Proteomics Analysis Reveals an Important Role for the PPAR Signaling Pathway in DBDCT-Induced Hepatotoxicity Mechanisms
Abstract
:1. Introduction
2. Results
2.1. Histopathology
2.2. Differentially Expressed Proteins in Rat Livers Exposed to DBDCT
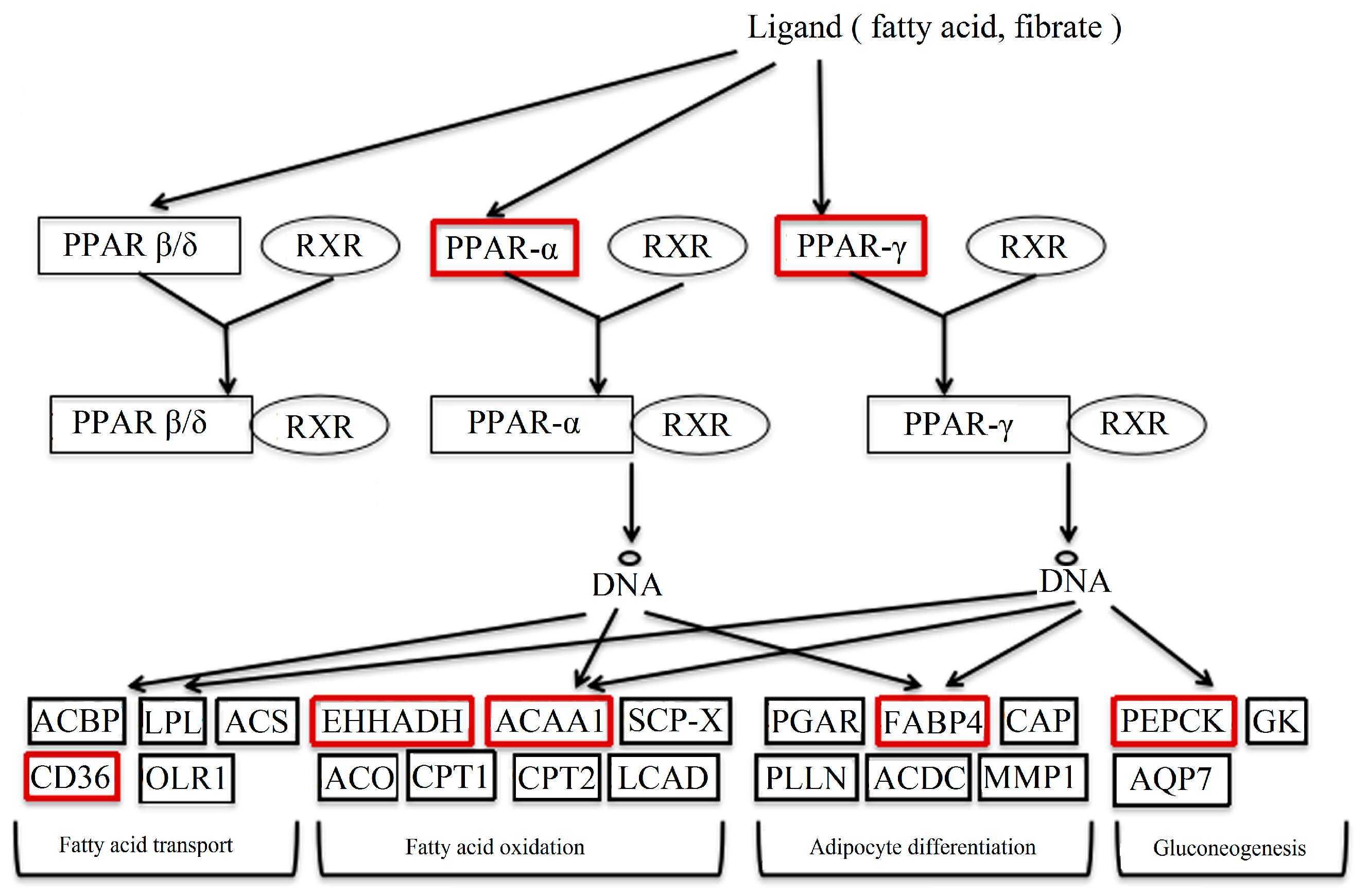
2.3. Pathway Data
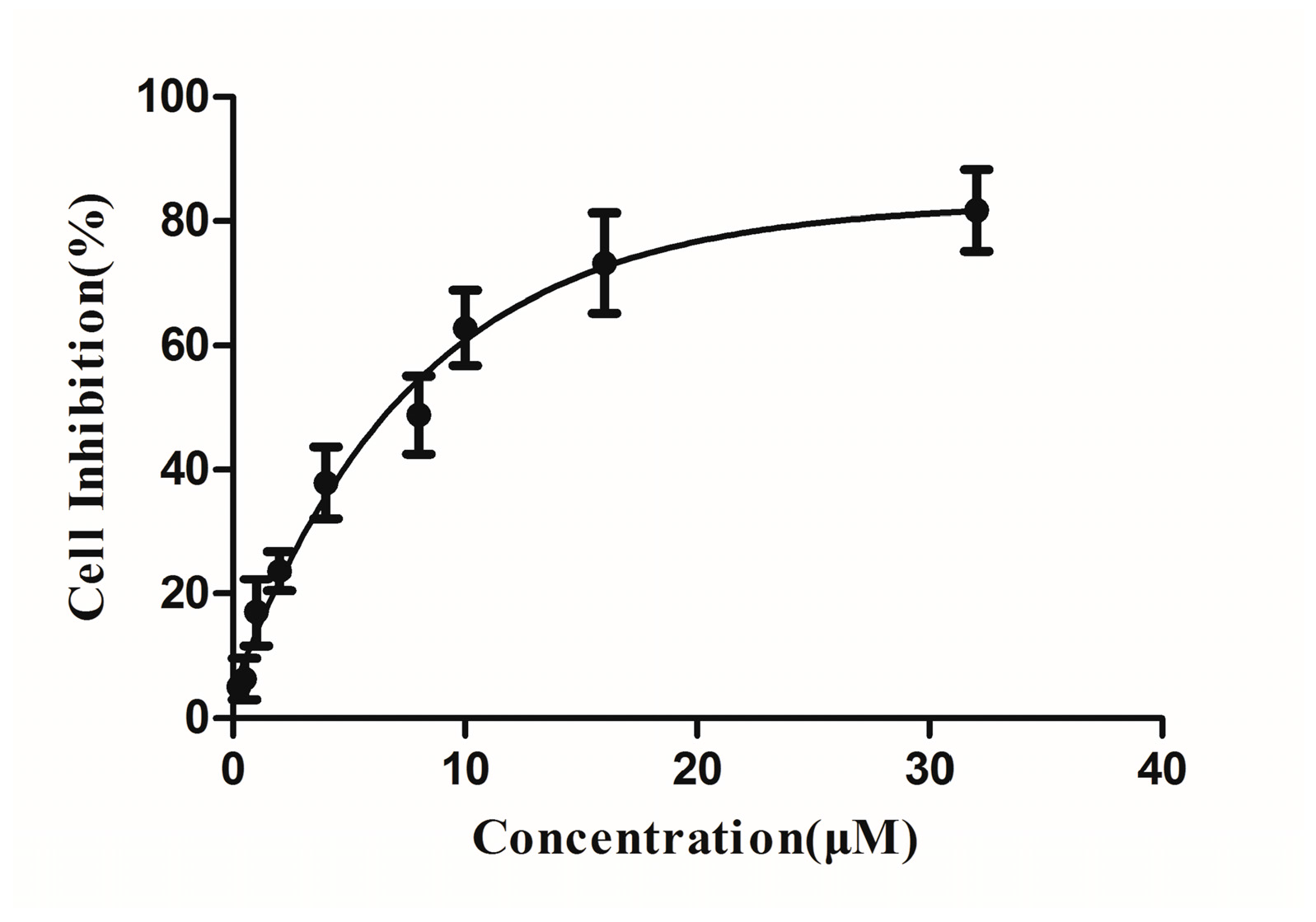
2.4. Cytotoxicities of DBDCT
2.5. DBDCT-Induced Apoptosis Mediated by Caspase Activation
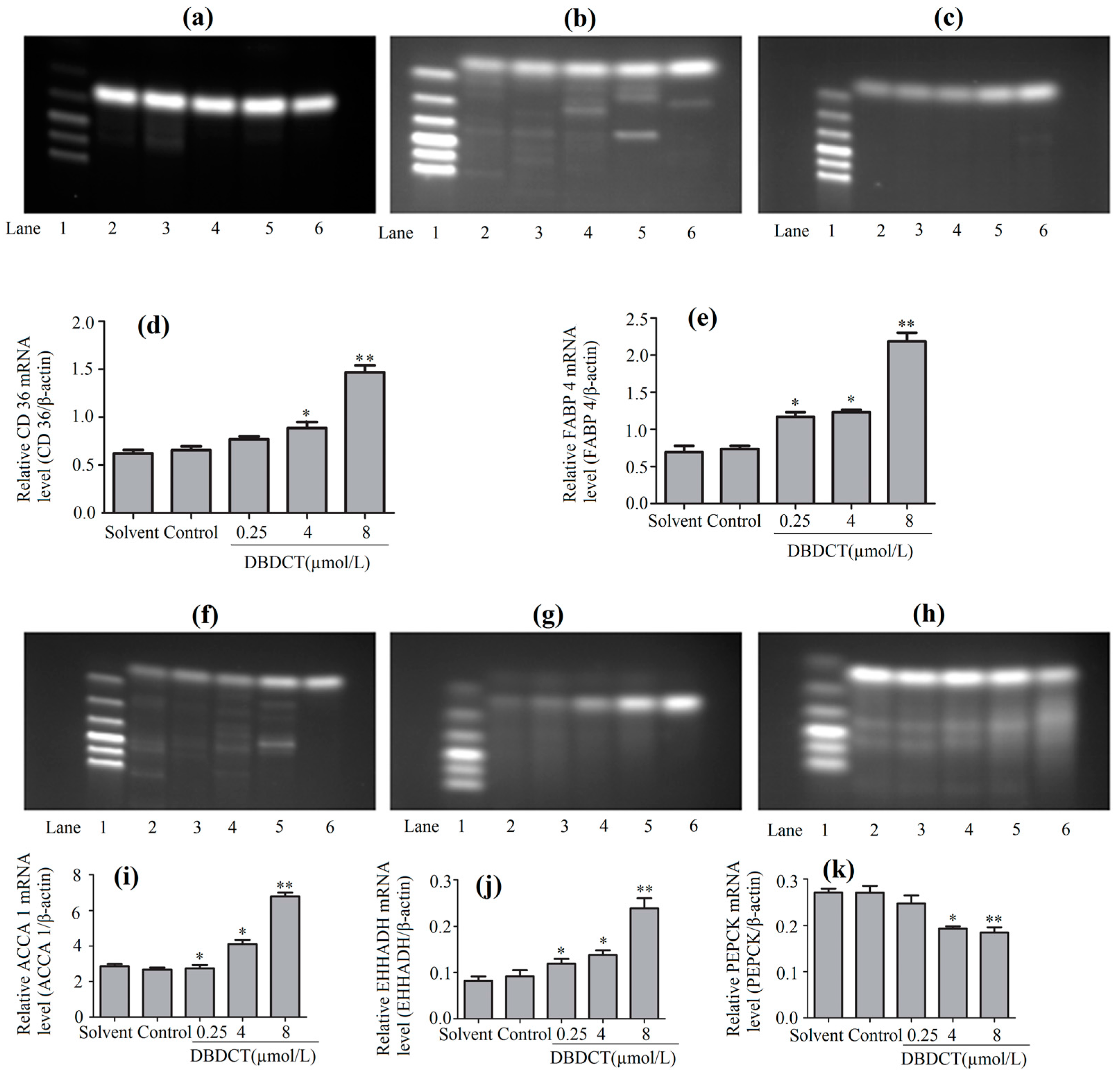
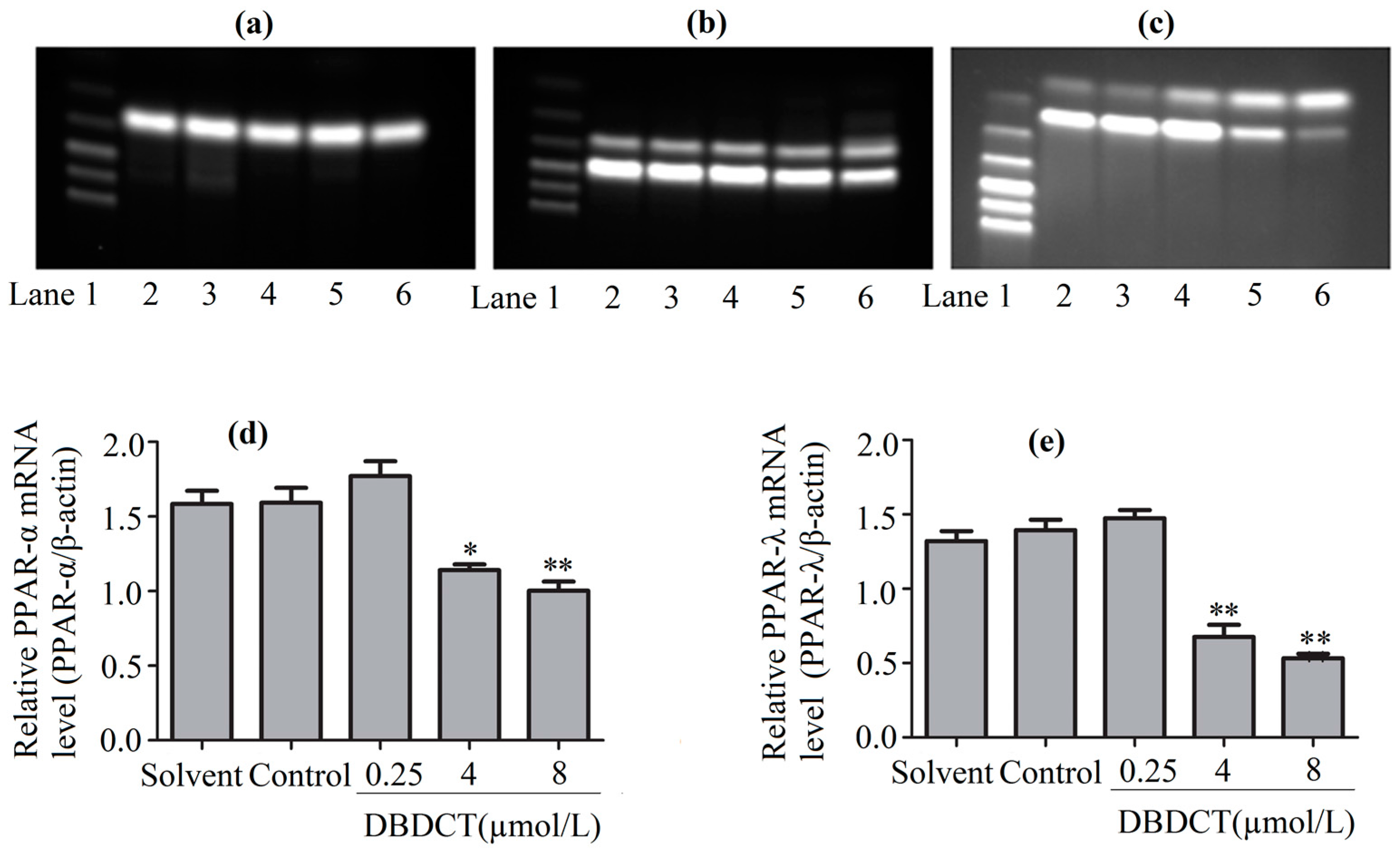
2.6. DBDCT Regulated the mRNA Levels of Downstream Targets in the PPAR Signaling Pathway
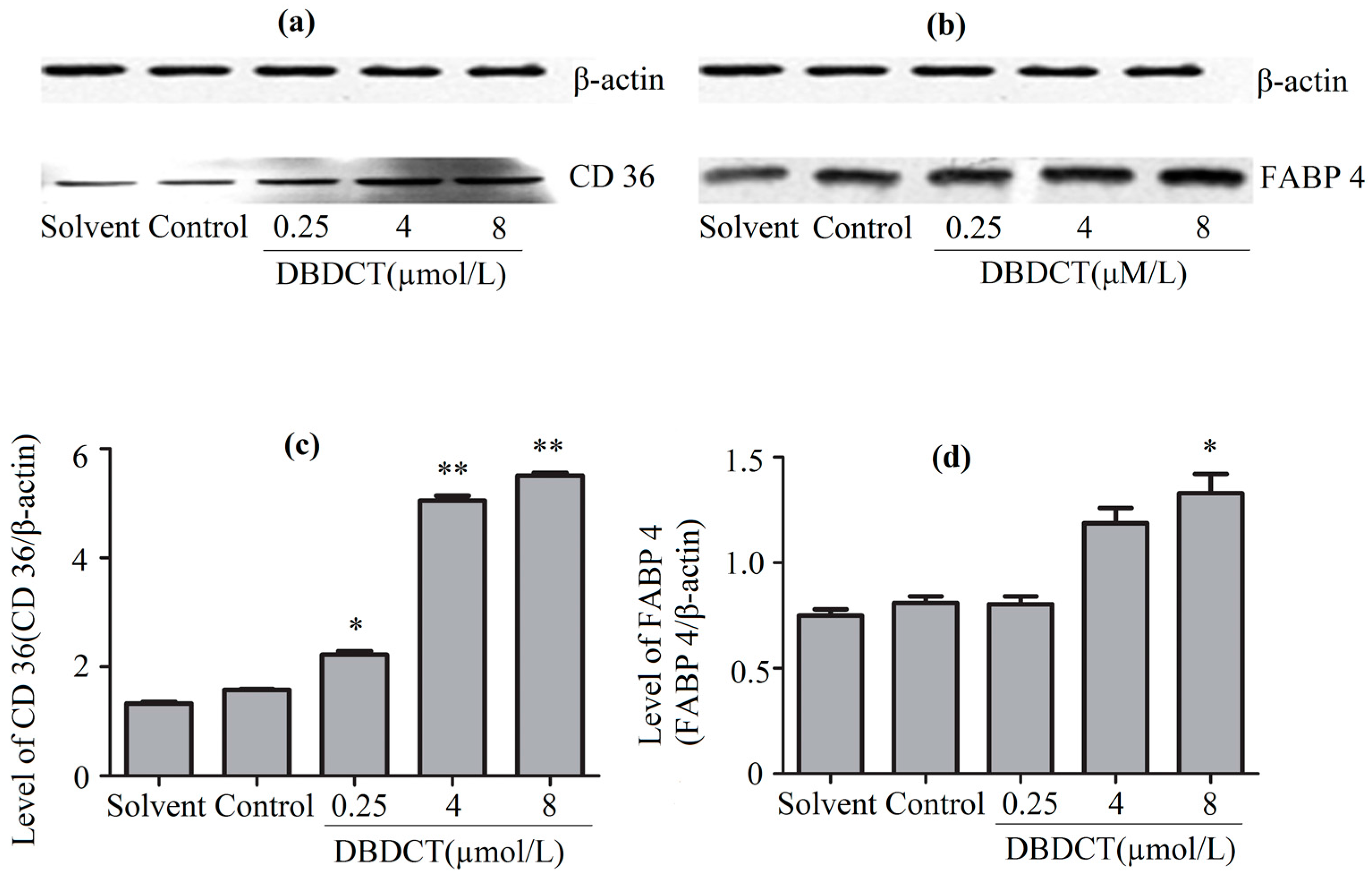
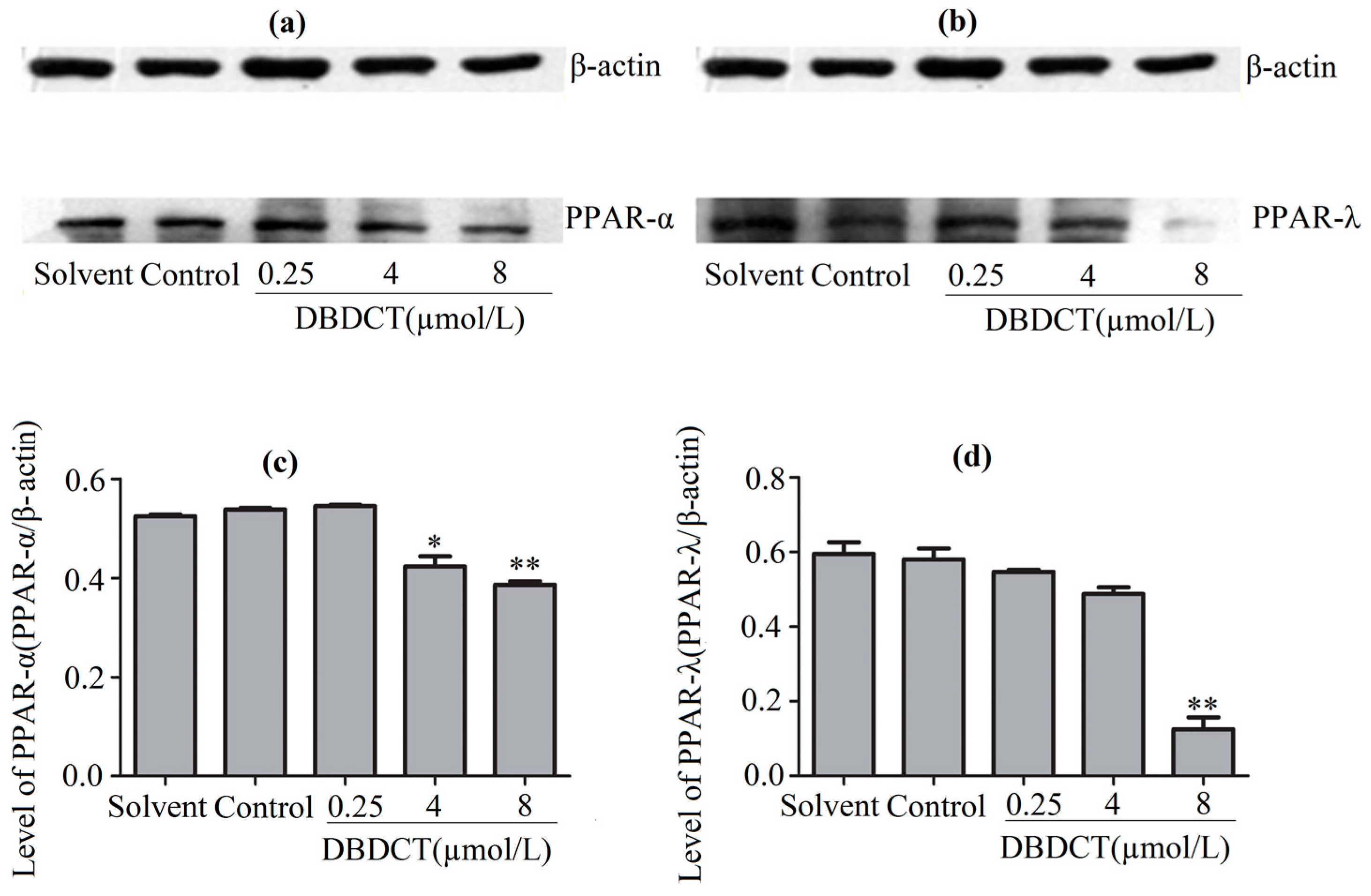
2.7. Effects of DBDCT on Expressions of PPAR-α and PPAR-λ in Protein and Gene Levels
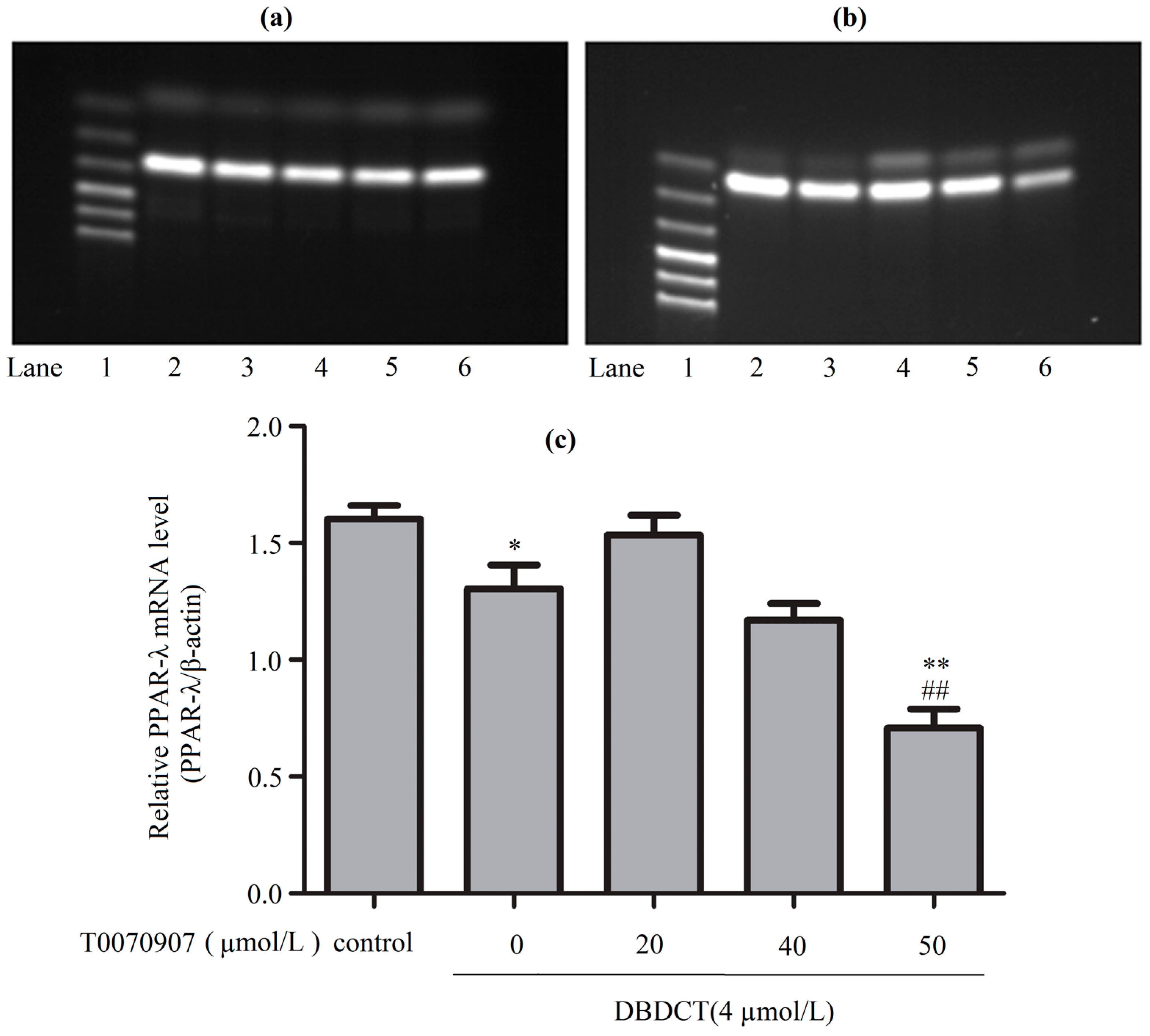
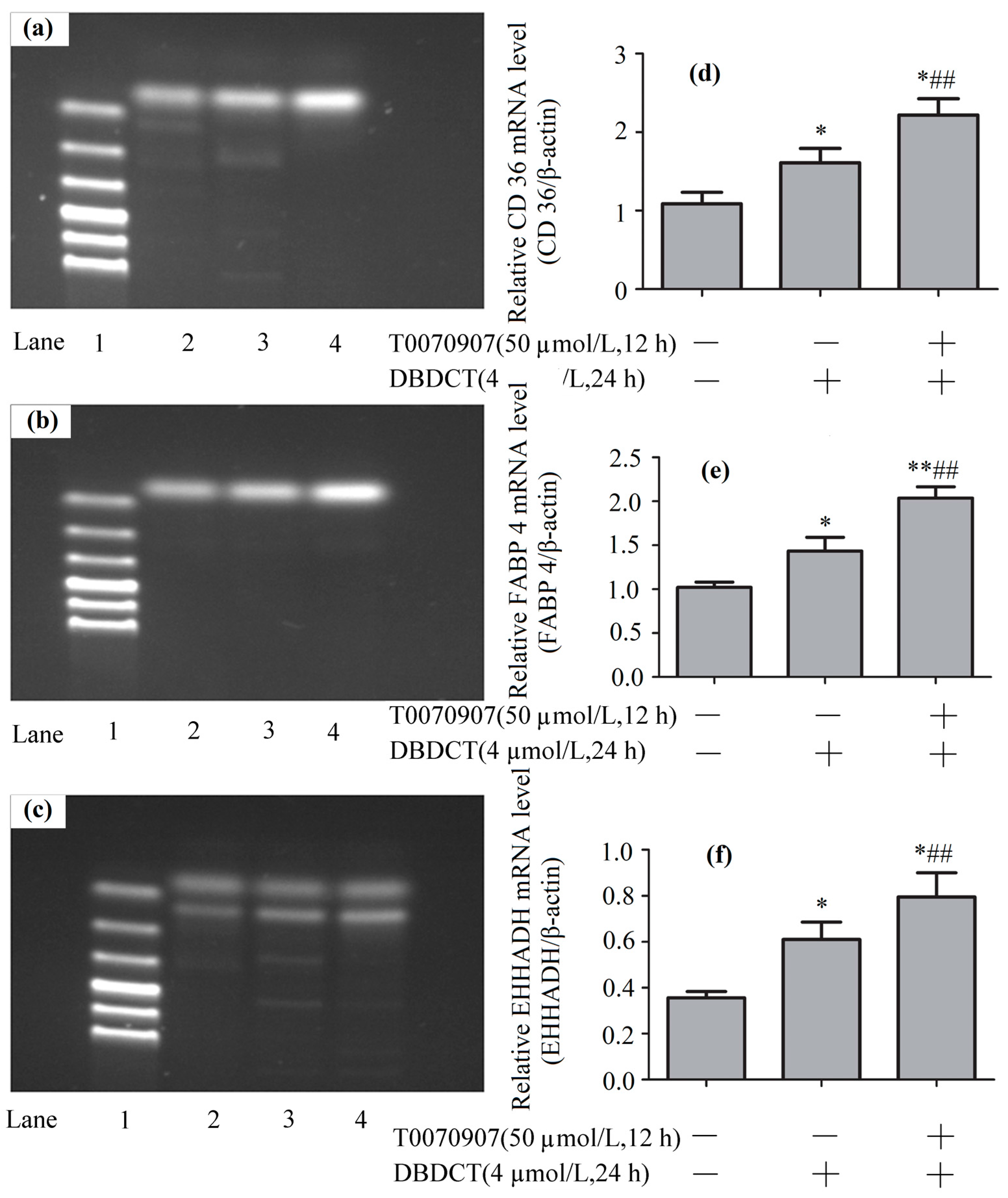
2.8. Effects of DBDCT on Expressions of PPAR-λ, CD36, FABP4 and EHHADH in Gene Levels
3. Discussion
4. Materials and Methods
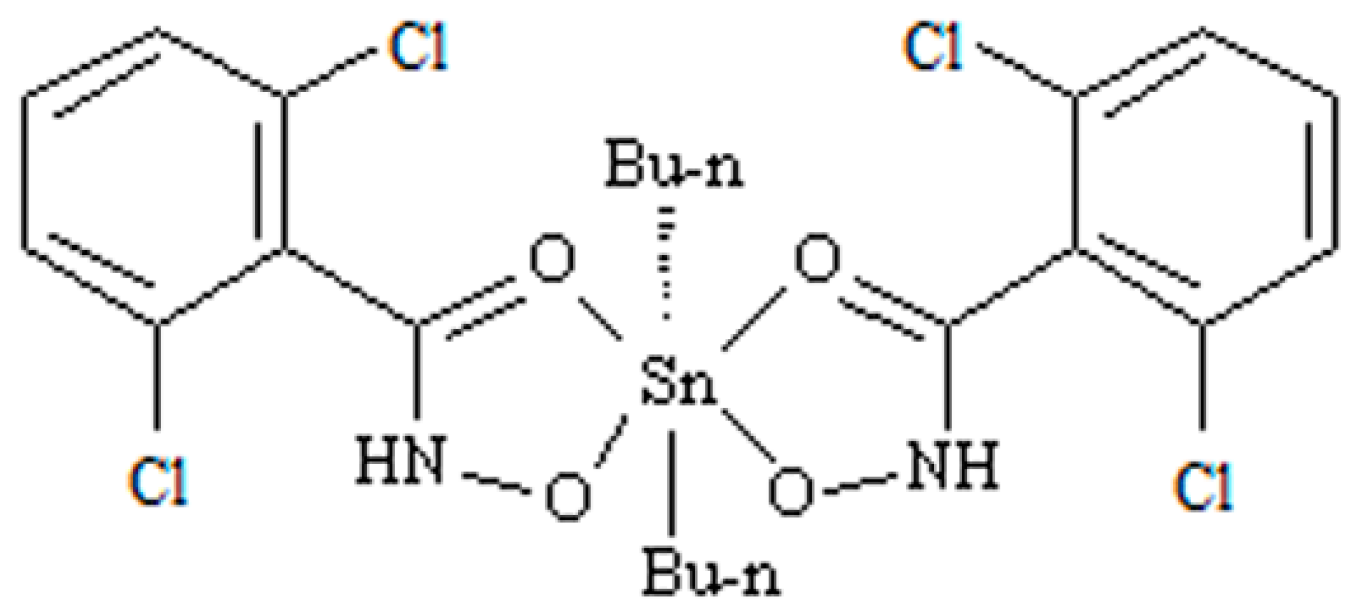
4.1. Reagents
4.2. Animals’ Treatment and Tissue Collection
4.3. Histological Evaluation of Liver Tissues
4.4. Sample Fractionation
4.5. Liquid Chromatography—Mass Spectrometry Analysis
4.6. Label-Free Protein Expression Data Processing
4.7. Cell Culture
4.8. Cytotoxicity Assay
4.9. Western Blot Analysis
4.10. RT-PCR Analysis
4.11. Statistical Analysis
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Huang, J.; Lou, G.; Zhang, Z.; Ju, Z.; Qi, C.; Zhang, Y.; Wang, C.; Li, J.; Yin, W.; Xu, Y.; et al. iTRAQ-proteomics and bioinformatics analyses of mammary tissue from cows with clinical mastitis due to natural infection with Staphylococci aureus. BMC Genom. 2014, 15, 839–853. [Google Scholar] [CrossRef] [PubMed]
- Dunston, C.R.; Griffiths, H.R.; Lambert, P.A.; Staddon, S.; Vernallis, A.B. Proteomic analysis of the anti-inflammatory action of minocycline. Proteomics 2011, 11, 42–51. [Google Scholar] [CrossRef] [PubMed]
- Rui, G.; Wenhua, M.; Yunlan, L.; Qingshan, L. Apoptosis induced neurotoxicity of di-n-butyl-di-(4-chlorobenzohydroxamato) tin (IV) via mitochondria-mediated pathway in PC12 cells. Toxicol. In Vitro 2013, 1, 92–102. [Google Scholar]
- Zhang, Y.; Li, Y.; Li, Q. Inhibition of cytochrome P450 3A in rat liver by the Diorganotin(IV) compound di-n-butyl-di-(4-chlorobenzo-hydroxamato)tin(IV) and its probable mechanism. Molecules 2012, 17, 10994–11009. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.L.; Wang, Z.W.; Guo, P.; Tang, L.; Ge, R.; Ban, S.R.; Chai, Q.Y.; Niu, L.; Li, Q.S. Diorganotin(IV) derivatives of substituted N-hydroxybenzamides with selective cytotoxicity in vitro and potent antitumor activity in vivo. J. Inorg. Biochem. 2014, 133, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.L.; Liu, J.J.; Li, Q.S. Mechanisms by which the antitumor compound di-n-butyl-di-(4-chlorobenzohydroxamato)tin(IV) induces apoptosis and the mitochondrial-mediated signaling pathway in human cancer SGC-7901 cells. J. Mol. Carcinogen. 2010, 49, 566–581. [Google Scholar] [CrossRef] [PubMed]
- Li, T. Investigation on Proteomics and Trx1-Mediated Oxidative Stress of DBDCT-Induced Hepatotoxicity. Ph.D. Thesis, Shanxi Medical University, Taiyuan, China, 2013. [Google Scholar]
- Agnieszka, L.; Konstantinos, V.; Manousos, M.; Julie, K.; William, M.; Mahmoud, A.; Konstantinos, S.; Ioannis, K.; Axel, S.M.; Jerome, Z.; et al. Comparative analysis of label-free and 8-Plex iTRAQ approach for quantitative tissue proteomic analysis. PLoS ONE 2015, 10, 1–25. [Google Scholar]
- Zihao, D. Isoniazid- and Rifampicin- Induced Hepatotoxicity and Its Mechanism. Ph.D. Thesis, Anhui Medical University, Hefei, China, 2010. [Google Scholar]
- Lieber, C.S. Alcoholic fatty liver: Its pathogenesis and mechanism of progression to inflammation and fibrosis. Alocohol 2004, 34, 9–19. [Google Scholar] [CrossRef] [PubMed]
- Le Maire, A.; Grimaldi, M.; Roecklin, D.; Dagnino, S.; Vivat-Hannah, V.; Balaguer, P.; Bourguet, W. Activation of RXR–PPAR heterodimers by organotin environmental endocrine disruptors. EMBO Rep. 2009, 10, 367–373. [Google Scholar] [CrossRef] [PubMed]
- Lyssimachou, A.; Santos, J.G.; André, A.; Soares, J.; Lima, D.; Guimarães, L.; Almeida, C.M.; Teixeira, C.; Castro, L.F.; Santos, M.M. The mammalian “obesogen” tributyltin targets hepatic triglyceride accumulation and the transcriptional regulation of lipid metabolism in the liver and brain of zebrafish. PLoS ONE 2015, 10, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Herreros, M.A.; Encinas, T.; Torres-Rovira, L.; Garcia-Fernandez, R.A.; Flores, J.M.; Ros, J.M.; Gonzalez-Bulnes, A. Exposure to the endocrine disruptor di(2-ethylhexyl)phthalate affects female reproductive features by altering pulsatile LH secretion. Environ. Toxicol. Pharmacol. 2013, 36, 1141–1149. [Google Scholar] [CrossRef] [PubMed]
- Boiteux, G.; Lascombe, I.; Roche, E.; Plissonnier, M.L.; Clairotte, A.; Bittard, H.; Fauconnet, S. A-FABP, a candidate progression marker of human transitional cell carcinoma of the bladder, is differentially regulated by PPAR in urothelial cancer cells. Int. J. Cancer 2009, 124, 1820–1828. [Google Scholar]
- Ivanova, E.A.; Parolari, A.; Myasoedova, V.; Melnichenko, A.A.; Bobryshev, Y.V.; Orekhov, A.N. Peroxisome proliferator-activated receptor (PPAR) gamma in cardiovascular disorders and cardiovascular surgery. J. Cardiol. 2015, 4, 271–278. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.X. PPARs: Diverse regulators in energy metabolism and metabolic diseases. Cell Res. 2010, 20, 124–137. [Google Scholar] [CrossRef] [PubMed]
- Habold, C.; Reichardt, F.; Le, M.Y.; Angel, F.; Liewig, N. Clay ingestion enhances intestinal triacylglycerol hydrolysis and non-esterified fatty acid absorption. Br. J. Nutr. 2009, 102, 249–257. [Google Scholar] [CrossRef] [PubMed]
- Bieghs, V.; Verheyen, F.; van Gorp, P.J.; Hendrikx, T.; Wouters, K.; Lütjohann, D.; Gijbels, M.J.; Febbraio, M.; Binder, C.J.; Hofker, M.H.; et al. Nternalization of modified lipids by CD36 and SR-A leads to hepatic inflammation and lysosomal cholesterol storage in Kupffer cells. PLoS ONE 2012, 7, e34378. [Google Scholar] [CrossRef] [PubMed]
- Ruixia, X.; Lei, G.; Weili, Z.; Wei, Z.; Guangchao, S.; Shangquan, G.; Guoqing, S. Analysis of FABP4 expression pattern in rump fat deposition and metabolism of Altay sheep. Hereditas 2015, 37, 174–182. [Google Scholar] [PubMed]
- Reza, J.Z.; Doosti, M.; Salehipour, M.; Packnejad, M.; Mojarrad, M.; Heidari, M. Modulation peroxisome proliferators activated receptor alpha (PPAR α) and acyl coenzyme A: Cholesterol acyltransferase1 (ACAT1) gene expression by fatty acids in foam cell. Lipids Health Dis. 2009, 8, 38. [Google Scholar] [CrossRef] [PubMed]
- Wan, Z.; Matravadia, S.; Holloway, G.P.; Wright, D.C. FAT/CD36 regulates PEPCK expression in adipose tissue. Am. J. Physiol. Cell Physiol. 2013, 304, 478–484. [Google Scholar] [CrossRef] [PubMed]
- Bo, M.; Huanhuan, Q.; Jing, L.; Hong, X.; Bo, C.; Jianwei, Z.; Lisha, Y.; Guohua, A.; Qi, Z. Triptolide disrupts fatty acids and peroxisome proliferator-activated receptor (PPAR) levels in male mice testes followed by testicular injury: A GC–MS based metabolomics study. Toxicology 2015, 336, 84–95. [Google Scholar]
- Kim, T.; Yang, Q. Peroxisome-proliferator-activated receptors regulate redox signaling in the cardiovascular system. World J. Cardiol. 2013, 5, 164–174. [Google Scholar] [CrossRef] [PubMed]
- Nakajima, T.; Kamijo, Y.; Tanaka, N.; Sugiyama, E.; Tanaka, E.; Kiyosawa, K.; Fukushima, Y.; Peters, J.M.; Gonzalez, F.J.; Aoyama, T. Peroxisome proliferator-activated receptor alpha protects against alcohol-induced liver damage. Hepatology 2004, 40, 972–980. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Wang, X.; Lü, X.; Jing, R.; Li, J.; Li, C.; Wang, D.; Bi, B.; Chen, X.; Wang, F.; et al. Unraveling molecular effects of ADAR1 overexpression in HEK293T cells by label-free quantitative proteomics. Cell Cycle 2016, 15, 1591–1601. [Google Scholar] [CrossRef] [PubMed]
- Rebhun, J.F.; Glynn, K.M.; Missler, S.R. Identification of glabridin as a bioactive compound in licorice (Glycyrrhiza glabra L.) extract that activates human peroxisome proliferator-activated receptor gamma (PPARγ). Fitoterapia 2015, 106, 55–61. [Google Scholar] [CrossRef] [PubMed]
- Vangaveti, V.N.; Shashidhar, V.M.; Rush, C.; Malabu, U.H.; Rasalam, R.R.; Collier, F.; Baune, B.T.; Kennedy, R.L. Hydroxyoctadecadienoic acids regulate apoptosis in human THP-1 cells in a PPARγ-dependent manner. Lipids 2014, 49, 1181–1192. [Google Scholar] [CrossRef] [PubMed]
- Datta, G.; Kramer, P.A.; Johnson, M.S.; Sawada, H.; Smythies, L.E.; Crossman, D.K.; Chacko, B.; Ballinger, S.W.; Westbrook, D.G.; Mayakonda, P.; et al. Bioenergetic programming of macrophages by the apolipoprotein A-I mimetic peptide 4F. Biochem. J. 2015, 467, 517–527. [Google Scholar] [CrossRef] [PubMed]
- Hah, Y.S.; Joo, H.H.; Kang, Y.H.; Park, B.W.; Hwang, S.C.; Kim, J.W.; Sung, I.Y.; Rho, G.J.; Woo, D.K.; Byun, J.H. Cultured human periosteal-derived cells have inducible adipogenic activity and can also differentiate into osteoblasts in a perioxisome proliferator-activated receptor-mediated fashion. Int. J. Med. Sci. 2014, 11, 1116–1128. [Google Scholar] [CrossRef] [PubMed]
- Parillo, F.; Maranesi, M.; Brecchia, G.; Gobbetti, A.; Boiti, C.; Zerani, M. In vivo chronic and in vitro acute effects of di(2-ethylhexyl) phthalate on pseudopregnant rabbit corpora lutea: Possible involvement of peroxisome proliferator-activated receptor gamma. Biol. Reprod. 2014, 90, 41. [Google Scholar] [CrossRef] [PubMed]
- Kong, L.; Ren, W.; Li, W.; Zhao, S.; Mi, H.; Wang, R.; Zhang, Y.; Wu, W.; Nan, Y.; Yu, J. Activation of peroxisome proliferator activated receptor alpha ameliorates ethanol induced steatohepatitis in mice. Lipids Health Dis. 2011, 10, 246. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of the compounds are not available from the authors. |

| Term | Count | p-Value |
|---|---|---|
| Oxidation reduction | 31 | 1.2 × 10−13 |
| Response to organic substance | 21 | 6.9 × 10−4 |
| Fatty acid metabolic process | 11 | 1.6 × 10−5 |
| Lipid biosynthetic process | 10 | 1.8 × 10−3 |
| Macromolecule catabolic process | 10 | 3.8 × 10−2 |
| Response to hormone stimulus | 10 | 6.4 × 10−2 |
| Protein transport | 10 | 9.2 × 10−2 |
| Establishment of protein localization | 10 | 9.6 × 10−2 |
| Response to drug | 9 | 1.5 × 10−2 |
| Cofactor metabolic process | 8 | 3.8 × 10−3 |
| Response to inorganic substance | 8 | 1.7 × 10−2 |
| Protein catabolic process | 8 | 4.6 × 10−2 |
| Lipid catabolic process | 7 | 2.1 × 10−3 |
| Di- and tri-valent inorganic cation transport | 7 | 3.3 × 10−3 |
| Steroid metabolic process | 7 | 6.3 × 10−3 |
| Cellular protein catabolic process | 7 | 8.9 × 10−2 |
| Organic acid biosynthetic process | 6 | 1.6 × 10−2 |
| Carboxylic acid biosynthetic process | 6 | 1.6 × 10−2 |
| Response to metal ion | 6 | 3.1 × 10−2 |
| Response to organic cyclic substance | 6 | 5.7 × 10−2 |
| Pathway | Count | p-Value |
|---|---|---|
| Drug metabolism | 13 | 2.8 × 10−11 |
| Metabolism of xenobiotics by cytochrome P450 | 11 | 1.4 × 10−9 |
| Retinol metabolism | 8 | 5.4 × 10−6 |
| Linoleic acid metabolism | 6 | 3.9 × 10−5 |
| PPAR signaling pathway | 7 | 1.8 × 10−4 |
| Arachidonic acid metabolism | 6 | 1.4 × 10−3 |
| Steroid biosynthesis | 4 | 1.7 × 10−2 |
| β-Alanine metabolism | 3 | 2.8 × 10−2 |
| Pyruvate metabolism | 3 | 7.9 × 10−2 |
| Pyrimidine metabolism | 4 | 9.7 × 10−2 |
| Protein ID | Protein Name | Fold Change | Protein Description |
|---|---|---|---|
| Q07969 | Platelet glycoprotein 4/CD36 | 3.56 | Binds long-chain fatty acids and may function in transport and/or as a regulator of fatty acid transport. As a coreceptor fortoll-like receptor 4-6 (TLR4-TLR6), promotes inflammation in monocytes/macrophages. Upon ligand binding, such as oxLDL or amyloid-β 42 binding, rapidly induces the formation of a heterodimer of TLR4 and TLR6, which is internalized and triggers inflammatory signals, leading to the NF-kappa-B-dependent production of CXCL1, CXCL2 and CCL9 cytokines via the MYD88 signaling pathway, and CCL5 cytokine via the TICAM1 signaling pathway, as well as IL1B secretion. |
| P70623 | Fatty acid binding protein, adipocyte 4 | 2.02 | Lipid transport protein in adipocytes. Binds both long-chain fatty acids and retinoic acid. Delivers long-chain fatty acids and retinoic acid to their cognate receptors in the nucleus. |
| P07896 | Peroxisomal bifunctional enzyme/enoyl-CoA hydratase | 3.16 | |
| P07871 | 3-ketoacyl-CoA thiolase B, peroxisomal/acetyl-CoA acyltransferase 1 | 2.28 | |
| P07379 | Phosphoenolpyruvate carboxykinase, cytosolic | 2.02 | Catalyzes the conversion of oxaloacetate (OAA) to phosphoenolpyruvate (PEP), the rate-limiting step in the metabolic pathway that produces glucose from lactate and other precursors derived from the citric acid cycle. |
| Isoforms | Primer Sequence | Product Size (bp) | PCR Conditions | ||
|---|---|---|---|---|---|
| Cycle | Denaturation | Annealing | |||
| β-actin | F:CTACAATGAGCTGCGTGTGGC R:CAGGTCCAGACGCAGGATGGC | 270 | 30 | 94 °C, 30 s | 55 °C, 30 s |
| PPAR-α | F:ACGATGCTGTCCTCCTTGATG R:GCGTCTGACTCGGTCTTCTTG | 407 | 30 | 94 °C, 30 s | 52 °C, 30 s |
| PPAR-λ | F:CCCTTTACCACGGTTGATTTCTC R:GCAGGCTCTACTTTGATCGCACD | 143 | 40 | 94 °C, 30 s | 59 °C, 30 s |
| CD36 | F:TGAATCCTAACGAAGATGAGCA R:GGCTTGACCAGTATGTTGACC | 106 | 40 | 94 °C, 30 s | 60 °C, 30 s |
| FABP4 | F:AGAAGTGGGAGTTGGCTTCG R:ACTCTCTGACCGGATGACGA | 103 | 40 | 94 °C, 30 s | 60 °C, 30 s |
| ACAA1 | F:TGGCATCAGAAATGGGTCTT R:AGGAATCAGGCAGTCTCTGG | 136 | 40 | 94 °C, 30 s | 60 °C, 30 s |
| EHHADH | F:CACGGTTATGAGCTTGTCCA R:TCTGGCTTGCTACCTTCCTC | 138 | 40 | 94 °C, 30 s | 60 °C, 30 s |
| PEPCK | F:CTGCCTCTCTCCACACCATT R:GCCTTCCACAAACTTCCTCA | 125 | 40 | 94 °C, 30 s | 60 °C, 30 s |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, Y.; Liu, X.; Niu, L.; Li, Q. Proteomics Analysis Reveals an Important Role for the PPAR Signaling Pathway in DBDCT-Induced Hepatotoxicity Mechanisms. Molecules 2017, 22, 1113. https://doi.org/10.3390/molecules22071113
Li Y, Liu X, Niu L, Li Q. Proteomics Analysis Reveals an Important Role for the PPAR Signaling Pathway in DBDCT-Induced Hepatotoxicity Mechanisms. Molecules. 2017; 22(7):1113. https://doi.org/10.3390/molecules22071113
Chicago/Turabian StyleLi, Yunlan, Xinxin Liu, Lin Niu, and Qingshan Li. 2017. "Proteomics Analysis Reveals an Important Role for the PPAR Signaling Pathway in DBDCT-Induced Hepatotoxicity Mechanisms" Molecules 22, no. 7: 1113. https://doi.org/10.3390/molecules22071113
APA StyleLi, Y., Liu, X., Niu, L., & Li, Q. (2017). Proteomics Analysis Reveals an Important Role for the PPAR Signaling Pathway in DBDCT-Induced Hepatotoxicity Mechanisms. Molecules, 22(7), 1113. https://doi.org/10.3390/molecules22071113




