Metal-Ligand Recognition Index Determination by NMR Proton Relaxation Study
Abstract
1. Introduction
2. Results and Discussion
2.1. Theory
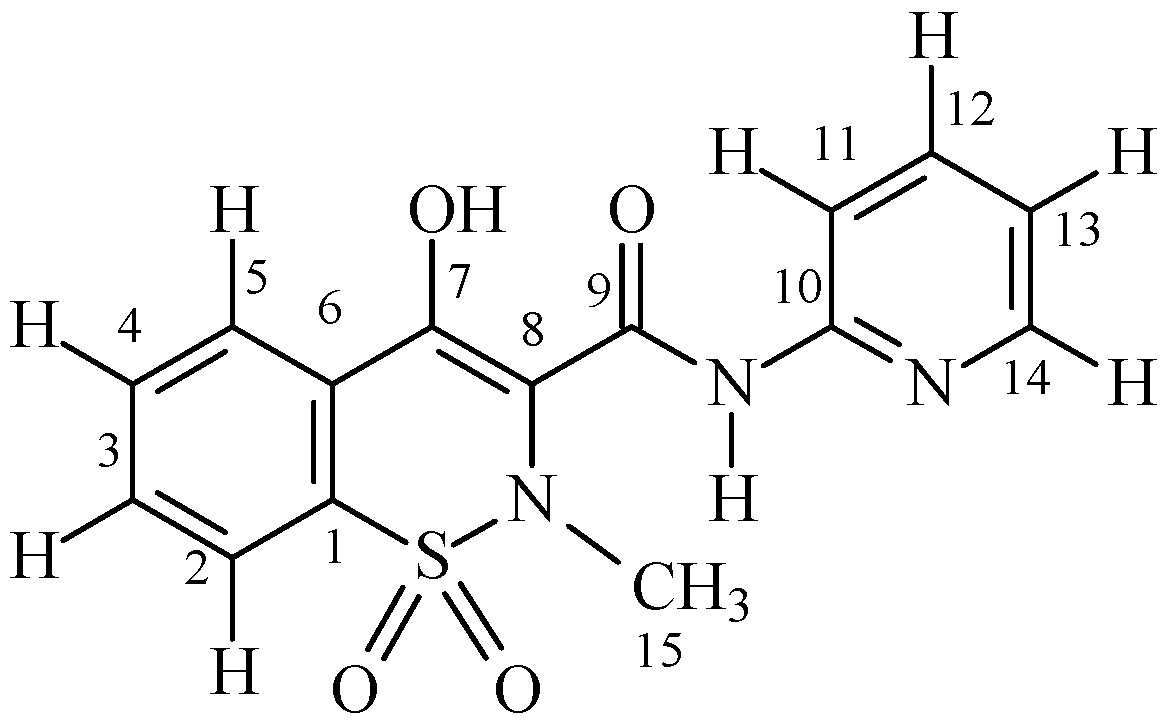
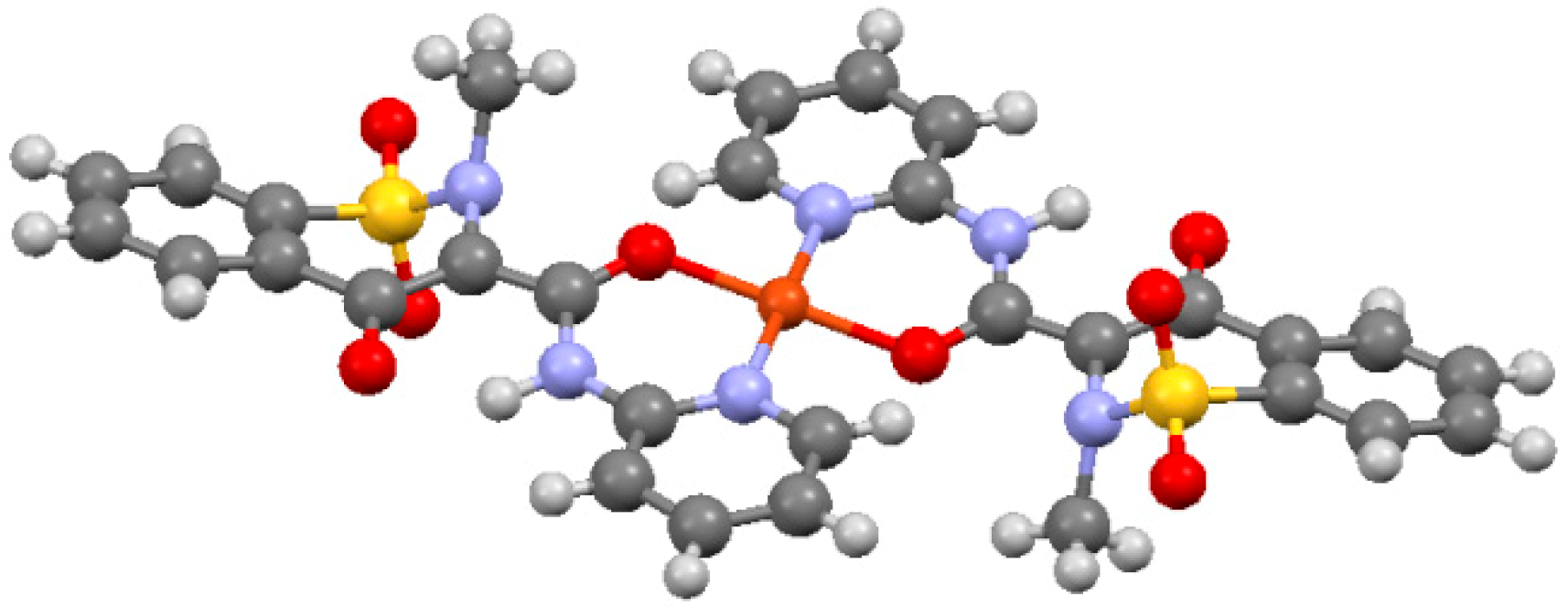
2.2. The case of Cu(II)-Piroxicam Complex
3. Materials and Methods
3.1. Materials
3.2. NMR Measurements
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Riccardi, M.L.; Genna, V.; De Vivo, M. Metal-ligand interaction in drug design. Nat. Rev. Chem. 2018, 2, 100–112. [Google Scholar] [CrossRef]
- Cook, S.A.; Borovik, A.S. Molecular designs for controlling the local environments around metal ions. Acc. Chem. Res. 2015, 48, 2407–2414. [Google Scholar] [CrossRef] [PubMed]
- Barone, C.R.; Cini, R.; de Pinto, S.; Di Masi, N.G.; Maresca, L.; Natile, G.; Tamasi, G. Coupling of cationic olefin complexes of platinum(II) with potential ambident nucleophiles. Inorg. Chim. Acta 2010, 363, 205–212. [Google Scholar] [CrossRef]
- Pontillo, N.; Ferraro, G.; Messori, L.; Tamasi, G.; Merlino, A. Ru-Based CO releasing molecules with azole ligands: Interaction with proteins and the CO release mechanism disclosed by X-ray crystallography. Dalton Trans. 2017, 46, 9621–9629. [Google Scholar] [CrossRef]
- Sun, L.-Z.; Zhang, D.; Chen, S.-J. Theory and modeling of RNA structure and interactions with metal ions and small molecules. Ann. Rev. Biophys. 2017, 46, 227–246. [Google Scholar] [CrossRef] [PubMed]
- Bush, A.I. Metal complexing agents as therapies for Alzheimer’s disease. Neurobiol. Aging 2002, 23, 1031–1038. [Google Scholar] [CrossRef]
- Tamasi, G.; Carpini, A.; Valensin, D.; Messori, L.; Pratesi, A.; Scaletti, F.; Jakupec, M.; Keppler, B.; Cini, R. {Ru(CO)x}-core complexes with selected azoles: Synthesis, X-ray structure, spectroscopy, DFT analysis and evaluation of cytotoxic activity against human cancer cells. Polyhedron 2014, 81, 227–237. [Google Scholar] [CrossRef]
- Tamasi, G.; Merlino, A.; Scaletti, F.; Heffeter, P.; Legin, A.A.; Jakupec, M.A.; Berger, W.; Messori, L.; Keppler, B.K.; Cini, R. {Ru(CO)x}-core complexes with benzimidazole ligands: Synthesis, X-ray structure and evaluation of anticancer activity in vivo. Dalton Trans. 2017, 46, 3025–3040. [Google Scholar] [CrossRef] [PubMed]
- Bruijnincx, P.C.A.; Sadler, P.J. New trends for metal complexes with anticancer activity. Curr. Opin. Chem. Biol. 2008, 12, 197–206. [Google Scholar] [CrossRef] [PubMed]
- Tamasi, G.; Bonechi, C.; Rossi, C.; Cini, R.; Magnani, A. Simulating the active sites of Copper trafficking proteins. Density Functional Structural and spectroscopy studies on Copper(I) complexes with thiols, carboxylato, amide and phenol ligands. J. Coord. Chem. 2016, 69, 404–424. [Google Scholar] [CrossRef]
- Tamasi, G.; Bernini, C.; Corbini, G.; Owens, N.; Messori, L.; Scaletti, F.; Massai, L.; Lo Giudice, P.; Cini, R. Synthesis, spectroscopic and DFT structural characterization of two novel Ruthenium(III) Oxicam complexes. In vivo evaluation of anti-inflammatory and gastric damaging activities. J. Inorg. Biochem. 2014, 134, 25–35. [Google Scholar] [CrossRef]
- Allardyce, C.S.; Dyson, P.J. Ruthenium in medicine: Current clinical uses and future prospects. Plat. Met. Rev. 2001, 45, 62–69. [Google Scholar]
- Tamasi, G.; Defazio, S.; Chiasserini, L.; Sega, A.; Cini, R. Ruthenium-thiobase complexes: Synthesis, spectroscopy, density functional studies for trans,cis,cis-[RuII(AsPh3)2(N,S-2-Thiopyrimidinato)2] and structural analysis of selected weak C–H⋯N and C–H⋯S interactions. Inorg. Chim. Acta 2009, 362, 1011–1021. [Google Scholar] [CrossRef]
- Zhang, C.X.; Lippard, S.J. New metal complexes as potential therapeutics. Curr. Opin. Chem. Biol. 2003, 7, 481–489. [Google Scholar] [CrossRef]
- Iniguez, E.; Sanchez, A.; Vasquez, M.A.; Martınez, A.; Olivas, J.; Sattler, A.; Sanchez-Delgado, R.A.; Maldonado, R.A. Metal-drug synergy: New ruthenium(II) complexes of ketoconazole are highly active against Leishmania major and Trypanosoma cruzi and nontoxic to human or murine normal cells. J. Biol. Inorg. Chem. 2013, 18, 779–790. [Google Scholar] [CrossRef] [PubMed]
- Adrees, M.; Ali, S.; Rizwan, M.; Zia-ur-Rehman, M.; Ibrahim, M.; Abbas, F.; Farid, M.; Qayyum, M.F.; Irshad, M.K. Mechanisms of silicon-mediated alleviation of heavy metal toxicity in plants: A review. Ecotoxicol. Environ. Saf. 2015, 119, 186–197. [Google Scholar] [CrossRef]
- Hernández, L.E.; Sobrino-Plata, J.; Montero-Palmero, M.B.; Carrasco-Gil, S.; Flores-Cáceres, M.L.; Ortega-Villasante, C.; Escobar, C. Contribution of glutathione to the control of cellular redox homeostasis under toxic metal and metalloid stress. J. Exp. Bot. 2015, 66, 2901–2911. [Google Scholar] [CrossRef]
- Fuertes, M.A.; Alonso, C.; Perez, J.M. Biochemical modulation of Cisplatin mechanisms of action: Enhancement of antitumor activity and circumvention of drug resistance. Chem. Rev. 2003, 103, 645–662. [Google Scholar] [CrossRef] [PubMed]
- Romero-Canelon, I.; Sadler, P.J. Next generation of metal anticancer complexes: Multitargeting via redox modulation. Inorg. Chem. 2013, 52, 12276–12291. [Google Scholar] [CrossRef] [PubMed]
- Graf, N.; Lippard, S.J. Redox activation of metal-based prodrugs as a strategy for drug delivery. Adv. Drug Del. Rev. 2012, 64, 993–1004. [Google Scholar] [CrossRef]
- Wu, M.X.; Yang, Y.W. Metal-Organic framework (MOF)-based drug/Cargo delivery and cancer therapy. Adv. Mater. 2017, 29, 1606134–1606153. [Google Scholar] [CrossRef] [PubMed]
- Tamasi, G.; Mangani, S.; Cini, R. Copper(I)-alkyl sulfide and -cysteine tri-nuclear clusters as models for metallo proteins: A structural density functional analysis. J. Biomol. Struct. Dyn. 2012, 30, 728–751. [Google Scholar] [CrossRef] [PubMed]
- Bonechi, C.; Donati, A.; Tamasi, G.; Leone, G.; Consumi, M.; Rossi, C.; Lamponi, S.; Magnani, A. Protective effect of quercetin and rutin encapsulated liposomes on induced oxidative stress. Biophys. Chem. 2018, 233, 55–63. [Google Scholar] [CrossRef]
- Leone, G.; Consumi, M.; Pepi, S.; Lamponi, S.; Bonechi, C.; Tamasi, G.; Donati, A.; Rossi, C.; Magnani, A. New formulations to enhance lovastatin release from Red Yeast Rice (RYR). J. Drug Deliv. Sci. Technol. 2016, 36, 110–119. [Google Scholar] [CrossRef]
- Moretti, E.; Mazzi, L.; Bonechi, C.; Salvatici, M.C.; Iacoponi, F.; Rossi, C.; Collodel, G. Effect of Quercetin-loaded liposomes on induced oxidative stress in human spermatozoa. Reprod. Toxicol. 2016, 60, 140–147. [Google Scholar] [CrossRef]
- Tardi, P.; Johnstone, S.; Webb, M.; Bally, M.; Abraham, S. Liposome Loading with Metal Ions. U.S. Patent 7,238,367 B2, 3 July 2007. [Google Scholar]
- Galvez-Llompart, M.; Zanni, R.; García-Domenech, R. Modeling natural anti-inflammatory compounds by molecular topology. Int. J. Mol. Sci. 2011, 12, 9481–9503. [Google Scholar] [CrossRef]
- Fylaktakidou, K.; Hadjipavlou-Litina, D.J.; Litinas, K.E.; Nicolaides, D.N. Natural and synthetic coumarin derivatives with anti-inflammatory/antioxidant activities. Curr. Pharm. Des. 2004, 10, 3813–3833. [Google Scholar] [CrossRef]
- Permender, R.; Hema, C.; Sushila, R.; Dharmender, R.; Vikash, K.; Kanchan, K. Mechanism of action of flavonoids as anti-inflammatory agents: A review. Inflamm. Allergy Drug Targets 2009, 8, 229–235. [Google Scholar]
- Bonechi, C.; Lamponi, S.; Donati, A.; Tamasi, G.; Consumi, M.; Leone, G.; Rossi, C.; Magnani, A. Effect of resveratrol on platelet aggregation by fibrinogen protection. Biophys. Chem. 2017, 222, 41–48. [Google Scholar] [CrossRef] [PubMed]
- Iwahara, J.; Schwieters, C.D.; Clore, G.M. Ensemble approach for NMR structure refinement against 1H paramagnetic relaxation enhancement data arising from a flexible paramagnetic group attached to a macromolecule. J. Am. Chem. Soc. 2004, 126, 5879–5896. [Google Scholar] [CrossRef]
- Kowalewski, J.; Nordenskiöld, L.; Benetis, N.; Westlund, P.-O. Theory of nuclear spin-relaxation in paramagnetic systems in solution. Prog. Nucl. Magn. Reson. Spectrosc. 1985, 17, 141–185. [Google Scholar] [CrossRef]
- Arnesano, F.; Banci, L.; Piccioli, M. NMR structures in paramagnetic metalloproteins. Quat. Rev. Biophys. 2005, 38, 167–219. [Google Scholar] [CrossRef] [PubMed]
- Gaggelli, E.; Rossi, C.; Tiezzi, E. Non covalent interactions and paramagnetic relaxation probe. Investigation of the correlation times. J. Phys. Chem. 1980, 84, 116–118. [Google Scholar] [CrossRef]
- Helm, L. Relaxivity in paramagnetic systems: Theory and mechanisms. Prog. Nucl. Magn. Reson. Spectrosc. 2006, 49, 45–64. [Google Scholar] [CrossRef]
- Niccolai, N.; Valensin, G.; Rossi, C.; Gibbons, W.A. The stereochemistry and dynamics of natural products and biopolymers from proton relaxation spectroscopy: Spin-label delineation of inner and outer protons of gramicidin S including hydrogen bonds. J. Am. Chem. Soc. 1982, 104, 1534–1537. [Google Scholar] [CrossRef]
- Bertini, I.; Pierattelli, R. Copper(II) proteins are amenable for NMR investigations. Pure Appl. Chem. 2004, 76, 321–333. [Google Scholar] [CrossRef]
- Clore, G.M.; Iwahara, J. Theory, practice and applications of paramagnetic relaxation enhancement for the characterization of transient low-population states of biological macromolecules and their complexes. Chem. Rev. 2009, 109, 4108–4139. [Google Scholar] [CrossRef]
- Sjodt, M.; Clubb, R.T. Nitroxide labeling of proteins and the determination of paramagnetic relaxation derived distance restraints for NMR studies. Bio-Protoc. 2017, 7, 1–17. [Google Scholar] [CrossRef]
- Clore, G.M. Practical aspects of paramagnetic relaxation enhancement in biological macromolecules. Method Enzymol. 2015, 564, 485–497. [Google Scholar]
- Otting, G. Protein NMR Using Paramagnetic Ions. Ann. Rev. Biophys. 2010, 39, 387–405. [Google Scholar] [CrossRef]
- D’Agostino, C.; Brauer, P.; Charoen-Rajapark, P.; Croucha, M.D.; Gladdena, L.F. Effect of paramagnetic species on T1, T2 and T1/T2 NMR relaxation times of liquids in porous CuSO4/Al2O3. RSC Adv. 2017, 7, 36163–36167. [Google Scholar] [CrossRef]
- Pilla, K.B.; Gaalswyk, K.; MacCallum, J.L. Molecular modelling of biomolecules by paramagnetic NMR and computational hybrid methods. Biochim. Biophys. Acta Proteins Proteom. 2017, 1865, 1654–1663. [Google Scholar] [CrossRef]
- Koehler, J.; Meiler, J. Expanding the utility of NMR restraints with paramagnetic compounds: Background and practical aspects. Prog. Nucl. Magn. Reson. Spectrosc. 2011, 59, 360–389. [Google Scholar] [CrossRef]
- Bertini, I.; Luchinat, C.; Parigi, G.; Pierattelli, R. Perspective in NMR metalloproteins. Dalton Trans. 2008, 3782–3790. [Google Scholar] [CrossRef]
- Huang, S.; Umemoto, R.; Tamura, Y.; Kofuku, Y.; Uyeda, T.Q.P.; Nishida, N.; Shimada, I. Utilization of paramagnetic relaxation enhancements for structural analysis of actin-binding proteins in complex with actin. Sci. Rep. 2016, 6, 33690. [Google Scholar] [CrossRef]
- Bonechi, C.; Martini, S.; Rossi, C. Interaction study of indigo carmine with albumin and dextran by NMR relaxation. J. Mater. Sci. 2011, 46, 2541–2547. [Google Scholar] [CrossRef]
- Goswami, S.; Sanyal, S.; Chakraborty, P.; Das, C.; Sarkar, M. Interaction of a common painkiller piroxicam and copper-piroxicam with chromatin causes structural alterations accompanied by modulation at the epigenomic/genomic level. Biochim. Biophys. Acta Gen. Subj. 2017, 1861, 2048–2059. [Google Scholar] [CrossRef] [PubMed]
- Cini, R.; Giorgi, G.; Cinquantini, A.; Rossi, C.; Sabat, M. Complexes of the antiinflammatory drug piroxicam. Inorg. Chem. 1990, 29, 5197–5200. [Google Scholar] [CrossRef]
- Tamasi, G.; Serinelli, F.; Consumi, M.; Magnani, A.; Casolaro, M.; Cini, R. Release studies from smart hydrogels as carriers for piroxicam and copper(II)-oxicam complexes as anti-inflammatory and anti-cancer drugs. X-ray structures of new copper(II)-piroxicam and -isoxicam complex molecules. J. Inorg. Biochem. 2008, 102, 1862–1873. [Google Scholar] [CrossRef]
- Hadadzadeh, H.; Salimi, M.; Weil, M.; Ahangari, R. The piroxicam complex of copper(II), trans-[Cu(Pir)2(THF)2], and its interaction with DNA. J. Mol. Struct. 2012, 1022, 172–180. [Google Scholar] [CrossRef]
- Ledesma-Olvera, L.G.; Agacino-Valdés, E.; Gómez-Balderas, R. Stability constants of Cu(II)-piroxicam complexes in solution: A DFT study. Theor. Chem. Acc. 2015, 135, 241–252. [Google Scholar] [CrossRef]
- Gehad, G.M.; El Gamel Nadia, E.A. Preparation and spectroscopic characterization of metal complexes of piroxicam. Vibr. Spectr. 2004, 36, 97–104. [Google Scholar] [CrossRef]
- Mildvan, A.S.; Cohn, M. Aspects of enzyme mechanisms studied by nuclear spin relaxation induced by paramagnetic probes. Adv. Enzymol. 1970, 33, 1–70. [Google Scholar] [PubMed]
- Solomon, I. Relaxation processes in a system of two spins. Phys. Rev. 1955, 99, 559–566. [Google Scholar] [CrossRef]
- Bloembergen, N. Proton relaxation times in paramagnetic solutions. Effects of electron spin relaxation. J. Chem. Phys. 1961, 34, 842–850. [Google Scholar] [CrossRef]
- Dwek, R.A. Nuclear Magnetic Resonance in Biochemistry; Clarendon Press: Oxford, UK, 1973. [Google Scholar]
- Esperson, W.G.; Hutton, W.C.; Chou, S.T.; Martin, R.B. Cautions concerning uses of Copper(II) selective broadening in nuclear magnetic resonance spectroscopy for determination of ligand binding sites. J. Am. Chem. Soc. 1974, 96, 8111–8112. [Google Scholar] [CrossRef]
- Fazakerley, G.V.; Jackson, G.E.; Phillips, M.A.; van Niekerk, L.C. A quantitative investigation of the interaction of Cu(II) and Mn(II) with some purine bases, nucleosides and nucleotides by NMR. Inorg. Chim. Acta 1979, 35, 151–160. [Google Scholar] [CrossRef]
- Banci, L.; Bertini, I.; Luchinat, C. Nuclear and Electron Relaxation: The Magnetic Nucleus-Unpaired Electron Coupling in Solution; VCH: New York, NY, USA, 1991. [Google Scholar]
Sample Availability: Samples of the compounds are not available from the authors. |
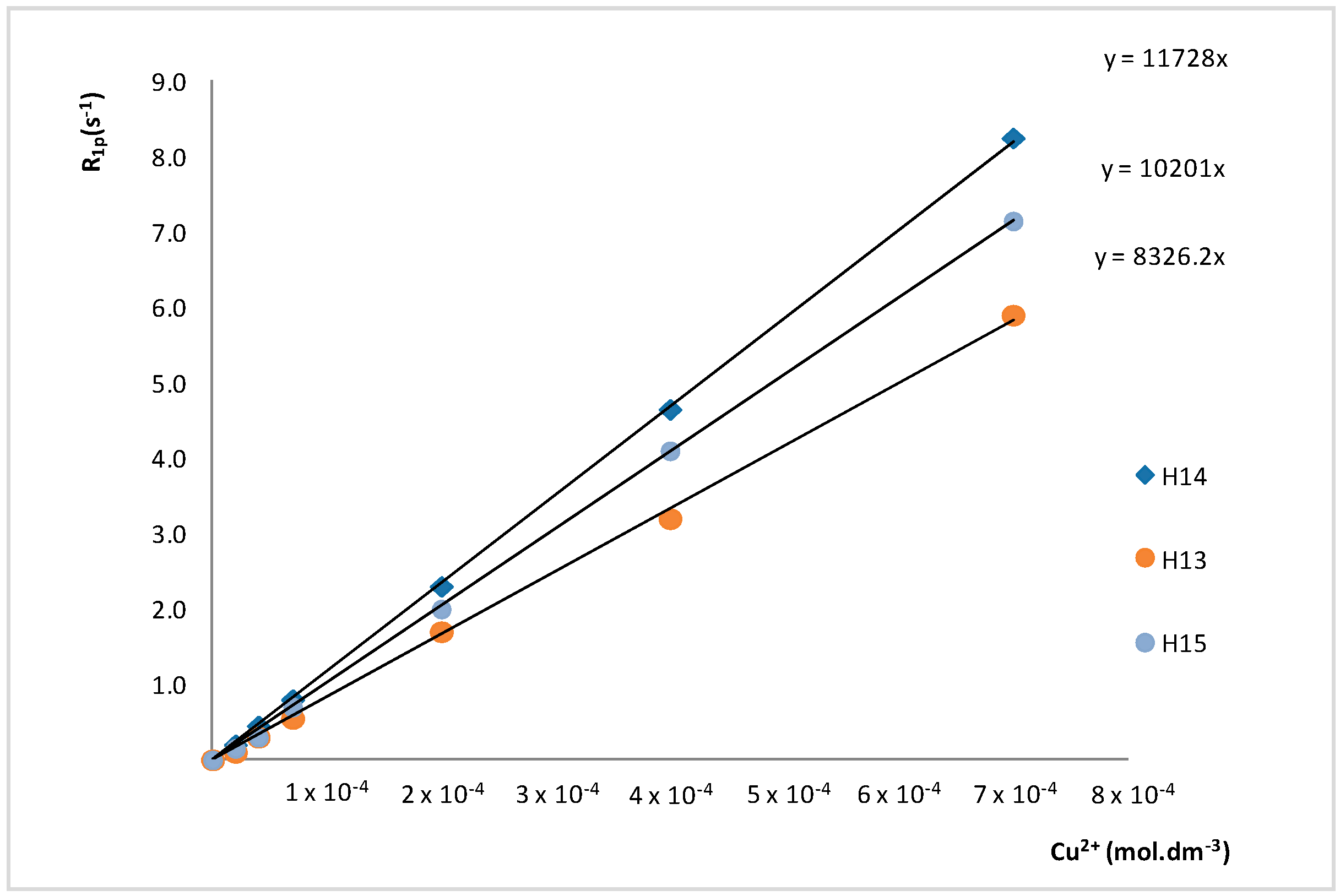
| Proton | δ (ppm) | [Cu] (mol L−1) | ||||||
|---|---|---|---|---|---|---|---|---|
| 0 | 2 × 10−5 | 4 × 10−5 | 7 × 10−5 | 2 × 10−4 | 4 × 10−4 | 7 × 10−4 | ||
| H14 | 8.42 | 0.40 | 0.60 | 0.85 | 1.20 | 2.70 | 5.05 | 8.65 |
| H2/H5 | 8.06 | 0.40 | 0.45 | 0.50 | 0.60 | 0.80 | 1.25 | 1.85 |
| H12 | 7.99 | 0.45 | 0.55 | 0.65 | 0.80 | 1.55 | 2.65 | 4.45 |
| H11/H3 | 7.89 | 0.50 | 0.55 | 0.56 | 0.70 | 0.95 | 1.40 | 2.27 |
| H4 | 7.86 | 0.55 | 0.60 | 0.65 | 0.85 | 0.95 | 1.25 | 1.85 |
| H13 | 7.28 | 0.45 | 0.55 | 0.75 | 1.00 | 2.15 | 3.65 | 6.35 |
| H15 | 2.87 | 1.10 | 1.25 | 1.40 | 1.80 | 3.10 | 5.20 | 8.25 |
| Proton | δ (ppm) | [Cu] (mol L−1) | |||||
|---|---|---|---|---|---|---|---|
| 2 × 10−5 | 4 × 10−5 | 7 × 10−5 | 2 × 10−4 | 4 × 10−4 | 7 × 10−4 | ||
| H14 | 8.42 | 0.20 | 0.45 | 0.80 | 2.30 | 4.65 | 8.25 |
| H2/H5 | 8.06 | 0.05 | 0.10 | 0.20 | 0.40 | 0.85 | 1.45 |
| H12 | 7.99 | 0.10 | 0.20 | 0.35 | 1.10 | 2.20 | 4.00 |
| H11/H3 | 7.89 | 0.05 | 0.10 | 0.20 | 0.45 | 0.90 | 1.60 |
| H4 | 7.86 | 0.05 | 0.10 | 0.30 | 0.40 | 0.70 | 1.30 |
| H13 | 7.28 | 0.10 | 0.30 | 0.55 | 1.70 | 3.20 | 5.90 |
| H15 | 2.87 | 0.15 | 0.30 | 0.70 | 2.00 | 4.10 | 7.15 |
| Proton | δ (ppm) | [Cu] (mol L−1) | ||||
|---|---|---|---|---|---|---|
| 0.25 | 0.20 | 0.15 | 0.10 | 5 × 10−5 | ||
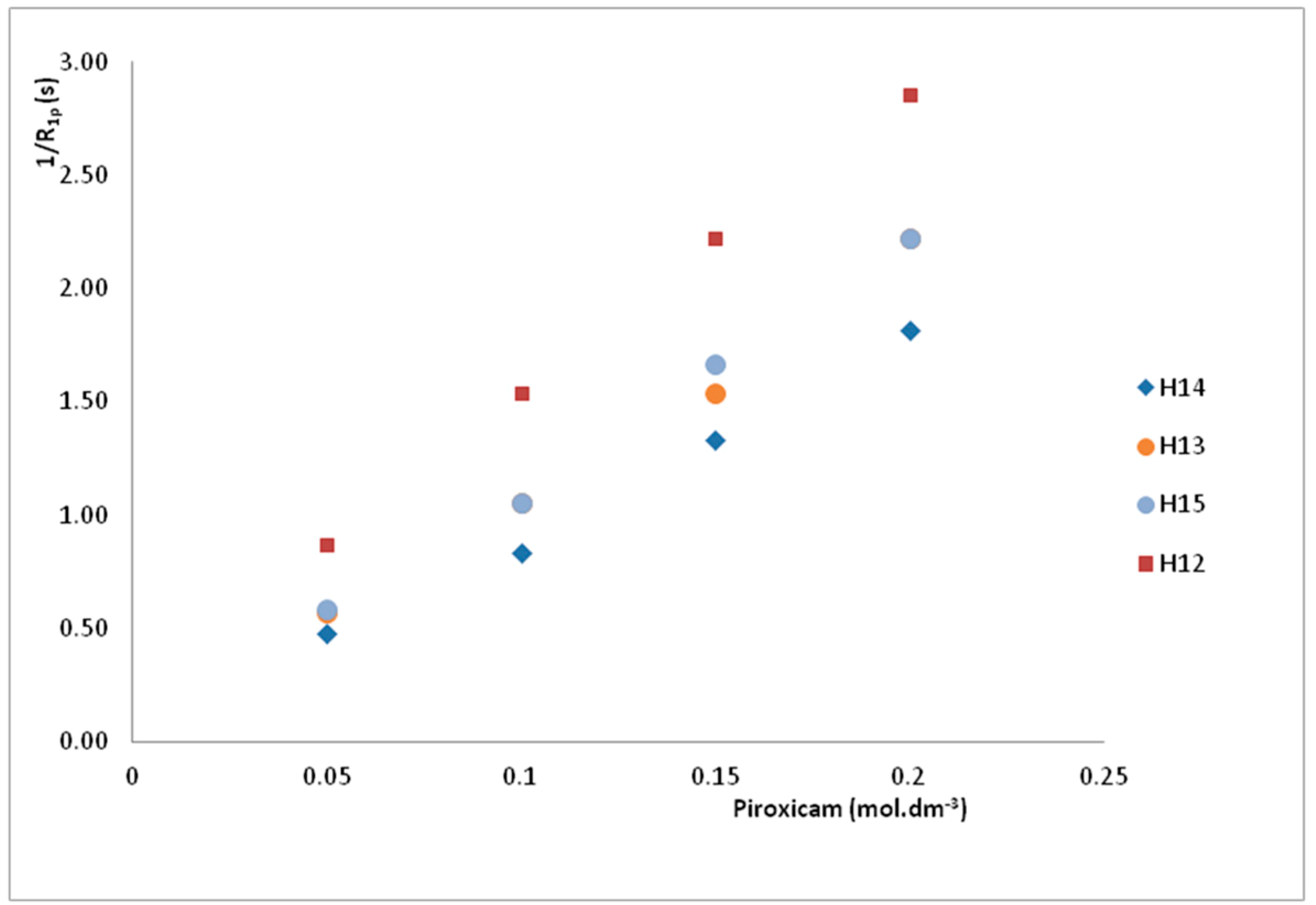
| H14 | 8.42 | 0.46 | 0.55 | 0.75 | 1.20 | 2.10 |
| H2/H5 | 8.06 | 0.12 | 0.15 | 0.20 | 0.35 | 0.45 |
| H12 | 7.99 | 0.30 | 0.35 | 0.45 | 0.65 | 1.15 |
| H11/H3 | 7.89 | 0.10 | 0.12 | 0.17 | 0.25 | 0.45 |
| H4 | 7.86 | 0.12 | 0.15 | 0.20 | 0.30 | 0.60 |
| H13 | 7.28 | 0.37 | 0.45 | 0.65 | 0.95 | 1.75 |
| H15 | 2.87 | 0.38 | 0.45 | 0.60 | 0.95 | 1.70 |
| Proton | δ (ppm) | R1m (s−1) |
|---|---|---|
| H14 | 8.42 | 1129 ± 55 |
| H12 | 7.99 | 730 ± 36 |
| H13 | 7.28 | 943 ± 46 |
| H15 | 2.87 | 917 ± 44 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bonechi, C.; Donati, A.; Tamasi, G.; Pardini, A.; Volpi, V.; Leone, G.; Consumi, M.; Magnani, A.; Rossi, C. Metal-Ligand Recognition Index Determination by NMR Proton Relaxation Study. Molecules 2019, 24, 1050. https://doi.org/10.3390/molecules24061050
Bonechi C, Donati A, Tamasi G, Pardini A, Volpi V, Leone G, Consumi M, Magnani A, Rossi C. Metal-Ligand Recognition Index Determination by NMR Proton Relaxation Study. Molecules. 2019; 24(6):1050. https://doi.org/10.3390/molecules24061050
Chicago/Turabian StyleBonechi, Claudia, Alessandro Donati, Gabriella Tamasi, Alessio Pardini, Vanessa Volpi, Gemma Leone, Marco Consumi, Agnese Magnani, and Claudio Rossi. 2019. "Metal-Ligand Recognition Index Determination by NMR Proton Relaxation Study" Molecules 24, no. 6: 1050. https://doi.org/10.3390/molecules24061050
APA StyleBonechi, C., Donati, A., Tamasi, G., Pardini, A., Volpi, V., Leone, G., Consumi, M., Magnani, A., & Rossi, C. (2019). Metal-Ligand Recognition Index Determination by NMR Proton Relaxation Study. Molecules, 24(6), 1050. https://doi.org/10.3390/molecules24061050









