Characterization of Nine Compounds Isolated from the Acid Hydrolysate of Lonicera fulvotomentosa Hsu et S. C. Cheng and Evaluation of Their In Vitro Activity towards HIV Protease
Abstract
1. Introduction
2. Results and Discussion
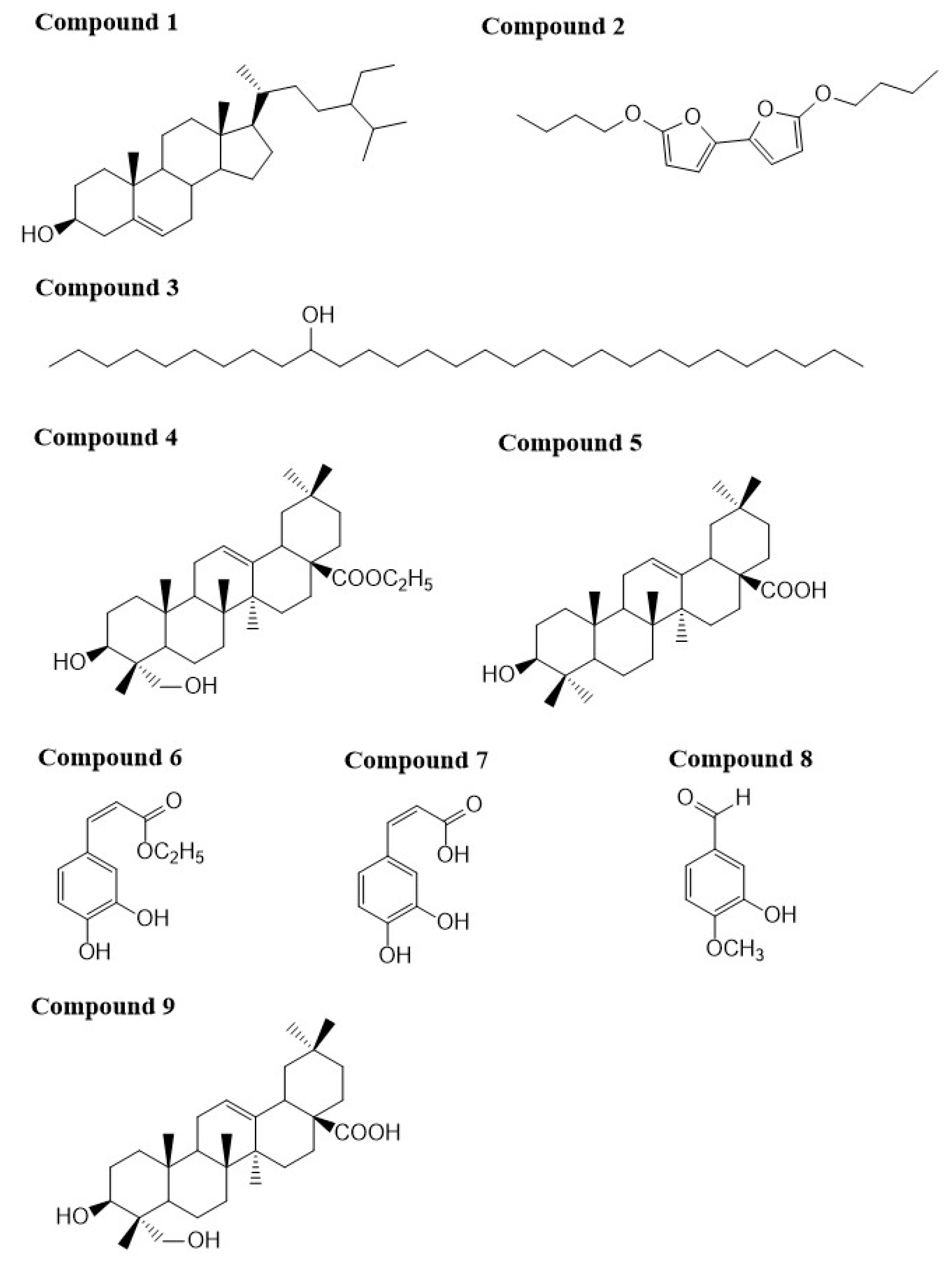
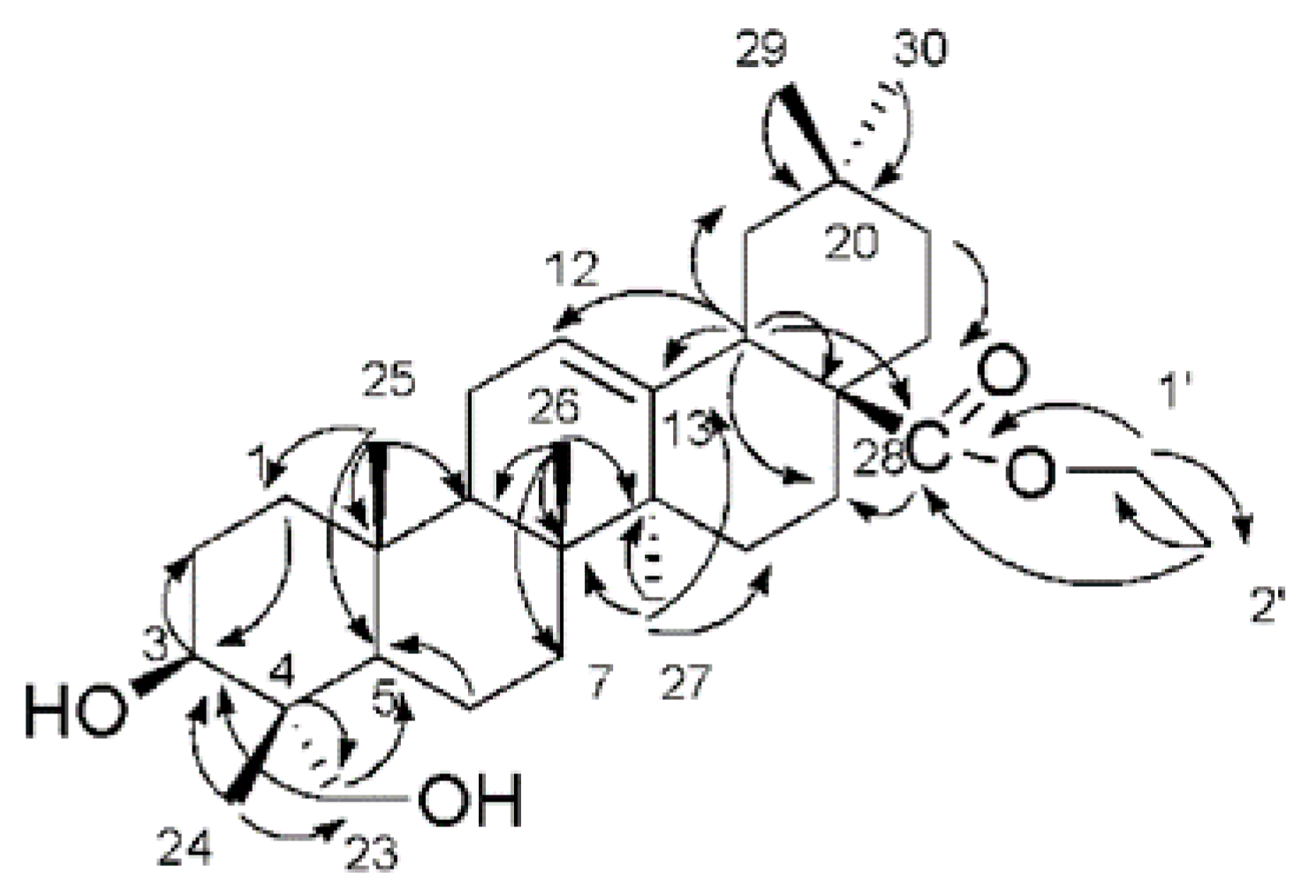
2.1. Characterization of the Compounds
2.2. Inhibitory Activity against HIV Protease
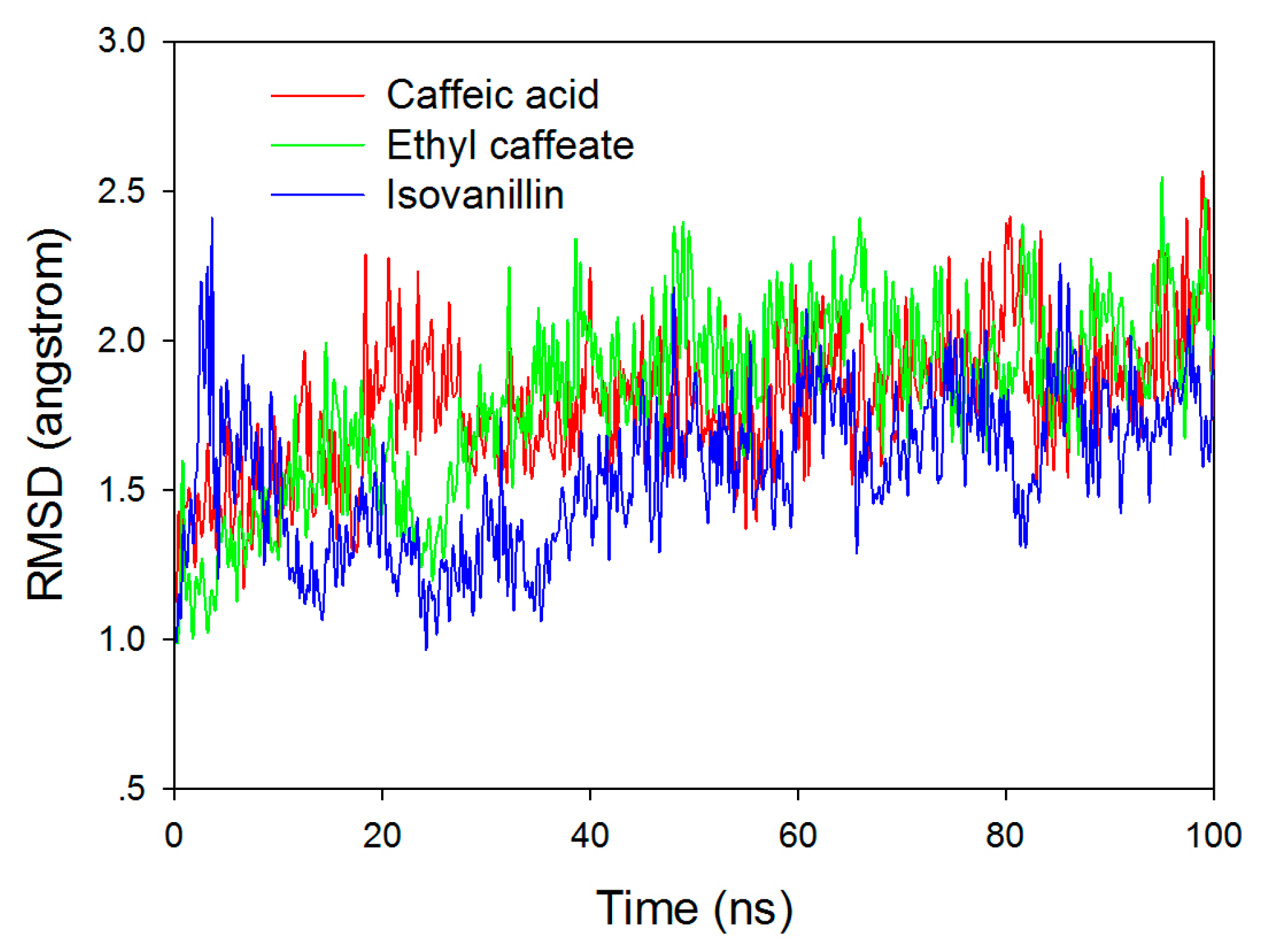
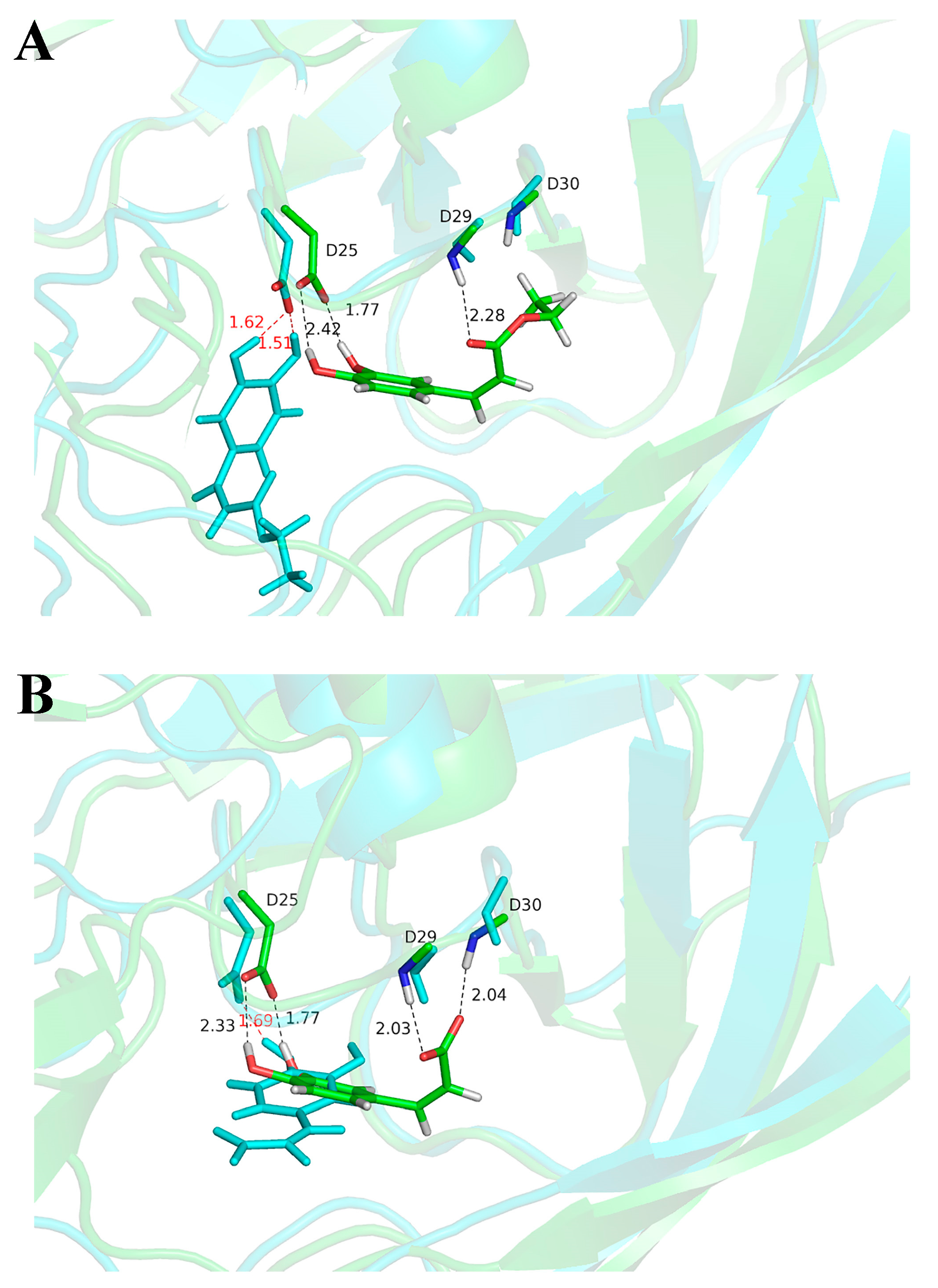
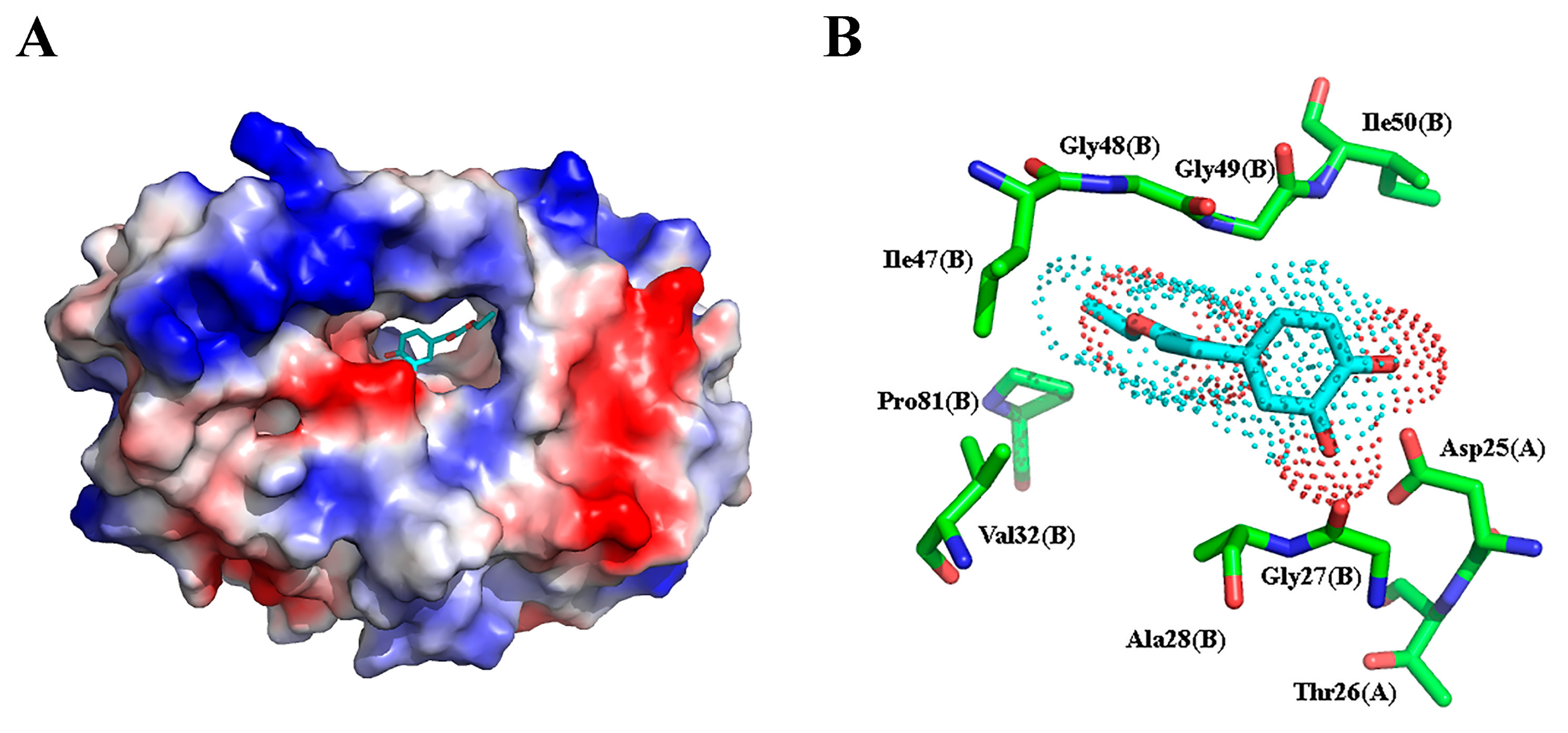
2.3. Molecular Docking and Molecular Dynamic Simulation
3. Materials and Methods
3.1. Materials
3.2. Extraction and Isolation
3.3. Compound Characterization
3.4. Anti-HIV Protease In Vitro Assay
3.5. Molecular Docking
3.6. Initial Structure Preparation and Molecular Dynamic Simulation
3.7. Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Chinese Pharmacopeia Commission. Pharmacopoeia of the People’s Republic of China; Chemistry Industry Publishing House: Beijing, China, 2015; p. 221. [Google Scholar]
- Available online: http://www.zysj.com.cn/lilunshuji/bencaoqiuzhen/619-11-3.html#hi-108293 (accessed on 9 December 2019).
- Dung, N.T.; Bajpai, V.K.; Yoon, J.I.; Kang, S.C.H. Phenolic contents, antioxidant and tyrosinase inhibitory activities of Lonicera japonica Thunb. J. Food Biochem. 2011, 35, 148–160. [Google Scholar] [CrossRef]
- Shang, X.; Pan, H.; Li, M.; Miao, X.; Ding, H. Lonicera japonicaThunb.: Ethnopharmacology, phytochemistry and pharmacology of an important traditional Chinese medicine. J. Ethnopharmacol. 2011, 138, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Editorial Board of Dictionary of Traditional Chinese Medicine Dictionary of Traditional Chinese Medicine; Shanghai Science and Technology Press: Shanghai, China, 1977; p. 1403.
- Xu, Y.; Oliverson, B.G.; Simmons, D.L. Trifunctional inhibition of COX-2 by extracts of Lonicera japonica: Direct inhibition, transcriptional and post-transcriptional down regulation. J. Ethnopharmacol. 2007, 111, 667–670. [Google Scholar] [CrossRef]
- Xiong, J.; Li, S.; Wang, W.; Hong, Y.; Tang, K.; Luo, Q. Screening and identification of the antibacterial bioactive compounds from Lonicera japonica Thunb. leaves. Food Chem. 2013, 138, 327–333. [Google Scholar] [CrossRef] [PubMed]
- Ko, H.; Wei, B.; Chiou, W. The effect of medicinal plants used in Chinese folk medicine on RANTES secretion by virus-infected human epithelial cells. J. Ethnopharmacol. 2006, 107, 205–210. [Google Scholar] [CrossRef] [PubMed]
- Peng, L.Y.; Mei, S.X.; Jiang, B.; Zhou, H.; Sun, H.D. Constituents from Lonicera japonica. Fitoterapia 2000, 71, 713–715. [Google Scholar] [CrossRef]
- Lin, L.M.; Zhang, X.G.; Zhu, J.J.; Gao, H.M.; Wang, Z.M.; Wang, W.H. Two new triterpenoid saponins from the flowers and buds of Lonicera japonica. J. Asian Nat. Prod. Res. 2008, 10, 925–929. [Google Scholar] [CrossRef]
- Zheng, Z.F.; Zhang, Q.J.; Chen, R.Y.; Yu, D.Q. Four new N-contained iridoid glycosides from flower buds of Lonicera japonica. J. Asian Nat. Prod. Res. 2012, 14, 729–737. [Google Scholar] [CrossRef]
- Yu, Y.; Jiang, Z.B.; Song, W.X.; Yang, Y.C.; Li, Y.H.; Jiang, J.D.; Shi, J.G. Glucosylated caffeoyl quinicacid derivatives from the flower buds of Lonicera japonica. Acta Pharm. Sin. B. 2015, 5, 210–214. [Google Scholar] [CrossRef]
- Kong, D.; Li, Y.; Bai, M.; Deng, Y.L.; Liang, G.X.; Wu, H. A comparative study of the dynamic accumulation of polyphenol components and the changes in their antioxidant activities in diploidand tetraploid Lonicera japonica. Plant Physiol. Biochem. 2017, 112, 87–96. [Google Scholar] [CrossRef]
- Kucharska, A.Z.; Sokół-Łętowska, A.; Oszmiański, J.; Piórecki, N.; Fecka, I. Iridoids, phenolic compounds and antioxidant activity of edible Honeysuckle Berries (Lonicera caerulea var. Kamtschatica Sevast.). Molecules 2017, 22, 405. [Google Scholar] [CrossRef] [PubMed]
- Robinson, W.E., Jr.; Cordeiro, M.; Abdel-Malek, S.; Jia, Q.; Chow, S.A.; Reinecke, M.G.; Mitchell, W.M. Dicaffeoylquinic acid inhibitors of human immunodeficiency virus integrase: Inhibition of the core catalytic domain of human immunodeficiency virus integrase. Mol. Pharmacol. 1996, 50, 846–855. [Google Scholar] [PubMed]
- Zhu, K.; Cordeiro, M.L.; Atienza, J.; Robinson, W.E., Jr.; Chow, S.A. Irreversible inhibition of human immunodeficiency virus type 1 integrase by dicaffeoylquinic acids. J. Virol. 1999, 73, 3309–3316. [Google Scholar] [PubMed]
- Crosby, D.C.; Lei, X.; Gibbs, C.G.; McDougall, B.R.; Robinson, W.E.; Reinecke, M.G. Design, synthesis, and biological evaluation of novel hybrid dicaffeoyltartaric/diketo acid and tetrazole-substituted L-chicoric acid analogue inhibitors of human immunodeficiency virus type 1 integrase. J. Med. Chem. 2010, 53, 8161–8175. [Google Scholar] [CrossRef] [PubMed]
- De Clercq, E. Current lead natural products for the chemotherapy of human immunodeficiency virus (HIV) infection. Med. Res. Rev. 2000, 20, 323–349. [Google Scholar] [CrossRef]
- Yang, B.; Meng, Z.Y.; Dong, J.X.; Yan, L.P.; Zou, L.B.; Tang, Z.M.; Dou, G.F. Metabolic profile of 1,5-dicaffeoylquinic acid in rats, an in vivo andin vitro study. Drug Metab. Dispos. 2005, 33, 930–936. [Google Scholar] [CrossRef]
- Choi, E.; Mallareddy, J.R.; Lu, D.; Kolluru, S. Recent advances in the discovery of small-molecule inhibitors of HIV-1 integrase. Future Sci. OA. 2018, 4, FSO338. [Google Scholar] [CrossRef]
- Marchand, C.; Maddali, K.; Métifiot, M.; Pommier, Y. HIV-1 IN inhibitors: 2010 update and perspectives. Curr. Top. Med. Chem. 2009, 9, 1016–1037. [Google Scholar] [CrossRef]
- Ding, Y.; Cao, Z.Y.; Cao, L.; Ding, G.; Wang, Z.Z.; Xiao, W. Antiviral activity of chlorogenic acid against influenza A (H1N1/H3N2) virus and its inhibition of neuraminidase. Sci. Rep. 2017, 7, 45723. [Google Scholar] [CrossRef]
- Meva’a, L.M.; Songue, J.L.; Wansi, J.D.; Waffo, A.F.K.; Dongo, E.; Mpondo, T.N.; Sewald, N. Acridone alkaloids and coumarins from the stem bark of Citropsis articulate (Rutaceae). Zeitschriftfür Naturforschung B. 2010, 65, 525–527. [Google Scholar] [CrossRef]
- Liu, J.; Xu, J.; Zhao, X.J.; Gao, W.Y.; Zhang, S.Z.; Guo, Y.Q. A new heterocyclic compound from Cyathula offifinalis Kuan. Chin. Chem. Lett. 2010, 21, 70–72. [Google Scholar] [CrossRef]
- Ishrat, N.; Saifullah; Khan, M.R. Nematicidal activity of nonacosane-10-ol and 23a-homostigmast-5-en-3β-ol isolated from the roots of Fumaria parviflora (Fumariaceae). J. Agric. Food Chem. 2013, 61, 5689–5695. [Google Scholar]
- Tori, K.; Seo, S.; Shimaoka, A.; Tomita, Y. Carbon-13C NMR spectra of olean-12-enes. Full signal assignments including quaternary carbon signals assigned by use of indirect 13C, 1H spin couplings. Tetrahedron Lett. 1974, 15, 4227–4230. [Google Scholar] [CrossRef]
- Li, Y.; Zhang, C.Y.; Wang, G.H.; Zeng, D.Q.; Guo, Z.J.; Zhou, X.H. Chemical constituents from Elephantopus tomentosus. Zhongguo Zhong Yao Za Zhi 2013, 38, 1751–1756. [Google Scholar]
- Jeong, C.H.; Jeong, H.R.; Choi, G.N.; Kim, D.O.; Lee, U.K.; Heo, H.J. Neuroprotective and anti-oxidant effects of caffeic acid isolated from Erigeron annuus leaf. Chin. Med. 2011, 6, 25–34. [Google Scholar] [CrossRef]
- Xu, J.J.; Tan, N.H.; Zeng, G.Z.; Han, H.J.; Huang, H.Q.; Ji, C.J.; Zhu, M.J.; Zhang, Y.M. Studies on chemical constituents in fruit of Alpinia oxyphylla. Zhongguo Zhong Yao Za Zhi 2009, 34, 990–993. [Google Scholar]
- He, W.D.; Van Puyvelde, L.; Maes, L.; Bosselaers, J.; De Kimpe, N. Antitrichoonas in vitro activity of Cussonia holstii Engl. Nat. Prod. Res. 2003, 17, 127–133. [Google Scholar] [CrossRef]
- Khalika, A.; Miyaseb, T.; El-Ashaalc, H.A.; Melekc, F.R. Triterpenoid saponins from Fagonia cretica. Phytochemistry 2000, 54, 853–859. [Google Scholar] [CrossRef]
- Wei, Y.; Ma, C.M.; Jiang, T.B.; Du, J.; Zhou, X.; Liu, G.Q.; Hattori, M. Synthesis of piscidinol A derivatives and their ability to inhibit HIV-1 protease. J. Asian Nat. Prod. Res. 2015, 17, 1079–1090. [Google Scholar] [CrossRef]
- Zhang, X.; Neamati, N.; Lee, Y.K.; Orr, A.; Brown, R.D.; Whitaker, N.; Pommier, Y.; Burke, T.R., Jr. Arylisothiocyanate-containing esters of caffeic acid designed as affinity ligands for HIV-1 integrase. Bioorg. Med. Chem. 2001, 9, 1649–1657. [Google Scholar] [CrossRef]
- Carrieri, M.P.; Protopopescu, C.; Marcellin, F.; Rosellini, S.; Wittkop, L.; Esterle, L.; Zucman, D.; Raffi, F.; Rosenthal, E.; Poizot-Martin, I.; et al. ANRS CO13 HEPAVIH Study Group. Protective effect of coffee consumption on all-cause mortality of French HIV-HCV co-infected patients. J. Hepatol. 2017, 67, 1157–1167. [Google Scholar] [CrossRef] [PubMed]
- Lam, P.Y.; Ru, Y.; Jadhav, P.K.; Aldrich, P.E.; DeLucca, G.V.; Eyermann, C.J.; Hodge, C.N. Cyclic HIV protease inhibitors: Synthesis, conformational analysis, P2/P2′ structure-activity relationship, and molecular recognition of cyclic ureas. J. Med. Chem. 1996, 39, 3514–3525. [Google Scholar] [CrossRef] [PubMed]
- Pokorná, J.; Machala, L.; Rezáčová, P.; Konvalinka, J. Current and novel inhibitors of HIV protease. Viruses 2009, 1, 1209–1239. [Google Scholar] [CrossRef] [PubMed]
- Rostkowski, M.; Olsson, M.H.M.; Sondergaard, C.R.; Jensen, J.H. Graphical analysis of pH-dependent properties of proteins predicted using PROPKA. BMC Struct. Biol. 2011, 11, 6. [Google Scholar] [CrossRef]
- Hehre, W.J.; Radom, L.; Schleyer, P.; Pople, J. Ab Inition Molecular Orbital Theory; John Wiley & Sons: New York, NY, USA, 1986. [Google Scholar]
- Wang, J.M.; Cieplak, P.; Kollman, P.A. How well does a restrained electrostatic potential (RESP) model perform in calculating conformational energies of organic and biological molecules? J. Comput. Chem. 2000, 21, 1049–1074. [Google Scholar] [CrossRef]
- Cornell, W.D.; Cieplak, P.; Bayly, C.I.; Gould, I.R.; Merz, K.M.; Ferguson, D.M.; Spellmeyer, D.C.; Fox, T.; Caldwell, J.W.; Kollman, P.A. A second generation force field for the simulation of proteins, nucleic acids, and organic molecules. J. Am. Chem. Soc. 1995, 117, 5179–5197. [Google Scholar] [CrossRef]
- Hornak, V.; Abel, R.; Okur, A.; Strockbine, B.; Roitberg, A.; Simmerling, C. Comparison of multiple amber force fields and development of improved protein backbone parameters. Proteins 2006, 65, 712–725. [Google Scholar] [CrossRef]
- Berendsen, H.J.C.; Postma, J.P.M.; Vangunsteren, W.F.; Dinola, A.; Haak, J.R. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 1984, 81, 3684–3690. [Google Scholar] [CrossRef]
- Darden, T.; York, D.; Pedersen, L. Particle mesh Ewald - An N∙log(N) method for Ewald sums in large systems. J. Chem. Phys. 1993, 98, 10089–10092. [Google Scholar] [CrossRef]
- Essmann, U.; Perera, L.; Berkowitz, M.L.; Darden, T.; Lee, H.; Pedersen, L.G. A smooth particle mesh Ewald method. J. Chem. Phys. 1995, 103, 8577–8593. [Google Scholar] [CrossRef]
- Case, D.A.; Betz, R.M.; Botello-Smith, W.; Cerutti, D.S.; Cheatham, T.E., III; Darden, I.T.A.; Duke, R.E.; Giese, T.J.; Gohlke, H.; Goetz, A.W.; et al. AMBER 2016; University of California: San Francisco, CA, USA, 2016. [Google Scholar]
| Position | δH | δC | Position | δH | δC |
|---|---|---|---|---|---|
| 1 | 0.90 (m) 1.44 (m) a | 38.1 | 17 | 46.4 | |
| 2 | 1.16 (m) b 1.98 (m) c | 26.6 | 18 | 2.86 (d, 8.0) | 41.7 |
| 3 | 3.63 (dd, 8.0, 16.0) | 76.8 | 19 | a, 1.12 (m) d b, 1.67 (m) | 46.1 |
| 4 | 41.2 | 20 | 30.6 | ||
| 5 | 1.13 (m) d | 49.7 | 21 | a, 1.08 (m) b, 1.33 (m) e | 33.1 |
| 6 | a, 1.30 (m) e b, 1.40 (m) | 18.3 | 22 | a, 1.73 (m) b, 1.84 (m) | 32.4 |
| 7 | a, 1.26 (m) b, 1.44 (m) a | 32.4 | 23 | a, 3.42 (d, 8.0) b, 3.72 (d, 8.0) | 72.1 |
| 8 | 39.3 | 24 | 0.89 (s) | 11.0 | |
| 9 | 1.55 (m) | 47.7 | 25 | 0.92 (s) | 15.8 |
| 10 | 36.9 | 26 | 0.74 (s) | 16.9 | |
| 11 | a, 1.35 (m) b, 1.95 (m) c | 25.8 | 27 | 1.13 (s) d | 25.8 |
| 12 | 5.28 (t, 3.5) | 122.5 | 28 | 177.5 | |
| 13 | 143.5 | 29 | 1.26 (s) | 29.6 | |
| 14 | 41.7 | 30 | 1.26 (s) | 29.6 | |
| 15 | a, 1.16 (m) b b, 1.66 (m) | 27.6 | 1’ | 4.08 (t, 8.0,16.0) | 60.2 |
| 16 | a, 1.96 (m) b, 2.04 (m) | 23.5 | 2’ | 1.23 (s) | 14.2 |
| Compound No. | Compound Name | 1.0 mg/mL | 0.1 mg/mL | 0.01 mg/mL | IC50 (μM) |
|---|---|---|---|---|---|
| 1 | β-Sitosterol | - | - | - | - |
| 2 | 5,5′-Dibutoxy-2,2′-bifuran | - | - | - | - |
| 3 | Nonacosane-10-ol | - | - | - | - |
| 4 | Ethyl (3β)-3,23-dihydroxyolean-12-en-28-oate | - | - | - | - |
| 5 | Oleanonic acid | - | - | - | - |
| 6 | Ethyl caffeate | 100 ± 12.8% | 26.8 ± 1.6% | - | 1.0 |
| 7 | Caffeic acid | 90.2 ± 7.3% | 17.3 ± 5.7% | - | 1.5 |
| 8 | Isovanillin | 61.2 ± 2.8% | 20.1 ± 4.6% | - | 3.5 |
| 9 | Hederagenin | - | - | - | - |
| Positive control | Pepstatin A | 53.2 ± 3.6% (0.02 μM) | 39.2 ± 1.8% (0.002 μM) | 29.1 ± 12.5% (0.0002 μM) | 0.016 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, X.; Wei, Y.; Tian, W.-Y.; Sakharkar, M.K.; Liu, Q.; Yang, X.; Zhou, Y.-Z.; Mou, C.-L.; Cai, G.-L.; Yang, J. Characterization of Nine Compounds Isolated from the Acid Hydrolysate of Lonicera fulvotomentosa Hsu et S. C. Cheng and Evaluation of Their In Vitro Activity towards HIV Protease. Molecules 2019, 24, 4526. https://doi.org/10.3390/molecules24244526
Wang X, Wei Y, Tian W-Y, Sakharkar MK, Liu Q, Yang X, Zhou Y-Z, Mou C-L, Cai G-L, Yang J. Characterization of Nine Compounds Isolated from the Acid Hydrolysate of Lonicera fulvotomentosa Hsu et S. C. Cheng and Evaluation of Their In Vitro Activity towards HIV Protease. Molecules. 2019; 24(24):4526. https://doi.org/10.3390/molecules24244526
Chicago/Turabian StyleWang, Xia, Ying Wei, Wei-Yi Tian, Meena Kishore Sakharkar, Qing Liu, Xin Yang, Yan-Zi Zhou, Cheng-Li Mou, Gui-Lan Cai, and Jian Yang. 2019. "Characterization of Nine Compounds Isolated from the Acid Hydrolysate of Lonicera fulvotomentosa Hsu et S. C. Cheng and Evaluation of Their In Vitro Activity towards HIV Protease" Molecules 24, no. 24: 4526. https://doi.org/10.3390/molecules24244526
APA StyleWang, X., Wei, Y., Tian, W.-Y., Sakharkar, M. K., Liu, Q., Yang, X., Zhou, Y.-Z., Mou, C.-L., Cai, G.-L., & Yang, J. (2019). Characterization of Nine Compounds Isolated from the Acid Hydrolysate of Lonicera fulvotomentosa Hsu et S. C. Cheng and Evaluation of Their In Vitro Activity towards HIV Protease. Molecules, 24(24), 4526. https://doi.org/10.3390/molecules24244526






