A Resolution-Free Parallel Algorithm for Image Edge Detection within the Framework of Enzymatic Numerical P Systems
Abstract
1. Introduction
2. MC and ENPS
- m is the number of membranes used.
- H is an alphabet that contains m symbols, and .
- is the membrane structure.
- is the set of variables from membrane i and are the initial values for these variables.
- is the set of rules in membrane i, composed of a production function and a repartition protocol. A typical rule is as follows.where is a variable from different from and . The rule can be executed at a time t only if . From the definition of ENPS, it is clear that with enzymes-like variables, the system can control multiple production functions to run in parallel in the same membrane deterministically [54]. Hence, it can overcome the disadvantages of traditional NPS that only run one rule nondeterministically at a time in a membrane. The ENPS with deterministic, parallel execution model has already been proved to be Turing universal [55,56]. In [57], it is shown that any ENPS working in all-parallel mode or one parallel model can be simulated by an equivalent one-membrane ENPS working in the same mode. Since the proposal of ENPS, this model has been successfully applied in a wide range of domains, such as robot control [58], big data field [59], and sequential minimal optimization [60] fields. In this paper, ENPS is used to solve the problem of gradient based image edge detection.
3. Problem Statement
| Algorithm 1: The pseudo code of GBED |
|
4. The EDENP Algorithm
4.1. Mathematical Model of EDENP
- .
- .
- .
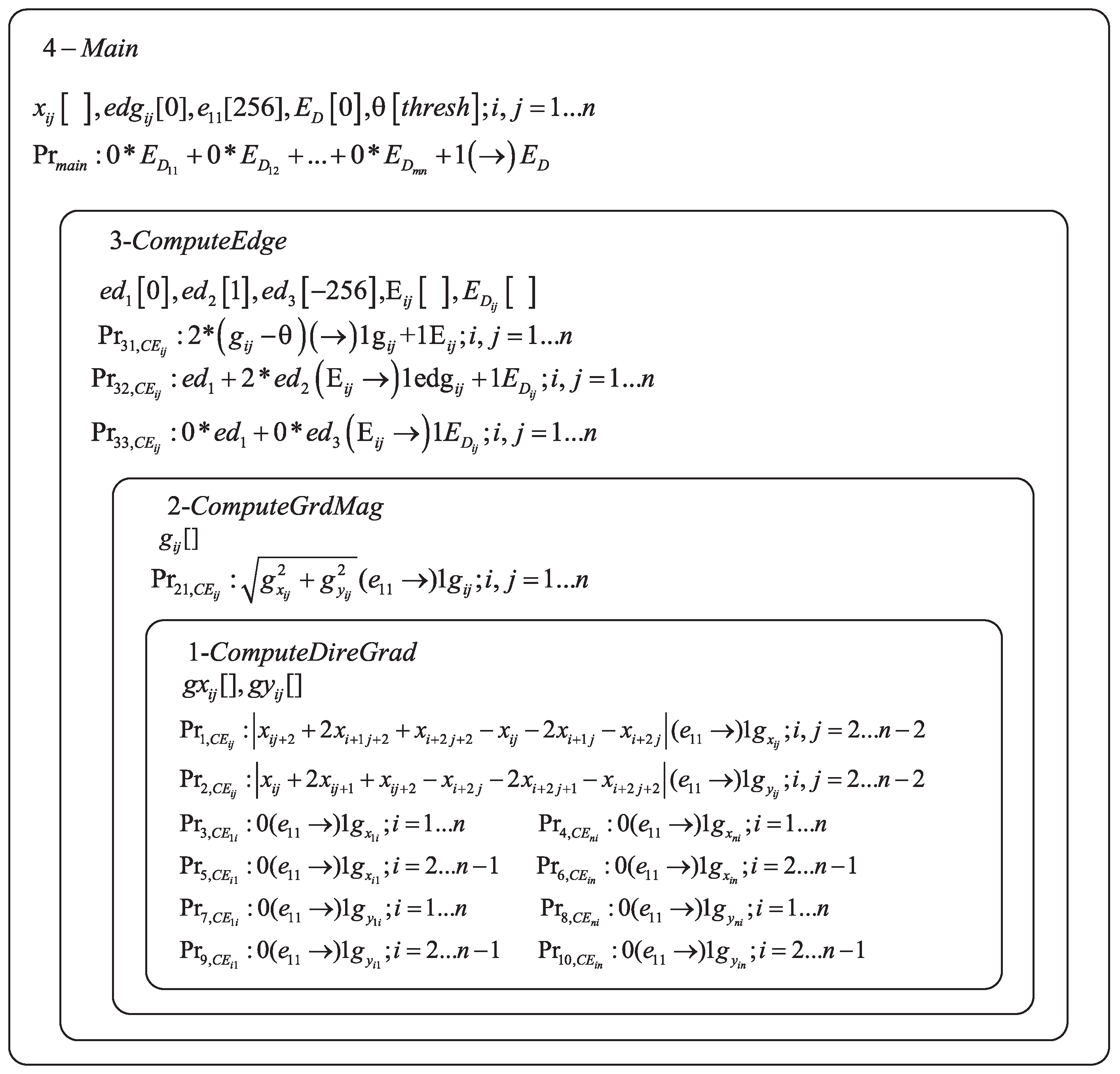
- , , , ., are the gray value of pixel with coordinate of on the source image plane., are the corresponding edge points of the source image with initial value 0., is a numerical variable which is used as the threshold value for edge detection, and the value of threshold should be predefined., are the horizontal derivative approximations at each pixel., are the vertical derivative approximations at each pixel., are the gradient magnitude approximations at each pixel., is a numerical variable with initial value 0, which is used as the background value of the edge image., is a numerical variable with initial value 1, which is used as the edge point value of the edge image., is a numerical variable with initial value −256, which is used as a intermediate variable.
- is a set of enzyme variables from membrane k, i.e., ,,,.
- is the set of programs (rules) in membrane k, composed of a production function and a repartition protocol.,,,,,,,.Those rules are used to execute Formula (1). The enzyme in must exist in enough amount so that the rules can be activated. Specifically, if the value of the enzyme is greater than variable , then rules are effective. Since variable is the gray value of image, the maximum value is 255. So, the initial value of is set to 256, such that the condition modeled by rule are satisfied. It is important to note that the number of rules are , and all the rules are executed in parallel.
- are the rules which are executed by Formula (5). Hence, after executing , the value of the variables , are obtained. The maximum value of and is 255, and the enzyme is 256. So the condition of execution for rules is satisfied. Hence, all rules are executed concurrently.
- are the rules which compute in Formula (6). After executing , the value of are obtained, which is equal to variables and in rule .
- ;and are rules for computing edge value as Formula (7). If is greater than or equal to 0, then and are executed. Because is 0, and is -256, so and . The execution condition of and is satisfied. If , only will be executed. Because and , only the execution condition of can be satisfied. After executing and , variables will be set to 1 if and every variable will be assigned.
- is a rule contained in membrane 4, which controls the stop condition of the P system. For pixel , if all the enzyme variables are assigned, the condition for is meet. Enzyme variable is set to 1 by rule , and the system stops running.
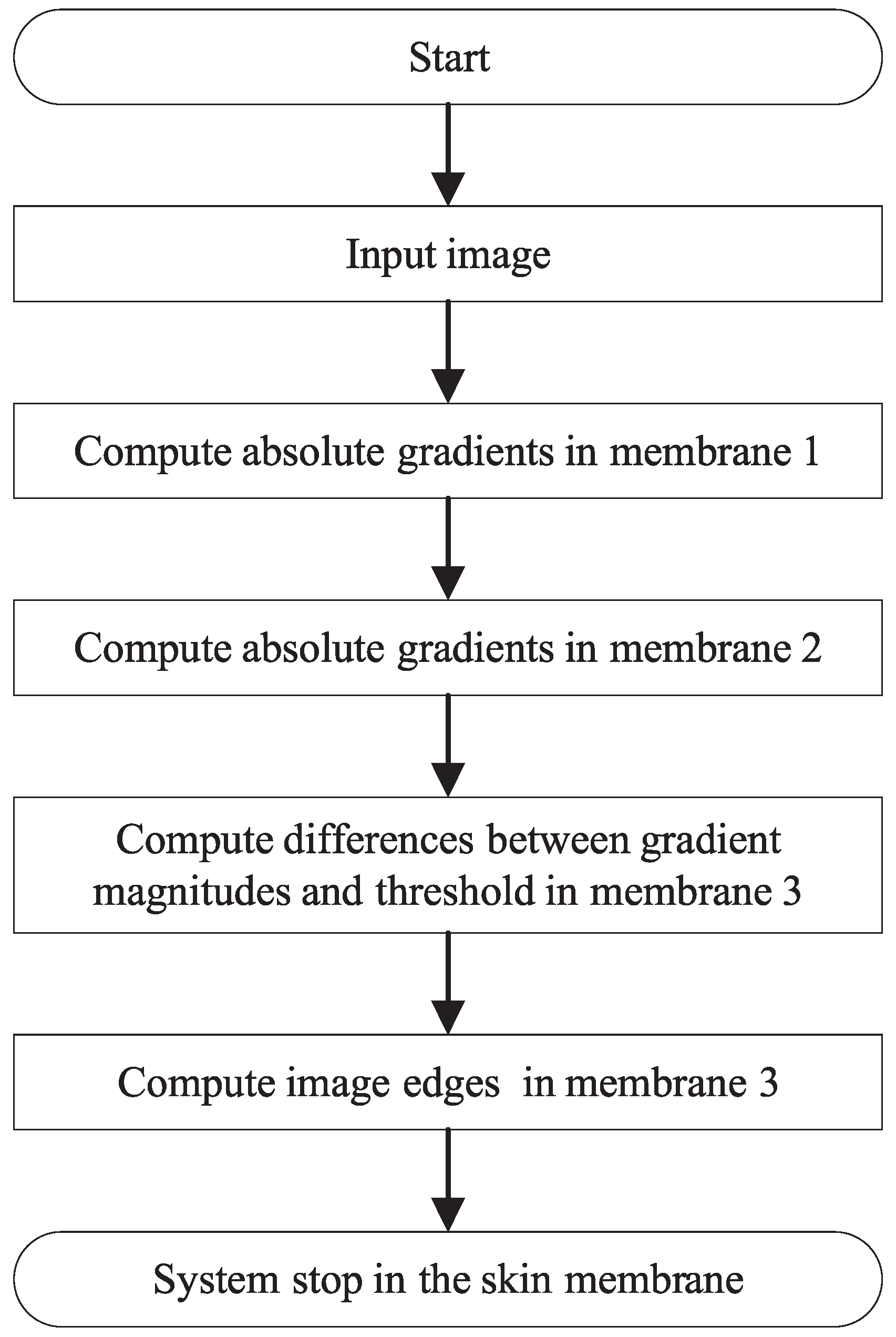
4.2. The Structure and Execution Processes of EDENP
4.3. Complexity and Resources Analysis
5. Experiments and Results
5.1. Performance Evaluation
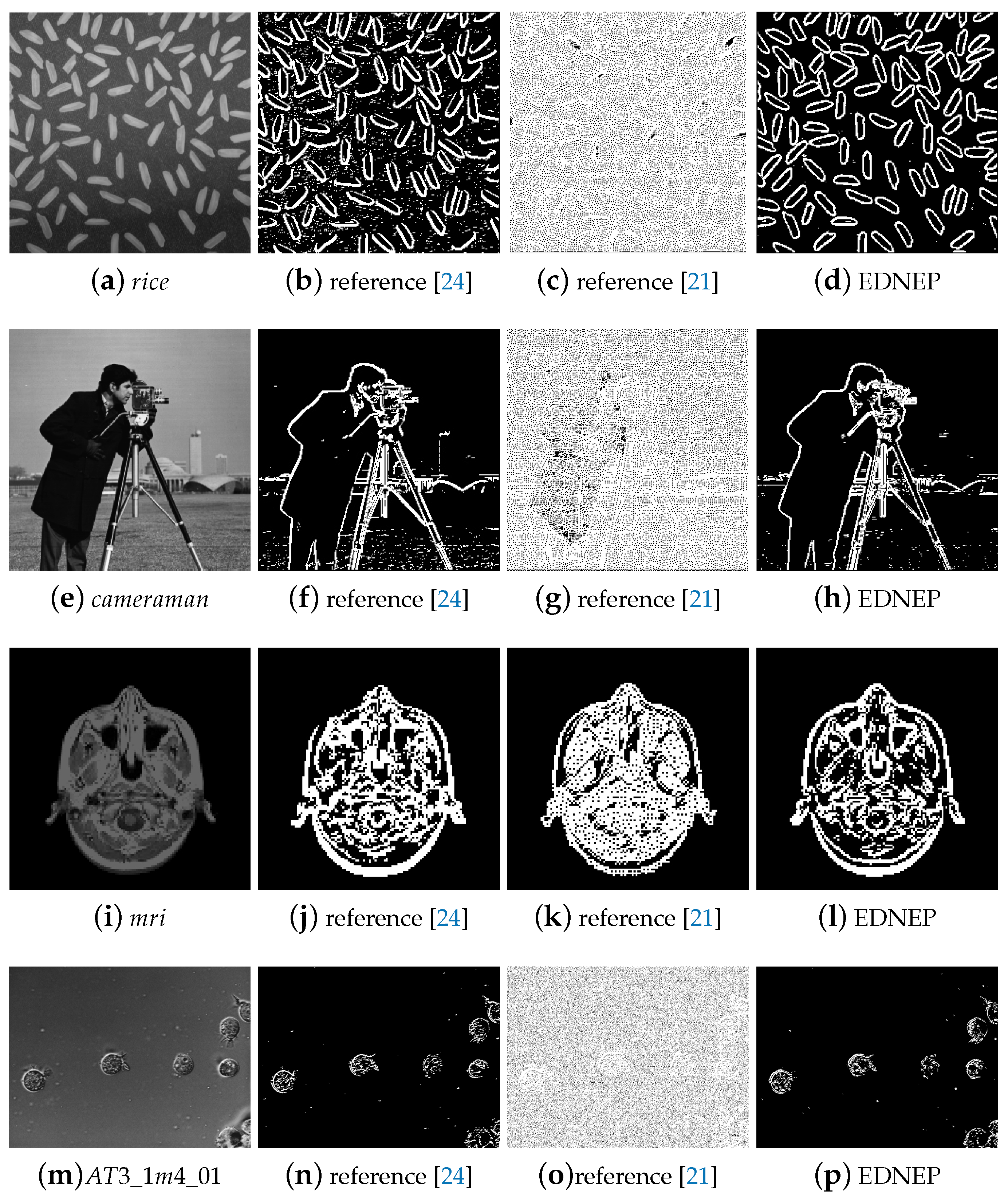
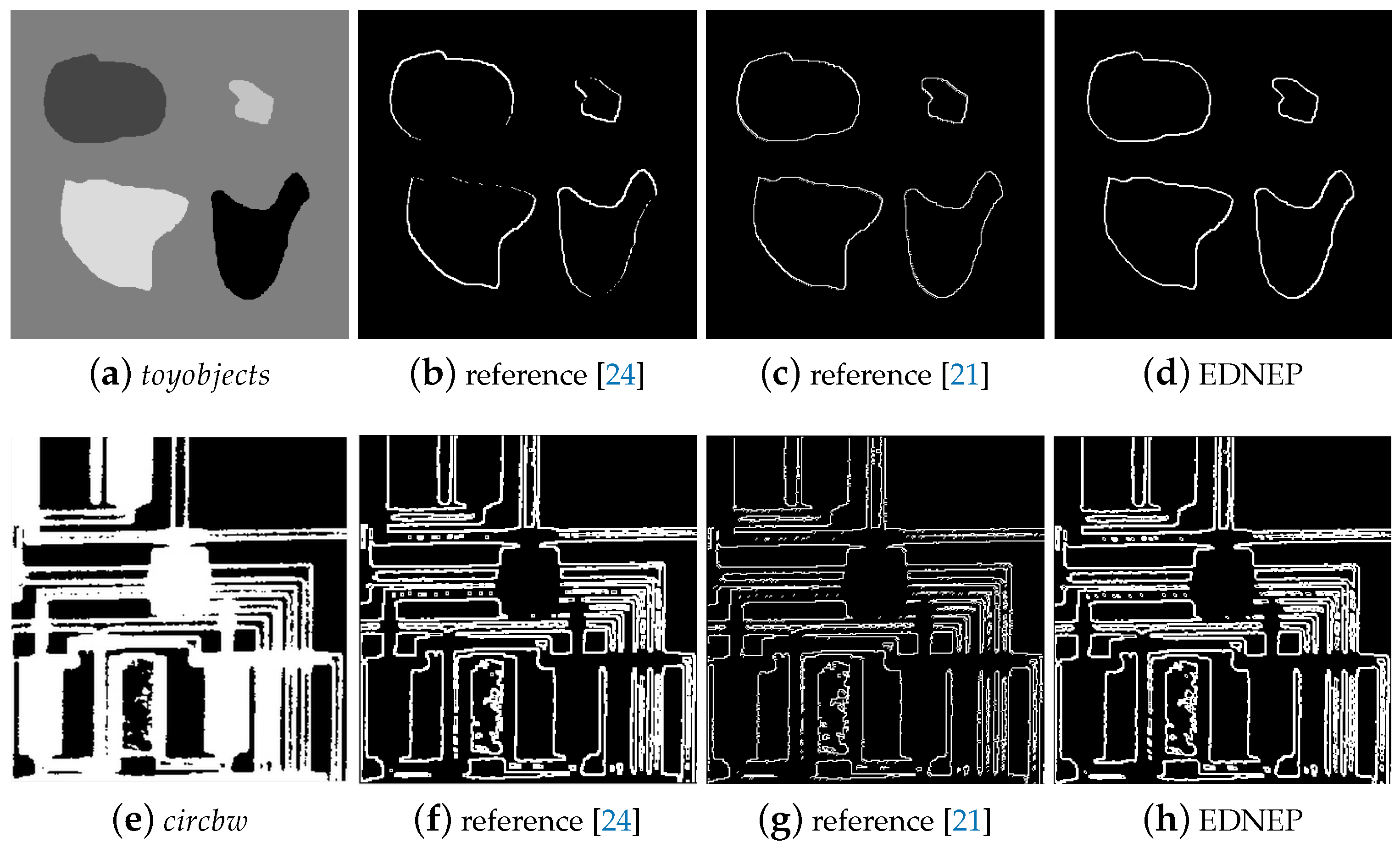
5.1.1. Qualitative Evaluation
5.1.2. Quantitative Evaluation
5.2. Efficiency Evaluation
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Zheng, S.; Yuille, A.; Tu, Z. Detecting object boundaries using low-, mid-, and high-level information. Comput. Vis. Image Underst. 2010, 114, 1055–1067. [Google Scholar] [CrossRef]
- Zhao, L.; Zhao, Q.; Liu, H.; Lv, P.; Gu, D. Structural sparse representation-based semi-supervised learning and edge detection proposal for visual tracking. Vis. Comput. 2017, 33, 1169–1184. [Google Scholar] [CrossRef]
- Hua, S.; Wen, H. Moment-preserving edge detection and its application to image data compression. Opt. Eng. 2013, 32, 1596–1608. [Google Scholar] [CrossRef]
- Satpathy, A.; Jiang, X.; Eng, H.L. LBP-based edge-texture features for object recognition. IEEE Trans. Image Process. 2014, 23, 1953–1964. [Google Scholar] [CrossRef]
- Saif, J.; Hammad, M.; Alqubati, I. Gradient based image edge detection. Int. J. Eng. Technol. 2016, 8, 153–156. [Google Scholar] [CrossRef]
- Jung, J.; Lee, H.; Lee, J.; Park, D. A novel template matching scheme for fast full-Search boosted by an integral image. IEEE Signal Proc. Lett. 2010, 17, 107–110. [Google Scholar] [CrossRef]
- Schweitzer, H.; Deng, R.; Anderson, R.F. A dual-bound algorithm for very fast and exact template matching. IEEE Trans. Pattern Anal. Mach. Intell. 2011, 33, 459–470. [Google Scholar] [CrossRef] [PubMed]
- Ghassabeh, Y.A.; Rudzicz, F.; Moghaddam, H.A. Fast incremental LDA feature extraction. Pattern Recognit. 2015, 48, 1999–2012. [Google Scholar] [CrossRef]
- Zhou, G.; Cichocki, A.; Zhang, Y.; Mandic, D.P. Group component analysis for multiblock data: common and individual feature extraction. IEEE Trans. Neural Netw. 2016, 27, 2426–2439. [Google Scholar] [CrossRef]
- Herout, A.; Josth, R.; Juranek, R.; Havel, J.; Hradis, M.; Zemcik, P. Real-time object detection on CUDA. J. Real-Time Image Process. 2011, 6, 159–170. [Google Scholar] [CrossRef]
- Jiang, B. Real-time multi-resolution edge detection with pattern analysis on graphics processing unit. J. Real-Time Image Process. 2018, 14, 293–321. [Google Scholar] [CrossRef]
- Zuo, H.; Zhang, Q.; Yong, X.; Zhao, R. Fast sobel edge detection algorithm based on GPU. Opto-Electron. Eng. 2009, 36, 8–12. [Google Scholar] [CrossRef]
- Jiang, J.; Liu, C.; Ling, S.R. An FPGA implementation for real-time edge detection. J. Real-Time Image Process. 2018, 15, 787–797. [Google Scholar] [CrossRef]
- Nausheen, N.; Seal, A.; Khanna, P.; Halder, S. A FPGA based implementation of Sobel edge detection. Microprocess Microsy 2018, 56, 84–91. [Google Scholar] [CrossRef]
- Paolo, B.; Sipp, D. Regulation: Sell help not hope. Nature 2014, 510, 336–337. [Google Scholar] [CrossRef]
- Jiang, N.; Dang, Y.; Wang, J. Quantum image matching. Quantum Inf. Process. 2016, 15, 3543–3572. [Google Scholar] [CrossRef]
- Tsaftaris, S.A.; Katsaggelos, A.K.; Pappas, T.N.; Papoutsakis, E.T. How can DNA computing be applied to digital signal processing? IEEE Signal Process. Mag. 2004, 21, 57–61. [Google Scholar] [CrossRef]
- Díaz-Pernil, D.; Gutierrez-Naranjo, M.; Peng, H. Membrane computing and image processing: A short survey. J. Membr. Comput. 2019. [Google Scholar] [CrossRef]
- Păun, Gh. Computing with membranes. J. Comput. Syst. Sci. 2000, 61, 108–143. [Google Scholar] [CrossRef]
- Alsalibi, B.; Venkat, I.; Subramanian, K.; Lebai, L.; De, W. The impact of bio-inspired approaches toward the advancement of face recognition. ACM Comput. Surv. 2015, 48, 1–33. [Google Scholar] [CrossRef]
- Christinalh, A.; Díaz-Pernil, D.; Real, P. Region-based segmentation of 2D and 3D images with tissue-like P systems. Pattern Recognit. Lett. 2011, 32, 2206–2212. [Google Scholar] [CrossRef]
- Díaz-Pernil, D.; Gutiérrez-Naranjo, M.; Molina-Abril, H.; Real, P. Designing a new software tool for digital imagery based on P systems. Nat. Comput. 2012, 11, 381–386. [Google Scholar] [CrossRef]
- Carnero, J.; Díaz-Pernil, D.; Molina-Abril, H.; Real, P. Image segmentation inspired by cellular models using hardware programming. In Proceedings of the 3rd International Workshop on Computational Topology in Image Context, Chipiona, Spain, 10–12 November 2010; Volume 1, pp. 143–150. [Google Scholar]
- Díaz-Pernil, D.; Berciano, A.; Pen̆a-Cantillana, F.; Gutiérrez-Naranjo, M. Segmenting images with gradient-based edge detection using membrane computing. Pattern Recognit. Lett. 2013, 34, 846–855. [Google Scholar] [CrossRef]
- Peña-Cantillana, F.; Díaz-Pernil, D.; Christinal, H.; Gutiérrez-Naranjo, M. Implementation on CUDA of the smoothing problem with tissue-like P systems. Int. J. Nat. Comput. Res. 2011, 2, 25–34. [Google Scholar] [CrossRef]
- Alsalibi, B.; Venkat, I.; Subramanian, K.; Christinal, H. A bio-inspired software for homology groups of 2D digital images. In Proceedings of the Asian Conference on Membrane Computing (ACMC), Coimbatore, India, 18–19 September 2014; IEEE Computer Society: Washington, DC, USA, 2014. [Google Scholar] [CrossRef]
- Díaz-Pernil, D.; Christinal, H.; Gutiérrez-Naranjo, M.; Real, P. Using membrane computing for effective homology. Appl. Algebr. Eng. Commun. 2012, 23, 233–249. [Google Scholar] [CrossRef]
- Ardelean, I.; Díaz-Pernil, D.; Gutiérrez-Naranjo, M.; Pen̆a-Cantillana, F.; Reina-Molina, R.; Sarchizian, I. Counting cells with tissue-like P systems. In Proceedings of the Tenth Brainstorming Week on Membrane Computing, Seville, Spain, 30 January–3 February 2012; Fénix Editora: Seville, Spain, 2012. [Google Scholar]
- Reina-Molina, R.; Díaz-Pernil, D.; Gutiérrez-Naranjo, M. Cell complexes and membrane computing for thinning 2D and 3D images. In Proceedings of the Tenth Brainstorming Week on Membrane Computing, Seville, Spain, 30 January–3 February 2012; Fénix Editora: Seville, Spain, 2012. [Google Scholar]
- Berciano, A.; Díaz-Pernil, D.; Christinal, H.; Venkat, I. First steps for a corner detection using membrane computing. In Proceedings of the Asian Conference on Membrane Computing, Coimbatore, India, 18–19 September 2014; IEEE Computer Society: Washington, DC, USA, 2014. [Google Scholar]
- Enguix, G. Preliminaries about some possible applications of P systems in linguistics. Lect. Notes Comput. Sci. 2002, 2597, 74–89. [Google Scholar] [CrossRef]
- Cabarle, F.; de la Cruz, R.; Zhang, X.; Jiang, M.; Liu, X.; Zeng, X. On string languages generated by spiking neural P systems with structural plasticity. IEEE Trans. Nanobiosci. 2018, 17, 560–566. [Google Scholar] [CrossRef]
- Song, T.; Zeng, X.; Zheng, P.; Jiang, M.; Rodríguez-Patǿn, A. A parallel workflow pattern modelling using spiking neural P systems with colored spikes. IEEE Trans. Nanobiosci. 2018, 17, 474–484. [Google Scholar] [CrossRef] [PubMed]
- Mayne, R.; Phillips, N.; Adamatzky, A. Towards experimental P-systems using multivesicular liposomes. J. Membr. Comput. 2019. [Google Scholar] [CrossRef]
- Mitrana, V. Polarization: A new communication protocol in networks of bio-inspired processors. J. Membr. Comput. 2019. published online. [Google Scholar] [CrossRef]
- Pan, L.; Păun, Gh.; Zhang, G.; Neri, F. Spiking Neural P Systems with Communication on Request. Int. J. Neural Syst. 2017, 27, 1750042. [Google Scholar] [CrossRef]
- Orellana-Martín, D.; Valencia-Cabrera, L.; Riscos-Núñez, A.; Pérez-Jiménez, M.J. P systems with proteins: A new frontier when membrane division disappears. J. Membr. Comput. 2019. [Google Scholar] [CrossRef]
- Sánchez-Karhunen, E.; Valencia-Cabrera, L. Modelling complex market interactions using PDP systems. J. Membr. Comput. 2019. [Google Scholar] [CrossRef]
- Zhang, G.; Rong, H.; Neri, F.; Pérez-Jiménez, M.J. An optimization spiking neural P system for approximately solving combinatorial optimization problems. Int. J. Neural Syst. 2014, 24, 1440006. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; Cheng, J.; Gheorghe, M.; Meng, Q. A hybrid approach based on differential evolution and tissue membrane systems for solving constrained manufacturing parameter optimization problems. Appl. Soft Comput. 2013, 13, 1528–1542. [Google Scholar] [CrossRef]
- Wang, T.; Zhang, G.; Zhao, J.; He, Z.; Wang, J.; Pérez-Jiménez, M.J. Fault diagnosis of electric power systems based on fuzzy reasoning spiking neural P systems. IEEE Trans. Power Syst. 2015, 30, 1182–1194. [Google Scholar] [CrossRef]
- Martín-Vide, C.; Păun, Gh.; Pazos, J.; Rodríguez-Patón, A. Tissue P systems. Theor. Comput. Sci. 2003, 296, 295–326. [Google Scholar] [CrossRef]
- Ionescu, M.; Păun, Gh.; Yokomori, T. Spiking neural P systems. Fund. Inform. 2006, 71, 279–308. [Google Scholar] [CrossRef]
- Zeng, X.; Pan, L.Q.; Pérez-Jiménez, M.J. Small universal simple spiking neural P systems with weights. Sci. China Inf. Sci. 2014, 57, 1–11. [Google Scholar] [CrossRef]
- Păun, G.; Păun, R. Membrane computing and economics: numerical P systems. Fund. Inform. 2006, 73, 213–227. [Google Scholar]
- Zhang, Z.; Wu, T.; Paun, A.; Pan, L. Numerical P systems with migrating variables. Theor. Comput. Sci. 2016, 641, 85–108. [Google Scholar] [CrossRef]
- Pan, L.; Zhang, Z.; Wu, T.; Xu, J. Numerical P systems with production thresholds. Theor. Comput. Sci. 2017, 673, 30–41. [Google Scholar] [CrossRef]
- Zhang, Z.; Pan, L. Numerical P systems with thresholds. Int. J. Comput. Commun. 2017, 11, 292–304. [Google Scholar] [CrossRef]
- Buiu, C.; Vasile, C.; Arsene, O. Development of membrane controllers for mobile robots. Inform. Sci. 2012, 187, 33–51. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, G.; Neri, F.; Jiang, T.; Zhao, J.; Gheorghe, M.; Ipate, F.; Lefticaru, R. Design and implementation of membrane controllers for trajectory tracking of nonholonomic wheeled mobile robots. Integr. Comput.-Aid Eng. 2016, 23, 15–30. [Google Scholar] [CrossRef]
- Zhang, G.; Gheorghe, M.; Pérez-Jiminez, M.J. Real-Life Applications with Membrane Computing; Springer International Publishing AG: Cham, Switzerland, 2017; pp. 130–141. [Google Scholar]
- Mahalingam, K.; Rama, R.; Sureshkumar, W. Robot motion planning inside a grid using membrane computing. Int. J. Imaging Robot. 2017, 17, 33–51. [Google Scholar]
- Pavel, A.; Arsene, O.; Buiu, C. Enzymatic numerical P systems—A new class of membrane computing systems. In Proceedings of the Fifth International Conference on Bio-Inspired Computing: Theories and Applications, Changsha, China, 23–26 September 2010; IEEE Computer Society: Washington, DC, USA, 2010. [Google Scholar] [CrossRef]
- Zhang, Z.; Wu, T.; Paun, A.; Pan, L. Universal enzymatic numerical P systems with small number of enzymatic variables. Sci. China Inf. Sci. 2018, 61, 38–49. [Google Scholar] [CrossRef]
- Vasile, C.; Pavel, A.; Dumitrache, I.; Păun, Gh. On the power of enzymatic numerical P system. ACTA Inform. 2012, 49, 395–412. [Google Scholar] [CrossRef]
- Vasile, C.; Pavel, A.; Dumitrache, I. Universality of enzymatic numerical P systems. Int. J. Comput. Math. 2013, 90, 869–879. [Google Scholar] [CrossRef]
- Leporati, A.; Porreca, A.; Zandron, C.; Mauri, G. Improved universality results for parallel enzymatic numerical P systems. Int. J. Unconv. Comput. 2013, 9, 385–404. [Google Scholar]
- Pavel, A.; Buiu, C. Using enzymatic numerical P systems for modeling mobile robot controllers. Nat. Comput. 2012, 11, 387–393. [Google Scholar] [CrossRef]
- Li, W.; Yang, J.; Zhang, J. Handling big data field with enzymatic numerical P System. J. Sichuan Univ. Nat. Sci. Ed. 2013, 45, 96–104. [Google Scholar]
- Pang, S.C.; Ding, T.; Rodriguez-Paton, A.; Song, T.; Zheng, P. A parallel bioinspired framework for numerical calculations using enzymatic P system with an enzymatic environment. IEEE Access 2018, 6, 65548–65556. [Google Scholar] [CrossRef]
- Cecilia, J.; García, J.; Guerrero, G.; Martínet-del-Amor, M.; Pérez-Hurtado, I.; Pérez-Jiménez, M. Simulation of P systems with active membranes on CUDA. Brief Bioinform. 2009, 11, 313–322. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Bi, J.; Wang, L.; Wang, X. A non-reference evaluation method for edge detection of wear particles in ferrograph images. Mech. Syst. Signal Process. 2018, 100, 863–876. [Google Scholar] [CrossRef]



| Term | Necessary Resources |
|---|---|
| Initial number of cells | 1 |
| Number of enzymatic variables | |
| Number of numerical variables | |
| Number of rules | |
| Execution steps | 5 |
| Term | Parameters |
|---|---|
| CPU model | Intel(R) Core(TM) i7-7700HQ |
| cache memory | 8 MB, 16-Way, 64 byte lines |
| main memory | 16 GB (2* DDR4 2400MHz) |
| hard disc | SSD, SK hynix SC308 SATA 128GB, 600 Mbps; MQ01ABD100, 1TB |
| GPU model | Nvidia GeForce GTX 1050 Ti (4 GB) |
| execution steps | 5 |
| Reference [24] | Reference [21] | EDENP | |
|---|---|---|---|
| rice | 0.75 | 0.56 | 0.84 |
| cameraman | 0.66 | 0.32 | 0.74 |
| mri | 0.63 | 0.56 | 0.68 |
| AT3_lm4_01 | 0.44 | 0.12 | 0.5 |
| toyobjects | 0.85 | 0.76 | 0.86 |
| circbw | 0.94 | 0.93 | 0.95 |
| text | 0.93 | 0.90 | 0.94 |
| testpart1 | 0.81 | 0.79 | 0.86 |
| Image Resolution | Platform | ||||||
|---|---|---|---|---|---|---|---|
| Elapsed time(ms) | 0.014 | 0.03 | 0.05 | 0.12 | 0.23 | 0.86 | GPU |
| Elapsed time(ms) | 3.5 | 9.1 | 4.4 | 9.6 | 41.9 | 72.8 | CPU |
| Speedup ratio | 250 | 303 | 88 | 80 | 182 | 130 |
| Image Resolution | |||||||
|---|---|---|---|---|---|---|---|
| rice | 79 | 121 | 79 | 101 | 136 | 82 | |
| mri | 60 | 80 | 77 | 62 | 73 | 66 | |
| AT3_lm4_01 | 80 | 90 | 102 | 172 | 71 | 75 | |
| toyobjects | 187 | 162 | 163 | 81 | 182 | 62 | |
| circbw | 193 | 213 | 262 | 210 | 176 | 66 | |
| text | 167 | 180 | 194 | 118 | 57 | 65 | |
| testpart1 | 53 | 76 | 100 | 161 | 87 | 64 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yuan, J.; Guo, D.; Zhang, G.; Paul, P.; Zhu, M.; Yang, Q. A Resolution-Free Parallel Algorithm for Image Edge Detection within the Framework of Enzymatic Numerical P Systems. Molecules 2019, 24, 1235. https://doi.org/10.3390/molecules24071235
Yuan J, Guo D, Zhang G, Paul P, Zhu M, Yang Q. A Resolution-Free Parallel Algorithm for Image Edge Detection within the Framework of Enzymatic Numerical P Systems. Molecules. 2019; 24(7):1235. https://doi.org/10.3390/molecules24071235
Chicago/Turabian StyleYuan, Jianying, Dequan Guo, Gexiang Zhang, Prithwineel Paul, Ming Zhu, and Qiang Yang. 2019. "A Resolution-Free Parallel Algorithm for Image Edge Detection within the Framework of Enzymatic Numerical P Systems" Molecules 24, no. 7: 1235. https://doi.org/10.3390/molecules24071235
APA StyleYuan, J., Guo, D., Zhang, G., Paul, P., Zhu, M., & Yang, Q. (2019). A Resolution-Free Parallel Algorithm for Image Edge Detection within the Framework of Enzymatic Numerical P Systems. Molecules, 24(7), 1235. https://doi.org/10.3390/molecules24071235






