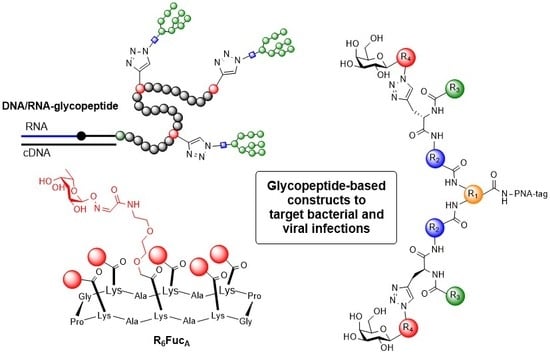
Glycopeptides and -Mimetics to Detect, Monitor and Inhibit Bacterial and Viral Infections: Recent Advances and Perspectives
Abstract
1. Introduction
2. Bacterial Infections
2.1. Escherichia coli (E. coli)
2.2. Pseudomonas aeruginosa
2.3. Burkholderia cepacia
2.4. Mycobacterium tuberculosis
3. Viral Infections
3.1. Human Immunodeficiency Virus Type-1 (HIV-1)
3.2. Epstein-Barr Virus (EBV)
3.3. Herpes Simplex Virus (HSV)
3.4. Influenza A Virus (IAV)
3.5. Norovirus (NoV)
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Bhatia, S.; Dimde, M.; Haag, R. Multivalent glycoconjugates as vaccines and potential drug candidates. Med. Chem. Commun. 2014, 5, 862–878. [Google Scholar] [CrossRef]
- Mammen, M.; Choi, S.K.; Whitesides, G.M. Polyvalent interactions in biological systems: Implications for design and use of multivalent ligands and inhibitors. Angew. Chem. Int. Ed. 1998, 37, 2754–2794. [Google Scholar] [CrossRef]
- Lundquist, J.J.; Toone, E.J. The cluster glycoside effect. Chem. Rev. 2002, 102, 555–578. [Google Scholar] [CrossRef]
- Dam, T.K.; Brewer, C.F. Effects of clustered epitopes in multivalent ligand–receptor interactions. Biochemistry 2008, 47, 8470–8476. [Google Scholar] [CrossRef] [PubMed]
- Fasting, C.; Schalley, C.A.; Weber, M.; Seitz, O.; Hecht, S.; Koksch, B.; Dernedde, J.; Graf, C.; Knapp, E.W.; Haag, R. Multivalency as a chemical organization and action principle. Angew. Chem. Int. Ed. 2012, 51, 10472–10498. [Google Scholar] [CrossRef] [PubMed]
- Sears, P.; Wong, C.H. Carbohydrate mimetics: A new strategy for tackling the problem of carbohydrate-mediated biological recognition. Angew. Chem. Int. Ed. 1999, 38, 2300–2324. [Google Scholar] [CrossRef]
- Ofek, I.; Hasty, D.L.; Sharon, N. Anti-adhesion therapy of bacterial diseases: Prospects and problems. FEMS Immunol. Med. Microbiol. 2003, 38, 181–191. [Google Scholar] [CrossRef]
- Germain, R.N. The biochemistry and cell biology of antigen presentation by MHC class I and class II molecules: Implications for development of combination vaccines. Ann. N. Y. Acad. Sci. 1995, 754, 114–125. [Google Scholar] [CrossRef] [PubMed]
- Avci, F.Y.; Li, X.; Tsuji, M.; Kasper, D.L. Carbohydrates and T cells: A sweet twosome. Semin. Immunol. 2013, 25, 146–151. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Middleton, D.R.; Wantuch, P.L.; Ozdilek, A.; Avci, F.Y. Carbohydrates as T-cell antigens with implications in health and disease. Glycobiology 2016, 26, 1029–1040. [Google Scholar] [CrossRef] [PubMed]
- Guttormsen, H.-K.; Sharpe, A.H.; Chandraker, A.K.; Brigtsen, A.K.; Sayegh, M.H.; Kasper, D.L. Cognate stimulatory B-cell–T-cell interactions are critical for T-cell help recruited by glycoconjugate vaccines. Infect. Immun. 1999, 67, 6375–6384. [Google Scholar] [PubMed]
- Avci, F.Y.; Kasper, D.L. How bacterial carbohydrates influence the adaptive immune system. Annu. Rev. Immunol. 2009, 28, 107–130. [Google Scholar] [CrossRef] [PubMed]
- WHO. Global Priority List of Antibiotic-Resistant Bacteria to Guide Research, Discovery, and Development of New Antibiotics. Available online: https://www.who.int/medicines/publications/WHO-PPL-Short_Summary_25Feb-ET_NM_WHO.pdf (accessed on 11 February 2019).
- Walsh, C.T.; Wencewicz, T.A. Prospects for new antibiotics: A molecule-centered perspective. J. Antibiot. 2014, 67, 7–22. [Google Scholar] [CrossRef]
- Luther, A.; Bisang, C.; Obrecht, D. Advances in macrocyclic peptide-based antibiotics. Bioorg. Med. Chem. 2018, 26, 2850–2858. [Google Scholar] [CrossRef] [PubMed]
- Blaskovich, M.A.; Hansford, K.A.; Butler, M.S.; Jia, Z.; Mark, A.E.; Cooper, M.A. Developments in glycopeptide antibiotics. ACS Infect. Dis. 2018, 4, 715–735. [Google Scholar] [CrossRef] [PubMed]
- Davis, B.G. Synthesis of Glycoproteins. Chem. Rev. 2002, 102, 579–602. [Google Scholar] [CrossRef] [PubMed]
- Payne, R.J.; Wong, C.-H. Advances in chemical ligation strategies for the synthesis of glycopeptides and glycoproteins. Chem. Commun. 2010, 46, 21–43. [Google Scholar] [CrossRef] [PubMed]
- Cecioni, S.; Oerthel, V.; Iehl, J.; Holler, M.; Goyard, D.; Praly, J.-P.; Imberty, A.; Nierengarten, J.-F.; Vidal, S. Synthesis of Dodecavalent Fullerene-Based Glycoclusters and Evaluation of Their Binding Properties towards a Bacterial Lectin. Chem. Eur. J. 2011, 17, 3252–3261. [Google Scholar] [CrossRef] [PubMed]
- Cecioni, S.; Lalor, R.; Blanchard, B.; Praly, J.-P.; Imberty, A.; Matthews, S.E.; Vidal, S. Achieving High Affinity towards a Bacterial Lectin through Multivalent Topological Isomers of Calix[4]arene Glycoconjugates. Chem. Eur. J. 2009, 15, 13232–13240. [Google Scholar] [CrossRef]
- Ramström, O.; Yan, M. Glyconanomaterials for Combating Bacterial Infections. Chem. Eur. J. 2015, 21, 16310–16317. [Google Scholar] [CrossRef]
- Rieger, J.; Stoffelbach, F.; Cui, D.; Imberty, A.; Lameignere, E.; Putaux, J.-L.; Jérôme, R.; Jérôme, C.; Auzély-Velty, R. Mannosylated Poly(ethylene oxide)-b-Poly(ε-caprolactone) Diblock Copolymers: Synthesis, Characterization, and Interaction with a Bacterial Lectin. Biomacromolecules 2007, 8, 2717–2725. [Google Scholar] [CrossRef] [PubMed]
- Ohlsen, K.; Oelschlaeger, T.; Hacker, J.; Khan, A.S. Carbohydrate receptors of bacterial adhesins: Implications and reflections. In Glycoscience and Microbial Adhesion; Lindhorst, T.K., Oscarson, S., Eds.; Springer: Berlin/Heidelberg, Germany, 2009; pp. 17–65. [Google Scholar]
- Hartmann, M.; Lindhorst, T.K. The bacterial lectin FimH, a target for drug discovery–carbohydrate inhibitors of type 1 fimbriae-mediated bacterial adhesion. Eur. J. Org. Chem. 2011, 2011, 3583–3609. [Google Scholar] [CrossRef]
- Ehrmann, S.; Chu, C.-W.; Kumari, S.; Silberreis, K.; Bӧttcher, C.; Dernedde, J.; Ravoo, B.J.; Haag, R. A Toolbox Approach for Multivalent Presentation of Ligand-Receptor Recognition on a Supramolecular Scaffold. J. Mater. Chem. B 2018, 6, 4216–4222. [Google Scholar] [CrossRef]
- Sperling, O.; Fuchs, A.; Lindhorst, T.K. Evaluation of the carbohydrate recognition domain of the bacterial adhesin FimH: Design, synthesis and binding properties of mannoside ligands. Org. Biomol. Chem. 2006, 4, 3913–3922. [Google Scholar] [CrossRef] [PubMed]
- Hung, C.S.; Bouckaert, J.; Hung, D.; Pinkner, J.; Widberg, C.; DeFusco, A.; Auguste, C.G.; Strouse, R.; Langermann, S.; Waksman, G.; et al. Structural basis of tropism of Escherichia coli to the bladder during urinary tract infection. Mol. Microbiol. 2002, 44, 903–915. [Google Scholar] [CrossRef] [PubMed]
- Bouckaert, J.; Berglund, J.; Schembri, M.; De Genst, E.; Cools, L.; Wuhrer, M.; Hung, C.S.; Pinkner, J.; Slättegård, R.; Zavialov, A.; et al. Receptor binding studies disclose a novel class of high-affinity inhibitors of the Escherichia coli FimH adhesin. Mol. Microbiol. 2005, 55, 441–455. [Google Scholar] [CrossRef] [PubMed]
- Wellens, A.; Garofalo, C.; Nguyen, H.; Van Gerven, N.; Slättegård, R.; Hernalsteens, J.-P.; Wyns, L.; Oscarson, S.; De Greve, H.; Hultgren, S.; et al. Intervening with urinary tract infections using anti-adhesives based on the crystal structure of the FimH–oligomannose-3 complex. PLoS ONE 2008, 3, e2040. [Google Scholar] [CrossRef]
- Knight, S.D.; Bouckaert, J. Structure, function, and assembly of type 1 fimbriae. In Glycoscience and Microbial Adhesion; Lindhorst, T.K., Oscarson, S., Eds.; Springer: Berlin/Heidelberg, Germany, 2009; pp. 67–107. [Google Scholar]
- Han, Z.; Pinkner, J.S.; Ford, B.; Obermann, R.; Nolan, W.; Wildman, S.A.; Hobbs, D.; Ellenberger, T.; Cusumano, C.K.; Hultgren, S.J.; et al. Structure-based drug design and optimization of mannoside bacterial FimH antagonists. J. Med. Chem. 2010, 53, 4779–4792. [Google Scholar] [CrossRef]
- Lahmann, M. Architectures of multivalent glycomimetics for probing carbohydrate–lectin interactions. In Glycoscience and Microbial Adhesion; Lindhorst, T.K., Oscarson, S., Eds.; Springer: Berlin/Heidelberg, Germay, 2009; pp. 165–183. [Google Scholar]
- Lindhorst, T.K.; Bruegge, K.; Fuchs, A.; Sperling, O. A bivalent glycopeptide to target two putative carbohydrate binding sites on FimH. Beilstein J. Org. Chem. 2010, 6, 801–809. [Google Scholar] [CrossRef]
- Schierholt, A.; Hartmann, M.; Schwekendiek, K.; Lindhorst, T.K. Cysteine-Based Mannoside Glycoclusters: Synthetic Routes and Antiadhesive Properties. Eur. J. Org. Chem. 2010, 2010, 3120–3128. [Google Scholar] [CrossRef]
- Wehner, J.W.; Hartmann, M.; Lindhorst, T.K. Are multivalent cluster glycosides a means of controlling ligand density of glycoarrays? Carbohydr. Res. 2013, 371, 22–31. [Google Scholar] [CrossRef] [PubMed]
- Schierholt, A.; Hartmann, M.; Lindhorst, T.K. Bi-and trivalent glycopeptide mannopyranosides as inhibitors of type 1 fimbriae-mediated bacterial adhesion: Variation of valency, aglycon and scaffolding. Carbohydr. Res. 2011, 346, 1519–1526. [Google Scholar] [CrossRef] [PubMed]
- Igde, S.; Röblitz, S.; Müller, A.; Kolbe, K.; Boden, S.; Fessele, C.; Lindhorst, T.K.; Weber, M.; Hartmann, L. Linear Precision Glycomacromolecules with Varying Interligand Spacing and Linker Functionalities Binding to Concanavalin A and the Bacterial Lectin FimH. Macromol. Biosci. 2017, 17, 1700198. [Google Scholar] [CrossRef]
- Gibson, R.L.; Burns, J.L.; Ramsey, B.W. Pathophysiology and management of pulmonary infections in cystic fibrosis. Am. J. Respir. Crit. Care Med. 2003, 168, 918–951. [Google Scholar] [CrossRef] [PubMed]
- Gellatly, S.L.; Hancock, R.E. Pseudomonas aeruginosa: New insights into pathogenesis and host defenses. Pathog. Dis. 2013, 67, 159–173. [Google Scholar] [CrossRef] [PubMed]
- Wagner, V.E.; Iglewski, B.H.P. aeruginosa biofilms in CF infection. Clin. Rev. Allergy Immunol. 2008, 35, 124–134. [Google Scholar] [CrossRef] [PubMed]
- Garber, N.; Guempel, U.; Belz, A.; Gilboa-Garber, N.; Doyle, R.J. On the specificity of the d-galactose-binding lectin (PA-I) of Pseudomonas aeruginosa and its strong binding to hydrophobic derivatives of d-galactose and thiogalactose. Biochim. Biophys. Acta 1992, 1116, 331–333. [Google Scholar] [CrossRef]
- Imberty, A.; Wimmerová, M.; Mitchell, E.P.; Gilboa-Garber, N. Structures of the lectins from Pseudomonas aeruginosa: Insights into the molecular basis for host glycan recognition. Microbes Infect. 2004, 6, 221–228. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.-P.; Song, S.-C.; Gilboa-Garber, N.; Chang, K.S.; Wu, A.M. Studies on the binding site of the galactose-specific agglutinin PA-IL from Pseudomonas aeruginosa. Glycobiology 1998, 8, 7–16. [Google Scholar] [CrossRef]
- Mitchell, E.; Houles, C.; Sudakevitz, D.; Wimmerova, M.; Gautier, C.; Pérez, S.; Wu, A.M.; Gilboa-Garber, N.; Imberty, A. Structural basis for oligosaccharide-mediated adhesion of Pseudomonas aeruginosa in the lungs of cystic fibrosis patients. Nat. Struct. Biol. 2002, 9, 918–921. [Google Scholar] [CrossRef]
- Wittmann, V.; Pieters, R.J. Bridging lectin binding sites by multivalent carbohydrates. Chem. Soc. Rev. 2013, 42, 4492–4503. [Google Scholar] [CrossRef]
- Huang, S.F.; Lin, C.H.; Lai, Y.T.; Tsai, C.L.; Cheng, T.J.R.; Wang, S.K. Development of Pseudomonas aeruginosa Lectin LecA Inhibitor by using Bivalent Galactosides Supported on Polyproline Peptide Scaffolds. Chem. Asian J. 2018, 13, 686–700. [Google Scholar] [CrossRef]
- Kadam, R.U.; Garg, D.; Schwartz, J.; Visini, R.; Sattler, M.; Stocker, A.; Darbre, T.; Reymond, J.-L. CH–π “T-Shape” Interaction with histidine explains binding of aromatic galactosides to Pseudomonas aeruginosa lectin LecA. ACS Chem. Biol. 2013, 8, 1925–1930. [Google Scholar] [CrossRef]
- Kadam, R.U.; Bergmann, M.; Hurley, M.; Garg, D.; Cacciarini, M.; Swiderska, M.A.; Nativi, C.; Sattler, M.; Smyth, A.R.; Williams, P.; et al. A glycopeptide dendrimer inhibitor of the galactose-specific lectin LecA and of Pseudomonas aeruginosa biofilms. Angew. Chem. Int. Ed. 2011, 50, 10631–10635. [Google Scholar] [CrossRef]
- Novoa, A.; Eierhoff, T.; Topin, J.; Varrot, A.; Barluenga, S.; Imberty, A.; Römer, W.; Winssinger, N. A LecA Ligand Identified from a Galactoside-Conjugate Array Inhibits Host Cell Invasion by Pseudomonas aeruginosa. Angew. Chem. Int. Ed. 2014, 126, 9031–9035. [Google Scholar] [CrossRef]
- Zambaldo, C.; Barluenga, S.; Winssinger, N. PNA-encoded chemical libraries. Curr. Opin. Chem. Biol. 2015, 26, 8–15. [Google Scholar] [CrossRef]
- Johansson, E.M.; Crusz, S.A.; Kolomiets, E.; Buts, L.; Kadam, R.U.; Cacciarini, M.; Bartels, K.-M.; Diggle, S.P.; Cámara, M.; Williams, P.; et al. Inhibition and dispersion of Pseudomonas aeruginosa biofilms by glycopeptide dendrimers targeting the fucose-specific lectin LecB. Chem. Biol. 2008, 15, 1249–1257. [Google Scholar] [CrossRef]
- Emma, M. Inhibition of Pseudomonas aeruginosa biofilms with a glycopeptide dendrimer containing d-amino acids. Med. Chem. Commun. 2011, 2, 418–420. [Google Scholar]
- Michaud, G.; Visini, R.; Bergmann, M.; Salerno, G.; Bosco, R.; Gillon, E.; Richichi, B.; Nativi, C.; Imberty, A.; Stocker, A.; et al. Overcoming antibiotic resistance in Pseudomonas aeruginosa biofilms using glycopeptide dendrimers. Chem. Sci. 2016, 7, 166–182. [Google Scholar] [CrossRef]
- Bergmann, M.; Michaud, G.; Visini, R.; Jin, X.; Gillon, E.; Stocker, A.; Imberty, A.; Darbre, T.; Reymond, J.-L. Multivalency effects on Pseudomonas aeruginosa biofilm inhibition and dispersal by glycopeptide dendrimers targeting lectin LecA. Org. Biomol. Chem. 2016, 14, 138–148. [Google Scholar] [CrossRef]
- Bücher, K.S.; Babic, N.; Freichel, T.; Kovacic, F.; Hartmann, L. Monodisperse Sequence-Controlled α-l-Fucosylated Glycooligomers and Their Multivalent Inhibitory Effects on LecB. Macromol. Biosci. 2018, 18, 1800337. [Google Scholar] [CrossRef] [PubMed]
- Bücher, K.S.; Konietzny, P.B.; Snyder, N.L.; Hartmann, L. Heteromultivalent glycooligomers as mimetics of blood group antigens. Chem. Eur. J. 2019, 25, 3301–3309. [Google Scholar] [CrossRef] [PubMed]
- Dumy, P.; Eggleston, I.; Cervigni, S.; Sila, U.; Sun, X.; Mutter, M. A convenient synthesis of cyclic peptides as regioselectively addressable functionalized templates (RAFT). Tetrahedron Lett. 1995, 36, 1255–1258. [Google Scholar] [CrossRef]
- Dumy, P.; Eggleston, I.M.; Esposito, G.; Nicula, S.; Mutter, M. Solution structure of regioselectively addressable functionalized templates: An NMR and restrained molecular dynamics investigation. Biopolymers 1996, 39, 297–308. [Google Scholar] [CrossRef]
- Grigalevicius, S.; Chierici, S.; Renaudet, O.; Lo-Man, R.; Dériaud, E.; Leclerc, C.; Dumy, P. Chemoselective assembly and immunological evaluation of multiepitopic glycoconjugates bearing clustered Tn antigen as synthetic anticancer vaccines. Bioconj. Chem. 2005, 16, 1149–1159. [Google Scholar] [CrossRef] [PubMed]
- Pifferi, C.; Berthet, N.; Renaudet, O. Cyclopeptide scaffolds in carbohydrate-based synthetic vaccines. Biomater. Sci. 2017, 5, 953–965. [Google Scholar] [CrossRef] [PubMed]
- Berthet, N.; Thomas, B.; Bossu, I.; Dufour, E.; Gillon, E.; Garcia, J.; Spinelli, N.; Imberty, A.; Dumy, P.; Renaudet, O. High affinity glycodendrimers for the lectin LecB from Pseudomonas aeruginosa. Bioconj. Chem. 2013, 24, 1598–1611. [Google Scholar] [CrossRef]
- Scoffone, V.C.; Chiarelli, L.R.; Trespidi, G.; Mentasti, M.; Riccardi, G.; Buroni, S. Burkholderia cenocepacia infections in cystic fibrosis patients: Drug resistance and therapeutic approaches. Front. Microbiol. 2017, 8, 1592. [Google Scholar] [CrossRef]
- Govan, J.R.; Deretic, V. Microbial pathogenesis in cystic fibrosis: Mucoid Pseudomonas aeruginosa and Burkholderia cepacia. Microbiol. Rev. 1996, 60, 539–574. [Google Scholar]
- Aaron, S.D.; Ferris, W.; Henry, D.A.; Speert, D.P.; MacDonald, N.E. Multiple combination bactericidal antibiotic testing for patients with cystic fibrosis infected with Burkholderia cepacia. Am. J. Respir. Crit. Care Med. 2000, 161, 1206–1212. [Google Scholar] [CrossRef]
- Lameignere, E.; Malinovská, L.; Sláviková, M.; Duchaud, E.; Mitchell, E.P.; Varrot, A.; Šedo, O.; Imberty, A.; Wimmerová, M. Structural basis for mannose recognition by a lectin from opportunistic bacteria Burkholderia cenocepacia. Biochem. J. 2008, 411, 307–318. [Google Scholar] [CrossRef]
- Lameignere, E.; Shiao, T.C.; Roy, R.; Wimmerova, M.; Dubreuil, F.; Varrot, A.; Imberty, A. Structural basis of the affinity for oligomannosides and analogs displayed by BC2L-A, a Burkholderia cenocepacia soluble lectin. Glycobiology 2009, 20, 87–98. [Google Scholar] [CrossRef]
- Marchetti, R.; Malinovska, L.; Lameignère, E.; Adamova, L.; de Castro, C.; Cioci, G.; Stanetty, C.; Kosma, P.; Molinaro, A.; Wimmerova, M. Burkholderia cenocepacia lectin A binding to heptoses from the bacterial lipopolysaccharide. Glycobiology 2012, 22, 1387–1398. [Google Scholar] [CrossRef]
- Pifferi, C.; Goyard, D.; Gillon, E.; Imberty, A.; Renaudet, O. Synthesis of Mannosylated Glycodendrimers and Evaluation against BC2L-A Lectin from Burkholderia Cenocepacia. ChemPlusChem 2017, 82, 390–398. [Google Scholar] [CrossRef]
- Goyard, D.; Baldoneschi, V.; Varrot, A.; Fiore, M.; Imberty, A.; Richichi, B.; Renaudet, O.; Nativi, C. Multivalent Glycomimetics with Affinity and Selectivity toward Fucose-Binding Receptors from Emerging Pathogens. Bioconj. Chem. 2017, 29, 83–88. [Google Scholar] [CrossRef]
- Vial, L.; Groleau, M.-C.; Lamarche, M.G.; Filion, G.; Castonguay-Vanier, J.; Dekimpe, V.; Daigle, F.; Charette, S.J.; Déziel, E. Phase variation has a role in Burkholderia ambifaria niche adaptation. ISME J. 2010, 4, 49–60. [Google Scholar] [CrossRef]
- WHO. Global Tuberculosis Report 2018. Available online: https://apps.who.int/iris/bitstream/handle/10665/274453/9789241565646-eng.pdf?ua=1 (accessed on 8 February 2019).
- Brennan, P.J. Structure, function, and biogenesis of the cell wall of Mycobacterium tuberculosis. Tuberculosis 2003, 83, 91–97. [Google Scholar] [CrossRef]
- Chan, J.; Fan, X.; Hunter, S.; Brennan, P.; Bloom, B. Lipoarabinomannan, a possible virulence factor involved in persistence of Mycobacterium tuberculosis within macrophages. Infect. Immun. 1991, 59, 1755–1761. [Google Scholar]
- Spencer, J.S.; Kim, H.J.; Wheat, W.H.; Chatterjee, D.; Balagon, M.V.; Cellona, R.V.; Tan, E.V.; Gelber, R.; Saunderson, P.; Duthie, M.S.J.C.; et al. Analysis of antibody responses to Mycobacterium leprae phenolic glycolipid I, lipoarabinomannan, and recombinant proteins to define disease subtype-specific antigenic profiles in leprosy. Clin. Vaccine Immunol. 2011, 18, 260–267. [Google Scholar] [CrossRef]
- Chou, Y.; Kitova, E.N.; Joe, M.; Brunton, R.; Lowary, T.L.; Klassen, J.S.; Derda, R.J.O. Genetically-encoded fragment-based discovery (GE-FBD) of glycopeptide ligands with differential selectivity for antibodies related to mycobacterial infections. Org. Biomol. Chem. 2018, 16, 223–227. [Google Scholar] [CrossRef]
- Rademacher, C.; Shoemaker, G.K.; Kim, H.-S.; Zheng, R.B.; Taha, H.; Liu, C.; Nacario, R.C.; Schriemer, D.C.; Klassen, J.S.; Peters, T. Ligand specificity of CS-35, a monoclonal antibody that recognizes mycobacterial lipoarabinomannan: A model system for oligofuranoside–protein recognition. J. Am. Chem. Soc. 2007, 129, 10489–10502. [Google Scholar] [CrossRef] [PubMed]
- Deiss, F.; Matochko, W.L.; Govindasamy, N.; Lin, E.Y.; Derda, R. Flow-through synthesis on teflon-patterned paper to produce peptide arrays for cell-based assays. Angew. Chem. Int. Ed. 2014, 53, 6374–6377. [Google Scholar] [CrossRef] [PubMed]
- Olofsson, S.; Blixt, O.; Bergström, T.; Frank, M.; Wandall, H.H. Viral O-GalNAc peptide epitopes: A novel potential target in viral envelope glycoproteins. Rev. Med. Virol. 2016, 26, 34–48. [Google Scholar] [CrossRef] [PubMed]
- Datema, R.; Olofsson, S.; Romero, P.A. Inhibitors of protein glycosylation and glycoprotein processing in viral systems. Pharmacol. Ther. 1987, 33, 221–286. [Google Scholar] [CrossRef]
- Olofsson, S.; Hansen, J.-E.S. Host cell glycosylation of viral glycoproteins-a battlefield for host defence and viral resistance. Scand. J. Infect. Dis. 1998, 30, 435–440. [Google Scholar] [PubMed]
- UNAIDS. Global HIV & AIDS Statistics—2018 Fact Sheet. Available online: http://www.unaids.org/sites/default/files/media_asset/UNAIDS_FactSheet_en.pdf (accessed on 11 February 2019).
- Burton, D.R.; Ahmed, R.; Barouch, D.H.; Butera, S.T.; Crotty, S.; Godzik, A.; Kaufmann, D.E.; McElrath, M.J.; Nussenzweig, M.C.; Pulendran, B. A blueprint for HIV vaccine discovery. Cell Host Microbe 2012, 12, 396–407. [Google Scholar] [CrossRef] [PubMed]
- Ward, A.B.; Wilson, I.A. Insights into the trimeric HIV-1 envelope glycoprotein structure. Trends Biochem. Sci. 2015, 40, 101–107. [Google Scholar] [CrossRef] [PubMed]
- Moore, J.P.; Sweet, R.W. The HIV gp120-CD4 interaction: A target for pharmacological or immunological intervention? Perspect. Drug Discov. Des. 1993, 1, 235–250. [Google Scholar] [CrossRef]
- Korber, B.; Gaschen, B.; Yusim, K.; Thakallapally, R.; Kesmir, C.; Detours, V. Evolutionary and immunological implications of contemporary HIV-1 variation. Br. Med. Bull. 2001, 58, 19–42. [Google Scholar] [CrossRef] [PubMed]
- Mizuochi, T.; Matthews, T.; Kato, M.; Hamako, J.; Titani, K.; Solomon, J.; Feizi, T. Diversity of oligosaccharide structures on the envelope glycoprotein gp 120 of human immunodeficiency virus 1 from the lymphoblastoid cell line H9. Presence of complex-type oligosaccharides with bisecting N-acetylglucosamine residues. J. Biol. Chem. 1990, 265, 8519–8524. [Google Scholar] [PubMed]
- Leonard, C.K.; Spellman, M.W.; Riddle, L.; Harris, R.J.; Thomas, J.N.; Gregory, T. Assignment of intrachain disulfide bonds and characterization of potential glycosylation sites of the type 1 recombinant human immunodeficiency virus envelope glycoprotein (gp120) expressed in Chinese hamster ovary cells. J. Biol. Chem. 1990, 265, 10373–10382. [Google Scholar] [PubMed]
- Reitter, J.N.; Means, R.E.; Desrosiers, R.C. A role for carbohydrates in immune evasion in AIDS. Nat. Med. 1998, 4, 679–684. [Google Scholar] [CrossRef] [PubMed]
- Earl, P.L.; Doms, R.W.; Moss, B. Oligomeric structure of the human immunodeficiency virus type 1 envelope glycoprotein. Proc. Natl. Acad. Sci. USA 1990, 87, 648–652. [Google Scholar] [CrossRef]
- Wilhelm, D.; Behnken, H.N.; Meyer, B. Glycosylation assists binding of HIV protein gp120 to human CD4 receptor. ChemBioChem 2012, 13, 524–527. [Google Scholar] [CrossRef]
- Walker, L.M.; Simek, M.D.; Priddy, F.; Gach, J.S.; Wagner, D.; Zwick, M.B.; Phogat, S.K.; Poignard, P.; Burton, D.R. A limited number of antibody specificities mediate broad and potent serum neutralization in selected HIV-1 infected individuals. PLoS Pathog. 2010, 6, e1001028. [Google Scholar] [CrossRef] [PubMed]
- Walker, L.M.; Huber, M.; Doores, K.J.; Falkowska, E.; Pejchal, R.; Julien, J.-P.; Wang, S.-K.; Ramos, A.; Chan-Hui, P.-Y.; Moyle, M.; et al. Broad neutralization coverage of HIV by multiple highly potent antibodies. Nature 2011, 477, 466–470. [Google Scholar] [CrossRef] [PubMed]
- Scanlan, C.N.; Pantophlet, R.; Wormald, M.R.; Saphire, E.O.; Stanfield, R.; Wilson, I.A.; Katinger, H.; Dwek, R.A.; Rudd, P.M.; Burton, D.R. The broadly neutralizing anti-human immunodeficiency virus type 1 antibody 2G12 recognizes a cluster of α1→ 2 mannose residues on the outer face of gp120. J. Virol. 2002, 76, 7306–7321. [Google Scholar] [CrossRef] [PubMed]
- Horiya, S.; Bailey, J.K.; Temme, J.S.; Guillen Schlippe, Y.V.; Krauss, I.J. Directed evolution of multivalent glycopeptides tightly recognized by HIV antibody 2G12. J. Am. Chem. Soc. 2014, 136, 5407–5415. [Google Scholar] [CrossRef] [PubMed]
- Bailey, J.K.; Nguyen, D.N.; Horiya, S.; Krauss, I.J. Synthesis of multivalent glycopeptide conjugates that mimic an HIV epitope. Tetrahedron 2016, 72, 6091–6098. [Google Scholar] [CrossRef] [PubMed]
- Walker, L.M.; Phogat, S.K.; Chan-Hui, P.-Y.; Wagner, D.; Phung, P.; Goss, J.L.; Wrin, T.; Simek, M.D.; Fling, S.; Mitcham, J.L.; et al. Broad and potent neutralizing antibodies from an African donor reveal a new HIV-1 vaccine target. Science 2009, 326, 285–289. [Google Scholar] [CrossRef] [PubMed]
- McLellan, J.S.; Pancera, M.; Carrico, C.; Gorman, J.; Julien, J.-P.; Khayat, R.; Louder, R.; Pejchal, R.; Sastry, M.; Dai, K.; et al. Structure of HIV-1 gp120 V1/V2 domain with broadly neutralizing antibody PG9. Nat. Biotechnol. 2011, 480, 336–343. [Google Scholar] [CrossRef] [PubMed]
- Pancera, M.; Shahzad-ul-Hussan, S.; Doria-Rose, N.A.; McLellan, J.S.; Bailer, R.T.; Dai, K.; Loesgen, S.; Louder, M.K.; Staupe, R.P.; Yang, Y.; et al. Structural basis for diverse N-glycan recognition by HIV-1–neutralizing V1–V2–directed antibody PG16. Nat. Struct. Mol. Biol. 2013, 20, 804–813. [Google Scholar] [CrossRef] [PubMed]
- Aussedat, B.; Vohra, Y.; Park, P.K.; Fernández-Tejada, A.; Alam, S.M.; Dennison, S.M.; Jaeger, F.H.; Anasti, K.; Stewart, S.; Blinn, J.H.; et al. Chemical Synthesis of Highly Congested gp120 V1V2 N-Glycopeptide Antigens for Potential HIV-1-Directed Vaccines. J. Am. Chem. Soc. 2013, 135, 13113–13120. [Google Scholar] [CrossRef]
- Alam, S.M.; Dennison, S.M.; Aussedat, B.; Vohra, Y.; Park, P.K.; Fernández-Tejada, A.; Stewart, S.; Jaeger, F.H.; Anasti, K.; Blinn, J.H.; et al. Recognition of synthetic glycopeptides by HIV-1 broadly neutralizing antibodies and their unmutated ancestors. Proc. Natl. Acad. Sci. USA 2013, 110, 18214–18219. [Google Scholar] [CrossRef] [PubMed]
- Amin, M.N.; McLellan, J.S.; Huang, W.; Orwenyo, J.; Burton, D.R.; Koff, W.C.; Kwong, P.D.; Wang, L.-X. Synthetic glycopeptides reveal the glycan specificity of HIV-neutralizing antibodies. Nat. Chem. Biol. 2013, 9, 521–526. [Google Scholar] [CrossRef]
- Toonstra, C.; Amin, M.N.; Wang, L.-X. Site-selective chemoenzymatic glycosylation of an HIV-1 polypeptide antigen with two distinct N-glycans via an orthogonal protecting group strategy. J. Org. Chem. 2016, 81, 6176–6185. [Google Scholar] [CrossRef] [PubMed]
- Longo, N.S.; Sutton, M.S.; Shiakolas, A.R.; Guenaga, J.; Jarosinski, M.C.; Georgiev, I.S.; McKee, K.; Bailer, R.T.; Louder, M.K.; O’Dell, S.; et al. Multiple antibody lineages in one donor target the glycan-V3 supersite of the HIV-1 envelope glycoprotein and display a preference for quaternary binding. J. Virol. 2016, 90, 10574–10586. [Google Scholar] [CrossRef]
- Orwenyo, J.; Cai, H.; Giddens, J.; Amin, M.N.; Toonstra, C.; Wang, L.-X. Systematic synthesis and binding study of HIV V3 glycopeptides reveal the fine epitopes of several broadly neutralizing antibodies. ACS Chem. Biol. 2017, 12, 1566–1575. [Google Scholar] [CrossRef]
- Kong, L.; Lee, J.H.; Doores, K.J.; Murin, C.D.; Julien, J.-P.; McBride, R.; Liu, Y.; Marozsan, A.; Cupo, A.; Klasse, P.-J.; et al. Supersite of immune vulnerability on the glycosylated face of HIV-1 envelope glycoprotein gp120. Nat. Struct. Mol. Biol. 2013, 20, 796–803. [Google Scholar] [CrossRef]
- Gristick, H.B.; von Boehmer, L.; West, A.P., Jr.; Schamber, M.; Gazumyan, A.; Golijanin, J.; Seaman, M.S.; Fätkenheuer, G.; Klein, F.; Nussenzweig, M.C.; et al. Natively glycosylated HIV-1 Env structure reveals new mode for antibody recognition of the CD4-binding site. Nat. Struct. Mol. Biol. 2016, 23, 906–915. [Google Scholar] [CrossRef]
- Cai, H.; Orwenyo, J.; Guenaga, J.; Giddens, J.; Toonstra, C.; Wyatt, R.T.; Wang, L.-X. Synthetic multivalent V3 glycopeptides display enhanced recognition by glycan-dependent HIV-1 broadly neutralizing antibodies. Chem. Commun. 2017, 53, 5453–5456. [Google Scholar] [CrossRef] [PubMed]
- Cai, H.; Zhang, R.; Orwenyo, J.; Giddens, J.; Yang, Q.; LaBranche, C.C.; Montefiori, D.C.; Wang, L.-X. Multivalent Antigen Presentation Enhances the Immunogenicity of a Synthetic Three-Component HIV-1 V3 Glycopeptide Vaccine. ACS Cent. Sci. 2018, 4, 582–589. [Google Scholar] [CrossRef] [PubMed]
- Cai, H.; Orwenyo, J.; Giddens, J.P.; Yang, Q.; Zhang, R.; LaBranche, C.C.; Montefiori, D.C.; Wang, L.-X. Synthetic Three-Component HIV-1 V3 Glycopeptide Immunogens Induce Glycan-Dependent Antibody Responses. Cell Chem. Biol. 2017, 24, 1513–1522. [Google Scholar] [CrossRef] [PubMed]
- Alam, S.M.; Aussedat, B.; Vohra, Y.; Meyerhoff, R.R.; Cale, E.M.; Walkowicz, W.E.; Radakovich, N.A.; Anasti, K.; Armand, L.; Parks, R.; et al. Mimicry of an HIV broadly neutralizing antibody epitope with a synthetic glycopeptide. Sci. Transl. Med. 2017, 9, eaai7521. [Google Scholar] [CrossRef] [PubMed]
- Cai, H.; Zhang, R.-S.; Orwenyo, J.; Giddens, J.; Yang, Q.; LaBranche, C.C.; Montefiori, D.C.; Wang, L.-X. Synthetic HIV V3 Glycopeptide Immunogen Carrying a N334 N-Glycan Induces Glycan-Dependent Antibodies with Promiscuous Site Recognition. J. Med. Chem. 2018, 61, 10116–10125. [Google Scholar] [CrossRef] [PubMed]
- Fera, D.; Lee, M.S.; Wiehe, K.; Meyerhoff, R.R.; Piai, A.; Bonsignori, M.; Aussedat, B.; Walkowicz, W.E.; Ton, T.; Zhou, J.O.; et al. HIV envelope V3 region mimic embodies key features of a broadly neutralizing antibody lineage epitope. Nat. Commun. 2018, 9, 1111. [Google Scholar] [CrossRef]
- Cohen, J.I. Epstein–Barr virus infection. N. Engl. J. Med. 2000, 343, 481–492. [Google Scholar] [CrossRef]
- Munch, M.; Riisom, K.; Christensen, T.; Møller-Larsen, A.; Haahr, S. The significance of Epstein—Barr virus seropositivity in multiple sclerosis patients? Acta Neurol. Scand. 1998, 97, 171–174. [Google Scholar] [CrossRef]
- Henle, G.; Henle, W.; Diehl, V. Relation of Burkitt’s tumor-associated herpes-ytpe virus to infectious mononucleosis. Proc. Natl. Acad. Sci. USA 1968, 59, 94–101. [Google Scholar] [CrossRef]
- Nemerow, G.; Mold, C.; Schwend, V.K.; Tollefson, V.; Cooper, N. Identification of gp350 as the viral glycoprotein mediating attachment of Epstein-Barr virus (EBV) to the EBV/C3d receptor of B cells: Sequence homology of gp350 and C3 complement fragment C3d. J. Virol. 1987, 61, 1416–1420. [Google Scholar]
- Ogembo, J.G.; Kannan, L.; Ghiran, I.; Nicholson-Weller, A.; Finberg, R.W.; Tsokos, G.C.; Fingeroth, J.D. Human complement receptor type 1/CD35 is an Epstein-Barr Virus receptor. Cell Rep. 2013, 3, 371–385. [Google Scholar] [CrossRef]
- D’Arrigo, I.; Cló, E.; Bergström, T.; Olofsson, S.; Blixt, O. Diverse IgG serum response to novel glycopeptide epitopes detected within immunodominant stretches of Epstein-Barr virus glycoprotein 350/220: Diagnostic potential of O-glycopeptide microarrays. Glycoconj. J. 2013, 30, 633–640. [Google Scholar] [CrossRef]
- Patel, A.; Patel, R. Recent insights into HSV infection and disease: Results of wider genome analysis. Curr. Opin. Infect. Dis. 2019, 32, 51–55. [Google Scholar] [CrossRef]
- Whitley, R.J.; Kimberlin, D.W.; Roizman, B. Herpes simplex viruses. Clin. Infect. Dis. 1998, 26, 541–553. [Google Scholar] [CrossRef]
- Spear, P.G. Herpes simplex virus: Receptors and ligands for cell entry. Cell. Microbiol. 2004, 6, 401–410. [Google Scholar] [CrossRef]
- Cló, E.; Kračun, S.K.; Nudelman, A.S.; Jensen, K.J.; Liljeqvist, J.-Å.; Olofsson, S.; Bergström, T.; Blixt, O. Characterization of the viral O-glycopeptidome: A novel tool of relevance for vaccine design and serodiagnosis. J. Virol. 2012, 86, 6268–6278. [Google Scholar] [CrossRef]
- Risinger, C.; Sørensen, K.K.; Jensen, K.J.; Olofsson, S.; Bergström, T.; Blixt, O. Linear Multiepitope (Glyco) peptides for Type-Specific Serology of Herpes Simplex Virus (HSV) Infections. ACS Infect. Dis. 2017, 3, 360–367. [Google Scholar] [CrossRef]
- Hay, A.; Wolstenholme, A.; Skehel, J.; Smith, M.H. The molecular basis of the specific anti-influenza action of amantadine. EMBO J. 1985, 4, 3021–3024. [Google Scholar] [CrossRef]
- Wang, C.; Takeuchi, K.; Pinto, L.; Lamb, R. Ion channel activity of influenza A virus M2 protein: Characterization of the amantadine block. J. Virol. 1993, 67, 5585–5594. [Google Scholar]
- Edmond, J.; Johnston, R.; Kidd, D.; Rylance, H.; Sommerville, R. The inhibition of neuraminidase and antiviral action. Br. J. Pharmacol. Chemother. 1966, 27, 415–426. [Google Scholar] [CrossRef]
- Wiley, D.C.; Skehel, J.J. The structure and function of the hemagglutinin membrane glycoprotein of influenza virus. Annu. Rev. Biochem 1987, 56, 365–394. [Google Scholar] [CrossRef]
- Kyo, Y.; Kato, K.; Park, Y.S.; Gajhate, S.; Umehara, T.; Lillehoj, E.P.; Suzaki, H.; Kim, K.C. Antiinflammatory Role of MUC1 Mucin during Infection with Nontypeable Haemophilus influenzae. Am. J. Respir. Cell Mol. Biol. 2012, 46, 149–156. [Google Scholar] [CrossRef] [PubMed]
- Lindén, S.K.; Sheng, Y.H.; Every, A.L.; Miles, K.M.; Skoog, E.C.; Florin, T.H.; Sutton, P.; McGuckin, M.A. MUC1 limits Helicobacter pylori infection both by steric hindrance and by acting as a releasable decoy. PLoS Pathog. 2009, 5, e1000617. [Google Scholar] [CrossRef] [PubMed]
- McAuley, J.; Corcilius, L.; Tan, H.; Payne, R.; McGuckin, M.; Brown, L. The cell surface mucin MUC1 limits the severity of influenza A virus infection. Mucosal Immunol. 2017, 10, 1581–1593. [Google Scholar] [CrossRef] [PubMed]
- Foundation, C. Global Burden of Norovirus and Prospects for Vaccine Development. Available online: https://www.cdc.gov/norovirus/downloads/global-burden-report.pdf (accessed on 6 Februrary 2019).
- Tan, M.; Jiang, X. The p domain of norovirus capsid protein forms a subviral particle that binds to histo-blood group antigen receptors. J. Virol. 2005, 79, 14017–14030. [Google Scholar] [CrossRef] [PubMed]
- Marionneau, S.; Ruvoën, N.; Le Moullac–Vaidye, B.; Clement, M.; Cailleau–Thomas, A.; Ruiz–Palacois, G.; Huang, P.; Jiang, X.; Le Pendu, J. Norwalk virus binds to histo-blood group antigens present on gastroduodenal epithelial cells of secretor individuals. Gastroenterology 2002, 122, 1967–1977. [Google Scholar] [CrossRef] [PubMed]
- Rydell, G.E.; Kindberg, E.; Larson, G.; Svensson, L. Susceptibility to winter vomiting disease: A sweet matter. Rev. Med. Virol. 2011, 21, 370–382. [Google Scholar] [CrossRef] [PubMed]
- Koromyslova, A.D.; Leuthold, M.M.; Bowler, M.W.; Hansman, G.S. The sweet quartet: Binding of fucose to the norovirus capsid. Virology 2015, 483, 203–208. [Google Scholar] [CrossRef]
- Bücher, K.S.; Yan, H.; Creutznacher, R.; Ruoff, K.; Mallagaray, A.; Grafmüller, A.; Dirks, J.S.; Kilic, T.; Weickert, S.; Rubailo, A.; et al. Fucose-functionalized precision glycomacromolecules targeting human norovirus capsid protein. Biomacromolecules 2018, 19, 3714–3724. [Google Scholar] [CrossRef] [PubMed]
- Celine, A.; Claudio, S. Converting a Peptide into a Drug: Strategies to Improve Stability and Bioavailability. Curr. Med. Chem. 2002, 9, 963–978. [Google Scholar]
- Brik, A. New Strategies for Glycopeptide, Neoglycopeptide, and Glycoprotein Synthesis. In Comprehensive Natural Products II; Liu, H.-W., Mander, L., Eds.; Elsevier: Oxford, UK, 2010; pp. 55–89. [Google Scholar]
- Di, L. Strategic approaches to optimizing peptide ADME properties. AAPS J. 2014, 17, 134–143. [Google Scholar] [CrossRef] [PubMed]
- Bruno, B.J.; Miller, G.D.; Lim, C.S. Basics and recent advances in peptide and protein drug delivery. Ther. Deliv. 2013, 4, 1443–1467. [Google Scholar] [CrossRef] [PubMed]
- Pudelko, M.; Bull, J.; Kunz, H. Chemical and Chemoenzymatic Synthesis of Glycopeptide Selectin Ligands Containing Sialyl Lewis X Structures. ChemBioChem 2010, 11, 904–930. [Google Scholar] [CrossRef] [PubMed]
- Chang, J.; Patton, J.T.; Sarkar, A.; Ernst, B.; Magnani, J.L.; Frenette, P.S. GMI-1070, a novel pan-selectin antagonist, reverses acute vascular occlusions in sickle cell mice. Blood 2010, 116, 1779–1786. [Google Scholar] [CrossRef] [PubMed]
- Telen, M.J.; Wun, T.; McCavit, T.L.; De Castro, L.M.; Krishnamurti, L.; Lanzkron, S.; Hsu, L.L.; Smith, W.R.; Rhee, S.; Magnani, J.L.; et al. Randomized phase 2 study of GMI-1070 in SCD: Reduction in time to resolution of vaso-occlusive events and decreased opioid use. Blood 2015, 125, 2656–2664. [Google Scholar] [CrossRef]
- Taylor-Papadimitriou, J.; Burchell, J.M.; Graham, R.; Beatson, R. Latest developments in MUC1 immunotherapy. Biochem. Soc. Trans. 2018, 46, 659–668. [Google Scholar] [CrossRef]
- Rivalland, G.; Loveland, B.; Mitchell, P. Update on Mucin-1 immunotherapy in cancer: A clinical perspective. Expert Opin. Biol. Ther. 2015, 15, 1773–1787. [Google Scholar] [CrossRef]


© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Behren, S.; Westerlind, U. Glycopeptides and -Mimetics to Detect, Monitor and Inhibit Bacterial and Viral Infections: Recent Advances and Perspectives. Molecules 2019, 24, 1004. https://doi.org/10.3390/molecules24061004
Behren S, Westerlind U. Glycopeptides and -Mimetics to Detect, Monitor and Inhibit Bacterial and Viral Infections: Recent Advances and Perspectives. Molecules. 2019; 24(6):1004. https://doi.org/10.3390/molecules24061004
Chicago/Turabian StyleBehren, Sandra, and Ulrika Westerlind. 2019. "Glycopeptides and -Mimetics to Detect, Monitor and Inhibit Bacterial and Viral Infections: Recent Advances and Perspectives" Molecules 24, no. 6: 1004. https://doi.org/10.3390/molecules24061004
APA StyleBehren, S., & Westerlind, U. (2019). Glycopeptides and -Mimetics to Detect, Monitor and Inhibit Bacterial and Viral Infections: Recent Advances and Perspectives. Molecules, 24(6), 1004. https://doi.org/10.3390/molecules24061004