Identification of Selected Antibiotic Resistance Genes in Two Different Wastewater Treatment Plant Systems in Poland: A Preliminary Study
Abstract
1. Introduction
2. Results and Discussion
2.1. Evaluation of Total DNA Extraction Kits
2.2. Occurrence and Removal of ARGs in WWTPs
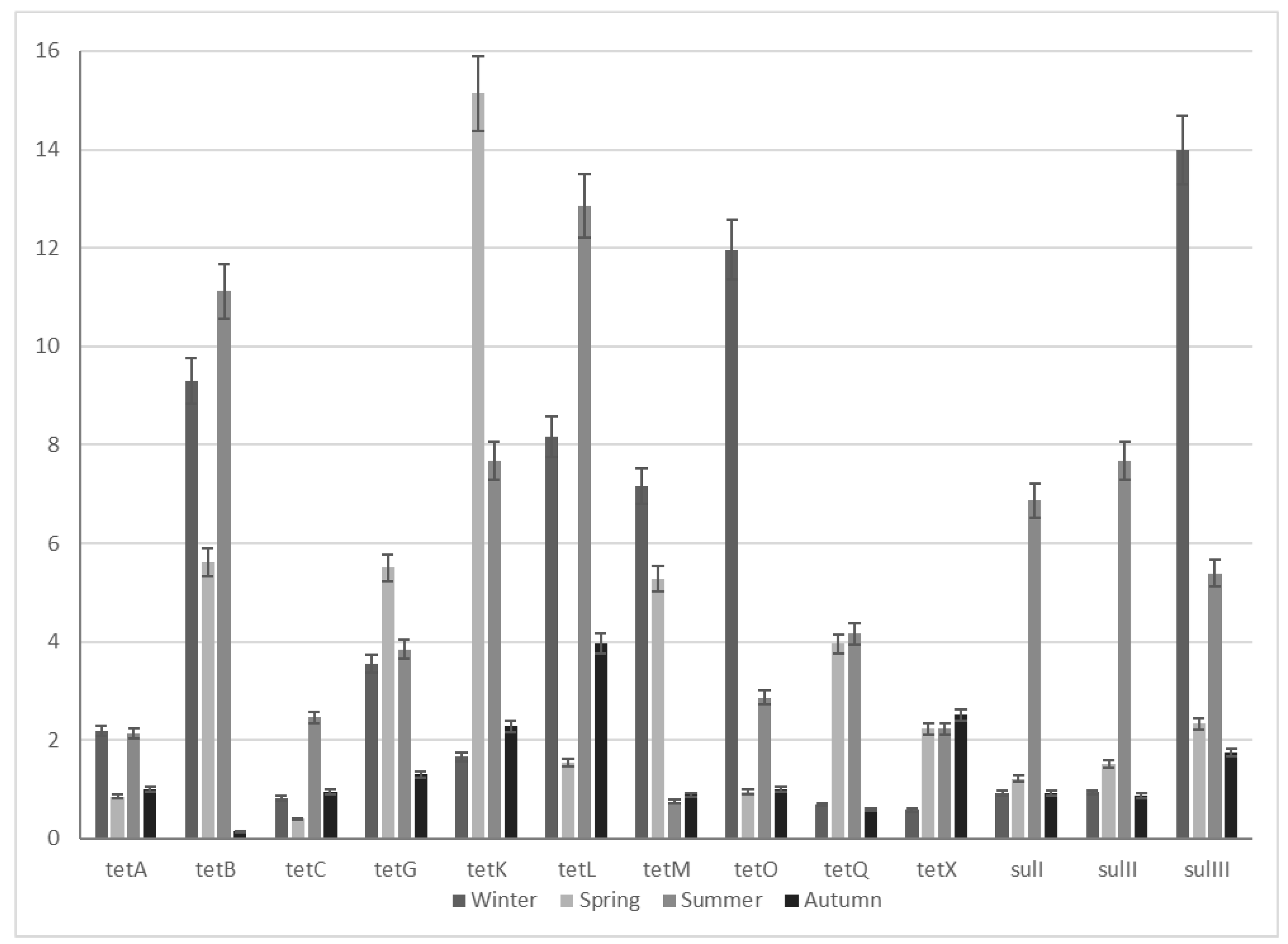
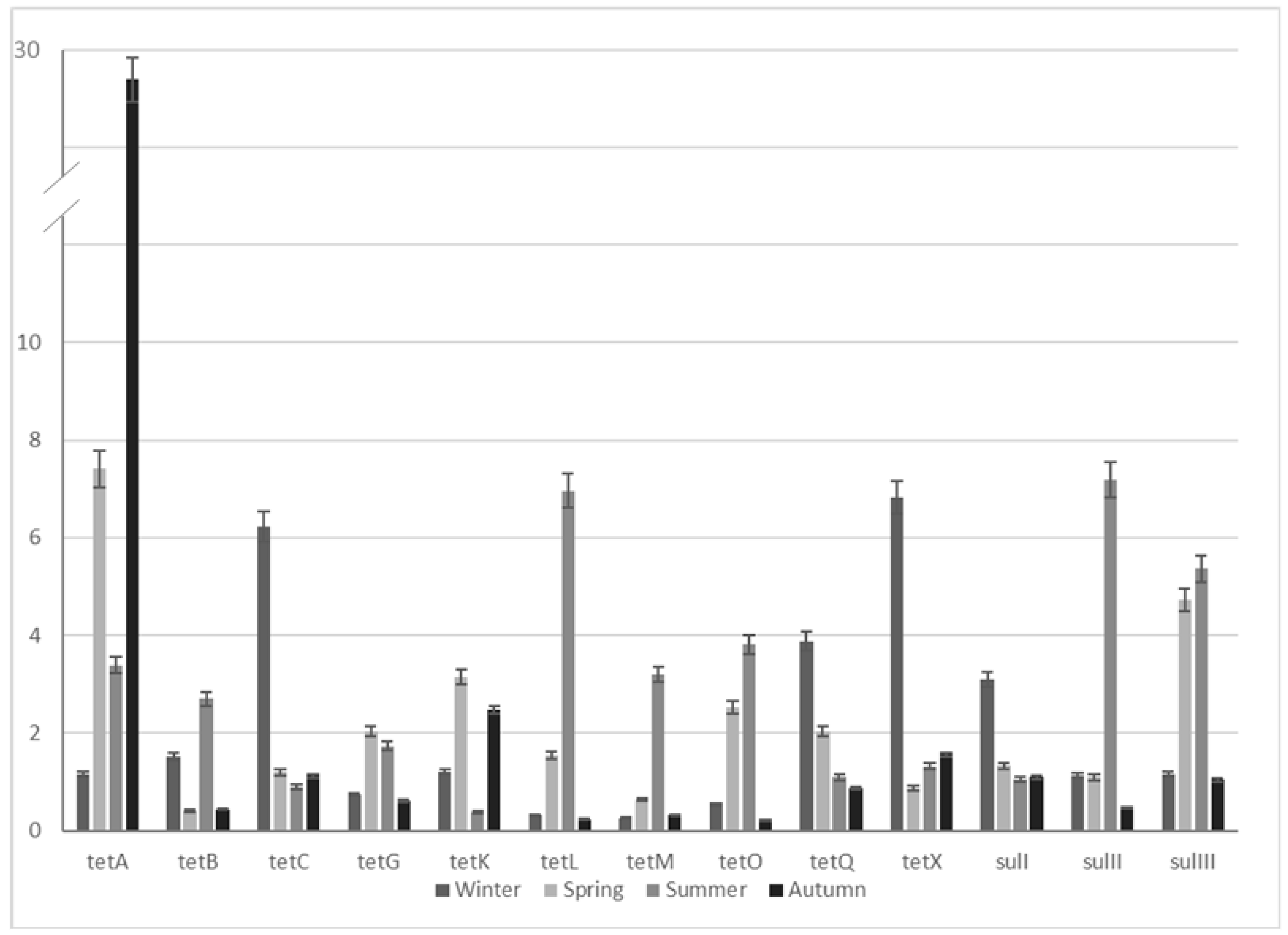
2.3. Seasonal ARG Changes
3. Materials and Methods
3.1. Characterization of Wastewater Treatment Plants
3.2. Samples Collection and Preparation
3.3. Selection of the Optimal DNA Isolation and Purification Method
3.4. Isolation and Purification of Total DNA in Influent and Effluent Samples
3.5. Quantitative PCR
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Alexander, J.; Bollmann, A.; Seitz, W.; Schwartz, T. Microbiological characterization of aquatic microbiomes targeting taxonomical marker genes and antibiotic resistance genes of opportunistic bacteria. Sci. Total. Environ. 2015, 512, 316–325. [Google Scholar] [CrossRef]
- Cassini, A.; Högberg, L.D.; Plachouras, D.; Quattrocchi, A.; Hoxha, A.; Simonsen, G.S.; Colomb-Cotinat, M.; Kretzschmar, M.E.; Devleesschauwer, B.; Cecchini, M.; et al. Faculty Opinions recommendation of Attributable deaths and disability-adjusted life-years caused by infections with antibiotic-resistant bacteria in the EU and the European Economic Area in 2015: A population-level modelling analysis. Fac. Opin. Post-Publ. Peer Rev. Biomed. Lit. 2019, 19, 56–66. [Google Scholar] [CrossRef]
- ECDC/EMEA. Technical Report. The Bacterial Challenge: Time to React; ECDC/EMEA: Stockholm, Sweden, 2009. [Google Scholar]
- Thorpe, K.E.; Joski, P.; Johnston, K.J. Antibiotic-Resistant Infection Treatment Costs Have Doubled Since 2002, Now Exceeding $2 Billion Annually. Health Aff. 2018, 37, 662–669. [Google Scholar] [CrossRef]
- Jimenez-Tototzintle, M.; Ferreira, I.J.; Duque, S.; Barrocas, P.R.G.; Saggioro, E.M. Removal of contaminants of emerging concern (CECs) and antibiotic resistant bacteria in urban wastewater using UVA/TiO2/H2O2 photocatalysis. Chemosphere 2018, 210, 449–457. [Google Scholar] [CrossRef]
- Gothwal, R.; Shashidhar, T. Antibiotic Pollution in the Environment: A Review. Clean Soil Air Water 2014, 43, 479–489. [Google Scholar] [CrossRef]
- Gullberg, E.; Cao, S.; Berg, O.G.; Ilbäck, C.; Sandegren, L.; Hughes, D.; Andersson, D.I. Selection of Resistant Bacteria at Very Low Antibiotic Concentrations. PLoS Pathog. 2011, 7, e1002158. [Google Scholar] [CrossRef] [PubMed]
- WHO. Antimicrobial Resistance: Global Report on Surveillance; WHO: Geneva, Switzerland, 2014. [Google Scholar]
- Wałaszek, M.; Różańska, A.; Bulanda, M.; Wojkowska-Mach, J. Alarming results of nosocomial bloodstream infections surveillance in Polish intensive care units. Przeglad Epidemiol. 2018, 72, 33–44. [Google Scholar]
- ECDC. European Surveillance of Antimicrobial Consumption Network (ESACNet). Available online: http://www.ecdc.europa.eu/en/activities/surveillance/ESAC-Net/Pages/index.aspx (accessed on 3 June 2020).
- ECDC. Quality Indicators for Antibiotic Consumption in the Community in Europe. Available online: http://ecdc.europa.eu/en/healthtopics/antimicrobial-resistance-and-consumption/antimicrobial_resistance/EARS-Net/Pages/EARS-Net.aspx (accessed on 3 June 2020).
- WHO. Antimicrobial Medicines Consumption (AMC) Network. 2017. Available online: http://www.euro.who.int/en/publications/abstracts/antimicrobial-medicines-consumption-amc-network.-amc-data-20112014-2017 (accessed on 3 June 2020).
- Wojkowska-Mach, J.; Godman, B.; Glassman, A.; Kurdi, A.; Pilc, A.; Różańska, A.; Skoczyński, S.; Wałaszek, M.; Bochenek, T. Antibiotic consumption and antimicrobial resistance in Poland; findings and implications. Antimicrob. Resist. Infect. Control. 2018, 7, 136. [Google Scholar] [CrossRef]
- ECDC. Antimicrobial Consumption—Annual Epidemiological Report for 2015; ECDC: Stockholm, Sweden, 2018. [Google Scholar]
- Pomorska-Wesołowska, M.; Różańska, A.; Sobońska, J.; Gryglewska, B.; Szczypta, A.; Dzikowska, M.; Chmielarczyk, A.; Wojkowska-Mach, J. Longevity and gender as the risk factors of methicillin-resistant Staphylococcus aureus infections in southern Poland. BMC Geriatr. 2017, 17, 51. [Google Scholar] [CrossRef] [PubMed]
- Chmielarczyk, A.; Pilarczyk-Zurek, M.; Kaminska, W.; Pobiega, M.; Romaniszyn, D.; Ziółkowski, G.; Wojkowska-Mach, J.; Bulanda, M. Molecular Epidemiology and Drug Resistance of Acinetobacter baumannii Isolated from Hospitals in Southern Poland: ICU as a Risk Factor for XDR Strains. Microb. Drug Resist. 2016, 22, 328–335. [Google Scholar] [CrossRef] [PubMed]
- Chmielarczyk, A.; Pomorska-Wesołowska, M.; Szczypta, A.; Romaniszyn, D.; Pobiega, M.; Wojkowska-Mach, J.; Agnieszka, C.; Monika, P.-W.; Anna, S.; Dorota, R.; et al. Molecular analysis of meticillin-resistant Staphylococcus aureus strains isolated from different types of infections from patients hospitalized in 12 regional, non-teaching hospitals in southern Poland. J. Hosp. Infect. 2017, 95, 259–267. [Google Scholar] [CrossRef]
- Romaniszyn, D.; Różańska, A.; Chmielarczyk, A.; Pobiega, M.; Adamski, P.; Helwich, E.; Lauterbach, R.; Borszewska-Kornacka, M.; Gulczyńska, E.; Kordek, A.; et al. Epidemiology, antibiotic consumption and molecular characterisation of Staphylococcus aureus infections—Data from the polish neonatology surveillance network, 2009–2012. BMC Infect Dis. 2015, 15, 1–9. [Google Scholar] [CrossRef] [PubMed]
- LaPara, T.M.; Burch, T.R.; McNamara, P.J.; Tan, D.T.; Yan, M.; Eichmiller, J.J. Tertiary-Treated Municipal Wastewater is a Significant Point Source of Antibiotic Resistance Genes into Duluth-Superior Harbor. Environ. Sci. Technol. 2011, 45, 9543–9549. [Google Scholar] [CrossRef] [PubMed]
- Pruden, A.; Larsson, D.G.J.; Amézquita, A.; Collignon, P.J.; Brandt, K.K.; Graham, D.W.; Lazorchak, J.M.; Suzuki, S.; Silley, P.; Snape, J.R.; et al. Management Options for Reducing the Release of Antibiotics and Antibiotic Resistance Genes to the Environment. Environ. Health Perspect. 2013, 121, 878–885. [Google Scholar] [CrossRef]
- Gulkowska, A.; Leung, H.; So, M.; Taniyasu, S.; Yamashita, N.; Yeung, L.W.; Richardson, B.J.; Lei, A.; Giesy, J.; Lam, P.K. Removal of antibiotics from wastewater by sewage treatment facilities in Hong Kong and Shenzhen, China. Water Res. 2008, 42, 395–403. [Google Scholar] [CrossRef]
- Minh, T.B.; Leung, H.W.; Loi, I.H.; Chan, W.H.; So, M.K.; Mao, J.; Choi, D.; Lam, J.C.W.; Zheng, G.; Martin, M.; et al. Antibiotics in the Hong Kong metropolitan area: Ubiquitous distribution and fate in Victoria Harbour. Mar. Pollut. Bull. 2009, 58, 1052–1062. [Google Scholar] [CrossRef]
- Peng, X.; Tan, J.; Tang, C.; Yu, Y.; Wang, Z. Multiresidue Determination of Fluoroquinolone, Sulfonamide, Trimethoprim, and Chloramphenicol Antibiotics in Urban Waters in China. Environ. Toxicol. Chem. 2008, 27, 73. [Google Scholar] [CrossRef]
- Batt, A.L.; Bruce, I.B.; Aga, D.S. Evaluating the vulnerability of surface waters to antibiotic contamination from varying wastewater treatment plant discharges. Environ. Pollut. 2006, 142, 295–302. [Google Scholar] [CrossRef] [PubMed]
- Łuczkiewicz, A.; Felis, E.; Ziembińska, A.; Gnida, A.; Kotlarska, E.; Olanczuk-Neyman, K.; Surmacz-Górska, J. Resistance of Escherichia coli and Enterococcus spp. to selected antimicrobial agents present in municipal wastewater. J. Water Health 2013, 11, 600–612. [Google Scholar] [CrossRef] [PubMed]
- Birosova, L.; Mackuľak, T.; Bodík, I.; Ryba, J.; Škubák, J.; Grabic, R. Pilot study of seasonal occurrence and distribution of antibiotics and drug resistant bacteria in wastewater treatment plants in Slovakia. Sci. Total. Environ. 2014, 490, 440–444. [Google Scholar] [CrossRef]
- Rizzo, L.; Manaia, C.M.; Merlin, C.; Schwartz, T.; Dagot, C.; Ploy, M.; Michael, I.; Fatta-Kassinos, D. Urban wastewater treatment plants as hotspots for antibiotic resistant bacteria and genes spread into the environment: A review. Sci. Total. Environ. 2013, 447, 345–360. [Google Scholar] [CrossRef] [PubMed]
- Storteboom, H.; Arabi, M.; Davis, J.G.; Crimi, B.; Pruden, A. Tracking Antibiotic Resistance Genes in the South Platte River Basin Using Molecular Signatures of Urban, Agricultural, And Pristine Sources. Environ. Sci. Technol. 2010, 44, 7397–7404. [Google Scholar] [CrossRef] [PubMed]
- Makowska, N.; Koczura, R.; Mokracka, J. Class 1 integrase, sulfonamide and tetracycline resistance genes in wastewater treatment plant and surface water. Chemosphere 2016, 144, 1665–1673. [Google Scholar] [CrossRef]
- Bengtsson-Palme, J.; Hammarén, R.; Pal, C.; Östman, M.; Björlenius, B.; Flach, C.-F.; Fick, J.; Kristiansson, E.; Tysklind, M.; Larsson, D.G.J. Elucidating selection processes for antibiotic resistance in sewage treatment plants using metagenomics. Sci. Total. Environ. 2016, 572, 697–712. [Google Scholar] [CrossRef] [PubMed]
- Szczepanowski, R.; Linke, B.; Krahn, I.; Gartemann, K.-H.; Gützkow, T.; Eichler, W.; Pühler, A.; Schlüter, A. Detection of 140 clinically relevant antibiotic-resistance genes in the plasmid metagenome of wastewater treatment plant bacteria showing reduced susceptibility to selected antibiotics. Microbiology 2009, 155, 2306–2319. [Google Scholar] [CrossRef]
- Pruden, A.; Arabi, M.; Storteboom, H.N. Correlation Between Upstream Human Activities and Riverine Antibiotic Resistance Genes. Environ. Sci. Technol. 2012, 46, 11541–11549. [Google Scholar] [CrossRef]
- Pazda, M.; Kumirska, J.; Stepnowski, P.; Mulkiewicz, E. Antibiotic resistance genes identified in wastewater treatment plant systems—A review. Sci. Total Environ. 2019, 697, 1–21. [Google Scholar] [CrossRef]
- Sharma, V.K.; Johnson, N.; Cizmas, L.; McDonald, T.J.; Kim, H. A review of the influence of treatment strategies on antibiotic resistant bacteria and antibiotic resistance genes. Chemosphere 2016, 150, 702–714. [Google Scholar] [CrossRef]
- Li, Y.; Zhu, G.; Ng, W.J.; Tan, S.K. A review on removing pharmaceutical contaminants from wastewater by constructed wetlands: Design, performance and mechanism. Sci. Total. Environ. 2014, 468, 908–932. [Google Scholar] [CrossRef]
- Chen, J.; Ying, G.-G.; Wei, X.-D.; Liu, Y.-S.; Liu, S.-S.; Hu, L.-X.; He, L.-Y.; Chen, Z.-F.; Chen, F.-R.; Yang, Y.-Q. Removal of antibiotics and antibiotic resistance genes from domestic sewage by constructed wetlands: Effect of flow configuration and plant species. Sci. Total. Environ. 2016, 571, 974–982. [Google Scholar] [CrossRef]
- Chen, J.; Wei, X.-D.; Liu, Y.-S.; Ying, G.-G.; Liu, S.-S.; He, L.-Y.; Su, H.-C.; Hu, L.-X.; Chen, F.-R.; Yang, Y.-Q. Removal of antibiotics and antibiotic resistance genes from domestic sewage by constructed wetlands: Optimization of wetland substrates and hydraulic loading. Sci. Total. Environ. 2016, 565, 240–248. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Zheng, J.; Liu, C.; Liu, L.; Liu, Y.; Fan, H. Removal of antibiotics and resistance genes from swine wastewater using vertical flow constructed wetlands: Effect of hydraulic flow direction and substrate type. Chem. Eng. J. 2017, 308, 692–699. [Google Scholar] [CrossRef]
- Gorito, A.M.; Ribeiro, A.R.; Almeida, C.M.R.; Silva, A.M. A review on the application of constructed wetlands for the removal of priority substances and contaminants of emerging concern listed in recently launched EU legislation. Environ. Pollut. 2017, 227, 428–443. [Google Scholar] [CrossRef]
- Liu, L.; Li, J.; Fan, H.; Huang, X.; Wei, L.; Liu, C. Fate of antibiotics from swine wastewater in constructed wetlands with different flow configurations. Int. Biodeterior. Biodegrad. 2019, 140, 119–125. [Google Scholar] [CrossRef]
- Zhang, S.; Lu, Y.-X.; Zhang, J.-J.; Liu, S.; Song, H.-L.; Yang, X.-L. Constructed Wetland Revealed Efficient Sulfamethoxazole Removal but Enhanced the Spread of Antibiotic Resistance Genes. Molecules 2020, 25, 834. [Google Scholar] [CrossRef] [PubMed]
- Aminov, R.; Garrigues-Jeanjean, N.; Mackie, R.I. Molecular Ecology of Tetracycline Resistance: Development and Validation of Primers for Detection of Tetracycline Resistance Genes Encoding Ribosomal Protection Proteins. Appl. Environ. Microbiol. 2001, 67, 22–32. [Google Scholar] [CrossRef]
- Wang, N.; Yang, X.; Jiao, S.; Zhang, J.; Ye, B.; Gao, S. Sulfonamide-Resistant Bacteria and Their Resistance Genes in Soils Fertilized with Manures from Jiangsu Province, Southeastern China. PLoS ONE 2014, 9, e112626. [Google Scholar] [CrossRef]
- Tuckman, M.; Petersen, P.J.; Howe, A.Y.M.; Orlowski, M.; Mullen, S.; Chan, K.; Bradford, P.A.; Jones, C.H. Occurrence of Tetracycline Resistance Genes among Escherichia coli Isolates from the Phase 3 Clinical Trials for Tigecycline. Antimicrob. Agents Chemother. 2007, 51, 3205–3211. [Google Scholar] [CrossRef]
- Du, J.; Geng, J.; Ren, H.; Ding, L.; Xu, K.; Zhang, Y. Variation of antibiotic resistance genes in municipal wastewater treatment plant with A2O-MBR system. Environ. Sci. Pollut. Res. 2014, 22, 3715–3726. [Google Scholar] [CrossRef]
- Zhang, T.; Shao, M.-F.; Ye, L. 454 Pyrosequencing reveals bacterial diversity of activated sludge from 14 sewage treatment plants. ISME J. 2011, 6, 1137–1147. [Google Scholar] [CrossRef]
- Park, K.Y.; Jang, H.M.; Park, M.-R.; Lee, K.; Kim, D.; Kim, Y.-M. Combination of different substrates to improve anaerobic digestion of sewage sludge in a wastewater treatment plant. Int. Biodeterior. Biodegrad. 2016, 109, 73–77. [Google Scholar] [CrossRef]
- Kim, Y.B.; Jeon, J.H.; Choi, S.; Shin, J.; Lee, Y.; Kim, Y.-M. Use of a filtering process to remove solid waste and antibiotic resistance genes from effluent of a flow-through fish farm. Sci. Total. Environ. 2018, 615, 289–296. [Google Scholar] [CrossRef] [PubMed]
- Zielińska, S.; Radkowski, P.; Blendowska, A.; Ludwig-Gałęzowska, A.; Łoś, J.M.; Łoś, M. The choice of the DNA extraction method may influence the outcome of the soil microbial community structure analysis. Microbiology 2017, 6, e00453. [Google Scholar] [CrossRef] [PubMed]
- Morgan, J.L.; Darling, A.E.; Eisen, J.A. Metagenomic Sequencing of an In Vitro-Simulated Microbial Community. PLoS ONE 2010, 5, e10209. [Google Scholar] [CrossRef]
- Biswal, B.K.; Mazza, A.; Masson, L.; Gehr, R.; Frigon, D. Impact of wastewater treatment processes on antimicrobial resistance genes and their co-occurrence with virulence genes in Escherichia coli. Water Res. 2014, 50, 245–253. [Google Scholar] [CrossRef]
- Rodriguez-Mozaz, S.; Chamorro, S.; Marti, E.; Huerta, B.; Gros, M.; Sànchez-Melsió, A.; Borrego, C.M.; Barceló, J.; Balcázar, J.L. Occurrence of antibiotics and antibiotic resistance genes in hospital and urban wastewaters and their impact on the receiving river. Water Res. 2015, 69, 234–242. [Google Scholar] [CrossRef] [PubMed]
- Amador, P.P.; Fernandes, R.M.; Prudencio, M.C.; Barreto, M.P.; Duarte, I.M. Antibiotic resistance in wastewater: Occurrence and fate of Enterobacteriaceae producers of class A and class C β-lactamases. J. Environ. Sci. Health A Tox. Hazard. Subst. Environ. Eng. 2015, 50, 26–39. [Google Scholar] [CrossRef]
- Auerbach, E.A.; Seyfried, E.E.; McMahon, K.D. Tetracycline resistance genes in activated sludge wastewater treatment plants. Water Res. 2007, 41, 1143–1151. [Google Scholar] [CrossRef]
- Guo, J.; Li, J.; Chen, H.; Bond, P.L.; Yuan, Z. Metagenomic analysis reveals wastewater treatment plants as hotspots of antibiotic resistance genes and mobile genetic elements. Water Res. 2017, 123, 468–478. [Google Scholar] [CrossRef]
- Laht, M.; Karkman, A.; Voolaid, V.; Ritz, C.; Tenson, T.; Virta, M.; Kisand, V. Abundances of Tetracycline, Sulphonamide and Beta-Lactam Antibiotic Resistance Genes in Conventional Wastewater Treatment Plants (WWTPs) with Different Waste Load. PLoS ONE 2014, 9, e103705. [Google Scholar] [CrossRef]
- Hultman, J.; Tamminen, M.; Pärnänen, K.M.M.; Cairns†, J.; Karkman, A.; Virta, M. Host range of antibiotic resistance genes in wastewater treatment plant influent and effluent. FEMS Microbiol. Ecol. 2018, 94, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Łuczkiewicz, A.; Jankowska, K.; Fudala-Ksiązek, S.; Olańczuk-Neyman, K. Antimicrobial resistance of fecal indicators in municipal wastewater treatment plant. Water Res. 2010, 44, 5089–5097. [Google Scholar] [CrossRef] [PubMed]
- Mao, D.; Yu, S.; Rysz, M.; Luo, Y.; Yang, F.; Li, F.; Hou, J.; Mu, Q.; Alvarez, P.J.J. Prevalence and proliferation of antibiotic resistance genes in two municipal wastewater treatment plants. Water Res. 2015, 85, 458–466. [Google Scholar] [CrossRef] [PubMed]
- Kotlarska, E.; Łuczkiewicz, A.; Pisowacka, M.; Burzyński, A. Antibiotic Resistance and Prevalence of Class 1 and 2 Integrons in Escherichia coli Isolated from Two Wastewater Treatment Plants, and Their Receiving Waters (Gulf of Gdansk, Baltic Sea, Poland). Wastewater Public Health 2015, 22, 67–98. [Google Scholar] [CrossRef]
- Nõlvak, H.; Truu, M.; Tiirik, K.; Oopkaup, K.; Sildvee, T.; Kaasik, A.; Mander, Ü.; Truu, J. Dynamics of antibiotic resistance genes and their relationships with system treatment efficiency in a horizontal subsurface flow constructed wetland. Sci. Total. Environ. 2013, 461, 636–644. [Google Scholar] [CrossRef]
- Liu, L.; Liu, C.; Zheng, J.; Huang, X.; Wang, Z.; Liu, Y.; Zhu, G. Elimination of veterinary antibiotics and antibiotic resistance genes from swine wastewater in the vertical flow constructed wetlands. Chemosphere 2013, 91, 1088–1093. [Google Scholar] [CrossRef]
- Liu, L.; Liu, Y.-H.; Wang, Z.; Liu, C.-X.; Huang, X.; Zhu, G.-F. Behavior of tetracycline and sulfamethazine with corresponding resistance genes from swine wastewater in pilot-scale constructed wetlands. J. Hazard. Mater. 2014, 278, 304–310. [Google Scholar] [CrossRef]
- Huang, X.; Liu, C.; Li, K.; Su, J.; Zhu, G.; Liu, L. Performance of vertical up-flow constructed wetlands on swine wastewater containing tetracyclines and tert genes. Water Res. 2015, 70, 109–117. [Google Scholar] [CrossRef]
- Chen, J.; Liu, Y.S.; Su, H.C.; Ying, G.G.; Liu, F.; Liu, S.S.; He, L.Y.; Chen, Z.F.; Yang, Y.Q.; Chen, F.R. Removal of antibiotics and antibiotic resistance henes in rural wastewater by an integrated constructed wetland. Environ. Sci. Pollut. Res. 2015, 22, 1794–1803. [Google Scholar] [CrossRef]
- Wolecki, D.; Caban, M.; Pazda, M.; Stepnowski, P.; Kumirska, J. Evaluation of the Possibility of Using Hydroponic Cultivations for the Removal of Pharmaceuticals and Endocrine Disrupting Compounds in Municipal Sewage Treatment Plants. Molecules 2019, 25, 162. [Google Scholar] [CrossRef]
- Nowrotek, M.; Kotlarska, E.; Łuczkiewicz, A.; Felis, E.; Sochacki, A.; Miksch, K. The treatment of wastewater containing pharmaceuticals in microcosm constructed wetlands: The occurrence of integrons (int1–2) and associated resistance genes (sul1–3, qacEΔ1). Environ. Sci. Pollut. Res. 2017, 24, 15055–15066. [Google Scholar] [CrossRef] [PubMed]
- Anderson, J.C.; Carlson, J.C.; Low, J.E.; Challis, J.K.; Wong, C.S.; Knapp, C.W.; Hanson, M. Performance of a constructed wetland in Grand Marais, Manitoba, Canada: Removal of nutrients, pharmaceuticals, and antibiotic resistance genes from municipal wastewater. Chem. Central J. 2013, 7, 54. [Google Scholar] [CrossRef] [PubMed]
- Song, H.L.; Zhang, S.; Guo, J.; Yang, Y.L.; Zhang, L.M.; Li, H.; Yang, X.L.; Liu, X. Vertical up-flow constructed wetlands exhibited efficient antibioticremoval but induced antibiotic resistance genes in effluent. Chemosphere 2018, 203, 434–441. [Google Scholar] [CrossRef] [PubMed]
- Goossens, H.; Ferech, M.; Vander Stichele, R. Outpatient antibiotic use in Europe and association with resistance: A crossnational database study. Lancet 2005, 365, 579–587. [Google Scholar] [CrossRef]
- Sun, L.; Klein, E.Y.; Laxminarayan, R. Seasonality and Temporal Correlation between Community Antibiotic Use and Resistance in the United States. Clin. Infect. Dis. 2012, 55, 687–694. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Q.-B.; Guo, M.-T.; Yang, J. Monitoring and assessing the impact of wastewater treatment on release of both antibiotic-resistant bacteria and their typical genes in a Chinese municipal wastewater treatment plant. Environ. Sci. Process. Impacts 2014, 16, 1930–1937. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Li, B.; Ju, F.; Zhang, T. Exploring Variation of Antibiotic Resistance Genes in Activated Sludge over a Four-Year Period through a Metagenomic Approach. Environ. Sci. Technol. 2013, 47, 10197–10205. [Google Scholar] [CrossRef]
- Börjesson, S.; Melin, S.; Matussek, A.; Lindgren, P.-E. A seasonal study of the mecA gene and Staphylococcus aureus including methicillin-resistant S. aureus in a municipal wastewater treatment plant. Water Res. 2009, 43, 925–932. [Google Scholar] [CrossRef]
- Börjesson, S.; Mattsson, A.; Lindgren, P.-E. Genes encoding tetracycline resistance in a full-scale municipal wastewater treatment plant investigated during one year. J. Water Health 2009, 8, 247–256. [Google Scholar] [CrossRef]
- Olańczuk-Neyman, K.; Geneja, M.; Quant, B.; Dembińska, M.; Kruczalak, K.; Kulbat, E.; Kulik-Kuziemska, I.; Mikołajski, S.; Gielert, M. Microbiological and Biological Aspects of the Wastewater Treatment Plant “Wschód” in Gdańsk. Pol. J. Environ. Stud. 2003, 12, 747–757. [Google Scholar]
- Suzuki, M.T.; Taylor, L.T.; Delong, E.F.; Burnett, S.L.; Chen, J.; Beuchat, L.R. Quantitative Analysis of Small-Subunit rRNA Genes in Mixed Microbial Populations via 5′-Nuclease Assays. Appl. Environ. Microbiol. 2000, 66, 4605–4614. [Google Scholar] [CrossRef] [PubMed]
- Ng, L.-K.; Martin, I.; Alfa, M.; Mulvey, M. Multiplex PCR for the detection of tetracycline resistant genes. Mol. Cell. Probes 2001, 15, 209–215. [Google Scholar] [CrossRef] [PubMed]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of the compounds are not available from the authors. |
| No. | DNA Extraction Kit | µg of Total DNA per 1 g of AS | A260/280 | A260/230 |
|---|---|---|---|---|
| 1 | Genomic Mini AX Stool (A&A Biotechnology, Gdynia, Poland) | 117.5 ± 0.3 | 1.90 ± 0.02 | 1.77 ± 0.03 |
| 2 | Genomic Mini AX Bacteria + (A&A Biotechnology, Gdynia, Poland) | 274.6 ± 1.6 | 1.92 ± 0.01 | 1.78 ± 0.04 |
| 3 | Exgene Soil DNA mini (GeneAll Biotechnology, Seoul, Korea) | 365.4 ± 4.5 | 2.00 ± 0.01 | 1.31 ± 0.62 |
| 4 | FastDNA SPIN Kit for Soil (MP Biomedicals, Solon, OH, USA) | 751.8 ± 3.0 | 1.91 ± 0.11 | 0.55 ± 0.01 |
| 5 | NucleoSpin Soil lysis buffer 1 (Macherey–Nagel, Düren, Germany) | 297.9 ± 16.9 | 1.98 ± 0.01 | 1.93 ± 0.08 |
| 6 | NucleoSpin Soil lysis buffer 2 (Macherey–Nagel, Düren, Germany) | 306.2 ± 9.8 | 1.97 ± 0.01 | 1.97 ± 0.04 |
| 7 | PowerSoil DNA Isolation Kit (Qiagen, Hilden, Germany) | 21.4 ± 1.4 | 1.76 ± 0.22 | 1.02 ± 0.31 |
| 8 | ZymoBIOMICS DNA Minikit (Zymo Research, Irvine, CA, USA) | 247.9 ± 0.5 | 1.89 ± 0.02 | 1.81 ± 0.02 |
| 9 | GeneMATRIX SOIL DNA Purification Kit (EURx, Gdańsk, Poland) | 274.6 ± 1.6 | 1.92 ± 0.01 | 1.78 ± 0.04 |
| 10 | GeneMATRIX Environmental DNA & RNA Purification Kit (EURx, Gdańsk, Poland) | 213.5 ± 0.8 | 2.18 ± 0.01 | 2.15 ± 0.01 |
| ARGs | ARG Enrichment | |||
|---|---|---|---|---|
| Winter | Spring | Summer | Autumn | |
| tetA | 2.2 | 0.9 | 2.1 | 1.0 |
| tetB | 9.3 | 5.6 | 11.1 | 0.2 |
| tetC | 0.8 | 0.4 | 2.5 | 1.0 |
| tetG | 3.6 | 5.5 | 3.8 | 1.3 |
| tetK | 1.7 | 15.1 | 7.7 | 2.3 |
| tetL | 8.2 | 1.6 | 12.9 | 4.0 |
| tetM | 7.2 | 5.3 | 0.8 | 0.9 |
| tetO | 12.0 | 1.0 | 2.9 | 1.0 |
| tetQ | 0.7 | 4.0 | 4.2 | 0.6 |
| tetX | 0.6 | 2.2 | 2.2 | 2.5 |
| sulI | 0.9 | 1.2 | 6.9 | 0.9 |
| sulII | 0.9 | 1.5 | 7.7 | 0.9 |
| sulIII | 14.0 | 2.3 | 5.4 | 1.8 |
| ARGs | ARG Enrichment | |||
|---|---|---|---|---|
| Winter | Spring | Summer | Autumn | |
| tetA | 1.2 | 7.4 | 3.4 | 28.8 |
| tetB | 1.5 | 0.4 | 2.7 | 0.4 |
| tetC | 6.2 | 1.2 | 0.9 | 1.1 |
| tetG | 0.7 | 2.0 | 1.7 | 0.6 |
| tetK | 1.2 | 3.1 | 0.4 | 2.5 |
| tetL | 0.3 | 1.6 | 7.0 | 0.2 |
| tetM | 0.3 | 0.6 | 3.2 | 0.3 |
| tetO | 0.5 | 2.5 | 3.8 | 0.2 |
| tetQ | 3.9 | 2.0 | 1.1 | 0.9 |
| tetX | 6.8 | 0.9 | 1.3 | 1.6 |
| sulI | 3.1 | 1.3 | 1.0 | 1.1 |
| sulII | 1.1 | 1.1 | 7.2 | 0.5 |
| sulIII | 1.2 | 4.7 | 5.4 | 1.0 |
| Parameter | Unit | WWTP1 | WWTP2 | ||
|---|---|---|---|---|---|
| Influent | Effluent | Influent | Effluent | ||
| BOD5 | mgO2/L | 446.0 | 3.7 | 449.0 | 2.6 |
| COD | mgO2/L | 1009.0 | 35.0 | 1066.0 | 32.5 |
| TSS | mg/L | 516.0 | 5.0 | 501.0 | 6.1 |
| TP | mg/L | 10.8 | 0.4 | 10.7 | 0.2 |
| TN | mg/L | 87.0 | 8.0 | 87.7 | 7.9 |
| Target Gene | Primer Sequence (5′-3′) | Amplicon Size (bp) | Source | |
|---|---|---|---|---|
| 16S rRNA | F | CGG TGA ATA CGT TCY CGG | 146 | [77] |
| R | GGW TAC CTT GTT ACG ACTT | |||
| tetA | F | GCT ACA TCC TGC TTG CCT TC | 210 | [78] |
| R | CAT AGA TCG CCG TGA AGA GG | |||
| tetB | F | TAC GTG AAT TTA TTG CTT CGG | 206 | [42] |
| R | ATA CAG CAT CCA AAG CGC AC | |||
| tetC | F | CTT GAG AGC CTT CAA CCC AG | 418 | [78] |
| R | ATG GTC GTC ATC TAC CTG CC | |||
| tetG | F | GCT CGG TGG TAT CTC TGC TC | 468 | [78] |
| R | AGC AAC AGA ATC GGG AAC AC | |||
| tetK | F | TCG ATA GGA ACA GCA GTA | 169 | [78] |
| R | CAG CAG ATC CTA CTC CTT | |||
| tetL | F | TCG TTA GCG TGC TGT CAT TC | 267 | [78] |
| R | GTA TCC CAC CAA TGT AGC CG | |||
| tetM | F | AAT AAA TCA TAA ACA GAA AGC TTA TTA TAT AAC | 171 | This study |
| R | AAT AAA TCA TAA TGG CGT GTC TAT GAT GTT CAC | |||
| tetO | F | AAC TTA GGC ATT CTG GCT CAC | 515 | [78] |
| R | TCC CAC TGT TCC ATA TCG TCA | |||
| tetQ | F | AGA ATC TGC TGT TTG CCA GTG | 169 | [42] |
| R | CGG AGT GTC AAT GAT ATT GCA | |||
| tetX | F | CAA TAA TTG GTG GTG GAC CC | 468 | [78] |
| R | TTC TTA CCT TGG ACA TCC CG | |||
| sulI | F | GAC GAG ATT GTG CGG TTC TT | 185 | [31] |
| R | GAG ACC AAT AGC GGA AGCC | |||
| sulII | F | GAC AGT TAT CAA CCC GCG AC | 147 | [31] |
| R | GTC TTG CAC CGA ATG CAT AA | |||
| sulIII | F | ACC ACC GAT AGT TTT TCC GA | 199 | [31] |
| R | TGC CTTT TTC TTT TAA AGCC | |||
| Reaction Stage | tet(B, K, L, M, Q, X), sulIII | tet(G, C), sulI, sulII | tet(A, O) | |||
|---|---|---|---|---|---|---|
| Pre-incubation | 95 °C, 5 min (4.4 °C/s) | 95 °C, 5 min (4.4 °C/s) | 95 °C, 5 min (4.4 °C/s) | |||
| Amplification | 95 °C, 10 s (4.4 °C/s) | 45 cycles | 95 °C, 10 s (4.4 °C/s) | 45 cycles | 95 °C, 10 s (4.4 °C/s) | 55 cycles |
| 62 °C, 30 s (2.2 °C/s) | 60 °C, 30 s (2.2 °C/s) | 60 °C, 30 s (2.2 °C/s) | ||||
| 72 °C, 30 s (4.4 °C/s) | 72 °C, 30 s (4.4 °C/s) | 72 °C, 30 s (4.4 °C/s) | ||||
| Melting curve | 95 °C, 5 s (4.4 °C/s) | 95 °C, 5 s (4.4 °C/s) | 95 °C, 5 s (4.4 °C/s) | |||
| 65 °C, 1 min (2.2 °C/s) | 65 °C, 1 min (2.2 °C/s) | 65 °C, 1 min (2.2 °C/s) | ||||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pazda, M.; Rybicka, M.; Stolte, S.; Piotr Bielawski, K.; Stepnowski, P.; Kumirska, J.; Wolecki, D.; Mulkiewicz, E. Identification of Selected Antibiotic Resistance Genes in Two Different Wastewater Treatment Plant Systems in Poland: A Preliminary Study. Molecules 2020, 25, 2851. https://doi.org/10.3390/molecules25122851
Pazda M, Rybicka M, Stolte S, Piotr Bielawski K, Stepnowski P, Kumirska J, Wolecki D, Mulkiewicz E. Identification of Selected Antibiotic Resistance Genes in Two Different Wastewater Treatment Plant Systems in Poland: A Preliminary Study. Molecules. 2020; 25(12):2851. https://doi.org/10.3390/molecules25122851
Chicago/Turabian StylePazda, Magdalena, Magda Rybicka, Stefan Stolte, Krzysztof Piotr Bielawski, Piotr Stepnowski, Jolanta Kumirska, Daniel Wolecki, and Ewa Mulkiewicz. 2020. "Identification of Selected Antibiotic Resistance Genes in Two Different Wastewater Treatment Plant Systems in Poland: A Preliminary Study" Molecules 25, no. 12: 2851. https://doi.org/10.3390/molecules25122851
APA StylePazda, M., Rybicka, M., Stolte, S., Piotr Bielawski, K., Stepnowski, P., Kumirska, J., Wolecki, D., & Mulkiewicz, E. (2020). Identification of Selected Antibiotic Resistance Genes in Two Different Wastewater Treatment Plant Systems in Poland: A Preliminary Study. Molecules, 25(12), 2851. https://doi.org/10.3390/molecules25122851








