Applications of Cancer Cell-Specific Aptamers in Targeted Delivery of Anticancer Therapeutic Agents
Abstract
:1. Introduction
2. Development of Cancer Cell-Targeting Aptamers
2.1. Development of Aptamers against Biomarkers Using SELEX Technology
2.2. Aptamers as Cancer Cell-Targeting Agents
3. Aptamer-Mediated Therapeutics against Cancer
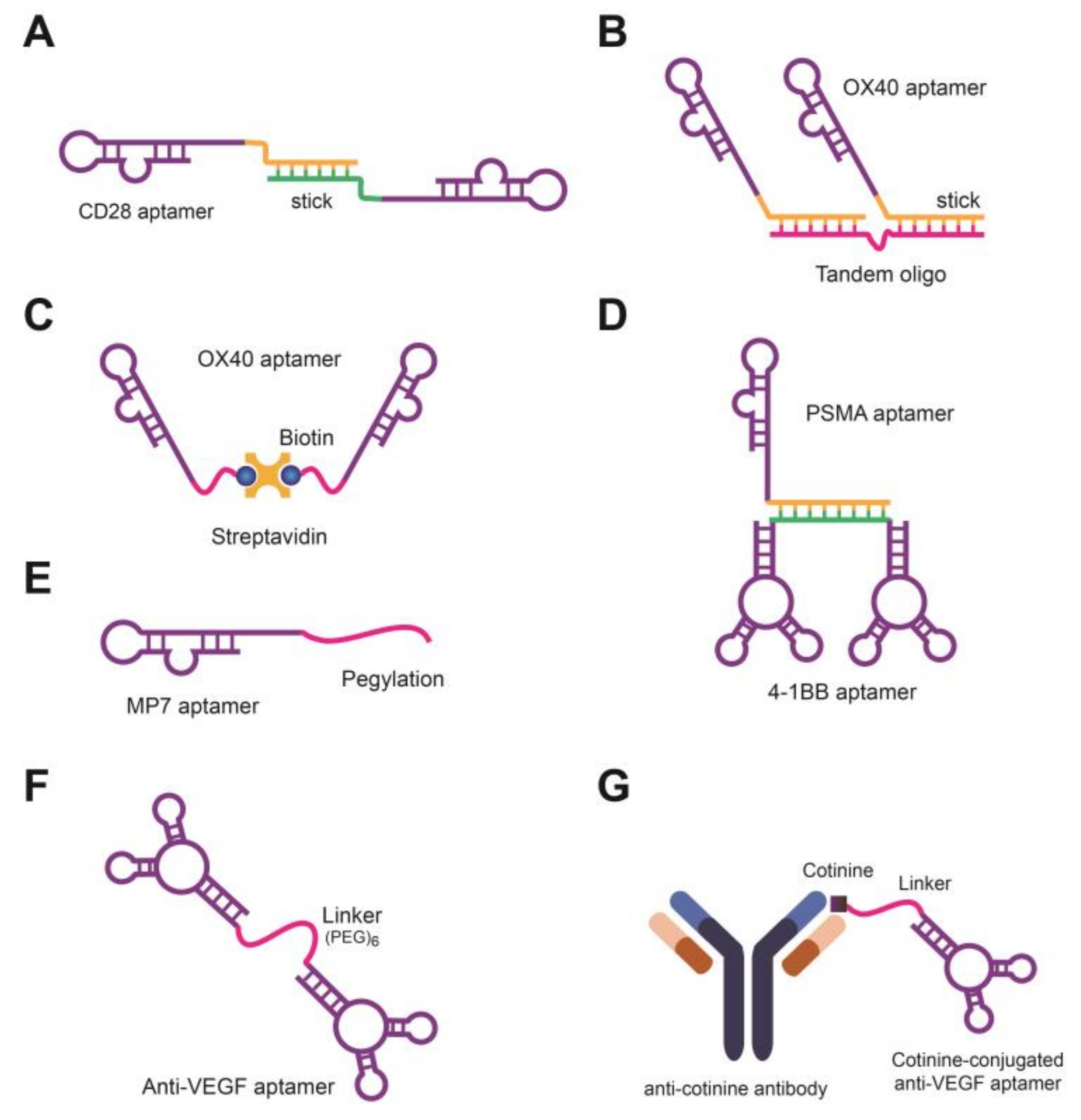
3.1. Aptamers as Cancer Cell Agonists and Antagonists
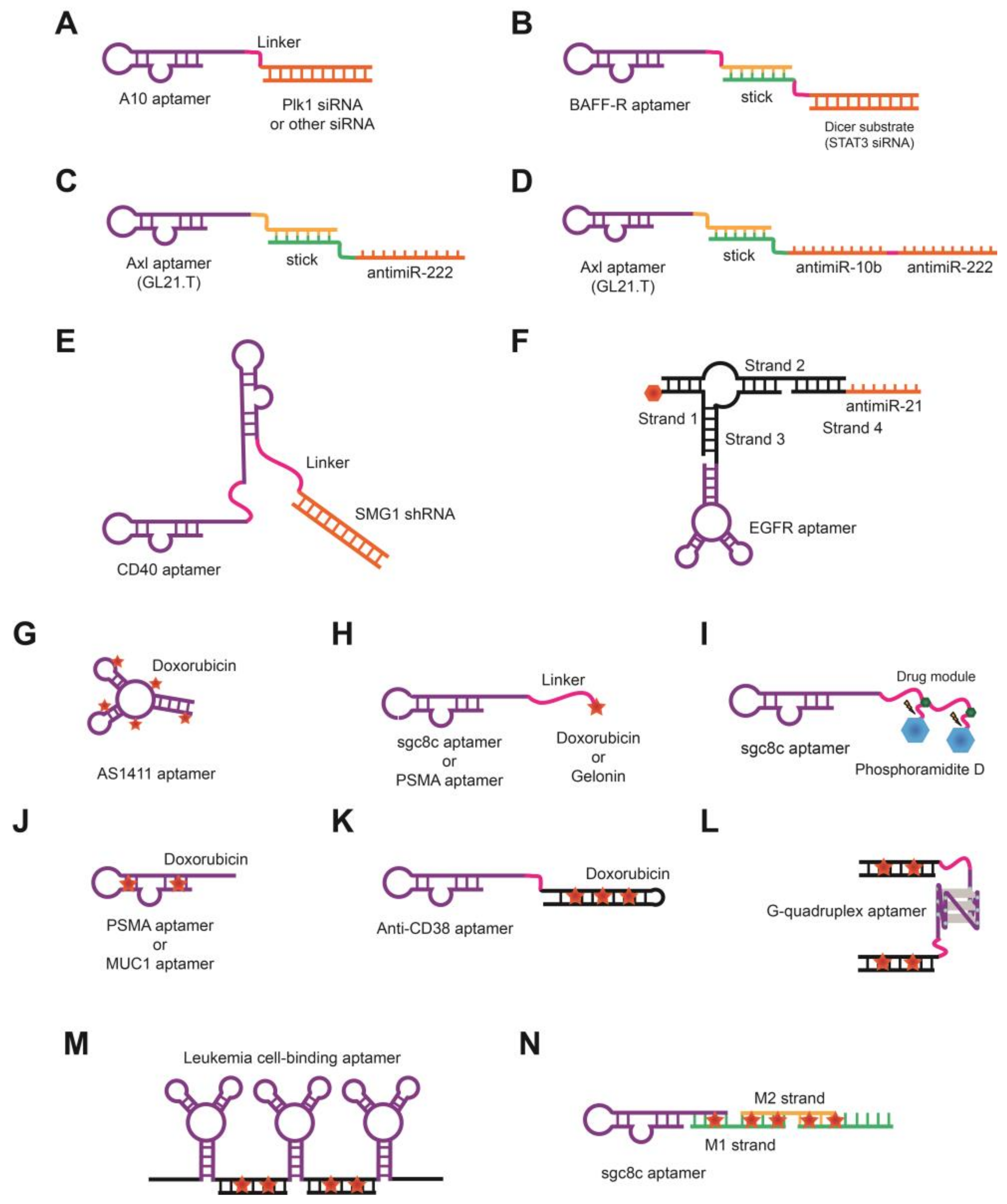
3.2. Aptamer-Drug Conjugates for Targeted Drug Delivery
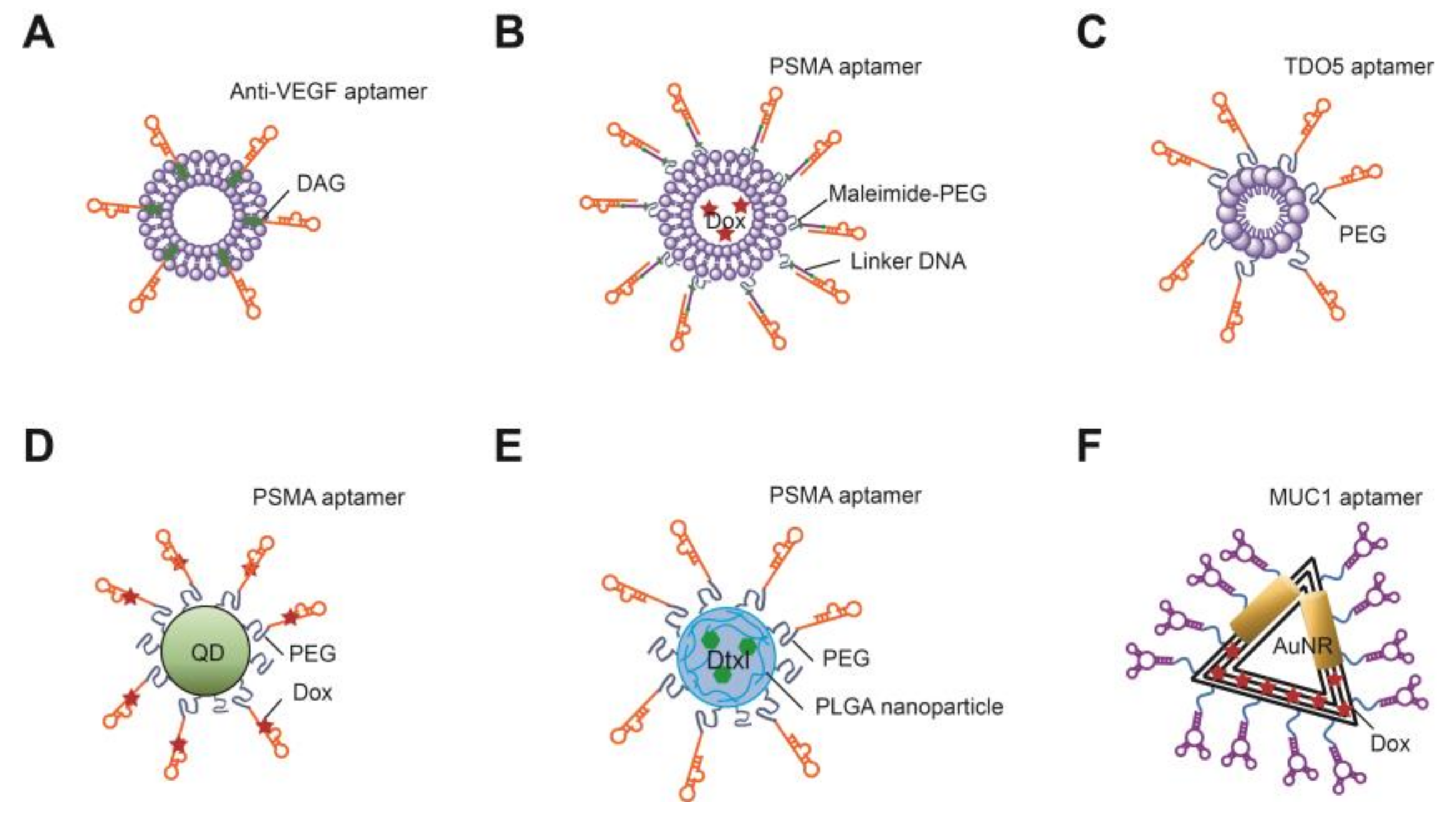
3.3. Aptamer-Conjugated Nano-Vehicles for Targeted Drug Delivery
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| MAPK | mitogen-activated protein kinase |
| PI3K | phosphoinositide 3-kinase |
| PKC | protein kinase C |
| STAT | signal transducer and activator of transcription |
| e-fabp | epidermal fatty acid binding protein |
| EMT | epithelial-to-mesenchymal transition |
References
- Hori, S.; Herrera, A.; Rossi, J.J.; Zhou, J.H. Current advances in aptamers for cancer diagnosis and therapy. Cancers 2018, 10, 9–41. [Google Scholar] [CrossRef] [PubMed]
- Shapira, A.; Livney, Y.D.; Broxterman, H.J.; Assaraf, Y.G. Nanomedicine for targeted cancer therapy: Towards the overcoming of drug resistance. Drug Resist. Updat. 2011, 14, 150–163. [Google Scholar] [CrossRef] [PubMed]
- Chabner, B.A.; Roberts, T.G., Jr. Timeline: Chemotherapy and the war on cancer. Nat. Rev. Cancer 2005, 5, 65–72. [Google Scholar] [CrossRef] [PubMed]
- DeVita, V.T., Jr.; Chu, E. A history of cancer chemotherapy. Cancer Res. 2008, 68, 8643–8653. [Google Scholar] [CrossRef] [PubMed]
- Gerhards, N.M.; Rottenberg, S. New tools for old drugs: Functional genetic screens to optimize current chemtherapy. Drug Resist. Updat. 2018, 36, 30–46. [Google Scholar] [CrossRef] [PubMed]
- Albini, A.; Pennesi, G.; Donatelli, F.; Cammarota, R.; De Flora, S.; Noonan, D.M. Cardiotoxicity of anticancer drugs: The need for cardio-oncology and cardio-oncological prevention. J. Natl. Cancer Inst. 2010, 102, 14–25. [Google Scholar] [CrossRef] [PubMed]
- Bae, Y.H.; Park, K. Targeted drug delivery to tumors: Myths, reality and possibility. J. Control. Release 2011, 153, 198–205. [Google Scholar] [CrossRef] [PubMed]
- Giljohann, D.A.; Seferos, D.S.; Daniel, W.L.; Massich, M.D.; Patel, P.C.; Mirkin, C.A. Gold nanoparticles for biology and medicine. Angew. Chem. Int. Ed. Engl. 2010, 49, 3280–3294. [Google Scholar] [CrossRef] [PubMed]
- Petros, R.A.; DeSimone, J.M. Strategies in the design of nanoparticles for therapeutic applications. Nat. Rev. Drug Discov. 2010, 9, 615–627. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Votruba, A.R.; Farokhzad, O.C.; Langer, R. Nanotechnology in drug delivery and tissue engineering: From discovery to applications. Nano Lett. 2010, 10, 3223–3230. [Google Scholar] [CrossRef] [PubMed]
- Ferrari, M. Cancer nanotechnology: Opportunities and challenges. Nat. Rev. Cancer 2005, 5, 161–171. [Google Scholar] [CrossRef] [PubMed]
- Noble, C.O.; Kirpotin, D.B.; Hayes, M.E.; Mamot, C.; Hong, K.; Park, J.W.; Benz, C.C.; Marks, J.D.; Drummond, D.C. Development of ligand-targeted liposomes for cancer therapy. Expert Opin. Ther. Targets 2004, 8, 335–353. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Zhang, Y.; Chen, C.; Yao, Q.; Li, M. Targeted drug delivery in pancreatic cancer. Biochim. Biophys. Acta 2010, 1805, 97–104. [Google Scholar] [CrossRef] [PubMed]
- Sapra, P.; Allen, T.M. Internalizing antibodies are necessary for improved therapeutic efficacy of antibody-targeted liposomal drugs. Cancer Res. 2002, 62, 7190–7194. [Google Scholar] [PubMed]
- Sapra, P.; Allen, T.M. Ligand-targeted liposomal anticancer drugs. Prog. Lipid Res. 2003, 42, 439–462. [Google Scholar] [CrossRef]
- Cao, Z.H.; Tong, R.; Mishra, A.; Xu, W.C.; Wong, G.C.L.; Cheng, J.J.; Lu, Y. Reversible cell-specific drug delivery with aptamer-functionalized liposomes. Angew. Chem. Int. Ed. 2009, 48, 6494–6498. [Google Scholar] [CrossRef] [PubMed]
- Kirpotin, D.B.; Drummond, D.C.; Shao, Y.; Shalaby, M.R.; Hong, K.L.; Nielsen, U.B.; Marks, J.D.; Benz, C.C.; Park, J.W. Antibody targeting of long-circulating lipidic nanoparticles does not increase tumor localization but does increase internalization in animal models. Cancer Res. 2006, 66, 6732–6740. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.G.; Zhu, X.; Lu, P.Y.; Rosato, R.R.; Tan, W.; Zu, Y.L. Oligonucleotide aptamers: New tools for targeted cancer therapy. Mol. Ther. Nucleic Acids 2014, 3, e182. [Google Scholar] [CrossRef] [PubMed]
- Tesmer, V.M.; Lennarz, S.; Mayer, G.; Tesmer, J.J.G. Molecular mechanism for inhibition of G protein-coupled receptor kinase 2 by a selective RNA aptamer. Structure 2012, 20, 1300–1309. [Google Scholar] [CrossRef] [PubMed]
- Fang, X.H.; Tan, W.H. Aptamers generated from cell-selex for molecular medicine: A chemical biology approach. Acc. Chem. Res. 2010, 43, 48–57. [Google Scholar] [CrossRef] [PubMed]
- Abeydeera, N.D.; Egli, M.; Cox, N.; Mercier, K.; Conde, J.N.; Pallan, P.S.; Mizurini, D.M.; Sierant, M.; Hibti, F.E.; Hassell, T.; et al. Evoking picomolar binding in RNA by a single phosphorodithioate linkage. Nucleic Acids Res. 2016, 44, 8052–8064. [Google Scholar] [CrossRef] [PubMed]
- Ellington, A.D.; Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 1990, 346, 818–822. [Google Scholar] [CrossRef] [PubMed]
- Jayasena, S.D. Aptamers: An emerging class of molecules that rival antibodies in diagnostics. Clin. Chem. 1999, 45, 1628–1650. [Google Scholar] [PubMed]
- Hermann, T.; Patel, D.J. Biochemistry—Adaptive recognition by nucleic acid aptamers. Science 2000, 287, 820–825. [Google Scholar] [CrossRef] [PubMed]
- Sefah, K.; Tang, Z.W.; Shangguan, D.H.; Chen, H.; Lopez-Colon, D.; Li, Y.; Parekh, P.; Martin, J.; Meng, L.; Phillips, J.A.; et al. Molecular recognition of acute myeloid leukemia using aptamers. Leukemia 2009, 23, 235–244. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.W.; Parekh, P.; Turner, P.; Moyer, R.W.; Tan, W.H. Generating aptamers for recognition of virus-infected cells. Clin. Chem. 2009, 55, 813–822. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.W.; Shangguan, D.; Wang, K.M.; Shi, H.; Sefah, K.; Mallikratchy, P.; Chen, H.W.; Li, Y.; Tan, W.H. Selection of aptamers for molecular recognition and characterization of cancer cells. Anal. Chem. 2007, 79, 4900–4907. [Google Scholar] [CrossRef] [PubMed]
- Shangguan, D.; Li, Y.; Tang, Z.W.; Cao, Z.H.C.; Chen, H.W.; Mallikaratchy, P.; Sefah, K.; Yang, C.Y.J.; Tan, W.H. Aptamers evolved from live cells as effective molecular probes for cancer study. Proc. Natl. Acad. Sci. USA 2006, 103, 11838–11843. [Google Scholar] [CrossRef] [PubMed]
- Bunka, D.H.J.; Stockley, P.G. Aptamers come of age—At last. Nat. Rev. Microbiol. 2006, 4, 588–596. [Google Scholar] [CrossRef] [PubMed]
- Dunn, M.R.; Jimenez, R.M.; Chaput, J.C. Analysis of aptamer discovery and technology. Nat. Rev. Chem. 2017, 1, 76–91. [Google Scholar] [CrossRef]
- Chen, T.; Shukoor, M.I.; Chen, Y.; Yuan, Q.A.; Zhu, Z.; Zhao, Z.L.; Gulbakan, B.; Tan, W.H. Aptamer-conjugated nanomaterials for bioanalysis and biotechnology applications. Nanoscale 2011, 3, 546–556. [Google Scholar] [CrossRef] [PubMed]
- Song, K.M.; Lee, S.; Ban, C. Aptamers and their biological applications. Sensors 2012, 12, 612–631. [Google Scholar] [CrossRef] [PubMed]
- Martin, D.F.; Klein, M.; Haller, J.; Adamis, A.; Gragoudas, E.; Miller, J.; Blumenkrantz, M.; Goldberg, M.; Yannuzzi, L.; Henninger, D.; et al. Preclinical and phase 1A clinical evaluation of an anti-VEGF pegylated aptamer (EYE001) for the treatment of exudative age-related macular degeneration. Retina J. Ret. Vit. Dis. 2002, 22, 143–152. [Google Scholar]
- Fish, G.; Haller, J.A.; Ho, A.C.; Klein, M.; Loewenstein, J.; Martin, D.; Orth, D.; Rosen, R.B.; Sanislo, S.; Schwartz, S.D.; et al. Anti-vascular endothelial growth factor therapy for subfoveal choroidal neovascularization secondary to age-related macular degeneration—Phase ii study results. Ophthalmology 2003, 110, 979–986. [Google Scholar]
- Ni, S.J.; Yao, H.Z.; Wang, L.L.; Lu, J.; Jiang, F.; Lu, A.P.; Zhang, G. Chemical modifications of nucleic acid aptamers for therapeutic purposes. Int. J. Mol. Sci. 2017, 18, 1683–1703. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.Y.; Liang, C.; Lv, Q.X.; Li, D.F.; Xu, X.G.; Liu, B.Q.; Lu, A.P.; Zhang, G. Molecular selection, modification and development of therapeutic oligonucleotide aptamers. Int. J. Mol. Sci. 2016, 17, 385–403. [Google Scholar] [CrossRef] [PubMed]
- Zhu, G.Z.; Ye, M.; Donovan, M.J.; Song, E.Q.; Zhao, Z.L.; Tan, W.H. Nucleic acid aptamers: An emerging frontier in cancer therapy. Chem. Commun. 2012, 48, 10472–10480. [Google Scholar] [CrossRef] [PubMed]
- Kourlas, H.; Schiller, D.S. Pegaptanib sodium for the treatment of neovascular age-related macular degeneration: A review. Clin. Ther. 2006, 28, 36–44. [Google Scholar] [CrossRef] [PubMed]
- Ng, E.W.M.; Shima, D.T.; Calias, P.; Cunningham, E.T.; Guyer, D.R.; Adamis, A.P. Pegaptanib, a targeted anti-VEGF aptamer for ocular vascular disease. Nat. Rev. Drug Discov. 2006, 5, 123–132. [Google Scholar] [CrossRef] [PubMed]
- Li, L.Y.; Hou, J.J.; Liu, X.J.; Guo, Y.J.; Wu, Y.; Zhang, L.H.; Yang, Z.J. Nucleolin-targeting liposomes guided by aptamer AS1411 for the delivery of siRNA for the treatment of malignant melanomas. Biomaterials 2014, 35, 3840–3850. [Google Scholar] [CrossRef] [PubMed]
- Soundararajan, S.; Wang, L.; Sridharan, V.; Chen, W.W.; Courtenay-Luck, N.; Jones, D.; Spicer, E.K.; Fernandes, D.J. Plasma membrane nucleolin is a receptor for the anticancer aptamer AS1411 in MV4-11 leukemia cells. Mol. Pharmacol. 2009, 76, 984–991. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, J.E.; Bambury, R.M.; Van Allen, E.M.; Drabkin, H.A.; Lara, P.N., Jr.; Harzstark, A.L.; Wagle, N.; Figlin, R.A.; Smith, G.W.; Garraway, L.A.; et al. A phase II trial of AS1411 (a novel nucleolin-targeted DNA aptamer) in metastatic renal cell carcinoma. Investig. New Drug 2014, 32, 178–187. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Yu, Y.Y.; Jiang, F.; Zhou, J.W.; Li, Y.S.; Liang, C.; Dang, L.; Lu, A.P.; Zhang, G. Development of cell-selex technology and its application in cancer diagnosis and therapy. Int. J. Mol. Sci. 2016, 17, 2079–2092. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.B.; Bassett, S.E.; Li, X.; Luxon, B.A.; Herzog, N.K.; Shope, R.E.; Aronson, J.; Prow, T.W.; Leary, J.F.; Kirby, R.; et al. Construction and selection of bead-bound combinatorial oligonucleoside phosphorothioate and phosphorodithioate aptamer libraries designed for rapid PCR-based sequencing. Nucleic Acids Res. 2002, 30, e132. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Huang, H.; Dong, S.W.; Ge, L.; Zhang, Y. Progress in aptamer-mediated drug delivery vehicles for cancer targeting and its implications in addressing chemotherapeutic challenges. Theranostics 2014, 4, 931–944. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pestourie, C.; Cerchia, L.; Gombert, K.; Aissouni, Y.; Boulay, J.; De Franciscis, V.; Libri, D.; Tavitian, B.; Duconge, F. Comparison of different strategies to select aptamers against a transmembrane protein target. Oligonucleotides 2006, 16, 323–335. [Google Scholar] [CrossRef] [PubMed]
- Chushak, Y.; Stone, M.O. In silico selection of RNA aptamers. Nucleic Acids Res. 2009, 37, e87. [Google Scholar] [CrossRef] [PubMed]
- Ahirwar, R.; Nahar, S.; Aggarwal, S.; Ramachandran, S.; Maiti, S.; Nahar, P. In silico selection of an aptamer to estrogen receptor alpha using computational docking employing estrogen response elements as aptamer-alike molecules. Sci. Rep. 2016, 6, 285–295. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.S.; Gu, M.B. Advances in aptamer screening and small molecule aptasensors. Adv. Biochem. Eng. Biotechnol. 2014, 140, 29–67. [Google Scholar] [PubMed]
- Dupont, D.M.; Larsen, N.; Jensen, J.K.; Andreasen, P.A.; Kjems, J. Characterisation of aptamer-target interactions by branched selection and high-throughput sequencing of selex pools. Nucleic Acids Res. 2015, 43, e139. [Google Scholar] [CrossRef] [PubMed]
- Thiel, W.H.; Thiel, K.W.; Flenker, K.S.; Bair, T.; Dupuy, A.J.; McNamara, J.O., 2nd; Miller, F.J.; Giangrande, P.H. Cell-internalization selex: Method for identifying cell-internalizing RNA aptamers for delivering sirnas to target cells. Methods Mol. Biol. 2015, 1218, 187–199. [Google Scholar] [PubMed]
- Sefah, K.; Shangguan, D.; Xiong, X.L.; O’Donoghue, M.B.; Tan, W.H. Development of DNA aptamers using cell-selex. Nat. Protoc. 2010, 5, 1169–1185. [Google Scholar] [CrossRef] [PubMed]
- Tolle, F.; Wilke, J.; Wengel, J.; Mayer, G. By-product formation in repetitive PCR amplification of DNA libraries during selex. PLoS ONE 2014, 9, e114693. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Drolet, D.W.; MoonMcDermott, L.; Romig, T.S. An enzyme-linked oligonucleotide assay. Nat. Biotechnol. 1996, 14, 1021–1025. [Google Scholar] [CrossRef] [PubMed]
- Nagarkatti, R.; de Araujo, F.F.; Gupta, C.; Debrabant, A. Aptamer based, non-PCR, non-serological detection of chagas disease biomarkers in Trypanosoma cruzi infected mice. PLOS Negl. Trop. Dis. 2014, 8, e2650. [Google Scholar] [CrossRef] [PubMed]
- McNamara, J.O.; Kolonias, D.; Pastor, F.; Mittler, R.S.; Chen, L.P.; Giangrande, P.H.; Sullenger, B.; Gilboa, E. Multivalent 4-1BB binding aptamers costimulate CD8(+) T cells and inhibit tumor growth in mice. J. Clin. Investig. 2008, 118, 376–386. [Google Scholar] [CrossRef] [PubMed]
- Sawyers, C.L. The cancer biomarker problem. Nature 2008, 452, 548–552. [Google Scholar] [CrossRef] [PubMed]
- Goossens, N.; Nakagawa, S.; Sun, X.C.; Hoshida, Y. Cancer biomarker discovery and validation. Transl. Cancer Res. 2015, 4, 256–269. [Google Scholar] [PubMed]
- Rucevic, M.; Hixson, D.; Josic, D. Mammalian plasma membrane proteins as potential biomarkers and drug targets. Electrophoresis 2011, 32, 1549–1564. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, C.S.M.; Matthews, C.S.; Missailidis, S. DNA aptamers that bind to muc1 tumour marker: Design and characterization of muc1-binding single-stranded DNA aptamers. Tumor Biol. 2006, 27, 289–301. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Duan, J.H.; Song, Y.M.; Ma, J.; Wang, F.D.; Lu, X.; Yang, X.D. Novel HER2 aptamer selectively delivers cytotoxic drug to HER2-positive breast cancer cells in vitro. J. Transl. Med. 2012, 148–157. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.H.B.; Chernis, G.A.; Hoang, V.Q.; Landgraf, R. Inhibition of heregulin signaling by an aptamer that preferentially binds to the oligomeric form of human epidermal growth factor receptor-3. Proc. Natl. Acad. Sci. USA 2003, 100, 9226–9231. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.L.; Zhu, Z.; An, Y.; Zhang, W.T.; Zhang, H.M.; Liu, D.; Yu, C.D.; Duan, W.; Yang, C.J. Selection of DNA aptamers against epithelial cell adhesion molecule for cancer cell imaging and circulating tumor cell capture. Anal. Chem. 2013, 85, 4141–4149. [Google Scholar] [CrossRef] [PubMed]
- Girvan, A.C.; Teng, Y.; Casson, L.K.; Thomas, S.D.; Juliger, S.; Ball, M.W.; Klein, J.B.; Pierce, W.M.; Barve, S.S.; Bates, P.J. AGRO100 inhibits activation of nuclear factor-kappa b (NF-κB) by forming a complex with NF-κB essential modulator (NEMO) and nucleolin. Mol. Cancer Ther. 2006, 5, 1790–1799. [Google Scholar] [CrossRef] [PubMed]
- Lupold, S.E.; Hicke, B.J.; Lin, Y.; Coffey, D.S. Identification and characterization of nuclease-stabilized RNA molecules that bind human prostate cancer cells via the prostate-specific membrane antigen. Cancer Res. 2002, 62, 4029–4033. [Google Scholar] [PubMed]
- Shigdar, S.; Lin, J.; Yu, Y.; Pastuovic, M.; Wei, M.; Duan, W. RNA aptamer against a cancer stem cell marker epithelial cell adhesion molecule. Cancer Sci. 2011, 102, 991–998. [Google Scholar] [CrossRef] [PubMed]
- Alshaer, W.; Hillaireau, H.; Vergnaud, J.; Ismail, S.; Fattal, E. Functionalizing liposomes with anti-CD44 aptamer for selective targeting of cancer cells. Bioconjug. Chem. 2015, 26, 1307–1313. [Google Scholar] [CrossRef] [PubMed]
- Prodeus, A.; Abdul-Wahid, A.; Fischer, N.W.; Huang, E.H.B.; Cydzik, M.; Gariepy, J. Targeting the PD-1/PD-l1 immune evasion axis with DNA aptamers as a novel therapeutic strategy for the treatment of disseminated cancers. Mol. Ther. Nucleic Acids 2015, 4, e237. [Google Scholar] [CrossRef] [PubMed]
- Pastor, F.; Kolonias, D.; McNamara, J.O.; Gilboa, E. Targeting 4-1BB costimulation to disseminated tumor lesions with bi-specific oligonucleotide aptamers. Mol. Ther. 2011, 19, 1878–1886. [Google Scholar] [CrossRef] [PubMed]
- Dollins, C.M.; Nair, S.; Boczkowski, D.; Lee, J.; Layzer, J.M.; Gilboa, E.; Sullenger, B.A. Assembling OX40 aptamers on a molecular scaffold to create a receptor-activating aptamer. Chem. Biol. 2008, 15, 675–682. [Google Scholar] [CrossRef] [PubMed]
- Green, L.S.; Jellinek, D.; Jenison, R.; Ostman, A.; Heldin, C.H.; Janjic, N. Inhibitory DNA ligands to platelet-derived growth factor B-chain. Biochemistry 1996, 35, 14413–14424. [Google Scholar] [CrossRef] [PubMed]
- Green, L.S.; Jellinek, D.; Bell, C.; Beebe, L.A.; Feistner, B.D.; Gill, S.C.; Jucker, F.M.; Janjic, N. Nuclease-resistant nucleic-acid ligands to vascular-permeability factor vascular endothelial growth-factor. Chem. Biol. 1995, 2, 683–695. [Google Scholar] [CrossRef]
- Hovanessian, A.G.; Soundaramourty, C.; El Khoury, D.; Nondier, I.; Svab, J.; Krust, B. Surface expressed nucleolin is constantly induced in tumor cells to mediate calcium-dependent ligand internalization. PLoS ONE 2010, 5, e15787. [Google Scholar] [CrossRef] [PubMed]
- Soundararajan, S.; Chen, W.W.; Spicer, E.K.; Courtenay-Luck, N.; Fernandes, D.J. The nucleolin targeting aptamer as1411 destabilizes BCL-2 messenger RNA in human breast cancer cells. Cancer Res. 2008, 68, 2358–2365. [Google Scholar] [CrossRef] [PubMed]
- Xing, H.; Tang, L.; Yang, X.J.; Hwang, K.; Wang, W.D.; Yin, Q.; Wong, N.Y.; Dobrucki, L.W.; Yasui, N.; Katzenellenbogen, J.A.; et al. Selective delivery of an anticancer drug with aptamer-functionalized liposomes to breast cancer cells in vitro and in vivo. J. Mater. Chem. B 2013, 1, 5288–5297. [Google Scholar] [CrossRef] [PubMed]
- Mongelard, F.; Bouvet, P. AS-1411, a guanosine-rich oligonucleotide aptamer targeting nucleolin for the potential treatment of cancer, including acute myeloid leukemia. Curr. Opin. Mol. Ther. 2010, 12, 107–114. [Google Scholar] [PubMed]
- Ristau, B.T.; O’Keefe, D.S.; Bacich, D.J. The prostate-specific membrane antigen: Lessons and current clinical implications from 20 years of research. Urol. Oncol. Semin. Orig. 2014, 32, 272–279. [Google Scholar] [CrossRef] [PubMed]
- Kolishetti, N.; Dhar, S.; Valencia, P.M.; Lin, L.Q.; Karnik, R.; Lippard, S.J.; Langer, R.; Farokhzad, O.C. Engineering of self-assembled nanoparticle platform for precisely controlled combination drug therapy. Proc. Natl. Acad. Sci. USA 2010, 107, 17939–17944. [Google Scholar] [CrossRef] [PubMed]
- Gu, F.; Zhang, L.; Teply, B.A.; Mann, N.; Wang, A.; Radovic-Moreno, A.F.; Langer, R.; Farokhzad, O.C. Precise engineering of targeted nanoparticles by using self-assembled biointegrated block copolymers. Proc. Natl. Acad. Sci. USA 2008, 105, 2586–2591. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Jeong, Y.Y.; Jon, S. A drug-loaded aptamer-gold nanoparticle bioconjugate for combined CT imaging and therapy of prostate cancer. ACS Nano 2010, 4, 3689–3696. [Google Scholar] [CrossRef] [PubMed]
- Baek, S.E.; Lee, K.H.; Park, Y.S.; Oh, D.K.; Oh, S.; Kim, K.S.; Kim, D.E. RNA aptamer-conjugated liposome as an efficient anticancer drug delivery vehicle targeting cancer cells in vivo. J. Control. Release 2014, 196, 234–242. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.J.; Cheng, F.F.; Zheng, T.T.; Zhu, J.J. Versatile aptasensor for electrochemical quantification of cell surface glycan and naked-eye tracking glycolytic inhibition in living cells. Biosens. Bioelectron. 2017, 89, 937–945. [Google Scholar] [CrossRef] [PubMed]
- Shen, C.C.; Zeng, K.; Luo, J.J.; Li, X.Q.; Yang, M.H.; Rasooly, A. Self-assembled DNA generated electric current biosensor for HER2 analysis. Anal. Chem. 2017, 89, 10264–10269. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Xiang, D.X.; Shigdar, S.; Yang, W.R.; Li, Q.; Lin, J.; Liu, K.X.; Duan, W. Epithelial cell adhesion molecule aptamer functionalized PLGA-lecithin-curcumin-peg nanoparticles for targeted drug delivery to human colorectal adenocarcinoma cells. Int. J. Nanomed. 2014, 9, 1083–1096. [Google Scholar]
- Huang, D.B.; Vu, D.; Cassiday, L.A.; Zimmerman, J.M.; Maher, L.J.; Ghosh, G. Crystal structure of NF-κB (p50)(2) complexed to a high-affinity RNA aptamer. Proc. Natl. Acad. Sci. USA 2003, 100, 9268–9273. [Google Scholar] [CrossRef] [PubMed]
- Tamboli, V.K.; Bhalla, N.; Jolly, P.; Bowen, C.R.; Taylor, J.T.; Bowen, J.L.; Allender, C.J.; Estrela, P. Hybrid synthetic receptors on MOSFET devices for detection of prostate specific antigen in human plasma. Anal. Chem. 2016, 88, 11486–11490. [Google Scholar] [CrossRef] [PubMed]
- Heydari-Bafrooei, E.; Shamszadeh, N.S. Electrochemical bioassay development for ultrasensitive aptasensing of prostate specific antigen. Biosens. Bioelectron. 2017, 91, 284–292. [Google Scholar] [CrossRef] [PubMed]
- Somasunderam, A.; Thiviyanathan, V.; Tanaka, T.; Li, X.; Neerathilingam, M.; Lokesh, G.L.R.; Mann, A.; Peng, Y.; Ferrari, M.; Klostergaard, J.; et al. Combinatorial selection of DNA thioaptamers targeted to the HA binding domain of human CD44. Biochemistry 2010, 49, 9106–9112. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.C.; Huang, Y.F.; Cao, Z.H.; Tan, W.H.; Chang, H.T. Aptamer-modified gold nanoparticles for colorimetric determination of platelet-derived growth factors and their receptors. Anal. Chem. 2005, 77, 5735–5741. [Google Scholar] [CrossRef] [PubMed]
- Willis, M.C.; Collins, B.D.; Zhang, T.; Green, L.S.; Sebesta, D.P.; Bell, C.; Kellogg, E.; Gill, S.C.; Magallanez, A.; Knauer, S.; et al. Liposome-anchored vascular endothelial, growth factor aptamers. Bioconjug. Chem. 1998, 9, 633. [Google Scholar] [CrossRef]
- Nonaka, Y.; Sode, K.; Ikebukuro, K. Screening and improvement of an anti-VEGF DNA aptamer. Molecules 2010, 15, 215–225. [Google Scholar] [CrossRef] [PubMed]
- Nonaka, Y.; Yoshida, W.; Abe, K.; Ferri, S.; Schulze, H.; Bachmann, T.T.; Ikebukuro, K. Affinity improvement of a VEGF aptamer by in silico maturation for a sensitive VEGF-detection system. Anal. Chem. 2013, 85, 1132–1137. [Google Scholar] [CrossRef] [PubMed]
- Ashrafuzzaman, M. Aptamers as both drugs and drug-carriers. Biomed. Res. Int. 2014, 2014, 21–42. [Google Scholar] [CrossRef] [PubMed]
- Pastor, F.; Soldevilla, M.M.; Villanueva, H.; Kolonias, D.; Inoges, S.; de Cerio, A.L.; Kandzia, R.; Klimyuk, V.; Gleba, Y.; Gilboa, E.; et al. CD28 aptamers as powerful immune response modulators. Mol. Ther. Nucleic Acids 2013, 2, e98. [Google Scholar] [CrossRef] [PubMed]
- Pratico, E.D.; Sullenger, B.A.; Nair, S.K. Identification and characterization of an agonistic aptamer against the T cell costimulatory receptor, OX40. Nucleic Acid Ther. 2013, 23, 35–43. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.P.; Han, X. Anti-PD-1/PD-l1 therapy of human cancer: Past, present, and future. J. Clin. Investig. 2015, 125, 3384–3391. [Google Scholar] [CrossRef] [PubMed]
- Akiyama, H.; Kachi, S.; Silva, R.L.E.; Umeda, N.; Hackett, S.F.; McCauley, D.; McCauley, T.; Zoltoski, A.; Epstein, D.M.; Campochiaro, P.A. Intraocular injection of an aptamer that binds PDGF-B: A potential treatment for proliferative retinopathies. J. Cell. Physiol. 2006, 207, 407–412. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Canny, M.D.; De Erkenez, A.; Krilleke, D.; Ng, Y.S.; Shima, D.T.; Pardi, A.; Jucker, F. A therapeutic aptamer inhibits angiogenesis by specifically targeting the heparin binding domain of VEGF165. Proc. Natl. Acad. Sci. USA 2005, 102, 18902–18907. [Google Scholar] [CrossRef] [PubMed]
- Heo, K.; Min, S.W.; Sung, H.J.; Kim, H.G.; Kim, H.J.; Kim, Y.H.; Choi, B.K.; Han, S.; Chung, S.; Lee, E.S.; et al. An aptamer-antibody complex (oligobody) as a novel delivery platform for targeted cancer therapies. J. Control. Release 2016, 229, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Dassie, J.P.; Liu, X.Y.; Thomas, G.S.; Whitaker, R.M.; Thiel, K.W.; Stockdale, K.R.; Meyerholz, D.K.; McCaffrey, A.P.; McNamara, J.O.; Giangrande, P.H. Systemic administration of optimized aptamer-siRNA chimeras promotes regression of PSMA-expressing tumors. Nat. Biotechnol. 2009, 27, 839–849. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.H.; Tiemann, K.; Chomchan, P.; Alluin, J.; Swiderski, P.; Burnett, J.; Zhang, X.Z.; Forman, S.; Chen, R.; Rossi, J. Dual functional BAFF receptor aptamers inhibit ligand-induced proliferation and deliver siRNAs to NHL cells. Nucleic Acids Res. 2013, 41, 4266–4283. [Google Scholar] [CrossRef] [PubMed]
- Catuogno, S.; Rienzo, A.; Di Vito, A.; Esposito, C.L.; de Franciscis, V. Selective delivery of therapeutic single strand antimirs by aptamer-based conjugates. J. Control. Release 2015, 210, 147–159. [Google Scholar] [CrossRef] [PubMed]
- Soldevilla, M.M.; Villanueva, H.; Bendandi, M.; Inoges, S.; de Cerio, A.L.D.; Pastor, F. 2-fluoro-RNA oligonucleotide CD40 targeted aptamers for the control of B lymphoma and bone-marrow aplasia. Biomaterials 2015, 67, 274–285. [Google Scholar] [CrossRef] [PubMed]
- Shu, D.; Li, H.; Shu, Y.; Xiong, G.F.; Carson, W.E.; Haque, F.; Xu, R.; Guo, P.X. Systemic delivery of anti-mirna for suppression of triple negative breast cancer utilizing RNA nanotechnology. ACS Nano 2015, 9, 9731–9740. [Google Scholar] [CrossRef] [PubMed]
- Le Trinh, T.; Zhu, G.Z.; Xiao, X.L.; Puszyk, W.; Sefah, K.; Wu, Q.F.; Tan, W.H.; Liu, C. A synthetic aptamer-drug adduct for targeted liver cancer therapy. PLoS ONE 2015, 10, e0136673. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.F.; Shangguan, D.H.; Liu, H.P.; Phillips, J.A.; Zhang, X.L.; Chen, Y.; Tan, W.H. Molecular assembly of an aptamer-drug conjugate for targeted drug delivery to tumor cells. ChemBioChem 2009, 10, 862–868. [Google Scholar] [CrossRef] [PubMed]
- Chu, T.C.; Marks, J.W.; Lavery, L.A.; Faulkner, S.; Rosenblum, M.G.; Ellington, A.D.; Levy, M. Aptamer: Toxin conjugates that specifically target prostate tumor cells. Cancer Res. 2006, 66, 5989–5992. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.W.; Zhu, G.Z.; Mei, L.; Xie, Y.; Ma, H.B.; Ye, M.; Qing, F.L.; Tan, W.H. Automated modular synthesis of aptamer-drug conjugates for targeted drug delivery. J. Am. Chem. Soc. 2014, 136, 2731–2734. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.H.; Rossi, J.J. Cell-type-specific, aptamer-functionalized agents for targeted disease therapy. Mol. Ther. Nucleic Acids 2014, 3, e169. [Google Scholar] [CrossRef] [PubMed]
- Bagalkot, V.; Farokhzad, O.C.; Langer, R.; Jon, S. An aptamer-doxorubicin physical conjugate as a novel targeted drug-delivery platform. Angew. Chem. Int. Ed. 2006, 45, 8149–8152. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Duan, J.H.; Zhan, Q.M.; Wang, F.D.; Lu, X.; Yang, X.D. Novel MUC1 aptamer selectively delivers cytotoxic agent to cancer cells in vitro. PLoS ONE 2012, 7, e31970. [Google Scholar] [CrossRef] [PubMed]
- Wen, J.; Tao, W.; Hao, S.; Iyer, S.P.; Zu, Y. A unique aptamer-drug conjugate for targeted therapy of multiple myeloma. Leukemia 2016, 30, 987–991. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Wei, T.; Zhao, J.; Huang, Y.Y.; Deng, H.; Kumar, A.; Wang, C.X.; Liang, Z.C.; Ma, X.W.; Liang, X.J. Multifunctional aptamer-based nanoparticles for targeted drug delivery to circumvent cancer resistance. Biomaterials 2016, 91, 44–56. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.Q.; Ali, M.M.; Eckert, M.A.; Kang, D.K.; Chen, Y.Y.; Sender, L.S.; Fruman, D.A.; Zhao, W.A. A polyvalent aptamer system for targeted drug delivery. Biomaterials 2013, 34, 9728–9735. [Google Scholar] [CrossRef] [PubMed]
- Zhu, G.Z.; Zheng, J.; Song, E.Q.; Donovan, M.; Zhang, K.J.; Liu, C.; Tan, W.H. Self-assembled, aptamer-tethered DNA nanotrains for targeted transport of molecular drugs in cancer theranostics. Proc. Natl. Acad. Sci. USA 2013, 110, 7998–8003. [Google Scholar] [CrossRef] [PubMed]
- Agudelo, D.; Bourassa, P.; Berube, G.; Tajmir-Riahi, H.A. Intercalation of antitumor drug doxorubicin and its analogue by DNA duplex: Structural features and biological implications. Int. J. Biol. Macromol. 2014, 66, 144–150. [Google Scholar] [CrossRef] [PubMed]
- Park, H.; Kim, D.M.; Baek, S.E.; Kim, K.S.; Kim, D.E. Comparison of drug delivery efficiency between doxorubicin intercalated in RNA aptamer and one encapsulated in RNA aptamer-conjugated liposome. Bull. Korean Chem. Soc. 2015, 36, 2494–2500. [Google Scholar] [CrossRef]
- Wu, Y.R.; Sefah, K.; Liu, H.P.; Wang, R.W.; Tan, W.H. DNA aptamer-micelle as an efficient detection/delivery vehicle toward cancer cells. Proc. Natl. Acad. Sci. USA 2010, 107, 5–10. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.F.; Lin, Y.W.; Lin, Z.H.; Chang, H.T. Aptamer-modified gold nanoparticles for targeting breast cancer cells through light scattering. J. Nanopart. Res. 2009, 11, 775–783. [Google Scholar] [CrossRef]
- Bagalkot, V.; Zhang, L.; Levy-Nissenbaum, E.; Jon, S.; Kantoff, P.W.; Langer, R.; Farokhzad, O.C. Quantum dot—Aptamer conjugates for synchronous cancer imaging, therapy, and sensing of drug delivery based on bi-fluorescence resonance energy transfer. Nano Lett. 2007, 7, 3065–3070. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.C.; Hu, Y.; Duan, J.H.; Yuan, W.; Wang, C.; Xu, H.Y.; Yang, X.D. Novel aptamer-nanoparticle bioconjugates enhances delivery of anticancer drug to MUC1-positive cancer cells in vitro. PLoS ONE 2011, 6, e24077. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.N.; Zhu, X.S.; Gao, J.; Liu, D.K.; Dong, C.X.; Jin, X. PLGA nanoparticles with CD133 aptamers for targeted delivery and sustained release of propranolol to hemangioma. Nanomed. Nanotechnol. 2017, 12, 2611–2624. [Google Scholar] [CrossRef] [PubMed]
- Zhu, G.Z.; Hu, R.; Zhao, Z.L.; Chen, Z.; Zhang, X.B.; Tan, W.H. Noncanonical self-assembly of multifunctional DNA nanoflowers for biomedical applications. J. Am. Chem. Soc. 2013, 135, 16438–16445. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.C.; Han, D.; Chen, T.; Peng, L.; Zhu, G.Z.; You, M.X.; Qiu, L.P.; Sefah, K.; Zhang, X.B.; Tan, W.H. Building a multifunctional aptamer-based DNA nanoassembly for targeted cancer therapy. J. Am. Chem. Soc. 2013, 135, 18644–18650. [Google Scholar] [CrossRef] [PubMed]
- Lu, Z.S.; Wang, Y.; Xu, D.; Pang, L. Aptamer-tagged DNA origami for spatially addressable detection of aflatoxin B1. Chem. Commun. 2017, 53, 941–944. [Google Scholar] [CrossRef] [PubMed]
- Song, L.L.; Jiang, Q.; Liu, J.B.; Li, N.; Liu, Q.; Dai, L.R.; Gao, Y.; Liu, W.L.; Liu, D.S.; Ding, B.Q. DNA origami/gold nanorod hybrid nanostructures for the circumvention of drug resistance. Nanoscale 2017, 9, 7750–7754. [Google Scholar] [CrossRef] [PubMed]
- Bhatia, S.; Frangioni, J.V.; Hoffman, R.M.; Iafrate, A.J.; Polyak, K. The challenges posed by cancer heterogeneity. Nat. Biotechnol. 2012, 30, 604–610. [Google Scholar] [CrossRef] [PubMed]
- Denison, T.A.; Bae, Y.H. Tumor heterogeneity and its implication for drug delivery. J. Control. Release 2012, 164, 187–191. [Google Scholar] [CrossRef] [PubMed]
- Wilting, R.H.; Dannenberg, J.H. Epigenetic mechanisms in tumorigenesis, tumor cell heterogeneity and drug resistance. Drug Resist. Updat. 2012, 15, 21–38. [Google Scholar] [CrossRef] [PubMed]
- Junttila, M.R.; de Sauvage, F.J. Influence of tumour micro-environment heterogeneity on therapeutic response. Nature 2013, 501, 346–354. [Google Scholar] [CrossRef] [PubMed]
- Hoey, T. Drug resistance, epigenetics, and tumor cell heterogeneity. Sci. Transl. Med. 2010, 2. [Google Scholar] [CrossRef] [PubMed]
- Min, K.; Jo, H.; Song, K.; Cho, M.; Chun, Y.S.; Jon, S.; Kim, W.J.; Ban, C. Dual-aptamer-based delivery vehicle of doxorubicin to both PSMA (+) and PSMA (−) prostate cancers. Biomaterials 2011, 32, 2124–2132. [Google Scholar] [CrossRef] [PubMed]
- Jo, H.; Her, J.; Ban, C. Dual aptamer-functionalized silica nanoparticles for the highly sensitive detection of breast cancer. Biosens. Bioelectron. 2015, 71, 129–136. [Google Scholar] [CrossRef] [PubMed]
- Taghdisi, S.M.; Danesh, N.M.; Ramezani, M.; Lavaee, P.; Jalalian, S.H.; Robati, R.Y.; Abnous, K. Double targeting and aptamer-assisted controlled release delivery of epirubicin to cancer cells by aptamers-based dendrimer in vitro and in vivo. Eur. J. Pharm. Biopharm. 2016, 102, 152–158. [Google Scholar] [CrossRef] [PubMed]
- Vermeulen, L.; Melo, F.D.E.; Richel, D.J.; Medema, J.P. The developing cancer stem-cell model: Clinical challenges and opportunities. Lancet Oncol. 2012, 13, e83–e89. [Google Scholar] [CrossRef]
Sample Availability: Samples of the compounds are not available from the authors. |
| Target | Known Expressing Cancer Type | Therapeutic Applications | Aptamer/Type (nts) | Reference |
|---|---|---|---|---|
| MUC1 (mucin 1) | ovarian, breast, lung, pancreatic cancers, multiple myeloma etc. | Prevent cancer cell invasion through beta catenin | S2.1/DNA (25) | [60] |
| Apt/DNA (25) | [82] | |||
| HER2 (human epidermal growth factor 2) | Breast, gastric, lung, colorectal, esophageal, ovarian cancers, etc. | Inhibition of tumorigenic signaling via MAPK, PI3K, PKC and STAT pathways | HB5/DNA (86) | [61] |
| Apt/DNA (31) | [83] | |||
| HER3 (human epidermal growth factor receptor 3) | Breast, lung, gastric, prostate, ovarian, pancreatic cancers etc. | Reduction of drug resistance in HER2+ cancer | A30/RNA (49) | [62] |
| EpCAM (epithelial cell adhesion molecule) | Bladder, breast, colon, lung, ovarian, pancreas, prostate cancers, etc. | Regulate gene expression of c-myc, e-fabp, cyclin, and modulate EMT | SYL3/DNA (80) | [63] |
| Apt/RNA (18) | [84] | |||
| EpDT3-DY647/RNA (19) | [66] | |||
| NF-kB (nuclear factor kappa-light-chain-enhancer of activated B cells) | Cervical, prostate, lung, breast cancers, etc. | Inhibit the genes that control cell proliferation and cell survival | Apt/RNA (29) | [85] |
| ARGO100/DNA (26) | [64] | |||
| PSMA (prostate specific membrane antigen) | Prostate, kidney, bladder cancers, etc. | Prevent hydrolysis of N-acetylaspartyl-glutamate for over-proliferation | xPSM-A10/RNA (40) | [64,78] |
| Apt/DNA (32) | [86,87] | |||
| CD44 | Breast, prostate, and cancer stem cells, etc. | Inhibit cell proliferation, differentiation, migration, and angiogenesis | TA1/DNA (30) | [67,88] |
| PD-1 (programmed death-1) | Colon cancer, carcinoma, etc. | Inhibiting immune response to cancer cells | MP5, MP7/DNA (75) | [68] |
| CD137 (4-1BB) | Prostate cancer etc. | Stimulating immune response to cancer | PSMA-4-1BB/RNA (293) | [69] |
| CD134 (OX40) | Melanoma tumor etc. | Stimulating immune response to cancer | Aptamer 9.8/RNA (80) | [70] |
| PDGF (platelet derived growth factor) | Ovarian, breast, thyroid, cervical, lung cancers, etc. | Inhibit tumor angiogenesis and development | 36t/DNA (39) | [71,89] |
| VEGF (vascular endothelial growth factor | Breast, brain, lung, colon, gastric, pancreatic, melanoma, myeloid, leukemia, etc. | Prevent neovascularization | NX-191/RNA (24) | [72] |
| NX-213/RNA (24) | [90] | |||
| Vap7, V7t1/DNA (25) | [91,92] | |||
| NCL (Nucleolin) | Leukemia, gastric, breast cancers etc. | Induce bcl-2 mRNA instability | AS1411/DNA (26) | [74] |
| Type | Name of Aptamer Drugs | Function in Cancer Therapy | Reference |
|---|---|---|---|
| Agonist & Antagonist | CD28Apt | Either reducing the T-cell tolerance by blocking the interaction with B7 or enhancing the vaccine-induced immune response | [94] |
| OX40 aptamer | Stimulating the T cell proliferation and cytokine production | [70,95] | |
| PSMA-4-1BB aptamer | Promoting the survival and expansion of activated CD8+ T cells | [69] | |
| PEG-MP7 | Inhibiting the PD-L1-mediated suppression of IL-2 secretion in T cells | [68] | |
| NX1838 aptamer | Binding to VEGF165 with high affinity and preventing blood vessel growth and arresting the progression | [98] | |
| Cot-pega oligobody | Inhibiting the Akt pathway that induces the cell survival, angiogenesis, differentiation, cell growth, proliferation | [99] | |
| Aptamer-drug conjugates | A10-Plk1 | Suppressing the expression of polo-like kinase 1 that pro-survival genes | [100] |
| BAFF-R-STAT3 siRNA | Blocking the BAFF-mediated proliferation of B-cell malignancies and suppressing the transcription factor STAT3 to inhibit the cell cycle progression, angiogenesis and tumor cell evasion of immune system | [101] | |
| GL21.T-222 | Inhibiting the receptor tyrosine kinases Axl and PDGFR β and reducing the level of miR-222 or miR-10b | [102] | |
| CD40-SMG1-shRNA chimera | Inhibiting SMG1 kinase that is essential for nonsense mRNA mediated decay initiation in tumor cells | [103] | |
| 3WJ-EGFRapt/anti-miR-21 | Inhibiting of tumor progression, invasion, and metastasis by suppressing of miR-21 | [104] | |
| AS1411-Dox | Inhibiting of tumor cell proliferation by inducing G2/M arrest | [105] | |
| Sgc8c-Dox | Recognizing the protein tyrosine kinase 7 and delivering Dox to the target CCRF-CEM (T-cell Acute Lymphoblastic Leukemia) cells | [106] | |
| ApDCs | Recognizing target cancer cells and release the Fluorouracil in a photocontrollable manner | [108] | |
| MA3 Apt-Dox | Selectively delivering the cytotoxic agent doxorubicin to MUC1-positivie adenocarcinomas cancer cells | [111] | |
| ApDC | Delivering the Dox to CD38-positive m1ultiple myeloma tumor cells and intracellular release of a high drug payload under a pH-controlled mechanism | [112] | |
| ApS&Dox | Targeting nucleolin molecule and circumventing Dox resistance by cell cycle arrest in S phase, effectively increased cell uptake | [113] | |
| Poly-Aptamer-Drug | Targeting and killing leukemia cells due to enhanced binding affinity and cell internalization via multivalent effects | [114] | |
| aptNTrs | Targeting human T-cell acute lymphocytic leukemia with high payload of drugs | [115] | |
| Aptamer-conjugated nano-vehicles | DAG-NX213-L | Inhibiting the VEGF-induced endothelial cell proliferation and vascular permeability increase and angiogenesis | [90] |
| Aptamosome | Selectively delivering the drug to PSMA-positive prostate cancer cells by Dox-encapsulating liposome conjugated with aptamers | [81] | |
| TDO5-micelle | Efficient delivering the drug to target cancer cells by aptamer-micelle assembly with high sensitivity and specificity in flow channel system | [118] | |
| QD-Apt | Delivering Dox to the prostate cancer cells and imaging the cancer cells by quantum dot | [120] | |
| NP-Apt | Suppressing the metastatic cancer progression and inducing the apoptosis of cancer cells | [79] | |
| MUC-1 Origami-Dox-AuNRs | Chemotherapeutically and photothermally killing the MUC1-overexpressed multidrug resistant breast cancer cells | [126] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kim, M.; Kim, D.-M.; Kim, K.-S.; Jung, W.; Kim, D.-E. Applications of Cancer Cell-Specific Aptamers in Targeted Delivery of Anticancer Therapeutic Agents. Molecules 2018, 23, 830. https://doi.org/10.3390/molecules23040830
Kim M, Kim D-M, Kim K-S, Jung W, Kim D-E. Applications of Cancer Cell-Specific Aptamers in Targeted Delivery of Anticancer Therapeutic Agents. Molecules. 2018; 23(4):830. https://doi.org/10.3390/molecules23040830
Chicago/Turabian StyleKim, Minhee, Dong-Min Kim, Keun-Sik Kim, Woong Jung, and Dong-Eun Kim. 2018. "Applications of Cancer Cell-Specific Aptamers in Targeted Delivery of Anticancer Therapeutic Agents" Molecules 23, no. 4: 830. https://doi.org/10.3390/molecules23040830
APA StyleKim, M., Kim, D.-M., Kim, K.-S., Jung, W., & Kim, D.-E. (2018). Applications of Cancer Cell-Specific Aptamers in Targeted Delivery of Anticancer Therapeutic Agents. Molecules, 23(4), 830. https://doi.org/10.3390/molecules23040830





