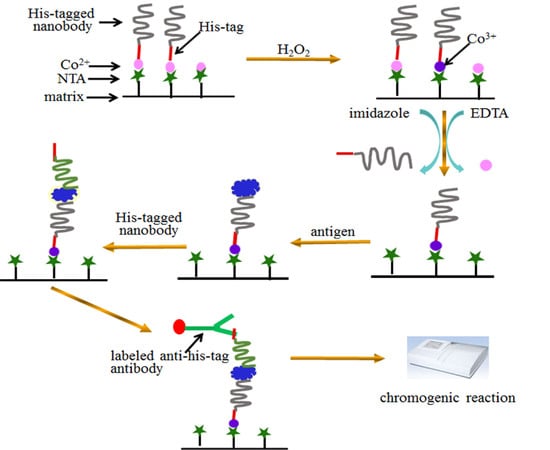
Oriented Immobilization and Quantitative Analysis Simultaneously Realized in Sandwich Immunoassay via His-Tagged Nanobody
Abstract
:1. Introduction
2. Results and Discussion
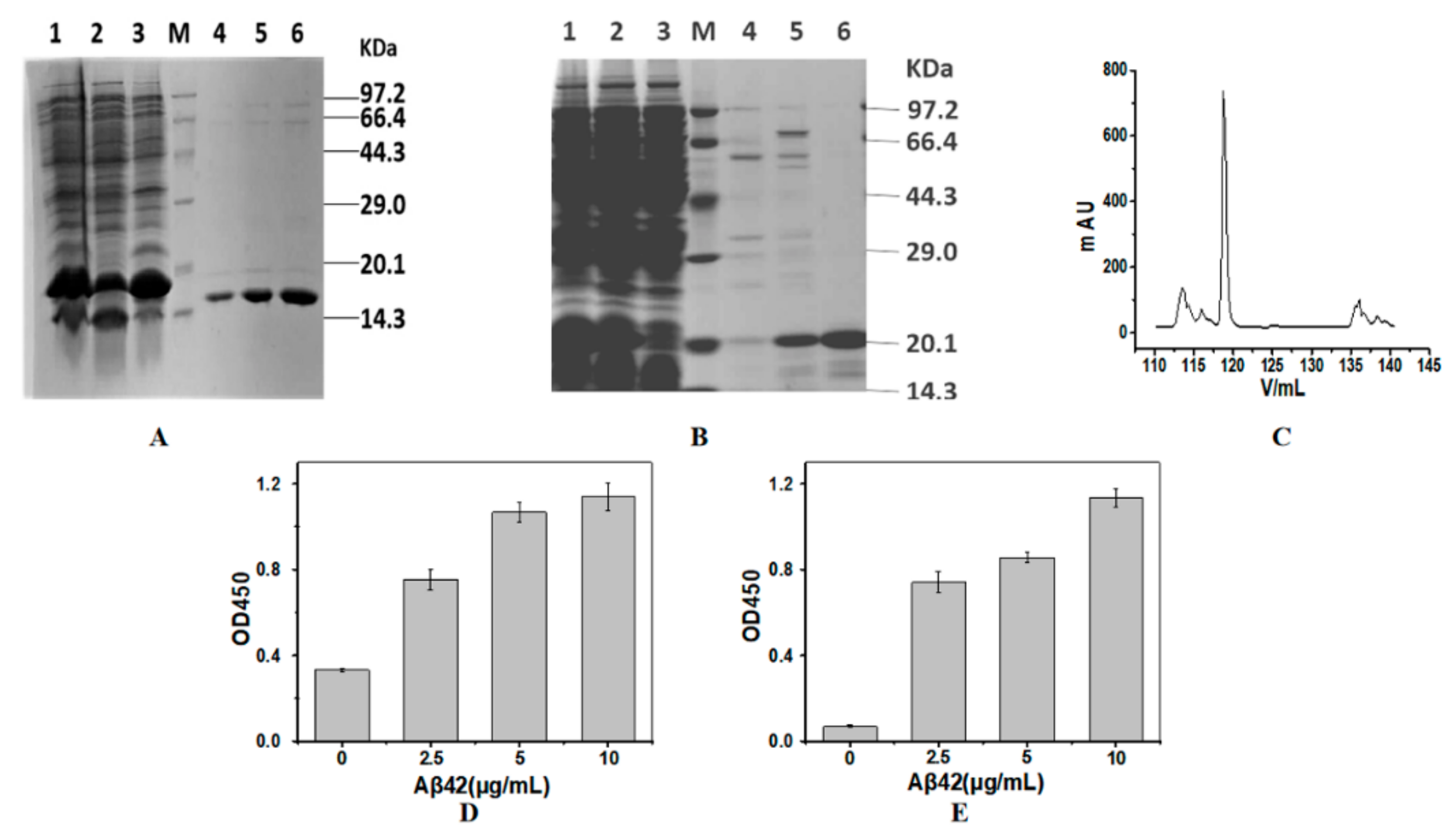
2.1. Expression and Purification of the Nanobodies
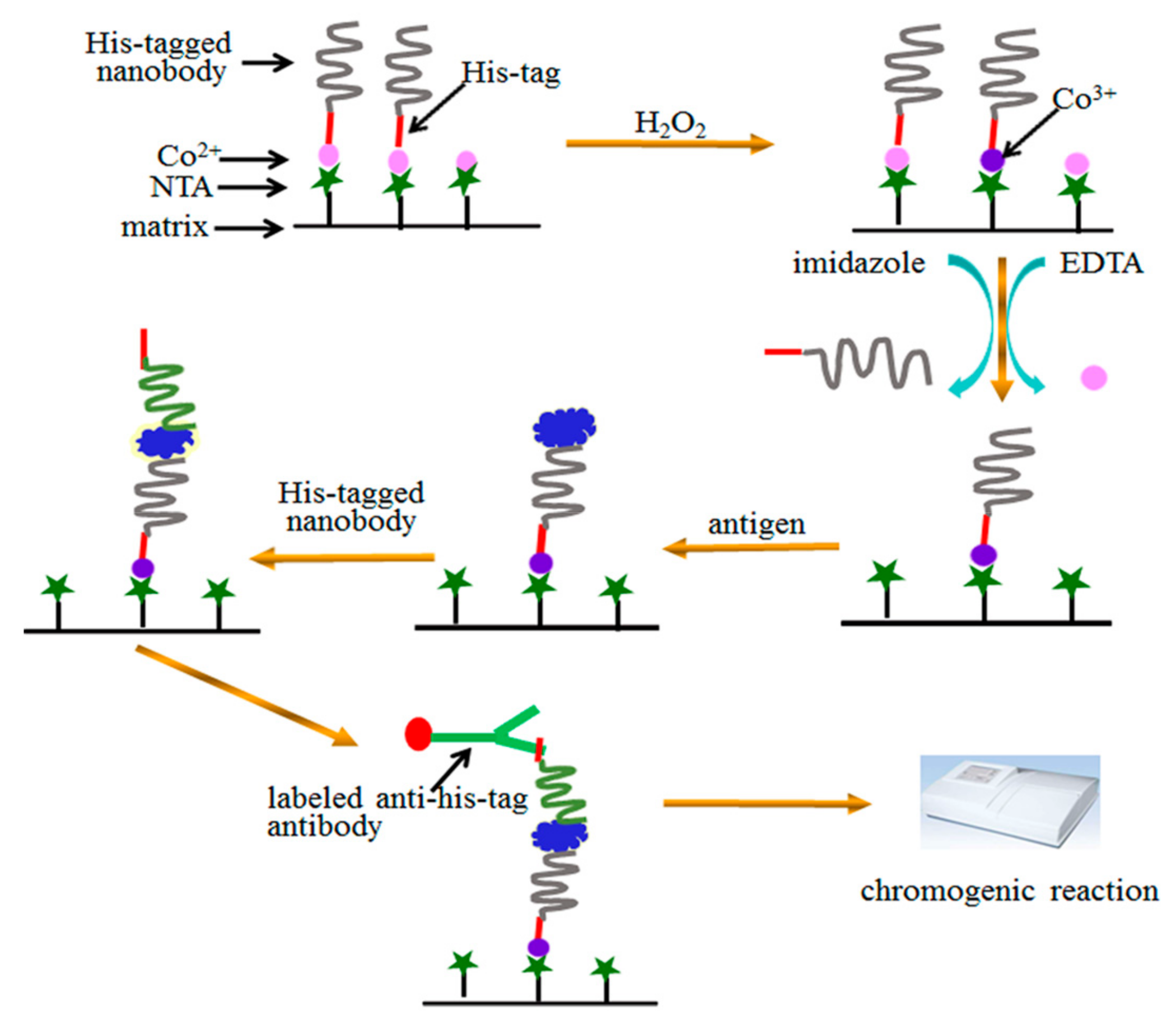
2.2. Immobilization of SH2
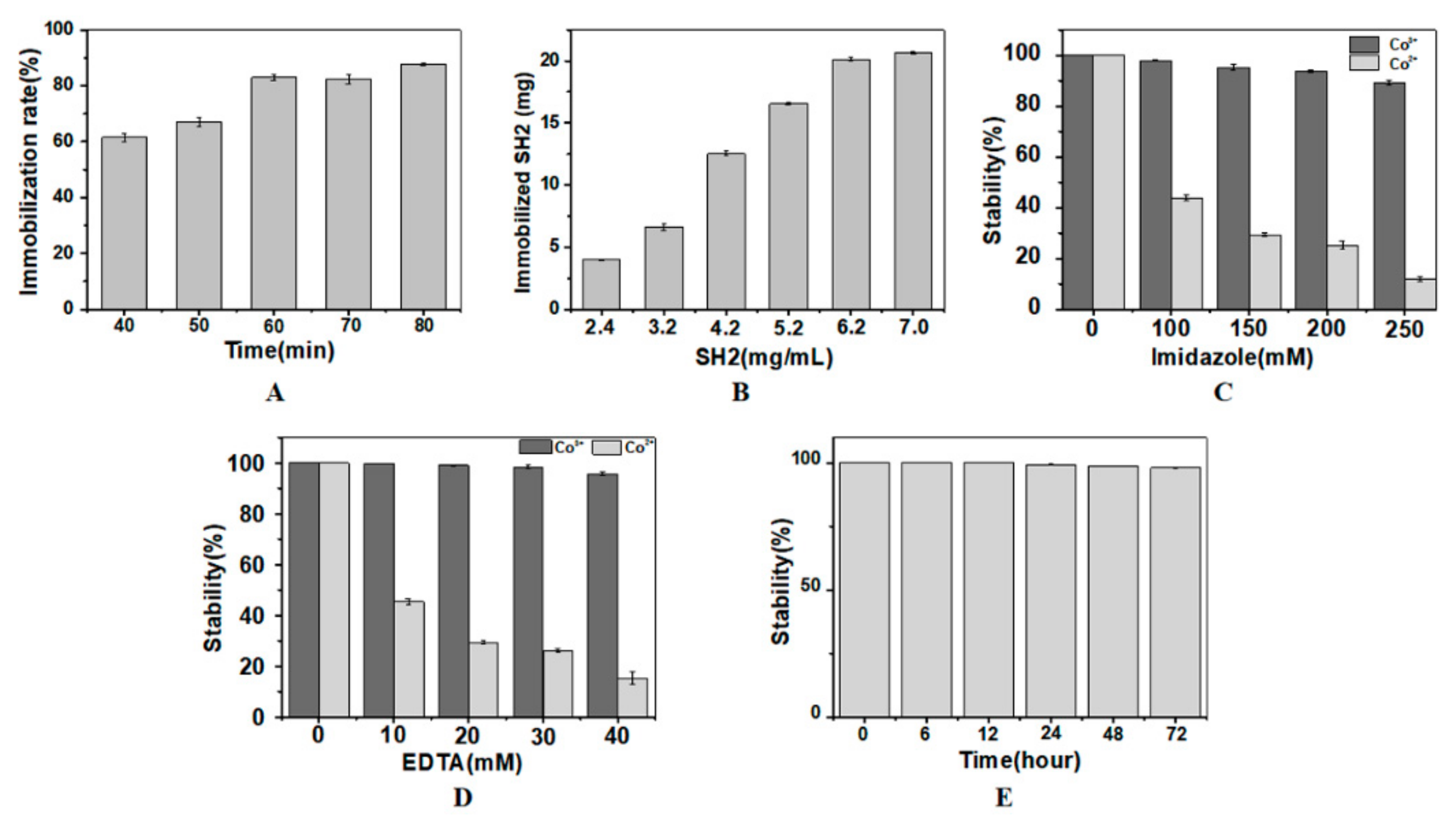
2.2.1. Effects of H2O2 Oxidation Time and SH2 Concentration on SH2 Immobilization
2.2.2. Effects of Imidazole and EDTA Elution on Immobilized SH2
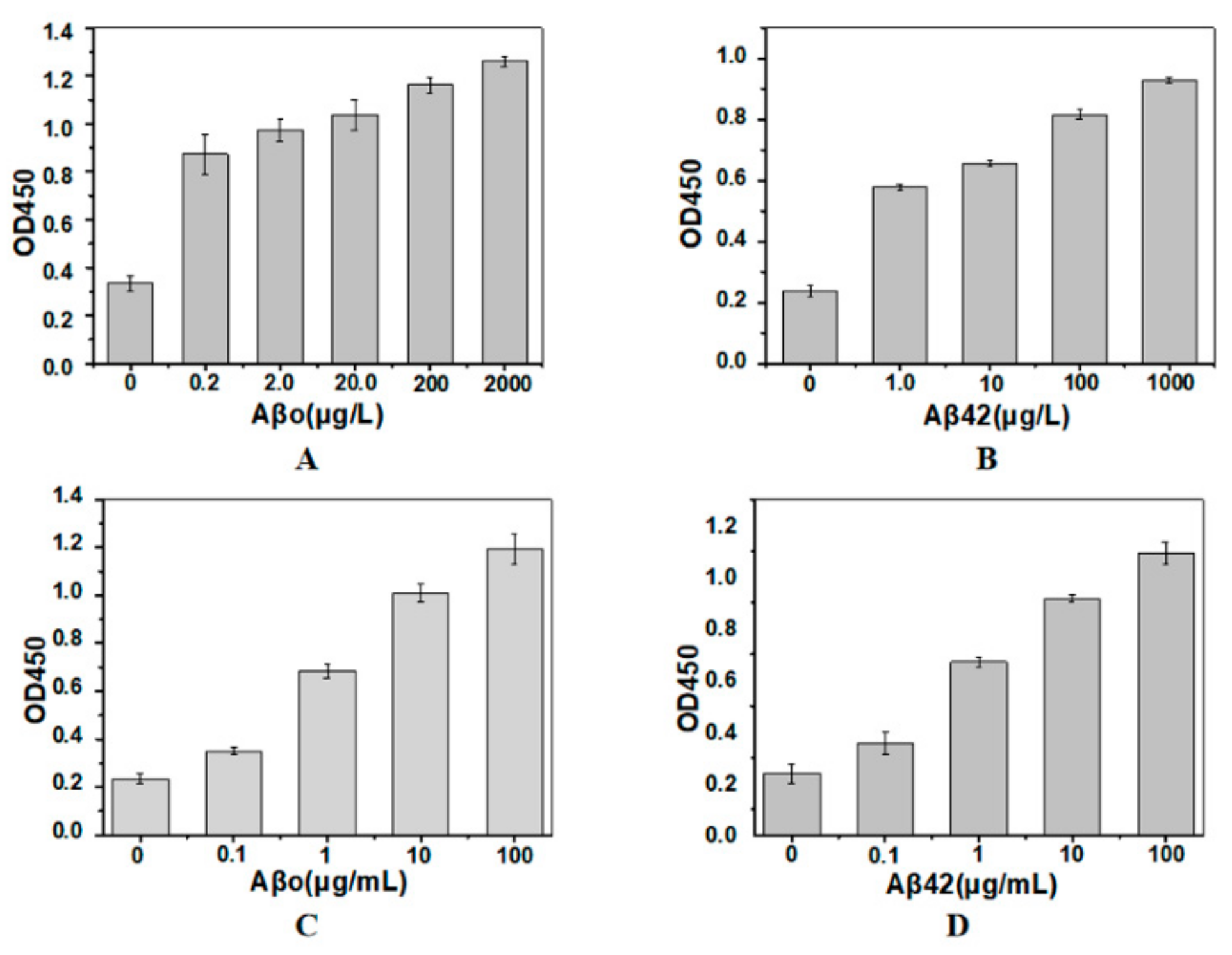
2.3. Sandwich Immunoassay Using the Immobilized Nanobody
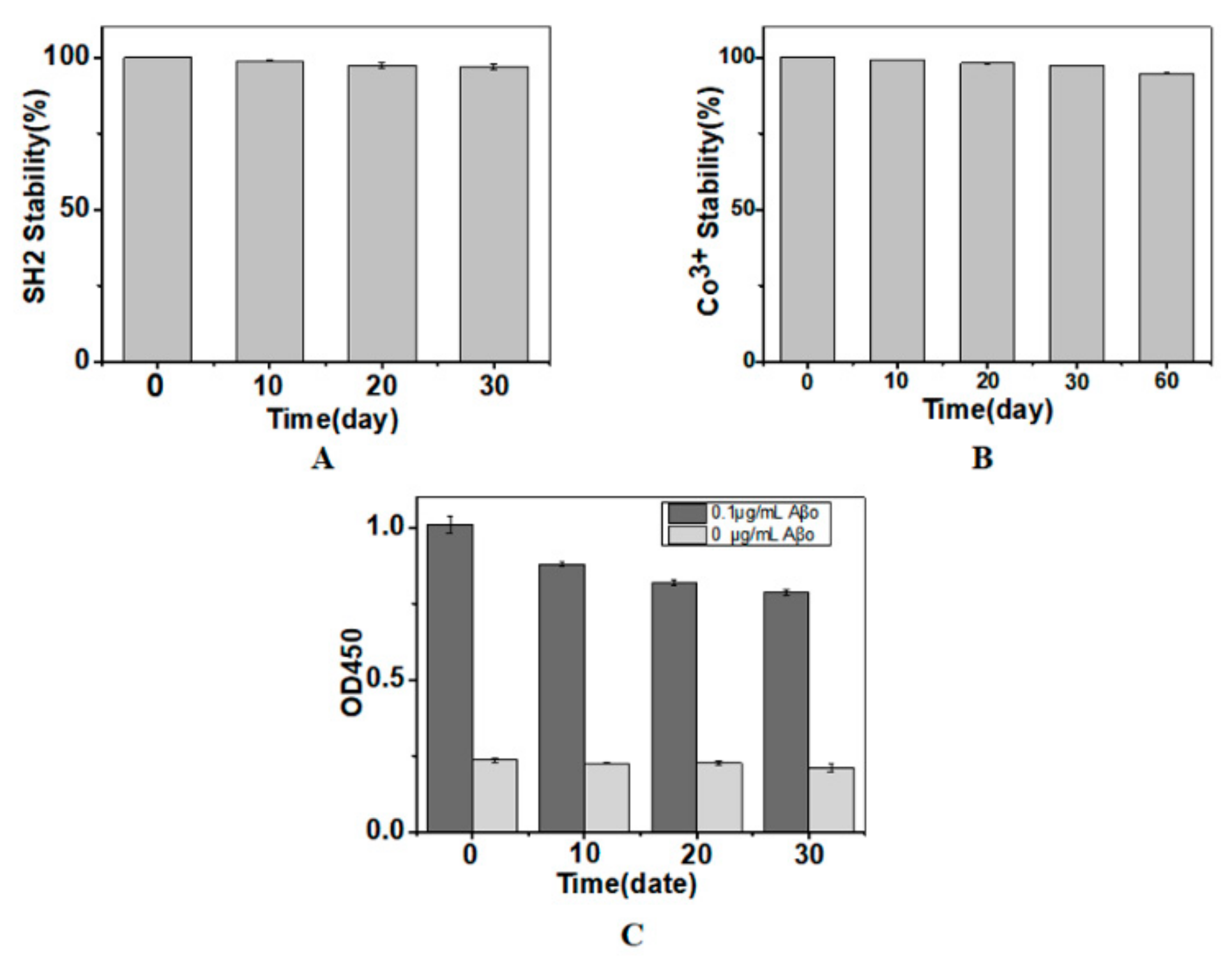
2.4. Storage Stability of the Immobilized SH2
3. Materials and Methods
3.1. Reagents and Materials
3.2. Preparation of Aβ Monomer and Aβo
3.3. Expression and Purification of Nanobodies
3.4. Indirect ELISA to Measure Activity of the Purified Nanobodies
3.5. Immobilization Nanobody on Nitrilotriacetic Acid Modified Beads
3.6. Sandwich ELISA
3.7. Storage Stability of the Immobilized Nanobody
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| NTA | nitrotriacetic acid |
| scFv | single chain antibody fragment |
| Aβ | β amyloid |
| Aβo | oligomer oligomer β amyloid |
| ELISA | enzyme linked immunosorbent assay |
References
- Harlow, E.; Lane, D. Using Antibodies: A Laboratory Manual; Cold Spring Harbor Laboratory Press: New York, NY, USA, 1999. [Google Scholar]
- Shepherd, P.; Dean, C. Monoclonal Antibodies; Oxford University Press: New York, NY, USA, 2000; pp. 297–300. [Google Scholar]
- Berson, S.A.; Yalow, R.S. Quantitative aspects of the reaction between insulin and insulin-binding antibody. J. Clin. Invest. 1959, 38, 1996–2016. [Google Scholar] [CrossRef]
- Surugiu, I.; Svitel, J.; Ye, L.; Haupt, K.; Danielsson, B. Development of a flow injection capillary chemiluminescent ELISA using an imprinted polymer instead of the antibody. Anal. Chem. 2001, 73, 4388–4392. [Google Scholar] [CrossRef]
- Li, Z.P.; Wang, Y.C.; Liu, C.H.; Li, Y.K. Development of chemiluminescence detection of gold nanoparticles in biological conjugates for immunoassay. Anal. Chim. Acta 2005, 551, 85–91. [Google Scholar] [CrossRef]
- Engvall, E.; Perlmann, P. Enzyme-linked immunosorbent assay (Elisa) quantitative assay of immunoglobulin-G. Immunochemistry 1971, 8, 871–874. [Google Scholar] [PubMed]
- Arakawa, H.; Maeda, M.; Tsuji, A. Chemi-luminescence enzyme immunoassay of cortisol using peroxidase as label. Anal. Biochem. 1979, 97, 248–254. [Google Scholar] [CrossRef]
- Klein, C.; Hagenah, J.; Landwehrmeyer, B.; Münte, T.; Klockgether, T. The presymptomatic stage of neurodegenerative disorders. Der Nervenarzt 2011, 82, 994–1001. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Wang, J.; Cui, L.; Zou, X.; Zhang, Y. Recombinant GST-I-A beta 28-induced efficient serum antibody against A beta 42. J. Neurosci. Methods 2010, 186, 52–59. [Google Scholar] [CrossRef] [PubMed]
- Hamers-Casterman, C.; Atarhouch, T.; Muyldermans, S.; Robinson, G.; Hamers, C.; Songa, E.B.; Bendahman, N.; Hamers, R. Naturally occurring antibodies devoid of light chains. Nature 1993, 363, 446. [Google Scholar] [CrossRef] [PubMed]
- Bruce, M.P.; Boyd, V.; Duch, C.; White, J.R. Dialysis-based bioreactor systems for the production of monoclonal antibodies-alternatives to ascites production in mice. J. Immunol. Methods 2002, 264, 59–68. [Google Scholar] [CrossRef]
- Zheng, M.Z.; Richard, J.J.; Binder, J. A review of rapid methods for the analysis of mycotoxins. Mycopathologia 2006, 161, 261–273. [Google Scholar] [CrossRef] [PubMed]
- Luo, L.; Zhang, Z.; Hou, L.; Wang, J.; Tian, W. The study of a chemiluminescence immunoassay using the peroxyoxalate chemiluminescent reaction and its application. Talanta 2007, 72, 1293–1297. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.S.; Song, H.P.; Chen, Q.; Yu, J.L.; Xian, M.; Nian, R.; Feng, D.X. Recent advances in the selection and identification of antigen-specific nanobodies. Mol. Immunol. 2018, 96, 37–47. [Google Scholar] [CrossRef]
- Hemdan, E.S.; Zhao, Y.J.; Sulkowski, E.; Porath, J. Surface topography of histidine residues: a facile probe by immobilized metal ion affinity chromatography. Proc. Natl. Acad. Sci. USA 1989, 86, 1811–1815. [Google Scholar] [CrossRef] [PubMed]
- Yip, T.T.; Hutchens, T.W. Immobilized metal ion affinity chromatography. Protein Expres. Purif. 1992, 3, 263–281. [Google Scholar]
- Kurztkowska, K.; Mielecki, M.; Grzelak, K.; Verwilst, P.; Dehaen, W.; Radecki, J.; Radecka, H. Immobilization of His-tagged kinase JAK2 onto the surface of a plasmon resonance gold disc modified with different copper (II) complexes. Talanta 2014, 130, 336–341. [Google Scholar] [CrossRef] [PubMed]
- Rusmini, F.; Zhong, Z.; Feijen, J. Protein immobilization strategies for protein biochips. Biomacromolecules 2007, 8, 1775–1789. [Google Scholar] [CrossRef]
- Valiokas, R.; Klenkar, G.; Tinazli, A.; Tampe, R.; Liedberg, B.; Piehler, J. Differential protein assembly on micropatterned surfaces with tailored molecular and surface multivalency. Chembiochem 2010, 7, 1285. [Google Scholar] [CrossRef]
- Park, S.J.; Kim, S.; Kim, S.H.; Park, K.M.; Hwang, B.H. His-tagged protein immobilization on cationic ferrite magnetic nanoparticles. Korean J. Chem. Eng. 2018, 35, 1297–1302. [Google Scholar] [CrossRef]
- Goldsmith, C.R.; Jaworski, J.; Sheng, M.; Lippard, S.J. Selective labeling of extracellular proteins containing polyhistidine sequences by a fluorescein-nitrilotriacetic acid conjugate. J. Am. Chem. Soc. 2006, 128, 418–419. [Google Scholar] [CrossRef]
- Kamoto, M.; Umezawa, N.; Kato, N.; Higuchi, T. Novel probes showing specific fluorescence enhancement on binding to a hexahistidine tag. Chem. Eur. J. 2010, 14, 8004–8012. [Google Scholar] [CrossRef]
- Gavutis, M.; Lata, S.; Piehler, J. Probing 2-dimensional protein-protein interactions on model membranes. Nat. Protoc. 2006, 1, 2091–2103. [Google Scholar] [CrossRef] [PubMed]
- Pires, M.M.; Ernenwein, D.; Chmielewski, J. Selective decoration and release of His-tagged proteins from metal-assembled collagen peptide microflorettes. Biomacromolecules 2011, 12, 2429–2433. [Google Scholar] [CrossRef] [PubMed]
- Wegner, S.V.; Spatz, J.P. Cobalt(III) as a stable and inert mediator ion between NTA and His6-tagged proteins. Angew Chem. Int. Edit. 2013, 52, 7593–7596. [Google Scholar] [CrossRef]
- Xu, L.; Wang, R.; Cao, H.Y.; Xu, T.; Han, L.L.; Huang, C.D.; Jia, L.Y. A facile method to oriented immobilization of His-tagged BirA on Co3+-NTA agarose beads. Enzyme Microb. Tech. 2019, 120, 36–42. [Google Scholar] [CrossRef]
- Racine, A.M.; Koscik, R.L.; Nicholas, C.R.; Clark, L.R.; Okonkwo, O.C.; Oh, J.M.; Hillmer, A.T.; Murali, D.; Barnhart, T.E.; Betthauser, T.J.; et al. Cerebrospinal fluid ratios with Abeta42 predict preclinical brain beta-amyloid accumulation. Alzheheimer’s Dement. 2016, 2, 27–38. [Google Scholar]
- Adamczuk, K.; Schaeverbeke, J.; Vanderstichele, H.M.; Lilja, J.; Nelissen, N.; Van Laere, K.; Dupont, P.; Hilven, K.; Poesen, K.; Vandenberghe, R. Diagnostic value of cerebrospinal floid Abeta ratios in preclinical Alzhemer’s disease. Alzhemer’s Res. Ther. 2015, 7, 75. [Google Scholar] [CrossRef]
- Shao, S.; Geng, J.; Ah Yi, H.; Gogia, S.; Neelamegham, S.; Jacobs, A.; Lovell, J.F. Functionalization of cobalt porphyrin-phospholipid bilayers with His-tagged ligands and antigens. Nat. Chem. 2015, 7, 438–446. [Google Scholar] [CrossRef] [PubMed]
- Auer, S.; Hellmann, F.; Krause, M.; Kurths, J. Stable immobilisation of His-tagged proteins on BLI biosensor surface using cobalt. Chaosl 2017, 243, 104–113. [Google Scholar] [CrossRef]
- Hernandez, K.; Berenguer-Murcia, A.; Rodrigues, R.C.; Fernandez-Lafuente, R. Hydrogen peroxide in biocatalysis. Curr. Org. Chem. 2012, 16, 2652–2672. [Google Scholar] [CrossRef]
- Stadtman, E.R.; Levine, R.L. Free radical-mediated oxidation of free amino acids and amino acid residues in proteins. Amino Acids 2003, 25, 207. [Google Scholar] [CrossRef]
- Wang, C.; Cheng, F.; Xu, L.; Jia, L.Y. HSA targets multiple Aβ42 species and inhibits the seeding-mediated aggregation and cytotoxicity of Aβ42 aggregates. RSC Adv. 2016, 6, 71165–71175. [Google Scholar] [CrossRef]
- Strittmatter, W.J.; Weisgraber, K.H.; Huang, D.Y.; Dong, L.M.; Salvesen, G.S.; Pericakvance, M.; Schmechel, D.; Saunders, A.M.; Goldgaber, D.; Roses, A.D. Binding of human apolipoprotein E to synthetic amyloid beta peptide: Isoform-specific effects and implications for late-onset Alzheimer’s disease. Proc. Natl. Acad. Sci. USA 1993, 90, 8098–8102. [Google Scholar] [CrossRef] [PubMed]
- Jan, A.; Hartley, D.M.; Lashuel, H.A. Preparation and characterization of toxic Abeta aggregates for structural and functional studies in Alzheimer’s disease research. Nat. Protoc. 2010, 5, 1186–1209. [Google Scholar] [CrossRef] [PubMed]
- Zor, T.; Selinger, Z. Linearization of the Bradford protein assay increases its sensitivity: Theoretical and experimental studies. Anal. Biochem. 1996, 236, 302–308. [Google Scholar] [CrossRef]
Sample Availability: Samples of the compounds are available from the authors, except the nanobody. |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xu, L.; Cao, H.; Huang, C.; Jia, L. Oriented Immobilization and Quantitative Analysis Simultaneously Realized in Sandwich Immunoassay via His-Tagged Nanobody. Molecules 2019, 24, 1890. https://doi.org/10.3390/molecules24101890
Xu L, Cao H, Huang C, Jia L. Oriented Immobilization and Quantitative Analysis Simultaneously Realized in Sandwich Immunoassay via His-Tagged Nanobody. Molecules. 2019; 24(10):1890. https://doi.org/10.3390/molecules24101890
Chicago/Turabian StyleXu, Li, Hanyu Cao, Chundong Huang, and Lingyun Jia. 2019. "Oriented Immobilization and Quantitative Analysis Simultaneously Realized in Sandwich Immunoassay via His-Tagged Nanobody" Molecules 24, no. 10: 1890. https://doi.org/10.3390/molecules24101890
APA StyleXu, L., Cao, H., Huang, C., & Jia, L. (2019). Oriented Immobilization and Quantitative Analysis Simultaneously Realized in Sandwich Immunoassay via His-Tagged Nanobody. Molecules, 24(10), 1890. https://doi.org/10.3390/molecules24101890