Abstract
Evidences from more than three decades of work support the function of non-duplex DNA structures called G-quadruplex (G4) in important processes like transcription and replication. In addition, G4 structures have been studied in connection with DNA base modifications and chromatin/nucleosome arrangements. Recent work, interestingly, shows promise of G4 structures, through interaction with G4 structure-interacting proteins, in epigenetics—in both DNA and histone modification. Epigenetic changes are found to be intricately associated with initiation as well as progression of cancer. Multiple oncogenes have been reported to harbor the G4 structure at regulatory regions. In this context, G4 structure-binding ligands attain significance as molecules with potential to modify the epigenetic state of chromatin. Here, using examples from recent studies we discuss the emerging role of G4 structures in epigenetic modifications and, therefore, the promise of G4 structure-binding ligands in epigenetic therapy.
1. Introduction
DNA-protein interactions package genomic DNA into globular chromatin. This along with modifications of nucleic acids—for example, methylation of cytosine residues—that otherwise do not affect the sequence of chromosomal DNA constitute the epigenetic state of the genome [1] Modifications of the epigenetic status are closely associated with several diseases including cancer, neurodegenerative and metabolic disorders and autoimmune diseases [2,3,4,5,6]. Therefore molecules that can alter or ‘correct’ aberrant epigenetic modifications are of importance as therapeutics—and are sometimes called ‘epigenetic drugs’ [5].
The non-duplex DNA secondary structure called G-quadruplex (commonly called G4), particularly molecules/ligands that specifically interact with G4 structures gain significance in this context. The biological role of G4 structures was first implicated when G-rich telomeric repeats were observed to adopt the four-stranded secondary structure through stacking interactions of guanine-tetrads (Figure 1) [7,8,9,10]. Interestingly, genome-wide analysis revealed a sequence capable of forming G4 structures was enriched in gene regulatory regions (Figure 1) [11,12,13]. This was initially observed through genome-wide analysis in bacteria including E. coli—based on which authors proposed a widespread gene regulatory role of G4 structures [11]. Prevalence and conservation within promoters of homologous genes in human, chimpanzee, mouse, and rat further implicated G4 structures in gene regulatory function (Figure 1) [11,14,15]—this was experimentally observed to be so using G4-binding ligands [16]. Gene regulatory functions, and in addition role of G4 structures in replication and recombination have been reviewed earlier [17,18].
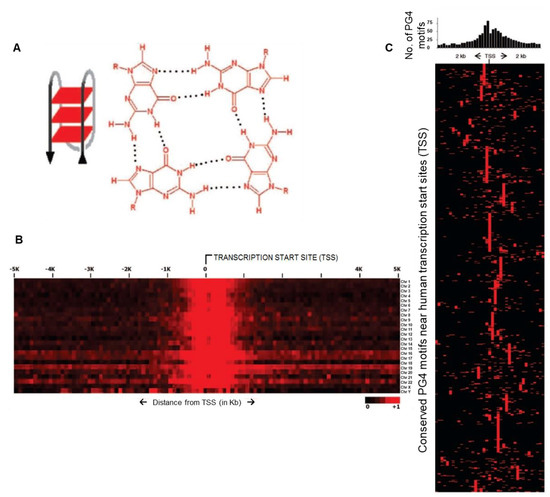
Figure 1.
The G4 structure and its relevance. (A) G-quadruplex (G4) structural illustration: left panel with G-tetrad planes forming an intramolecular G4 stem, right panel shows Hoogsteen base-pairing of guanines making a G-tetrad. (B) Heat map showing averaged relative enrichment of potential G4 (PG4) sequences near TSS across all chromosomes in the human genome (density of PG4s in 100 base windows). (C) Heat map of conserved promoter PG4s across organisms: upper panel shows enriched PG4 motifs near TSS, lower panel shows conservation of PG4 motif clusters between human and ‘orthologous’ mouse and rat promoters (red boxes for PG4 motifs per 100 bp window, each row displays individual promoters); 773 human promoters containing 1414 PG4 motifs shown here. Reprinted (adapted) with permission from (Verma, A. et al. Genome-Wide Computational and Expression Analyses Reveal G-Quadruplex DNA Motifs as Conserved cis-Regulatory Elements in Human and Related Species. J. Med. Chem. 51, 5641–5649 (2008)). Copyright (2008) American Chemical Society.
The involvement of G4 structures in epigenetic functions, though noted in early work, has received more direct attention in a recent review, where G4 structures have been implicated as structural mediators of epigenetic modifications in chromatin [19]. The authors have focused on human telomerase reverse transcriptase (hTERT) promoter transcription factor binding sites and telomerase reactivation in cancer as a case study for epigenetic regulation mediated by G4 structures. Therefore, G4 structure-binding ligands, including ones available as nutrient molecules might be important in epigenetic regulation/modifications—particularly in conditions with established epigenetic aberrations. G4 structure binding ligands have been previously characterized from natural sources (e.g., berberine [20], sanguinarine [21]) and are also artificially synthesized (e.g., ligand 360A [22], pyridostatin [23]). Several of such ligands were found to affect gene regulation through possible epigenetic mechanisms implicated in cancers as well other disorders.
Role of epigenetics in cancer has gained significance as multiple genes and microRNAs related to cancer initiation and progression were reported to exhibit epigenetic abnormalities [24]. Most of these were results of differential regulation of genes coding for epigenetic modifiers itself, leading to silencing of tumor suppressor genes, activation of oncogenes, and altered expression of microRNAs. Several reviews cover this aspect in substantial detail [3,25,26].
Attempts to find out any possible association of G4 structures with genes reported to undergo epigenetic modifications in various cancer types, yielded evidence that several of these gene promoters exhibit potential G4 sequence (PG4), for example, hTERT [27], H19 [28], KRAS [29] BCL-2 [30], RET [31,32], PARP-1 [33,34]. Interestingly, these epigenetic modifications were shown to be regulated by epigenetic modifiers such as DNA methyltransferases (DNMTs) and polycomb group (PcG) proteins, like EZH2, which has been reported to bind to G4 structures in vitro and in vivo respectively [35,36,37] (vide infra). This indicates the possibility that G4 structures could potentially recruit epigenetic modifiers.
With these in mind, we herein focus reviewing literature on how G4 structure-binding molecules and proteins might be important for development of epigenetic therapeutic interventions in future, particularly in cancer.
2. G4 Structures Impact Local Chromatin at Telomeric and Extra-Telomeric Sites
2.1. The G4 Structure and DNA Base Modifications
Methylation at the C5 position of cytosine within (GGGGCC)8•(GGCCCC)8 repeats—associated with two neurodegenerative diseases amyotrophic lateral sclerosis (ALS) and fronto-temporal dementia (FTD)—was reported to influence stability of G4 structures in the promoter as well as coding region of the C9orf72 gene [38]. In addition, cytosine methylation of dCGG repeats in the FMR1 gene, which expand during progression of fragile X-mental retardation syndrome, were reported to result in stabilization of G4 structures formed by the dCGG repeats in vitro [39]. Increased G4 structure stability upon methylation of d(CGG)n oligomers was therefore implicated in repression of FMR1 in fragile X syndrome [39]. In similar lines, C5 methylation within the G-rich promoter region of the B-cell lymophoma (BCL-2) gene, which forms G4 structure, was observed to lead to repression of BCL-2 known to be abnormally overexpressed in many cancers [30,40]. Results showing that C5 methylation stabilized folding of the G4 structure-forming oligomer further implicated role of DNA methylation-dependent stability of the G4 structure in epigenetic regulation of BCL-2 [30]. Recently, a CTCF binding site located in the first exon of the human telomerase hTERT gene was reported to be disrupted due to the formation of a stable G4 structure following C5 methylation. This was found to result in marked reactivation of hTERT—the enzyme essential for telomere synthesis found to over-expressed in more than 90% of human cancers [41,42]. Furthermore, 8-oxoguanine (8oxoG) modification of DNA—from oxidation through reactive oxygen species—was shown to affect stability of promoter G4 structures resulting in altered expression of multiple genes like c-myc, VEGF, NTHL1, and KRAS (Figure 2) [29,43,44,45,46].
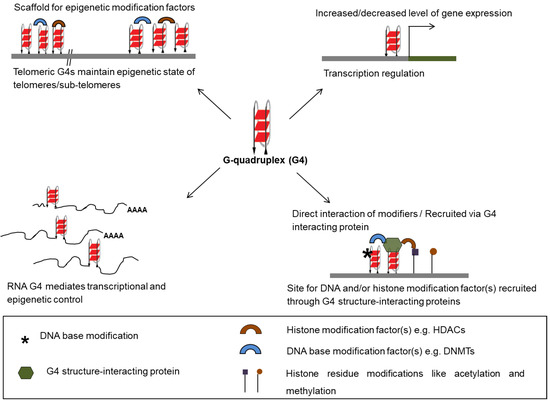
Figure 2.
Regulatory roles of G-quadruplex. G4 structures in DNA and RNA are involved in the transcriptional and epigenetic regulation of the genome by acting as anchor sites for recruitment of transcription factors at promoters. G4 structure-interacting proteins recruit epigenetic modifiers upon binding to G4 structures at telomeres and extra-telomeric sites.
2.2. G4 Structures and Histone Protein Modifications
Recent work further reveal the possible role of the G4 structure in arrangement/modification of histones—proteins required to package chromatin—which help determine the epigenetic state of the genome [47,48,49]. It was noted that the absence of REV1, a helicase that resolves the G4 structure, resulted in replication-associated errors [47]. Because of this, after replication, in cells without REV1 the β-globin gene locus was found to lose the K9-dimethylated variant of histone H3 critical for maintaining the repressed state of chromatin. This resulted in de-repression of the β-globin gene. The role of the G4 structure was studied further through artificial insertion a G4 structure in the lysozyme C gene, which otherwise did not have a G4 structure and therefore was unaffected by the absence of REV1 [47]: artificial insertion of the G4 structure resulted in activation of lysozyme expression in cells without REV1. In addition, it was also found that the presence or absence of the G4 structure affected histone H3 modifications (K4-trimethylation and K9/K14-acetylation) at the BU-1 promoter, which was dependent on the presence of the G4-helicase REV1 [49].
2.3. G4 Structures Engage Epigenetic Factors through G4 Binding Proteins
It has been noted that modifications of DNA and histones can cooperate to engage or disrupt binding of regulatory factors [50,51]. Therefore, the role of G4 structures in DNA/histone modifications are expected to impact association of regulatory factors. This was further supported when the binding of epigenetic factors was observed to be dependent on the promoter G4 structure within the cyclin-dependent kinase p21 and telomerase (hTERT) promoters [22,27]. Interestingly, at the p21 promoter this was through the recently discovered function of the telomeric protein TRF2 as a transcription factor. Recruitment of the epigenetic repressor complex of proteins including REST/co-REST/LSD1 was through TRF2—where TRF2 binding required presence of the p21 promoter G4 structure [22]. Similarly, in another study authors noted that critical histone modifications for hTERT repression in normal adult cells required binding of the metastasis suppressor factor NME2 [27]. Occupancy of NME2 on the hTERT promoter depended on the promoter G4 structure—consistent with NME2-G4 association noted earlier [52]—thereby making the hTERT histone modifications and expression G4-dependent (Figure 2) [27].
Furthermore, high-affinity binding of G4 structures with factors that methylate DNA called DNA methyltransferases (DNMT) was reported recently [53]. Along with earlier work implicating association between G4 structures, global DNA methylation and DNMTs these further supports the possible role of G4 structures in epigenetic modifications [35,54].
2.4. G4 Structures Formed by RNA: Role in Epigenetic Modifications
Biological role of G4 structures formed by RNA sequences (RNA-G4) in transcription/translation [55,56,57,58], including epigenetic regulation [59] and the potential of RNA-G4 structures as targets for small molecule-based therapies has been reviewed (Figure 2) [60,61]. Mature human microRNAs were recently discovered to contain RNA-G4 structures that were implicated in miRNA-mRNA-based transcriptional regulation [62]. Multiple studies show the telomeric repeat-containing RNA (TERRA), a long non-coding RNA molecule (lncRNA) that forms G4 structures (RNA-G4), to be important in this context [63,64]. Interestingly, it was noted that RNA-G4 structures formed by TERRA bind to lysine-specific histone demethylase1 (LSD-1)—a histone modifier protein—and this catalyzes the removal of methyl groups from histone 3 at lysine 4 and lysine 9 (H3K4/9) in metazoans [65].
2.5. Telomeric G4 Structures and Epigenetic Modifiers
Formation of G4 structures by (TTAGGG)n telomeric repeats in vertebrates has been implicated in the activity of the telomere synthesizing protein telomerase [66]. Relatively recent work reveal telomeric G4 structures might be involved in maintaining the chromatin state of the telomeric/subtelomeric regions (Figure 2) [67,68]. RNA-G4 binding proteins like TLS/FUS and EWS bind TERRA as well as telomeric G4 structures forming a ternary RNA-DNA G4 complex [69,70,71,72]. This complex of proteins was observed to recruit the methyltransferase Suv4-20h2, which tri-methylated K20 residues of histone H4 one of the prime histone modifiers at telomeres [72]. In addition, association of TERRA with the G4 structure-binding RGG3 domain of TLS/FUS mediates K9 tri-methylation of histone H3, which is an essential heterochromatin mark at telomeres [73].
Interestingly, interaction of ATRX, an epigenetic modifier of SWI2/SNF2 family, with telomeric G4 structures was shown to be important in maintaining the ‘dynamic’ state of telomeric chromatin in undifferentiated pluripotent cells [74]. Binding of CBX5 (chromobox homolog 5) along with ATRX at telomeres was involved in inducing the repressed chromatin state. At the same time, ATRX bound to TTAGGG repeats interacted with K4 of H3.3 histones imparting features of open chromatin. In differentiated cells telomeres are predominantly in a closed conformation. Therefore the ATRX-G4 interaction mediated cell cycle-specific ‘open/closed’ telomeric state in undifferentiated pluripotent cells appears to be of significance [74].
3. Promise of G4 Structure Binding Molecules in Epigenetics Based Therapeutics
3.1. G4 Structure Binding Ligands as Potential Modifiers of the Epigenetic State
Epigenetic drugs include compounds that bind to proteins that affect chromatin organization, such as histone methylation/demethylation inhibitors, bromo-domain inhibitors, HAT inhibitors, HDAC inhibitors, and DNA methyltransferase inhibitors: many of which are at different stages of clinical trials as anti-cancer molecules [75,76,77]. Ligand(s) that bind to G4 structures in DNA/RNA and thereby modulate changes in the chromatin in ways described above, therefore, could be of importance as ‘epigenetic modifiers’. With this in mind, in the following sections we focus on G4 structure-binding ligands that could be relevant in epigenetics. The role of G4 ligands as potential anticancer agents and in antiviral therapy through functions other than epigenetic mechanisms have been reviewed earlier [78,79].
Berberine, a plant alkaloid known to bind G4 structures [20,21], was found to induce hypomethylation of the TP53 promoter leading to apoptosis in the human multiple myeloma U266 cells [80]. In addition, berberine has been shown to down-regulate histone deacetlyases (HDACs) [81]; up-regulate histone acetyltransferases, demethylases, and methyltransferases, resulting in wide spread changes in methylation of lysine K4/K27/K36 of histone H3 (i.e., H3K4me3, H3K27me3, and H3K36me3) [82]; and, interestingly, affect interaction of DNMTs with microRNAs during malignant transformation of colorectal cancer cells [83]. Although G4-berberine interaction was not directly studied, together these studies implicate berberine in epigenetic functions that could be through G4 structures (Table 1). Similarly, sanguinarine, another molecule obtained from plants, that binds the telomeric and c-myc promoter G4 structures [21,84] was noted to epigenetically modify chromatin by inducing altered histone methylation [85].

Table 1.
G4 structure binding ligands and their biological roles including in epigenetics.
Based on the effect of the G4 structure observed in replication (described above) small molecules derived from modification of the well-known G4 binding ligand pyridine 2,6-dicarboximide (PDC) was screened using the BU-1 locus in DT-40 chicken cells [105]. This resulted in several ligands (e.g., PDC12, 14, 22, 23, 25, and 40) that induced transcriptional reprogramming of the BU-1 locus. This was found to be through the loss of trimethylated-K4 of histone H3 (H3K4me3) and interestingly, cytosine methylation in the BU-1 gene. Together these suggested the role of the G4 structure in re(placement) of histone marks, a hallmark of epigenetic regulation [105].
As mentioned earlier, epigenetic reorganization of the hTERT promoter through interaction of NME2 with the hTERT promoter G4 structures results in repression of abnormally overexpressed hTERT in cancer cells [27]. Prompted by this authors checked several known G4 binding ligands. Many of these like 9A, 9B, and Bis-ANON (acridine based), JD59 (bis-indole carboxamide) and RR110 (pyridostatin based) showed more than 50% reduction in hTERT expression, which was shown to be dependent on presence of the hTERT promoter G4 structure [27]. In addition to this, several other G4 ligands have been reported to repress hTERT expression [106]. These findings could be useful in development of G4 based epigenetic therapeutic interventions for restricting hTERT overexpression as seen in cancer cells.
Transcription regulation of p21—activation of which results in growth arrest of cancer cells on treatment with anticancer drugs—was dependent on TRF2-G4 interactions that induced epigenetic modifications [22]. Anti-cancer drug resistance often results from ineffective p21 activation [107]. The role of the G4 structure in p21 epigenetic regulation was tested using the pyridine derivative G4 ligand 360A [22,108]. Authors showed that aggressive MDAMB-231 breast cancer cells, otherwise resistant to the anti-cancer drug doxorubicin, regained doxorubicin-sensitivity in presence of 360A. This was through 360A-mediated de-repression of p21 in MDAMB-231 cells suggesting the potential function of G4 ligands in modification of cellular epigenetic mechanisms (Figure 3) [22].
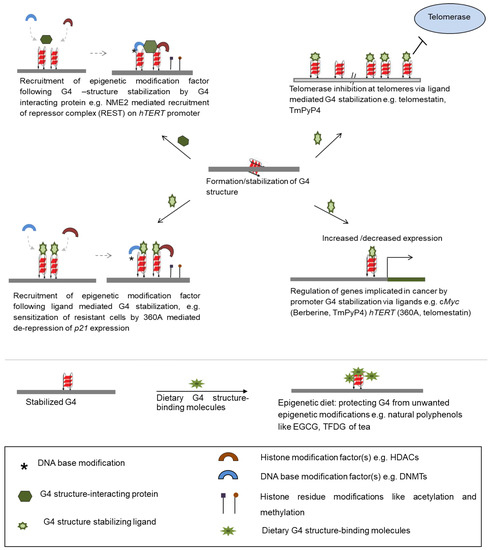
Figure 3.
Ways of therapeutic intervention through G4 structure-binding small molecules. Stabilization of G4 structure by means of ligands inhibits telomerase activity at telomeres, regulates expression of genes at transcriptional and epigenetic levels. Proteins stabilizing G4 structure upon binding, allow epigenetic modifiers to dock at the site, further regulating gene expression. Several dietary components protect G4 structure from unwanted modifications by binding to the secondary structure. HDACs: histone deacetlyases; DNMTs: DNA methyltransferases.
3.2. Dietary G4 Ligands Can Affect Epigenetic Modifications
Dietary molecules that affect epigenetics and resulting changes in gene regulation include tea polyphenols like ellagic acid [109], epigallocatechin gallate [110], curcumin [111], genistein [112], resveratrol [113], and sulforaphane [114]. Amongst these, epigallocatechin gallate and theaflavin-3,3′-digallate (TFDG) from green tea and black tea, and resveratrol from berries were reported to bind telomeric G4 structures with high affinity [115,116]. Curcumin and ellagic acid were also shown to bind KRAS G4 sequences in vitro [117]. Deficiency of the dietary component folate, a methyl group donor metabolite, was observed to result in global hypomethylation of CpG islands and increased G4 structure formation in HeLa cells [118], consistent with decreased methylation within CpG islands that harbor potential G4 structures in a genome wide study [54]. Berberine was found to impair parasitic infections from Eimeria sp. through epigenetic modifications in cells of the gastrointestinal tract in mouse models showing potential as a food supplement in animal husbandry [119]. Furthermore, ROS-induced oxidative stress is known to result in 8-oxo-guanine modifications of Guanine base. As described above such modifications have been reported to affect stability of the G4 structure leading to altered function [44,45,120,121]. Therefore, the effect of dietary anti-oxidants on G4 structures and related epigenetics could be interesting to consider in future (Figure 3).
3.3. G4 Structure-Binding Epigenetic Modifier Proteins: Potential for Development of Epigenetic Intervention Agents
Nucleolin, possibly the first protein noted to interact with G-rich oligonucleotides that adopt G4 structure was found to be involved in epigenetic modification of histone H1 implicated in decondensation of chromatin [122,123,124].
Interestingly, in 2009, a metastasis suppressor factor NME2 was found to not only associate with the promoter G4 structure of the oncogene c-myc but also important for transcription regulation of c-myc suggesting transcription regulatory roles of G4 structures in association with regulatory factors [52]. More recently, NME2 was shown to be involved in epigenetic regulation of hTERT through association with the G4 structure in the promoter of hTERT [27].
Epigenetic modifiers like DNMT3A and 3B, EZH2, and ATRX (as discussed earlier) bind to G4 structures where epigenetic regulatory functions mediated through such interactions might be of clinical significance [35,37,53,74]. In addition, interaction of TRF2 with G4 structures and/or G-rich binding sites might be important because of epigenetic regulation of genes like p21 and several other [22], which interestingly was also noted to be dependent on telomere length [125]. The TRF2-mediated epigenetic regulation of p21 appears to be of added significance in aggressive as well as commonly encountered drug resistant cancer cells.
Somewhat in line with these studies a large scale screening for G4 structure interacting factors using protein microarrays comprising >9000 human proteins found several factors that are involved in binding nucleosomes [126]. It is also likely that function of the G4 structure helicases like FANCJ [127], BLM [128], WRN [129], and REV-1 [49] would be important in epigenetic modifications in a replication-dependent manner (as demonstrated for REV-1) [49]. Similarly factors that bind to RNA G4 structures like the polycomb repressive complex 2 (PRC2) [37], TLS/FUS [72,73], EWS [71], and hnRNP A1 [130,131] suggest further importance of G4 structure-protein interactions in epigenetic regulation [37,71,72,73]. It is of interest to note here that many of the G4 structure interacting proteins possess the positively charged Arg-Gly-Gly (RGG/RG) motif containing domain, which is noted to be important for G4 structure binding (Figure 3) [132,133,134].
4. Conclusions and Future Perspectives
For more than a decade G4 structures have been implicated in epigenetic modifications that might impact state of chromatin resulting in altered gene regulation. Recent studies through more direct studies show how G4 structures modify chromatin by not only change in histone and/or DNA modification but also during replication. A growing number of reports suggest that G4 structure has significant role to play in epigenetic control of genes involved in various neurological disorders as well as cancer. Herein we have focused on these studies. This is discussed along with studies that have focused on design and characterization of different classes of small molecule ligands that specifically bind to G4 structures. However, the epigenetic effects of these ligands remain to be confirmed in more physiologically relevant settings, such as in animal models.
Together, these bring forth the promise of the G4 structure binding ligands, including dietary molecules, in affecting epigenetic mechanisms. This becomes particularly notable in cases where changes in epigenetic pattern have been shown to play a role in diseases such as cancer and neurodegenerative disorders. It is possible, therefore, that ligands that bind to G4 structures reinstate/rescue aberrant epigenetic modifications in chromatin and thereby enable therapeutic interventions. Mitochondrial DNA (mtDNA) G4 structures are another promising avenue for small molecule therapeutics; although there is a lack of sufficient data in the field currently mtDNA G4 structures are increasingly under consideration as targets for therapeutic intervention in mitochondrial diseases [135]. A recent study reported that RHPS4, a G4-binding ligand thought to localize to nuclear G4s showed preferential binding to mtDNA G4 structures in both cancerous and non-cancerous cell lines thereby opening new avenues to study mtDNA transcriptional and epigenetic regulation using G4-binding molecules specific to mtDNA [136]. Although intracellular G4 structures are primarily right-handed in orientation, left-handed G-quadruplexes have been observed in vitro [137]; however, there is lack of sufficient evidence to validate their formation by nuclear or mitochondrial G-rich sequences in cellulo. Both left and right handed G4 structures have been shown to form from the same nucleic acid sequence mediated by small molecule binding [138]. Questions about the effect of G4 orientation and ‘handedness’ on genome structural dynamics need to be addressed to improve on the structural sensitivity of G4-binding small molecules.
Although several G4 helicases and G4 binding proteins are known to be associated with genetic diseases not much has been explored for therapeutic interventions. The current arsenal of G4 related therapeutics comprise of G4 selective ligands, which are being attempted to be upgraded to locus specific targeting and G4 DNA aptamers which can bind and inhibit G4 interacting proteins [139]. Aptamers based on promoter G4s are being focused on to serve as G4 decoys in several cases and also being considered as a drug delivery tool as in case of AS1411-drug conjugate nanoparticles [140].
The multitude of data on the biological significance of G4 and G4 structure interacting proteins could also be utilized to design novel drug molecules. Small peptides or peptidomimetics with better stability could be designed to bind and stabilize G4 structures as well as mediate epigenetic changes at locus of interest. This strategy combined with conventional G4 ligands or alone, could be effective in inducing desired epigenetic modification to counter a particular disease state. It could also be specifically delivered to cancer cells using above mentioned aptamer based delivery systems [140]. However, detailed knowledge of protein structure and the interacting G4 structures is still required to develop molecules which can both bind and recruit epigenetic factors. Perhaps tailoring this for a specific locus would be equally important. In conclusion, the G4 structure has been deemed as a promising target in anti-cancer therapy for long now—its emerging role in epigenetic control of pharmacogenes could be a new-found angle in this battle.
Author Contributions
A.S., A.G. and S.C. wrote the manuscript. A.S. and A.G. made the figures and S.C. conceptualized and supervised the overall preparation of the manuscript.
Funding
SC research group is supported by CSIR, was partly supported by the Wellcome Trust/DBT India Alliance Fellowship (grant number 500127/Z/09/Z) awarded to SC from 2011–2017.
Acknowledgments
A.S. and A.G. acknowledge CSIR and DST-SERB for research fellowship, respectively. All members of S.C. group are acknowledged for engaging discussions and support.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Zentner, G.E.; Henikoff, S. Regulation of nucleosome dynamics by histone modifications. Nat. Struct. Mol. Biol. 2013, 20, 259–266. [Google Scholar] [CrossRef] [PubMed]
- Feinberg, A.P.; Ohlsson, R.; Henikoff, S. The epigenetic progenitor origin of human cancer. Nat. Rev. Genet. 2006, 7, 21–33. [Google Scholar] [CrossRef] [PubMed]
- Esteller, M. Epigenetics in Cancer. N. Engl. J. Med. 2008, 358, 1148–1159. [Google Scholar] [CrossRef] [PubMed]
- Dawson, M.A.; Kouzarides, T. Cancer Epigenetics: From Mechanism to Therapy. Cell 2012, 150, 12–27. [Google Scholar] [CrossRef] [PubMed]
- Prachayasittikul, V.; Prathipati, P.; Pratiwi, R.; Phanus-umporn, C.; Malik, A.A.; Schaduangrat, N.; Seenprachawong, K.; Wongchitrat, P.; Supokawej, A.; Prachayasittikul, V.; et al. Exploring the epigenetic drug discovery landscape. Expert Opin. Drug Discov. 2017, 12, 345–362. [Google Scholar] [CrossRef]
- Berson, A.; Nativio, R.; Berger, S.L.; Bonini, N.M. Epigenetic Regulation in Neurodegenerative Diseases. Trends Neurosci. 2018, 41, 587–598. [Google Scholar] [CrossRef] [PubMed]
- Sen, D.; Gilbert, W. Formation of parallel four-stranded complexes by guanine-rich motifs in DNA and its implications for meiosis. Nature 1988, 334, 364–366. [Google Scholar] [CrossRef] [PubMed]
- Sundquist, W.I.; Klug, A. Telomeric DNA dimerizes by formation of guanine tetrads between hairpin loops. Nature 1989, 342, 825–829. [Google Scholar] [CrossRef]
- Balagurumoorthy, P.; Brahmachari, S.K. Structure and stability of human telomeric sequence. J. Biol. Chem. 1994, 269, 21858–21869. [Google Scholar]
- Lipps, H.J.; Rhodes, D. G-quadruplex structures: In vivo evidence and function. Trends Cell Biol. 2009, 19, 414–422. [Google Scholar] [CrossRef]
- Rawal, P.; Kummarasetti, V.B.R.; Ravindran, J.; Kumar, N.; Halder, K.; Sharma, R.; Mukerji, M.; Das, S.K.; Chowdhury, S. Genome-wide prediction of G4 DNA as regulatory motifs: Role in Escherichia coli global regulation. Genome Res. 2006, 14, 644–655. [Google Scholar] [CrossRef] [PubMed]
- Huppert, J.L.; Balasubramanian, S. G-quadruplexes in promoters throughout the human genome. Nucleic Acids Res. 2007, 35, 406–413. [Google Scholar] [CrossRef] [PubMed]
- Patel, D.J.; Phan, A.T.; Kuryavyi, V. Human telomere, oncogenic promoter and 5′-UTR G-quadruplexes: Diverse higher order DNA and RNA targets for cancer therapeutics. Nucleic Acids Res. 2007, 35, 7429–7455. [Google Scholar] [CrossRef] [PubMed]
- Verma, A.; Halder, K.; Halder, R.; Yadav, V.K.; Rawal, P.; Thakur, R.K.; Mohd, F.; Sharma, A.; Chowdhury, S. Genome-wide computational and expression analyses reveal G-quadruplex DNA motifs as conserved cis-regulatory elements in human and related species. J. Med. Chem. 2008, 51, 5641–5649. [Google Scholar] [CrossRef] [PubMed]
- Yadav, V.K.; Abraham, J.K.; Mani, P.; Kulshrestha, R.; Chowdhury, S. QuadBase: Genome-wide database of G4 DNA—Occurrence and conservation in human, chimpanzee, mouse and rat promoters and 146 microbes. Nucleic Acids Res. 2008, 36, D381–D385. [Google Scholar] [CrossRef] [PubMed]
- Verma, A.; Yadav, V.K.; Basundra, R.; Kumar, A.; Chowdhury, S. Evidence of genome-wide G4 DNA-mediated gene expression in human cancer cells. Nucleic Acids Res. 2009, 37, 4194–4204. [Google Scholar] [CrossRef] [PubMed]
- Postberg, J.; Tsytlonok, M.; Sparvoli, D.; Rhodes, D.; Lipps, H.J. A telomerase-associated RecQ protein-like helicase resolves telomeric G-quadruplex structures during replication. Gene 2012, 497, 147–154. [Google Scholar] [CrossRef] [PubMed]
- Rhodes, D.; Lipps, H.J. G-quadruplexes and their regulatory roles in biology. Nucleic Acids Res. 2015, 43, 8627–8637. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, A.K.; Sharma, S.; Chowdhury, S. Non-duplex G-Quadruplex Structures Emerge as Mediators of Epigenetic Modifications. Trends Genet. 2019, 35, 129–144. [Google Scholar] [CrossRef]
- Franceschin, M.; Rossetti, L.; D’Ambrosio, A.; Schirripa, S.; Bianco, A.; Ortaggi, G.; Savino, M.; Schultes, C.; Neidle, S. Natural and synthetic G-quadruplex interactive berberine derivatives. Bioorg. Med. Chem. Lett. 2006, 16, 1707–1711. [Google Scholar] [CrossRef]
- Bhadra, K.; Kumar, G.S. Interaction of berberine, palmatine, coralyne, and sanguinarine to quadruplex DNA: A comparative spectroscopic and calorimetric study. Biochim. Biophys. Acta Gen. Subj. 2011, 1810, 485–496. [Google Scholar] [CrossRef] [PubMed]
- Hussain, T.; Saha, D.; Purohit, G.; Kar, A.; Mukherjee, A.K.; Sharma, S.; Sengupta, S.; Dhapola, P.; Maji, B.; Vedagopuram, S.; et al. Transcription regulation of CDKN1A (p21/CIP1/WAF1) by TRF2 is epigenetically controlled through the REST repressor complex. Sci. Rep. 2017, 7, 11541. [Google Scholar] [CrossRef] [PubMed]
- Sadhukhan, R.; Chowdhury, P.; Ghosh, S.; Ghosh, U. Expression of Telomere-Associated Proteins is Interdependent to Stabilize Native Telomere Structure and Telomere Dysfunction by G-Quadruplex Ligand Causes TERRA Upregulation. Cell Biochem. Biophys. 2018, 76, 311–319. [Google Scholar] [CrossRef] [PubMed]
- Banno, K.; Kisu, I.; Yanokura, M.; Tsuji, K.; Masuda, K.; Ueki, A.; Kobayashi, Y.; Yamagami, W.; Nomura, H.; Tominaga, E.; et al. Epimutation and cancer: A new carcinogenic mechanism of Lynch syndrome. Int. J. Oncol. 2012, 41, 793–797. [Google Scholar] [CrossRef] [PubMed]
- Esteller, M. Cancer epigenomics: DNA methylomes and histone-modification maps. Nat. Rev. Genet. 2007, 8, 286. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.A.; Issa, J.-P.J.; Baylin, S. Targeting the cancer epigenome for therapy. Nat. Rev. Genet. 2016, 17, 630–641. [Google Scholar] [CrossRef] [PubMed]
- Saha, D.; Singh, A.; Hussain, T.; Srivastava, V.; Sengupta, S.; Kar, A.; Dhapola, P.; Dhople, V.; Ummanni, R.; Chowdhury, S. Epigenetic suppression of human telomerase (hTERT) is mediated by the metastasis suppressor NME2 in a G-quadruplex-dependent fashion. J. Biol. Chem. 2017, 292, 15205–15215. [Google Scholar] [CrossRef]
- Fukuhara, M.; Ma, Y.; Nagasawa, K.; Toyoshima, F. A G-quadruplex structure at the 5′ end of the H19 coding region regulates H19 transcription. Sci. Rep. 2017, 7, 45815. [Google Scholar] [CrossRef]
- Cogoi, S.; Ferino, A.; Miglietta, G.; Pedersen, E.B.; Xodo, L.E. The regulatory G4 motif of the Kirsten ras (KRAS) gene is sensitive to guanine oxidation: Implications on transcription. Nucleic Acids Res. 2018, 46, 661–676. [Google Scholar] [CrossRef]
- Lin, J.; Hou, J.; Xiang, H.; Yan, Y.; Gu, Y.; Tan, J.; Li, D.; Gu, L.; Ou, T.; Huang, Z. Stabilization of G-quadruplex DNA by C-5-methyl-cytosine in bcl-2 promoter: Implications for epigenetic regulation. Biochem. Biophys. Res. Commun. 2013, 433, 368–373. [Google Scholar] [CrossRef]
- Shin, Y.J.; Kumarasamy, V.; Camacho, D.; Sun, D. Involvement of G-quadruplex structures in regulation of human RET gene expression by small molecules in human medullary thyroid carcinoma TT cells. Oncogene 2015, 34, 1292. [Google Scholar] [CrossRef]
- Draht, M.X.G.; Smits, K.M.; Jooste, V.; Tournier, B.; Vervoort, M.; Ramaekers, C.; Chapusot, C.; Weijenberg, M.P.; van Engeland, M.; Melotte, V. Analysis of RET promoter CpG island methylation using methylation-specific PCR (MSP), pyrosequencing, and methylation-sensitive high-resolution melting (MS-HRM): Impact on stage II colon cancer patient outcome. Clin. Epigenet. 2016, 8, 44. [Google Scholar] [CrossRef] [PubMed]
- Gong, C.; Tao, G.; Yang, L.; Liu, J.; Liu, Q.; Li, W.; Zhuang, Z. Methylation of PARP-1 promoter involved in the regulation of nano-SiO 2-induced decrease of PARP-1 mRNA expression. Toxicol. Lett. 2012, 209, 264–269. [Google Scholar] [CrossRef] [PubMed]
- Sengar, A.; Jeya Vandana, J.; Chambers, V.S.; Di Antonio, M.; Winnerdy, F.R.; Balasubramanian, S.; Phan, A.T. Structure of a (3+1) hybrid G-quadruplex in the PARP1 promoter. Nucleic Acids Res. 2018. [Google Scholar] [CrossRef] [PubMed]
- Smith, S.S.; Kan, J.L.; Baker, D.J.; Kaplan, B.E.; Dembek, P. Recognition of unusual DNA structures by human DNA (cytosine-5)methyltransferase. J. Mol. Biol. 1991, 217, 39–51. [Google Scholar] [CrossRef]
- Baylin, S.B.; Jones, P.A. A decade of exploring the cancer epigenome-biological and translational implications. Nat. Rev. Cancer 2011, 11, 726. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Goodrich, K.J.; Gooding, A.R.; Naeem, H.; Archer, S.; Paucek, R.D.; Youmans, D.T.; Cech, T.R.; Davidovich, C. Targeting of Polycomb Repressive Complex 2 to RNA by Short Repeats of Consecutive Guanines. Mol. Cell 2017, 65, 1056–1067. [Google Scholar] [CrossRef]
- Zamiri, B.; Mirceta, M.; Bomsztyk, K.; Macgregor, R.B.; Pearson, C.E. Quadruplex formation by both G-rich and C-rich DNA strands of the C9orf72 (GGGGCC)8•(GGCCCC)8 repeat: Effect of CpG methylation. Nucleic Acids Res. 2015, 43, 10055–10064. [Google Scholar]
- Fry, M.; Loeb, L.A. The fragile X syndrome d(CGG)n nucleotide repeats form a stable tetrahelical structure. Proc. Natl. Acad. Sci. USA 1994, 91, 4950–4954. [Google Scholar] [CrossRef]
- Reed, J.C. Regulation of apoptosis by bcl-2 family proteins and its role in cancer and chemoresistance. Curr. Opin. Oncol. 1995, 7, 541–546. [Google Scholar] [CrossRef]
- Kim, N.W.; Piatyszek, M.A.; Prowse, K.R.; Harley, C.B.; West, M.D.; Ho, P.L.; Coviello, G.M.; Wright, W.E.; Weinrich, S.L.; Shay, J.W. Specific association of human telomerase activity with immortal cells and cancer. Science 1994, 266, 2011–2015. [Google Scholar] [CrossRef] [PubMed]
- Li, P.-T.; Wang, Z.-F.; Chu, I.-T.; Kuan, Y.-M.; Li, M.-H.; Huang, M.-C.; Chiang, P.-C.; Chang, T.-C.; Chen, C.-T. Expression of the human telomerase reverse transcriptase gene is modulated by quadruplex formation in its first exon due to DNA methylation. J. Biol. Chem. 2017, 292, 20859–20870. [Google Scholar] [CrossRef] [PubMed]
- Beckett, J.; Burns, J.; Broxson, C.; Tornaletti, S. Spontaneous DNA lesions modulate DNA structural transitions occurring at nuclease hypersensitive element III(1) of the human c-myc proto-oncogene. Biochemistry 2012, 51, 5257–5268. [Google Scholar] [CrossRef] [PubMed]
- Cheong, V.V.; Heddi, B.; Lech, C.J.; Phan, A.T. Xanthine and 8-oxoguanine in G-quadruplexes: Formation of a G·G·X·O tetrad. Nucleic Acids Res. 2015, 43, 10506–10514. [Google Scholar] [CrossRef] [PubMed]
- Fleming, A.M.; Ding, Y.; Burrows, C.J. Oxidative DNA damage is epigenetic by regulating gene transcription via base excision repair. Proc. Natl. Acad. Sci. USA 2017, 114, 2604–2609. [Google Scholar] [CrossRef] [PubMed]
- Fleming, A.M.; Zhu, J.; Ding, Y.; Burrows, C.J. 8-Oxo-7,8-dihydroguanine in the Context of a Gene Promoter G-Quadruplex Is an On-Off Switch for Transcription. ACS Chem. Biol. 2017, 12, 2417–2426. [Google Scholar] [CrossRef]
- Sarkies, P.; Reams, C.; Simpson, L.J.; Sale, J.E. Epigenetic Instability due to Defective Replication of Structured DNA. Mol. Cell 2010, 40, 703–713. [Google Scholar] [CrossRef]
- Sarkies, P.; Murat, P.; Phillips, L.G.; Patel, K.J.; Balasubramanian, S.; Sale, J.E. FANCJ coordinates two pathways that maintain epigenetic stability at G-quadruplex DNA. Nucleic Acids Res. 2012, 40, 1485–1498. [Google Scholar] [CrossRef]
- Schiavone, D.; Guilbaud, G.; Murat, P.; Papadopoulou, C.; Sarkies, P.; Prioleau, M.-N.; Balasubramanian, S.; Sale, J.E. Determinants of G quadruplex-induced epigenetic instability in REV1-deficient cells. EMBO J. 2014, 33, 2507–2520. [Google Scholar] [CrossRef]
- Bartke, T.; Vermeulen, M.; Xhemalce, B.; Robson, S.C.; Mann, M.; Kouzarides, T. Nucleosome-interacting proteins regulated by DNA and histone methylation. Cell 2010, 143, 470–484. [Google Scholar] [CrossRef]
- Borgel, J.; Tyl, M.; Schiller, K.; Pusztai, Z.; Dooley, C.M.; Deng, W.; Wooding, C.; White, R.J.; Warnecke, T.; Leonhardt, H.; et al. KDM2A integrates DNA and histone modification signals through a CXXC/PHD module and direct interaction with HP1. Nucleic Acids Res. 2017, 45, 1114–1129. [Google Scholar] [CrossRef] [PubMed]
- Thakur, R.K.; Kumar, P.; Halder, K.; Verma, A.; Kar, A.; Parent, J.-L.; Basundra, R.; Kumar, A.; Chowdhury, S. Metastases suppressor NM23-H2 interaction with G-quadruplex DNA within c-MYC promoter nuclease hypersensitive element induces c-MYC expression. Nucleic Acids Res. 2009, 37, 172–183. [Google Scholar] [CrossRef] [PubMed]
- Cree, S.L.; Fredericks, R.; Miller, A.; Pearce, F.G.; Filichev, V.; Fee, C.; Kennedy, M.A. DNA G-quadruplexes show strong interaction with DNA methyltransferases in vitro. FEBS Lett. 2016, 590, 2870–2883. [Google Scholar] [CrossRef] [PubMed]
- Halder, R.; Halder, K.; Sharma, P.; Garg, G.; Sengupta, S.; Chowdhury, S. Guanine quadruplex DNA structure restricts methylation of CpG dinucleotides genome-wide. Mol. Biosyst. 2010, 6, 2439–2447. [Google Scholar] [CrossRef] [PubMed]
- Agarwala, P.; Pandey, S.; Maiti, S. The tale of RNA G-quadruplex. Org. Biomol. Chem. 2015, 13, 5570–5585. [Google Scholar] [CrossRef] [PubMed]
- Song, J.; Perreault, J.-P.; Topisirovic, I.; Richard, S. RNA G-quadruplexes and their potential regulatory roles in translation. Translation 2016, 4, e1244031. [Google Scholar] [CrossRef] [PubMed]
- Fay, M.M.; Lyons, S.M.; Ivanov, P. RNA G-Quadruplexes in Biology: Principles and Molecular Mechanisms. J. Mol. Biol. 2017, 429, 2127–2147. [Google Scholar] [CrossRef] [PubMed]
- Cammas, A.; Millevoi, S. RNA G-quadruplexes: Emerging mechanisms in disease. Nucleic Acids Res. 2017, 45, 1584–1595. [Google Scholar] [CrossRef]
- Armas, P.; Calcaterra, N.B. G-quadruplex in animal development: Contribution to gene expression and genomic heterogeneity. Mech. Dev. 2018, 154, 64–72. [Google Scholar] [CrossRef]
- Rizvi, N.F.; Smith, G.F. RNA as a small molecule druggable target. Bioorg. Med. Chem. Lett. 2017, 27, 5083–5088. [Google Scholar] [CrossRef]
- Donlic, A.; Hargrove, A.E. Targeting RNA in mammalian systems with small molecules. Wiley Interdiscip. Rev. RNA 2018, 9, e1477. [Google Scholar] [CrossRef] [PubMed]
- Chan, K.L.; Peng, B.; Umar, M.I.; Chan, C.-Y.; Sahakyan, A.B.; Le, M.T.N.; Kwok, C.K. Structural analysis reveals the formation and role of RNA G-quadruplex structures in human mature microRNAs. Chem. Commun. 2018, 54, 10878–10881. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Kaminaga, K.; Komiyama, M. G-Quadruplex Formation by Human Telomeric Repeats-Containing RNA in Na+ Solution. J. Am. Chem. Soc. 2008, 130, 11179–11184. [Google Scholar] [CrossRef]
- Xu, Y.; Suzuki, Y.; Ito, K.; Komiyama, M. Telomeric repeat-containing RNA structure in living cells. Proc. Natl. Acad. Sci. USA 2010, 107, 14579–14584. [Google Scholar] [CrossRef] [PubMed]
- Hirschi, A.; Martin, W.J.; Luka, Z.; Loukachevitch, L.V.; Reiter, N.J. G-quadruplex RNA binding and recognition by the lysine-specific histone demethylase-1 enzyme. RNA 2016, 22, 1250–1260. [Google Scholar] [CrossRef] [PubMed]
- Zahler, A.M.; Williamson, J.R.; Cech, T.R.; Prescott, D.M. Inhibition of telomerase by G-quartet DNA structures. Nature 1991, 350, 718–720. [Google Scholar] [CrossRef]
- Benetti, R.; García-Cao, M.; Blasco, M.A. Telomere length regulates the epigenetic status of mammalian telomeres and subtelomeres. Nat. Genet. 2007, 39, 243–250. [Google Scholar] [CrossRef]
- Maguire, D.; Neytchev, O.; Talwar, D.; McMillan, D.; Shiels, P. Telomere Homeostasis: Interplay with Magnesium. Int. J. Mol. Sci. 2018, 19, 157. [Google Scholar] [CrossRef]
- Xu, Y. Chemistry in human telomere biology: Structure, function and targeting of telomere DNA/RNA. Chem. Soc. Rev. 2011, 40, 2719–2740. [Google Scholar] [CrossRef]
- Xu, Y.; Suzuki, Y.; Komiyama, M. Click chemistry for the identification of g-quadruplex structures: Discovery of a DNA-RNA g-quadruplex. Angew. Chem. Int. Ed. 2009, 48, 3281–3284. [Google Scholar] [CrossRef]
- Takahama, K.; Kino, K.; Arai, S.; Kurokawa, R.; Oyoshi, T. Identification of Ewing’s sarcoma protein as a G-quadruplex DNA- and RNA-binding protein. FEBS J. 2011, 278, 988–998. [Google Scholar] [CrossRef] [PubMed]
- Takahama, K.; Takada, A.; Tada, S.; Shimizu, M.; Sayama, K.; Kurokawa, R.; Oyoshi, T. Regulation of telomere length by G-quadruplex telomere DNA- and TERRA-binding protein TLS/FUS. Chem. Biol. 2013, 20, 341–350. [Google Scholar] [CrossRef] [PubMed]
- Takahama, K.; Miyawaki, A.; Shitara, T.; Mitsuya, K.; Morikawa, M.; Hagihara, M.; Kino, K.; Yamamoto, A.; Oyoshi, T. G-Quadruplex DNA- and RNA-Specific-Binding Proteins Engineered from the RGG Domain of TLS/FUS. ACS Chem. Biol. 2015, 10, 2564–2569. [Google Scholar] [CrossRef]
- Wong, L.H.; McGhie, J.D.; Sim, M.; Anderson, M.A.; Ahn, S.; Hannan, R.D.; George, A.J.; Morgan, K.A.; Mann, J.R.; Choo, K.H.A. ATRX interacts with H3.3 in maintaining telomere structural integrity in pluripotent embryonic stem cells. Genome Res. 2010. [Google Scholar] [CrossRef] [PubMed]
- Szyf, M. Epigenetics, DNA Methylation, and Chromatin Modifying Drugs. Annu. Rev. Pharmacol. Toxicol. 2009, 49, 243–263. [Google Scholar] [CrossRef] [PubMed]
- Nebbioso, A.; Carafa, V.; Benedetti, R.; Altucci, L. Trials with ‘epigenetic’ drugs: An update. Mol. Oncol. 2012, 6, 657–682. [Google Scholar] [CrossRef] [PubMed]
- Heerboth, S.; Lapinska, K.; Snyder, N.; Leary, M.; Rollinson, S.; Sarkar, S. Use of Epigenetic Drugs in Disease: An Overview. Genet. Epigenet. 2014, 6, GEG.S12270. [Google Scholar] [CrossRef] [PubMed]
- Han, H.; Hurley, L.H. G-quadruplex DNA: A potential target for anti-cancer drug design. Trends Pharmacol. Sci. 2000, 21, 136–142. [Google Scholar] [CrossRef]
- Ruggiero, E.; Richter, S.N. G-quadruplexes and G-quadruplex ligands: Targets and tools in antiviral therapy. Nucleic Acids Res. 2018, 46, 3270–3283. [Google Scholar] [CrossRef]
- Qing, Y.; Hu, H.; Liu, Y.; Feng, T.; Meng, W.; Jiang, L.; Sun, Y.; Yao, Y. Berberine induces apoptosis in human multiple myeloma cell line U266 through hypomethylation of p53 promoter: Berberine induces hypomethylation of p53 in U266. Cell Biol. Int. 2014, 38, 563–570. [Google Scholar] [CrossRef]
- Kalaiarasi, A.; Anusha, C.; Sankar, R.; Rajasekaran, S.; John Marshal, J.; Muthusamy, K.; Ravikumar, V. Plant Isoquinoline Alkaloid Berberine Exhibits Chromatin Remodeling by Modulation of Histone Deacetylase to Induce Growth Arrest and Apoptosis in the A549 Cell Line. J. Agric. Food Chem. 2016, 64, 9542–9550. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Liu, Y.; Xue, Y.; Hu, H.; Ye, J.; Li, X.; Lu, Z.; Meng, F.; Liang, S. Berberine acts as a putative epigenetic modulator by affecting the histone code. Toxicol. Vitr. 2016, 36, 10–17. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Liu, H.; Gong, X.-L.; Wu, L.-Y.; Wen, B. Effect of evodiamine and berberine on the interaction between DNMTs and target microRNAs during malignant transformation of the colon by TGF-β1. Oncol. Rep. 2017, 37, 1637–1645. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S.; Pradhan, S.K.; Kar, A.; Chowdhury, S.; Dasgupta, D. Molecular basis of recognition of quadruplexes human telomere and c-myc promoter by the putative anticancer agent sanguinarine. Biochim. Biophys. Acta Gen. Subj. 2013, 1830, 4189–4201. [Google Scholar] [CrossRef]
- Selvi, B.R.; Pradhan, S.K.; Shandilya, J.; Das, C.; Sailaja, B.S.; Shankar, G.N.; Gadad, S.S.; Reddy, A.; Dasgupta, D.; Kundu, T.K. Sanguinarine Interacts with Chromatin, Modulates Epigenetic Modifications, and Transcription in the Context of Chromatin. Chem. Biol. 2009, 16, 203–216. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Chang, X.; Song, X.; Chen, C.; Chen, H.; Lu, Z.; Gao, X.; Lu, D. Berberine reverses abnormal expression of L-type pyruvate kinase by DNA demethylation and histone acetylation in the livers of the non-alcoholic fatty disease rat. Int. J. Clin. Exp. Med. 2015, 8, 7535–7543. [Google Scholar] [PubMed]
- Zhang, L.; Miao, X.-J.; Wang, X.; Pan, H.-H.; Li, P.; Ren, H.; Jia, Y.-R.; Lu, C.; Wang, H.-B.; Yuan, L.; et al. Antiproliferation of berberine is mediated by epigenetic modification of constitutive androstane receptor (CAR) metabolic pathway in hepatoma cells. Sci. Rep. 2016, 6, 28116. [Google Scholar] [CrossRef]
- Minhas, G.S.; Pilch, D.S.; Kerrigan, J.E.; LaVoie, E.J.; Rice, J.E. Synthesis and G-quadruplex stabilizing properties of a series of oxazole-containing macrocycles. Bioorg. Med. Chem. Lett. 2006, 16, 3891–3895. [Google Scholar] [CrossRef] [PubMed]
- Tauchi, T.; Shin-ya, K.; Sashida, G.; Sumi, M.; Okabe, S.; Ohyashiki, J.H.; Ohyashiki, K. Telomerase inhibition with a novel G-quadruplex-interactive agent, telomestatin: In vitro and in vivo studies in acute leukemia. Oncogene 2006, 25, 5719–5725. [Google Scholar] [CrossRef]
- Qin, Y.; Fortin, J.S.; Tye, D.; Gleason-Guzman, M.; Brooks, T.A.; Hurley, L.H. Molecular cloning of the human platelet-derived growth factor receptor beta (PDGFR-beta) promoter and drug targeting of the G-quadruplex-forming region to repress PDGFR-beta expression. Biochemistry 2010, 49, 4208–4219. [Google Scholar] [CrossRef]
- Hewitt, K.J.; Shamis, Y.; Knight, E.; Smith, A.; Maione, A.; Alt-Holland, A.; Sheridan, S.D.; Haggarty, S.J.; Garlick, J.A. PDGFRβ expression and function in fibroblasts derived from pluripotent cells is linked to DNA demethylation. J. Cell Sci 2012, 125, 2276–2287. [Google Scholar] [CrossRef] [PubMed]
- Bay, D.H.; Busch, A.; Lisdat, F.; Iida, K.; Ikebukuro, K.; Nagasawa, K.; Karube, I.; Yoshida, W. Identification of G-quadruplex structures that possess transcriptional regulating functions in the Dele and Cdc6 CpG islands. BMC Mol. Biol. 2017, 18, 17. [Google Scholar] [CrossRef] [PubMed]
- Sparapani, S.; Haider, S.M.; Doria, F.; Gunaratnam, M.; Neidle, S. Rational design of acridine-based ligands with selectivity for human telomeric quadruplexes. J. Am. Chem. Soc. 2010, 132, 12263–12272. [Google Scholar] [CrossRef] [PubMed]
- Guo, Q.-L.; Su, H.-F.; Wang, N.; Liao, S.-R.; Lu, Y.-T.; Ou, T.-M.; Tan, J.-H.; Li, D.; Huang, Z.-S. Synthesis and evaluation of 7-substituted-5,6-dihydrobenzo[c]acridine derivatives as new c-KIT promoter G-quadruplex binding ligands. Eur. J. Med. Chem. 2017, 130, 458–471. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, J.; Pereira, E.; Marquevielle, J.; Campello, M.P.C.; Mergny, J.-L.; Paulo, A.; Salgado, G.F.; Queiroz, J.A.; Cruz, C. Fluorescent light-up acridine orange derivatives bind and stabilize KRAS-22RT G-quadruplex. Biochimie 2018, 144, 144–152. [Google Scholar] [CrossRef] [PubMed]
- Sun, D.; Liu, W.-J.; Guo, K.; Rusche, J.J.; Ebbinghaus, S.; Gokhale, V.; Hurley, L.H. The proximal promoter region of the human vascular endothelial growth factor gene has a G-quadruplex structure that can be targeted by G-quadruplex–interactive agents. Mol. Cancer Ther. 2008, 7, 880–889. [Google Scholar] [CrossRef] [PubMed]
- Cammas, A.; Dubrac, A.; Morel, B.; Lamaa, A.; Touriol, C.; Teulade-Fichou, M.-P.; Prats, H.; Millevoi, S. Stabilization of the G-quadruplex at the VEGF IRES represses cap-independent translation. RNA Biol. 2015, 12, 320–329. [Google Scholar] [CrossRef]
- Siddiqui-Jain, A.; Grand, C.L.; Bearss, D.J.; Hurley, L.H. Direct evidence for a G-quadruplex in a promoter region and its targeting with a small molecule to repress c-MYC transcription. Proc. Natl. Acad. Sci. USA 2002, 99, 11593–11598. [Google Scholar] [CrossRef]
- Pérez-Arnaiz, C.; Busto, N.; Santolaya, J.; Leal, J.M.; Barone, G.; García, B. Kinetic evidence for interaction of TMPyP4 with two different G-quadruplex conformations of human telomeric DNA. Biochim. Biophys. Acta. Gen. Subj. 2018, 1862, 522–531. [Google Scholar] [CrossRef]
- Alniss, H.; Zamiri, B.; Khalaj, M.; Pearson, C.E.; Macgregor, R.B. Thermodynamic and spectroscopic investigations of TMPyP4 association with guanine- and cytosine-rich DNA and RNA repeats of C9orf72. Biochem. Biophys. Res. Commun. 2018, 495, 2410–2417. [Google Scholar] [CrossRef]
- Zhao, Y.; Uhler, J.P. Identification of a G-quadruplex forming sequence in the promoter of UCP1. Acta Biochim. Biophys. Sin. 2018, 50, 718–722. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Tan, W.; Chen, H.; Zhou, J.; Xu, M.; Yuan, G. Up- and downregulation of mature miR-1587 function by modulating its G-quadruplex structure and using small molecules. Int. J. Biol. Macromol. 2019, 121, 127–134. [Google Scholar] [CrossRef] [PubMed]
- Shan, C.; Yan, J.-W.; Wang, Y.-Q.; Che, T.; Huang, Z.-L.; Chen, A.-C.; Yao, P.-F.; Tan, J.-H.; Li, D.; Ou, T.-M.; et al. Design, Synthesis, and Evaluation of Isaindigotone Derivatives to Downregulate c-myc Transcription via Disrupting the Interaction of NM23-H2 with G-Quadruplex. J. Med. Chem. 2017, 60, 1292–1308. [Google Scholar] [CrossRef] [PubMed]
- Verma, S.P.; Das, P. Novel splicing in IGFN1 intron 15 and role of stable G-quadruplex in the regulation of splicing in renal cell carcinoma. PLoS ONE 2018, 13, e0205660. [Google Scholar] [CrossRef] [PubMed]
- Guilbaud, G.; Murat, P.; Recolin, B.; Campbell, B.C.; Maiter, A.; Sale, J.E.; Balasubramanian, S. Local epigenetic reprograming induced by G-quadruplex ligands. Nat. Chem. 2017, 9, 1110–1117. [Google Scholar] [CrossRef] [PubMed]
- Palumbo, S.L.; Ebbinghaus, S.W.; Hurley, L.H. Formation of a unique end-to-end stacked pair of G-quadruplexes in the hTERT core promoter with implications for inhibition of telomerase by G-quadruplex-interactive ligands. J. Am. Chem. Soc. 2009, 131, 10878–10891. [Google Scholar] [CrossRef] [PubMed]
- Lüpertz, R.; Wätjen, W.; Kahl, R.; Chovolou, Y. Dose- and time-dependent effects of doxorubicin on cytotoxicity, cell cycle and apoptotic cell death in human colon cancer cells. Toxicology 2010, 271, 115–121. [Google Scholar] [CrossRef]
- Pennarun, G.; Granotier, C.; Gauthier, L.R.; Gomez, D.; Hoffschir, F.; Mandine, E.; Riou, J.F.; Mergny, J.L.; Mailliet, P.; Boussin, F.D. Apoptosis related to telomere instability and cell cycle alterations in human glioma cells treated by new highly selective G-quadruplex ligands. Oncogene 2005, 24, 2917. [Google Scholar] [CrossRef]
- Ruthrotha, B.; Batta, K.; Kishore, A.H.; Mantelingu, K.; Varier, R.A.; Balasubramanyam, K.; Kalyan Pradhan, S.; Dasgupta, D.; Sriram, S.; Agrawal, S.; et al. Identification of a Novel Inhibitor of Coactivator-associated Arginine Methyltransferase 1 (CARM1)-mediated Methylation of Histone H3 Arg-17. J. Biol. Chem. 2009, 285, 7143–7152. [Google Scholar]
- Berletch, J.B.; Liu, C.; Love, W.K.; Andrews, L.G.; Katiyar, S.K.; Tollefsbol, T.O. Epigenetic and genetic mechanisms contribute to telomerase inhibition by EGCG. J. Cell. Biochem. 2008, 103, 509–519. [Google Scholar] [CrossRef]
- Liu, H.L.; Chen, Y.; Cui, G.H.; Zhou, J.F. Curcumin, a potent anti-tumor reagent, is a novel histone deacetylase inhibitor regulating B-NHL cell line Raji proliferation. Acta Pharmacol. Sin. 2005, 26, 603. [Google Scholar] [CrossRef] [PubMed]
- Majid, S.; Kikuno, N.; Nelles, J.; Noonan, E.; Tanaka, Y.; Kawamoto, K.; Hirata, H.; Li, L.C.; Zhao, H.; Okino, S.T.; et al. Genistein Induces the p21WAF1/CIP1 and p16INK4a Tumor Suppressor Genes in Prostate Cancer Cells by Epigenetic Mechanisms Involving Active Chromatin Modification. Cancer Res. 2008, 68, 2736–2744. [Google Scholar] [CrossRef] [PubMed]
- Kala, R.; Shah, H.N.; Martin, S.L.; Tollefsbol, T.O. Epigenetic-based combinatorial resveratrol and pterostilbene alters DNA damage response by affecting SIRT1 and DNMT enzyme expression, including SIRT1-dependent γ-H2AX and telomerase regulation in triple-negative breast cancer. BMC Cancer 2015, 15, 672. [Google Scholar] [CrossRef]
- Myzak, M.C.; Dashwood, W.M.; Orner, G.A.; Ho, E.; Dashwood, R.H. Sulforaphane inhibits histone deacetylase in vivo and suppresses tumorigenesis in Apcmin mice. FASEB J. 2006, 20, 506–508. [Google Scholar] [CrossRef]
- Mikutis, G.; Karaköse, H.; Jaiswal, R.; LeGresley, A.; Islam, T.; Fernandez-Lahore, M.; Kuhnert, N. Phenolic promiscuity in the cell nucleus – epigallocatechingallate (EGCG) and theaflavin-3,3′-digallate from green and black tea bind to model cell nuclear structures including histone proteins, double stranded DNA and telomeric quadruplex DNA. Food Funct. 2013, 4, 328–337. [Google Scholar] [CrossRef] [PubMed]
- Platella, C.; Guida, S.; Bonmassar, L.; Aquino, A.; Bonmassar, E.; Ravagnan, G.; Montesarchio, D.; Roviello, G.N.; Musumeci, D.; Fuggetta, M.P. Antitumour activity of resveratrol on human melanoma cells: A possible mechanism related to its interaction with malignant cell telomerase. Biochim. Biophys. Acta Gen. Subj. 2017, 1861, 2843–2851. [Google Scholar] [CrossRef] [PubMed]
- Pattanayak, R.; Basak, P.; Sen, S.; Bhattacharyya, M. Interaction of KRAS G-quadruplex with natural polyphenols: A spectroscopic analysis with molecular modeling. Int. J. Biol. Macromol. 2016, 89, 228–237. [Google Scholar] [CrossRef]
- François, M.; Leifert, W.R.; Tellam, R.; Fenech, M.F. Folate deficiency and DNA-methyltransferase inhibition modulate G-quadruplex frequency. Mutagenesis 2016, 31, 409–416. [Google Scholar] [CrossRef]
- Dkhil, M.A.; Metwaly, M.S.; Al-Quraishy, S.; Sherif, N.E.; Delic, D.; Al Omar, S.Y.; Wunderlich, F. Anti-Eimeria activity of berberine and identification of associated gene expression changes in the mouse jejunum infected with Eimeria papillata. Parasitol. Res. 2015, 114, 1581–1593. [Google Scholar] [CrossRef]
- Cheong, V.V.; Lech, C.J.; Heddi, B.; Phan, A.T. Inverting the G-Tetrad Polarity of a G-Quadruplex by Using Xanthine and 8-Oxoguanine. Angew. Chem. Int. Ed. Engl. 2016, 55, 160–163. [Google Scholar] [CrossRef]
- Fleming, A.M.; Burrows, C.J. 8-Oxo-7,8-dihydroguanine, friend and foe: Epigenetic-like regulator versus initiator of mutagenesis. DNA Repair 2017, 56, 75–83. [Google Scholar] [CrossRef] [PubMed]
- Erard, M.S.; Belenguer, P.; Caizergues-Ferrer, M.; Pantaloni, A.; Amalric, F. A major nucleolar protein, nucleolin, induces chromatin decondensation by binding to histone {H}1. Eur. J. Biochem. 1988, 175, 525–530. [Google Scholar] [CrossRef] [PubMed]
- Dapić, V.; Abdomerović, V.; Marrington, R.; Peberdy, J.; Rodger, A.; Trent, J.O.; Bates, P.J. Biophysical and biological properties of quadruplex oligodeoxyribonucleotides. Nucleic Acids Res. 2003, 31, 2097–2107. [Google Scholar] [CrossRef] [PubMed]
- González, V.; Guo, K.; Hurley, L.; Sun, D. Identification and characterization of nucleolin as a c-myc G-quadruplex-binding protein. J. Biol. Chem. 2009, 284, 23622–23635. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, A.K.; Sharma, S.; Sengupta, S.; Saha, D.; Kumar, P.; Hussain, T.; Srivastava, V.; Roy, S.D.; Shay, J.W.; Chowdhury, S. Telomere length-dependent transcription and epigenetic modifications in promoters remote from telomere ends. PLoS Genet. 2018, 14, e1007782. [Google Scholar] [CrossRef] [PubMed]
- Vlasenok, M.; Levchenko, O.; Basmanov, D.; Klinov, D.; Varizhuk, A.; Pozmogova, G. Data set on G4 DNA interactions with human proteins. Data Brief 2018, 15, 348–359. [Google Scholar] [CrossRef] [PubMed]
- Bharti, S.K.; Awate, S.; Banerjee, T.; Brosh, R.M. Getting ready for the dance: FANCJ Irons out DNA wrinkles. Genes 2016, 7, 31. [Google Scholar] [CrossRef]
- Chatterjee, S.; Zagelbaum, J.; Savitsky, P.; Sturzenegger, A.; Huttner, D.; Janscak, P.; Hickson, I.D.; Gileadi, O.; Rothenberg, E. Mechanistic insight into the interaction of BLM helicase with intra-strand G-quadruplex structures. Nat. Commun. 2014, 5, 5556. [Google Scholar] [CrossRef]
- Mohaghegh, P. The Bloom’s and Werner’s syndrome proteins are DNA structure-specific helicases. Nucleic Acids Res. 2001, 29, 2843–2849. [Google Scholar] [CrossRef]
- Hudson, J.S.; Ding, L.; Le, V.; Lewis, E.; Graves, D. Recognition and Binding of Human Telomeric G-Quadruplex DNA by Unfolding Protein 1. Biochemistry 2014, 53, 3347–3356. [Google Scholar] [CrossRef]
- Liu, X.; Ishizuka, T.; Bao, H.L.; Wada, K.; Takeda, Y.; Iida, K.; Nagasawa, K.; Yang, D.; Xu, Y. Structure-Dependent Binding of hnRNPA1 to Telomere RNA. J. Am. Chem. Soc. 2017, 139, 7533–7539. [Google Scholar] [CrossRef] [PubMed]
- Thandapani, P.; O’Connor, T.R.; Bailey, T.L.; Richard, S. Defining the RGG/RG Motif. Mol. Cell 2013, 50, 613–623. [Google Scholar] [CrossRef] [PubMed]
- McRae, E.K.S.; Booy, E.P.; Padilla-Meier, G.P.; McKenna, S.A. On Characterizing the Interactions between Proteins and Guanine Quadruplex Structures of Nucleic Acids. J. Nucleic Acids 2017, 2017, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Brázda, V.; Cerveň, J.; Bartas, M.; Mikysková, N.; Coufal, J.; Pečinka, P. The amino acid composition of quadruplex binding proteins reveals a shared motif and predicts new potential quadruplex interactors. Molecules 2018, 23, 2341. [Google Scholar] [CrossRef] [PubMed]
- Falabella, M.; Fernandez, R.J.; Johnson, B.; Kaufman, B.A. Potential roles for G-quadruplexes in mitochondria. Curr. Med. Chem. 2018, 25. [Google Scholar] [CrossRef] [PubMed]
- Falabella, M.; Kolesar, J.E.; Xiang, I.M.; Wang, T.; Horne, W.; Wallace, C.; Sun, L.; Taguchi, Y.; Wang, C.; Turek-Herman, J.; et al. G-quadruplex dynamics contribute to epigenetic regulation of mitochondrial function. bioRxiv 2018. [Google Scholar] [CrossRef]
- Jun Chung, W.; Heddi, B.; Schmitt, E.; Wai Lim, K.; Mechulam, Y.; Phan, A.T. Structure of a left-handed DNA G-quadruplex. Proc. Natl. Acad. Sci. USA 2015. [Google Scholar] [CrossRef]
- Fu, B.; Huang, J.; Chen, Y.; Wang, Y.; Xue, T.; Xu, G.; Wang, S.; Zhou, X. Right-handed and left-handed G-quadruplexes have the same DNA sequence: Distinct conformations induced by an organic small molecule and potassium. Chem. Commun. 2016, 52, 10052–10055. [Google Scholar] [CrossRef]
- Varizhuk, A.; Ilyinsky, N.; Smirnov, I.; Pozmogova, G. G4 Aptamers: Trends in Structural Design. J. Clin. Microbiol. 2015, 16, 1321–1329. [Google Scholar]
- Liu, J.; Wei, T.; Zhao, J.; Huang, Y.; Deng, H.; Kumar, A.; Wang, C.; Liang, Z.; Ma, X.; Liang, X.J. Multifunctional aptamer-based nanoparticles for targeted drug delivery to circumvent cancer resistance. Biomaterials 2016, 91, 44–56. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).