We propose to leverage a graph-based representation of a computationally probed protein structure space as follows. A nearest-neighbor graph, which we define below, is used to embed computed structures of a protein molecule of interest. Utilizing ideas and techniques from network science and network community detection, the graph is then subjected to community detection methods. The latter group/organize structures into communities. In this paper, we first evaluate the obtained communities and then demonstrate how various ranking-based techniques that leverage properties of identified communities perform in automatically selecting communities more likely to contain functionally relevant structures in the context of decoy selection. We now describe the main steps of the proposed approach.
4.1. A Graph-Based Embedding of Decoys
We first note that the ideas laid out in this paper are general and extend beyond molecular structures. However, to directly connect with our objective of evaluating these ideas in the context of decoy selection, we refer to the data (the molecular structures) where we seek an underlying organization as decoys.
As summarized above, a nearest-neighbor graph (nngraph) is employed to represent the structure space probed via computation. The graph encodes the proximity of decoys in this space. Let us denote the nngraph where the set of decoys are embedded as
. The decoys populate the vertex set
V. A local neighborhood structure is inferred for each decoy to populate the edge set
E. This is based on proximity, measuring the distance between two decoys via root-mean-squared-deviation (RMSD). First, each decoy is superimposed over a decoy selected as reference (we arbitrarily choose as reference the first decoy). The superimposition minimizes differences due to rigid-body motions, as described in [
26]. After this superimposition, the RMSD is then measured between every pair of decoys. Please note that superimposing all decoys to a reference decoy and then performing pairwise RMSD computations saves computational time. In contrast, seeking an optimal superimposition for each pair of decoys would result in quadratic (rather than linear) running time. Once such distances are available for every pair of decoys, the neighbors of each vertex
are other vertices
such that
, where
is a user-defined parameter that controls the radius of the neighborhood. A vertex is connected via an edge to each of its neighbors determined in this manner. We note that proximity query data structures (such as kd-trees and others) allow efficiently extracting the nearest neighbors of a vertex.
It is worth noting that the value of is an important consideration. A small value may result in a disconnected graph. This can be remedied by initializing to some initial value and then increasing it by over a maximum number iterations, while at the same time controlling the density of the nngraph via a parameter k. This parameter specifies the maximum number of neighbors allowed per vertex. In this way, only vertices with no more than k neighbors gain neighbors after each iterative increment of . Please note that k allows controlling the density of the graph.
We note that alternative constructions of nngraphs use
k directly rather than
. Indeed,
Section 2 presents an evaluation in terms of
k. However, in molecular structure data that may be the result of non-uniform sampling, specifying
k may result in connecting structures that are very different from each-other (that is, not in proximity).
We note that in what is described above, the nngraph is undirected. Directionality can be easily added to additionally include the role of energy in the embedding of a decoy dataset in a discrete structure, such as a graph. An edge can be directed from a vertex corresponding to a decoy with higher energy to a neighboring vertex corresponding to a decoy with lower energy.
Whether undirected or directed, the nngraph can now be investigated for its organization via community detection methods. We consider 7 representative, state-the-art methods, described below.
4.2. Community Detection Methods
Edge betweenness (Girvan-Newman): This approach was introduced to sidestep the drawbacks of hierarchical clustering. It operates based on the intuition that edges linking the communities are anticipated to possess high edge betweenness, which generalizes Freeman’s betweenness centrality [
30] from vertices to edges. To reveal the underlying community structure of the network, the Girvan-Newman method successively removes edges with high edge betweenness. Measuring edge betweenness takes
time. Since this step has to be carried out repeatedly (for each edge), the entire approach runs in
time.
Leading Eigenvector (LE): The prime objective of this method is modularity maximization (in terms of the eigenspectrum of modularity matrix) across possible subdivisions of a network [
31]. With repeated divisions, the method discovers a leading eigenvector that partitions the graph into two subgroups; the goal of maximal improvement of modularity is achieved at every step. This process terminates when modification of modularity in the sub-network starts being negative. In fact, the method is associated with additional outcomes: a spectral measure of bipartite architecture in the network and a centrality measure to detect the vertices holding nuclear positions in communities. In general, the partitioning step takes
time.
Walktrap (WT): This method employs random walks to take into account the architectural resemblance between vertices (or groups of vertices). The underlying intuition is that vertices that are within the same community are supposed to have shorter distance for random walks [
32]. The methods administers an agglomerative approach which starts from
communities (reduced to singleton clusters) and hierarchically merges two adjacent communities at each step. This is an effective approach to handle dense subgraphs of sparse graphs, which is most often the case for real-world complex networks. The method runs in time
and space
in the worst case.
Label Propagation (LP): This method is based on the intuition that each vertex in the network is supposed to follow the majority of its neighbors while joining a community [
33]. The method aims robust use of the network infrastructure instead of a predefined objective function (to optimize) or
a-priori information on the communities. At the beginning, a unique label is assigned to each vertex; that is, the method initializes
singleton communities. In progressive steps, adoption of a label comes into play for each vertex depending on the label possessed by the majority of its neighbors at that instant. This iterative process effectively performs the task of label propagation through the network and helps to form a consensus on a unique label for densely connected vertices. The process halts when each vertex and most its neighbors have an identical label. The algorithm takes linear time in the number of edges (
).
Louvain (Lo): This is a heuristic-based method focusing on modularity optimization [
34]. The method consists of iterative repetition of two stages. The first stage deals with the initial partition, where each vertex is assigned to a unique community (singleton communities). Modularity gain is measured by assigning a vertex to a neighbor community so as to exclusively search for the way to maximize positive gain. The order in which vertices are explored does not affect modularity but may increase computation time. The second stage commences with the construction of a new weighted network, whose vertices are the communities generated by the first phase. The above process continues until maximum modularity is achieved.
InfoMap (IM): This method identifies communities by using random walks along with information flow analysis [
35]. The vertices and their connections are decomposed into modules to represent the network in such a way that maximizes the amount of information in the actual network. The method tries to assign codewords to vertices; the process is efficient in terms of the dynamics on the network. A signal is transmitted to a decoder (via a limited capacity channel) who tries to decode the message, as well as to form viable candidates for the actual network. The lower the number of candidates, the more information about the actual network has been transmitted. The method runs in
time.
Greedy Modularity Maximization (GMM): This is a hierarchical agglomeration method that makes use of a greedy optimization approach. The underlying assumption is that high values of modularity are associated with good communities [
36]. Initially, each vertex itself forms a community. Then, vertices of two communities are combined together in a way that yields maximum modularity gain. This step is repeated
times. The process is represented as a hierarchical tree-like structure (a dendrogram), whose end-nodes represent the vertices of the actual network, and the internal vertices correspond to the connections; that is, the dendrogram shows a hierarchical decomposition (level-wise) of the network into communities. The method runs in
time, where
d is the depth of the dendrogram representing the network’s community architecture.
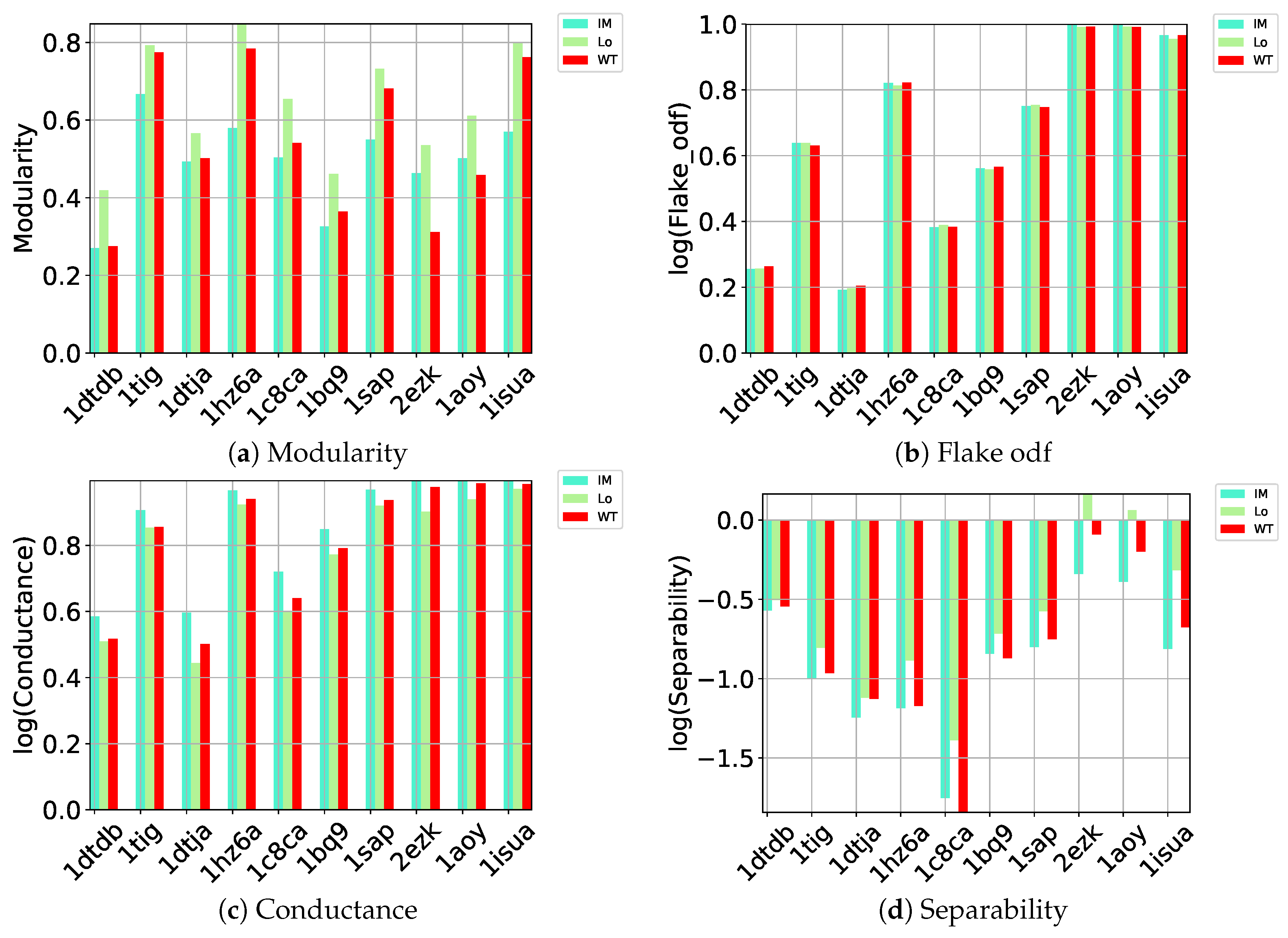
4.3. Metrics for Evaluating Community Detection Methods
A comprehensive list consisting of 15 community-recommended metrics has been considered to assess the community detection methods summarized above [
37]. We note that the following metrics are scoring functions that perform mathematical formalization of the community-wise connectivity structure of a provided set of vertices and identify communities as high-scored sets. To summarize these metrics, let us consider a graph
with
vertices and
edges, and a community is defined as a set
S of
vertices and
edges.
Fraction Over Median Degree (fomd): Let the degree of u for each vertex be denoted by , and let be the median across the degrees . Then, fomd is determined as the fraction of vertices in S with an internal degree greater than ; that is, . The denser and more cohesive the communities, the higher the associated fomd scores.
Max odf (out degree fraction): Max odf evaluates the maximum ratio of edges of a vertex in community S which point outward from S. That is, . According to Max odf, a community is characterized as a set of vertices that connect to more vertices within the set than to vertices outside of it. As a result, better communities are associated with lower Max odf scores.
Triangle(Triad) Participation Ratio (tpr): Let denotes the number of vertices which form a triangle in S. The tpr metric measures the ratio of vertices belonging to a triangle and can be formulated as: . Better community clustering yields higher tpr scores.
Internal Edge Density: For a set S, let us denote the maximum number of possible edges by . The internal edge density is the ratio of the edges that are actually in S, denoted by , over ; that is, . This metric represents the internal connectivity of a cluster (community) and a higher score indicates that there are more connections within the vertices of that community.
Average Internal Degree: This metric determines the average internal degree of the members of set S and can be formulated as: . The denser a community, the higher its average internal degree score.
Cut Ratio: Let denotes the edges that are going outward from a set S. The cut ratio score measures the ratio of over all possible edges and is defined as: . Better communities are associated with lower scores.
Expansion: This metric calculates the number of edges (for each vertex) going out of a set S and can be formulated as: . Lower scores correspond to better communities.
Edges Inside: This metric measures the internal connectivity of a set S as . Better communities are related with higher scores.
Conductance: This metric is based on the combination of internal and external connectivity and is measured as: . Lower scores relate with well-separated communities.
Normalized Cut: This metric is defined as: . The metric has the special property that concurrently meets the two following objectives: maximization of dissimilarity across communities and minimization of overall similarity (eschewing the unnatural bias for breaking up small sets). Lower values of normalized cut maintain balance between these two objectives.
Coverage: This metric measures the ratio of the number of intra-community edges to the number of edges in the graph and is defined as: . Here, . Higher coverage values indicate that there are more connections within communities rather than edges linking various communities. In fact, the ideal scenario is that communities are completely separated from one another, which would correspond to a coverage of 1 (the maximum possible value).
Average odf This metric provides the average ratio of edges that point outward of S over vertices in S and is defined as: . Lower values of average odf relate with better communities.
Modularity: This metric is based on the network model and determines the difference between the number of edges within S and the anticipated number of such edges in a random graph of exactly the same degree sequence. Modularity can be defined as: . Higher values of modularity correspond to denser connections within a community than anticipated at random.
Flake odf: This metric combines internal and external connectivity and determines the fraction of the number of vertices with fewer connections within the community than with the outside. Flake odf is defined as: . Better communities are associated with higher values.
Separability: This is a community-goodness metric [
37] based on the intuition that good communities are well-separated (have relatively few edges from set
S to the rest of the network). Separability finds the ratio between edges pointing in and outside of the set
S and is defined as:
. Higher value indicate better communities.
We note that these metrics can be grouped into four major classes [
37]: metrics based on internal connectivity (fraction over median degree, triangle participation ratio, internal edge density, average internal degree, edges inside), metrics based on external connectivity (cut ratio, expansion), metrics based on internal-external connectivity (conductance, normalized cut, max odf, average odf, flake odf), and metrics based on the network model (modularity).
4.4. Community Selection for Decoy Selection
The methods described above organize decoys into communities (more generally, groupings) by using ideas from network community detection. In previous work [
27], we have used ideas from statistics and computational geometry to group molecular structures into energy basins, leveraging the concept of the energy landscape. In that work, ranking-based methods are also debuted that evaluate groupings based on group-level characteristics and use these characteristics to rank groupings. Rankings provide a straightforward approach for decoy selection, and the quality of top-rank, selected groupings can be evaluated in the specific context of decoy selection.
Here, building on work in [
27], we summarize what group-level characteristics can be associated with communities. They fall into three categories: size, energy, and hybrid characteristics. The size of a community is the number of decoys/vertices in it. Energy can also be associated with a community. We note that the decoys we consider here are generated from template-free methods, which pursue an optimization approach that seeks to minimize the interatomic energy in a structure via a selected energy function. The result is that each decoy has an associated energy value. Given the energies of decoys in a community, the energy of a community can be defined as the minimum energy over all decoys in it or the average over the energies of the decoys in it.
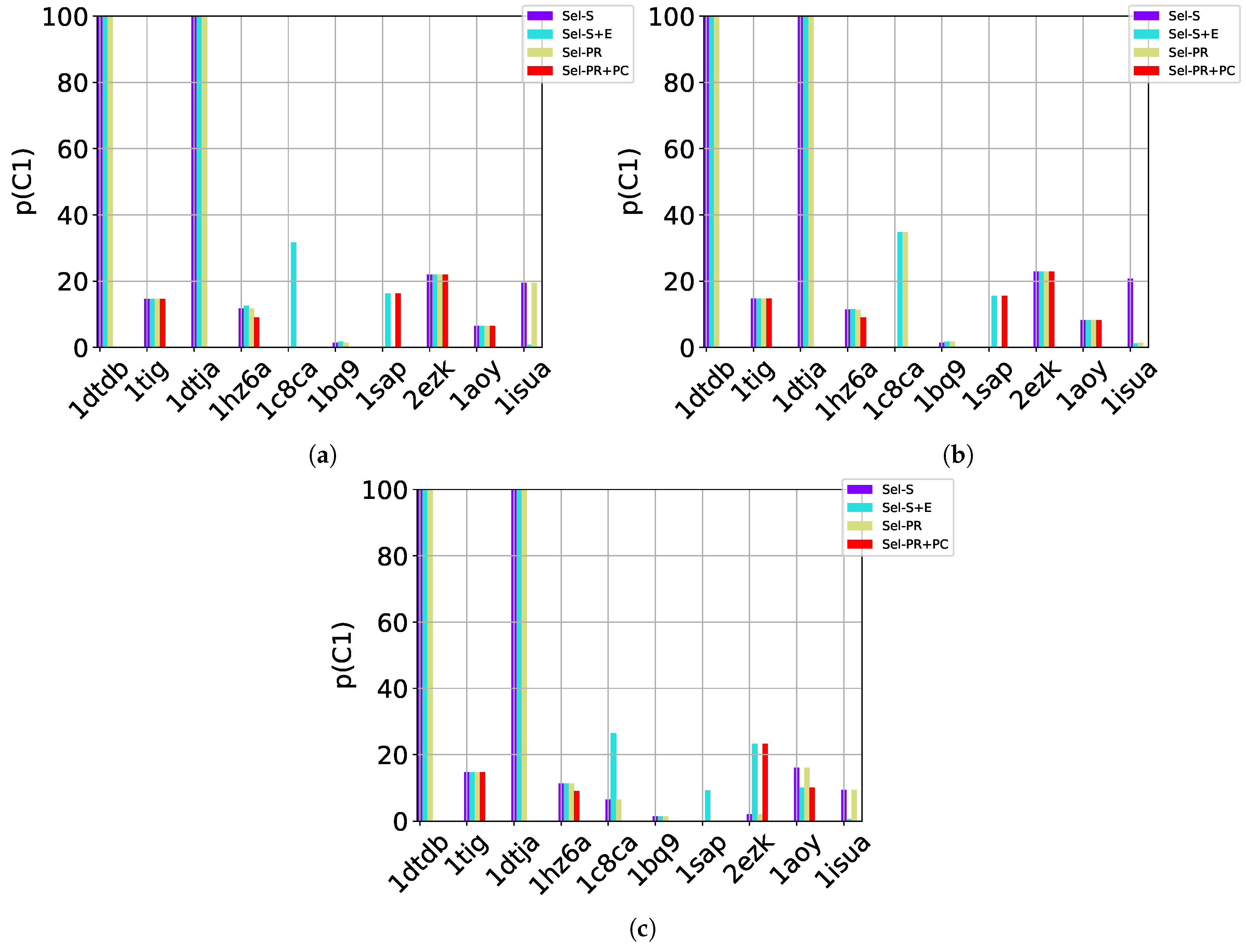
Whether size or energy, a selection technique ranks (via sorting) the communities and selects the
c top-rank communities, offering them as “prediction” for where native and near-native structures reside. We refer to the size-based ranking/selection strategy as Sel-S. We recall that considering only energy would promote a significant number of false positives, as it is well known that protein energy functions are inherently inaccurate (which is the reason decoy selection remains challenging, as summarized in
Section 1). Therefore, we consider both size and energy together in a second selection strategy to which we refer as Sel-S+E; we consider the
largest communities and then re-sort them from lowest to highest energy, selecting the top
c of them for prediction.
Hybrid characteristics consider both size and energy but additionally take into account the possibility that size and energy are possibly conflicting optimization criteria. Since solutions minimizing all conflicting objectives simultaneously are typically non-existent, Pareto-optimal solutions are sought. A Pareto-optimal solution cannot be improved in one objective without sacrificing the quality of at least one other objective. That is, a solution Pareto-dominates another solution if the following two conditions are satisfied: (1) For all optimization objectives i, ; (2) For at least one optimization objective i, .
Based on the above, two additional quantities, Pareto Rank (PR) and Pareto Count (PC), can be associated with each community C. These two quantities employ the concept of dominance, summarized above. PR(C) is the number of communities that dominate C. PC(C) is the number of communities that C dominates. It is now straightforward to use these two new, hybrid characteristics, in the same ranking-based manner. We just note that in Sel-PR, the communities are sorted by low to high PR values; in Sel-PR+PC, PC is additionally considered as follows: Communities with the same PR value are additionally sorted from high to low PCs.
4.5. Evaluating Selected Communities
We note that the ranking-based techniques described above complete an unsupervised learning framework, where we first organize decoys into communities and then automatically select a set
S of decoys from the top
c selected communities to offer as “prediction”. As we relate in
Section 2,
c can be varied so as to evaluate not only the top community but to additionally extend the analysis to the top
communities.
The set
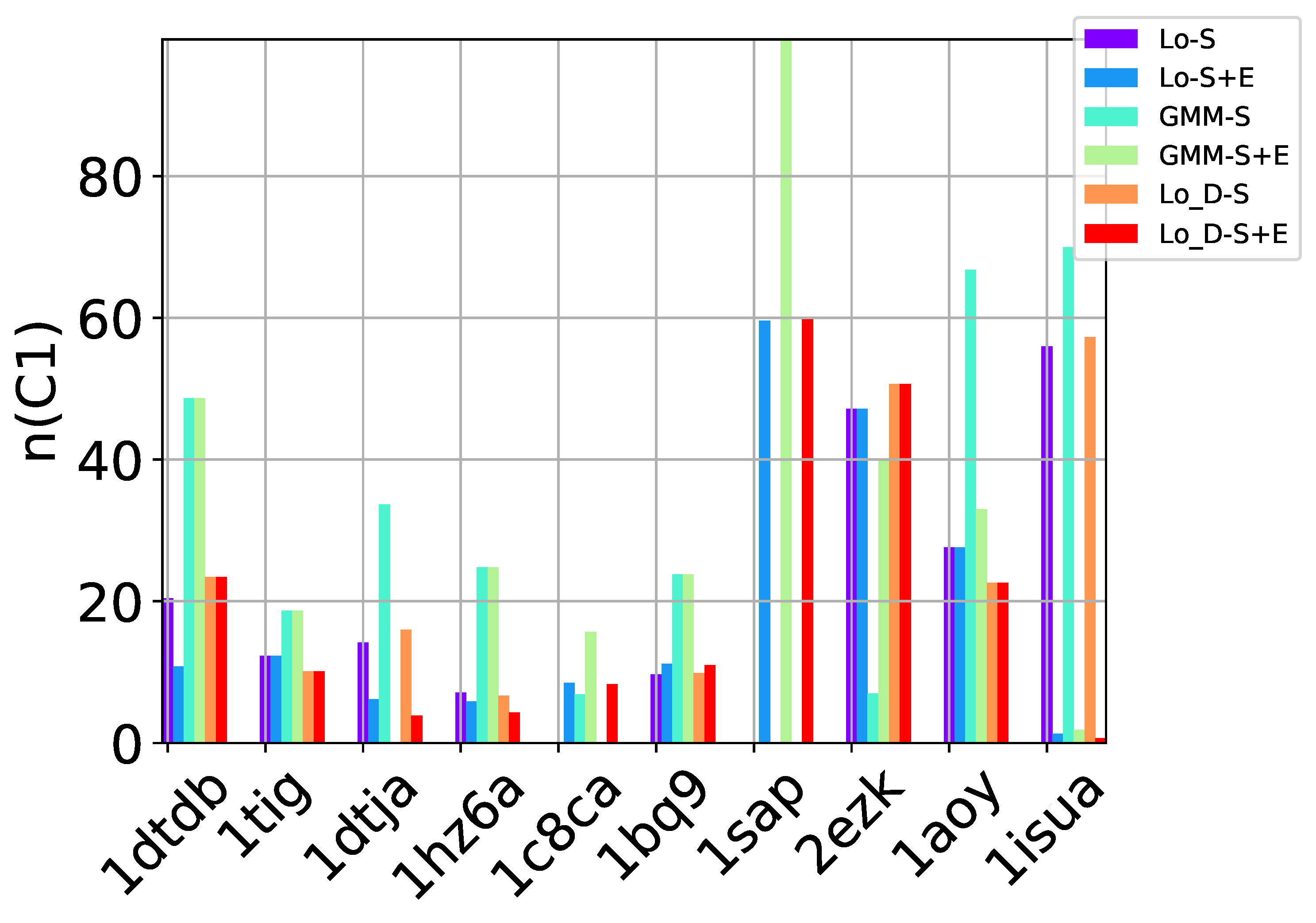
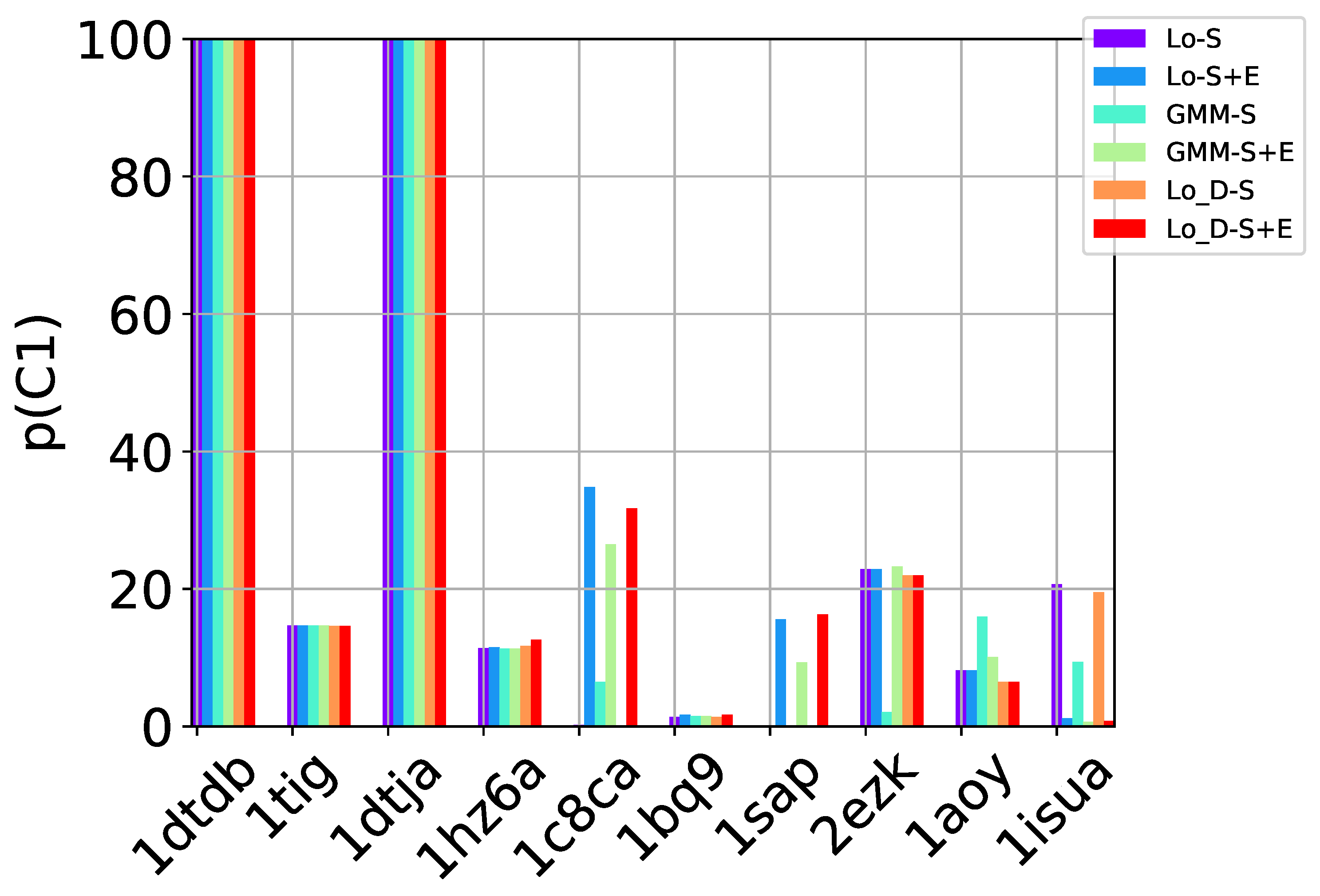
S of decoys offered as prediction can now be evaluated in the presence of a known native structure. The native structure is treated as the ground truth. Decoys within a dist_thresh RMSD of the native structure are considered near-native and are labeled as true positives (TP). We delay details on the selection of dist_thresh and its impact on the evaluation. Two metrics inspired from machine learning can be associated with a selected set
S of decoys [
27]: (1)
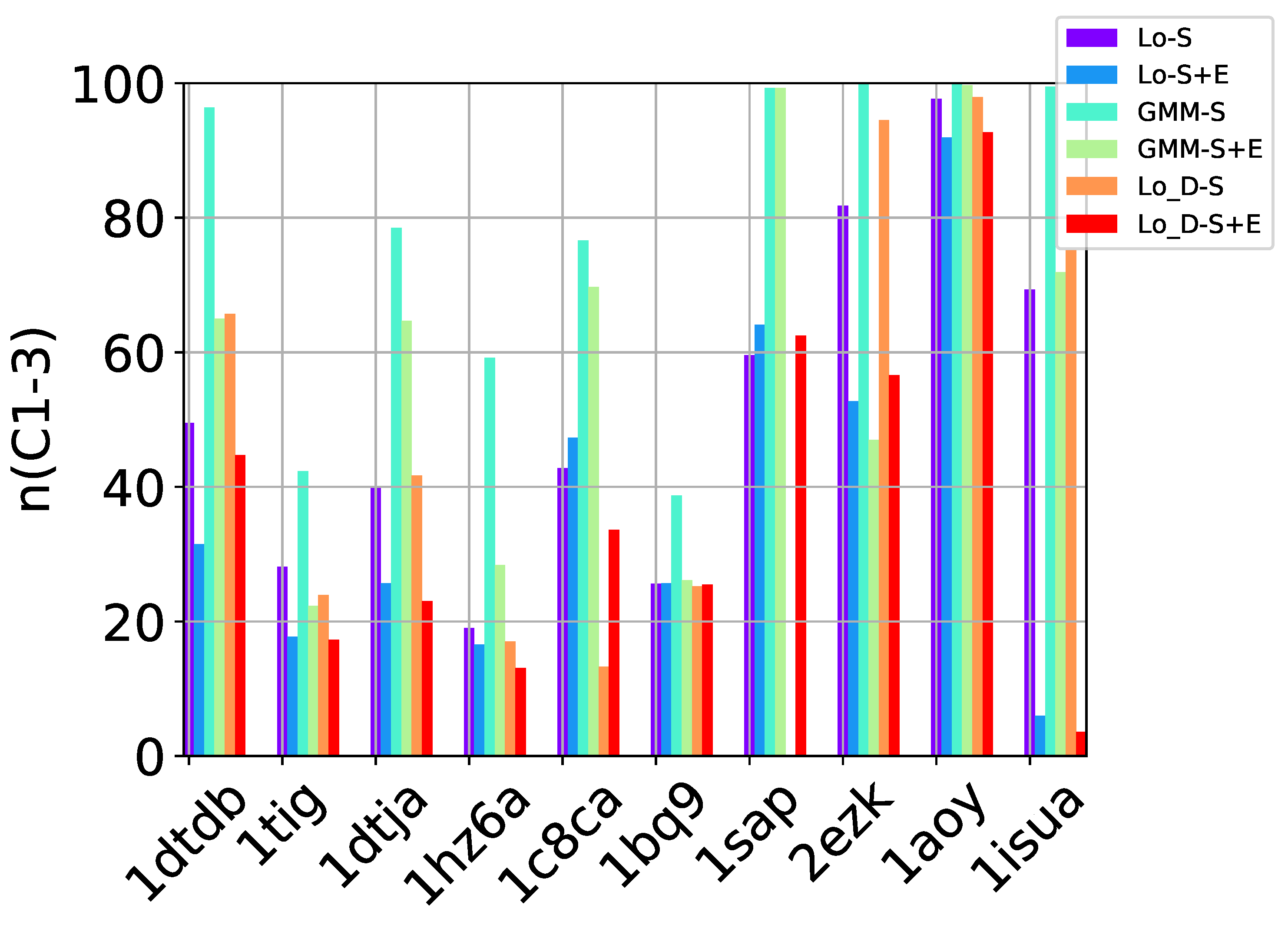
n (
number) and
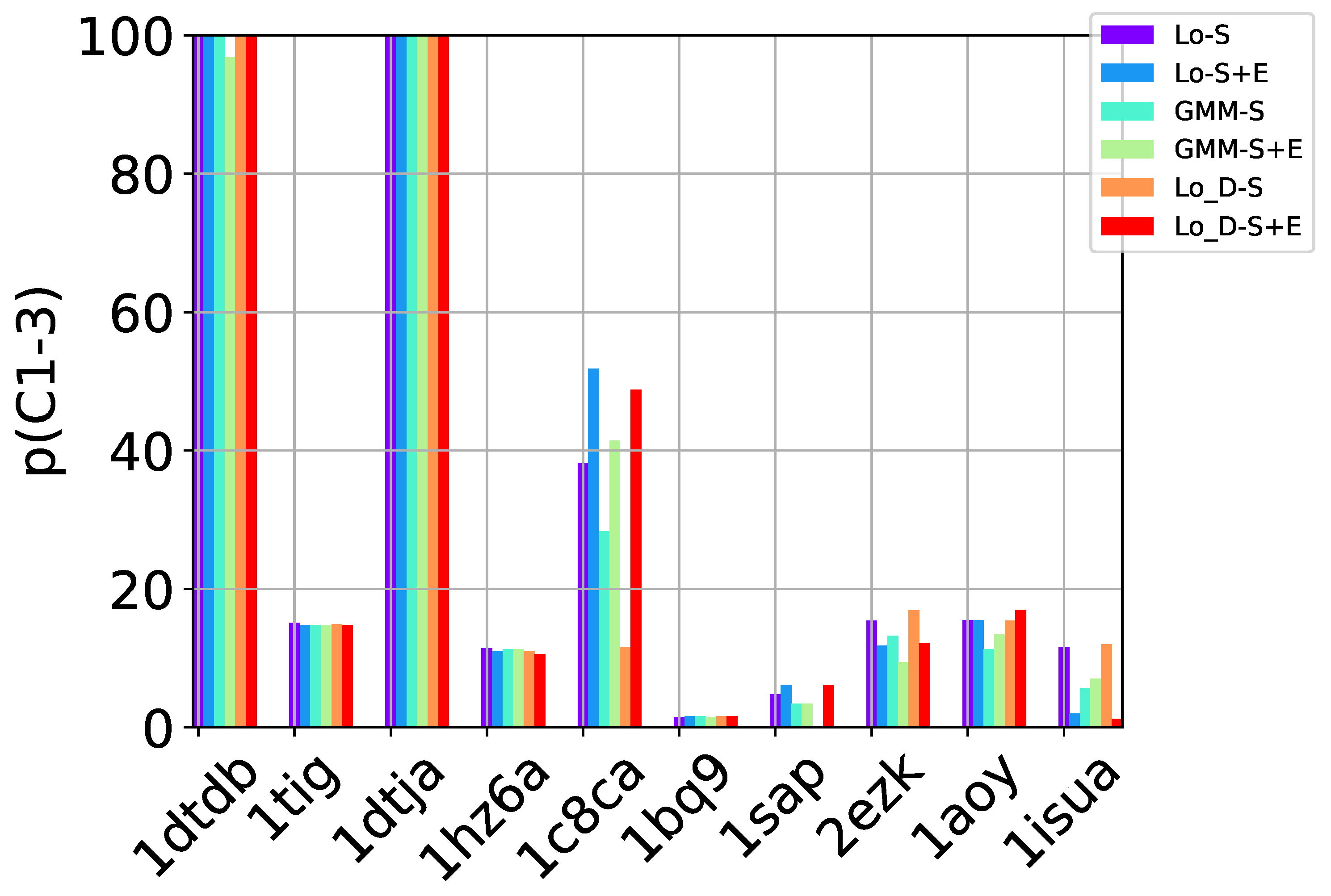
p (
purity). The first measures the percentage of near-native decoys in
S relative to the overall number of near-native decoys in the entire decoy dataset. The second measures the percentage of near-native decoys in
S over the number of decoys in
S itself. We note that
p penalizes
S if
S contains a high number of false positives (non-native decoys) and so is a useful metric for decoy selection. If a set of decoys is presented to contain the true answer but the majority of decoys are false positives, then the ratio of signal to noise is too low to be useful for automated decoy selection. In contrast, a set with more near-native decoys is a better “prediction,” as the likelihood of selecting a near-native decoy uniformly at random from it is higher when the number of false positives is low.
We note that selecting a threshold dist_thresh RMSD allows populating the positive data set and carry out further evaluation. The threshold dist_thresh is set on a per-target basis, as there are protein targets on which the quality of generated decoys suffers from either the size and/or fold of the protein under investigation. We have taken care to consider proteins that are easy, medium, and hard in their difficulty for template-free structure prediction methods, such as Rosetta. For instance, we include in our evaluation cases where Rosetta does not get close to 3 Å of the known native structure. As we have done in previous work [
27], we consider the following thresholds: If the lowest lRMSD min_dist (over all decoys) from a given native structure is ≤0.7 (these are considered easy cases), dist_thresh is set to 2 Å. Otherwise, dist_thresh is set to the minimum value that results in a non-zero number of near-native decoys in the largest-size cluster obtained via leader clustering; the latter is used as a baseline in our evaluation of community-based decoy selection. For medium-difficulty proteins (
Å< min_dist
Å), dist_thresh varies between 2−
Å. We set dist_thresh to 6 Å if min_dist
Å (these are the hard cases). This ensures a non-zero number of near-native decoys for evaluation. A detailed analysis of the impact of this threshold on cluster-based decoy selection is related in [
27].