Towards Understanding of Polymorphism of the G-rich Region of Human Papillomavirus Type 52
Abstract
:1. Introduction
2. Results
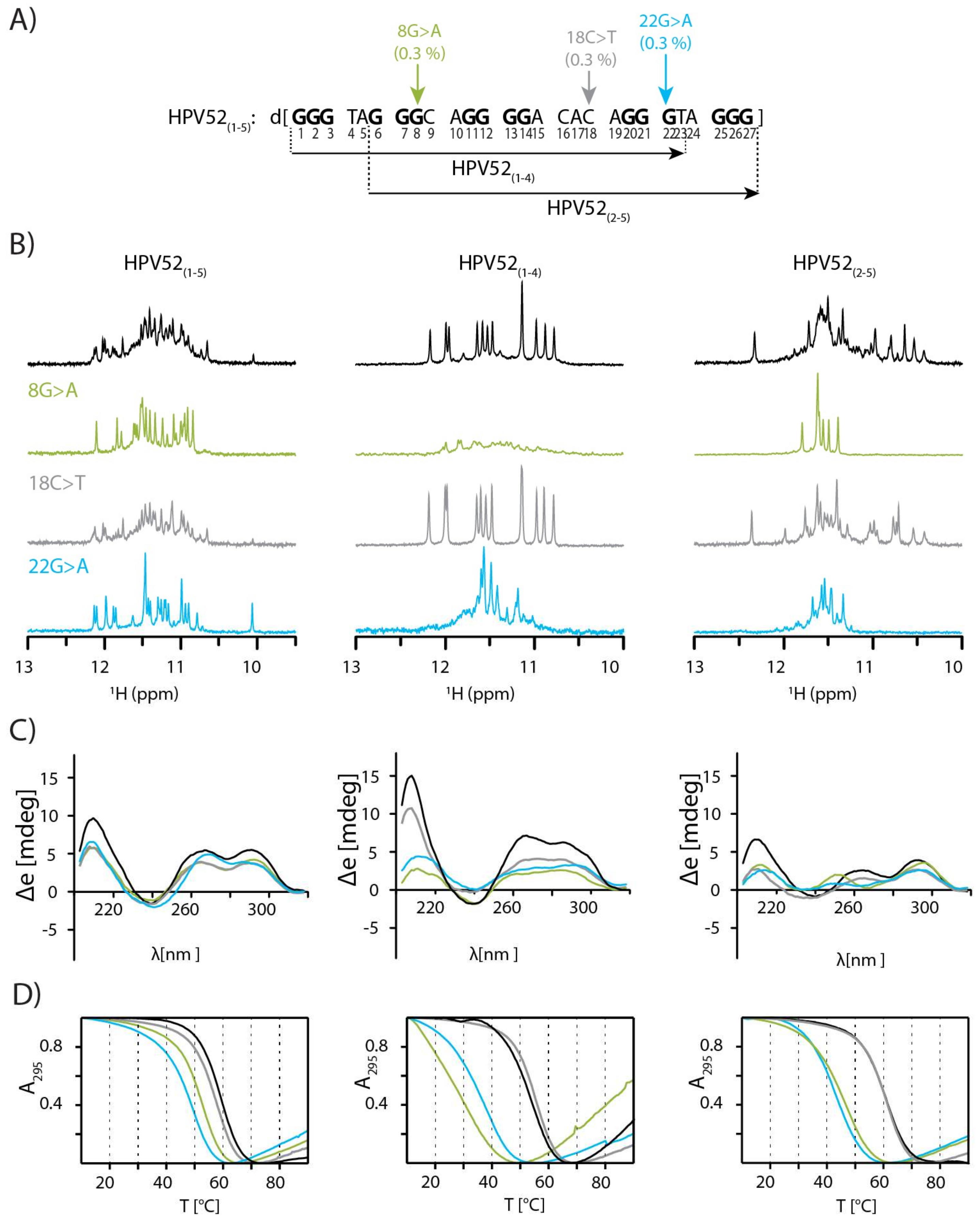
2.1. SNPs Reduce Polymorphism of HPV52(1–5) and Assist Identification of Predominant Species
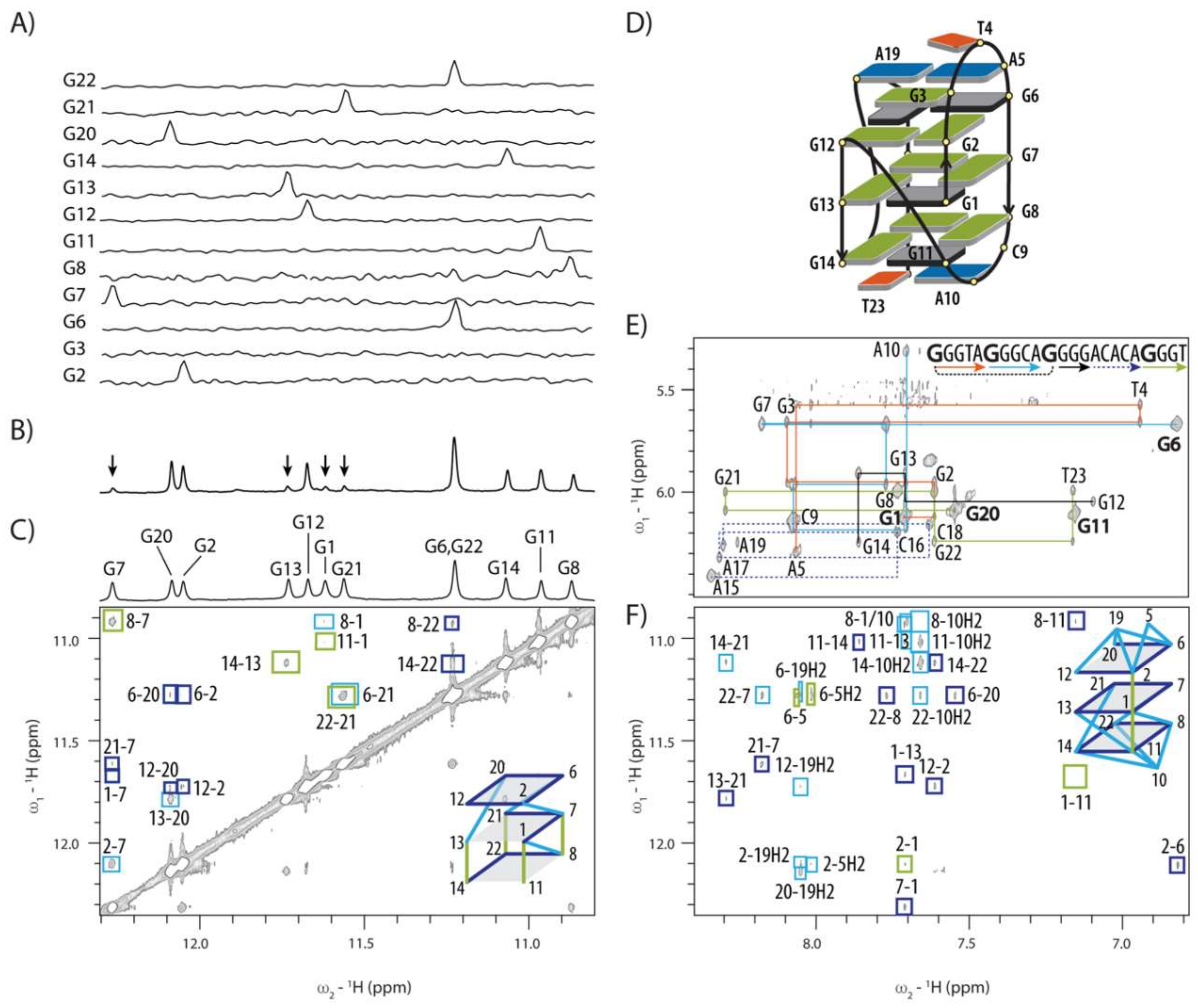
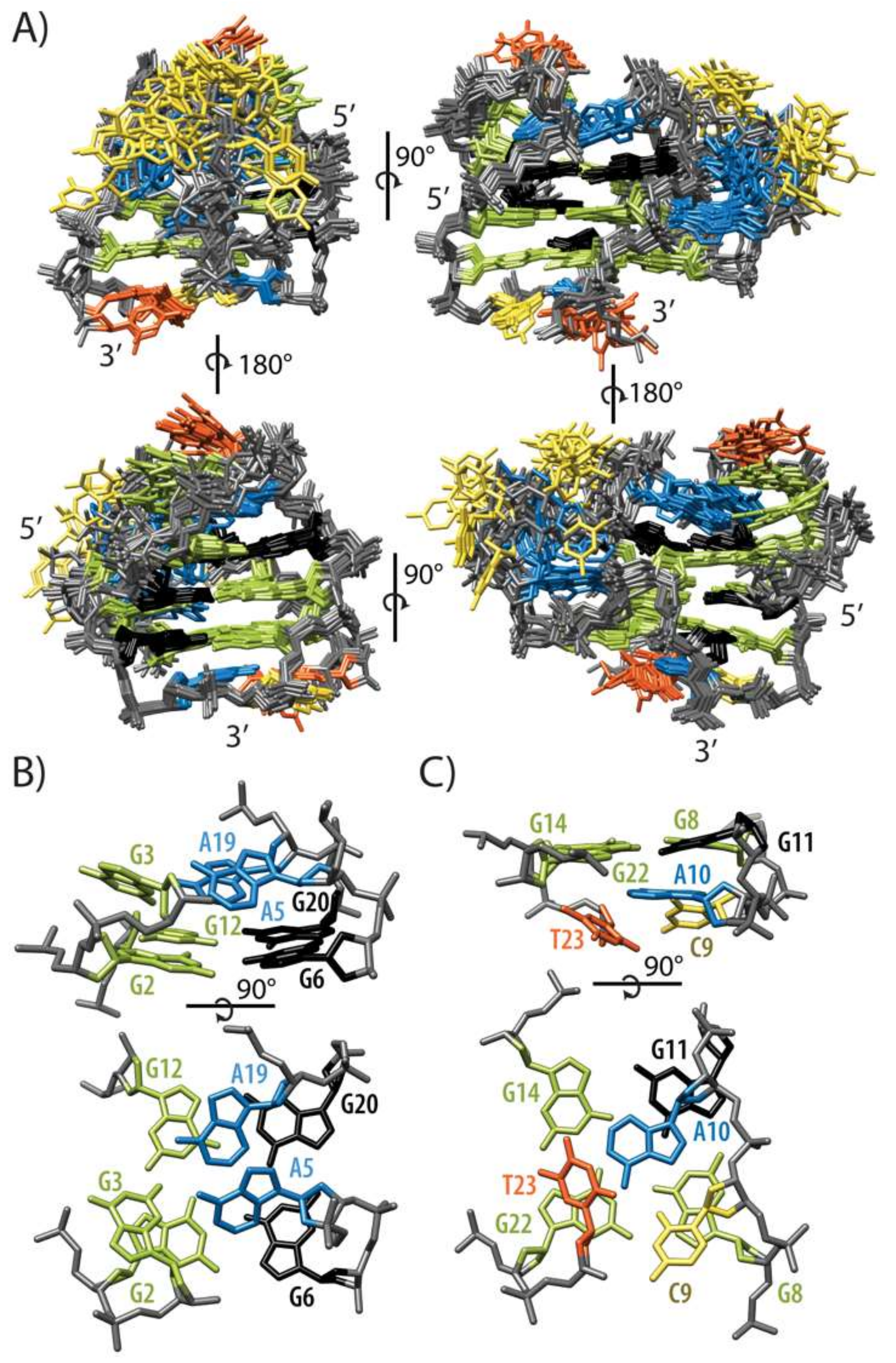
2.2. Topology of G-quadruplex Adopted by HPV52(1–4)
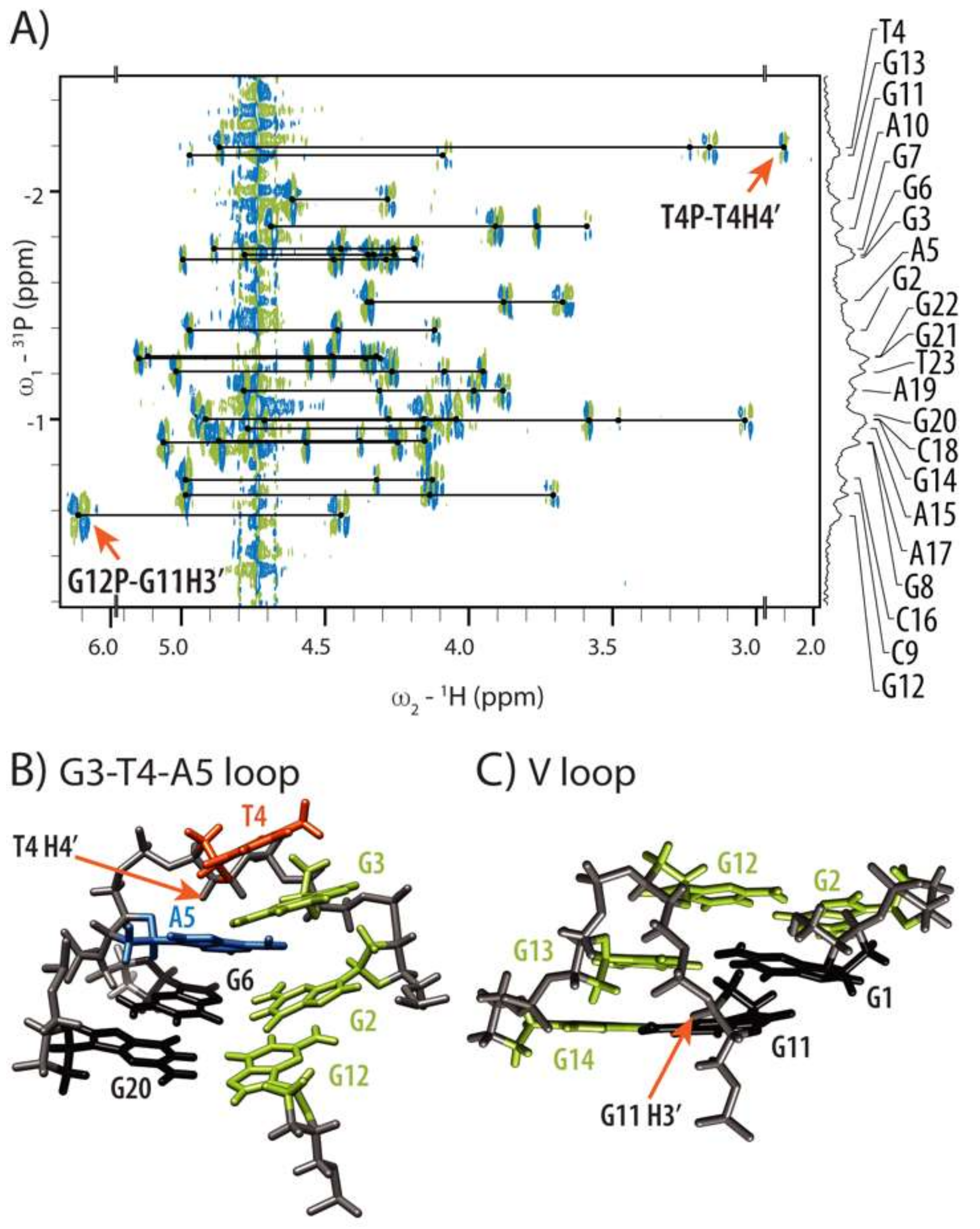
2.3. Conformations of Edgewise G3-T4-A5 and A No-Residue V Loop Justify Unusual Chemical Shifts
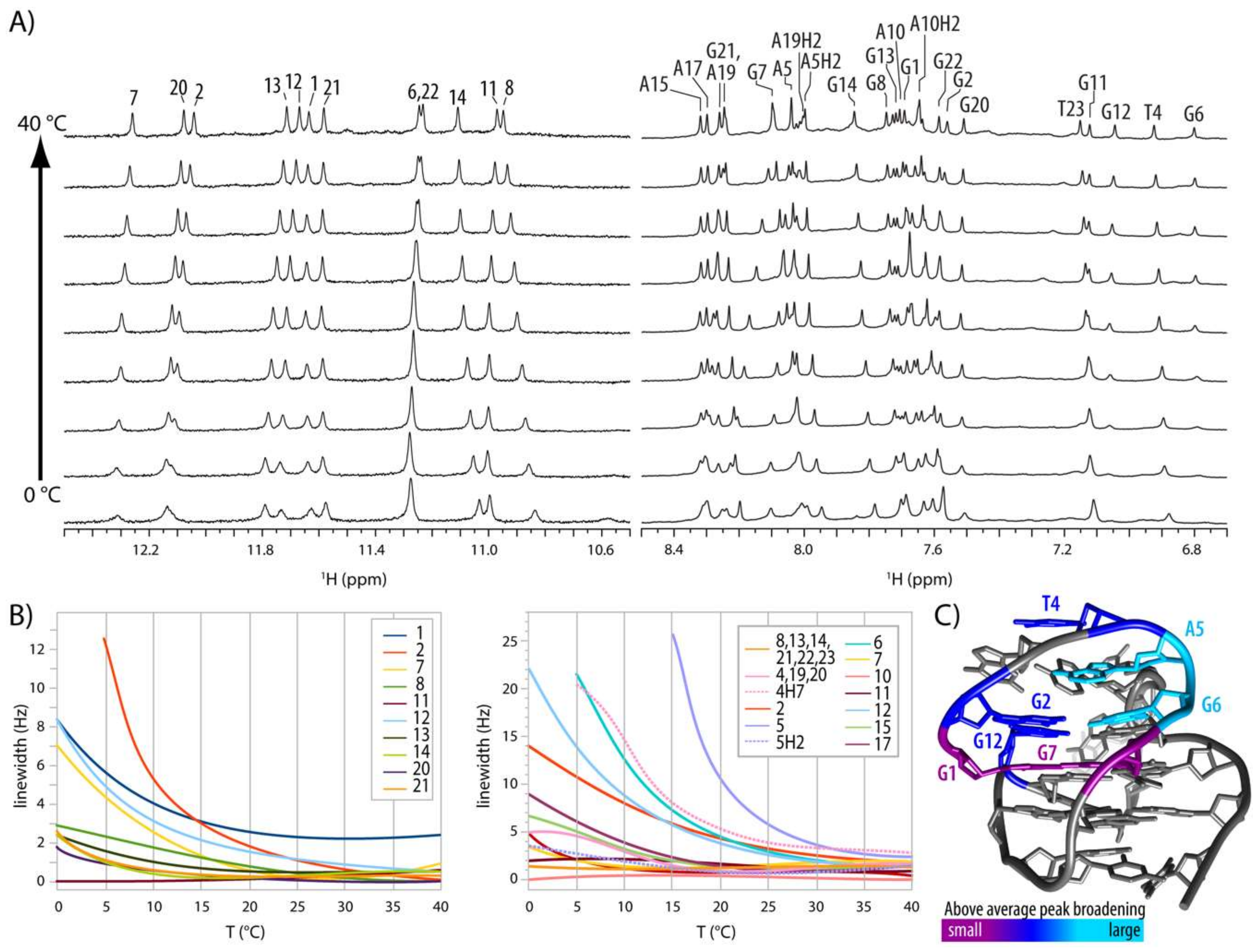
2.4. Internal Motion of G3-T4-A5 Edgewise Loop
2.5. GNA and Four Guanine Tract Are Crucial Structural Elements That Guide Folding of HPV52(1–4)
3. Discussion
4. Materials and Methods
4.1. Sample Preparation
4.2. CD Spectroscopy
4.3. UV Spectroscopy
4.4. Native Gel Electrophoresis
4.5. NMR Spectroscopy
4.6. Structure Calculations
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hpvcenter—International Human Papillomavirus Reference Center. Available online: http://www.nordicehealth.se/hpvcenter/ (accessed on 24 January 2019).
- Hong, S.; Laimins, L.A. Regulation of the life cycle of HPVs by differentiation and the DNA damage response. Future Microbiol. 2013, 8, 1547–1557. [Google Scholar] [CrossRef] [PubMed]
- Winer, R.L.; Kiviat, N.B.; Hughes, J.P.; Adam, D.E.; Lee, S.-K.; Kuypers, J.M.; Koutsky, L.A. Development and duration of human papillomavirus lesions, after initial infection. J. Infect. Dis. 2005, 191, 731–738. [Google Scholar] [CrossRef]
- Doorbar, J.; Quint, W.; Banks, L.; Bravo, I.G.; Stoler, M.; Broker, T.R.; Stanley, M.A. The biology and life-cycle of human papillomaviruses. Vaccine 2012, 30 (Suppl. 5), F55–F70. [Google Scholar] [CrossRef]
- Roman, A.; Munger, K. The papillomavirus E7 proteins. Virology 2013, 445, 138–168. [Google Scholar] [CrossRef] [PubMed]
- Vande Pol, S.B.; Klingelhutz, A.J. Papillomavirus E6 oncoproteins. Virology 2013, 445, 115–137. [Google Scholar] [CrossRef]
- Bosch, F.X.; Broker, T.R.; Forman, D.; Moscicki, A.-B.; Gillison, M.L.; Doorbar, J.; Stern, P.L.; Stanley, M.; Arbyn, M.; Poljak, M.; et al. Comprehensive Control of Human Papillomavirus Infections and Related Diseases. Vaccine 2013, 31 (Suppl. 7), H1–H31. [Google Scholar] [CrossRef]
- Doorbar, J.; Egawa, N.; Griffin, H.; Kranjec, C.; Murakami, I. Human papillomavirus molecular biology and disease association. Rev. Med. Virol. 2015, 25 (Suppl. 1), 2–23. [Google Scholar] [CrossRef]
- Burk, R.D.; Harari, A.; Chen, Z. Human papillomavirus genome variants. Virology 2013, 445, 232–243. [Google Scholar] [CrossRef] [PubMed]
- Marušič, M.; Hošnjak, L.; Krafčikova, P.; Poljak, M.; Viglasky, V.; Plavec, J. The effect of single nucleotide polymorphisms in G-rich regions of high-risk human papillomaviruses on structural diversity of DNA. Biochim. Biophys. Acta BBA Gen. Subj. 2017, 1861, 1229–1236. [Google Scholar] [CrossRef]
- Gellert, M.; Lipsett, M.N.; Davies, D.R. Helix formation by guanylic acid. Proc. Natl. Acad. Sci. USA 1962, 48, 2013–2018. [Google Scholar] [CrossRef] [PubMed]
- Paeschke, K.; Bochman, M.L.; Garcia, P.D.; Cejka, P.; Friedman, K.L.; Kowalczykowski, S.C.; Zakian, V.A. Pif1 family helicases suppress genome instability at G-quadruplex motifs. Nature 2013, 497, 458–462. [Google Scholar] [CrossRef]
- Rodriguez, R.; Miller, K.M.; Forment, J.V.; Bradshaw, C.R.; Nikan, M.; Britton, S.; Oelschlaegel, T.; Xhemalce, B.; Balasubramanian, S.; Jackson, S.P. Small-molecule–induced DNA damage identifies alternative DNA structures in human genes. Nat. Chem. Biol. 2012, 8, 301–310. [Google Scholar] [CrossRef] [PubMed]
- Wolfe, A.L.; Singh, K.; Zhong, Y.; Drewe, P.; Rajasekhar, V.K.; Sanghvi, V.R.; Mavrakis, K.J.; Jiang, M.; Roderick, J.E.; Van der Meulen, J.; et al. RNA G-quadruplexes cause eIF4A-dependent oncogene translation in cancer. Nature 2014, 513, 65–70. [Google Scholar] [CrossRef]
- Siddiqui-Jain, A.; Grand, C.L.; Bearss, D.J.; Hurley, L.H. Direct evidence for a G-quadruplex in a promoter region and its targeting with a small molecule to repress c-MYC transcription. Proc. Natl. Acad. Sci. USA 2002, 99, 11593–11598. [Google Scholar] [CrossRef] [PubMed]
- Lavezzo, E.; Berselli, M.; Frasson, I.; Perrone, R.; Palù, G.; Brazzale, A.R.; Richter, S.N.; Toppo, S. G-quadruplex forming sequences in the genome of all known human viruses: A comprehensive guide. PLoS Comput. Biol. 2018, 14, e1006675. [Google Scholar] [CrossRef] [PubMed]
- Ruggiero, E.; Richter, S.N. G-quadruplexes and G-quadruplex ligands: Targets and tools in antiviral therapy. Nucleic Acids Res. 2018, 46, 3270–3283. [Google Scholar] [CrossRef]
- Métifiot, M.; Amrane, S.; Litvak, S.; Andreola, M.-L. G-quadruplexes in viruses: Function and potential therapeutic applications. Nucleic Acids Res. 2014, 42, 12352–12366. [Google Scholar] [CrossRef]
- Baran, N. The SV40 large T-antigen helicase can unwind four stranded DNA structures linked by G-quartets. Nucleic Acids Res. 1997, 25, 297–303. [Google Scholar] [CrossRef]
- Madireddy, A.; Purushothaman, P.; Loosbroock, C.P.; Robertson, E.S.; Schildkraut, C.L.; Verma, S.C. G-quadruplex-interacting compounds alter latent DNA replication and episomal persistence of KSHV. Nucleic Acids Res. 2016, 44, 3675–3694. [Google Scholar] [CrossRef]
- Murat, P.; Zhong, J.; Lekieffre, L.; Cowieson, N.P.; Clancy, J.L.; Preiss, T.; Balasubramanian, S.; Khanna, R.; Tellam, J. G-quadruplexes regulate Epstein-Barr virus–encoded nuclear antigen 1 mRNA translation. Nat. Chem. Biol. 2014, 10, 358–364. [Google Scholar] [CrossRef]
- Perrone, R.; Nadai, M.; Frasson, I.; Poe, J.A.; Butovskaya, E.; Smithgall, T.E.; Palumbo, M.; Palù, G.; Richter, S.N. A dynamic G-quadruplex region regulates the HIV-1 long terminal repeat promoter. J. Med. Chem. 2013, 56, 6521–6530. [Google Scholar] [CrossRef]
- Plyler, J.; Jasheway, K.; Tuesuwan, B.; Karr, J.; Brennan, J.S.; Kerwin, S.M.; David, W.M. Real-time Investigation of SV40 Large T-antigen Helicase Activity Using Surface Plasmon Resonance. Cell Biochem. Biophys. 2009, 53, 43–52. [Google Scholar] [CrossRef] [PubMed]
- Shen, W.; Gorelick, R.J.; Bambara, R.A. HIV-1 Nucleocapsid Protein Increases Strand Transfer Recombination by Promoting Dimeric G-quartet Formation. J. Biol. Chem. 2011, 286, 29838–29847. [Google Scholar] [CrossRef]
- Huppert, J.L.; Balasubramanian, S. Prevalence of quadruplexes in the human genome. Nucleic Acids Res. 2005, 33, 2908–2916. [Google Scholar] [CrossRef] [PubMed]
- Todd, A.K.; Johnston, M.; Neidle, S. Highly prevalent putative quadruplex sequence motifs in human DNA. Nucleic Acids Res. 2005, 33, 2901–2907. [Google Scholar] [CrossRef] [PubMed]
- Chambers, V.S.; Marsico, G.; Boutell, J.M.; Di Antonio, M.; Smith, G.P.; Balasubramanian, S. High-throughput sequencing of DNA G-quadruplex structures in the human genome. Nat. Biotechnol. 2015, 33, 877–881. [Google Scholar] [CrossRef]
- Hänsel-Hertsch, R.; Beraldi, D.; Lensing, S.V.; Marsico, G.; Zyner, K.; Parry, A.; Di Antonio, M.; Pike, J.; Kimura, H.; Narita, M.; et al. G-quadruplex structures mark human regulatory chromatin. Nat. Genet. 2016, 48, 1267–1272. [Google Scholar] [CrossRef] [PubMed]
- Huppert, J.L. Four-stranded DNA: Cancer, gene regulation and drug development. Philos. Trans. R. Soc. Lond. Math. Phys. Eng. Sci. 2007, 365, 2969–2984. [Google Scholar] [CrossRef] [PubMed]
- Tlučková, K.; Marušič, M.; Tóthová, P.; Bauer, L.; Šket, P.; Plavec, J.; Viglasky, V. Human Papillomavirus G-Quadruplexes. Biochemistry 2013, 52, 7207–7216. [Google Scholar] [CrossRef] [PubMed]
- Bedrat, A.; Lacroix, L.; Mergny, J.-L. Re-evaluation of G-quadruplex propensity with G4Hunter. Nucleic Acids Res. 2016, 44, 1746–1759. [Google Scholar] [CrossRef] [PubMed]
- Kikin, O.; D’Antonio, L.; Bagga, P.S. QGRS Mapper: A web-based server for predicting G-quadruplexes in nucleotide sequences. Nucleic Acids Res. 2006, 34, W676–W682. [Google Scholar] [CrossRef] [PubMed]
- Chan, P.K.S.; Ho, W.C.S.; Chan, M.C.W.; Wong, M.C.S.; Yeung, A.C.M.; Chor, J.S.Y.; Hui, M. Meta-Analysis on Prevalence and Attribution of Human Papillomavirus Types 52 and 58 in Cervical Neoplasia Worldwide. PLoS ONE 2014, 9, e107573. [Google Scholar] [CrossRef]
- Benson, D.A.; Cavanaugh, M.; Clark, K.; Karsch-Mizrachi, I.; Lipman, D.J.; Ostell, J.; Sayers, E.W. GenBank. Nucleic Acids Res. 2013, 41, D36–D42. [Google Scholar] [CrossRef]
- Guédin, A.; Gros, J.; Alberti, P.; Mergny, J.-L. How long is too long? Effects of loop size on G-quadruplex stability. Nucleic Acids Res. 2010, 38, 7858–7868. [Google Scholar] [CrossRef]
- Karsisiotis, A.I.; Hessari, N.M.; Novellino, E.; Spada, G.P.; Randazzo, A.; Webba da Silva, M. Topological Characterization of Nucleic Acid G-Quadruplexes by UV Absorption and Circular Dichroism. Angew. Chem. Int. Ed. 2011, 50, 10645–10648. [Google Scholar] [CrossRef]
- Del Villar-Guerra, R.; Trent, J.O.; Chaires, J.B. G-Quadruplex Secondary Structure Obtained from Circular Dichroism Spectroscopy. Angew. Chem. Int. Ed. Engl. 2018, 57, 7171–7175. [Google Scholar] [CrossRef]
- Webba da Silva, M. Geometric Formalism for DNA Quadruplex Folding. Chem. Eur. J. 2007, 13, 9738–9745. [Google Scholar] [CrossRef]
- Greene, K.L.; Wang, Y.; Live, D. Influence of the glycosidic torsion angle on 13C and 15N shifts in guanosine nucleotides: Investigations of G-tetrad models with alternating syn and anti bases. J. Biomol. NMR 1995, 5, 333–338. [Google Scholar] [CrossRef] [PubMed]
- Fonville, J.M.; Swart, M.; Vokáčová, Z.; Sychrovský, V.; Šponer, J.E.; Šponer, J.; Hilbers, C.W.; Bickelhaupt, F.M.; Wijmenga, S.S. Chemical Shifts in Nucleic Acids Studied by Density Functional Theory Calculations and Comparison with Experiment. Chem. Eur. J. 2012, 18, 12372–12387. [Google Scholar] [CrossRef] [PubMed]
- Dickerhoff, J.; Weisz, K. Flipping a G-Tetrad in a Unimolecular Quadruplex without Affecting Its Global Fold. Angew. Chem. Int. Ed. 2015, 54, 5588–5591. [Google Scholar] [CrossRef] [PubMed]
- Ulrich, E.L.; Akutsu, H.; Doreleijers, J.F.; Harano, Y.; Ioannidis, Y.E.; Lin, J.; Livny, M.; Mading, S.; Maziuk, D.; Miller, Z.; et al. BioMagResBank. Nucleic Acids Res. 2008, 36, D402–D408. [Google Scholar] [CrossRef]
- Chou, S.-H.; Zhu, L.; Gao, Z.; Cheng, J.-W.; Reid, B.R. Hairpin Loops Consisting of Single Adenine Residues Closed by Sheared A·A and G·G Pairs Formed by the DNA Triplets AAA and GAG: Solution Structure of the d(GTACAAAGTAC) Hairpin. J. Mol. Biol. 1996, 264, 981–1001. [Google Scholar] [CrossRef]
- Hirao, I.; Kawai, G.; Yoshizawa, S.; Nishimura, Y.; Ishido, Y.; Watanabe, K.; Miura, K. Most compact hairpin-turn structure exerted by a short DNA fragment, d(GCGAAGC) in solution: An extraordinarily stable structure resistant to nucleases and heat. Nucleic Acids Res. 1994, 22, 576–582. [Google Scholar] [CrossRef]
- Kettani, A.; Gorin, A.; Majumdar, A.; Hermann, T.; Skripkin, E.; Zhao, H.; Jones, R.; Patel, D.J. A dimeric DNA interface stabilized by stacked A · (G · G · G · G) · A hexads and coordinated monovalent cations. J. Mol. Biol. 2000, 297, 627–644. [Google Scholar] [CrossRef]
- Matsugami, A.; Ouhashi, K.; Kanagawa, M.; Liu, H.; Kanagawa, S.; Uesugi, S.; Katahira, M. An intramolecular quadruplex of (GGA)4 triplet repeat DNA with a G:G:G:G tetrad and a G(:A):G(:A):G(:A):G heptad, and its dimeric interaction. J. Mol. Biol. 2001, 313, 255–269. [Google Scholar] [CrossRef] [PubMed]
- Yoshizawa, S.; Kawai, G.; Watanabe, K.; Miura, K.; Hirao, I. GNA Trinucleotide Loop Sequences Producing Extraordinarily Stable DNA Minihairpins. Biochemistry 1997, 36, 4761–4767. [Google Scholar] [CrossRef]
- Zhu, L.; Chou, S.H.; Xu, J.; Reid, B.R. Structure of a single-cytidine hairpin loop formed by the DNA triplet GCA. Nat. Struct. Biol. 1995, 2, 1012–1017. [Google Scholar] [CrossRef] [PubMed]
- Bončina, M.; Vesnaver, G.; Chaires, J.B.; Lah, J. Unraveling the Thermodynamics of the Folding and Interconversion of Human Telomere G-Quadruplexes. Angew. Chem. Int. Ed. Engl. 2016, 55, 10340–10344. [Google Scholar] [CrossRef] [PubMed]
- Kuryavyi, V.; Patel, D.J. Solution Structure of a Unique G-Quadruplex Scaffold Adopted by a Guanosine-Rich Human Intronic Sequence. Structure 2010, 18, 73–82. [Google Scholar] [CrossRef] [PubMed]
- Cang, X.; Šponer, J.; Cheatham, T.E. Insight into G-DNA structural polymorphism and folding from sequence and loop connectivity through free energy analysis. J. Am. Chem. Soc. 2011, 133, 14270–14279. [Google Scholar] [CrossRef] [PubMed]
- Brčić, J.; Plavec, J. Solution structure of a DNA quadruplex containing ALS and FTD related GGGGCC repeat stabilized by 8-bromodeoxyguanosine substitution. Nucleic Acids Res. 2015, 43, 8590–8600. [Google Scholar] [CrossRef] [PubMed]
- Crnugelj, M.; Sket, P.; Plavec, J. Small change in a G-rich sequence, a dramatic change in topology: New dimeric G-quadruplex folding motif with unique loop orientations. J. Am. Chem. Soc. 2003, 125, 7866–7871. [Google Scholar] [CrossRef]
- Delaglio, F.; Grzesiek, S.; Vuister, G.W.; Zhu, G.; Pfeifer, J.; Bax, A. NMRPipe: A multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 1995, 6, 277–293. [Google Scholar] [CrossRef] [PubMed]
- Goddard, T.D.; Kneller, D.G. SPARKY 3; University of California: San Francisco, CA, USA, 2008. [Google Scholar]
- Lee, W.; Tonelli, M.; Markley, J.L. NMRFAM-SPARKY: Enhanced software for biomolecular NMR spectroscopy. Bioinformatics 2015, 31, 1325–1327. [Google Scholar] [CrossRef]
- Clore, G.M.; Gronenborn, A.M. Interproton distance measurements in solution for a double-stranded DNA undecamer comprising a portion of the specific target site for the cyclic AMP receptor protein in the gal operon: A nuclear Overhauser enhancement study. FEBS Lett. 1984, 175, 117–123. [Google Scholar] [CrossRef]
- Wijmenga, S.S.; van Buuren, B.N.M. The use of NMR methods for conformational studies of nucleic acids. Prog. Nucl. Magn. Reson. Spectrosc. 1998, 32, 287–387. [Google Scholar] [CrossRef]
- Webba da Silva, M. Experimental Demonstration of T:(G:G:G:G):T Hexad and T:A:A:T Tetrad Alignments within a DNA Quadruplex Stem. Biochemistry 2005, 44, 3754–3764. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, J.T.; Arar, K.; Petersen, M. NMR solution structures of LNA (locked nucleic acid) modified quadruplexes. Nucleic Acids Res. 2006, 34, 2006–2014. [Google Scholar] [CrossRef]
- Kim, S.-G.; Lin, L.-J.; Reid, B.R. Determination of nucleic acid backbone conformation by proton NMR. Biochemistry 1992, 31, 3564–3574. [Google Scholar] [CrossRef]
- Pikkemaat, J.A.; Altona, C. Fine Structure of the P–H5′ Cross-Peak in 31P–1H Correlated 2D NMR Spectroscopy. An Efficient Probe for the Backbone Torsion Angles β and γ in Nucleic Acids. Magn. Reson. Chem. 1996, 34, S33–S39. [Google Scholar] [CrossRef]
- Roongta, V.A.; Jones, C.R.; Gorenstein, D.G. Effect of distortions in the deoxyribose phosphate backbone conformation of duplex oligodeoxyribonucleotide dodecamers containing GT, GG, GA, AC, and GU base-pair mismatches on 31P NMR spectra. Biochemistry 1990, 29, 5245–5258. [Google Scholar] [CrossRef]
- Case, D.A.; Babin, V.; Berryman, J.T.; Betz, R.M.; Cai, Q.; Cerutti, D.S.; Cheatham, T.E.; Darden, T.A.; Duke, R.E.; Gohlke, H.; et al. AMBER 14; University of California: San Francisco, CA, USA, 2014. [Google Scholar]
- Aduri, R.; Psciuk, B.T.; Saro, P.; Taniga, H.; Schlegel, H.B.; SantaLucia, J. AMBER Force Field Parameters for the Naturally Occurring Modified Nucleosides in RNA. J. Chem. Theory Comput. 2007, 3, 1464–1475. [Google Scholar] [CrossRef]
- Krepl, M.; Zgarbová, M.; Stadlbauer, P.; Otyepka, M.; Banáš, P.; Koča, J.; Cheatham, T.E.; Jurečka, P.; Sponer, J. Reference simulations of noncanonical nucleic acids with different χ variants of the AMBER force field: Quadruplex DNA, quadruplex RNA and Z-DNA. J. Chem. Theory Comput. 2012, 8, 2506–2520. [Google Scholar] [CrossRef] [PubMed]
- Zgarbová, M.; Luque, F.J.; Šponer, J.; Cheatham, T.E.; Otyepka, M.; Jurečka, P. Toward Improved Description of DNA Backbone: Revisiting Epsilon and Zeta Torsion Force Field Parameters. J. Chem. Theory Comput. 2013, 9, 2339–2354. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Chen, J.; Ju, H.; Zhou, J. Thermal denaturation profile: A straightforward signature to characterize parallel G-quadruplexes. Biochimie 2019, 157, 22–25. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of the compounds are not available from the authors. |
| NMR Distance and Torsion Angle Restraints | |||
|---|---|---|---|
| NOE-derived distance restraints | Non-exch. | Exch. | All |
| Total | 322 | 55 | 377 |
| Intra-residue | 215 | 0 | 215 |
| Inter-residue | 107 | 55 | 162 |
| Sequential | 88 | 15 | 103 |
| Long-range | 19 | 40 | 59 |
| Chemical shift derived distance restraints | 4 | ||
| Hydrogen bond restraints | 24 | ||
| Hydrogen bonds non-observed | 3 | ||
| Torsion angle restraints | 82 | ||
| G-quartet planarity restraints | 36 | ||
| Structure statistics | |||
| Violations | |||
| Mean NOE restraint violation (Å) | 0.14 ± 0.001 | ||
| Max. NOE restraint violation (Å) | 0.33 | ||
| Max torsion angle restraint violation (°) | 6.768 | ||
| Deviation from idealized geometry | |||
| Bonds (Å) | 0.012 ± 0.000 | ||
| Angles (°) | 2.43 ± 0.03 | ||
| Pairwise heavy atom RMSD (Å) | |||
| Overall | 1.779 | ||
| G1-G14 + G20-G23 | 0.987 | ||
| G-quartets | 0.751 | ||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Marušič, M.; Plavec, J. Towards Understanding of Polymorphism of the G-rich Region of Human Papillomavirus Type 52. Molecules 2019, 24, 1294. https://doi.org/10.3390/molecules24071294
Marušič M, Plavec J. Towards Understanding of Polymorphism of the G-rich Region of Human Papillomavirus Type 52. Molecules. 2019; 24(7):1294. https://doi.org/10.3390/molecules24071294
Chicago/Turabian StyleMarušič, Maja, and Janez Plavec. 2019. "Towards Understanding of Polymorphism of the G-rich Region of Human Papillomavirus Type 52" Molecules 24, no. 7: 1294. https://doi.org/10.3390/molecules24071294
APA StyleMarušič, M., & Plavec, J. (2019). Towards Understanding of Polymorphism of the G-rich Region of Human Papillomavirus Type 52. Molecules, 24(7), 1294. https://doi.org/10.3390/molecules24071294







