New Perspectives on the Use of Phytochemicals as an Emergent Strategy to Control Bacterial Infections Including Biofilms
Abstract
:1. Introduction
2. Clinical Multidrug-Resistant Bacteria—The Beginning of the Post-Antibiotic Era
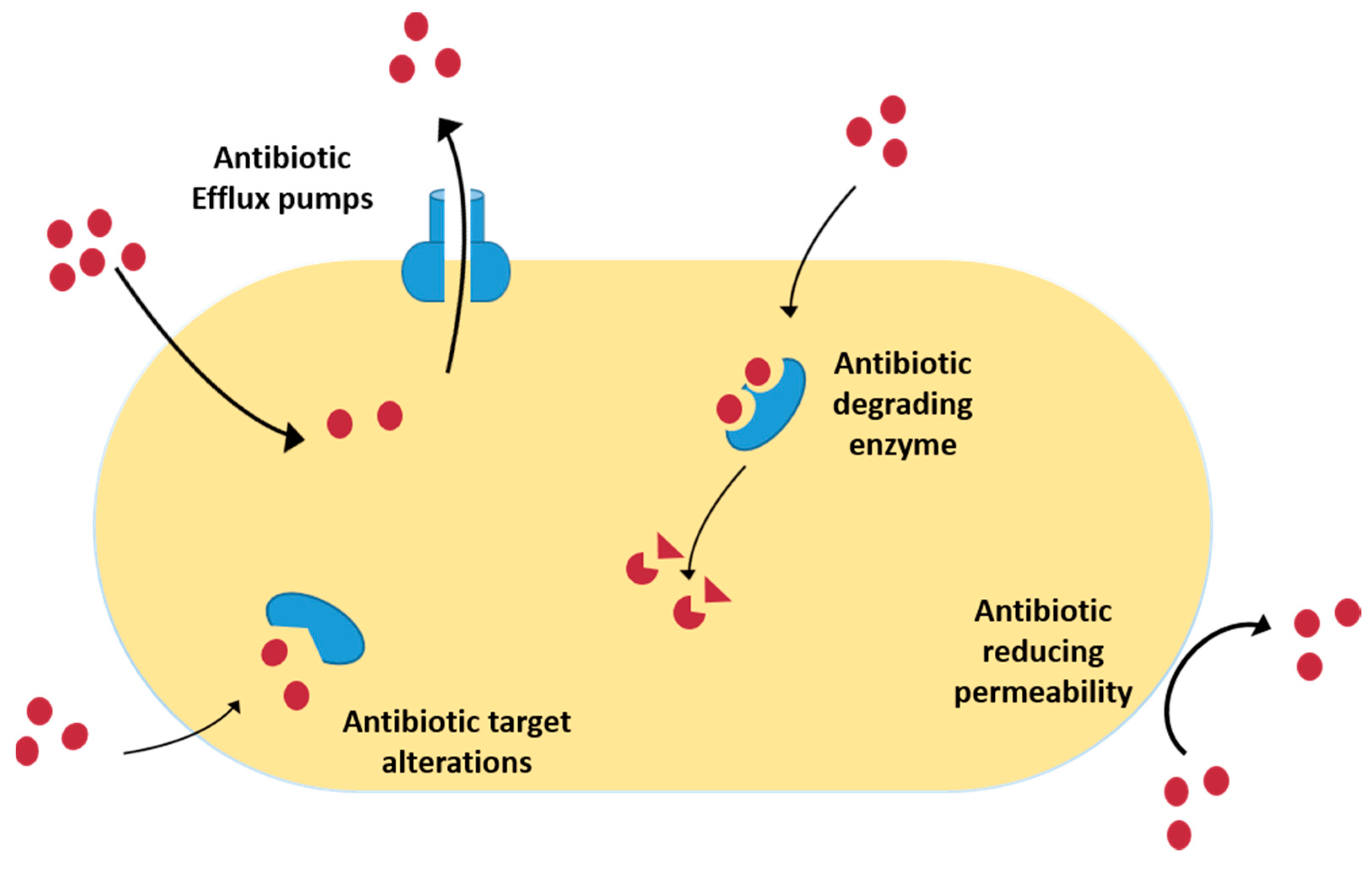
2.1. Mechanisms Explaining Bacterial Resistance to Antibiotics
- Prevention of access to target by reducing permeability—Microorganisms are surrounded by a cell envelop that constitutes a selective permeability barrier and thus can effectively offer protection from drug molecules in the extracellular environment, while providing sufficient nutrients to the cell [27]. Most of the antibiotics target intracellular processes and must be able to penetrate the bacterial membrane [28]. Reduction of the concentration of the antibiotic into the cell, due to modification of the cell surface, limits the interaction with the drug (e.g., lipid A modification) or reduces the number of entry channels (e.g., porins) [27]. In Gram-negative bacteria the outer membrane serves as a physical and functional barrier. Such an outer membrane contains an inner layer that has phospholipids and an outer layer that has lipid A. This composition reduces the permeability to many drug [25]. For bacteria belonging to the Enterobacteriaceae family, the majority of the porins are thought to function as non-specific channels; an example of this is the outer-membrane proteins OmpF and OmpC of Escherichia coli [15].
- Increased efflux pumps—Several energy-dependent systems that reduce the intracellular concentration of toxic substances were identified in bacteria. EPs are transport proteins, localized in the cytoplasmatic membrane, found in both Gram-negative and -positive bacteria as well as in eukaryotic organisms. These pump systems remove toxic compounds out of the bacterial cell in a process that does not comprise the alteration or degradation of the molecules [27]. EPs can be specific for one substrate or transport a variety of structurally dissimilar substances, such as these latter pumps associated with the occurrence of MDR bacteria [29]. Currently, the identification and characterization of EPs remains one of the major problems [30]. There are five major families of EP transporters: RND (resistance-nodulation-division); MF (major facilitator); MATE (multidrug and toxic efflux); SMR (small multidrug resistance); and ABC (ATP binding cassette). In most of the bacteria, EP genes are localized in an operon controlled by a regulatory gene [31]. In the last years, new EPs have been described, such as MdeA in Streptococcus mutans, KexD in Klebsiella pneumoniae and Lmrs in Staphylococcus aureus [32,33,34].
- Degradation and modification of antibiotics—Another means by which bacteria can be resistant is by destroying the active component of the antibiotics. There are three mechanisms through which bacteria inactivate the antibiotics: enzymatic hydrolysis; group transfer; or redox process [30]. Among them, antibiotic inactivation catalysed by enzymes is the main mechanism of resistance. Various antibiotics have hydrolytically susceptible chemical bonds, e.g., esters and amides, whose integrity is central to biological activity [35]. Thousands of enzymes are known to degrade and modify antibiotics of different classes, including β-lactams, aminoglycosides, phenicols and macrolides. The expansion of antibiotic classes to include derivatives that have improved properties has been reflected by the emergence of hydrolytic enzymes that have altered spectres of activity [36,37,38,39].The β-lactamases are broadly prevalent and clinically important resistant enzymes [40]. The β-lactamases can be classified using two systems: Ambler and Bush-Jacoby-Medeiros [24]. These enzymes are the most important mechanisms of resistance in Gram-negative bacteria and can be coded on plasmids and chromosomes [25]. The genes that codify β-lactamases can be transferred by transposons but can be found in the composition of integrons [41]. There are two distinct chemical mechanisms employed by β-lactamases to hydrolytically cleave the ring of β-lactam antibiotics: formation of a covalent enzyme intermediate followed by hydrolysis or meta-activation of nucleophilic water molecules via the Zn2+ centre [30,42]. Serine β-lactamase cephalosporinases are found in Enterobacter spp. and Pseudomonas spp., penicilinases in strains of S. aureus [24,43,44,45,46]. The enzyme metallo-β-lactamase, found in Pseudomonas aeruginosa, K. pneumoniae, E. coli, and on the Gram-positive bacteria Proteus mirabilis and Enterobacter spp., is responsible for resistance to imipenem, and the new generation of cephalosporins and penicillins [25,43]. Extended-spectrum β-lactamases (ESBLs) that are active against first-generation β-lactams can be encoded in large plasmids but can also be transferred by transposon insertion [47,48,49,50,51]. Other examples of hydrolytic enzymes include esterases, which have been linked to macrolide antibiotic resistance and fosfomycin resistance ring-opening epoxidases [25]. Hydrolytic enzymes inactivate the antibiotics before the molecule can reach their target in the bacteria. Because these enzymes require only water as a co-substrate, they can often be excreted by the bacteria [30]. The most varied family of resistance enzymes is the transferases group that inactivates aminoglycosides, chloramphenicol, streptogramin, macrolides or rifampicin. The inactivation of antimicrobial agents occurs by binding adenylyl, phosphoryl, or acetyl groups to the periphery of the molecule. Phosphoryltransferases, nucleotidyltransferases, or adenylyltransferases and acetyltransferases are aminoglycoside neutralizer enzymes [52]. These enzymes can be found in S. aureus, Enterococcus faecalis and Streptococcus pneumoniae [53]. Oxidation and reduction are also used by pathogenic bacteria as a mechanism of antibiotic resistance [52,54].
- Modification of the molecular target—The majority of the antibiotics specifically bind to their targets with high affinity, so even a small mutation in a target molecule is sufficient to influence antibiotic binding to the target [52]. Alteration of the natural antibiotic target may arise from a spontaneous chromosomal mutation resulting in single or multiple amino acid modification, or from homologous recombination with exogenous DNA containing gene segments that encode proteins with low antibiotic binding affinity [30]. In clinical pathogenic bacteria, several genes that encode for target modification of the same antibiotic were already described. One example is the methicillin resistance in S. aureus [24,52,55].
2.2. Biofilms as a Contributing Factor for the Increased Resistance to Antibacterials
2.3. Drug-Resistant Microorganisms—from Environment to Clinic
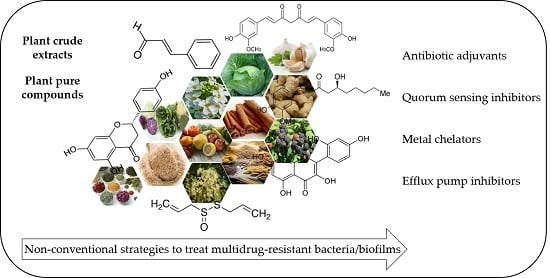
3. Non-Conventional Strategies to Treat Drug-Resistant Bacteria
4. The Use of Plants as Sources of Antibiotic Adjuvants
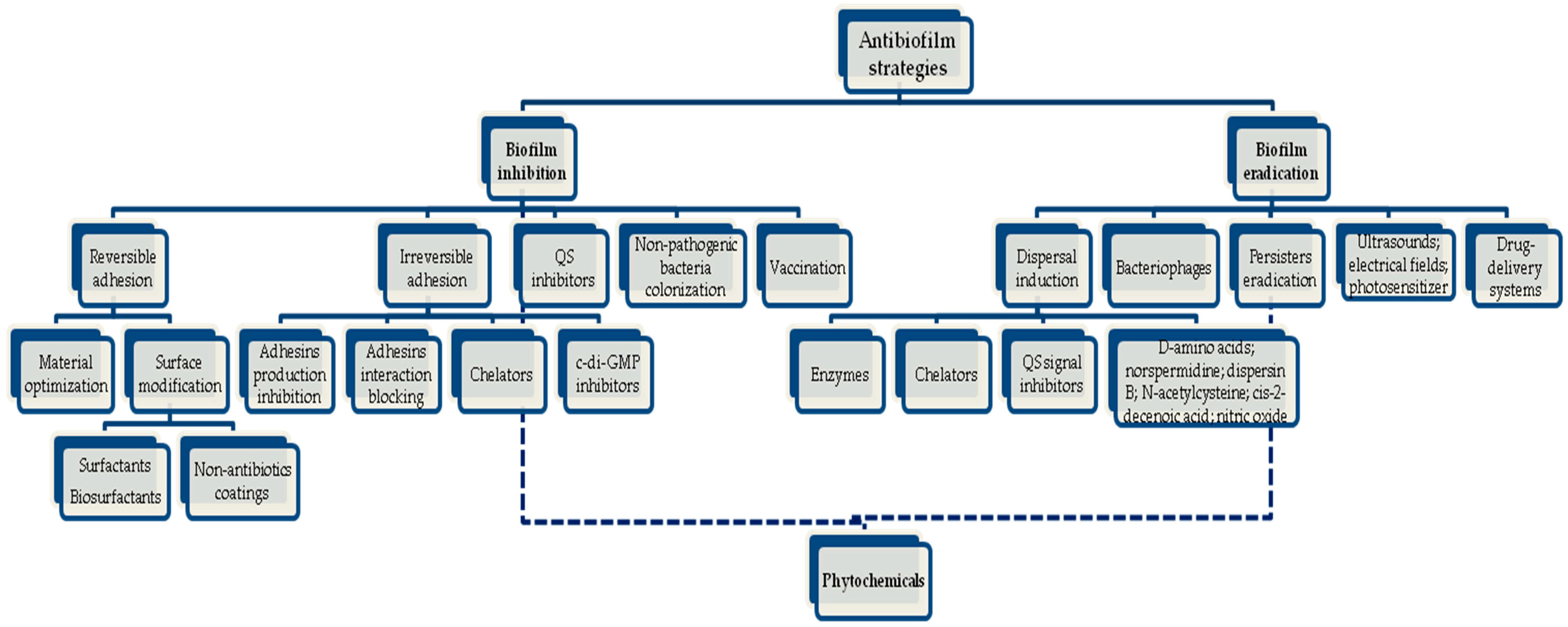
5. Non-Conventional Strategies to Treat Bacteria Growing within a Biofilm
- (a)
- Inhibition of bacterial reversible adhesion by optimization of the physicochemical properties of the materials used in medical devices and implants, and surface modification using surfactants/biosurfactants and other types of non-antibiotic coatings;
- (b)
- Inhibition of irreversible adhesion by interference with the production of adhesins, blocking of the adhesins’ interaction with their receptors, use of chelating agents that inhibit the transport of essential metals to the interior of cells, thus stopping biochemical pathways that are crucial for biofilm formation, and inhibition of nucleotide signalling biosynthesis such as cyclic diguanosine monophosphate (c-di-GMP), which can maintain bacteria in the planktonic state;
- (c)
- Interference with the bacterial communication through the use of QS inhibitors (QSI); the use of non-pathogenic bacteria which can compete with pathogens by producing toxins (e.g., bacteriocins) or other substances, thus preventing colonization; and vaccination in order to produce antibodies against antigens of bacterial biofilms, preventing the evolution of infection.
- (a)
- Induction of dispersal by the application of enzymes (e.g., DNAse I, proteinase K, tripsin, lysostaphin, amylase, lyase and lactonase), divalent metal chelators, QS signal inhibitors, and other molecules such as d-amino acids (e.g., d-leucine, d-methionine, d-tyrosine and d-tryptophan), norspermidine, dispersin B, N-acetylcysteine, cis-2-decenoic acid, and nitric oxide;
- (b)
- The use of bacteriophages;
- (c)
- Eradication of persister cells (e.g., combined application of sugars or silver with antibiotics and/or an increase in the production of reactive oxygen species).
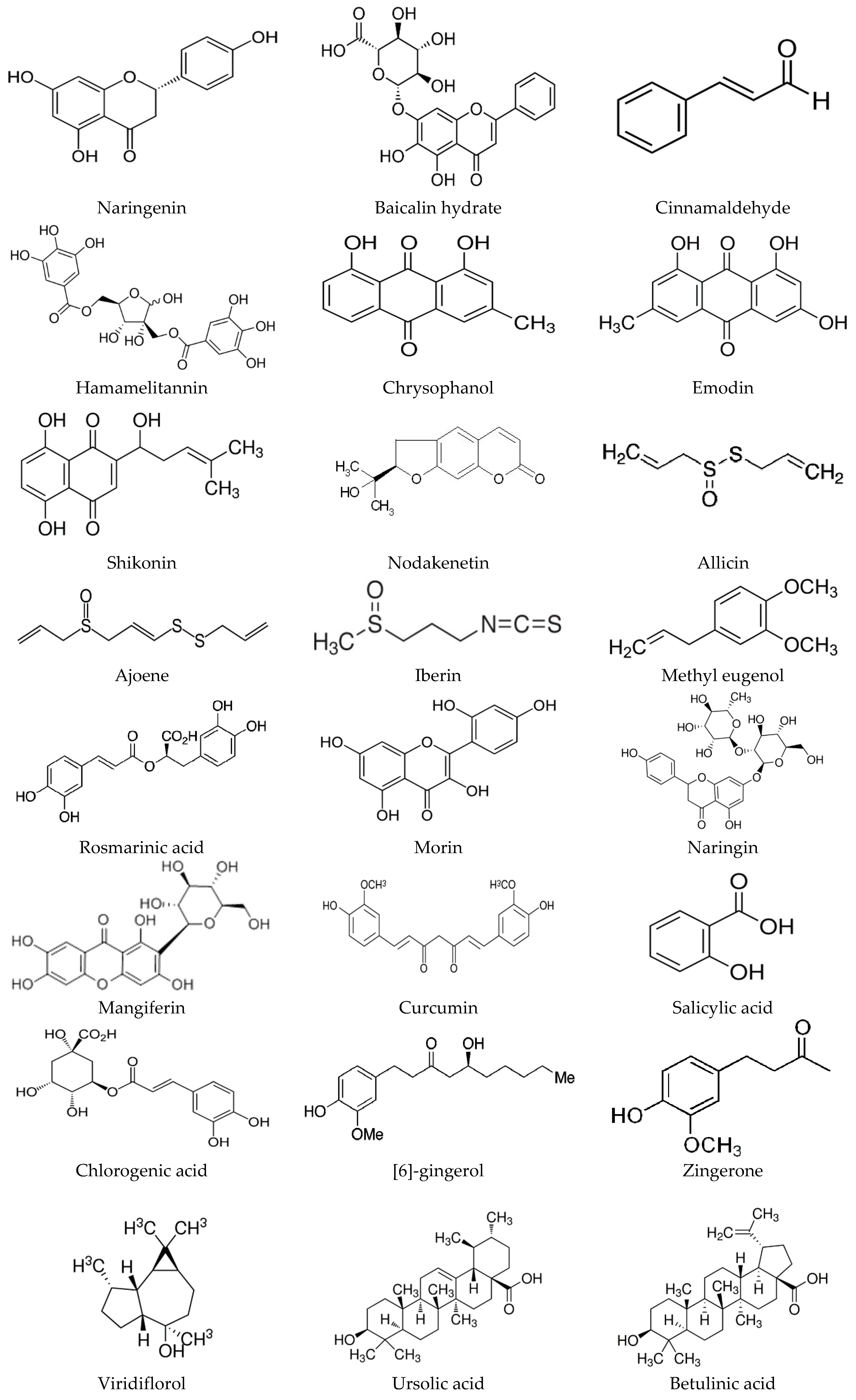
6. Plant-Based Strategies to Deal with Unwanted Biofilms
6.1. Phytochemicals as Quorum-Sensing Inhibitors
6.2. Phytochemicals as Biofilm Metal Chelators
6.3. Phytochemicals as Biofilm Efflux Pump Inhibitors
7. Concluding Remarks and Future Perspectives
Acknowledgments
Conflicts of Interest
Abbreviations
| ABC | ATP binding cassette |
| agr | accessory gene regulator |
| AHLs | N-Acyl homoserine lactones |
| c-di-GMP | Cyclic diguanosine monophosphate |
| EPS | Extracellular polymeric substances |
| ESBLs | Extended-spectrum β-lactamases |
| EPs | Efflux pumps |
| EPIs | Efflux pump inhibitors |
| EPS | Extracellular polymeric substances |
| FDA | Food and Drug Administration |
| MF | Major facilitator |
| MIC | Minimum inhibitory concentration |
| MATE | Multidrug and toxic efflux |
| MDR | Multidrug resistant |
| MRSA | Methicillin-resistant S. aureus |
| PGG | 1,2,3,4,6-Penta-O-galloyl-b-d-glucopyranose |
| QS | Quorum sensing |
| Quorum-quenching | |
| QSI | QS inhibitors |
| RND | Resistance-nodulation-division |
| SMR | Small multidrug resistance |
References
- Iriti, M.; Faoro, F. Chemical diversity and defence metabolism: How plants cope with pathogens and ozone pollution. Int. J. Mol. Sci. 2009, 10, 3371–3399. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, N.; Rivero, R.M.; Shulaev, V.; Blumwald, E.; Mittler, R. Abiotic and biotic stress combinations. New Phytol. 2014, 203, 32–43. [Google Scholar] [CrossRef] [PubMed]
- Rausher, M.D. Co-evolution and plant resistance to natural enemies. Nature 2001, 411, 857–864. [Google Scholar] [CrossRef] [PubMed]
- Dangl, J.L.; Jones, J.D.G. Plant pathogens and integrated defence responses to infection. Nature 2001, 411, 826–833. [Google Scholar] [CrossRef] [PubMed]
- War, A.R.; Paulraj, M.G.; Ahmad, T.; Buhroo, A.A.; Hussain, B.; Ignacimuthu, S.; Sharma, H.C. Mechanisms of plant defense against insect herbivores. Plant Signal. Behav. 2012, 7, 1306–1320. [Google Scholar] [CrossRef] [PubMed]
- Fürstenberg-Hägg, J.; Zagrobelny, M.; Bak, S. Plant defense against insect herbivores. Int. J. Mol. Sci. 2013, 14, 10242–10297. [Google Scholar] [CrossRef] [PubMed]
- VanEtten, H.D.; Mansfield, J.W.; Bailey, J.A.; Farmer, E.E. Two classes of plant antibiotics: Phytoalexins versus “phytoanticipins”. Plant Cell 1994, 6, 1191–1192. [Google Scholar] [CrossRef] [PubMed]
- González-Lamothe, R.; Mitchell, G.; Gattuso, M.; Diarra, M.S.; Malouin, F.; Bouarab, K. Plant antimicrobial agents and their effects on plant and human pathogens. Int. J. Mol. Sci. 2009, 10, 3400–3419. [Google Scholar] [CrossRef] [PubMed]
- Hemaiswarya, S.; Kruthiventi, A.K.; Doble, M. Synergism between natural products and antibiotics against infectious diseases. Phytomedicine 2008, 15, 639–652. [Google Scholar] [CrossRef] [PubMed]
- Dehghan, H.; Sarrafi, Y.; Salehi, P. Antioxidant and antidiabetic activities of 11 herbal plants from Hyrcania region, Iran. J. Food Drug Anal. 2016, 24, 179–188. [Google Scholar] [CrossRef]
- Akram, M.; Hamid, A.; Khalil, A.; Ghaffar, A.; Tayyaba, N.; Saeed, A.; Ali, M.; Naveed, A. Review on medicinal uses, pharmacological, phytochemistry and immunomodulatory activity of plants. Int. J. Immunopathol. Pharmacol. 2014, 27, 313–319. [Google Scholar] [PubMed]
- da Silva, J.A.T.; Yonekura, L.; Kaganda, J.; Mookdasanit, J.; Nhut, D.T.; Afach, G. Important secondary metabolites and essential oils of species within the Anthemideae (Asteraceae). J. Herbs Spices Med. Plants 2005, 11, 1–46. [Google Scholar] [CrossRef]
- Lewis, K. Platforms for antibiotic discovery. Nat. Rev. Drug Discov. 2013, 12, 371–387. [Google Scholar] [CrossRef] [PubMed]
- Tegos, G.; Stermitz, F.R.; Lomovskaya, O.; Lewis, K. Multidrug pump inhibitors uncover remarkable activity of plant antimicrobials. Antimicrob. Agents Chemother. 2002, 46, 3133–3141. [Google Scholar] [CrossRef] [PubMed]
- Blair, J.M.A.; Webber, M.A.; Baylay, A.J.; Ogbolu, D.O.; Piddock, L.J.V. Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 2015, 13, 42–51. [Google Scholar] [CrossRef] [PubMed]
- Marti, E.; Variatza, E.; Balcazar, J.L. The role of aquatic ecosystems as reservoirs of antibiotic resistance. Trends Microbiol. 2014, 22, 36–41. [Google Scholar] [CrossRef] [PubMed]
- Coates, A.R.M.; Halls, G.; Hu, Y.M. Novel classes of antibiotics or more of the same? Br. J. Pharmacol. 2011, 163, 184–194. [Google Scholar] [CrossRef] [PubMed]
- Lorenzi, V.; Muselli, A.; Bernardini, A.F.; Berti, L.; Pagès, J.-M.; Amaral, L.; Bolla, J.-M. Geraniol restores antibiotic activities against multidrug-resistant isolates from Gram-negative species. Antimicrob. Agents Chemother. 2009, 53, 2209–2211. [Google Scholar] [CrossRef] [PubMed]
- French, G.L. Clinical impact and relevance of antibiotic resistance. Adv. Drug Deliv. Rev. 2005, 57, 1514–1527. [Google Scholar] [CrossRef] [PubMed]
- Walsh, F. Investigating antibiotic resistance in non-clinical environments. Front. Microbiol. 2013, 4. [Google Scholar] [CrossRef] [PubMed]
- Dantas, G.; Sommer, M.O.A.; Oluwasegun, R.D.; Church, G.M. Bacteria subsisting on antibiotics. Science 2008, 320, 100–103. [Google Scholar] [CrossRef] [PubMed]
- Fiamegos, Y.C.; Kastritis, P.L.; Exarchou, V.; Han, H.; Bonvin, A.M.J.J.; Vervoort, J.; Lewis, K.; Hamblin, M.R.; Tegos, G.P. Antimicrobial and efflux pump inhibitory activity of caffeoylquinic acids from Artemisia absinthium against gram-positive pathogenic bacteria. PLoS ONE 2011, 6, e18127. [Google Scholar] [CrossRef] [PubMed]
- Holmes, A.H.; Moore, L.S.P.; Sundsfjord, A.; Steinbakk, M.; Regmi, S.; Karkey, A.; Guerin, P.J.; Piddock, L.J.V. Understanding the mechanisms and drivers of antimicrobial resistance. Lancet 2016, 387, 176–187. [Google Scholar] [CrossRef]
- Alekshun, M.N.; Levy, S.B. Molecular mechanisms of antibacterial multidrug resistance. Cell 2007, 128, 1037–1050. [Google Scholar] [CrossRef] [PubMed]
- Giedraitiene, A.; Vitkauskiene, A.; Naginiene, R.; Pavilonis, A. Antibiotic resistance mechanisms of clinically important bacteria. Medicina 2011, 47, 137–146. [Google Scholar] [PubMed]
- Raghunath, D. Emerging antibiotic resistance in bacteria with special reference to India. J. Biosci. 2008, 33, 593–603. [Google Scholar] [PubMed]
- Fernandez, L.; Hancock, R.E.W. Adaptive and mutational resistance: Role of porins and efflux pumps in drug resistance. Clin. Microbiol. Rev. 2012, 25, 661. [Google Scholar] [CrossRef] [PubMed]
- Delcour, A.H. Outer membrane permeability and antibiotic resistance. Biochim. Biophys. Acta 2009, 1794, 808–816. [Google Scholar] [CrossRef] [PubMed]
- Piddock, L.J.V. Clinically relevant chromosomally encoded multidrug resistance efflux pumps in bacteria. Clin. Microbiol. Rev. 2006, 19, 382–402. [Google Scholar] [CrossRef] [PubMed]
- Wright, G.D. Molecular mechanisms of antibiotic resistance. Chem. Commun. 2011, 47, 4055–4061. [Google Scholar] [CrossRef] [PubMed]
- Webber, M.A.; Piddock, L.J.V. The importance of efflux pumps in bacterial antibiotic resistance. J. Antimicrob. Chemother. 2003, 51, 9–11. [Google Scholar] [CrossRef] [PubMed]
- Floyd, J.L.; Smith, K.P.; Kumar, S.H.; Floyd, J.T.; Varela, M.F. LmrS Is a multidrug efflux pump of the major facilitator superfamily from Staphylococcus aureus. Antimicrob. Agents Chemother. 2010, 54, 5406–5412. [Google Scholar] [CrossRef] [PubMed]
- Hu, R.M.; Liao, S.T.; Huang, C.C.; Huang, Y.W.; Yang, T.C. An inducible fusaric acid tripartite efflux pump contributes to the fusaric acid resistance in Stenotrophomonas maltophilia. PLoS ONE 2012, 7. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, W.; Onishi, M.; Ni, R.T.; Tsuchiya, T.; Kuroda, T. Functional study of the novel multidrug efflux pump KexD from Klebsiella pneumoniae. Gene 2012, 498, 177–182. [Google Scholar] [CrossRef] [PubMed]
- De Pascale, G.; Wright, G.D. Antibiotic resistance by enzyme inactivation: From mechanisms to solutions. ChemBioChem 2010, 11, 1325–1334. [Google Scholar] [CrossRef] [PubMed]
- Livermore, D.M. Defining an extended-spectrum beta-lactamase. Clin. Microbiol. Infect. 2008, 14, 3–10. [Google Scholar] [CrossRef] [PubMed]
- Nordmann, P.; Poirel, L.; Walsh, T.R.; Livermore, D.M. The emerging NDM carbapenemases. Trends Microbiol. 2011, 19, 588–595. [Google Scholar] [CrossRef] [PubMed]
- Voulgari, E.; Poulou, A.; Koumaki, V.; Tsakris, A. Carbapenemase-producing Enterobacteriaceae: Now that the storm is finally here, how will timely detection help us fight back? Future Microbiol. 2013, 8, 27–39. [Google Scholar] [CrossRef] [PubMed]
- Woodford, N.; Turton, J.F.; Livermore, D.M. Multiresistant Gram-negative bacteria: The role of high-risk clones in the dissemination of antibiotic resistance. FEMS Microbiol. Rev. 2011, 35, 736–755. [Google Scholar] [CrossRef] [PubMed]
- Bush, K.; Jacoby, G.A. Updated functional classification of beta-Lactamases. Antimicrob. Agents Chemother. 2010, 54, 969–976. [Google Scholar] [CrossRef] [PubMed]
- Jacoby, G.A.; Munoz-Price, L.S. Mechanisms of disease: The new beta-lactamases. N. Engl. J. Med. 2005, 352, 380–391. [Google Scholar] [CrossRef] [PubMed]
- Drawz, S.M.; Bonomo, R.A. Three decades of beta-lactamase inhibitors. Clin. Microbiol. Rev. 2010, 23, 160–201. [Google Scholar] [CrossRef] [PubMed]
- Thomson, J.M.; Bonomo, R.A. The threat of antibiotic resistance in Gram-negative pathogenic bacteria: Beta-lactams in peril! Curr. Opin. Microbiol. 2005, 8, 518–524. [Google Scholar] [CrossRef] [PubMed]
- Bush, K.; Jacoby, G.A.; Medeiros, A.A. A functional classification scheme for beta-lactamases and its correlation with molecular-structure. Antimicrob. Agents Chemother. 1995, 39, 1211–1233. [Google Scholar] [CrossRef] [PubMed]
- Garau, G.; Garcia-Saez, I.; Bebrone, C.; Anne, C.; Mercuri, P.; Galleni, M.; Frere, J.M.; Dideberg, O. Update of the standard numbering scheme for class B beta-lactamases. Antimicrob. Agents Chemother. 2004, 48, 2347–2349. [Google Scholar] [CrossRef] [PubMed]
- Lambert, P.A. Mechanisms of antibiotic resistance in Pseudomonas aeruginosa. J. R. Soc. Med. 2002, 95, 22–26. [Google Scholar] [PubMed]
- Johnson, A.P.; Woodford, N. Global spread of antibiotic resistance: The example of New Delhi metallo-beta-lactamase (NDM)-mediated carbapenem resistance. J. Med. Microbiol. 2013, 62, 499–513. [Google Scholar] [CrossRef] [PubMed]
- Livermore, D.M. Beta-lactamases in laboratory and clinical resistance. Clin. Microbiol. Rev. 1995, 8, 557–584. [Google Scholar] [PubMed]
- Bonnet, R. Growing group of extended-spectrum beta-lactamases: The CTX-M enzymes. Antimicrob. Agents Chemother. 2004, 48, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Livermore, D.M.; Woodford, N. The beta-lactamase threat in Enterobacteriaceae, Pseudomonas and Acinetobacter. Trends Microbiol. 2006, 14, 413–420. [Google Scholar] [CrossRef] [PubMed]
- Paterson, D.L.; Bonomo, R.A. Extended-spectrum beta-lactamases: A clinical update. Clin. Microbiol. Rev. 2005, 18, 657. [Google Scholar] [CrossRef] [PubMed]
- Dzidic, S.; Suskovic, J.; Kos, B. Antibiotic resistance mechanisms in bacteria: Biochemical and genetic aspects. Food Technol. Biotechnol. 2008, 46, 11–21. [Google Scholar]
- Martinez, J.L.; Baquero, F. Interactions among strategies associated with bacterial infection: Pathogenicity, epidemicity, and antibiotic resistance. Clin. Microbiol. Rev. 2002, 15, 647. [Google Scholar] [CrossRef] [PubMed]
- Hawkey, P.M. The origins and molecular basis of antibiotic resistance. Br. Med. J. 1998, 317, 657–660. [Google Scholar] [CrossRef]
- Hiramatsu, K.; Cui, L.; Kuroda, M.; Ito, T. The emergence and evolution of methicillin-resistant Staphylococcus aureus. Trends Microbiol. 2001, 9, 486–493. [Google Scholar] [CrossRef]
- Borges, A.; Saavedra, M.J.; Simões, M. Insights on antimicrobial resistance, biofilms and the use of phytochemicals as new antimicrobial agents. Curr. Med. Chem. 2015, 22, 2590–2614. [Google Scholar] [CrossRef] [PubMed]
- Stewart, P.S. Mechanisms of antibiotic resistance in bacterial biofilms. Int. J. Med. Microbiol. 2002, 292, 107–113. [Google Scholar] [CrossRef] [PubMed]
- Stewart, P.S.; Costerton, J.W. Antibiotic resistance of bacteria in biofilms. Antibiot. Resist. Bact. Biofilms 2001, 358, 135–138. [Google Scholar] [CrossRef]
- Singh, P.K.; Schaefer, A.L.; Parsek, M.R.; Moninger, T.O.; Welsh, M.J.; Greenberg, E.P. Quorum-sensing signals indicate that cystic fibrosis lungs are infected with bacterial biofilms. Nature 2000, 407, 762–764. [Google Scholar] [CrossRef] [PubMed]
- De la Fuente-Nunez, C.; Reffuveille, F.; Fernandez, L.; Hancock, R.E. Bacterial biofilm development as a multicellular adaptation: Antibiotic resistance and new therapeutic strategies. Curr. Opin. Microbiol. 2013, 16, 580–589. [Google Scholar] [CrossRef] [PubMed]
- Van, H.R.; Michiels, C.W. Biofilm formation and the food industry, a focus on the bacterial outer surface. J. Appl. Microbiol. 2010, 109, 1117–1131. [Google Scholar]
- Bordi, C.; de Bentzmann, S. Hacking into bacterial biofilms: A new therapeutic challenge. Ann. Intensive Care 2011, 1, 19. [Google Scholar] [CrossRef] [PubMed]
- Lewis, K. Riddle of biofilm resistance. Antimicrob. Agents Chemother. 2001, 45, 999–1007. [Google Scholar] [CrossRef] [PubMed]
- Mah, T.F.; O'Toole, G.A. Mechanisms of biofilm resistance to antimicrobial agents. Trends Microbiol. 2001, 9, 34–39. [Google Scholar] [CrossRef]
- Brooun, A.; Liu, S.; Lewis, K. A dose-response study of antibiotic resistance in Pseudomonas aeruginosa biofilms. Antimicrob. Agents Chemother. 2000, 44, 640–646. [Google Scholar] [CrossRef] [PubMed]
- Driffield, K.; Miller, K.; Bostock, J.M.; O’Neill, A.J.; Chopra, I. Increased mutability of Pseudomonas aeruginosa in biofilms. J. Antimicrob. Chemother. 2008, 61, 1053–1056. [Google Scholar] [CrossRef] [PubMed]
- Molin, S.; Tolker-Nielsen, T. Gene transfer occurs with enhanced efficiency in biofilms and induces enhanced stabilisation of the biofilm structure. Curr. Opin. Biotechnol. 2003, 14, 255–261. [Google Scholar] [CrossRef]
- Hoiby, N.; Bjarnsholt, T.; Givskov, M.; Molin, S.; Ciofu, O. Antibiotic resistance of bacterial biofilms. Int. J. Antimicrob. Agents 2010, 35, 322–332. [Google Scholar] [CrossRef] [PubMed]
- Donlan, R.M.; Costerton, J.W. Biofilms: Survival mechanisms of clinically relevant microorganisms. Clin. Microbiol. Rev. 2002, 15, 167–193. [Google Scholar] [CrossRef] [PubMed]
- Donlan, R.M. Biofilms: Microbial life on surfaces. Emerg. Infect. Dis. 2002, 8, 881–890. [Google Scholar] [CrossRef] [PubMed]
- Flemming, H.C.; Wingender, J. The biofilm matrix. Nat. Rev. Microbiol. 2010, 8, 623–633. [Google Scholar] [CrossRef] [PubMed]
- Suci, P.A.; Mittelman, M.W.; Yu, F.P.; Geesey, G.G. Investigation of ciprofloxacin penetration into Pseudomonas aeruginosa biofilms. J. Antimicrob. Chemother. 1994, 38, 2125–2133. [Google Scholar] [CrossRef]
- Hoyle, B.D.; Wong, C.K.; Costerton, J.W. Disparate efficacy of tobramycin on Ca(2+)-, Mg(2+)-, and HEPES-treated Pseudomonas aeruginosa biofilms. Can. J. Microbiol. 1992, 38, 1214–1218. [Google Scholar] [CrossRef] [PubMed]
- Patel, R. Biofilms and antimicrobial resistance. Clin. Orthop. Relat. Res. 2005, 437, 41–47. [Google Scholar] [CrossRef] [PubMed]
- Dunne, W.M., Jr. Bacterial adhesion: Seen any good biofilms lately? Clin. Microbiol. Rev. 2002, 15, 155–166. [Google Scholar] [CrossRef] [PubMed]
- Evans, D.J.; Allison, D.G.; Brown, M.R.; Gilbert, P. Susceptibility of Pseudomonas aeruginosa and Escherichia coli biofilms towards ciprofloxacin: Effect of specific growth rate. J. Antimicrob. Chemother. 1991, 27, 177–184. [Google Scholar] [CrossRef] [PubMed]
- Duguid, I.G.; Evans, E.; Brown, M.R.; Gilbert, P. Growth-rate-independent killing by ciprofloxacin of biofilm-derived Staphylococcus epidermidis; evidence for cell-cycle dependency. J. Antimicrob. Chemother. 1992, 30, 791–802. [Google Scholar] [CrossRef] [PubMed]
- Duguid, I.G.; Evans, E.; Brown, M.R.; Gilbert, P. Effect of biofilm culture upon the susceptibility of Staphylococcus epidermidis to tobramycin. J. Antimicrob. Chemother. 1992, 30, 803–810. [Google Scholar] [CrossRef] [PubMed]
- Lewis, K. Persister cells and the riddle of biofilm survival. Biochemesrty 2005, 70, 267–274. [Google Scholar] [CrossRef]
- Keren, I.; Kaldalu, N.; Spoering, A.; Wang, Y.; Lewis, K. Persister cells and tolerance to antimicrobials. FEMS Microbiol. Lett. 2004, 230, 13–18. [Google Scholar] [CrossRef]
- Malik, A.; Grohmann, E.; Akhtar, R. Environmental Deterioration and Human Health: Natural and Anthropogenic Determinants; Springer Science & Business Media: Heidelberg, Germany, 2013. [Google Scholar]
- Berglund, B. Environmental dissemination of antibiotic resistance genes and correlation to anthropogenic contamination with antibiotics. Infect. Ecol. Epidemiol. 2015, 5. [Google Scholar] [CrossRef] [PubMed]
- Dancer, S.J.; Shears, P.; Platt, D.J. Isolation and characterization of coliforms from glacial ice and water in Canada’s High Arctic. J. Appl. Microbiol. 1997, 82, 597–609. [Google Scholar] [CrossRef] [PubMed]
- Wright, G.D. Antibiotic resistance in the environment: A link to the clinic? Curr. Opin. Microbiol. 2010, 13, 589–594. [Google Scholar] [CrossRef] [PubMed]
- Allen, H.K.; Donato, J.; Wang, H.H.; Cloud-Hansen, K.A.; Davies, J.; Handelsman, J. Call of the wild: Antibiotic resistance genes in natural environments. Nat. Rev. Microbiol. 2010, 8, 251–259. [Google Scholar] [CrossRef] [PubMed]
- D’Costa, V.M.; McGrann, K.M.; Hughes, D.W.; Wright, G.D. Sampling the antibiotic resistome. Science 2006, 311, 374–377. [Google Scholar] [CrossRef] [PubMed]
- Forsberg, K.J.; Reyes, A.; Wang, B.; Selleck, E.M.; Sommer, M.O.A.; Dantas, G. The shared antibiotic resistome of soil bacteria and human pathogens. Science 2012, 337, 1107–1111. [Google Scholar] [CrossRef] [PubMed]
- Riesenfeld, C.S.; Goodman, R.M.; Handelsman, J. Uncultured soil bacteria are a reservoir of new antibiotic resistance genes. Environ. Microbiol. 2004, 6, 981–989. [Google Scholar] [CrossRef] [PubMed]
- Donato, J.J.; Moe, L.A.; Converse, B.J.; Smart, K.D.; Berklein, F.C.; McManus, P.S.; Handelsman, J. Metagenomic analysis of apple orchard soil reveals antibiotic resistance genes encoding predicted bifunctional proteins. Appl. Environ. Microbiol. 2010, 76, 4396–4401. [Google Scholar] [CrossRef] [PubMed]
- Lang, K.S.; Anderson, J.M.; Schwarz, S.; Williamson, L.; Handelsman, J.; Singer, R.S. Novel florfenicol and chloramphenicol resistance gene discovered in Alaskan soil by using functional metagenomics. Appl. Environ. Microbiol. 2010, 76, 5321–5326. [Google Scholar] [CrossRef] [PubMed]
- Brown, M.G.; Balkwill, D.L. Antibiotic resistance in bacteria isolated from the deep terrestrial subsurface. Microb. Ecol. 2008, 57, 484–493. [Google Scholar] [CrossRef] [PubMed]
- Chatterjee, M.; Anju, C.P.; Biswas, L.; Anil Kumar, V.; Gopi Mohan, C.; Biswas, R. Antibiotic resistance in Pseudomonas aeruginosa and alternative therapeutic options. Int. J. Med. Microbiol. 2016, 306, 48–58. [Google Scholar] [CrossRef] [PubMed]
- Demanèche, S.; Sanguin, H.; Poté, J.; Navarro, E.; Bernillon, D.; Mavingui, P.; Wildi, W.; Vogel, T.M.; Simonet, P. Antibiotic-resistant soil bacteria in transgenic plant fields. Proc. Natl. Acad. Sci. USA 2008, 105, 3957–3962. [Google Scholar] [CrossRef] [PubMed]
- Yap, P.S.X.; Yiap, B.C.; Ping, H.C.; Lim, S.H.E. Essential oils, a new horizon in combating bacterial antibiotic resistance. Open Microbiol. J. 2014, 8, 6–14. [Google Scholar] [CrossRef] [PubMed]
- Davies, J.; Davies, D. Origins and evolution of antibiotic resistance. Microbiol. Mol. Biol. Rev. 2010, 74, 417–433. [Google Scholar] [CrossRef] [PubMed]
- Toussaint, K.A.; Gallagher, J.C. β-lactam/β-lactamase inhibitor combinations: From then to now. Ann. Pharmacother. 2015, 49, 86–98. [Google Scholar] [CrossRef] [PubMed]
- Bolla, J.-M.; Alibert-Franco, S.; Handzlik, J.; Chevalier, J.; Mahamoud, A.; Boyer, G.; Kieć-Kononowicz, K.; Pagès, J.-M. Strategies for bypassing the membrane barrier in multidrug resistant Gram-negative bacteria. FEBS Lett. 2011, 585, 1682–1690. [Google Scholar] [CrossRef] [PubMed]
- Pagès, J.-M.; Amaral, L. Mechanisms of drug efflux and strategies to combat them: Challenging the efflux pump of Gram-negative bacteria. Biochim. Biophys. Acta 2009, 1794, 826–833. [Google Scholar] [CrossRef] [PubMed]
- Bhardwaj, A.K.; Mohanty, P. Bacterial efflux pumps involved in multidrug resistance and their inhibitors: Rejuvinating the antimicrobial chemotherapy. Recent Pat. Antiinfect. Drug Discov. 2012, 7, 73–89. [Google Scholar] [CrossRef] [PubMed]
- Kalan, L.; Wright, G.D. Antibiotic adjuvants: Multicomponent anti-infective strategies. Expert Rev. Mol. Med. 2011, 13, e5. [Google Scholar] [CrossRef] [PubMed]
- Zechini, B.; Versace, I. Inhibitors of multidrug resistant efflux systems in bacteria. Recent Pat. Antiinfect. Drug Discov. 2009, 4, 37–50. [Google Scholar] [CrossRef] [PubMed]
- Mahmood, H.Y.; Jamshidi, S.; Mark Sutton, J.; Rahman, K.M. Current advances in developing inhibitors of bacterial multidrug efflux pumps. Curr. Med. Chem. 2016, 23, 1062–1081. [Google Scholar] [CrossRef] [PubMed]
- Gill, E.E.; Franco, O.L.; Hancock, R. Antibiotic adjuvants: Diverse strategies for controlling drug-resistant pathogens. Chem. Biol. Drug Des. 2015, 85, 56–78. [Google Scholar] [CrossRef] [PubMed]
- Marquez, B. Bacterial efflux systems and efflux pumps inhibitors. Biochimie 2005, 87, 1137–1147. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Deng, Z.; Yan, A. Bacterial multidrug efflux pumps: Mechanisms, physiology and pharmacological exploitations. Biochem. Biophys. Res. Commun. 2014, 453, 254–267. [Google Scholar] [CrossRef] [PubMed]
- Cragg, G.M.; Newman, D.J. Natural products: A continuing source of novel drug leads. Biochim. Biophys. Acta 2013, 1830, 3670–3695. [Google Scholar] [CrossRef] [PubMed]
- Ettefagh, K.A.; Burns, J.T.; Junio, H.A.; Kaatz, G.W.; Cech, N.B. Goldenseal (Hydrastis canadensis L.) extracts synergistically enhance the antibacterial activity of berberine via efflux pump inhibition. Planta Med. 2011, 77, 835–840. [Google Scholar] [CrossRef] [PubMed]
- Berdy, J. Thoughts and facts about antibiotics: Where we are now and where we are heading. J. Antibiot. 2012, 65, 385–395. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Di, H.; Wang, Y.; Cao, Q.; Xu, B.; Zhang, X.; Yang, N.; Liu, G.; Yang, C.-G.; Xu, Y.; et al. Small-molecule targeting of a diapophytoene desaturase inhibits S. aureus virulence. Nat. Chem. Biol. 2016, 12, 174–179. [Google Scholar] [CrossRef] [PubMed]
- Aiyegoro, O.A.; Okoh, A.I. Use of bioactive plant products in combination with standard antibiotics: Implications in antimicrobial chemotherapy. J. Med. Plants Res. 2009, 3, 1147–1152. [Google Scholar]
- Wagner, H.; Ulrich-Merzenich, G. Synergy research: Approaching a new generation of phytopharmaceuticals. Phytomedicine 2009, 16, 97–110. [Google Scholar] [CrossRef] [PubMed]
- Abreu, A.C.; McBain, A.J.; Simoes, M. Plants as sources of new antimicrobials and resistance-modifying agents. Nat. Prod. Rep. 2012, 29, 1007–1021. [Google Scholar] [CrossRef] [PubMed]
- Sibanda, T.; Okoh, A.I. The challenges of overcoming antibiotic resistance: Plant extracts as potential sources of antimicrobial and resistance modifying agents. Afr. J. Biotechnol. 2007, 6, 2886–2896. [Google Scholar]
- Betoni, J.E.C.; Mantovani, R.P.; Barbosa, L.N.; Di Stasi, L.C.; Fernandes Junior, A. Synergism between plant extract and antimicrobial drugs used on Staphylococcus aureus diseases. Mem. Inst. Oswaldo Cruz 2006, 101, 387–390. [Google Scholar] [CrossRef] [PubMed]
- Jayaraman, P.; Sakharkar, M.K.; Lim, C.S.; Tang, T.H.; Sakharkar, K.R. Activity and interactions of antibiotic and phytochemical combinations against Pseudomonas aeruginosa in vitro. Int. J. Biol. Sci. 2010, 6, 556–568. [Google Scholar] [CrossRef] [PubMed]
- Abreu, A.C.; Serra, S.C.; Borges, A.; Saavedra, M.J.; McBain, A.J.; Salgado, A.J.; Simões, M. Combinatorial activity of flavonoids with antibiotics against drug-resistant Staphylococcus aureus. Microb. Drug Res. 2015, 21, 600–609. [Google Scholar] [CrossRef] [PubMed]
- Hoffman, S.B. Mechanisms of antibiotic resistance. Compendium 2001, 23, 464–473. [Google Scholar]
- Stavri, M.; Piddock, L.J.V.; Gibbons, S. Bacterial efflux pump inhibitors from natural sources. J. Antimicrob. Chemother. 2007, 59, 1247–1260. [Google Scholar] [CrossRef] [PubMed]
- Savoia, D. Plant-derived antimicrobial compounds: Alternatives to antibiotics. Future Microbiol. 2012, 7, 979–990. [Google Scholar] [CrossRef] [PubMed]
- Garvey, M.I.; Rahman, M.M.; Gibbons, S.; Piddock, L.J.V. Medicinal plant extracts with efflux inhibitory activity against Gram-negative bacteria. Int. J. Antimicrob. Agents 2011, 37, 145–151. [Google Scholar] [CrossRef] [PubMed]
- Darwish, R.M.; Aburjai, T.; Al-Khalil, S.; Mahafzah, A. Screening of antibiotic resistant inhibitors from local plant materials against two different strains of Staphylococcus aureus. J. Ethnopharmacol. 2002, 79, 359–364. [Google Scholar] [CrossRef]
- Ahmad, I.; Aqil, F. In vitro efficacy of bioactive extracts of 15 medicinal plants against ESβL-producing multidrug-resistant enteric bacteria. Microbiol. Res. 2007, 162, 264–275. [Google Scholar] [CrossRef] [PubMed]
- Touani, F.K.; Seukep, A.J.; Djeussi, D.E.; Fankam, A.G.; Noumedem, J.A.K.; Kuete, V. Antibiotic-potentiation activities of four Cameroonian dietary plants against multidrug-resistant Gram-negative bacteria expressing efflux pumps. BMC Complement. Altern. Med. 2014, 14, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Barreto, H.M.; Coelho, K.M.R.N.; Ferreira, J.H.L.; dos Santos, B.H.C.; de Abreu, A.P.L.; Coutinho, H.D.M.; da Silva, R.A.C.; de Sousa, T.O.; Citó, A.M.D.G.L.; Lopes, J.A.D. Enhancement of the antibiotic activity of aminoglycosides by extracts from Anadenanthera colubrine (Vell.) Brenan var. cebil against multi-drug resistant bacteria. Nat. Prod. Res. 2015, 1–4. [Google Scholar] [CrossRef]
- Barreto, H.M.; de Lima, I.S.; Coelho, K.M.R.N.; Osório, L.R.; de Almeida Mourão, R.; Santos, B.H.C.D.; Coutinho, H.D.M.; de Abreu, A.P.L.; de Medeiros, M.D.G.F.; Citó, A.M.D.G.L.; et al. Effect of Lippia origanoides H.B.K. essential oil in the resistance to aminoglycosides in methicillin resistant Staphylococcus aureus. Eur. J. Integr. Med. 2014, 6, 560–564. [Google Scholar] [CrossRef]
- Aumeeruddy-Elalfi, Z.; Gurib-Fakim, A.; Mahomoodally, F. Antimicrobial, antibiotic potentiating activity and phytochemical profile of essential oils from exotic and endemic medicinal plants of Mauritius. Ind. Crop. Prod. 2015, 71, 197–204. [Google Scholar] [CrossRef]
- Fankam, A.G.; Kuiate, J.R.; Kuete, V. Antibacterial and antibiotic resistance modifying activity of the extracts from Alanblackia gabonensis, Combretum molle and Gladiolus quartinianus against Gram-negative bacteria including multi-drug resistant phenotypes. BMC Complement. Altern. Med. 2015, 15, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Tankeo, S.B.; Tane, P.; Kuete, V. In vitro antibacterial and antibiotic-potentiation activities of the methanol extracts from Beilschmiedia acuta, Clausena anisata, Newbouldia laevis and Polyscias fulva against multidrug-resistant Gram-negative bacteria. BMC Complement. Altern. Med. 2015, 15, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Alves, M.J.; Ferreira, I.C.F.R.; Lourenço, I.; Castro, A.; Pereira, L.; Martins, A.; Pintado, M. Wild mushroom extracts potentiate the action of standard antibiotics against multiresistant bacteria. J. Appl. Microbiol. 2014, 116, 32–38. [Google Scholar] [CrossRef] [PubMed]
- Bjarnsholt, T.; Ciofu, O.; Molin, S.; Givskov, M.; Høiby, N. Applying insights from biofilm biology to drug development - can a new approach be developed? Nat. Rev. Drug Discov. 2013, 12, 791–808. [Google Scholar] [CrossRef] [PubMed]
- Lynch, A.S.; Abbanat, D. New antibiotic agents and approaches to treat biofilm-associated infections. Expert Opin. Ther. Pat. 2010, 20, 1373–1387. [Google Scholar] [CrossRef] [PubMed]
- Francolini, I.; Donelli, G. Prevention and control of biofilm-based medical-device-related infections. FEMS Immunol. Med. Microbiol. 2010, 59, 227–238. [Google Scholar] [CrossRef] [PubMed]
- Simões, M.; Bennett, R.N.; Rosa, E.A. Understanding antimicrobial activities of phytochemicals against multidrug resistant bacteria and biofilms. Nat. Prod. Rep. 2009, 26, 746–757. [Google Scholar] [CrossRef] [PubMed]
- Blackledge, M.S.; Worthington, R.J.; Melander, C. Biologically inspired strategies for combating bacterial biofilms. Curr. Opin. Pharmacol. 2013, 13, 699–706. [Google Scholar] [CrossRef] [PubMed]
- Lebeaux, D.; Ghigo, J.-M.; Beloin, C. Biofilm-related infections: Bridging the gap between clinical management and fundamental aspects of recalcitrance toward antibiotics. Microbiol. Mol. Biol. Rev. 2014, 78, 510–543. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Moser, C.; Wang, H.-Z.; Høiby, N.; Song, Z.-J. Strategies for combating bacterial biofilm infections. Int. J. Oral Sci. 2015, 7, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Taraszkiewicz, A.; Fila, G.; Grinholc, M.; Nakonieczna, J. Innovative strategies to overcome biofilm resistance. Biomed. Res. Int. 2013, 2013, 13. [Google Scholar] [CrossRef] [PubMed]
- Oppenheimer-Shaanan, Y.; Steinberg, N.; Kolodkin-Gal, I. Small molecules are natural triggers for the disassembly of biofilms. Trends Microbiol. 2013, 21, 594–601. [Google Scholar] [CrossRef] [PubMed]
- Smith, A.W. Biofilms and antibiotic therapy: Is there a role for combating bacterial resistance by the use of novel drug delivery systems? Adv. Drug Deliv. Rev. 2005, 57, 1539–1550. [Google Scholar] [CrossRef] [PubMed]
- Martin, C.; LiLow, W.; Gupta, A.; Cairul Iqbal Mohd Amin, M.; Radecka, I.; T Britland, S.; Raj, P. Strategies for antimicrobial drug delivery to biofilm. Curr. Pharm. Des. 2015, 21, 43–66. [Google Scholar] [CrossRef] [PubMed]
- LaSarre, B.; Federle, M.J. Exploiting quorum sensing to confuse bacterial pathogens. Microbiol. Mol. Biol. Rev. 2013, 77, 73–111. [Google Scholar] [CrossRef] [PubMed]
- Rasko, D.A.; Sperandio, V. Anti-virulence strategies to combat bacteria-mediated disease. Nat. Rev. Drug Discov. 2010, 9, 117–128. [Google Scholar] [CrossRef] [PubMed]
- Jakobsen, T.H.; van Gennip, M.; Phipps, R.K.; Shanmugham, M.S.; Christensen, L.D.; Alhede, M.; Skindersoe, M.E.; Rasmussen, T.B.; Friedrich, K.; Uthe, F.; et al. Ajoene, a sulfur-rich molecule from garlic, inhibits genes controlled by quorum sensing. Antimicrob. Agents Chemother. 2012, 56, 2314–2325. [Google Scholar] [CrossRef] [PubMed]
- Borges, A.; Serra, S.; Cristina Abreu, A.; Saavedra, M.J.; Salgado, A.; Simões, M. Evaluation of the effects of selected phytochemicals on quorum sensing inhibition and in vitro cytotoxicity. Biofouling 2014, 30, 183–195. [Google Scholar] [CrossRef] [PubMed]
- Qian, P.-Y.; Chen, L.; Xu, Y. Mini-review: Molecular mechanisms of antifouling compounds. Biofouling 2013, 29, 381–400. [Google Scholar] [CrossRef] [PubMed]
- Jakobsen, T.H.; Bragason, S.K.; Phipps, R.K.; Christensen, L.D.; van Gennip, M.; Alhede, M.; Skindersoe, M.; Larsen, T.O.; Høiby, N.; Bjarnsholt, T.; et al. Food as a source for quorum sensing inhibitors: Iberin from horseradish revealed as a quorum sensing inhibitor of Pseudomonas aeruginosa. Appl. Environ. Microbiol. 2012, 78, 2410–2421. [Google Scholar] [CrossRef] [PubMed]
- Welsh, M.A.; Eibergen, N.R.; Moore, J.D.; Blackwell, H.E. Small molecule disruption of quorum sensing cross-regulation in Pseudomonas aeruginosa causes major and unexpected alterations to virulence phenotypes. J. Am. Chem. Soc. 2015, 137, 1510–1519. [Google Scholar] [CrossRef] [PubMed]
- Kalia, V.C. Quorum sensing inhibitors: An overview. Biotechnol. Adv. 2013, 31, 224–245. [Google Scholar] [CrossRef] [PubMed]
- Davies, D.G.; Parsek, M.R.; Pearson, J.P.; Iglewski, B.H.; Costerton, J.W.; Greenberg, E.P. The involvement of cell-to-cell signals in the development of a bacterial biofilm. Science 1998, 280, 295–298. [Google Scholar] [CrossRef] [PubMed]
- Jayaraman, A.; Wood, T.K. Bacterial quorum sensing: Signals, circuits, and implications for biofilms and disease. Annu. Rev. Biomed. Eng. 2008, 10, 145–167. [Google Scholar] [CrossRef] [PubMed]
- Bjarnsholt, T.; Givskov, M. Quorum-sensing blockade as a strategy for enhancing host defences against bacterial pathogens. Philos. Trans. R. Soc. B Biol. Sci. 2007, 362, 1213–1222. [Google Scholar] [CrossRef] [PubMed]
- Pan, J.; Ren, D. Quorum sensing inhibitors: A patent overview. Expert Opin. Ther. Pat. 2009, 19, 1581–1601. [Google Scholar] [CrossRef] [PubMed]
- Castillo-Juárez, I.; Maeda, T.; Mandujano-Tinoco, E.A.; Tomás, M.; Pérez-Eretza, B.; García-Contreras, S.J.; Wood, T.K.; García-Contreras, R. Role of quorum sensing in bacterial infections. World J. Clin. Cases 2015, 3, 575. [Google Scholar] [PubMed]
- Vattem, D.A.; Mihalik, K.; Crixell, S.H.; McLean, R.J.C. Dietary phytochemicals as quorum sensing inhibitors. Fitoterapia 2007, 78, 302–310. [Google Scholar] [CrossRef] [PubMed]
- Rasmussen, T.B.; Givskov, M. Quorum-sensing inhibitors as anti-pathogenic drugs. Int. J. Med. Microbiol. 2006, 296, 149–161. [Google Scholar] [CrossRef] [PubMed]
- Musthafa, K.S.; Ravi, A.V.; Annapoorani, A.; Packiavathy, I.S.V.; Pandian, S.K. Evaluation of anti-quorum-sensing activity of edible plants and fruits through inhibition of the N-acyl-homoserine lactone system in Chromobacterium violaceum and Pseudomonas aeruginosa. Chemotherapy 2010, 56, 333–339. [Google Scholar] [CrossRef] [PubMed]
- Koh, K.H.; Tham, F.Y. Screening of traditional Chinese medicinal plants for quorum-sensing inhibitors activity. J. Microbiol. Immunol. Infect. 2011, 44, 144–148. [Google Scholar] [CrossRef] [PubMed]
- Issac Abraham, S.V.P.; Palani, A.; Ramaswamy, B.R.; Shunmugiah, K.P.; Arumugam, V.R. Antiquorum sensing and antibiofilm potential of Capparis spinosa. Arch. Med. Res. 2011, 42, 658–668. [Google Scholar] [CrossRef] [PubMed]
- Quave, C.L.; Plano, L.R.; Bennett, B.C. Quorum sensing inhibitors of Staphylococcus aureus from Italian medicinal plants. Planta Med. 2011, 77, 188–195. [Google Scholar] [CrossRef] [PubMed]
- Al-Sohaibani, S.; Murugan, K. Anti-biofilm activity of Salvadora persica on cariogenic isolates of Streptococcus mutans: In vitro and molecular docking studies. Biofouling 2012, 28, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Hasan, S.; Danishuddin, M.; Adil, M.; Singh, K.; Verma, P.K.; Khan, A.U. Efficacy of E. officinalis on the cariogenic properties of Streptococcus mutans: A novel and alternative approach to suppress quorum-sensing mechanism. PLoS ONE 2012, 7, e40319. [Google Scholar] [CrossRef] [PubMed]
- Masurkar, S.; Chaudhari, P.; Shidore, V.; Kamble, S. Effect of biologically synthesised silver nanoparticles on Staphylococcus aureus biofilm quenching and prevention of biofilm formation. Int. J. Pharm. Bio. Sci. 2012, 6, 110–114. [Google Scholar] [CrossRef] [PubMed]
- Cech, N.B.; Junio, H.A.; Ackermann, L.W.; Kavanaugh, J.S.; Horswill, A.R. Quorum quenching and antimicrobial activity of goldenseal (Hydrastis canadensis) against methicillin-resistant Staphylococcus aureus (MRSA). Planta Med. 2012, 78, 1556–1561. [Google Scholar] [CrossRef] [PubMed]
- Murugan, K.; Sekar, K.; Sangeetha, S.; Ranjitha, S.; Sohaibani, S. Antibiofilm and quorum sensing inhibitory activity of Achyranthes aspera on cariogenic Streptococcus mutans: An in vitro and in silico study. Pharm. Biol. 2013, 51, 728–736. [Google Scholar] [CrossRef] [PubMed]
- Castillo-Juárez, I.; García-Contreras, R.; Velázquez-Guadarrama, N.; Soto-Hernández, M.; Martínez-Vázquez, M. Amphypterygium adstringens anacardic acid mixture inhibits quorum sensing-controlled virulence factors of Chromobacterium violaceum and Pseudomonas aeruginosa. Arch. Med. Res. 2013, 44, 488–494. [Google Scholar] [CrossRef] [PubMed]
- Damte, D.; Gebru, E.; Lee, S.-J.; Suh, J.-W.; Park, S.-C. Evaluation of anti-quorum sensing activity of 97 indigenous plant extracts from Korea through bioreporter bacterial strains Chromobacterium violaceum and Pseudomonas aeruginosa. J. Microb. Biochem. Technol. 2013, 2013. [Google Scholar] [CrossRef]
- Sarabhai, S.; Sharma, P.; Capalash, N. Ellagic acid derivatives from Terminalia chebula Retz. downregulate the expression of quorum sensing genes to attenuate Pseudomonas aeruginosa PAO1 virulence. PLoS ONE 2013, 8, e53441. [Google Scholar] [CrossRef] [PubMed]
- Rasamiravaka, T.; Jedrzejowski, A.; Kiendrebeogo, M.; Rajaonson, S.; Randriamampionona, D.; Rabemanantsoa, C.; Andriantsimahavandy, A.; Rasamindrakotroka, A.; Duez, P.; El Jaziri, M. Endemic Malagasy Dalbergia species inhibit quorum sensing in Pseudomonas aeruginosa PAO1. Microbiology 2013, 159, 924–938. [Google Scholar] [CrossRef] [PubMed]
- Vasavi, H.S.; Arun, A.B.; Rekha, P.D. Inhibition of quorum sensing in Chromobacterium violaceum by Syzygium cumini L. and Pimenta dioica L. Asian Pac. J. Trop. Biomed. 2013, 3, 954–959. [Google Scholar] [CrossRef]
- Vasavi, H.S.; Arun, A.B.; Rekha, P.-D. Anti-quorum sensing activity of Psidium guajava L. flavonoids against Chromobacterium violaceum and Pseudomonas aeruginosa PAO1. Microbiol. Immunol. 2014, 58, 286–293. [Google Scholar] [CrossRef] [PubMed]
- Vasavi, H.S.; Arun, A.B.; Rekha, P.D. Anti-quorum sensing activity of flavonoid-rich fraction from Centella asiatica L. against Pseudomonas aeruginosa PAO1. J. Microbiol. Immunol. Infect. 2014, 49, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Vasavi, H.S.; Arun, A.B.; Rekha, P.-D. Anti-quorum sensing potential of Adenanthera pavonina. Pharmacogn. Res. 2015, 7, 105. [Google Scholar]
- Zhang, J.; Rui, X.; Wang, L.; Guan, Y.; Sun, X.; Dong, M. Polyphenolic extract from Rosa rugosa tea inhibits bacterial quorum sensing and biofilm formation. Food Control 2014, 42, 125–131. [Google Scholar] [CrossRef]
- Ta, C.A.; Freundorfer, M.; Mah, T.-F.; Otárola-Rojas, M.; Garcia, M.; Sanchez-Vindas, P.; Poveda, L.; Maschek, J.A.; Baker, B.J.; Adonizio, A.L. Inhibition of bacterial quorum sensing and biofilm formation by extracts of neotropical rainforest plants. Planta Med. 2014, 80, 343–350. [Google Scholar] [CrossRef] [PubMed]
- González-Ortiz, G.; Quarles Van Ufford, H.; Halkes, S.B.A.; Cerdà-Cuéllar, M.; Beukelman, C.J.; Pieters, R.J.; Liskamp, R.M.; Pérez, J.F.; Martín-Orue, S.M. New properties of wheat bran: Anti-biofilm activity and interference with bacteria quorum-sensing systems. Environ. Microbiol. 2014, 16, 1346–1353. [Google Scholar] [CrossRef] [PubMed]
- Kazemian, H.; Ghafourian, S.; Heidari, H.; Amiri, P.; Yamchi, J.K.; Shavalipour, A.; Houri, H.; Maleki, A.; Sadeghifard, N. Antibacterial, anti-swarming and anti-biofilm formation activities of Chamaemelum nobile against Pseudomonas aeruginosa. Rev. Soc. Bras. Med. Trop. 2015, 48, 432–436. [Google Scholar] [CrossRef] [PubMed]
- Sarkar, R.; Mondal, C.; Bera, R.; Chakraborty, S.; Barik, R.; Roy, P.; Kumar, A.; Yadav, K.K.; Choudhury, J.; Chaudhary, S.K. Antimicrobial properties of Kalanchoe blossfeldiana: A focus on drug resistance with particular reference to quorum sensing-mediated bacterial biofilm formation. J. Pharm. Pharmacol. 2015, 67, 951–962. [Google Scholar] [CrossRef] [PubMed]
- Rahman, M.R.T.; Lou, Z.; Yu, F.; Wang, P.; Wang, H. Anti-quorum sensing and anti-biofilm activity of Amomum tsaoko (Amommum tsao-ko Crevost et Lemarie) on foodborne pathogens. Saudi J. Biol. Sci. 2015. [Google Scholar] [CrossRef]
- Salini, R.; Pandian, S.K. Interference of quorum sensing in urinary pathogen Serratia marcescens by Anethum graveolens. Pathog. Dis. 2015, 73, ftv038. [Google Scholar] [CrossRef] [PubMed]
- Mutungwa, W.; Alluri, N.; Majumdar, M. Anti-quorum sensing activity of some commonly used traditional indian spices. Int. J. Pharm. Pharm. Sci. 2015, 7, 80–83. [Google Scholar]
- Quave, C.L.; Lyles, J.T.; Kavanaugh, J.S.; Nelson, K.; Parlet, C.P.; Crosby, H.A.; Heilmann, K.P.; Horswill, A.R. Castanea sativa (European chestnut) leaf extracts rich in ursene and oleanene derivatives block Staphylococcus aureus virulence and pathogenesis without detectable resistance. PLoS ONE 2015, 10, e0136486. [Google Scholar] [CrossRef] [PubMed]
- Tan, X.; Yang, D.; Yang, G.; Chen, J.; Dong, W.; Shi, J.; Jia, A. The investigation of inhibiting quorum sensing and methicillin-resistant Staphylococcus aureus biofilm formation from Liriodendron hybrid. Pak. J. Pharm. Sci. 2015, 28, 903–908. [Google Scholar] [PubMed]
- Singh, B.N.; Prateeksha; Pandey, G.; Jadaun, V.; Singh, S.; Bajpai, R.; Nayaka, S.; Naqvi, A.H.; Singh Rawat, A.K.; Upreti, D.K.; et al. Development and characterization of a novel Swarna-based herbo-metallic colloidal nano-formulation - inhibitor of Streptococcus mutans quorum sensing. RSC Adv. 2015, 5, 5809–5822. [Google Scholar] [CrossRef]
- Hasan, S.; Danishuddin, M.; Khan, A.U. Inhibitory effect of Zingiber officinale towards Streptococcus mutans virulence and caries development: In vitro and in vivo studies. BMC Microbiol. 2015, 15, 1. [Google Scholar] [CrossRef] [PubMed]
- Bhargava, N.; Singh, S.P.; Sharma, A.; Sharma, P.; Capalash, N. Attenuation of quorum sensing-mediated virulence of Acinetobacter baumannii by Glycyrrhiza glabra flavonoids. Future Microbiol. 2015, 10, 1953–1968. [Google Scholar] [CrossRef] [PubMed]
- Shukla, V.; Bhathena, Z. Broad spectrum anti-quorum sensing activity of tannin-rich crude extracts of indian medicinal plants. Scientifica 2016, 2016, 8. [Google Scholar] [CrossRef] [PubMed]
- Datta, S.; Jana, D.; Maity, T.R.; Samanta, A.; Banerjee, R. Piper betle leaf extract affects the quorum sensing and hence virulence of Pseudomonas aeruginosa PAO1. 3 Biotech 2016, 6, 1–6. [Google Scholar] [CrossRef]
- Oliveira, B.D.Á.; Rodrigues, A.C.; Cardoso, B.M.I.; Ramos, A.L.C.C.; Bertoldi, M.C.; Taylor, J.G.; da Cunha, L.R.; Pinto, U.M. Antioxidant, antimicrobial and anti-quorum sensing activities of Rubus rosaefolius phenolic extract. Ind. Crops Prod. 2016, 84, 59–66. [Google Scholar] [CrossRef]
- Bezek, K.; Kurinčič, M.; Knauder, E.; Klančnik, A.; Raspor, P.; Bucar, F.; Smole Možina, S. Attenuation of adhesion, biofilm formation and quorum sensing of Campylobacter jejuni by Euodia ruticarpa. Phytother. Res. 2016. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Liu, Y.; Sternberg, C.; Molin, S. Evaluation of enoyl-acyl carrier protein reductase inhibitors as Pseudomonas aeruginosa quorum-quenching reagents. Molecules 2010, 15, 780–792. [Google Scholar] [CrossRef] [PubMed]
- Vandeputte, O.M.; Kiendrebeogo, M.; Rajaonson, S.; Diallo, B.; Mol, A.; El Jaziri, M.; Baucher, M. Identification of catechin as one of the flavonoids from Combretum albiflorum bark extract that reduces the production of quorum-sensing-controlled virulence factors in Pseudomonas aeruginosa PAO1. J. Appl. Microbiol. 2010, 76, 243–253. [Google Scholar] [CrossRef] [PubMed]
- Matsunaga, T.; Nakahara, A.; Minnatul, K.M.; Noiri, Y.; Ebisu, S.; Kato, A.; Azakami, H. The inhibitory effects of catechins on biofilm formation by the periodontopathogenic bacterium, Eikenella corrodens. Biosci. Biotechnol. Biochem. 2010, 74, 2445–2450. [Google Scholar] [CrossRef] [PubMed]
- Song, Z.; Kong, K.; Wu, H.; Maricic, N.; Ramalingam, B.; Priestap, H.; Schneper, L.; Quirke, J.; Høiby, N.; Mathee, K. Panax ginseng has anti-infective activity against opportunistic pathogen Pseudomonas aeruginosa by inhibiting quorum sensing, a bacterial communication process critical for establishing infection. Phytomedicine 2010, 17, 1040–1046. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Lee, B.; Yang, L.; Wang, H.; Givskov, M.; Molin, S.; Høiby, N.; Song, Z. Effects of ginseng on Pseudomonas aeruginosa motility and biofilm formation. FEMS Immunol. Med. Microbiol. 2011, 62, 49–56. [Google Scholar] [CrossRef] [PubMed]
- Brackman, G.; Cos, P.; Maes, L.; Nelis, H.J.; Coenye, T. Quorum sensing inhibitors increase the susceptibility of bacterial biofilms to antibiotics in vitro and in vivo. Antimicrob. Agents Chemother. 2011, 55, 2655–2661. [Google Scholar] [CrossRef] [PubMed]
- Ding, X.; Yin, B.; Qian, L.; Zeng, Z.; Yang, Z.; Li, H.; Lu, Y.; Zhou, S. Screening for novel quorum-sensing inhibitors to interfere with the formation of Pseudomonas aeruginosa biofilm. J. Med. Microbiol. 2011, 60, 1827–1834. [Google Scholar] [CrossRef] [PubMed]
- Christensen, L.D.; van Gennip, M.; Jakobsen, T.H.; Alhede, M.; Hougen, H.P.; Høiby, N.; Bjarnsholt, T.; Givskov, M. Synergistic antibacterial efficacy of early combination treatment with tobramycin and quorum-sensing inhibitors against Pseudomonas aeruginosa in an intraperitoneal foreign-body infection mouse model. J. Antimicrob. Chemother. 2012, 67, 1198–1206. [Google Scholar] [CrossRef] [PubMed]
- Packiavathy, I.A.S.V.; Agilandeswari, P.; Musthafa, K.S.; Karutha Pandian, S.; Veera Ravi, A. Antibiofilm and quorum sensing inhibitory potential of Cuminum cyminum and its secondary metabolite methyl eugenol against Gram negative bacterial pathogens. Food Res. Int. 2012, 45, 85–92. [Google Scholar] [CrossRef]
- Annapoorani, A.; Umamageswaran, V.; Parameswari, R.; Pandian, S.K.; Ravi, A.V. Computational discovery of putative quorum sensing inhibitors against LasR and RhlR receptor proteins of Pseudomonas aeruginosa. J. Comput. Aided Mol. Des. 2012, 26, 1067–1077. [Google Scholar] [CrossRef] [PubMed]
- Packiavathy, I.; Sasikumar, P.; Pandian, S.; Veera Ravi, A. Prevention of quorum-sensing-mediated biofilm development and virulence factors production in Vibrio spp. by curcumin. Appl. Microbiol. Biotechnol. 2013, 97, 10177–10187. [Google Scholar] [CrossRef] [PubMed]
- Packiavathy, I.A.S.V.; Priya, S.; Pandian, S.K.; Ravi, A.V. Inhibition of biofilm development of uropathogens by curcumin—An anti-quorum sensing agent from Curcuma longa. Food Chem. 2014, 148, 453–460. [Google Scholar] [CrossRef] [PubMed]
- Brango-Vanegas, J.; Costa, G.M.; Ortmann, C.F.; Schenkel, E.P.; Reginatto, F.H.; Ramos, F.A.; Arévalo-Ferro, C.; Castellanos, L. Glycosylflavonoids from Cecropia pachystachya Trécul are quorum sensing inhibitors. Phytomedicine 2014, 21, 670–675. [Google Scholar] [CrossRef] [PubMed]
- Lemos, M.; Borges, A.; Teodósio, J.; Araújo, P.; Mergulhão, F.; Melo, L.; Simões, M. The effects of ferulic and salicylic acids on Bacillus cereus and Pseudomonas fluorescens single- and dual-species biofilms. Int. Biodeterior. Biodegrad. 2014, 86 Part A, 42–51. [Google Scholar] [CrossRef]
- Chang, C.-Y.; Krishnan, T.; Wang, H.; Chen, Y.; Yin, W.-F.; Chong, Y.-M.; Tan, L.Y.; Chong, T.M.; Chan, K.-G. Non-antibiotic quorum sensing inhibitors acting against N-acyl homoserine lactone synthase as druggable target. Sci. Rep. 2014, 4. [Google Scholar] [CrossRef] [PubMed]
- Vijendra Kumar, N.; Murthy, P.S.; Manjunatha, J.R.; Bettadaiah, B.K. Synthesis and quorum sensing inhibitory activity of key phenolic compounds of ginger and their derivatives. Food Chem. 2014, 159, 451–457. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.-S.; Lee, S.-H.; Byun, Y.; Park, H.-D. 6-Gingerol reduces Pseudomonas aeruginosa biofilm formation and virulence via quorum sensing inhibition. Sci. Rep. 2015, 5. [Google Scholar] [CrossRef] [PubMed]
- Kumar, L.; Chhibber, S.; Kumar, R.; Kumar, M.; Harjai, K. Zingerone silences quorum sensing and attenuates virulence of Pseudomonas aeruginosa. Fitoterapia 2015, 102, 84–95. [Google Scholar] [CrossRef] [PubMed]
- Kumar, L.; Chhibber, S.; Harjai, K. Zingerone inhibit biofilm formation and improve antibiofilm efficacy of ciprofloxacin against Pseudomonas aeruginosa PAO1. Fitoterapia 2013, 90, 73–78. [Google Scholar] [CrossRef] [PubMed]
- Gilabert, M.; Marcinkevicius, K.; Andujar, S.; Schiavone, M.; Arena, M.E.; Bardón, A. Sesqui- and triterpenoids from the liverwort Lepidozia chordulifera inhibitors of bacterial biofilm and elastase activity of human pathogenic bacteria. Phytomedicine 2015, 22, 77–85. [Google Scholar] [CrossRef] [PubMed]
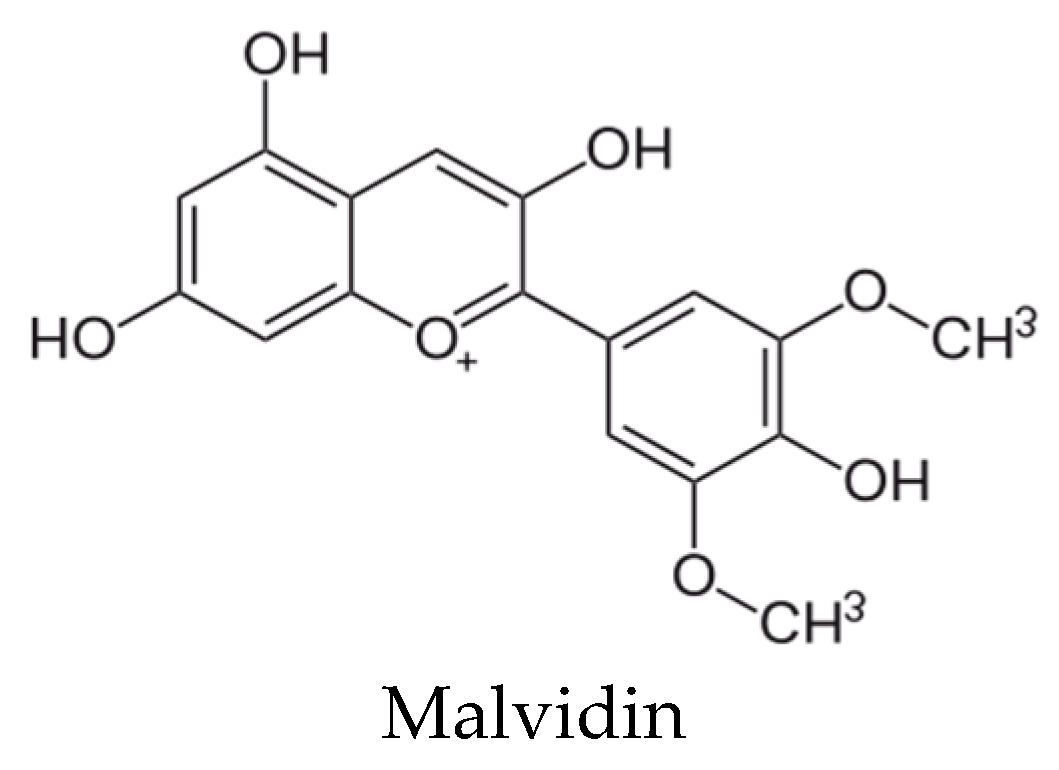
- Gopu, V.; Kothandapani, S.; Shetty, P.H. Quorum quenching activity of Syzygium cumini (L.) Skeels and its anthocyanin malvidin against Klebsiella pneumoniae. Microb. Pathog. 2015, 79, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Brackman, G.; Breyne, K.; De Rycke, R.; Vermote, A.; van Nieuwerburgh, F.; Meyer, E.; Van Calenbergh, S.; Coenye, T. The Quorum sensing inhibitor hamamelitannin increases antibiotic susceptibility of Staphylococcus aureus biofilms by affecting peptidoglycan biosynthesis and eDNA release. Sci. Rep. 2016, 6, 20321. [Google Scholar] [CrossRef] [PubMed]
- Husain, F.M.; Ahmad, I.; Asif, M.; Tahseen, Q. Influence of clove oil on certain quorum-sensing-regulated functions and biofilm of Pseudomonas aeruginosa and Aeromonas hydrophila. J. Biosci. 2013, 38, 835–844. [Google Scholar] [CrossRef] [PubMed]
- Bai A, J.; Vittal, R.R. Quorum sensing inhibitory and anti-biofilm activity of essential oils and their in vivo efficacy in food systems. Food Biotechnol. 2014, 28, 269–292. [Google Scholar] [CrossRef]
- Kalia, M.; Yadav, V.K.; Singh, P.K.; Sharma, D.; Pandey, H.; Narvi, S.S.; Agarwal, V. Effect of cinnamon oil on quorum sensing-controlled virulence factors and biofilm formation in Pseudomonas aeruginosa. PLoS ONE 2015, 10, e0135495. [Google Scholar] [CrossRef] [PubMed]
- Husain, F.M.; Ahmad, I.; Khan, M.S.; Ahmad, E.; Tahseen, Q.; Khan, M.S.; Alshabib, N.A. Sub-MICs of Mentha piperita essential oil and menthol inhibits AHL mediated quorum sensing and biofilm of Gram-negative bacteria. Front. Microbiol. 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Jayalekshmi, H.; Omanakuttan, A.; Pandurangan, N.; Vargis, V.S.; Maneesh, M.; Nair, B.G.; Kumar, G.B. Clove bud oil reduces kynurenine and inhibits pqs A gene expression in P. aeruginosa. Appl. Microbiol. Biotechnol. 2016, 100, 3681–3692. [Google Scholar]
- Zhou, L.; Zheng, H.; Tang, Y.; Yu, W.; Gong, Q. Eugenol inhibits quorum sensing at sub-inhibitory concentrations. Biotechnol. Lett. 2013, 35, 631–637. [Google Scholar] [CrossRef] [PubMed]
- Burt, S.A.; Ojo-Fakunle, V.T.A.; Woertman, J.; Veldhuizen, E.J.A. The natural antimicrobial carvacrol inhibits quorum sensing in Chromobacterium violaceum and reduces bacterial biofilm formation at sub-lethal concentrations. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Borges, A.; Abreu, A.; Malheiro, J.; Saavedra, M.J.; Simões, M. Biofilm prevention and control by dietary phytochemicals. In Microbial Pathogens and Strategies for Combating Them: Science, Technology and Education, 2013th ed.; Microbiology Book Series; Méndez-Vilas, A., Ed.; Formatex Research Center: Badajoz, Spain, 2013; Volume 1, pp. 32–41, ISBN-13: 978-84-939843-9-7. [Google Scholar]
- Bjarnsholt, T.; Jensen, P.; Burmølle, M.; Hentzer, M.; Haagensen, J.A.; Hougen, H.P.; Calum, H.; Madsen, K.G.; Moser, C.; Molin, S.; et al. Pseudomonas aeruginosa tolerance to tobramycin, hydrogen peroxide and polymorphonuclear leukocytes is quorum-sensing dependent. Microbiology 2005, 151, 373–383. [Google Scholar] [PubMed]
- Bjarnsholt, T.; Jensen, P.Ø.; Rasmussen, T.B.; Christophersen, L.; Calum, H.; Hentzer, M.; Hougen, H.-P.; Rygaard, J.; Moser, C.; Eberl, L.; et al. Garlic blocks quorum sensing and promotes rapid clearing of pulmonary Pseudomonas aeruginosa infections. Microbiology 2005, 151, 3873–3880. [Google Scholar] [CrossRef] [PubMed]
- Slusarenko, A.; Patel, A.; Portz, D. Control of plant diseases by natural products: Allicin from garlic as a case study. Eur. J. Plant Pathol. 2008, 121, 313–322. [Google Scholar] [CrossRef]
- Ankri, S.; Mirelman, D. Antimicrobial properties of allicin from garlic. Microbes Infect. 1999, 1, 125–129. [Google Scholar] [CrossRef]
- Porcheron, G.; Garénaux, A.; Proulx, J.; Sabri, M.; Dozois, C.M. Iron, copper, zinc, and manganese transport and regulation in pathogenic Enterobacteria: Correlations between strains, site of infection and the relative importance of the different metal transport systems for virulence. Front. Cell. Infect. Microbiol. 2013, 3, 172–194. [Google Scholar] [CrossRef] [PubMed]
- Banin, E.; Vasil, M.L.; Greenberg, E.P. Iron and Pseudomonas aeruginosa biofilm formation. Proc. Natl. Acad. Sci. USA 2005, 102, 11076–11081. [Google Scholar] [CrossRef] [PubMed]
- O'Toole, G.A.; Kolter, R. Initiation of biofilm formation in Pseudomonas fluorescens WCS365 proceeds via multiple, convergent signalling pathways: A genetic analysis. Mol. Microbiol. 1998, 28, 449–461. [Google Scholar] [CrossRef] [PubMed]
- Singh, P.K.; Parsek, M.R.; Greenberg, E.P.; Welsh, M.J. A component of innate immunity prevents bacterial biofilm development. Nature 2002, 417, 552–555. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Barken, K.B.; Skindersoe, M.E.; Christensen, A.B.; Givskov, M.; Tolker-Nielsen, T. Effects of iron on DNA release and biofilm development by Pseudomonas aeruginosa. Microbiology 2007, 153, 1318–1328. [Google Scholar] [CrossRef] [PubMed]
- Patriquin, G.M.; Banin, E.; Gilmour, C.; Tuchman, R.; Greenberg, E.P.; Poole, K. Influence of quorum sensing and iron on twitching motility and biofilm formation in Pseudomonas aeruginosa. J. Bacteriol. 2008, 190, 662–671. [Google Scholar] [CrossRef] [PubMed]
- Johnson, M.; Cockayne, A.; Williams, P.H.; Morrissey, J.A. Iron-responsive regulation of biofilm formation in Staphylococcus aureus involves fur-dependent and fur-independent mechanisms. J. Bacteriol. 2005, 187, 8211–8215. [Google Scholar] [CrossRef] [PubMed]
- Musk, D.J.; Banko, D.A.; Hergenrother, P.J. Iron salts perturb biofilm formation and disrupt existing biofilms of Pseudomonas aeruginosa. Chem. Biol. 2005, 12, 789–796. [Google Scholar] [CrossRef] [PubMed]
- Sathyanarayanan, M.B.; Balachandranath, R.; Genji Srinivasulu, Y.; Kannaiyan, S.K.; Subbiahdoss, G. The effect of gold and iron-oxide nanoparticles on biofilm-forming pathogens. ISRN Microbiol. 2013, 2013, 272086. [Google Scholar] [CrossRef] [PubMed]
- Raad, I.I.; Fang, X.; Keutgen, X.M.; Jiang, Y.; Sherertz, R.; Hachem, R. The role of chelators in preventing biofilm formation and catheter-related bloodstream infections. Curr. Opin. Infect. Dis. 2008, 21, 385–392. [Google Scholar] [CrossRef] [PubMed]
- Körstgens, V.; Flemming, H.-C.; Wingender, J.; Borchard, W. Influence of calcium ions on the mechanical properties of a model biofilm of mucoid Pseudomonas aeruginosa. Wood Sci. Technol. 2001, 43, 49–57. [Google Scholar]
- Chen, X.; Stewart, P. Role of electrostatic interactions in cohesion of bacterial biofilms. Appl. Microbiol. Biotechnol. 2002, 59, 718–720. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Yang, L.; Molin, S. Synergistic activities of an efflux pump inhibitor and iron chelators against Pseudomonas aeruginosa growth and biofilm formation. Antimicrob. Agents Chemother. 2010, 54, 3960–3963. [Google Scholar] [CrossRef] [PubMed]
- Banin, E.; Brady, K.M.; Greenberg, E.P. Chelator-induced dispersal and killing of Pseudomonas aeruginosa cells in a biofilm. Appl. Environ. Microbiol. 2006, 72, 2064–2069. [Google Scholar] [CrossRef] [PubMed]
- Bergan, T.; Klaveness, J.; Aasen, A.J. Chelating agents. Chemotherapy 2001, 47, 10–14. [Google Scholar] [CrossRef] [PubMed]
- Che, Y.; Sanderson, K.; Roddam, L.F.; Kirov, S.M.; Reid, D.W. Iron-binding compounds impair Pseudomonas aeruginosa biofilm formation, especially under anaerobic conditions. J. Med. Microbiol. 2009, 58, 765–773. [Google Scholar]
- Bosma, J.W.; Siegert, C.E.; Peerbooms, P.G.; Weijmer, M.C. Reduction of biofilm formation with trisodium citrate in haemodialysis catheters: A randomized controlled trial. Nephrol. Dial. Transplant. 2010, 25, 1213–1217. [Google Scholar] [CrossRef] [PubMed]
- Mladěnka, P.; Macáková, K.; Filipský, T.; Zatloukalová, L.; Jahodář, L.; Bovicelli, P.; Silvestri, I.P.; Hrdina, R.; Saso, L. In vitro analysis of iron chelating activity of flavonoids. J. Inorg. Biochem. 2011, 105, 693–701. [Google Scholar] [CrossRef] [PubMed]
- Jayasinghe, S.; Siriwardhana, A.; Karunaratne, V. Natural iron sequestering agents: Their roles in nature and therapeutic potential. Int. J. Pharm. Pharm. Sci. 2015, 7, 8–12. [Google Scholar]
- Hatcher, H.C.; Singh, R.N.; Torti, F.M.; Torti, S.V. Synthetic and natural iron chelators: Therapeutic potential and clinical use. Future Med. Chem. 2009, 1, 1643–1670. [Google Scholar] [CrossRef] [PubMed]
- Lin, M.-H.; Shu, J.-C.; Huang, H.-Y.; Cheng, Y.-C. Involvement of iron in biofilm formation by Staphylococcus aureus. PLoS ONE 2012, 7, e34388. [Google Scholar] [CrossRef] [PubMed]
- Lin, B.; Johnson, B.J.; Rubin, R.A.; Malanoski, A.P.; Ligler, F.S. Iron chelation by cranberry juice and its impact on Escherichia coli growth. Biofactors 2011, 37, 121–130. [Google Scholar] [CrossRef] [PubMed]
- Abouelhassan, Y.; Garrison, A.T.; Burch, G.M.; Wong, W.; Norwood, V.M.; Huigens, R.W. Discovery of quinoline small molecules with potent dispersal activity against methicillin-resistant Staphylococcus aureus and Staphylococcus epidermidis biofilms using a scaffold hopping strategy. Bioorg. Med. Chem. Lett. 2014, 24, 5076–5080. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.-H.; Kim, Y.-G.; Ryu, S.Y.; Lee, J. Calcium-chelating alizarin and other anthraquinones inhibit biofilm formation and the hemolytic activity of Staphylococcus aureus. Sci. Rep. 2016, 6, 19267. [Google Scholar] [CrossRef] [PubMed]
- Raad, I.; Rosenblatt, J.; Reitzel, R.; Jiang, Y.; Dvorak, T.; Hachem, R. Chelator-based catheter lock solutions in eradicating organisms in biofilm. Antimicrob. Agents Chemother. 2013, 57, 586–588. [Google Scholar] [CrossRef] [PubMed]
- Lebeaux, D.; Leflon-Guibout, V.; Ghigo, J.-M.; Beloin, C. In vitro activity of gentamicin, vancomycin or amikacin combined with EDTA or l-arginine as lock therapy against a wide spectrum of biofilm-forming clinical strains isolated from catheter-related infections. J. Antimicrob. Chemother. 2015, 70, 1704–1712. [Google Scholar] [CrossRef] [PubMed]
- Raad, I.; Chatzinikolaou, I.; Chaiban, G.; Hanna, H.; Hachem, R.; Dvorak, T.; Cook, G.; Costerton, W. In vitro and ex vivo activities of minocycline and EDTA against microorganisms embedded in biofilm on catheter surfaces. Antimicrob. Agents Chemother. 2003, 47, 3580–3585. [Google Scholar] [CrossRef] [PubMed]
- Górska, A.; Sloderbach, A.; Marszałł, M.P. Siderophore–drug complexes: Potential medicinal applications of the ‘Trojan horse’strategy. Trends Pharmacol. Sci. 2014, 35, 442–449. [Google Scholar] [CrossRef] [PubMed]
- Soto, S.M. Role of efflux pumps in the antibiotic resistance of bacteria embedded in a biofilm. Virulence 2013, 4, 223–229. [Google Scholar] [CrossRef] [PubMed]
- Van Acker, H.; Coenye, T. The role of efflux and physiological adaptation in biofilm tolerance and resistance. J. Biol. Chem. 2016, 291, 12565–12572. [Google Scholar] [CrossRef] [PubMed]
- Folsom, J.P.; Richards, L.; Pitts, B.; Roe, F.; Ehrlich, G.D.; Parker, A.; Mazurie, A.; Stewart, P.S. Physiology of Pseudomonas aeruginosa in biofilms as revealed by transcriptome analysis. BMC Microbiol. 2010, 10, 294. [Google Scholar] [CrossRef] [PubMed]
- Stewart, P.S.; Franklin, M.J.; Williamson, K.S.; Folsom, J.P.; Boegli, L.; James, G.A. Contribution of stress responses to antibiotic tolerance in Pseudomonas aeruginosa biofilms. J. Antimicrob. Chemother. 2015, 59, 3838–3847. [Google Scholar] [CrossRef] [PubMed]
- Chan, Y.Y.; Chua, K.L. The Burkholderia pseudomallei BpeAB-OprB efflux pump: Expression and impact on quorum sensing and virulence. J. Bacteriol. 2005, 187, 4707–4719. [Google Scholar] [CrossRef] [PubMed]
- Gupta, G.; Wren, M.; Ganguly, K.; Pardington, P. Multi-drug resistance efflux pumps confer additional resistance against host innate immune defense via induction of genes for biofilm formation and virulence (MPF3P. 803). J. Immunol. 2014, 192, 132.3. [Google Scholar]
- Yamasaki, S.; Wang, L.-Y.; Hirata, T.; Hayashi-Nishino, M.; Nishino, K. Multidrug efflux pumps contribute to Escherichia coli biofilm maintenance. Int. J. Antimicrob. Agents 2015, 45, 439–441. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Mah, T.-F. Involvement of a novel efflux system in biofilm-specific resistance to antibiotics. J. Bacteriol. 2008, 190, 4447–4452. [Google Scholar] [CrossRef] [PubMed]
- Pagès, J.-M.; Masi, M.; Barbe, J. Inhibitors of efflux pumps in Gram-negative bacteria. Trends Mol. Med. 2005, 11, 382–389. [Google Scholar] [CrossRef] [PubMed]
- Matsumura, K.; Furukawa, S.; Ogihara, H.; Morinaga, Y. Roles of multidrug efflux pumps on the biofilm formation of Escherichia coli K-12. Biocontrol Sci. 2011, 16, 69–72. [Google Scholar] [CrossRef] [PubMed]
- Baugh, S.; Ekanayaka, A.S.; Piddock, L.J.; Webber, M.A. Loss of or inhibition of all multidrug resistance efflux pumps of Salmonella enterica serovar Typhimurium results in impaired ability to form a biofilm. J. Antimicrob. Chemother. 2012, 67, 2409–2417. [Google Scholar] [CrossRef] [PubMed]
- Pantel, A.; Dunyach-Remy, C.; Ngba Essebe, C.; Mesureur, J.; Sotto, A.; Pagès, J.-M.; Nicolas-Chanoine, M.-H.; Lavigne, J.-P. Modulation of membrane influx and efflux in Escherichia coli sequence type 131 has an impact on bacterial motility, biofilm formation, and virulence in a Caenorhabditis elegans model. J. Antimicrob. Chemother. 2016, 60, 2901–2911. [Google Scholar] [CrossRef] [PubMed]
- Moore, J.D.; Gerdt, J.P.; Eibergen, N.R.; Blackwell, H.E. Active efflux influences the potency of quorum sensing inhibitors in Pseudomonas aeruginosa. ChemBioChem 2014, 15, 435–442. [Google Scholar] [CrossRef] [PubMed]
- Aybey, A.; Usta, A.; Demirkan, E. Effects of psychotropic drugs as bacterial efflux pump inhibitors on quorum sensing regulated behaviors. J. Microbiol. Biotechnol. Food Sci. 2014, 4, 128. [Google Scholar] [CrossRef]
- Poole, K.; Krebes, K.; McNally, C.; Neshat, S. Multiple antibiotic resistance in Pseudomonas aeruginosa: Evidence for involvement of an efflux operon. J. Bacteriol. 1993, 175, 7363–7372. [Google Scholar] [PubMed]
- Modarresi, F.; Azizi, O.; Shakibaie, M.R.; Motamedifar, M.; Valibeigi, B.; Mansouri, S. Effect of iron on expression of efflux pump (adeABC) and quorum sensing (luxI, luxR) genes in clinical isolates of Acinetobacter baumannii. APMIS 2015, 123, 959–968. [Google Scholar] [CrossRef] [PubMed]
- Kvist, M.; Hancock, V.; Klemm, P. Inactivation of efflux pumps abolishes bacterial biofilm formation. Appl. Environ. Microbiol. 2008, 74, 7376–7382. [Google Scholar] [CrossRef] [PubMed]
- Baugh, S.; Phillips, C.R.; Ekanayaka, A.S.; Piddock, L.J.V.; Webber, M.A. Inhibition of multidrug efflux as a strategy to prevent biofilm formation. J. Antimicrob. Chemother. 2014, 69, 673–681. [Google Scholar] [CrossRef] [PubMed]
- Magesh, H.; Kumar, A.; Alam, A.; Priyam, S.U.; Sumantran, V.; Vaidyanathan, R. Identification of natural compounds which inhibit biofilm formation in clinical isolates of Klebsiella pneumoniae. Indian J. Exp. Biol. 2013, 51, 764–772. [Google Scholar] [PubMed]
- Abouelhassan, Y.; Garrison, A.T.; Bai, F.; Norwood, V.M.; Nguyen, M.T.; Jin, S.; Huigens, R.W. A phytochemical–halogenated quinoline combination therapy strategy for the treatment of pathogenic bacteria. ChemMedChem 2015, 10, 1157–1162. [Google Scholar] [CrossRef] [PubMed]
- Koul, S.; Prakash, J.; Mishra, A.; Kalia, V.C. Potential emergence of multi-quorum sensing inhibitor resistant (MQSIR) bacteria. Indian J. Microbiol. 2016, 56, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Kalia, V.C.; Wood, T.K.; Kumar, P. Evolution of resistance to quorum-sensing inhibitors. Microb. Ecol. 2014, 68, 13–23. [Google Scholar] [CrossRef] [PubMed]


| Source | Effect(s) | Bacteria | Reference(s) |
|---|---|---|---|
| Plant Crude Extracts | |||
| Aqueous extracts from Ananas comosus, Musa paradiciaca, Manilkara zapota and Ocimum sanctum | -Inhibition of biofilm formation and virulence factor production (pyocyanin, staphylolytic protease and elastase); Reduction of N-Acyl homoserine lactones (AHLs)-mediated violacein production | P. aeruginosa PAO1; Chromobacterium violaceum CV12472 | [156] |
| Extracts from Prunus armeniaca, Prunella vulgaris, Nelumbo nucífera, Panax notoginseng, Punica granatum, Areca catechu and Imperata cylindrica | -Inhibition of violacein production and swarming motility | C. violaceum CV026; P. aeruginosa PAO1 | [157] |
| Methanolic extract from Capparis spinosa Linn. | -Inhibition of biofilm formation and motility (swimming and swarming); Decreases the biosurfactant and EPS production; Inhibition of violacein production | P. aeruginosa PAO1; E. coli ATCC 10536; P. mirabilis ATCC 7002; Serratia marcescens FJ584421; C. violaceum CV026/CV12472 | [158] |
| Ethanolic extracts of Italian medicinal plants Ballota nigra, Castanea sativa and Sambucus ebulus | -Inhibition of δ-haemolysin production through of the interference with agr (accessory gene regulator) locus | MRSA (NRS385) clinical isolate | [159] |
| Methanolic, ethanolic, chloroformic, acetonic and aqueous extracts from Salvadora persica | -Inhibition of initial adhesion and biofilm formation; Interferes with QS regulators, Streptococcus OmpP and Staphylococcus Lux proteins | S. mutans cariogenic isolates | [160] |
| Crude extract and ethanolic fraction from Emblica officinalis | -Reduces cell adherence, cell-surface hydrophobicity, glucan synthesis and biofilm formation; Suppresses the expression of genes involved in biofilm formation; Obliterates biofilm structure | S. mutans MTCC 497 | [161] |
| Extract of Cymbopogan citratus (lemongrass) | -Inhibition of biofilm formation; eradication of pre-formed biofilms | S. aureus NCIM 5022 | [162] |
| Ethanolic extracts from Hydrastis canadensis L. (Ranunculaceae) | -Anti-QS activity by attenuation of signal transduction through the AgrCA component system; Inhibit toxin (alpha-toxin) production and prevents keratinocyte damage | S. aureus (CA-MRSA USA300 TCH1516, LAC, AH1263 and SA502A) | [163] |
| Methanolic extracts from Achyranthes aspera L. (Amaranthaceae) | -Inhibition of QS through of the interaction with OmpR QS regulator; Prevent glycosyltransferase (EPS synthesizing enzyme) expression; Inhibit biofilm formation | S. mutans cariogenic isolates | [164] |
| Hexane extract from Amphypterygium adstringens | -Inhibition of pyocyanin and rhamnolipid production; Decreases the elastolytic activity; Inhibition of violacein production | P. aeruginosa PA14; C. violaceum CV12472 | [165] |
| Methanolic extracts from Securinega suffruticosa, Angelica dahurica, Rodgersia podophylla, Viburnum carlesii, Nymphaea tetragona var. Angusta and Mallotus japonicus | -Inhibition of swarming motility; Inhibition of violacein production | P. aeruginosa PAO1; C. violaceum CV12472 | [166] |
| Methanolic extract rich in ellagic acid derivatives from Terminalia chebula Retz. | -Downregulation of lasIR and rhlIR genes expression; Attenuation of virulence factor production (pyocyanin, elastase, rhamnolipid and protease); Reduction of alginate production; Inhibition of biofilm formation and enhanced sensitivity of biofilm towards tobramycin; Enhanced survival of Caenorhabditis elegans | P. aeruginosa PAO1 | [167] |
| n-hexane extract of Dalbergia trichocarpa | -Inhibition of biofilm formation, motility (swarming and twitching) and virulence factor production (pyocyanin, elastase and proteases); Increases the effectiveness of biofilm-encapsulated to tobramycin | P. aeruginosa PAO1 | [168] |
| Ethyl acetate fraction of ethanol extract from Syzygium cumini L., Pimentadioica L., Centella asiatica L. and Adenanthera pavonina L.; Flavonoid fraction from leaves of Psidium guajava L. | -Inhibition of several QS-regulated phenotypes, namely: pyocyanin production, elastolytic and proteolytic activities, swarming motility and biofilm formation; Inhibition of violacein production | P. aeruginosa PAO1; C. violaceum CV026/CV12472 | [169,170,171,172] |
| Polyphenol rich extract from Rosa rugosa tea | -Inhibition of swarming motility and biofilm formation; Inhibition of violacein production | E. coli K-12; P. aeruginosa PAO1; C. violaceum CV026/CV12472 | [173] |
| Extracts of neotropical rainforest plants from Meliaceae, Melastomataceae, Lepidobotryaceae, and Sapindaceae families | -Biofilm formation and violacein production inhibition | P. aeruginosa PA14; C. violaceum CV12472 | [174] |
| Extract from wheat bran | -Interferes with QS by degrading AHLs; Inhibition of biofilm formation; Eradication of pre-formed biofilms | Pseudomonas fluorescens (P3/pME6863 and P3/pME6000); S. aureus (BMA/FR/0.32/0074) clinical isolate | [175] |
| Extracts from Chamaemelum nobile (Chamomile) | -Inhibit biofilm formation, swarming motility | P. aeruginosa PAO1 and clinical isolates from different types of infections | [176] |
| Methanolic extract from Kalanchoe blossfeldiana | -Reduces virulence factors secretion (protease and pyoverdine) and cytokine formation in lipopolysaccharide-stimulated peripheral blood mononuclear cells; Inhibition of biofilm production | P. aeruginosa MTCC 2453 | [177] |
| Ethanolic extract from Amomum tsaoko | -Inhibition of violacein production, swarming motility and biofilm formation | C. violaceum ATCC 12472; S. aureus ATCC 6538; Salmonella enterica serovar Typhimurium ATCC 50013; P. aeruginosa ATCC 9027 | [178] |
| Methanolic extract from Anethum graveolens and one of its principle active, 3-O-methyl ellagic acid | -Downregulation of QS-regulated genes (fimC, bsmA and flhD) crucial for initial adhesion and motility; Reduction of biofilm formation and virulence factors (prodigiosin and protease) production | S. marcescens MG1/MG44 and clinical isolates | [179] |
| Methanolic extract from Sygygium aromaticum | -Inhibition of biofilm formation; Inhibition of EPS and pyocyanin production, proteolytic activity and swimming motility | C. violaceum ATCC 12472; P. aeruginosa clinical isolates | [180] |
| Methanolic extracts rich in ursene and oleanene derivatives (pentacyclic triterpenes) from Castanea sativa (European chestnut) | -Inhibition of haemolytic activity, harmful exotoxins (e.g., δ-toxin) production and biofilm formation (to a lesser extent) as results of the agr-mediated QS blockage; Attenuate skin abscesses in an in vivo animal model | S. aureus (AH408, SA502A, AH430, AH845, AH1263, LAC, AH1677, AH1747, AH1872, AH2759 and AH3052) | [181] |
| n-hexane and dichloromethane extracts of Liriodendron hybrid barks | -Inhibition of violacein production; Inhibition of biofilm formation | C. violaceum ATCC 12472/CV026; MRSA clinical isolates | [182] |
| Extract rich in polyphenols (orcinol, arabitol, apigenin, and usnic acid) from Usnea longissimi (Beard lichen) | -Inhibition of violacein production; Reduction of virulence factor secretion (acid production, ATPase, enolase, lactate dehydrogenase, protease, total exopolysaccharide content and glucosidase); Inhibition of biofilm formation; Improvement of the susceptibility to conventional antibiotics | C. violaceum CV12472; S. mutans MTCC 0497 clinical strain | [183] |
| Crude extract and methanolic fraction from Z. officinale | -Inhibition of biofilm formation; Reduce the insoluble glucan synthesis and sucrose-dependent adherence; Induce the dispersal of biofilm cells; Reduce caries development using an in vivo mouse model | S. mutans UA159 | [184] |
| Extract rich in flavonoids (licoricone, glycyrin and glyzarin) from Glycyrrhiza glabra | -Reduce the production of QS-regulated virulence factors (e.g., motility, biofilm formation and production of antioxidant enzymes); Downregulation of the autoinducer (AI) synthase gene (abaI) expression | A. baumannii ATCC 19606 and ATCC 17978, and clinical isolates (M2, M2 (abal::Km), M2 (Pabal-lacZ) and C1-C4) | [185] |
| Methanolic extracts rich in tannin from Phyllanthus emblica, Terminalia bellirica, Terminalia chebula, Punica granatum, Syzygium cumini, and Mangifera indica | -β-lactamase inhibition as a result of the interference with agr expression; Inhibition of violacein production | S. aureus agrP3::blaZ RN6390 pRN8826; C. violaceum CV12472 | [186] |
| Ethanolic extract from Piper betle | -Reduces swarming, swimming, and twitching; Inhibition of biofilm formation and pyocyanin production | P. aeruginosa PAO1 | [187] |
| Phenolic extract from Rubus rosaefolius (wild strawberry) | -Inhibition of several QS-regulated phenotypes, namely: violacein production, swarming motility and biofilm formation | C. violaceum ATCC 6357; S. marcescens UFOP-001; Aeromonas hydrophila IOC/FDA110 | [188] |
| Ethanol solution extract rich in evodiamine, rutaecarpine and evocarpine from Euodia ruticarpa | -Inhibition of AI-2 production; Inhibition of cell adhesion and biofilm formation | Campylobacter jejuni | [189] |
| Plant pure compounds | |||
| Epigallocatechin gallate from green tea | -Inhibition of biofilm formation and swarming motility; Synergistic activity with ciprofloxacin in the treatment of biofilm infections | P. aeruginosa PAO1 | [190] |
| Catechin and naringenin from Combretum albiflorum | -Interfere with the pyocyanin and elastase production; Affect the AIs perception; Biofilm formation inhibition | P. aeruginosa PAO1 | [191] |
| Catechins with a galloyl moiety (e.g., epichatechin gallate and epigallocatechin gallate) | -Affect AI-2 and inhibit biofilm formation | Eikenella corrodens 1073 | [192] |
| Saponins, ginsenosides, and polysaccharides fom Panax ginseng | -Suppression of the production of LasA and LasB; Downregulation of AHLs synthesis; Clearance of pulmonary infections in animal studies by biofilm disruption | P. aeruginosa PAO1 and its isogenic mucoid variant (PAOmucA22) | [193,194] |
| Baicalin hydrate, cinnamaldehyde and hamamelitannin | -Increase biofilm susceptibility to treatment with antibiotics (e.g., tobramycin, clindamycin, vancomycin); Enhance the survival of infected C. elegans and Galleria mellonella; Reduce the microbial load in the lungs of infected BALB/c mice | P. aeruginosa (PAO1, ATCC 9027, MH340 and MH710); Burkholderia cenocepacia (LMG 16656, LMG 16659 and LMG 18828); S. aureus (LMG 10147 and Mu50–MRSA) and clinical isolates (CS1) | [195] |
| Chrysophanol, nodakenetin, shikonin and emodin from tradicional Chinese herbs (Rheum officinale Baill, Peucedanum decursivum (Miq). Maxim, Lithospermum erythrorhizon Sieb, Rheum palmatum L.) | -Inhibition of biofilm formation; Potentiation of the ampicillin activity; Proteolysis of the QS signal receptor TraR | Stenotrophomonas maltophilia GIMT1.118; P. aeruginosa PAO1; E. coli BL21(DE3) | [196] |
| Allicin and ajoene from Allium sativum | -Reduction of QS-controlled virulence genes expression; Attenuation of the rhamnolipid production; Synergistic activity with tobramycin on biofilms; Cessation of the polymorphonuclear leukocytes lytic necrosis; Enable the clearance of pulmonary infections in mouse models | P. aeruginosa PAO1 | [143,197] |
| Iberin from Armoracia rusticana (horseradish) | -Blockage of the QS-regulated genes expression by targeting LasIR and RhlIR QS networks; Downregulation of rhamnolipid production | P. aeruginosa PAO1 | [146] |
| Methyl eugenol from Cuminum cyminum | -Inhibition of biofilm formation, motility (swimming and swarming) and EPS production | P. aeruginosa PAO1; P. mirabilis ATCC 7002; S. marcescens FJ584421 | [198] |
| Rosmarinic acid, naringin, chlorogenic acid, morin and mangiferin | -Inhibition of biofilm formation and virulence factor production (protease, elastase and haemolysin) | P. aeruginosa PAO1; P. aeruginosa AS1 and AS2 | [199] |
| Curcumin from Curcuma longa L. | -Inhibition of biofilm formation and attenuation of QS-dependent factors (exopolysaccharide and alginate production); Inhibition of swimming and swarming motility; Biofilm susceptibility enhancement to antibiotics; Enhanced survival rate of Artemia nauplii | E. coli ATCC 10536; P. aeruginosa PAO1; P. mirabilis ATCC 7002; S. marcescens FJ584421; Vibrio harveyi MTCC 3438, Vibrio parahaemolyticus ATCC 17802 and Vibrio vulnificus MTCC 1145 | [200,201] |
| Glycosylflavonoids (chlorogenic acid, isoorientin, orientin, isovitexin, vitexin, and rutin) from Cecropia pachystachya Trécul | -Inhibition of violacein production; Inhibition of bioluminescence production | C. violaceum ATCC 31532; E. coli pSB403 | [202] |
| Salicylic acid | -Inhibit swimming motility; Dual-species biofilms enhancement to a second exposure to salicylic acid | Bacillus cereus isolated from a disinfectant solution; P. fluorescens ATCC 13525 | [203] |
| Salicylic acid, tannic acid and trans-cinnamaldehyde | -Inhibition of AHLs and pyocyanin production | P. aeruginosa PAO1 | [204] |
| [6]-gingerol, [6]-shogaol and zingerone from Zingiber officinale Roscoe | -Inhibition of biofilm formation, violacein and pyocyanin production | C. violaceum MTCC 2656; P. aeruginosa MTCC 2297/PA14 | [205,206] |
| Zingerone from ginger root | Decreases swimming, swarming and twitching motility; Reduces biofilm-forming capacity; Interferes with the production of virulence factors including rhamnolipid, elastase, protease, pyocyanin; Improves the antibiofilm efficacy of ciprofloxacin | P. aeruginosa PAO1 | [207,208] |
| Sesquiterpenoid viridiflorol and triterpenoids, ursolic and betulinic acids, from the liverwort Lepidozia chordulifera | -Inhibition of biofilm formation and elastolytic activity | P. aeruginosa ATCC27853; S. aureus ATTC6538 | [209] |
| Malvidin of methanolic extract from Syzygium cumini | -Inhibition of violacein production, EPS synthesis and biofilm formation; Potentiation of the susceptibility to conventional antibiotics | C. violaceum CV026/MTCC 2656; K. pneumoniae PUFST23 | [210] |
| Hamamelitannin | -Increases the in vitro biofilm susceptibility to vancomycin treatment through the TraP receptor by affecting cell wall biosynthesis (peptidoglycan) and extracellular DNA release; Increases the in vivo susceptibility to antibiotic treatment using C. elegans and mouse (mammary gland infection) models | MRSA Mu50 | [211] |
| Essential oils and components | |||
| Clove essential oil | -Inhibition of LasB, total protease, chitinase, pyocyanin and exopolysaccharide production; Swimming motility and biofilm formation reduction | P. aeruginosa PAO1; A. hydrophila WAF-38 | [212] |
| Essential oil from Murraya koenigii | -Inhibition of violacein production; Biofilm formation inhibition; Reduces cell adhesion, metabolic activity and EPS production; Prevents biofilm maturation | C. violaceum CV026/CV12472; P. aeruginosa PAO1 | [213] |
| Cinnamon oil | -Inhibits biofilm formation and virulence factors production (pyocyanin, rhamnolipid, and protease); Reduces alginate and EPS production, and swarming motility | P. aeruginosa PAO1 | [214] |
| Mentha piperita essential oil (peppermint) and menthol | -Inhibition of violacein production; Biofilm formation, EPS production and swarming motility inhibition; Affect QS regulate virulence factors production (elastase, total protease, pyocyanin and chitinase); Interference with las and pqs QS systems; Enhanced survival of C. elegans | C. violaceum CV026; P.aeruginosa; A. hydrophila; E. coli (MG4/pKDT17 and pEAL08-2) | [215] |
| Clove bud oil | -Attenuation of extracellular DNA, exopolysaccharides and pigment production; Decreases the transcription of pqsA gene; Biofilm inhibition and dispersal | P. aeruginosa PAO1 | [216] |
| Eugenol | -Inhibition of violacein production, elastase, pyocyanin and biofilm formation; Interference with las and pqs QS systems | C. violaceum CV026; P. aeruginosa PAO1/PAO-MW1; E. coli (MG4/pKDT17 and pEAL08-2) | [217] |
| Carvacrol | -Inhibition of biofilm formation; Reduction of the expression of cviI gene, production of violacein and chitinase activity | C. violaceum ATCC 12472; S. enterica serovar Typhimurium DT104; S. aureus 0074 | [218] |
© 2016 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Borges, A.; Abreu, A.C.; Dias, C.; Saavedra, M.J.; Borges, F.; Simões, M. New Perspectives on the Use of Phytochemicals as an Emergent Strategy to Control Bacterial Infections Including Biofilms. Molecules 2016, 21, 877. https://doi.org/10.3390/molecules21070877
Borges A, Abreu AC, Dias C, Saavedra MJ, Borges F, Simões M. New Perspectives on the Use of Phytochemicals as an Emergent Strategy to Control Bacterial Infections Including Biofilms. Molecules. 2016; 21(7):877. https://doi.org/10.3390/molecules21070877
Chicago/Turabian StyleBorges, Anabela, Ana Cristina Abreu, Carla Dias, Maria José Saavedra, Fernanda Borges, and Manuel Simões. 2016. "New Perspectives on the Use of Phytochemicals as an Emergent Strategy to Control Bacterial Infections Including Biofilms" Molecules 21, no. 7: 877. https://doi.org/10.3390/molecules21070877
APA StyleBorges, A., Abreu, A. C., Dias, C., Saavedra, M. J., Borges, F., & Simões, M. (2016). New Perspectives on the Use of Phytochemicals as an Emergent Strategy to Control Bacterial Infections Including Biofilms. Molecules, 21(7), 877. https://doi.org/10.3390/molecules21070877