New Arctic Bacterial Isolates with Relevant Enzymatic Potential
Abstract
1. Introduction
2. Results and Discussion
3. Materials and Methods
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Grinnell, J. With frontispiece and eleven other illustrations by Hilda Wood Grinnell. The Condor Volume XLII. 1940, p. 1. Available online: https://sora.unm.edu/sites/default/files/journals/condor/v042n01/p0003-p0034.pdf (accessed on 1 July 2020).
- Youn, U.J.; Kim, M.J.; Han, S.J.; Yim, J.H. Isolation of secondary metabolites from an Arctic bacterium, Pseudomonas aeruginosa and their antimicrobial activities. Kor. J. Microb. 2016, 52, 415–420. [Google Scholar] [CrossRef][Green Version]
- Han, S.J.; Park, H.; Kim, S.; Kim, D.; Park, H.J.; Yim, J.H. Enhanced production of protease by Pseudoalteromonas arctica PAMC 21717 via statistical optimization of mineral components and fed-batch fermentation. Prep. Biochem. Biotech. 2016, 46, 328–335. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.S.; You, Y.-P.; Han, Y.H.; Xiang, P.; Zhang, W.X.; Zhou, Z.H.; Chen, D.L.; Li, M. Siderophores produces by Pseudomonas sp. Strain 4-05: Identification, purification and ferrous- dependent production traits. Fresenius Envir. Bull. 2016, 25, 5980–5988. [Google Scholar]
- Meyer, J.M.; Abdallah, M.A. The Fluorescent Pigment of Pseudomonas fluorescens: Biosynthesis, Purification and Physicochemical Properties. J. G. Microb. 1997, 107, 319–328. [Google Scholar] [CrossRef]
- Budzikiewicz, H. Siderophores of the Pseudomonas sensu strict (fluorescent and non-fluorescent Pseudomonas ssp.). Fortschr. Chem. Org. Naturst. 2004, 87, 81–237. [Google Scholar]
- Hohnadel, D.; Meyer, J.M. Specifity of pyoverdine- mediated iron uptake among fluorescent Pseudomonas strains. J. Bacteriol. 1988, 170, 4865–4873. [Google Scholar] [CrossRef]
- Jankiewicz, U. Charakterystyka i znaczenie piowerdyn bakterii z rodzaju Pseudomonas. Post. Mikrobiol. 2009, 48, 243–254. [Google Scholar]
- Gleaser, S.P.; Kampfer, P. Multilocus sequence analysis (MLSA) in prokaryotic taxonomy. Syst. Appl. Microb. 2015, 38, 237–245. [Google Scholar] [CrossRef]
- Garrio-Sanz, D.; Meier-Kolthoff, J.P.; Goker, M.; Martin, M.; Rivilla, R.; Redondo-Nieeto, M. Genomic ad Genetic Diversity with the Pseudomonas fluorescens Complex. PLoS ONE 2016, 11, e0150183. [Google Scholar] [CrossRef]
- Andersen, S.M.; Johnsen, K.; Sørensen, J.; Nielsen, P.; Jacobsen, C.S. P. frederiksbergensis sp. nov., isolated from soil at a coal gasification site. Int. J. Syst. Evol. Microbiol. 2000, 50, 1957–1964. [Google Scholar] [CrossRef]
- Reddy, G.S.N.; Matsumoto, G.I.; Schumann, P.; Stackebrandt, E.; Shivaji, S. Psychrophilic pseudomonads from Antarctica: Pseudomonas antarctica sp. nov., Pseudomonas. meridiana sp. nov. and Pseudomonas proteolytica sp. nov. Int. J. Syst. and Evol. Microbiol. 2004, 54, 713–719. [Google Scholar] [CrossRef] [PubMed]
- Achouak, W.; Conrod, S.; Cohen, V.; Heulin, T. Phenotypic variation of Pseudomonas brassicacearum as a plant root-colonization strategy. Mol. Plant Microbe. Interact. 2004, 17, 872–879. [Google Scholar] [CrossRef] [PubMed]
- Munsch, P.; Geoggroy, V.A.; Alatossava, T.; Meyer, J.-M. Application of Siderotyping for Characterization of Pseudomonas tolaasii and “Pseudomonas reactans” Isolates Associated with Brown Blotch Disease of Cultivated Mushrooms. Appl. Environ. Microb. 2000, 66, 4834–4841. [Google Scholar] [CrossRef]
- Achouak, W.; Sutra, L.; Heulin, T.; Meyer, J.-M.; Fromin, N.; Degraeve, S.; Christen, R.; Gardan, L. Pseudomonas brassicacearum sp. nov. and Pseudomonas thivervalensis sp. nov., two root-associated bacteria isolated from Brassica napus and Arabidopsis thaliana. Int. J. Syst. Evol. Microbiol. 2000, 50, 9–18. [Google Scholar] [CrossRef] [PubMed]
- Verhille, S.; Baida, N.; Dabboussi, F.; Izard, D.; Leclerc, H. Taxonomic study of bacteria isolated from natural mineral waters: Proposal of Pseudomonas jessenii sp. nov. and Pseudomonas mandelii sp. nov. Syst. Appl. Microb. 1999, 22, 45–58. [Google Scholar] [CrossRef]
- Wieser, M.; Denner, E.B.M.; Kampfer, P.; Schumann, P.; Tindall, B.; Steiner, U.; Vybiral, D.; Lubitz, W.; Maszenan, A.; Patel, M.; et al. Emended descriptions of the genus Micrococcus, Micrococcus luteus (Cohn 1872) and Micrococcus lylae (Kloos et al. 1974). Int. J. Syst. Evol. Microbiol. 2002, 52, 629–637. [Google Scholar] [CrossRef]
- Mulet, M.; Lalucat, J.; García-Valdés, E. DNA sequence-based analysis of the Pseudomonas species. Environ. Microbiol. 2010, 12, 1513–1530. [Google Scholar]
- Daffonchio, D.; Raddadi, N.; Merabishvili, M.; Cherif, A.; Carmagnola, L.; Brusetti, L.; Rizzi, A.; Chanishvili, N.; Visca, P.; Sharp, R.; et al. Strategy for identification of Bacillus cereus and Bacillus thuringiensis strains closely related to Bacillus anthracis. Appl. Environ. Microbiol. 2006, 72, 1295–1301. [Google Scholar] [CrossRef]
- Liu, Y.; Lai, Q.; Göker, M.; Meier-Kolthoff, J.P.; Wang, M.; Sun, Y.; Wang, L.; Shao, Z. Genomic insights into the taxonomic status of the Bacillus cereus group. Sci. Rep. 2015, 5, 14082. [Google Scholar] [CrossRef]
- Spilker, T.; Baldwin, A.; Bumford, A.; Dowson, C.G.; Mahenthiralingam, E.; LiPuma, J.J. Expanded multilocus sequence typing for Burkholderia species. J. Clin. Microbiol. 2009, 47, 2607–2610. [Google Scholar] [CrossRef]
- Urwyler, S.K.; Glaubitz, J. Advantage of MALDI-TOF-MS over biochemical-based phenotoping for microbial identification illustrated on industrial applications. Lett. Apll. Microb. 2015, 62, 130–137. [Google Scholar] [CrossRef] [PubMed]
- Aksentijević, K.; Ašanin, J.; Milivojević, D.; Čolović, S.; Butorac, A.; Cindrić, M.; Mišić, D. Differentiation between Pseudomonas and Stenotrophomonas Species Isolated from Fish Using Molecular and MALDI-TOF Method. Acta Veterinaria 2016, 66, 304–316. [Google Scholar] [CrossRef]
- Timperio, A.M.; Gorrasi, S.; Zolla, L.; Fenice, M. Evaluation of MALDI-TOF mass spectrometry and MALDI BioTyper in comparison to 16S rDNA sequencing for the identification of bacteria isolated from Arctic sea water. PLoS ONE 2017, 12, e0181860. [Google Scholar] [CrossRef] [PubMed]
- Strejcek, M.; Smrhova, T.; Junkova, P.; Uhlik, O. Whole-Cell MALDI-TOF MS versus 16S rRNA Gene Analysis for Identification and Dereplication of Recurrent Bacterial Isolates. Fron. Microb. 2018, 9, 1294–1302. [Google Scholar] [CrossRef] [PubMed]
- Ferres, I.; Amarelle, V.; Noya, F.; Fabiano, E. Identification of Antarctic culturable bacteria able to produce diverse enzymes of potential biotechnological interest. Adv. Polar Sci. 2015, 25, 71–79. [Google Scholar]
- Meyer, J.M.; Stintzi, A. Iron metabolism and siderophores in Pseudomonas and related species. In Pseudomonas. Biotechnology Handbooks 10 Plenum Publishing; Montie, T.C., Ed.; Springer Science + Business Media, LLC: New York, NY, USA, 1998; pp. 201–243. [Google Scholar]
- Visca, P.; Ciervo, A.; Orsi, N. Cloning and nucleotide sequenceof the pvdA gene encoding the pyoverdin biosynthetic enzyme L-ornithine N5-oxygenase in Pseudomonas aeruginosa. J. Bacteriol. 1994, 176, 1128–1140. [Google Scholar] [CrossRef]
- Kačániová, M.; Klūga, A.; Kántor, A.; Medo, J.; Žiarovská, J.; Puchalski, C.; Trentjeva, M. Comparison of MALDI-TOF MS Biotyper and 16S rDNA sequencing for the identification of Pseudomonas species isolated from fish. Microb Pathog. 2019, 132, 313–318. [Google Scholar] [CrossRef]
- Laba, W.; Choinska, A.; Rodziewicz, A.; Piegza, M. Keratinolytic abilities of Micrococcus luteus from poultry waste. Braz. J. Microbiol. 2015, 46, 691–700. [Google Scholar] [CrossRef]
- Piegza, M.; Witkowska, D.; Stempniewicz, R. Enzymatic and molecular characteristics of Geotrichum candidum strains as a starter culture for malting. J. Inst. Brew. 2014, 120, 341–346. [Google Scholar] [CrossRef]
- Blacket, M.J.; Robin, C.; Good, R.T.; Lee, S.F.; Miller, A.D. Universal primers for fluorescent labelling of PCR fragments—An efficient and cost-effective approach to genotyping by fluorescence. Mol. Ecol. Res. 2012, 12, 456–463. [Google Scholar] [CrossRef]
Sample Availability: Samples of the compounds are not available from the authors. |
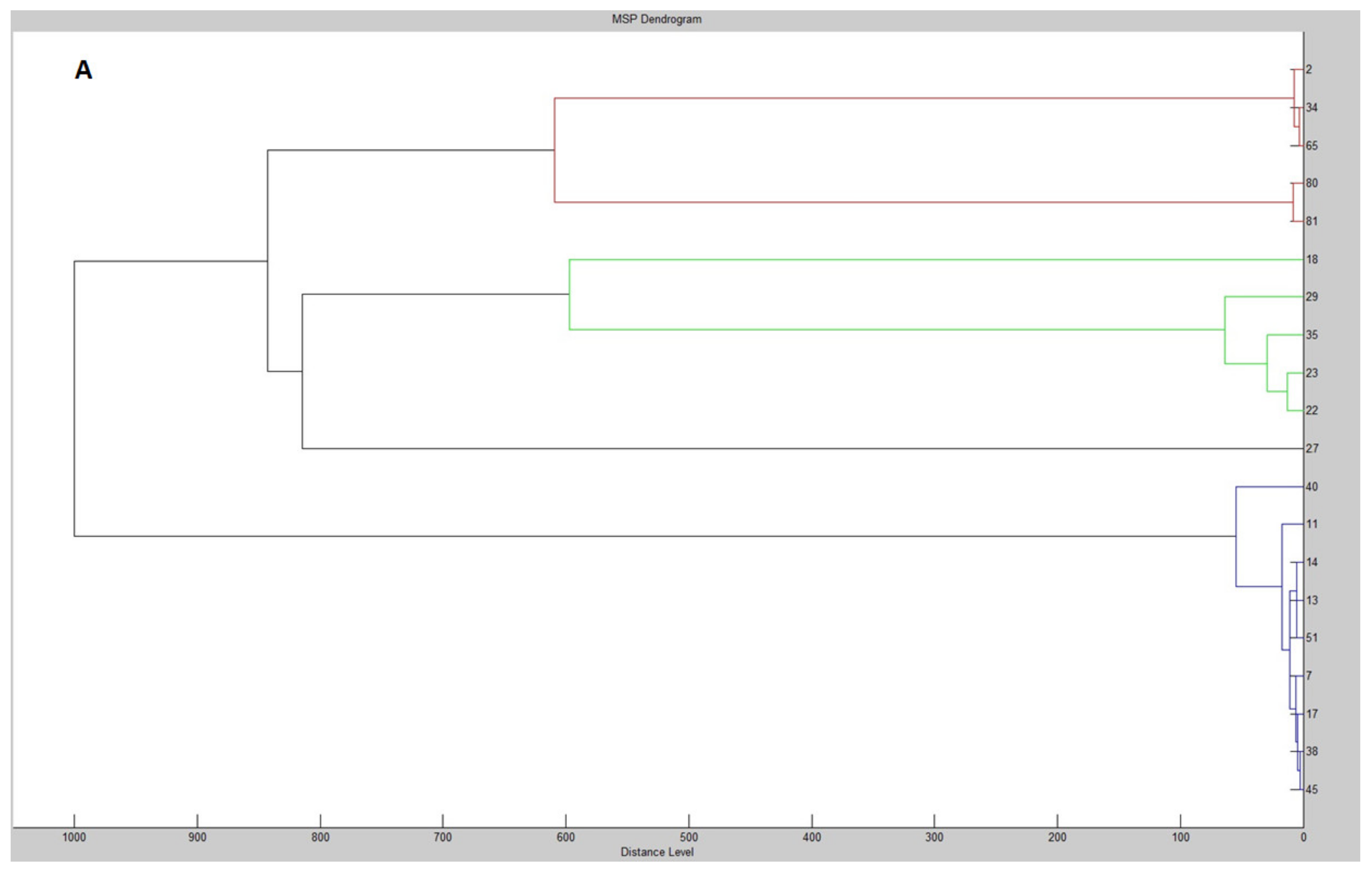

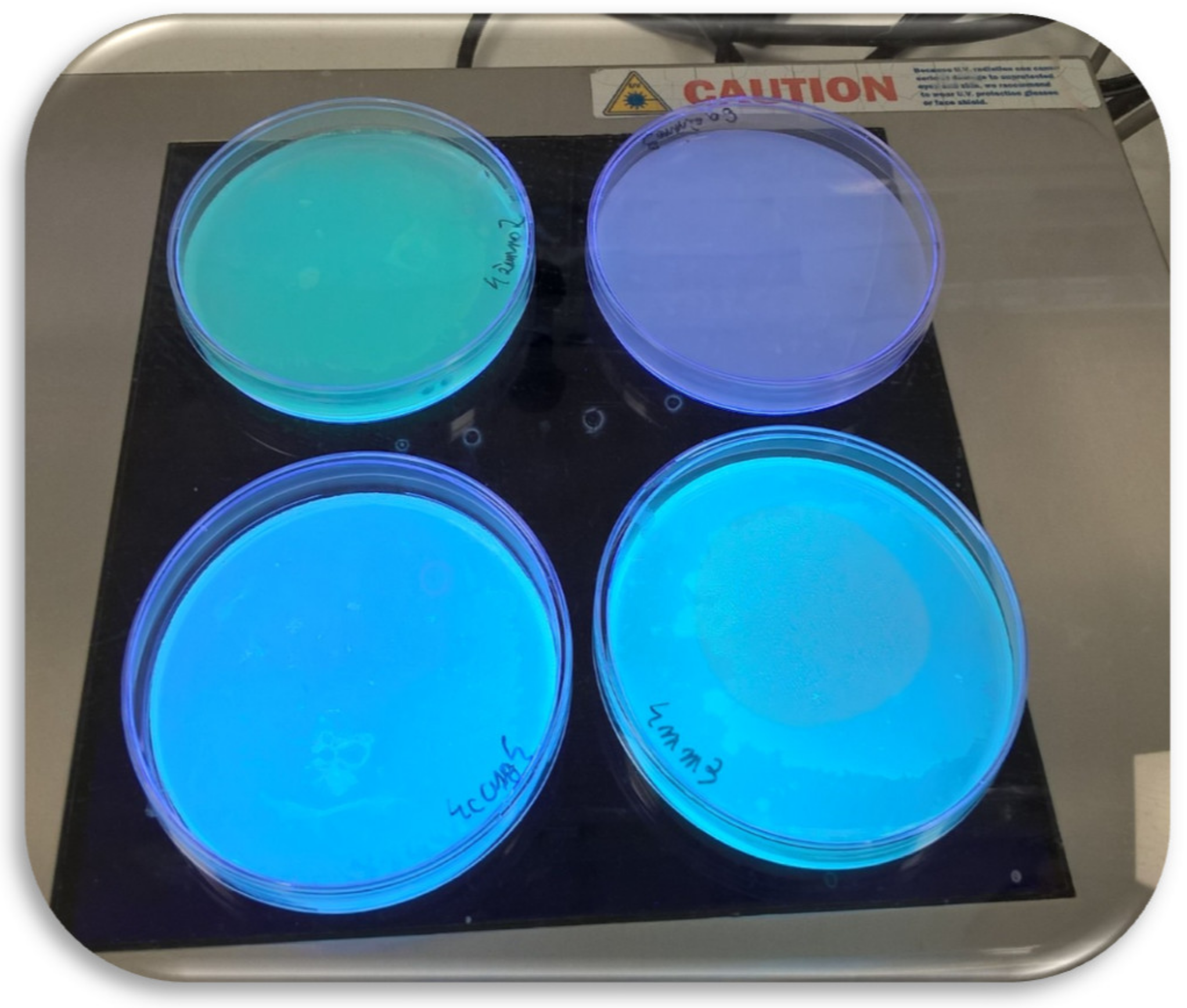

| MALDI-TOF MS Biotyper | 16S rDNA | CEL | XYL | PG | PROT | FL | |
|---|---|---|---|---|---|---|---|
| WG 1 | Arthrobacter oryzae | Arthrobacter oryzae | 0 | 0.18 | 0.06 | 0 | |
| WG 2 | Arthobacter citreus | Arthrobacter citreus | 0.05 | 0.49 | 0.12 | 0 | |
| WG 3 | Arthrobacter sp. | Arthrobacter sp. | 0.03 | 0 | 0.12 | 0 | |
| WG 4 | Bacillus cereus | Bacillus sp. | 0 | 0 | 0 | 0 | |
| WG 5 | Pseudomonas sp. | Pseudomonas sp. | 0.06 | 0.18 | 0 | 0.01 | |
| WG 7 | Pseudomonas frederiksbergensis | Pseudomonas sp. | 0 | 0.02 | 0.07 | 0.02 | + |
| WG 8 | Bacillus weihenstephanensis/mycoides | Bacillus mycoides | 0 | 0 | 0 | 0 | + |
| WG 9 | Pseudomonas frederiksbergensis | Pseudomonas frederiksbergensis | 0.03 | 0 | 0 | 0 | |
| WG 11 | Pseudomonas frederiksbergensis | Pseudomonas frederiksbergensis | 0 | 0 | 0 | 0.01 | |
| WG 12 | Mycobacterium sp. | Massilia sp. | 0.15 | 0.20 | 0.04 | 0 | |
| WG 13 | Pseudomonas frederiksbergensis | Pseudomonas sp. | 0.10 | 0.59 | 0.31 | 0 | |
| WG 14 | Pseudomonas frederiksbergensis | Pseudomonas sp. | 0 | 0 | 0 | 0 | + |
| WG 15 | Rhodococcus erythopolis | Rhodococcus erythropolis | 0 | 0.04 | 0 | 0 | |
| WG 16 | Pseudomonas sp. | Pseudomonas frederiksbergensis | 0.11 | 0.39 | 0.17 | 0 | |
| WG 17 | Pseudomonas frederiksbergensis | Pseudomonas frederiksbergensis | 0.19 | 0 | 0 | 0.06 | |
| WG 18 | Rhodococcus sp. | Rhodococcus erythropolis | 0 | 0 | 0.09 | 0 | |
| WG 19 | Rhodococcus erythopolis | Rhodococcus erythopolis | |||||
| WG 20 | Rhodococcus erythopolis | Rhodococcus erythropolis | 0 | 0.01 | 0.06 | 0.03 | |
| WG 21 | Pseudomonas sp. | Pseudomonas sp. | 0 | 0.05 | 0 | 0 | |
| WG 22 | Rhodococcus erythopolis | Rhodococcus erythropolis | 0 | 0 | 0 | 0 | |
| WG 23 | Rhodococcus erythopolis | Rhodococcus erythropolis | 0 | 0 | 0 | 0 | |
| WG 25 | Pseudomonas sp. | Pseudomonas sp. | 0.06 | 0.42 | 0 | 0 | |
| WG 26 | Arthobacter citreus | Arthobacter citreus | 0 | 0.05 | 0.07 | 0.01 | |
| WG 27 | Rhodococcus erythopolis | Pseudomonas frederiksbergensis | 0 | 0.42 | 0.44 | 0 | |
| WG 28 | Acinetobacter sp. | Acinetobacter sp. | 0.08 | 0 | 0.01 | 0 | |
| WG 29 | Rhodococcus erythopolis | Rhodococcus erythropolis | 0.02 | 0.04 | 0.01 | 0.05 | |
| WG 30 | Rhodococcus erythopolis | Rhodococcus erythopolis | |||||
| WG 34 | Arthobacter citreus | Arthrobacter citreus | 0.02 | 0.03 | 0.05 | 0.02 | |
| WG 35 | Rhodococcus erythopolis | Rhodococcus erythropolis | 0.01 | 0 | 0.01 | 0.02 | |
| WG 36 | Bacillus sp. | Sporosarcina aquimarina | 0 | 0.26 | 0.11 | 0.08 | |
| WG 38 | Pseudomonas sp. | Pseudomonas frederiksbergensis | 0 | 0 | 0.05 | 0 | + |
| WG 40 | Pseudomonas antarctica | Pseudomonas antarctica | 0.02 | 0.01 | 0.05 | 0 | + |
| WG 43 | Pseudomonas arsenicoxydans | 0 | 0 | 0.20 | 0 | ||
| WG 44 | Pseudomonas frederiksbergensis | Pseudomonas frederiksbergensis | 0.08 | 0 | 0 | 0.12 | + |
| WG 45 | Pseudomonas frederiksbergensis | Pseudomonas frederiksbergensis | 0 | 0.77 | 0 | 0.01 | |
| WG 46 | Sphingomonas sp. | 0 | 0.3 | 0 | 0 | ||
| WG 47 | Rhodococcus erythopolis | Rhodococcus erythopolis | 0 | 0.07 | 0.03 | 0 | |
| WG 51 | Pseudomonas sp. | Pseudomonas sp. | 0.09 | 0.08 | 0.30 | 0 | + |
| WG 53 | Pseudomonas frederiksbergensis | Pseudomonas frederiksbergensis | 0 | 0.01 | 0.10 | 0 | |
| WG 54 | Pseudomonas brassicacearum | Pseudomonas brassicacearum | 0 | 0.01 | 0.01 | 0 | |
| WG 56 | Pseudomonas koreensis | Pseudomonas koreensis | 0.01 | 0 | 0.09 | 0 | |
| WG 60 | Pseudomonas sp. | Pseudomonas sp. | 0.02 | 0.05 | 0.18 | 0.02 | |
| WG 61 | Pseudomonas frederiksbergensis | Pseudomonas frederiksbergensis | 0 | 0 | 0 | 0 | +/− |
| WG 62 | Pseudomonas corrugata | Pseudomonas corrugata | 0 | 0.04 | 0.09 | 0 | |
| WG 64 | Pseudomonas mandelii | Pseudomonas mandelii | 0 | 0.02 | 0 | 0.01 | + |
| WG 65 | Arthobacter citreus | Arthrobacter citreus | 0.20 | 0 | 0.09 | 0 | |
| WG 67 | Burkholderia sordidicola | 0.04 | 0 | 0.28 | 0 | ||
| WG 69 | Bacillus psychrosaccharolyticus | Bacillus psychrosaccharolyticus | 0 | 0 | 0 | 0.01 | |
| WG 71 | Pseudomonas sp. | Pseudomonas sp. | 0 | 0.03 | 0.58 | 0 | |
| WG 72 | Burkholderia sp. | 0.16 | 0 | 0 | 0.05 | ||
| WG 76 | Lactobacillus sp. | Lactobacillus sp. | 0.011 | 0.39 | 0.04 | 0 | |
| WG 77 | Micrococcus luteus | Micrococcus luteus | 0.09 | 0.01 | 0.06 | 0 | |
| WG 79 | Arthrobacter oxydans | 0.22 | 0.47 | 0 | 0 | ||
| WG 80 | Micrococcus luteus | Micrococcus luteus | 0 | 0.13 | 0 | 0 | |
| WG 81 | Micrococcus luteus | Micrococcus luteus | 0 | 0.92 | 0 | 0 | + |
| WG 82 | Micrococcus luteus | Micrococcus luteus | 0 | 0 | 1.60 | 0 | |
| WG 83 | Micrococcus luteus | Micrococcus luteus | 0.02 | 0.03 | 0.33 | 0.01 | |
| WG 85 | Arthrobacter sp. | 0.07 | 0 | 0.03 | 0 | ||
| WG 89 | Arthobacter polychromogenes | Burkholderia sp. | 0.01 | 0.16 | 0.17 | 0 | |
| WG 93 | Burkholderia sp. | 0 | 0 | 0.77 | 0 | ||
| WG 94 | Aerococcus vividans | Sphingomonas sp. | 0.03 | 0 | 0.03 | 0 | |
| WG 95 | Sphingomonas sp. | 0.01 | 0 | 0.05 | 0 | ||
| WG 99 | Cupriavidus metallidurans | Sphingomonas sp. | 0.05 | 0 | 0.28 | 0.01 | |
| WG 100 | Arthobacter sufonivorans | Arthobacter sufonivorans | 0 | 0.2 | 0.03 | 0 | |
| WG 101 | Cupriavidus metallidurans | Sphingomonas sp. | 0 | 0.05 | 0.07 | 0 | |
| WG 103 | Mycobacterium sp. | Caulobacter sp. | 0 | 0.9 | 0 | 0.02 | |
| WG 104 | Arthobacter sp. | Arthobacter sp. | 0 | 0 | 0.03 | 0.05 | |
| WG 108 | Arthobacter ramosus | Arthrobacter sp. | 0 | 0 | 0.05 | 0 | |
| WG 113 | Mycobacterium sp. | Mycobacterium sp. | 0 | 0 | 0 | 0 | |
| WG 114 | Bacillus simplex | Bacillus simplex | 0.08 | 0 | 0 | 0 | |
| WG 116 | Acinetobacter baumanii | Frondihabitans sp. | 0.01 | 0.02 | 0.05 | 0.01 | |
| WG 117 | Paenibacillus pabuli | Streptococcus salivarius | 0.01 | 0.04 | 0.03 | 0.01 |
| Fluc I | Fluc II | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| STRAINS | STRAINS | ||||||||||
| 11 | 40 | 54 | 64 | 81 | 939 | 11 | 40 | 54 | 64 | 81 | 939 |
| 2262 | 2045 | ||||||||||
| 1647 | 1516 | ||||||||||
| 1207 | 1244 | ||||||||||
| 1000 | 1047 | 1192 | 1131 | ||||||||
| 988 | 994 | 934 | |||||||||
| 889 | 856 | 805 | |||||||||
| 771 | 767 | 729 | |||||||||
| 676 | 682 | 668 | 691 | ||||||||
| 517 | 500 | 593 | 586 | 593 | 520 | 582 | 589 | 589 | |||
| 500 | 434 | 488 | 488 | ||||||||
| 438 | 461 | 303 | 310 | 303 | 339 | 362 | |||||
| 308 | 376 | 367 | 376 | 344 | 202 | 245 | |||||
| 270 | 280 | 292 | 275 | 187 | 74 | 96 | |||||
| 232 | 216 | ||||||||||
| 174 | 179 | 167 | 155 | 80 | 147 | ||||||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Piegza, M.; Łaba, W.; Kačániová, M. New Arctic Bacterial Isolates with Relevant Enzymatic Potential. Molecules 2020, 25, 3930. https://doi.org/10.3390/molecules25173930
Piegza M, Łaba W, Kačániová M. New Arctic Bacterial Isolates with Relevant Enzymatic Potential. Molecules. 2020; 25(17):3930. https://doi.org/10.3390/molecules25173930
Chicago/Turabian StylePiegza, Michał, Wojciech Łaba, and Miroslava Kačániová. 2020. "New Arctic Bacterial Isolates with Relevant Enzymatic Potential" Molecules 25, no. 17: 3930. https://doi.org/10.3390/molecules25173930
APA StylePiegza, M., Łaba, W., & Kačániová, M. (2020). New Arctic Bacterial Isolates with Relevant Enzymatic Potential. Molecules, 25(17), 3930. https://doi.org/10.3390/molecules25173930







