LC-MS-Based Metabolomics for the Chemosystematics of Kenyan Dodonaea viscosa Jacq (Sapindaceae) Populations
Abstract
1. Introduction
2. Results and Discussion
2.1. Identification of Compounds
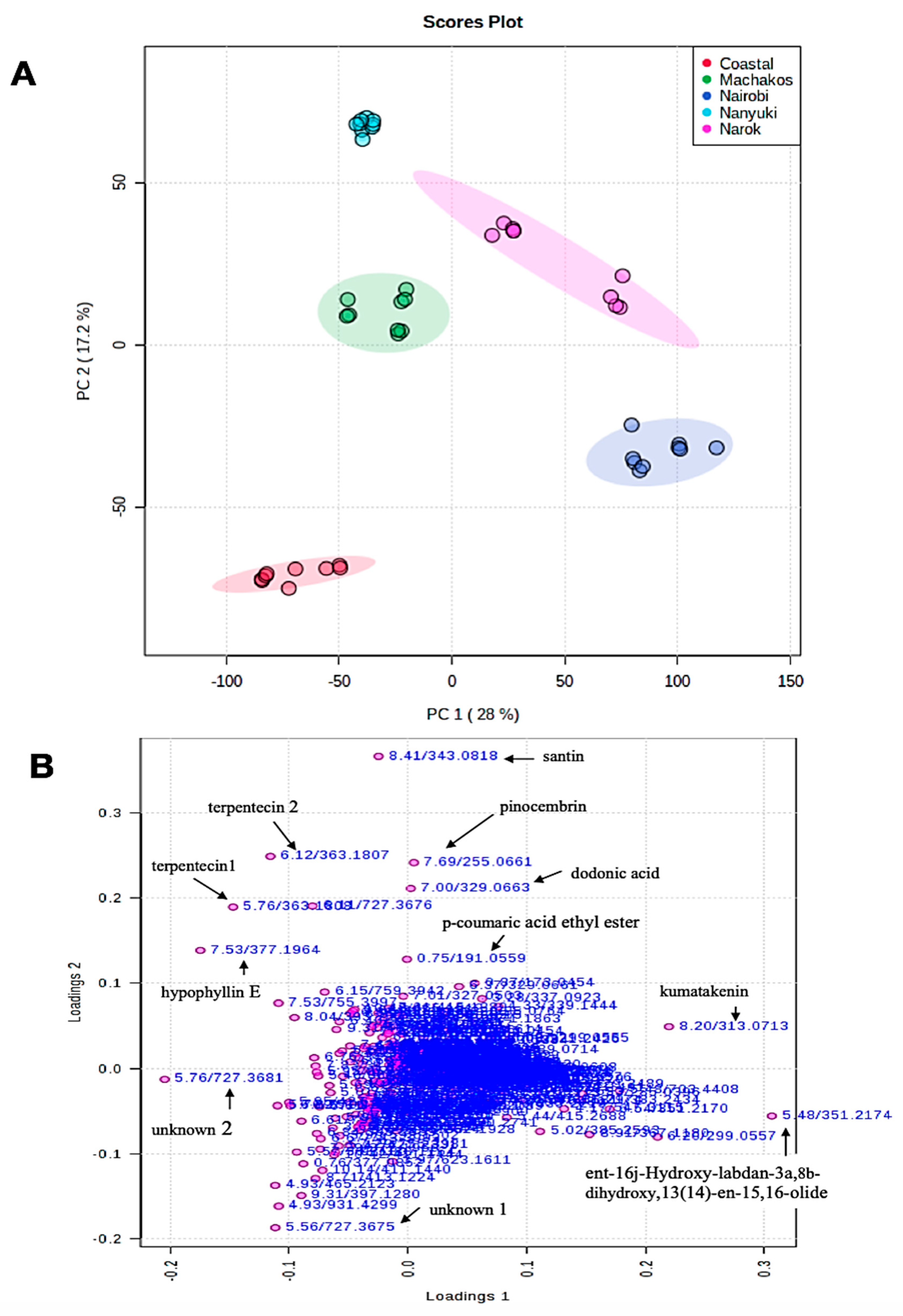
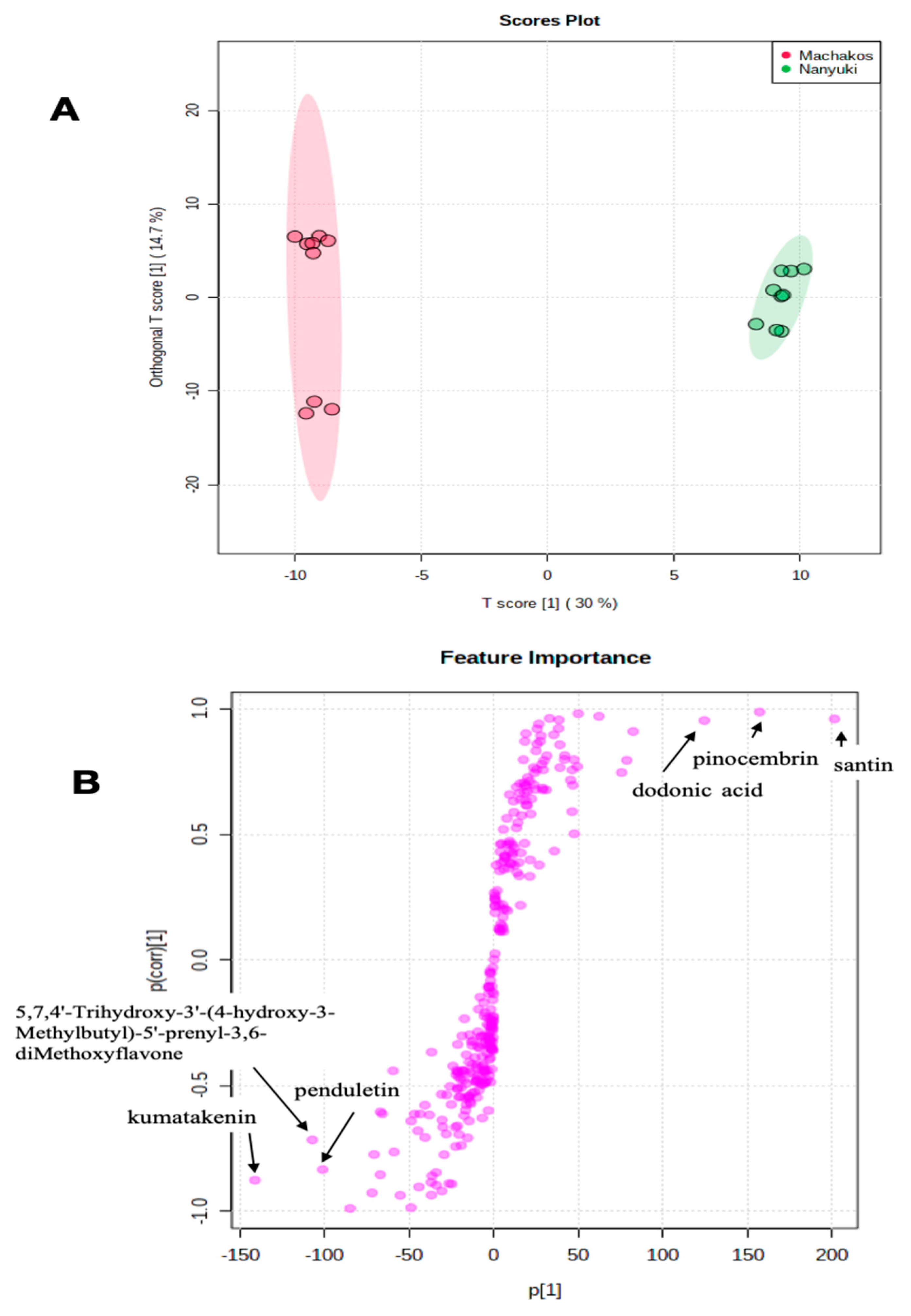
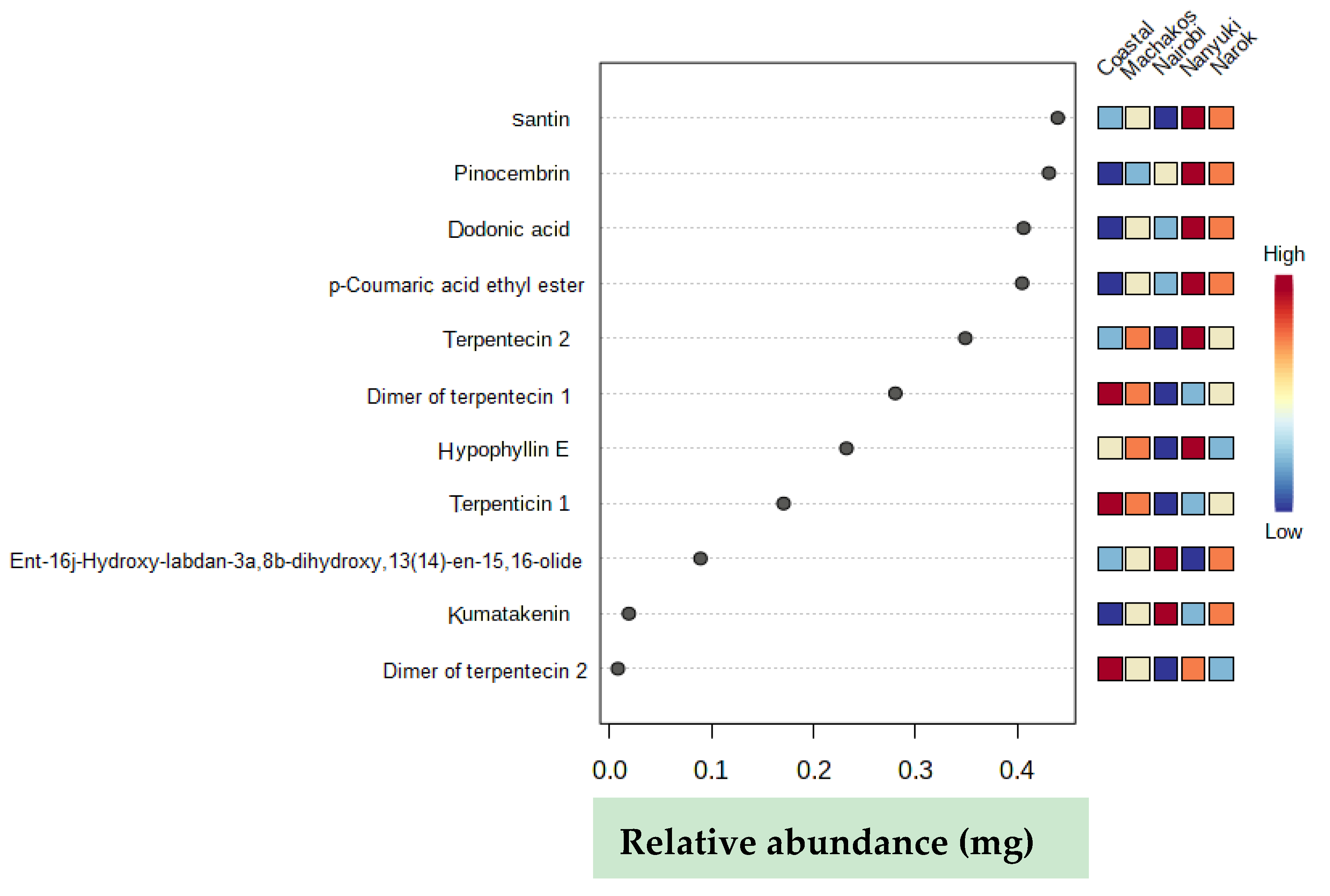
2.2. Chemotype Variation
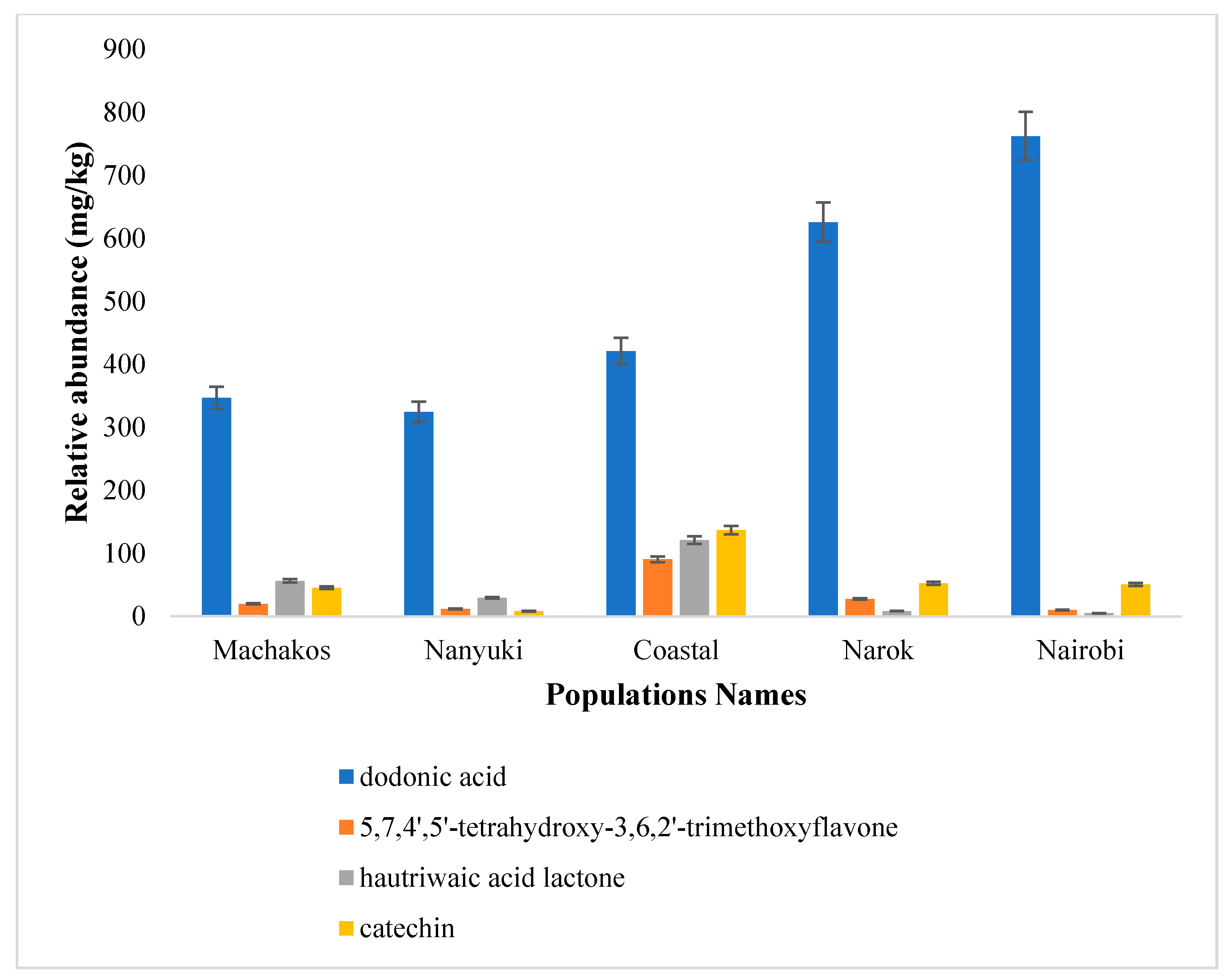
2.3. Secondary Metabolites Isolated from Gazi Coastal Population of Dodonaea viscosa
2.4. Antimicrobial Activity of the Isolated Compounds and the Extracts
3. Materials and Methods
3.1. Collection of Plant Materials
3.2. Extraction for Metabolite Profiling
Standards
3.3. Liquid Chromatography Mass Spectrometry (LC-MS) Analysis
3.4. Extraction and Isolation of Compounds: Gazi Coastal Population
3.5. Characterization of Pure Compounds
3.6. Antimicrobial Activity of the Isolated Compounds
3.7. Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Makunga, N.P.; Engelbrecht, A.M. Method and Composition for Treating Breast Cancer. Google Patents 0,188,689, 29 January 2019. [Google Scholar]
- Anandan, M.; Prabu, H.G. Dodonaea viscosa leaf extract assisted synthesis of gold nanoparticles: Characterization and cytotoxicity against A549 NSCLC cancer cells. J. Inorg. Organomet Polym. Mater. 2018, 28, 932–941. [Google Scholar] [CrossRef]
- Venkatesh, S.; Reddy, Y.S.R.; Ramesh, M.; Swamy, M.M.; Mahadevan, N.; Suresh, B. Pharmacognostical studies on Dodonaea viscosa leaves. African J. Pharm. Pharmacol. 2008, 2, 83–88. [Google Scholar]
- Tauchen, J.; Doskocil, I.; Caffi, C.; Lulekal, E.; Marsik, P.; Havlik, J.; Van Damme, P.; Kokoska, L. In vitro antioxidant and anti-proliferative activity of Ethiopian medicinal plant extracts. Ind. Crops Prod. 2015, 74, 671–679. [Google Scholar] [CrossRef]
- Rojas, A.; Cruz, S.; Ponce-Monter, H.; Mata, R. Smooth muscle relaxing compounds from Dodonaea viscosa. Planta. Med. 1996, 62, 154–159. [Google Scholar] [CrossRef]
- Getie, M.; Gebre-Mariam, T.; Rietz, R.; Höhne, C.; Huschka, C.; Schmidtke, M.; Abate, A.; Neubert, R.H. Evaluation of the anti-microbial and anti-inflammatory activities of the medicinal plants Dodonaea viscosa, Rumex nervosus and Rumex abyssinicus. Fitoterapia 2003, 74, 139–143. [Google Scholar] [CrossRef]
- Khalil, N.M.; Sperotto, J.S.; Manfron, M.P. Antiinflammatory activity and acute toxicity of Dodonaea viscosa. Fitoterapia 2006, 77, 478–480. [Google Scholar] [CrossRef]
- Al-Snafi, A.E. A review on Dodonaea viscosa: A potential medicinal plant. IOSR J. Pharm. 2017, 7, 10–21. [Google Scholar] [CrossRef]
- Rani, M.S.; Pippalla, R.S.; Mohan, K. Dodonaea viscosa Linn.—An overview. Asian J. Pharm. Res. Heal. Care. 2009, 1, 97–112. [Google Scholar]
- Mostafa, A.E.; Atef, A.; Mohammad, A.E.; Jacob, M.; Cutler, S.J.; Ross, S.A. New secondary metabolites from Dodonaea viscosa. Phytochem. Lett. 2014, 8, 10–15. [Google Scholar] [CrossRef]
- Muhammad, A.; Tel-Cayan, G.; Öztürk, M.; Nadeem, S.; Duru, M.E.; Anis, I.; Ng, S.W.; Shah, M.R. Biologically active flavonoids from Dodonaea viscosa and their structure-activity relationships. Ind. Crops Prod. 2015, 78, 66–72. [Google Scholar] [CrossRef]
- Harrington, M.G.; Gadek, P.A. A species well travelled–the Dodonaea viscosa (Sapindaceae) complex based on phylogenetic analyses of nuclear ribosomal ITS and ETSf sequences. J. Biogeogr. 2009, 36, 2313–2323. [Google Scholar] [CrossRef]
- Christmas, M.J.; Biffin, E.; Lowe, A.J. Transcriptome sequencing, annotation and polymorphism detection in the hop bush, Dodonaea viscosa. BMC Genom. 2015, 16, 803. [Google Scholar] [CrossRef] [PubMed]
- Beentje, H.; Adamson, J.; Bhanderi, D. Kenya Trees, Shrubs, and Lianas; National Museums of Kenya: Nairobi, Kenya, 1994. [Google Scholar]
- Kindt, R.; Groen, T.; de Leeuw, J.; Lilleso, J.P. Distribution and ecology of trees in Eastern Africa drylands. In Treesilience: An Assessment of the Resilience Provided by Trees in the Drylands of Eastern Africa; de Leeuw, J., Ed.; The World Agroforestry Centre (ICRAF): Nairobi, Kenya, 2014; pp. 24–34. [Google Scholar]
- Omosa, L.K.; Midiwo, J.O.; Derese, S.; Yenesew, A.; Peter, M.G.; Heydenreich, M. neo-Clerodane diterpenoids from the leaf exudate of Dodonaea angustifolia. Phytochem. Lett. 2010, 3, 217–220. [Google Scholar] [CrossRef]
- Omosa, L.K.; Amugune, B.; Ndunda, B.; Milugo, T.K.; Heydenreich, M.; Yenesew, A.; Midiwo, J.O. Antimicrobial flavonoids and diterpenoids from Dodonaea angustifolia. South. Afr. J. Bot. 2014, 91, 58–62. [Google Scholar] [CrossRef]
- Martucci, M.E.P.; De Vos, R.C.H.; Carollo, C.A.; Gobbo-Neto, L. Metabolomics as a potential chemotaxonomical tool: Application in the genus Vernonia Schreb. PLoS ONE 2014, 9, e93149. [Google Scholar] [CrossRef]
- Sandasi, M.; Kamatou, G.P.P.; Viljoen, A.M. An untargeted metabolomic approach in the chemotaxonomic assessment of two Salvia species as a potential source of α-bisabolol. Phytochemistry 2012, 84, 94–101. [Google Scholar] [CrossRef]
- Padilla-González, G.F.; Diazgranados, M.; Da Costa, F.B. Biogeography shaped the metabolome of the genus Espeletia: A phytochemical perspective on an Andean adaptive radiation. Sci. Rep. 2017, 7, 8835. [Google Scholar] [CrossRef]
- Chagas-Paula, D.A.; Oliveira, T.B.; Zhang, T.; Edrada-Ebel, R.; Da Costa, F.B. Prediction of anti-inflammatory plants and discovery of their biomarkers by machine learning algorithms and metabolomic studies. Planta. Med. 2015, 81, 450–458. [Google Scholar] [CrossRef]
- Cox, D.G.; Oh, J.; Keasling, A.; Colson, K.L.; Hamann, M.T. The utility of metabolomics in natural product and biomarker characterization. Biochim. Biophys. Acta. (BBA)-Gen. Subj. 2014, 1840, 3460–3474. [Google Scholar] [CrossRef]
- Albrecht, C.F.; Stander, M.A.; Grobbelaar, M.C.; Colling, J.; Kossmann, J.; Hills, P.N.; Makunga, N.P. LC–MS-based metabolomics assists with quality assessment and traceability of wild and cultivated plants of Sutherlandia frutescens (Fabaceae). South Afr. J. Bot. 2012, 82, 33–45. [Google Scholar] [CrossRef]
- Yuliana, N.D.; Jahangir, M.; Verpoorte, R.; Choi, Y.H. Metabolomics for the rapid dereplication of bioactive compounds from natural sources. Phytochem. Rev. 2013, 12, 293–304. [Google Scholar] [CrossRef]
- Niu, H.M.; Zeng, D.Q.; Long, C.L.; Peng, Y.H.; Wang, Y.H.; Luo, J.F.; Wang, H.S.; Shi, Y.N.; Tang, G.H.; Zhao, F.W. Clerodane diterpenoids and prenylated flavonoids from Dodonaea viscosa. J. Asian Nat. Prod. Res. 2010, 12, 7–14. [Google Scholar] [CrossRef] [PubMed]
- Dreyer, D.L. Kaempferol methyl ethers from flowers of Dodonaea viscosa. Rev. Lat. Am. Quim. 1978, 9, 97–98. [Google Scholar]
- Hamadi, S.S. Chemical study of Dodonaea viscosa planting in Iraq. Int. J. Adv. Chem. Eng. Biol. Sci. 2017, 4, 121–125. [Google Scholar]
- de Oliveira, S.Q.; de Almeida, M.T.R.; Maraslis, F.; Silva, I.T.; Sincero, T.C.M.; Palermo, J.A.; Cabrera, G.M.; Caro, M.S.; Simões, C.M.; Schenkel, E.P. Isolation of three new ent-labdane diterpenes from Dodonaea viscosa Jacquin (Sapindaceae): Preliminary evaluation of antiherpes activity. Phytochem. Lett. 2012, 5, 500–505. [Google Scholar] [CrossRef]
- Manjulatha, K. A Comparative Study of Different Parts of Flavonoid Rich Plant, Dodonaea Viscosa for Antimicrobial and Antioxidant Potential; Lambert Academic Publishing: Rīgā, Latvia, 2012; Available online: https//wwwlappublishingcom/catalog/details//store/gb/book/978-3-659-28793-0/Dodonaea-viscosa-flavonoid-rich-plantand-its-biological-potential (accessed on 28 April 2020).
- Wiklund, S. Multivariate Data Analysis for Omics; Umea Umetrics: Umeå, Sweden, 2008. [Google Scholar]
- Barton, N.H. The role of hybridization in evolution. Mol. Ecol. 2001, 10, 551–568. [Google Scholar] [CrossRef]
- Boccard, J.; Rutledge, D.N. A consensus orthogonal partial least squares discriminant analysis (OPLS-DA) strategy for multiblock Omics data fusion. Anal. Chim. Acta. 2013, 769, 30–39. [Google Scholar] [CrossRef]
- Kooke, R.; Keurentjes, J.J.B. Epigenetic variation contributes to environmental adaptation of Arabidopsis thaliana. Plant Signal. Behav. 2015, 10, e1057368. [Google Scholar] [CrossRef]
- Verma, N.; Shukla, S. Impact of various factors responsible for fluctuation in plant secondary metabolites. J. Appl. Res. Med. Aromat. Plants 2015, 2, 105–113. [Google Scholar] [CrossRef]
- Newman, D.J.; Cragg, G.M. Natural products as sources of new drugs over the last 25 years. J. Nat. Prod. 2007, 70, 461–477. [Google Scholar] [CrossRef]
- Mazid, M.; Khan, T.A.; Mohammad, F. Role of secondary metabolites in defense mechanisms of plants. Biol. Med. 2011, 3, 232–249. [Google Scholar]
- Speranza, C.I. Drought coping and adaptation strategies: Understanding adaptations to climate change in agro-pastoral livestock production in Makueni district, Kenya. Eur. J. Dev. Res. 2010, 22, 623–642. [Google Scholar] [CrossRef]
- Bhattacharjee, S. Reactive oxygen species and oxidative burst: Roles in stress, senescence and signal transducation in plants. Curr. Sci. 2005, 1113–1121. [Google Scholar]
- Akula, R.; Ravishankar, G.A. Influence of abiotic stress signals on secondary metabolites in plants. Plant Signal. Behav. 2011, 6, 1720–1731. [Google Scholar] [CrossRef] [PubMed]
- Smirnoff, N.; Stewart, G.R. Stress metabolites and their role in coastal plants. In Ecology of Coastal Vegetation; Springer: Haamstede, The Netherlands, 1985; pp. 273–278. [Google Scholar]
- Christmas, M.J. Assessing Genomic Variation in the Hopbush, Dodonaea viscosa, to Investigate Micro-Evolution and Adaptation. Ph.D. Thesis, The University of Adelaide, Adelaide, South Australia, December 2016. [Google Scholar]
- Yu, S.; Fang, N.; Mabry, T.J. Flavonoids from Gymnosperma glutinosum. Phytochemistry 1988, 27, 171–177. [Google Scholar] [CrossRef]
- Sachdev, K.; Kulshreshtha, D.K. Flavonoids from Dodonaea viscosa. Phytochemistry 1983, 22, 1253–1256. [Google Scholar] [CrossRef]
- Compean, K.L.; Ynalvez, R.A. Antimicrobial activity of plant secondary metabolites: A review. Res. J. Med. Plants 2014, 8, 204–213. [Google Scholar]
- Tiwari, R.; Rana, C.S. Plant secondary metabolites: A review. Int. J. Eng. Res. Gen. Sci. 2015, 3, 661–670. [Google Scholar]
- Pirzada, A.J.; Shaikh, W.; Usmanghani, K.; Mohiuddin, E. Antifungal activity of Dodonaea viscosa Jacq extract on pathogenic fungi isolated from super ficial skin infection. Pak. J. Pharm. Sci. 2010, 23, 337–340. [Google Scholar]
- Ramamurthy, V.; Rajeswari, D.M.; Gowri, R.; Vadivazhagi, M.K.; Jayanthi, G.; Raveendran, S. Study of the phytochemical analysis and antimicrobial activity of Dodonaea viscosa. J. Pure Appl. Zool. 2013, 1, 178–184. [Google Scholar]
- Naidoo, R.; Patel, M.; Gulube, Z.; Fenyvesi, I. Inhibitory activity of Dodonaea viscosa var. angustifolia extract against Streptococcus mutans and its biofilm. J. Ethnopharmacol. 2012, 144, 171–174. [Google Scholar] [CrossRef] [PubMed]
- Rajamanickam, V.; Rajasekaran, A.; Anandarajagopal, K.; Sridharan, D.; Selvakumar, K.; Rathinaraj, B.S. Anti-diarrheal activity of Dodonaea viscosa root extracts. Int. J. Pharma. Bio. Sci. 2010, 1, 182–185. [Google Scholar]
- Kabouche, A.; Kabouche, Z.; Öztürk, M.; Kolak, U.; Topçu, G. Antioxidant abietane diterpenoids from Salvia barrelieri. Food Chem. 2007, 102, 1281–1287. [Google Scholar] [CrossRef]
- Dickson, R.A.; Houghton, P.J.; Hylands, P.J. Antibacterial and antioxidant cassane diterpenoids from Caesalpinia benthamiana. Phytochemistry 2007, 68, 1436–1441. [Google Scholar] [CrossRef] [PubMed]
- Neerman, M.F. Sesquiterpene lactones: A diverse class of compounds found in essential oils possessing antibacterial and antifungal properties. Int. J. Aromather. 2003, 13, 114–120. [Google Scholar] [CrossRef]
- Srivastava, J.K.; Gupta, S. Extraction, characterization, stability and biological activity of flavonoids isolated from chamomile flowers. Mol. Cell. Pharmacol. 2009, 1, 138. [Google Scholar] [CrossRef]
- Braca, A.; Sortino, C.; Politi, M.; Morelli, I.; Mendez, J. Antioxidant activity of flavonoids from Licania licaniaeflora. J. Ethnopharmacol. 2002, 79, 379–381. [Google Scholar] [CrossRef]
- Veluri, R.; Weir, T.L.; Bais, H.P.; Stermitz, F.R.; Vivanco, J.M. Phytotoxic and antimicrobial activities of catechin derivatives. J. Agric. Food Chem. 2004, 52, 1077–1082. [Google Scholar] [CrossRef]
- Bais, H.P.; Walker, T.S.; Stermitz, F.R.; Hufbauer, R.A.; Vivanco, J.M. Enantiomeric-dependent phytotoxic and antimicrobial activity of (±)-catechin. A rhizosecreted racemic mixture from spotted knapweed. Plant Physiol. 2002, 128, 1173–1179. [Google Scholar] [CrossRef]
- Muthuswamy, S.; Rupasinghe, H.P.V. Fruit phenolics as natural antimicrobial agents: Selective antimicrobial activity of catechin, chlorogenic acid and phloridzin. J. Food Agric. Environ. 2005, 5, 81–85. [Google Scholar]
- Xie, Y.; Yang, W.; Tang, F.; Chen, X.; Ren, L. Antibacterial activities of flavonoids: Structure-activity relationship and mechanism. Curr. Med. Chem. 2015, 22, 132–149. [Google Scholar] [CrossRef] [PubMed]
- Wayne, P.A. Reference method for broth dilution antifungal susceptibility testing of yeasts, approved standard. CLSI Doc. 2002, M27-A2. [Google Scholar]
- Wikler, M.A. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically: Approved standard. CLSI 2006, 26, M7-A7. [Google Scholar]
- Balouiri, M.; Sadiki, M.; Ibnsouda, S.K. Methods for in vitro evaluating antimicrobial activity: A review. J. Pharm. Anal. 2016, 6, 71–79. [Google Scholar] [CrossRef]
- Goodacre, R.; Broadhurst, D.; Smilde, A.K.; Kristal, B.S.; Baker, J.D.; Beger, R.; Bessant, C.; Connor, S.; Capuani, G.; Craig, A.; et al. Proposed minimum reporting standards for data analysis in metabolomics. Metabolomics 2007, 3, 231–241. [Google Scholar] [CrossRef]
- Heyman, H.M.; Meyer, J.J.M. NMR-based metabolomics as a quality control tool for herbal products. South. Afr. J. Bot. 2012, 82, 21–32. [Google Scholar] [CrossRef]
- Carazzone, C.; Mascherpa, D.; Gazzani, G.; Papetti, A. Identification of phenolic constituents in red chicory salads (Cichorium intybus) by high-performance liquid chromatography with diode array detection and electrospray ionisation tandem mass spectrometry. Food Chem. 2013, 138, 1062–1071. [Google Scholar] [CrossRef]
Sample Availability: Samples of the compounds (1–4) are available from the authors. |
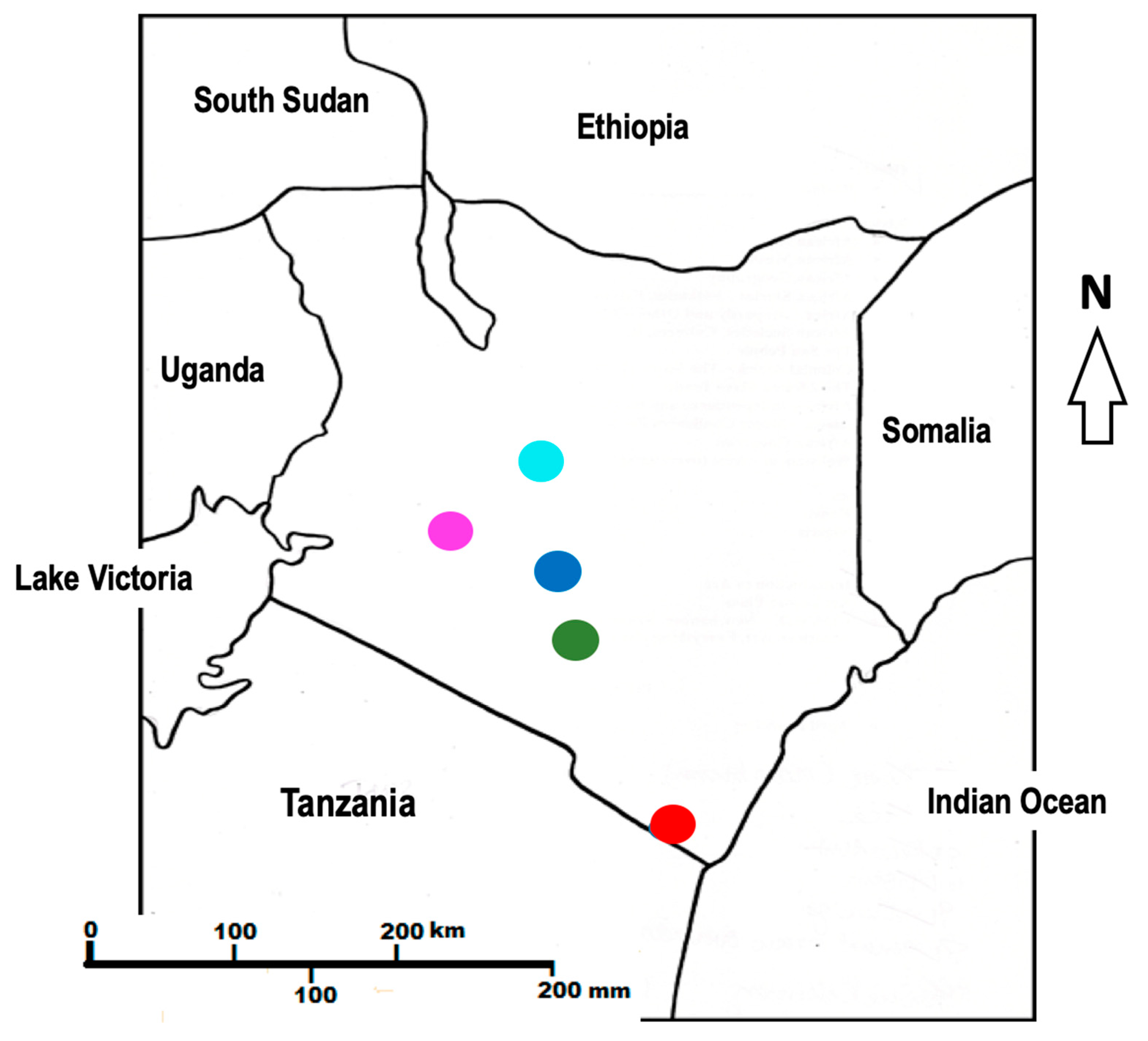

| No. | Locality Name | Specific Location | GPS (S) | GPS (E) | Voucher No |
|---|---|---|---|---|---|
| 1. | Machakos | Kaani ka itheu | 1° 29′ 55.8″ | 37° 21′ 58.2″ | MK2/2018 |
| 2. | Nanyuki | Kahurura | 0° 02′ 06.0″ | 37° 07′ 49.4″ | MK4/2018 |
| 3. | Coast | Gazi | 4° 25′ 29.1″ | 39° 30′ 22.5″ | MK5/2018 |
| 4. | Nairobi | Karura Forest | 1° 14′ 43.6″ | 36° 50′ 17.4″ | MK8/2018 |
| 5. | Narok | Maasai Mara Reserve | 1° 28′ 36.5″ | 35° 05′ 33.3″ | MK9/2018 |
| Experimental m/z [M − H]− | Retention Time (min) | Formula | PPM Error | MSE Fragments | UV (nm) | Proposed Compound | References | |
|---|---|---|---|---|---|---|---|---|
| 1. | 341.1081 | 0.75 | C21H25O4 | 1.2 | 341.1078, 173.0419, 377.0883,515.1646 | weak | Methyl dodovisate B | [25] |
| 2. | 191.0559 | 0.75 | C10H7O4 | 191.0557, 377.0834, 379.0815, 719.1973 | weak | p-Coumaric acid ethyl ester | [26] | |
| 3. | 315.0507 | 1.32 | C16H12O7 | 0 | 315.0504, 300.0272, 151.0038, 107.0134 | weak | Isorhamnetin | [8,17] |
| 4. | 315.0711 | 2.41 | C13H15O9 | −1.9 | 315.0612, 153.0178, 109.0285 | weak | Protocatechuic acid 4-O-glucoside | First report * |
| 5. | 315.0718 | 2.41 | C20H27O3 | 0.8 | 315.0724, 316.0713, 327.0630, 463.0933 | weak | Hardwickiic acid | [25] |
| 6. | 207.0292 | 2.66 | C10H7O5 | −1.4 | 207.0287, 163.0394, 119.049, 165.0393 | 282 | Fraxetin | First report * |
| 7. | 353.0865 | 2.98 | C16H17O9 | −2.3 | 353.0872, 191.0555, 173.0447, 119.0487 | 322 | Chlorogenic acid | First report * |
| 8. | 577.1351 | 3.04 | C30H25O12 | 0.9 | 577.1346, 191.0556, 125.0232, 289.071 | 280 | Procyanidin dimer B5 | First report * |
| 9. | 289.0719 | 3.21 | C15H13O6 | 1.4 | 289.0715, 105.0195, 161.0249, 267.0509 | 279 | Catechin | [8] |
| 10. | 337.0923 | 3.28 | C16H17O8 | −1.2 | 337.0915,173.045, 191.0553, 289.0701 | 281 | p-Coumaric acid | [9] |
| 11. | 313.0234 | 3.42 | C20H25O3 | 1.9 | 313.2361, 309.2002, 311.2331, 311.2128 | 279 | Hautriwaic acid lactone | [25] |
| 12. | 431.1901 | 3.43 | C27H27O5 | −6.9 | 382.119, 300.0346, 153.0962 | weak | 5,7-Dimethoxy-2,2-dimethyl-10-(3-methylbut-2-enyl)-8-phenyl-6-pyrano[3 ,2-g][1]benzopyranone | First report * |
| 13. | 609.1458 | 3.7 | C27H29O16 | 0.3 | 609.1367, 301.0345, 300.0277, 477.0683 | weak | Rutin | [27] |
| 14. | 593.1502 | 3.9 | C27H29O15 | −1.0 | 593.1522, 285.0408 | 351 | Luteolin 7-rutinoside | First report * |
| 15. | 623.1612 | 4.02 | C28H31O16 | 1 | 315.0509, 300.028, 271.0279, 243.0317 | weak | Isorhamnetin 3-O-glucoside 7-O-rhamnoside | First report * |
| 16. | 287.0551 | 4.32 | C15H12O6 | −1.9 | 287.2227, 329.2318, 327.2163, 288.2239 | 361 | Aromadendrin | [11] |
| 17. | 339.1449 | 4.36 | C17H23O7 | 2.13 | 287.0554, 176.0869, 121.0287 | weak | 3-β-acetoxy-5-α-(2-α-hydroxyethyl)acryloyloxy-7-hydroxycarvotacetone | First report * |
| 18. | 465.1913 | 4.93 | C24H33O9 | 1.09 | 379.1883, 285.1477, 241.1612, 119.0352 | 340 | Terpene lactone 1 | First report * |
| 19. | 931 | 4.95 | C48H67O18 | −1.6 | 931,4309, 465.212, 241.1595, 285.1487 | weak | Terpene lactone 1 dimer | First report * |
| 20. | 385.267 | 5.04 | C21H37O6 | −0.5 | 385.2659, 325.2376, 285.1486, 177.0921 | weak | Cryptomeridiol-11-rhamnoside | First report * |
| 21. | 361.1647 | 5.46 | C20H25O6 | 0.72 | 351.2182,3 17.17, 307.23, 243.18, 126.03 | 250 | Diterpene lactone | First report * |
| 22. | 351.2171 | 5.48 | C20H31O5 | 0.6 | 351.2170, 307.2278, 315.0502, 249.1857 | weak | ent-16j-Hydroxy-labdan-3a,8b-dihydroxy,13(14)-en-15,16-olide | [28] |
| 23. | 363.181 | 5.76 | C20H27O6 | 0.6 | 363.1711, 319.1913, 275.201, 259.1695 | weak | Terpentecin 1 | First report * |
| 24. | 727.3684 | 5.76 | C40H55O12 | −1.4 | 727.3680,363.180, 364.1838,275.2008 | weak | Dimer of terpentecin | First report * |
| 25. | 285.0399 | 5.87 | C15H9O6 | 0 | 285.0397, 241.1589, 151.0029, 242.162 | 266, 366 | Kaempferol | [16] |
| 26. | 301.0711 | 5.93 | C15H9O7 | 3.0 | 301.0700, 609.1830, 610.1893, 302.0850 | 251 | Quercetin | [8] |
| 27. | 315.0021 | 5.97 | C30 H25O12 | 1.1 | 315.2560, 293.2096, 316.2598, 249.1506 | weak | Quercetin 3′-O-methyl ether | [9] |
| 28. | 363.1807 | 6.12 | C20H27O6 | 0.6 | 363.1805, 319.1913, 275.201, 259.1695 | weak | Terpentecin 2 | First report * |
| 29. | 429.2489 | 6.13 | C23H26O8 | 2.7 | 429.2496, 351.2181, 299.0559, 285.0429 | 261 | Aliarin | [26] |
| 30. | 299.0557 | 6.2 | C16H11O6 | 1.6 | 299.0557, 375.1808, 347.1865, 300.0591 | weak | Isokaempferide | [17,27,29] |
| 31. | 327.2164 | 6.3 | C18H16O6 | 1.8 | 327.2166, 328.2183, 325.1966, 313.1455 | weak | Kaempferol 5,7,4′-trimethyl ether | [28] |
| 32. | 359.0764 | 6.58 | C21H27O5 | 0.8 | 359.0974, 266.0666, 197.0446, 435.1606 | weak | Methoxymkapwanin | [27] |
| 33. | 375.0723 | 6.88 | C18H16O9 | 1.6 | 375.0722, 361.1641, 351.2161, 551.2051 | 280 | 5,7,4′,5′-Tetrahydroxy-3,6,2′–trimethoxyflavone | First report * |
| 34. | 329.0663 | 7.00 | C20H25O4 | 0.2 | 329.0654, 361.1648, 347.1866, 418.2239 | weak | Dodonic acid | [17] |
| 35. | 329.0661 | 7.07 | C17H13O7 | 1.5 | 329.0660, 271.0249, 314.0432, 275.2003 | 277,337 | Rhamnazin | [17] |
| 36. | 375.1808 | 7.13 | C21H28O6 | −4.46 | 345.17, 319.2, 259.17, 116.93 | 280 | Terpene lactone 1 | First report * |
| 37. | 343.0822 | 7.43 | C18H15O7 | −1.3 | 343.0817, 344.0853, 393.1919, 345.1689 | 279 | Penduletin | [28] |
| 38. | 377.1957 | 7.56 | C21H29O6 | −1.9 | 377.1957, 345.1701, 301.1799, 189.1278 | 350 | Hypophyllin E | First report * |
| 39. | 285.0764 | 7.62 | C16H12O5 | 1.5 | 285.0403, 286.0438, 533.2025, 571.0843 | 261 | Sakuranetin | [27] |
| 40. | 255.0662 | 7.78 | C15H11O4 | 1.2 | 255.0660, 151.0035, 213.0554, 107.0135 | 288 | Pinocembrin | [17] |
| 41. | 347.1857 | 7.91 | C20H27O5 | −0.3 | 347.1850, 303.1965, 285.1488, 241.1591 | weak | (A)-6a-Hydroxy-5a,8a,9a,10a-cleroda-3,13-dien-16,15-olid-18-oicacid | [10] |
| 42. | 299.0560 | 7.96 | C16H11O6 | 1.3 | 299.055, 271.0605, 65.0193, 284.0329 | 266,365 | Rhamnocitrin | [17] |
| 43. | 313.0713 | 8.20 | C17H13O6 | 0.7 | 313.0712, 314.0749 377.1953, 298.0486 | 361 | Kumatakenin | [17] |
| 44. | 343.0812 | 8.41 | C18H15O7 | −1.7 | 343.0812, 313.0346, 301.1798, 270.0161 | 270,340 | Santin | [17] |
| 45. | 413.1236 | 8.76 | C22H21O8 | 0.5 | 413.1231, 368.0905, 331.1908, 161.0144 | 351 | Picropodophyllin | First report * |
| 46. | 367.1180 | 8.91 | C21H19O6 | −0.5 | 352.0954, 323.0915, 297.0328, 269.044 | 267 | 7-O-methylluteone | First report * |
| 47. | 313.0720 | 8.98 | C17H13O6 | 2.6 | 313.0720, 283.0252, 255.0296, 161.0255 | 267,347 | 3,5-Dihydroxy-4′,7-dimethoxyfla-vone | [17,27] |
| 48. | 331.1910 | 9.19 | C20H27O4 | 0.3 | 331.1914, 397.1290, 332.1949, 398.1334 | 340 | Hautriwaic acid | [17] |
| 49. | 397.1280 | 9.31 | C22H21O7 | −0.4 | 397.1291, 1169.585, 331.1901, 1156.5643 | weak | 5,7,3′-Trihydroxy-3,5′-dimethoxy-2′-(3′-methylbut-2-enyl)flavone | [26] |
| 50. | 483.2021 | 9.35 | C27H31O8 | 0.2 | 483.2018, 453.1683,331.1906, 397.1288 | 341 | 5,7,4′-Trihydroxy-3′-(4-hydroxy-3-Methylbutyl)-5′-prenyl-3,6-diMethoxyflavone | [26] |
| 51. | 293.2117 | 9.58 | C18H29O3 | −1.4 | 293.2111,161.0242, 152.9944, 265.1489 | 267,347 | 17-Hydroxylinolenic acid | First report * |
| 52. | 321.2443 | 10.08 | C20H33O3 | −2.43 | 277.2537, 116.938 | weak | diterpenoid | First report * |
| 53. | 411.1444 | 10.19 | C23H23O7 | 0 | 411.1439, 396.1214, 331.1898, 265.1464 | 340 | Viscosol | [28] |
| 54. | 435.1808 | 10.36 | C26H27O6 | −1.6 | 435.1806, 255.2322, 365.102, 161.0247 | weak | Artocarpin | First report * |
| 55. | 277.2168 | 11.33 | C18H29O2 | 0 | 277.2163, 116.9271, 265.1465, 152.9955 | weak | Punicic acid | First report * |
| Retention Time (min) | ESI negative [M − H]− (m/z) | Elemental Composition | Name | Population with Highest Concentration | |
|---|---|---|---|---|---|
| 1. | 0.75 | 191.0559 | C10H7O4 | p-coumaric acid ethyl ester | Nanyuki |
| 2. | 5.48 | 351.2174 | C20H31O5 | ent-16j-hydroxy-labdan-3a,8b-dihydroxy,13(14)-en-15,16-olide | Nairobi |
| 3. | 5.76 | 363.1808 | C20H27O6 | terpentecin 1 | coastal (Gazi) |
| 4. | 5.76 | 727.3681 | C40H55O12 | dimer of terpentecin 1 | coastal (Gazi) |
| 5. | 6.11 | 727.3676 | C40H55O12 | dimer of terpentecin 2 | coastal (Gazi) |
| 6. | 6.12 | 363.1807 | C20H27O6 | terpentecin 2 | Nanyuki |
| 7. | 7.00 | 329.0663 | C20H25O4 | dodonic acid | Nanyuki |
| 8. | 7.53 | 377.1964 | C21H29O6 | hypophyllin E | Nanyuki |
| 9. | 7.69 | 255.0661 | C15H11O4 | pinocembrin | Nanyuki |
| 10. | 8.20 | 313.07 | C17H13O6 | kumatakenin | Nairobi |
| 11. | 8.41 | 343.0818 | C18H15O7 | santin | Nanyuki |
| Sample Name | Microbial Names | |||
|---|---|---|---|---|
| MRSA | S. aureus | E. coli | C. albicans | |
| Dodonic acid (1) | >1000 | 500 | 500 | >1000 |
| 5,7,4′,5′-Tetrahydroxy-3,6,2′ trimethoxyflavone (2) | >1000 | >1000 | >1000 | >1000 |
| Hautriwaic acid lactone (3) | 62.50 | 1.95 | 1.95 | 7.81 |
| Catechin (4) | 7.81 | 3.91 | 7.81 | 3.91 |
| Coastal (Gazi) | 62.50 | 3.91 | 7.81 | 15.62 |
| Machakos | 125 | 7.81 | 15.62 | 31.20 |
| Nairobi | >1000 | 31.25 | 62.50 | >1000 |
| Nanyuki | 62.50 | 7.81 | 15.62 | 7.81 |
| Narok | 125 | 15.62 | 31.25 | 62.5 |
| Omacilin | 0.98 | 0.49 | 0.98 | - |
| Fluconazole | 1.95 | |||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kaigongi, M.M.; Lukhoba, C.W.; Ochieng‘, P.J.; Taylor, M.; Yenesew, A.; Makunga, N.P. LC-MS-Based Metabolomics for the Chemosystematics of Kenyan Dodonaea viscosa Jacq (Sapindaceae) Populations. Molecules 2020, 25, 4130. https://doi.org/10.3390/molecules25184130
Kaigongi MM, Lukhoba CW, Ochieng‘ PJ, Taylor M, Yenesew A, Makunga NP. LC-MS-Based Metabolomics for the Chemosystematics of Kenyan Dodonaea viscosa Jacq (Sapindaceae) Populations. Molecules. 2020; 25(18):4130. https://doi.org/10.3390/molecules25184130
Chicago/Turabian StyleKaigongi, Magrate M., Catherine W. Lukhoba, Purity J. Ochieng‘, Malcolm Taylor, Abiy Yenesew, and Nokwanda P. Makunga. 2020. "LC-MS-Based Metabolomics for the Chemosystematics of Kenyan Dodonaea viscosa Jacq (Sapindaceae) Populations" Molecules 25, no. 18: 4130. https://doi.org/10.3390/molecules25184130
APA StyleKaigongi, M. M., Lukhoba, C. W., Ochieng‘, P. J., Taylor, M., Yenesew, A., & Makunga, N. P. (2020). LC-MS-Based Metabolomics for the Chemosystematics of Kenyan Dodonaea viscosa Jacq (Sapindaceae) Populations. Molecules, 25(18), 4130. https://doi.org/10.3390/molecules25184130






