Curcumin and Resveratrol Regulate Intestinal Bacteria and Alleviate Intestinal Inflammation in Weaned Piglets
Abstract
1. Introduction
2. Results
2.1. The Level of Serum and Intestinal Interleukin and Immunoglobulin
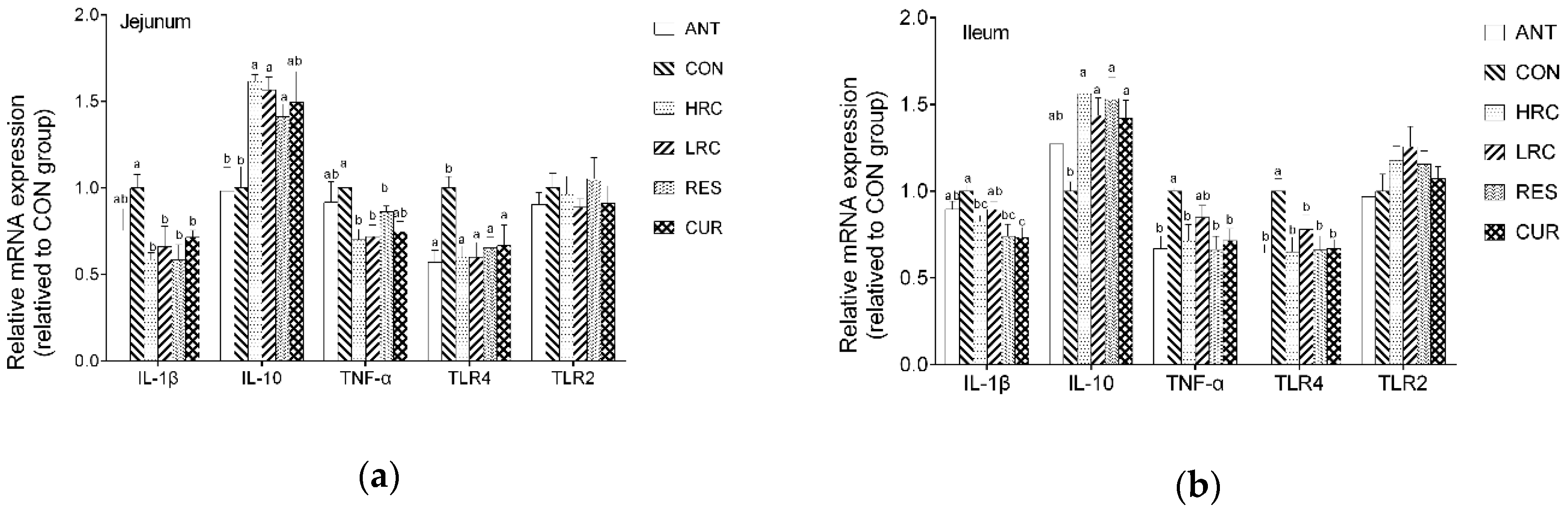
2.2. Messenger RNA Expression
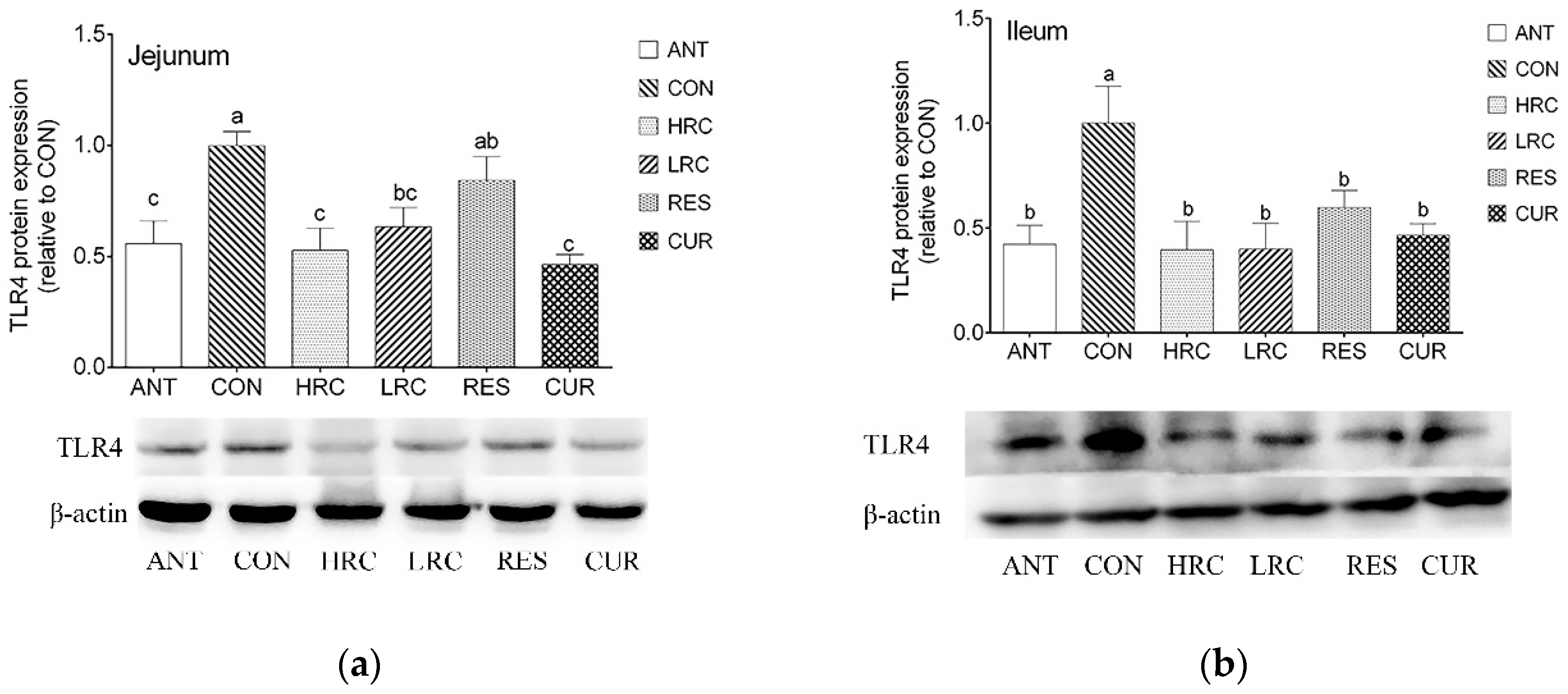
2.3. The Expression of TLR4 Protein
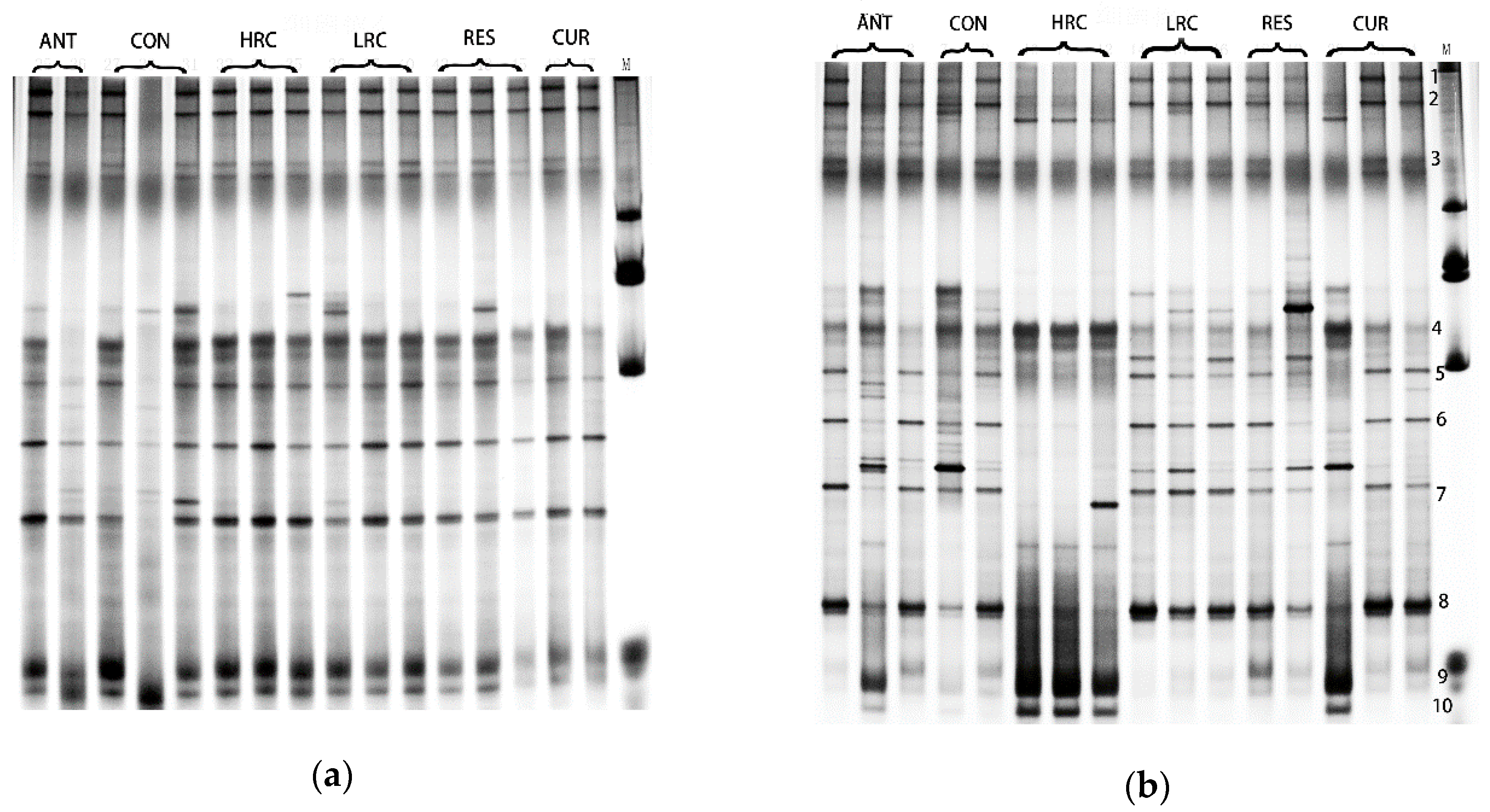
2.4. DGGE Analysis
2.5. DGGE Band Sequencing
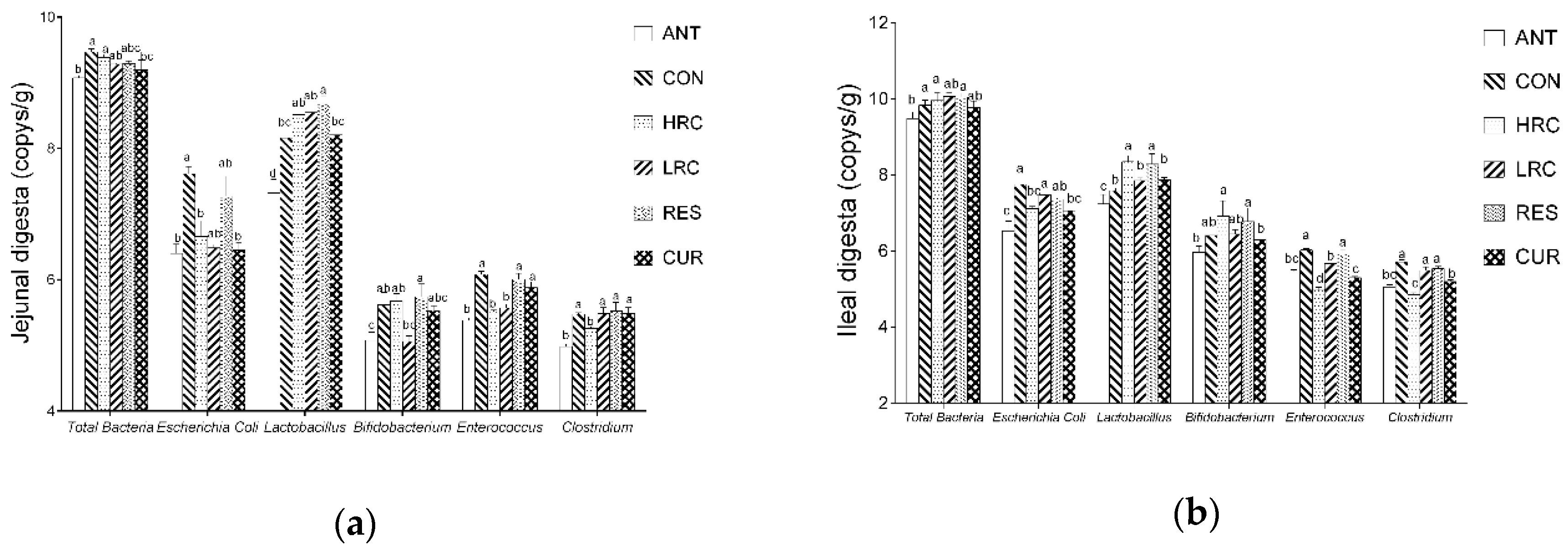
2.6. Bacteria Copy Numbers and Community
3. Discussion
4. Materials and Methods
4.1. Animals and Experimental Design
4.2. Sample Collection
4.3. Serum and Intestinal Cytokines
4.4. Total RNA Extraction and REAL-Time PCR
4.5. Western Blotting
4.6. DNA Isolation and Microbial Quantitative PCR
4.7. PCR Amplification and DGGE
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Mestecky, J.; Russell, M.W.; Elson, C.O. Intestinal IgA: Novel views on its function in the defence of the largest mucosal surface. Gut 1999, 44, 2–5. [Google Scholar] [CrossRef] [PubMed]
- Ouwehand, A.; Isolauri, E.; Salminen, S. The role of the intestinal microflora for the development of the immune system in early childhood. Eur. J. Nutr. 2002, 41, i32–i37. [Google Scholar] [CrossRef] [PubMed]
- Kau, A.L.; Ahern, P.P.; Griffin, N.W.; Goodman, A.L.; Gordon, J.I. Human nutrition, the gut microbiome and the immune system. Nature 2011, 474, 327–336. [Google Scholar] [CrossRef] [PubMed]
- Cebra, J.J. Influences of microbiota on intestinal immune system development. Amer. J. Clin. Nutr. 1999, 69, 1046s–1051s. [Google Scholar] [CrossRef]
- Hooper, L.V.; Littman, D.R.; Macpherson, A.J. Interactions between the microbiota and the immune system. Science 2012, 336, 1268–1273. [Google Scholar] [CrossRef] [PubMed]
- Nolan, J.P. The role of intestinal endotoxin in liver injury: A long and evolving history. Hepatology 2010, 52, 1829–1835. [Google Scholar] [CrossRef]
- Turner, J.R. Intestinal mucosal barrier function in health and disease. Nature Rev. Immunol. 2009, 9, 799–809. [Google Scholar] [CrossRef] [PubMed]
- Moeser, A.J.; Ryan, K.A.; Nighot, P.K.; Blikslager, A.T. Gastrointestinal dysfunction induced by early weaning is attenuated by delayed weaning and mast cell blockade in pigs. Am. J. Physiol. Gastrointest. Liver Physiol. 2007, 293, G413. [Google Scholar] [CrossRef] [PubMed]
- Kloubert, V.; Blaabjerg, K.; Dalgaard, T.S.; Poulsen, H.D.; Rink, L.; Wessels, I. Influence of zinc supplementation on immune parameters in weaned pigs. J. Trace Elements Med. Biol. 2018, 44, 231–240. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.B.; Piao, X.S.; Kim, S.W.; Wang, L.; Liu, P.; Yoon, I.; Zhen, Y.G. Effects of yeast culture supplementation on growth performance, intestinal health, and immune response of nursery pigs. J. Animal Sci. 2009, 87, 2614–2624. [Google Scholar] [CrossRef]
- Resende, J.A.; Silva, V.L.; Oliveira, T.L.R.D.; Fortunato, S.D.O.; Carneiro, J.D.C.; Otenio, M.H.; Diniz, C.G. Prevalence and persistence of potentially pathogenic and antibiotic resistant bacteria during anaerobic digestion treatment of cattle manure. Bioresour. Technol. 2014, 153, 284–291. [Google Scholar] [CrossRef] [PubMed]
- Martin, B. Antibiotic overuse: Stop the killing of beneficial bacteria. Nature 2011, 476, 393–394. [Google Scholar]
- Jung, C.M.; Heinze, T.M.; Schnackenberg, L.K.; Mullis, L.B.; Elkins, S.A.; Elkins, C.A.; Steele, R.S.; Sutherland, J.B. Interaction of dietary resveratrol with animal-associated bacteria. Fems Microbiol. Lett. 2010, 297, 266–273. [Google Scholar] [CrossRef] [PubMed]
- Iyori, M.; Kataoka, H.; Shamsul, H.M.; Kiura, K.; Yasuda, M.; Nakata, T.; Hasebe, A.; Shibata, K. Resveratrol modulates phagocytosis of bacteria through an NF-kappaB-dependent gene program. Anti. Agents Chemother. 2008, 52, 121–127. [Google Scholar] [CrossRef] [PubMed]
- Mun, S.H.; Joung, D.K.; Kim, Y.S.; Kang, O.H.; Kim, S.B.; Seo, Y.S.; Kim, Y.C.; Lee, D.S.; Shin, D.W.; Kweon, K.T. Synergistic antibacterial effect of curcumin against methicillin-resistant Staphylococcus aureus. Phytomedicine 2013, 20, 714–718. [Google Scholar] [CrossRef] [PubMed]
- Bisht, K.; Wagner, K.H.; Bulmer, A.C. Curcumin, resveratrol and flavonoids as anti-inflammatory, cyto- and DNA-protective dietary compounds. Toxicology 2010, 278, 88–100. [Google Scholar] [CrossRef] [PubMed]
- Moghadamtousi, S.Z.; Kadir, H.A.; Hassandarvish, P.; Tajik, H.; Abubakar, S.; Zandi, K. A Review on Antibacterial, Antiviral, and Antifungal Activity of Curcumin. Biomed. Res. Int. 2014, 2014, 186864. [Google Scholar] [PubMed]
- Amri, A.; Chaumeil, J.C.; Sfar, S.; Charrueau, C. Administration of resveratrol: What formulation solutions to bioavailability limitations? J. Control. Release Off. J. Control. Release Soc. 2012, 158, 182–193. [Google Scholar] [CrossRef] [PubMed]
- Anand, P.; Kunnumakkara, A.B.; Newman, R.A.; Aggarwal, B.B. Bioavailability of curcumin: Problems and promises. J. Mol. Pharm. 2007, 4, 807–818. [Google Scholar] [CrossRef]
- Briskey, D.; Sax, A.; Mallard, A.R.; Rao, A. Increased bioavailability of curcumin using a novel dispersion technology system (LipiSperse®). Eur. J. Nutr. 2018, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Mohammadi, F.; Moeeni, M. Study on the interactions of trans-resveratrol and curcumin with bovine α-lactalbumin by spectroscopic analysis and molecular docking. Mater. Sci. Eng. C Mater. Biol. Appl. 2015, 50, 358–366. [Google Scholar] [CrossRef] [PubMed]
- Mehdi, S.; Ali, M.; Constanze, B. Curcumin synergizes with resveratrol to stimulate the MAPK signaling pathway in human articular chondrocytes in vitro. Genes Nutr. 2011, 6, 171. [Google Scholar]
- Laura, M.; Enrica, D.S.; Massimo, F.; Rosanna, M.; Monica, B.; Laura, M.; Pamela, S.; Chiara, F.; Roberta, B.; Ilaria, T. Resveratrol potentiates the in vitro and in vivo anti-tumoral effects of curcumin in head and neck carcinomas. Oncotarget 2014, 5, 10745–10762. [Google Scholar]
- Pandey, K.B.; Rizvi, S.I. Resveratrol may protect plasma proteins from oxidation under conditions of oxidative stress in vitro. J. Brazil. Chem. Soc. 2010, 21, 909–913. [Google Scholar] [CrossRef]
- De, K.S.; Tobin, M.C.; Forsyth, C.B.; Keshavarzian, A.; Landay, A.L. Regulation of Intestinal Immune Responses through TLR Activation: Implications for Pro- and Prebiotics. J. Front Immunol. 2014, 5, 60. [Google Scholar]
- Frazier, T.H.; DiBaise, J.K.; McClain, C.J. Gut microbiota, intestinal permeability, obesity-induced inflammation, and liver injury. JPEN J. Parent. Enteral Nutr. 2011, 35, 14s–20s. [Google Scholar] [CrossRef] [PubMed]
- Lu, N.; Li, X.; Yu, J.; Li, Y.; Zhong, X. Curcumin Attenuates Lipopolysaccharide-Induced Hepatic Lipid Metabolism Disorder by Modification of m 6 A RNA Methylation in Piglets. Lipids 2018, 53, 53–63. [Google Scholar] [CrossRef]
- Sánchez-Fidalgo, S.; Cárdeno, A.; Villegas, I.; Talero, E.; Lastra, C.A.D.L. Dietary supplementation of resveratrol attenuates chronic colonic inflammation in mice. Eur. J. Pharm. 2010, 633, 78–84. [Google Scholar] [CrossRef] [PubMed]
- Antonio Ramón, M.; Isabel, V.; Marina, S.H.; Catalina Alarcón, D.L.L. The effects of resveratrol, a phytoalexin derived from red wines, on chronic inflammation induced in an experimentally induced colitis model. Br. J. Pharmacol. 2010, 147, 873–885. [Google Scholar]
- Bereswill, S.; Muñoz, M.; Fischer, A.; Plickert, R.; Haag, L.M.; Otto, B.; Kühl, A.A.; Loddenkemper, C.; Göbel, U.B.; Heimesaat, M.M. Anti-inflammatory effects of resveratrol, curcumin and simvastatin in acute small intestinal inflammation. PLoS ONE 2010, 5, e15099. [Google Scholar] [CrossRef] [PubMed]
- Song, W.B.; Wang, Y.Y.; Meng, F.S.; Zhang, Q.H.; Zeng, J.Y.; Xiao, L.P.; Yu, X.P.; Peng, D.D.; Su, L.; Xiao, B. Curcumin protects intestinal mucosal barrier function of rat enteritis via activation of MKP-1 and attenuation of p38 and NF-κB activation. PLoS ONE 2010, 5, e12969. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Xu, Y.; Chen, X.; Fang, C.; Zhao, L.; Chen, F. The Maturing Development of Gut Microbiota in Commercial Piglets during the Weaning Transition. Front. Microbiol. 2017, 8, 1688. [Google Scholar] [CrossRef] [PubMed]
- van Deventer, S.J.; ten Cate, J.W.; Tytgat, G.N. Intestinal endotoxemia. Clinical significance. Gastroenterology 1988, 94, 825–831. [Google Scholar] [CrossRef]
- Nguyen, T.V.; Van, P.L.; Huy, C.L.; Gia, K.N.; Weintraub, A. Detection and characterization of diarrheagenic Escherichia coli from young children in Hanoi, Vietnam. J. Clin. Microbiol. 2005, 43, 755–760. [Google Scholar] [CrossRef] [PubMed]
- Tsai, C.H.; Chen, Y.Y.; Wang, K.G.; Chen, C.Y.; Chen, C.P. Characteristics of early-onset neonatal sepsis caused by Escherichia coli. Taiwanese J. Obstetr. Gynecol. 2012, 51, 26–30. [Google Scholar] [CrossRef] [PubMed]
- Kristich, C.J.; Rice, L.B.; Arias, C.A. Enterococcal Infection-Treatment and Antibiotic Resistance. In Enterococci: From Commensals to Leading Causes of Drug Resistant Infection; Gilmore, M.S., Clewell, D.B., Ike, Y., Shankar, N., Eds.; Massachusetts Eye and Ear Infirmary: Boston, MA, USA, 2014. [Google Scholar]
- Tamara, S.; Michiel, W.; Justyna, P.; Christian, C.; Sylvain, B.; Jos, B.; Van Hylckama Vlieg, J.E.; Siezen, R.J. Lactobacillus paracasei comparative genomics: Towards species pan-genome definition and exploitation of diversity. PLoS ONE 2013, 8, e68731. [Google Scholar]
- Mccann, K.S.J.N. The diversity–stability debate. Nature 2000, 405, 228–233. [Google Scholar] [CrossRef]
- Walle, T. Bioavailability of resveratrol. Ann. N. Y. Acad. Sci. 2011, 1215, 9–15. [Google Scholar] [CrossRef]
- Purpura, M.; Lowery, R.P.; Wilson, J.M.; Mannan, H.; Munch, G.; Razmovski-Naumovski, V. Analysis of different innovative formulations of curcumin for improved relative oral bioavailability in human subjects. Eur. J. Nutr. 2018, 57, 929–938. [Google Scholar] [CrossRef] [PubMed]
- Katie, F.; Carol, P. The ecology, epidemiology and virulence of Enterococcus. Microbiology 2009, 155, 1749–1757. [Google Scholar]
- Ryan, K.J. Sherris Medical Microbiology: An Introduction to Infectious Diseases; Elsevier: Amsterdam, The Netherlands, 1990. [Google Scholar]
- Mylonakis, E.; Ryan, E.T.; Calderwood, S.B. Clostridium difficile--Associated diarrhea: A review. Arch. Int. Med. 2001, 161, 525–533. [Google Scholar] [CrossRef]
- Mencin, A.; Kluwe, J.; Schwabe, R.F. Toll-like receptors as targets in chronic liver diseases. Gut 2009, 58, 704–720. [Google Scholar] [CrossRef] [PubMed]
- Kaszowska, M. Chemical structure and biosynthesis of lipopolysaccharide—Important component of the cell envelope of Gram-negative bacteria. Postepy higieny i medycyny doswiadczalnej (Online) 2004, 58, 333–342. [Google Scholar]
- Dong, H.H.; Tang, X.D.; Zhang, Z.Y.; Dong, C.J. Structural insight into lipopolysaccharide transport from the Gram-negative bacterial inner membrane to the outer membrane. Bio. Et Biophy. Acta-Mol. Cell Biol. Lipids 2017, 1862, 1461–1467. [Google Scholar] [CrossRef] [PubMed]
- Galili, U.; Mandrell, R.E.; Hamadeh, R.M.; Shohet, S.B.; Griffiss, J.M. Interaction between human natural anti-alpha-galactosyl immunoglobulin G and bacteria of the human flora. J. Infect. 1988, 56, 1730. [Google Scholar]
- Heimesaat, M.M.; Fischer, A.; Jahn, H.K.; Niebergall, J.; Freudenberg, M.; Blaut, M.; Liesenfeld, O.; Schumann, R.R.; Göbel, U.B.; Bereswill, S. Exacerbation of murine ileitis by Toll-like receptor 4 mediated sensing of lipopolysaccharide from commensal Escherichia coli. Gut 2007, 56, 941–948. [Google Scholar] [CrossRef] [PubMed]
- Lenka, F.; Pavel, D.; Pavel, R.; Klara, K.; Helena, T.H. Expression of Toll-like receptor 2 (TLR2), TLR4, and CD14 in biopsy samples of patients with inflammatory bowel diseases: Upregulated expression of TLR2 in terminal ileum of patients with ulcerative colitis. J. Histochem. 2008, 56, 267–274. [Google Scholar]
- Youn, H.S.; Lee, J.Y.; Fitzgerald, K.A.; Young, H.A.; Shizuo, A.; Hwang, D.H. Specific inhibition of MyD88-independent signaling pathways of TLR3 and TLR4 by resveratrol: Molecular targets are TBK1 and RIP1 in TRIF complex. J. Immunol. 2005, 175, 3339–3346. [Google Scholar] [CrossRef]
- Fu, Y.; Gao, R.; Cao, Y.; Guo, M.; Wei, Z.; Zhou, E.; Li, Y.; Yao, M.; Yang, Z.; Zhang, N. Curcumin attenuates inflammatory responses by suppressing TLR4-mediated NF-κB signaling pathway in lipopolysaccharide-induced mastitis in mice. Int. Immunopharmacol. 2014, 20, 54–58. [Google Scholar] [CrossRef] [PubMed]
- Winters, B.D.; Eberlein, M.; Leung, J.; Needham, D.M.; Pronovost, P.J.; Sevransky, J.E. Long-term mortality and quality of life in sepsis: A systematic review. Critical Care Med. 2010, 38, 1276. [Google Scholar] [CrossRef]
- Starke, R.M.; Raper, D.M.; Ding, D.; Chalouhi, N.; Owens, G.K.; Hasan, D.M.; Medel, R.; Dumont, A.S. Tumor necrosis factor-alpha modulates cerebral aneurysm formation and rupture. Trans. Stroke Res. 2014, 5, 269–277. [Google Scholar] [CrossRef]
- Ma, T.Y.; Iwamoto, G.K.; Hoa, N.T.; Akotia, V.; Pedram, A.; Boivin, M.A.; Said, H.M. TNF-alpha-induced increase in intestinal epithelial tight junction permeability requires NF-kappa B activation. Am. J. Physiol. Gastrointest Liver Phys. 2004, 286, G367–G376. [Google Scholar] [CrossRef]
- Al-Sadi, R.M.; Ma, T.Y. IL-1beta causes an increase in intestinal epithelial tight junction permeability. J. Immunol. 2007, 178, 4641–4649. [Google Scholar] [CrossRef]
- Ai, X.L.; Yao, F.; Wang, X.J.; Duan, D.B.; Li, K.; Hu, Z.Y.; Yin, G.; Wang, M.; Wu, B.Y. Role of allograft inflammatory factor-1 in regulating the proliferation, migration and apoptosis of colorectal cancer cells. Nan Fang yi ke da xue xue bao = J. Southern Med. Univ. 2018, 38, 511–519. [Google Scholar]
- Su, W.; Zhang, H.; Ying, Z.; Li, Y.; Zhou, L.; Wang, F.; Zhang, L.; Wang, T. Effects of dietary L-methionine supplementation on intestinal integrity and oxidative status in intrauterine growth-retarded weanling piglets. Eur. J. Nutr. 2017, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Han, G.Q.; Yu, B.; Chen, D.W.; Mao, X.B.; Chen, H.; Mao, Q.; Huang, Z.Q. Effects of different starch sources on Bacillus spp. in intestinal tract and expression of intestinal development related genes of weanling piglets. Mol. Biol. Rep. 2012, 39, 1869–1876. [Google Scholar] [CrossRef]
- Rinttil, T.; Kassinen, A.; Malinen, E.; Krogius, L.; Palva, A. Development of an extensive set of 16S rDNA-targeted primers for quantification of pathogenic and indigenous bacteria in faecal samples by real-time PCR. J. Appl. Microbiol. 2010, 97, 1166–1177. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Mao, X.; He, J.; Yu, B.; Huang, Z.; Yu, J.; Zheng, P.; Chen, D. Dietary fibre affects intestinal mucosal barrier function and regulates intestinal bacteria in weaning piglets. Br. J. Nutr. 2013, 110, 1837–1848. [Google Scholar] [CrossRef]
- Sanguinetti, C.J.; Dias Neto, E.; Simpson, A.J. Rapid silver staining and recovery of PCR products separated on polyacrylamide gels. Biotechniques 1994, 17, 914–921. [Google Scholar]
- Keylock, C. Simpson diversity and the Shannon-Wiener index as special cases of a generalized entropy. Oikos 2010, 109, 203–207. [Google Scholar] [CrossRef]
Sample Availability: Samples of the compounds curcumin and resveratrol are available from the author X.Z. |
| Items | ANT | CON | HRC | LRC | RES | CUR |
|---|---|---|---|---|---|---|
| Jejunum | ||||||
| IL-1β (ng/g prot) | 28.46 ± 3.05 ab | 31.53 ± 2.18 a | 18.77 ± 1.08 c | 25.18 ± 2.71 c | 22.69 ± 0.92 bc | 23.44 ± 1.13 bc |
| TNF-α (pg/mg prot) | 157.05 ± 7.34 a | 167.52 ± 1.96 a | 123.82 ± 5.79 b | 172.10 ± 4.51 a | 121.56 ± 4.16 b | 122.51 ± 12.89 b |
| IL-10 (pg/mg prot) | 78.85 ± 2.20 c | 88.99 ± 4.03 bc | 92.38 ± 2.35 ab | 97.57 ± 4.29 a | 90.92 ± 3.98 ab | 91.21 ± 1.28 ab |
| IgG (μg/mg prot) | 16.10 ± 0.55 | 15.74 ± 0.69 | 15.89 ± 0.50 | 15.84 ± 0.74 | 15.78 ± 0.37 | 15.19 ± 0.27 |
| IgA (pg/mg prot) | 80.56 ± 2.03 | 78.98 ± 1.89 | 79.22 ± 1.57 | 80.24 ± 2.41 | 78.15 ± 1.66 | 76.66 ± 0.77 |
| Ileum | ||||||
| IL-1β (ng/g prot) | 30.33 ± 2.77 b | 40.15 ± 2.44 a | 31.44 ± 1.06 b | 39.84 ± 2.21 a | 35.52 ± 1.08 ab | 34.49 ± 2.46 ab |
| TNF-α (pg/mg prot) | 163.56 ± 4.81 b | 198.63 ± 5.25 a | 121.40 ± 8.08 c | 131.00 ± 3.39 c | 167.21 ± 3.80 b | 176.78 ± 7.84 b |
| IL-10(pg/mg prot) | 115.90 ± 4.59 ab | 109.47 ± 4.89 b | 119.94 ± 4.24 ab | 134.31 ± 6.36 a | 122.49 ± 9.59 ab | 127.26 ± 8.07 ab |
| IgG (μg/mg prot) | 15.25±0.61 a | 12.41 ± 0.59 b | 16.39 ± 0.57 a | 15.66 ± 0.93 a | 14.99 ± 0.55 a | 15.73 ± 0.50 a |
| IgA (pg/mg prot) | 81.23 ± 2.16 a | 74.80 ± 0.95 b | 83.24 ± 2.67 a | 80.55 ± 2.33 a | 74.25 ± 1.27 b | 74.25 ± 1.27 b |
| Serum | ||||||
| IgA (pg/mL) | 25.38 ± 0.2 b | 24.53 ± 0.16 c | 26.54 ± 0.20 a | 25.42 ± 0.26 b | 26.10 ± 0.17 a | 25.45 ± 0.14 b |
| IgG (pg/mL) | 131.17 ± 1.64 bc | 122.21 ± 1.86 d | 137.49 ± 0.95 a | 129.08 ± 0.98 c | 136.43 ± 3.89 ab | 128.11 ± 1.42 cd |
| Items | H (Shannon Index) | E (Evenness) | S (Species Richness) |
|---|---|---|---|
| Jejunum | |||
| ANT | 2.38 | 0.97 | 11 |
| CON | 2.96 | 0.97 | 14 |
| HRC | 2.79 | 0.98 | 14 |
| LRC | 2.60 | 0.99 | 14 |
| RES | 2.89 | 0.99 | 13 |
| CUR | 2.73 | 0.97 | 13 |
| Ileum | |||
| ANT | 2.61 | 0.97 | 15 |
| CON | 2.86 | 0.97 | 20 |
| HRC | 2.95 | 0.98 | 22 |
| LRC | 2.84 | 0.98 | 20 |
| RES | 3.05 | 0.96 | 24 |
| CUR | 2.73 | 0.97 | 17 |
| Band Number | Closest Relative Bacteria | NCBI Accession Number | Identity |
|---|---|---|---|
| Band 1 | Nostoc punctiforme | NR_074317.1 | 92% |
| Band 2 | Streptococcus ferus | NR_115276.1 | 100% |
| Band 3 | Uncultured bacterium | JX183833.1 | 100% |
| Band 4 | Methanocaldococcus bathoardescens | NR_134839.1 | 97% |
| Band 5 | Enterococcus alcedinis | NR_109727.1 | 98% |
| Band 6 | Enterococcus hirae | NR_114783.2 | 99% |
| Band 7 | Clostridium bornimense | NR_134005.1 | 99% |
| Band 8 | Barnesiella viscericola | NR_121773.2 | 95% |
| Band 9 | Chroococcidiopsis thermalis | NR_102464.1 | 98% |
| Band 10 | Lactobacillus johnsonii | NR_075064.1 | 97% |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gan, Z.; Wei, W.; Li, Y.; Wu, J.; Zhao, Y.; Zhang, L.; Wang, T.; Zhong, X. Curcumin and Resveratrol Regulate Intestinal Bacteria and Alleviate Intestinal Inflammation in Weaned Piglets. Molecules 2019, 24, 1220. https://doi.org/10.3390/molecules24071220
Gan Z, Wei W, Li Y, Wu J, Zhao Y, Zhang L, Wang T, Zhong X. Curcumin and Resveratrol Regulate Intestinal Bacteria and Alleviate Intestinal Inflammation in Weaned Piglets. Molecules. 2019; 24(7):1220. https://doi.org/10.3390/molecules24071220
Chicago/Turabian StyleGan, Zhending, Wenyao Wei, Yi Li, Jiamin Wu, Yongwei Zhao, Lili Zhang, Tian Wang, and Xiang Zhong. 2019. "Curcumin and Resveratrol Regulate Intestinal Bacteria and Alleviate Intestinal Inflammation in Weaned Piglets" Molecules 24, no. 7: 1220. https://doi.org/10.3390/molecules24071220
APA StyleGan, Z., Wei, W., Li, Y., Wu, J., Zhao, Y., Zhang, L., Wang, T., & Zhong, X. (2019). Curcumin and Resveratrol Regulate Intestinal Bacteria and Alleviate Intestinal Inflammation in Weaned Piglets. Molecules, 24(7), 1220. https://doi.org/10.3390/molecules24071220








