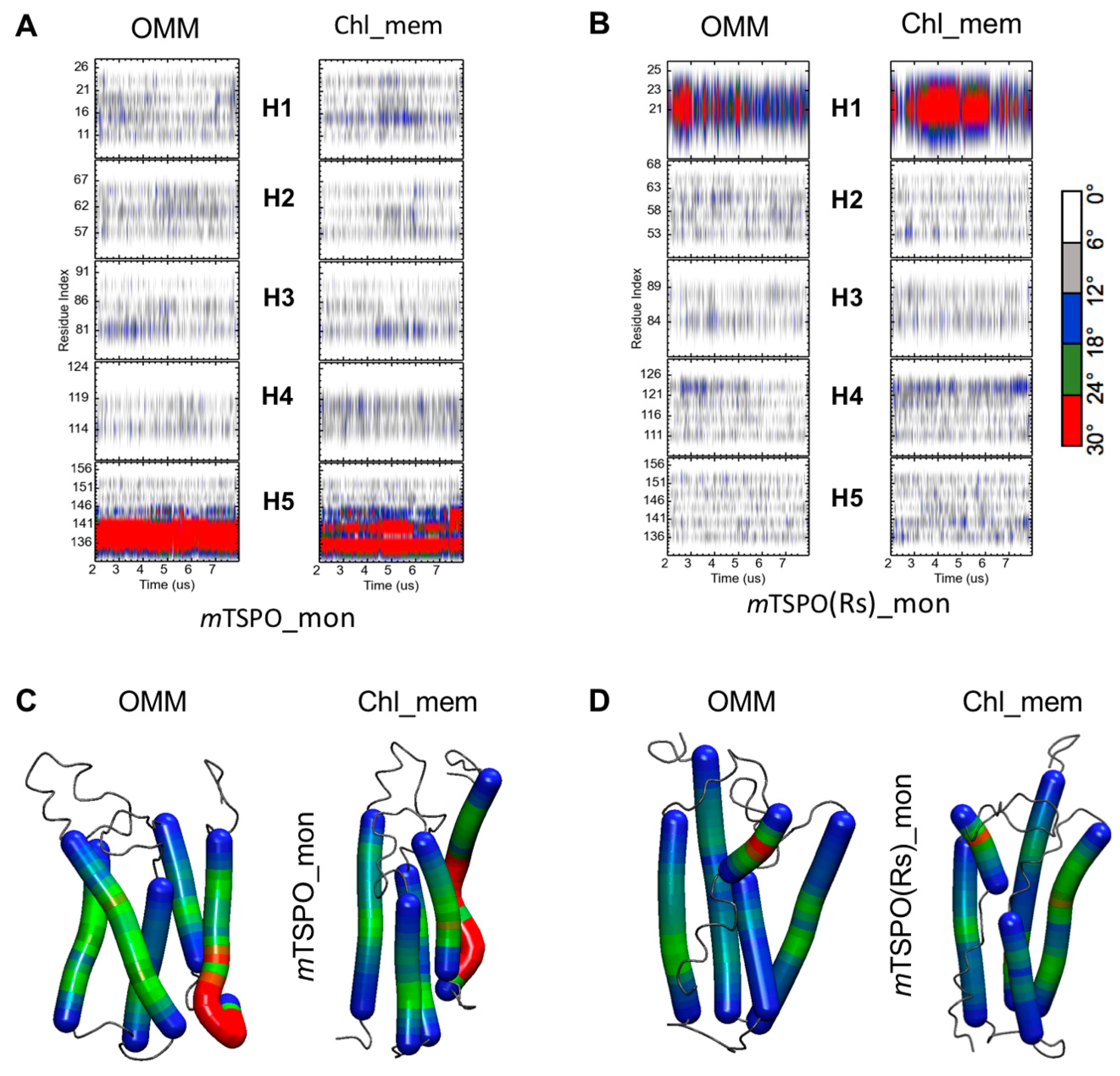
2.2.2. Structural Features
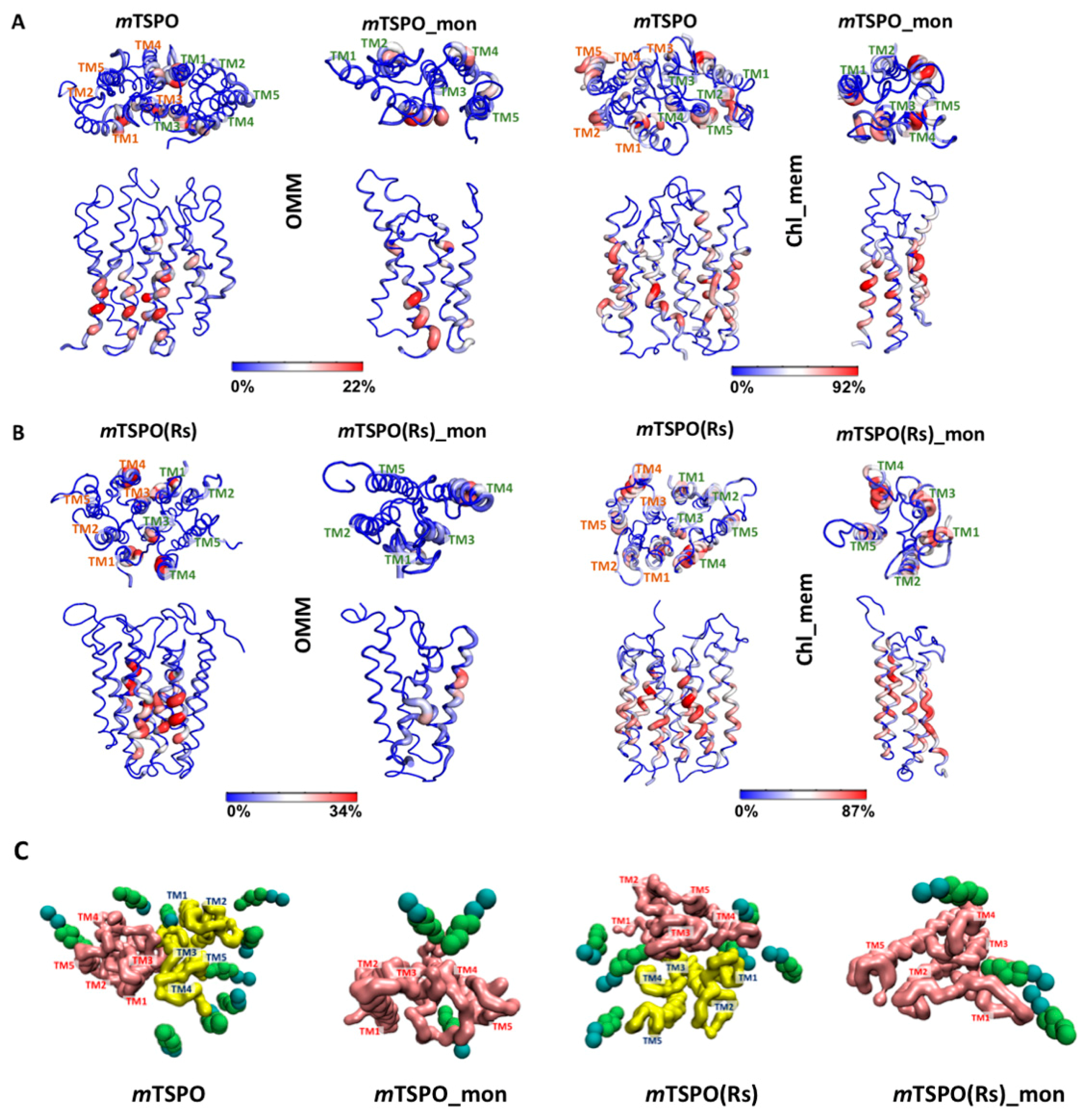
2.3. Cholesterol Occupancy
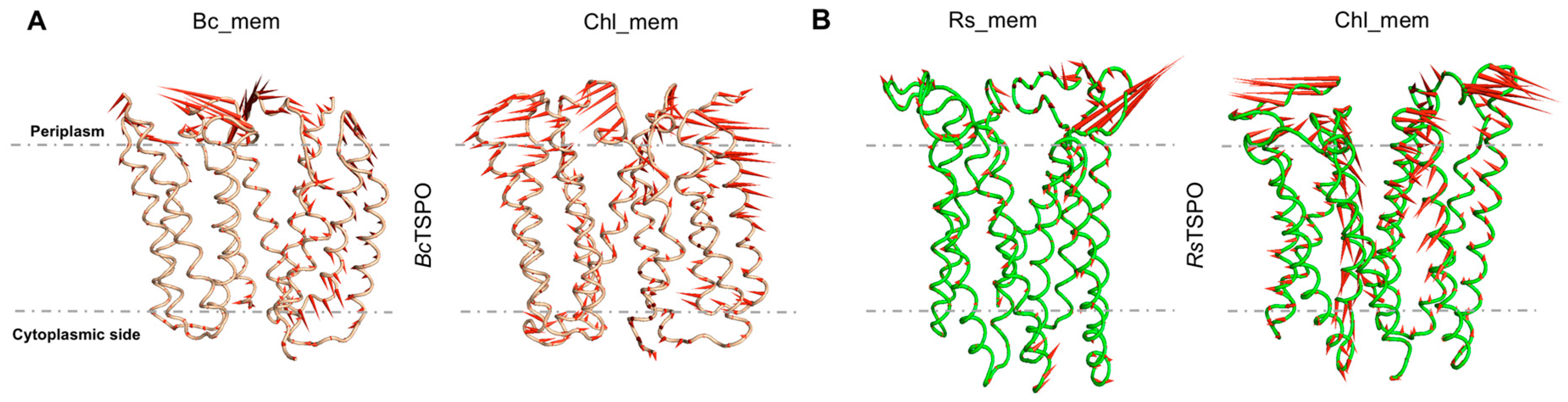
2.4. Bacterial TSPO Dimers
3. Discussion
4. Materials and Methods
4.1. System Setup of Coarse-Grained Models
4.2. Coarse-Grained Simulation Parameters
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| mTSPO | Dimeric mouse TSPO model based on NMR structure (PDBiD: 2MGY) |
| mTSPO_mon | Monomeric mouse TSPO model based on NMR structure (PDBiD: 2MGY) |
| mTSPO(Rs) | Dimeric mouse TSPO model based on RsTSPO dimer (PDBiD: 4UC1) |
| mTSPO(Rs)_mon | Monomeric mouse TSPO model based on RsTSPO dimer (PDBiD: 4UC1) |
| RsTSPO | Rhodobacter sphaeroides dimer (PDBiD: 4UC3) |
| BcTSPO | Bacillus cereus dimer (PDBiD: 4RYI) |
| OMM | Mammalian Outer Mitochondrial Membrane |
| Chl_mem | Cholesterol rich membrane |
| Rs_mem | Rs bacterial membrane |
| Bc_mem | Bc bacterial membrane |
References
- Fan, J.; Lindemann, P.; GJ Feuilloley, M.; Papadopoulos, V. Structural and functional evolution of the translocator protein (18 kDa). Curr. Mol. Med. 2012, 12, 369–386. [Google Scholar]
- Veenman, L.; Vainshtein, A.; Yasin, N.; Azrad, M.; Gavish, M. Tetrapyrroles as endogenous TSPO ligands in eukaryotes and prokaryotes: Comparisons with synthetic ligands. Int. J. Mol. Sci. 2016, 17, 880. [Google Scholar] [CrossRef] [PubMed]
- Balsemão-Pires, E.; Jaillais, Y.; Olson, B.J.; Andrade, L.R.; Umen, J.G.; Chory, J.; Sachetto-Martins, G. The Arabidopsis translocator protein (AtTSPO) is regulated at multiple levels in response to salt stress and perturbations in tetrapyrrole metabolism. BMC Plant Biol. 2011, 11, 108. [Google Scholar] [CrossRef] [PubMed]
- Yeliseev, A.A.; Krueger, K.E.; Kaplan, S. A mammalian mitochondrial drug receptor functions as a bacterial “oxygen” sensor. Proc. Natl. Acad. Sci. 1997, 94, 5101–5106. [Google Scholar] [CrossRef] [PubMed]
- Rupprecht, R.; Papadopoulos, V.; Rammes, G.; Baghai, T.C.; Fan, J.; Akula, N.; Groyer, G.; Adams, D.; Schumacher, M. Translocator protein (18 kDa)(TSPO) as a therapeutic target for neurological and psychiatric disorders. Nat. Rev. Drug Discov. 2010, 9, 971. [Google Scholar] [CrossRef]
- Li, F.; Liu, J.; Garavito, R.M.; Ferguson-Miller, S. Evolving understanding of translocator protein 18 kDa (TSPO). Pharmacol. Res. 2015, 99, 404–409. [Google Scholar] [CrossRef]
- Lee, Y.; Park, Y.; Nam, H.; Lee, J.-W.; Yu, S.-W. Translocator protein (TSPO): The new story of the old protein in neuroinflammation. BMB Rep. 2020, 53, 20. [Google Scholar] [CrossRef]
- Kim, E.-J.; Yu, S.-W. Translocator protein 18 kDa (TSPO): Old dogma, new mice, new structure, and new questions for neuroprotection. Neural Regen. Res. 2015, 10, 878. [Google Scholar]
- Li, F.; Liu, J.; Zheng, Y.; Garavito, R.M.; Ferguson-Miller, S. Crystal structures of translocator protein (TSPO) and mutant mimic of a human polymorphism. Science 2015, 347, 555–558. [Google Scholar] [CrossRef]
- Li, F.; Xia, Y.; Meiler, J.; Ferguson-Miller, S. Characterization and modeling of the oligomeric state and ligand binding behavior of purified translocator protein 18 kDa from Rhodobacter sphaeroides. Biochemistry 2013, 52, 5884–5899. [Google Scholar] [CrossRef]
- Guo, Y.; Kalathur, R.C.; Liu, Q.; Kloss, B.; Bruni, R.; Ginter, C.; Kloppmann, E.; Rost, B.; Hendrickson, W.A. Structure and activity of tryptophan-rich TSPO proteins. Science 2015, 347, 551–555. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Liu, J.; Liu, N.; Kuhn, L.A.; Garavito, R.M.; Ferguson-Miller, S. Translocator protein 18 kDa (TSPO): an old protein with new functions? Biochemistry 2016, 55, 2821–2831. [Google Scholar] [CrossRef] [PubMed]
- Delavoie, F.; Li, H.; Hardwick, M.; Robert, J.-C.; Giatzakis, C.; Péranzi, G.; Yao, Z.-X.; Maccario, J.; Lacapere, J.-J.; Papadopoulos, V. In vivo and in vitro peripheral-type benzodiazepine receptor polymerization: Functional significance in drug ligand and cholesterol binding. Biochemistry 2003, 42, 4506–4519. [Google Scholar] [CrossRef] [PubMed]
- Boujrad, N.; Vidic, B.; Papadopoulos, V. Acute action of choriogonadotropin on Leydig tumor cells: Changes in the topography of the mitochondrial peripheral-type benzodiazepine receptor. Endocrinology 1996, 137, 5727–5730. [Google Scholar] [CrossRef] [PubMed]
- Rone, M.B.; Midzak, A.S.; Issop, L.; Rammouz, G.; Jagannathan, S.; Fan, J.; Ye, X.; Blonder, J.; Veenstra, T.; Papadopoulos, V. Identification of a dynamic mitochondrial protein complex driving cholesterol import, trafficking, and metabolism to steroid hormones. Mol. Endocrinol. 2012, 26, 1868–1882. [Google Scholar] [CrossRef] [PubMed]
- Jaremko, L.; Jaremko, M.; Giller, K.; Becker, S.; Zweckstetter, M. Structure of the Mitochondrial Translocator Protein in Complex with a Diagnostic Ligand. Science 2014, 343, 1363–1366. [Google Scholar] [CrossRef]
- Corazzi, L.; Roberti, R. Lipids of brain mitochondria. In Handbook of Neurochemistry and Molecular Neurobiology; Springer: Berlin, Germany, 2010. [Google Scholar]
- Li, H.; Papadopoulos, V. Peripheral-type benzodiazepine receptor function in cholesterol transport. Identification of a putative cholesterol recognition/interaction amino acid sequence and consensus pattern. Endocrinology 1998, 139, 4991–4997. [Google Scholar] [CrossRef]
- Jaipuria, G.; Leonov, A.; Giller, K.; Vasa, S.K.; Jaremko, Á.; Jaremko, M.; Linser, R.; Becker, S.; Zweckstetter, M. Cholesterol-mediated allosteric regulation of the mitochondrial translocator protein structure. Nat. Commun. 2017, 8, 14893. [Google Scholar] [CrossRef]
- Fantini, J.; Di Scala, C.; Evans, L.S.; Williamson, P.T.; Barrantes, F.J. A mirror code for protein-cholesterol interactions in the two leaflets of biological membranes. Sci. Rep. 2016, 6, 21907. [Google Scholar] [CrossRef]
- Jamin, N.; Neumann, J.-M.; Ostuni, M.A.; Vu, T.K.N.; Yao, Z.-X.; Murail, S.; Robert, J.-C.; Giatzakis, C.; Papadopoulos, V.; Lacapere, J.-J. Characterization of the cholesterol recognition amino acid consensus sequence of the peripheral-type benzodiazepine receptor. Mol. Endocrinol. 2005, 19, 588–594. [Google Scholar] [CrossRef]
- Li, F.; Liu, J.; Valls, L.; Hiser, C.; Ferguson-Miller, S. Identification of a key cholesterol binding enhancement motif in translocator protein 18 kDa. Biochemistry 2015, 54, 1441–1443. [Google Scholar] [CrossRef] [PubMed]
- Rohmer, M.; Bouvier-Nave, P.; Ourisson, G. Distribution of Hopanoid Triterpenes in Prokaryotes. Microbiology 1984, 130, 1137–1150. [Google Scholar] [CrossRef]
- Zhang, X.; Tamot, B.; Hiser, C.; Reid, G.E.; Benning, C.; Ferguson-Miller, S. Cardiolipin deficiency in Rhodobacter sphaeroides alters the lipid profile of membranes and of crystallized cytochrome oxidase, but structure and function are maintained. Biochemistry 2011, 50, 3879–3890. [Google Scholar] [CrossRef] [PubMed]
- Epand, R.F.; Pollard, J.E.; Wright, J.O.; Savage, P.B.; Epand, R.M. Depolarization, bacterial membrane composition, and the antimicrobial action of ceragenins. Antimicrob. Agents Chemother. 2010, 54, 3708–3713. [Google Scholar] [CrossRef]
- de Jong, D.H.; Singh, G.; Bennett, W.D.; Arnarez, C.; Wassenaar, T.A.; Schäfer, L.V.; Periole, X.; Tieleman, D.P.; Marrink, S.J. Improved parameters for the martini coarse-grained protein force field. J. Chem. Theory Comput. 2013, 9, 687–697. [Google Scholar] [CrossRef]
- Monticelli, L.; Kandasamy, S.K.; Periole, X.; Larson, R.G.; Tieleman, D.P.; Marrink, S.-J. The MARTINI coarse-grained force field: Extension to proteins. J. Chem. Theory Comput. 2008, 4, 819–834. [Google Scholar] [CrossRef]
- Zeng, J.; Guareschi, R.; Damre, M.; Cao, R.; Kless, A.; Neumaier, B.; Bauer, A.; Giorgetti, A.; Carloni, P.; Rossetti, G. Structural prediction of the dimeric form of the mammalian translocator membrane protein TSPO: a key target for brain diagnostics. Int. J. Mol. Sci. 2018, 19, 2588. [Google Scholar] [CrossRef]
- Xia, Y.; Ledwitch, K.; Kuenze, G.; Duran, A.; Li, J.; Sanders, C.R.; Manning, C.; Meiler, J. A unified structural model of the mammalian translocator protein (TSPO). J. Biomol. NMR 2019, 73, 347–364. [Google Scholar] [CrossRef]
- Rao, R.M.; Diharce, J.; Dugué, B.; Ostuni, M.A.; Cadet, F.; Etchebest, C. Versatile dimerisation process of translocator protein (TSPO) revealed by an extensive sampling based on a coarse-grained dynamics study. J. Chem. Inf. Modeling 2020. [Google Scholar] [CrossRef]
- Wassenaar, T.A.; Pluhackova, K.; Böckmann, R.A.; Marrink, S.J.; Tieleman, D.P. Going backward: A flexible geometric approach to reverse transformation from coarse grained to atomistic models. J. Chem. Theory Comput. 2014, 10, 676–690. [Google Scholar] [CrossRef]
- Dahl, A.C.E.; Chavent, M.; Sansom, M.S. Bendix: Intuitive helix geometry analysis and abstraction. Bioinformatics 2012, 28, 2193–2194. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Wolf, R.M.; Caldwell, J.W.; Kollman, P.A.; Case, D.A. Development and testing of a general amber force field. J. Comput. Chem. 2004, 25, 1157–1174. [Google Scholar] [CrossRef] [PubMed]
- Lin, T.-Y.; Gross, W.S.; Auer, G.K.; Weibel, D.B. Cardiolipin alters Rhodobacter sphaeroides cell shape by affecting peptidoglycan precursor biosynthesis. Mbio 2019, 10. [Google Scholar] [CrossRef] [PubMed]
- Leneveu-Jenvrin, C.; Connil, N.; Bouffartigues, E.; Papadopoulos, V.; Feuilloley, M.G.; Chevalier, S. Structure-to-function relationships of bacterial translocator protein (TSPO): A focus on Pseudomonas. Front. Microbiol. 2014, 5, 631. [Google Scholar] [CrossRef]
- Karshikoff, A.; Nilsson, L.; Ladenstein, R. Rigidity versus flexibility: The dilemma of understanding protein thermal stability. FEBS J. 2015, 282, 3899–3917. [Google Scholar] [CrossRef]
- Selvaraj, V.; Stocco, D.M.; Tu, L.N. Minireview: Translocator protein (TSPO) and steroidogenesis: A reappraisal. Mol. Endocrinol. 2015, 29, 490–501. [Google Scholar] [CrossRef]
- Šali, A.; Blundell, T.L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 1993, 234, 779–815. [Google Scholar] [CrossRef]
- Shen, M.y.; Sali, A. Statistical potential for assessment and prediction of protein structures. Protein Sci. 2006, 15, 2507–2524. [Google Scholar] [CrossRef]
- John, B.; Sali, A. Comparative protein structure modeling by iterative alignment, model building and model assessment. Nucleic Acids Res. 2003, 31, 3982–3992. [Google Scholar] [CrossRef]
- Harayama, T.; Riezman, H. Understanding the diversity of membrane lipid composition. Nat. Ecol. Evol. 2018, 1–18. [Google Scholar] [CrossRef]
- Hicks, A.M.; DeLong, C.J.; Thomas, M.J.; Samuel, M.; Cui, Z. Unique molecular signatures of glycerophospholipid species in different rat tissues analyzed by tandem mass spectrometry. Biochim. Et Biophys. Acta (BBA) Mol. Cell Biol. Lipids 2006, 1761, 1022–1029. [Google Scholar] [CrossRef] [PubMed]
- Kiebish, M.A.; Han, X.; Seyfried, T.N. Examination of the brain mitochondrial lipidome using shotgun lipidomics. In Lipidomics; Springer: Berlin, Germany, 2009; pp. 3–18. [Google Scholar]
- Olofsson, G.; Sparr, E. Ionization constants pK a of cardiolipin. PLoS ONE 2013, 8, e73040. [Google Scholar] [CrossRef] [PubMed]
- Van Der Spoel, D.; Lindahl, E.; Hess, B.; Groenhof, G.; Mark, A.E.; Berendsen, H.J. GROMACS: Fast, flexible, and free. J. Comput. Chem. 2005, 26, 1701–1718. [Google Scholar] [CrossRef]
- Bussi, G.; Donadio, D.; Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 2007, 126, 014101. [Google Scholar] [CrossRef]
- Martoňák, R.; Laio, A.; Parrinello, M. Predicting crystal structures: The Parrinello-Rahman method revisited. Phys. Rev. Lett. 2003, 90, 075503. [Google Scholar] [CrossRef]
- Van Aalten, D.M.; De Groot, B.L.; Findlay, J.B.; Berendsen, H.J.; Amadei, A. A comparison of techniques for calculating protein essential dynamics. J. Comput. Chem. 1997, 18, 169–181. [Google Scholar] [CrossRef]
- DeLano, W.L. The PyMOL Molecular Graphics System; Schrödinger: New York, NY, USA, 2002. [Google Scholar]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- Lerner, M.; Carlson, H. APBS Plugin for PyMOL; University of Michigan: Ann Arbor, MI, USA, 2006. [Google Scholar]
- Song, W.; Duncan, A.; Corey, R.; Ansell, B. PyLipID. Available online: https://github.com/wlsong/PyLipID (accessed on 17 September 2020).
- Zhao, A.H.; Tu, L.N.; Mukai, C.; Sirivelu, M.P.; Pillai, V.V.; Morohaku, K.; Cohen, R.; Selvaraj, V. Mitochondrial Translocator Protein (TSPO) Function Is Not Essential for Heme Biosynthesis. J. Biol. Chem. 2016, 291, 1591–1603. [Google Scholar] [CrossRef]
- Papadopoulos, V.; Fan, J.; Zirkin, B. Translocator protein (18 kDa): An update on its function in steroidogenesis. J. Neuroendocrinol. 2017, 30, e12500. [Google Scholar] [CrossRef]
- Taylor, E.B. Functional Properties of the Mitochondrial Carrier System. Trends Cell Biol. 2017, 27, 633–644. [Google Scholar] [CrossRef]
- Da Pozzo, E.; Costa, B.; Martini, C. Translocator protein (TSPO) and neurosteroids: Implications in psychiatric disorders. Curr. Mol. Med. 2012, 12, 426–442. [Google Scholar] [CrossRef]
Sample Availability: Models of the systems reported are available from the authors. |

| Region | Inferred by Experiment | mTSPO | mTSPO(Rs) |
|---|---|---|---|
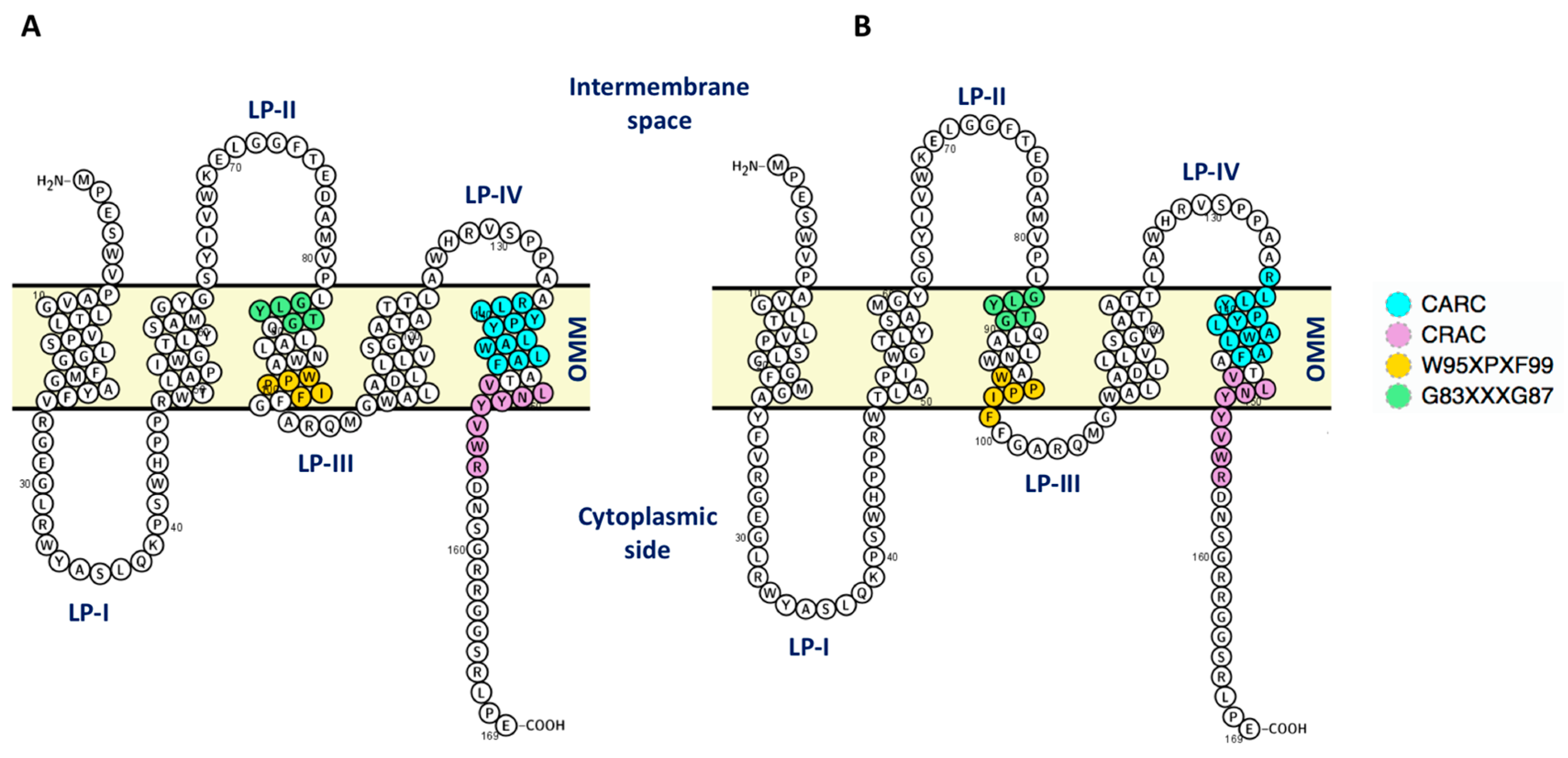
| Dimer interface | V80, G83, Q88, N92, W93, W95, I98, F100, G101, A102, D111, V118 | Initial structures | |
| F74, T75, E76, D77, M79, V80, P81, G83, L84, T86, G87, Q88, A90, L91 | V6, P7, G10, L11, L13, V14, L17, G18, F20, M21, Y24 V26, R27, M79, V80, L82, G83, L84, Y85, T86, G87, A90, L91, W93, A94, P97, I98, A102, Q104, W107, A108, A110, D111, L114, V118, A121, A125 | ||
| Structures after equilibration in outer mitochondrial membrane (OMM) | |||
| M1, P2, S4, W5, P7, A8, G10, L11, T12, L13, V14, P15, L17, G18, M21, G22, F25, V26, E29, Y34, L37, K39, P40, R46, V67, W68, E70, L71, D77, A78, M79, V80, P81, L82, G83, Y85, T86, G87, Q88, L89, A90, L91, N92, W93, A94, W95, P96, P97, I98, F99, F100, G101, A102, R103, Q104, M105, G106, W107, A108, L109, A110, D111, L113, L114, V115, S116, G117, V118, A121, T122, L124, A125, W126, R128, V129, S130, P131, N151, R166, L167, A168, E169 | S4, V6, P7, V9, G10, L11, T12, L13, V14, P15, L17, G18, F20, M21, G22, Y24, F25, V26, R27, G28, E29, G30, L31, R32, W33, W47, W53, Y57, M60, S64, Y65, V67, W68, K69, E70, L71, G72, G73, F74, T75, E76, A78, M79, V80, P81, L82, G83, L84, Y85, T86, G87, L89, A90, L91, W93, A94, W95, P96, P97, I98, F100, G101, R103, Q104, M105, G106, W107, A110, D111, L114, V118, A121, W126 | ||
| System | RMSD (nm) | Radius of Gyration (nm) |
|---|---|---|
| mTSPO_mon, OMM | 1.2 ± 0.05 | 1.7 ± 0.03 |
| mTSPO_mon, chl_mem | 0.6 ± 0.05 | 1.8 ± 0.03 |
| mTSPO(Rs)_mon, OMM | 0.8 ± 0.05 | 1.7 ± 0.02 |
| mTSPO(Rs)_mon, chl_mem | 0.6 ± 0.05 | 1.7 ± 0.02 |
| mTSPO, chain A, OMM | 1.1 ± 0.05 | 1.7 ± 0.02 |
| mTSPO, chain B, OMM | 1.0 ± 0.05 | 1.8 ± 0.02 |
| mTSPO, chain A, chl_mem | 1.2 ± 0.04 | 1.7 ± 0.01 |
| mTSPO, chain B, chl_mem | 1.1 ± 0.06 | 1.8 ± 0.02 |
| mTSPO(Rs), chain A, OMM | 0.8 ± 0.05 | 1.8 ± 0.02 |
| mTSPO(Rs), chain B, OMM | 0.6 ± 0.05 | 1.7 ± 0.01 |
| mTSPO(Rs), chain A, chl_mem | 0.6 ± 0.04 | 1.7 ± 0.02 |
| mTSPO(Rs), chain B, chl_mem | 0.5 ± 0.05 | 1.7 ± 0.01 |
| BcTSPO, chain A, Bc_mem | 0.6 ± 0.01 | 1.6 ± 0.01 |
| BcTSPO, chain B, Bc_mem | 0.6 ± 0.02 | 1.6 ± 0.01 |
| BcTSPO, chain A, chl_mem | 0.5 ± 0.03 | 1.6 ± 0.01 |
| BcTSPO, chain B, chl_mem | 0.6 ± 0.02 | 1.6 ± 0.01 |
| RsTSPO, chain A, Rs_mem | 0.7 ± 0.04 | 1.7 ± 0.01 |
| RsTSPO, chain B, Rs_mem | 0.5 ± 0.02 | 1.7 ± 0.01 |
| RsTSPO_chain A, chl_mem | 0.6 ± 0.04 | 1.7 ± 0.01 |
| RsTSPO_chain B, chl_mem | 0.6 ± 0.03 | 1.7 ± 0.02 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Si Chaib, Z.; Marchetto, A.; Dishnica, K.; Carloni, P.; Giorgetti, A.; Rossetti, G. Impact of Cholesterol on the Stability of Monomeric and Dimeric Forms of the Translocator Protein TSPO: A Molecular Simulation Study. Molecules 2020, 25, 4299. https://doi.org/10.3390/molecules25184299
Si Chaib Z, Marchetto A, Dishnica K, Carloni P, Giorgetti A, Rossetti G. Impact of Cholesterol on the Stability of Monomeric and Dimeric Forms of the Translocator Protein TSPO: A Molecular Simulation Study. Molecules. 2020; 25(18):4299. https://doi.org/10.3390/molecules25184299
Chicago/Turabian StyleSi Chaib, Zeineb, Alessandro Marchetto, Klevia Dishnica, Paolo Carloni, Alejandro Giorgetti, and Giulia Rossetti. 2020. "Impact of Cholesterol on the Stability of Monomeric and Dimeric Forms of the Translocator Protein TSPO: A Molecular Simulation Study" Molecules 25, no. 18: 4299. https://doi.org/10.3390/molecules25184299
APA StyleSi Chaib, Z., Marchetto, A., Dishnica, K., Carloni, P., Giorgetti, A., & Rossetti, G. (2020). Impact of Cholesterol on the Stability of Monomeric and Dimeric Forms of the Translocator Protein TSPO: A Molecular Simulation Study. Molecules, 25(18), 4299. https://doi.org/10.3390/molecules25184299






