Anti-Bacterial Effect and Cytotoxicity Assessment of Lipid 430 Isolated from Algibacter sp.
Abstract
1. Introduction
2. Results
2.1. High Troughput Screening and Identification of the Strains
2.2. Bioactivity Screening and Dereplication
2.3. Isolation of Lipid 430
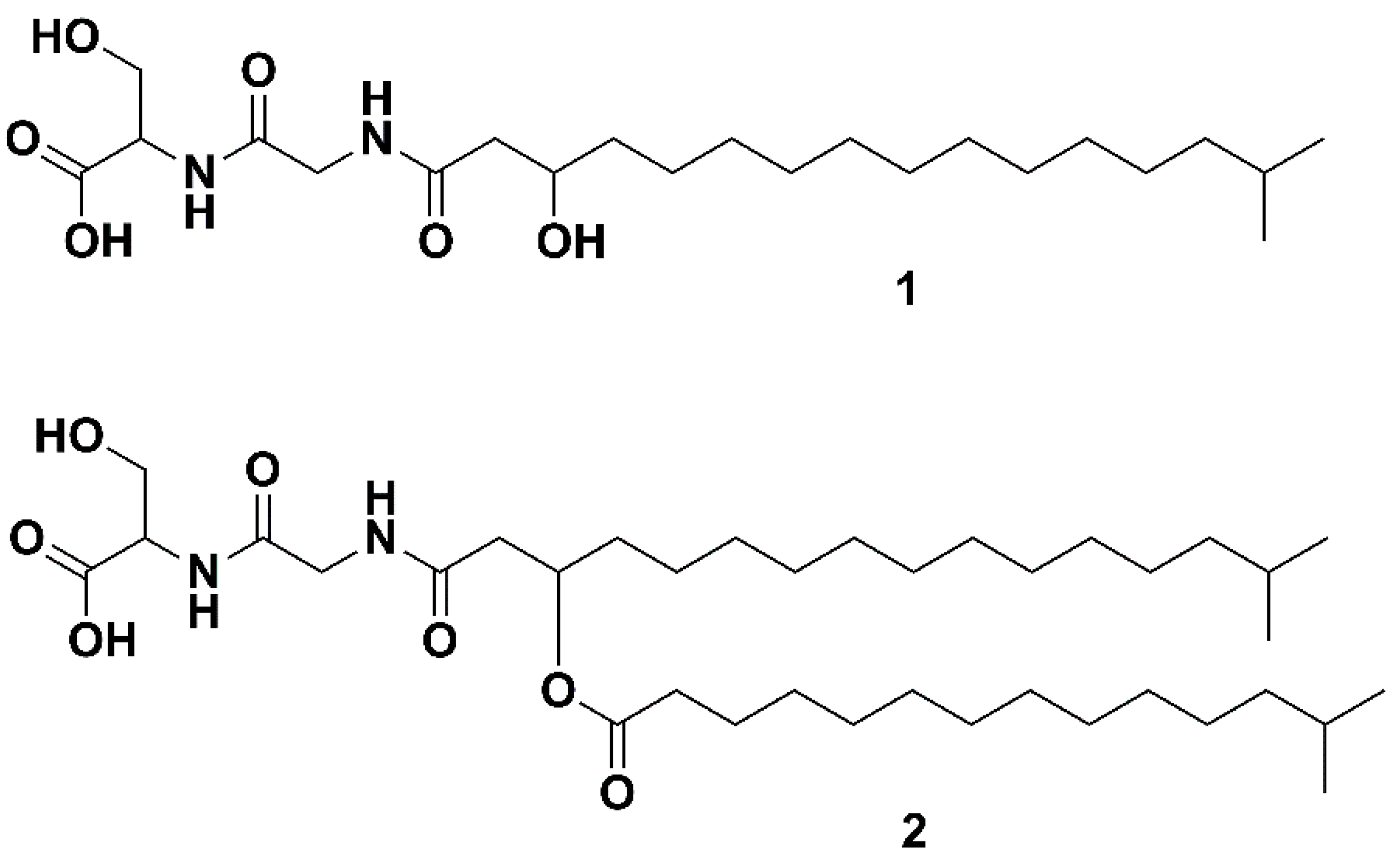
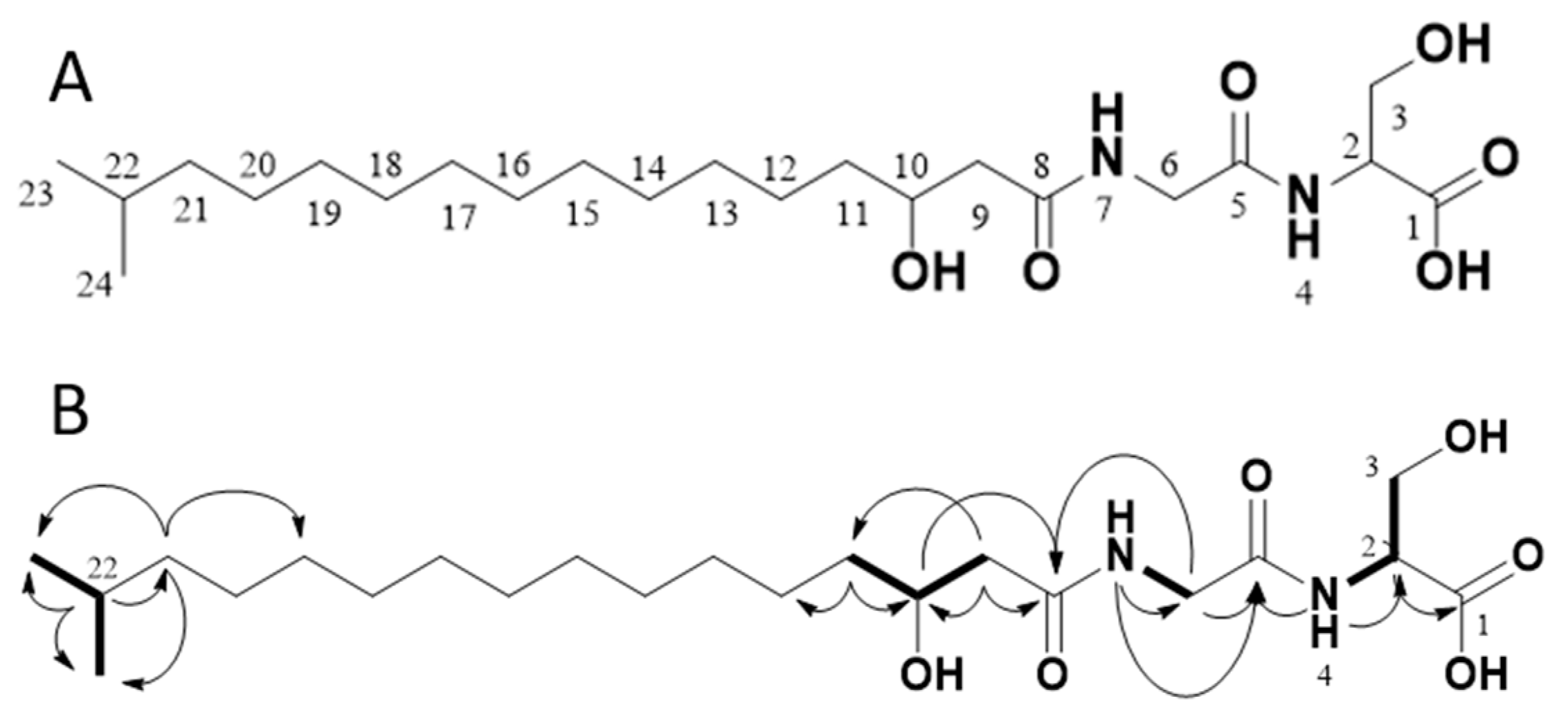
2.4. Structure Elucidation via NMR and MS/MS Analysis
2.5. Anti-microbial effect of Lipid 430
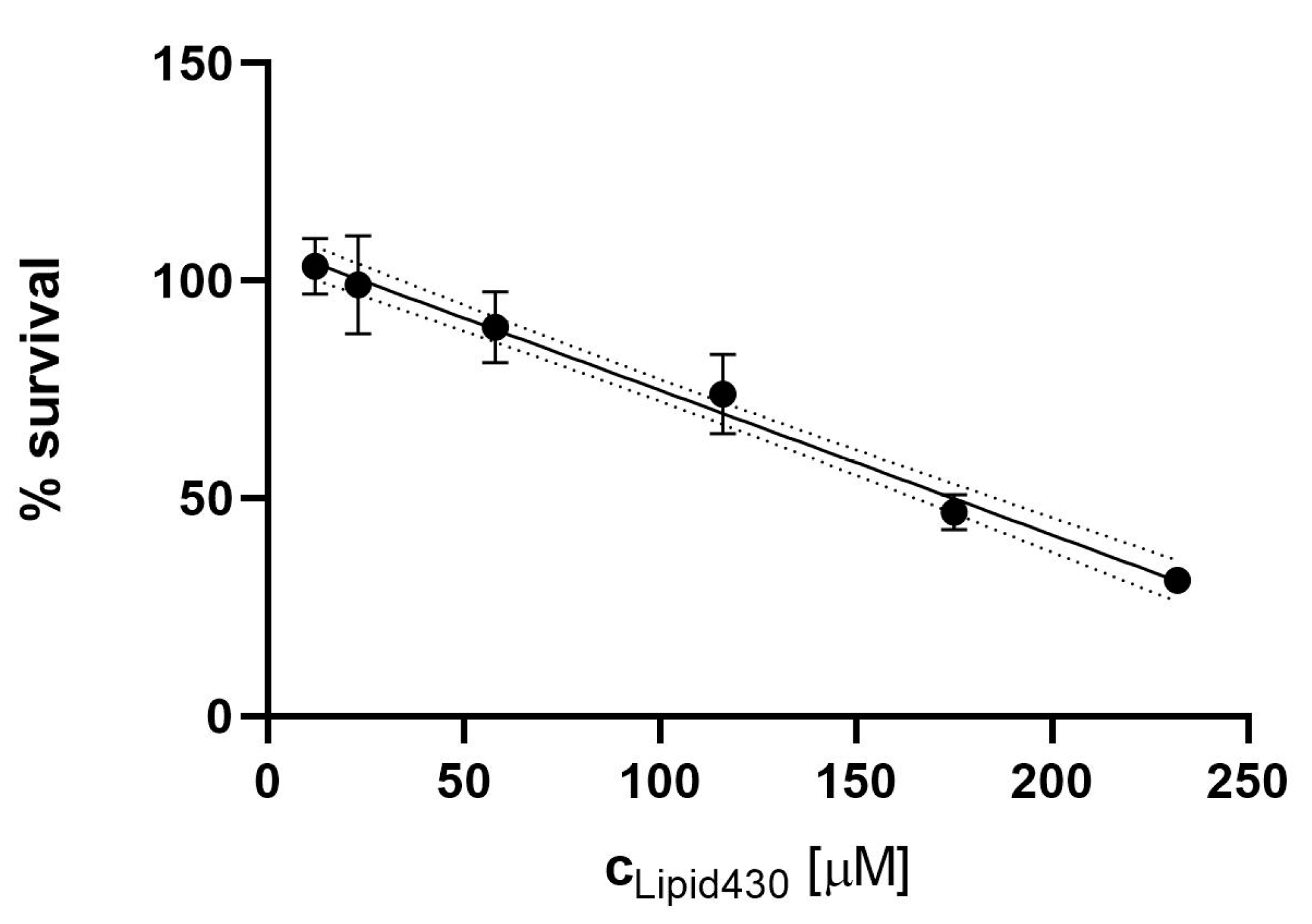
2.6. Cytotoxic Effect of 1
2.6.1. Cytotoxicity Assay
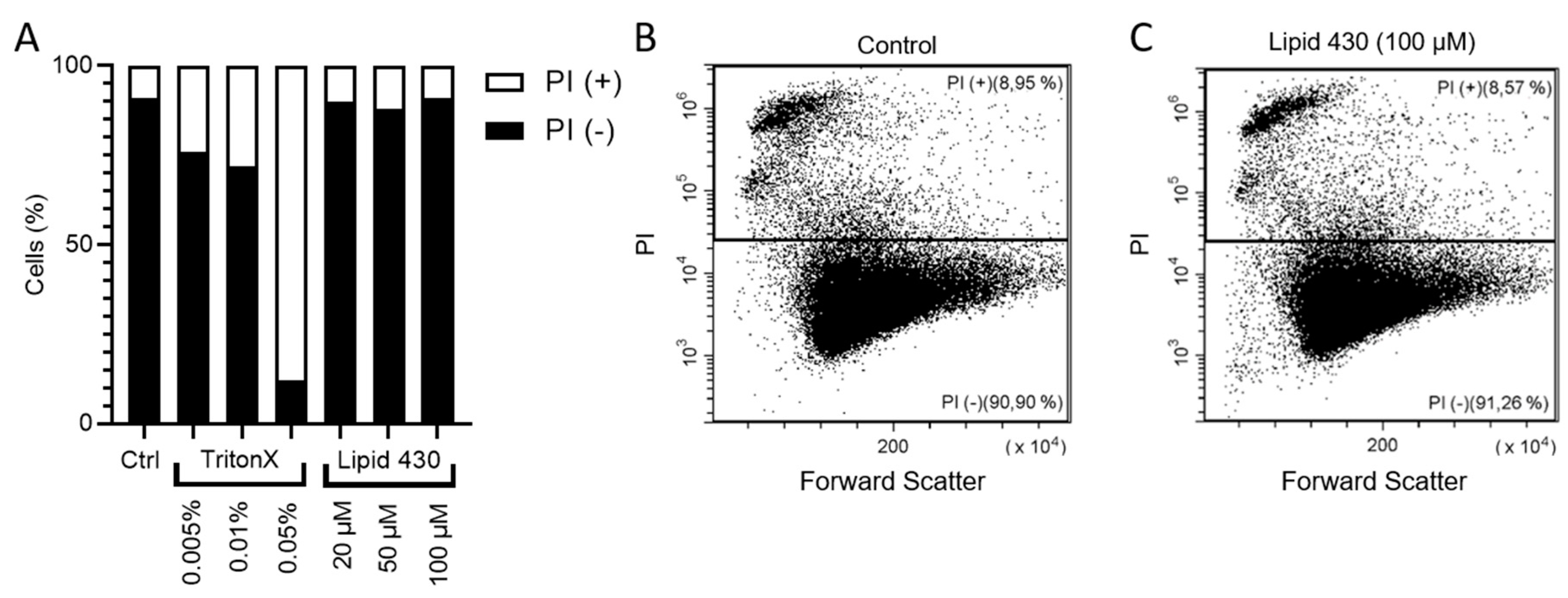
2.6.2. Propidium Iodide Staining and Flow Cytometry
2.6.3. Microscopic Investigation of the Melanoma Cell Line A2058
2.7. Lipid Isolation, Detection of Lipid 654
3. Discussion
4. Materials and Methods
4.1. Bacterial Isolates
4.2. PCR and Identification of the Strains
4.3. Fermentation and Extraction of Algibacter Cultures
4.4. Fractionation
4.5. UHPLC-HR-MS and Dereplication
4.6. Isolation of Lipid 430
4.7. NMR Spectroscopy
4.8. Lipid Extraction
4.9. Anti-Microbial Assays
4.9.1. Growth Inhibition Assay
4.9.2. Bactericidal Assay
4.10. Cell Proliferation Assay
4.11. Mode of Action Studies
4.11.1. Flow Cytometry
4.11.2. Microscopic Investigation of Melanoma Cells
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
| Strain ID | Genus | 16s rRNA sequence |
| M09B557 | Algibacter sp. | TTGGGTTTAAGGGTCCGTAGGTGGATAATTAAGTCAGAGGTGAAAGTTTGCAGCTCAACTGTAAAATTGCCTTTGATACTGGTTATCTTGAATCATTATGAAGTGGTTAGAATATGTAGTGTAGCGGTGAAATGCATAGATATTACATAGAATACCAATTGCGAAGGCAGATCACTAATAATGTATTGACACTGATGGACGAAAGCGTGGGGAGCGAACAGGATTAGATACCCTGGTAGTCCACGCCGTAAACGATGGATACTAGCTGTTCGGAACTTGTTTCTGAGTGGCTAAGCGAAAGTGATAAGTATCCCACCTGGGGAGTACGTTCGCAAGAATGAAACTCAAAGGAATTGACGGGGGCCCGCACAAGCGGTGGAGCATGTGGTTTAATTCGATGATACGCGAGGAACCTTACCAGGGCTTAAATGTAGATTGACAGGACTAGAGATAGTTTTTTCTTCGGACAATTTACAAGGTGCTGCATGGTTGTCGTCAGCTCGTGCCGTGAGGTGTCAGGTTAAGTCCTATAACGAGCGCAACCCCTGTTGTTAGTTGCCAGCGAGTCAAGTCGGGAACTCTAACAAGACTGCCAGTGCAAACTGTGAGGAAGGTGGGGATGACGTCAAATCATCACGGCCCTTACGTCCTGGGCTACACACGTGCTACAATGGTAGGGACAGAGAGCAGCCACTGGGCGACCAGGAGCGAATCTATAAACCCTATCACAGTTCGGATCGGAGTCTGCAACTCGACTCCGTGAAGCTGGAATCGCTAGTAATCGCATATCAGCCATGATGCGGNGAATACGTTCCCGGGNNNT |
| M09B045 | Algibacter sp. | TGANNGTTTGCAGCTCANNNNNNAAATTGCCTTTGATACNNGTTATCTTGAATCATTATGANNNNNNTAGANTNNGNANNNNNGCGGTGAAATGCATAGATATTACATAGAATACCAATTGCGAAGGCAGATCACTAATAATGTATTGACACTGATGGACGAAAGCGTGGGGAGCGAACAGGATTAGATACCCTGGTAGTCCACGCCGTAAACGATGGATACTAGCTGTTCGGAACTTGTTTCTGAGTGGCTAAGCGAAAGTGATAAGTATCCCACCTGGGGAGTACGTTCGCAAGAATGAAACTCAAAGGAATTGACGGGGGCCCGCACAAGCGGTGGAGCATGTGGTTTAATTCGATGATACGCGAGGAACCTTACCAGGGCTTAAATGTAGATTGACAGGACTAGAGATAGTTTTTTCTTCGGACAATTTACAAGGTGCTGCATGGTTGTCGTCAGCTCGTGCCGTGAGGTGTCAGGTTAAGTCCTATAACGAGCGCAACCCCTGTTGTTAGTTGCCAGCGAGTCATGTCGGGAACTCTAACAAGACTGCCAGTGCAAACTGTGAGGAAGGGGGGGGGATGACGTCAAATCATCACGGCCCTTACGTCCTGGGCTACACACGTGCTACAATGGTAGGGACAGAGAGCAGCCACTGGGCGACCAGGAGCGAATCTATAAACCCTATCACAGTTCGGATCGGAGTCTGCAACTCGACTCCGTGAAGCTGGAATCGCTAGTAATCGCATATCAGCCATGATGCGGTGAATACGTTCCCGGGCCTTGTACACACCGCCCGTCAAGCCATGGAAGCTGGGANTGNCTGAAGTCCGTCACCGTAAGGGAGCGGGC |
References
- Kirchman, D.L. The ecology of Cytophaga-Flavobacteria in aquatic environments. FEMS Microb. Ecol. 2002, 39, 91–100. [Google Scholar] [CrossRef]
- Fernández-Gómez, B.; Richter, M.; Schüler, M.; Pinhassi, J.; Acinas, S.G.; González, J.M.; Pedrós-Alió, C. Ecology of marine Bacteroidetes: A comparative genomics approach. ISME J. 2013, 7, 1026–1037. [Google Scholar] [CrossRef]
- Gómez-Pereira, P.R.; Fuchs, B.M.; Alonso, C.; Oliver, M.J.; Beusekom, J.E.E.; Amann, R. Distinct flavobacterial communities in contrasting water masses of the north Atlantic Ocean. ISME J. 2010, 4, 472–487. [Google Scholar] [CrossRef]
- Nedashkovskaya, O.I.; Kim, S.B.; Han, S.K.; Rhee, M.S.; Lysenko, A.M.; Rohde, M.; Zhukova, N.V.; Frolova, G.M.; Mikhailkov, V.V.; Bae, K.S. Algibacter lectus gen. nov., sp. nov., a novel member of the family Flavobacteriaceae isolated from green algae. Int. J. System. Evol. Microbiol. 2004, 54, 1257–1261. [Google Scholar] [CrossRef]
- Park, S.C.; Hwang, Y.M.; Choe, H.N.; Baik, K.S.; Kim, H.; Seong, C.N. Algibacter aquimarinus sp. nov., isolated from a marine environment, and reclassification of Pontirhabdus pectinivorans as Algibacter pectinivorans comb. nov. Int. J. System. Evol. Microbiol. 2013, 63, 2038–2042. [Google Scholar] [CrossRef]
- Park, S.C.; Hwang, Y.M.; Lee, J.H.; Baik, K.S.; Seong, C.N. Algibacter agarivorans sp. nov. and Algibacter agarilyticus sp. nov., isolated from seawater, reclassification of Marinivirga aestuarii as Algibacter aestuarii comb. nov. and emended description of the genus Algibacter. Int. J. System. Evol. Microbiol. 2013, 63, 3494–3500. [Google Scholar] [CrossRef]
- Nedashkovskaya, O.I.; Vancanneyt, M.; Kim, S.B.; Hoste, B.; Bae, K.S. Algibacter mikhailovii sp. nov., a novel marine bacterium of the family Flavobacteriaceae, and emended description of the genus Algibacter. Int. J. System. Evol. Microbiol. 2007, 57, 2147–2150. [Google Scholar] [CrossRef]
- Park, S.; Jung, Y.T.; Yoon, J.H. Algibacter miyuki sp. nov., a member of the family Flavobacteriaceae isolated from leachate of a brown algae reservoir. Antonie Van Leeuwenhoek 2013, 104, 253–260. [Google Scholar] [CrossRef]
- Yoon, J.H.; Park, S. Algibacter wandonensis sp nov., isolated from sediment around a brown algae (Undaria pinnatifida) reservoir. Int. J. System. Evol. Microbiol. 2013, 63, 4771–4776. [Google Scholar] [CrossRef]
- Sun, C.; Fu, G.; Zhang, C.; Hu, J.; Xu, L.; Wang, R.; Su, Y.; Han, S.; Yu, X.; Cheng, H.; et al. Isolation and complete genome sequence of Algibacter alginolytica sp. nov., a novel seaweed-degrading Bacteroidetes bacterium with diverse putative polysaccharide utilization loci. Appl. Environ. Microbiol. 2016, 82, 2975–2987. [Google Scholar] [CrossRef]
- Kawai, Y.; Yano, I.; Kaneda, K. Various kinds of lipoamino acids including a novel serine-containing lipid in an opportunistic pathogen Flavobacterium. Eur. J. Biochem. 1988, 171, 73–80. [Google Scholar] [CrossRef]
- Shiozaki, M.; Deguchi, N.; Mochizuki, T.; Wakabayashi, T.; Ishikawa, T.; Haruyama, H.; Kawai, Y.; Nishijima, M. Revised structure and synthesis of Flavolipin. Tetrahedron 1998, 54, 11861–11876. [Google Scholar] [CrossRef]
- Ratner, H. Flavobacterium meningosepticum. Inf. Contr. 1984, 5, 237–239. [Google Scholar] [CrossRef]
- Kawai, Y.; Akagawa, K. Macrophage activation by an ornithine-containing lipid or a serine-containing lipid. Inf. Immun. 1989, 57, 2086–2091. [Google Scholar]
- Gomi, K.; Kawasaki, K.; Kawai, Y.; Shiozaki, M.; Nishijima, M. Toll-like receptor 4-MD-2 complex mediates the signal transduction induced by Flavolipin, an amino acid-containing lipid unique to Flavobacterium meningosepticum. J. Immun. 2002, 168, 2939–2943. [Google Scholar] [CrossRef]
- Clark, B.R.; Cervantes, J.L.; Maciejewski, M.W.; Farrokhi, V.; Nemati, R.; Yao, X.; Anstadt, E.; Fujiwara, M.; Wright, K.T.; Riddle, C.; et al. Serine lipids of Porphyromonas gingivalis are human and mouse toll-like receptor 2 ligands. Inf. Immun. 2013, 81, 3479–3489. [Google Scholar] [CrossRef]
- Dietz, C.; Hart, T.K.; Nemati, R.; Yao, X.; Nichols, F.C.; Smith, M.B. Structural verification via convergent total synthesis of dipeptide–lipids isolated from Porphyromonas gingivalis. Tetrahedron 2016, 72, 7557–7569. [Google Scholar] [CrossRef]
- Wang, Y.H.; Jiang, J.; Zhu, Q.; AlAnezi, A.Z.; Clark, R.B.; Jiang, X.; Rowe, D.W.; Nichols, F.C. Porphyromonas gingivalis lipids inhibit osteoblastic differentiation and function. Infect. Immun. 2010, 78, 3726–3735. [Google Scholar] [CrossRef]
- Nichols, F.C.; Housley, W.J.; O’Conor, C.A.; Manning, T.; Wu, S.; Clark, R.B. Unique lipids from a common human bacterium represent a new class of toll-like receptor 2 ligands capable of enhancing autoimmunity. Ame. J. Pathol. 2009, 175, 2430–2438. [Google Scholar] [CrossRef]
- Wang, Y.H.; Nemati, R.; Anstadt, E.; Liu, Y.; Son, Y.; Zhu, Q.; Yao, X.; Clark, R.B.; Rowe, D.W.; Nichols, F.C. Serine dipeptide lipids of Porphyromonas gingivalis inhibit osteoblast differentiation: Relationship to Toll-like receptor 2. Bone 2015, 81, 654–661. [Google Scholar] [CrossRef]
- Olsen, I.; Nichols, F.C. Are Sphingolipids and serine dipeptide lipids underestimated virulence factors of Porphyromonas gingivalis? Infect. Immun. 2018, 86, e00035-18. [Google Scholar] [CrossRef]
- Nemati, R.; Dietz, C.; Anstadt, E.J.; Cervantes, J.; Liu, Y.; Dewhirst, F.E.; Clark, R.B.; Finegold, S.; Gallagher, J.J.; Smith, M.B.; et al. Deposition and hydrolysis of serine dipeptide lipids of Bacteroidetes bacteria in human arteries: Relationship to atherosclerosis. J. Lipid Res. 2017, 58, 1999–2007. [Google Scholar] [CrossRef]
- Nemati, R.; Dietz, C.; Anstadt, E.; Clark, R.; Smith, M.; Nichols, F.; Yao, X. Simultaneous determination of absolute configuration and quantity of lipopeptides using chiral liquid chromatography/mass spectrometry and diastereomeric internal standards. Anal. Chem. 2017, 89, 3583–3589. [Google Scholar] [CrossRef]
- Kristoffersen, V.; Rämä, T.; Isaksson, J.; Andersen, J.H.; Gerwick, W.H.; Hansen, E. Characterization of Rhamnolipids produced by an arctic marine bacterium from the Pseudomonas fluorescence group. Mar. Drugs 2018, 16, 163. [Google Scholar] [CrossRef]
- Sasaki, D.T.; Dumas, S.E.; Engleman, E.G. Discrimination of viable and non-viable cells using propidium iodide in two color immunofluorescence. Cytometry 1987, 8, 413–420. [Google Scholar] [CrossRef]
- Pietkiewicz, S.; Schmidt, J.H.; Lavrik, I.N. Quantification of apoptosis and necroptosis at the single cell level by a combination of imaging flow cytometry with classical annexin V/propidium iodide staining. J. Immun. Methods 2015, 423, 99–103. [Google Scholar] [CrossRef]
- Mascotti, K.; McCullough, J.; Burger, S.R. HPC viability measurement: Trypan blue versus acridine orange and propidium iodide. Transfusion 2000, 40, 693–696. [Google Scholar] [CrossRef]
- Hamley, I.W. Lipopeptides: From self-assembly to bioactivity. Chem. Commun. 2015, 51, 8574–8583. [Google Scholar] [CrossRef]
- Raaijmakers, J.M.; De Bruijn, I.; Nybroe, O.; Ongena, M. Natural functions of lipopeptides from Bacillus and Pseudomonas: More than surfactants and antibiotics. FEMS Microbiol. Rev. 2010, 34, 1037–1062. [Google Scholar] [CrossRef]
- Seydlová, G.; Svobodová, J. Review of surfactin chemical properties and the potential biomedical applications. Cent. Eur. J. Med. 2008, 3, 123–133. [Google Scholar] [CrossRef]
- Baltz, R.H. Daptomycin: Mechanisms of action and resistance, and biosynthetic engineering. Curr. Opin. Chem. Biol. 2009, 13, 144–151. [Google Scholar] [CrossRef] [PubMed]
- Mor, A.; Gurwitz, D. Peptide-based antibiotics: A potential answer to raging antimicrobial resistance. Drug Dev. Res. 2000, 50, 440–447. [Google Scholar] [CrossRef]
- Humphries, R.M.; Pollett, S.; Sakoulas, G. A current perspective on Daptomycin for the clinical microbiologist. Clin. Microbiol. Rev. 2013, 26, 759–780. [Google Scholar] [CrossRef] [PubMed]
- Debono, M.; Abbott, B.J.; Molloy, R.M.; Fukuda, D.S.; Hunt, A.H.; Daupert, V.M.; Counter, F.T.; Ott, J.L.; Carrell, C.B.; Howard, L.C.; et al. Enzymatic and chemical modifications of lipopeptide antibiotic A21978C: The synthesis and evaluation of Daptomycin (LY146032). J. Antibiot. 1988, 41, 1093–1105. [Google Scholar] [CrossRef] [PubMed]
- Peypoux, F.; Bonmatin, J.M.; Wallach, J. Recent trends in the biochemistry of surfactin. Appl. Microbiol. Biotech. 1999, 51, 553–563. [Google Scholar] [CrossRef] [PubMed]
- Tareq, F.S.; Lee, M.A.; Lee, H.-S.; Lee, J.-S.; Lee, Y.-J.; Shin, H.J. Gageostatins A-C, antimicrobial linear lipopeptides from a marine Bacillus Subtilis. Mar. Drugs 2014, 12, 871–885. [Google Scholar] [CrossRef] [PubMed]
- Makovitzki, A.; Avrahami, D.; Shai, Y. Ultrashort antibacterial and antifungal lipopeptides. Proc. Natl. Acad. Sci. USA 2006, 103, 15997–16002. [Google Scholar] [CrossRef]
- Chrzanowski, Ł.; Ławniczak, Ł.; Czaczyk, K. Why do microorganisms produce rhamnolipids? World J. Microbiol. Biotechnol. 2012, 28, 401–419. [Google Scholar] [CrossRef]
- Abdel-Mawgoud, A.; Lépine, F.; Déziel, E. Rhamnolipids: Diversity of structures, microbial origins and roles. Appl. Microbiol. Biotechnol. 2010, 86, 1323–1336. [Google Scholar] [CrossRef]
Sample Availability: Samples of the compound 1 are available from the authors. |


| Lipid 430 (1) | ||
|---|---|---|
| Position | δC, Type | δH (J in Hz) |
| 1 | 173.8, C | |
| 2 | 56.5, CH | 4.47, dt (8.2, 4.2) |
| 3a | 63.2, CH2 | 3.89, dd (9.3, 5.2) ) c |
| 3b | 3.82, dd (11.3, 3.9) | |
| 4 | 7.99, d (7.8) | |
| 5 | 171.5, C | |
| 6a | 43.6, CH2 | 3.98, dd (16.7, 5.9) b |
| 6b | 3.89, dd (9.3, 5.2) c | |
| 7 | 8.31, t (5.9) | |
| 8 | 175.0, C | |
| 9a | 44.8, CH2 | 2.40, dd (13.9, 4.1) |
| 9b | 2.33, dd (14.0, 8.8) | |
| 10 | 70.0, CH | 3.98, dd (16.7, 5.9) b |
| 11 | 38.3, CH2 | 1.50–1.47, m |
| 12 | 26.5, CH2 | 1.47–1.42, m |
| 13 | 30.9–30.6 a, CH2 | 1.29, p (6.1, 5.4) e |
| 14 | 30.9–30.6 a, CH2 | 1.29, p (6.1, 5.4) e |
| 15 | 30.9–30.6 a, CH2 | 1.29, p (6.1, 5.4) e |
| 16 | 30.9–30.6 a, CH2 | 1.29, p (6.1, 5.4) e |
| 17 | 30.9–30.6 a, CH2 | 1.29, p (6.1, 5.4) e |
| 18 | 30.9–30.6 a, CH2 | 1.29, p (6.1, 5.4) e |
| 19 | 30.9–30.6 a, CH2 | 1.29, p (6.1, 5.4) e |
| 20 | 28.4, CH2 | 1.29, p (6.1, 5.4) e |
| 21 | 40.1, CH2 | 1.16, q (7.1, 6.7) |
| 22 | 29.0, CH | 1.56–1.50, m |
| 23 | 22.9, CH3 | 0.87, d (6.6) d |
| 24 | 22.9, CH3 | 0.87, d (6.6) d |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Schneider, Y.K.-H.; Ø. Hansen, K.; Isaksson, J.; Ullsten, S.; H. Hansen, E.; Hammer Andersen, J. Anti-Bacterial Effect and Cytotoxicity Assessment of Lipid 430 Isolated from Algibacter sp. Molecules 2019, 24, 3991. https://doi.org/10.3390/molecules24213991
Schneider YK-H, Ø. Hansen K, Isaksson J, Ullsten S, H. Hansen E, Hammer Andersen J. Anti-Bacterial Effect and Cytotoxicity Assessment of Lipid 430 Isolated from Algibacter sp. Molecules. 2019; 24(21):3991. https://doi.org/10.3390/molecules24213991
Chicago/Turabian StyleSchneider, Yannik K.-H., Kine Ø. Hansen, Johan Isaksson, Sara Ullsten, Espen H. Hansen, and Jeanette Hammer Andersen. 2019. "Anti-Bacterial Effect and Cytotoxicity Assessment of Lipid 430 Isolated from Algibacter sp." Molecules 24, no. 21: 3991. https://doi.org/10.3390/molecules24213991
APA StyleSchneider, Y. K.-H., Ø. Hansen, K., Isaksson, J., Ullsten, S., H. Hansen, E., & Hammer Andersen, J. (2019). Anti-Bacterial Effect and Cytotoxicity Assessment of Lipid 430 Isolated from Algibacter sp. Molecules, 24(21), 3991. https://doi.org/10.3390/molecules24213991






