A Sorghum MYB Transcription Factor Induces 3-Deoxyanthocyanidins and Enhances Resistance against Leaf Blights in Maize
Abstract
:1. Introduction
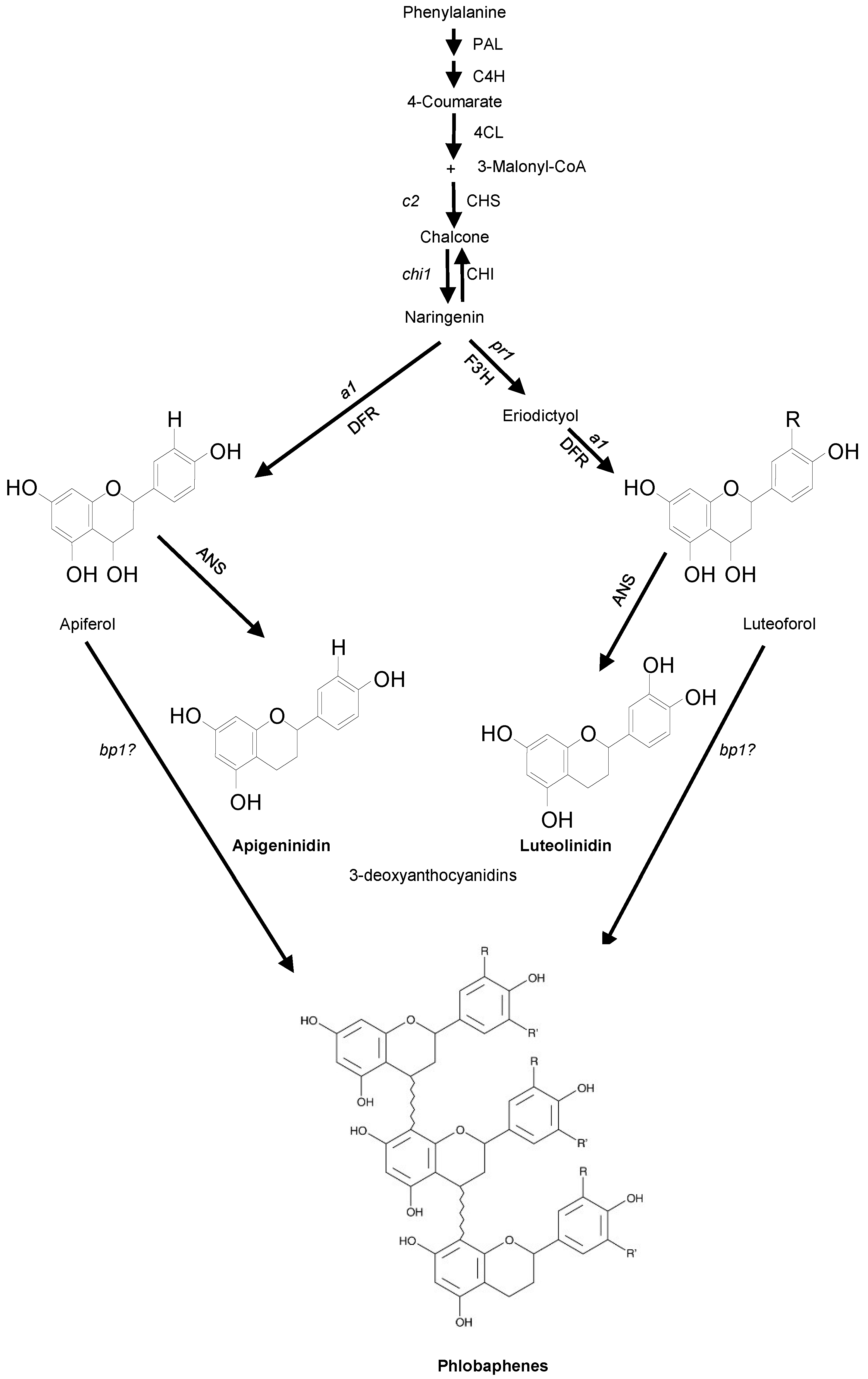
2. Results and Discussion
2.1. y1 Transgenes Phenocopy p1 Pigmentation Patterns in Maize
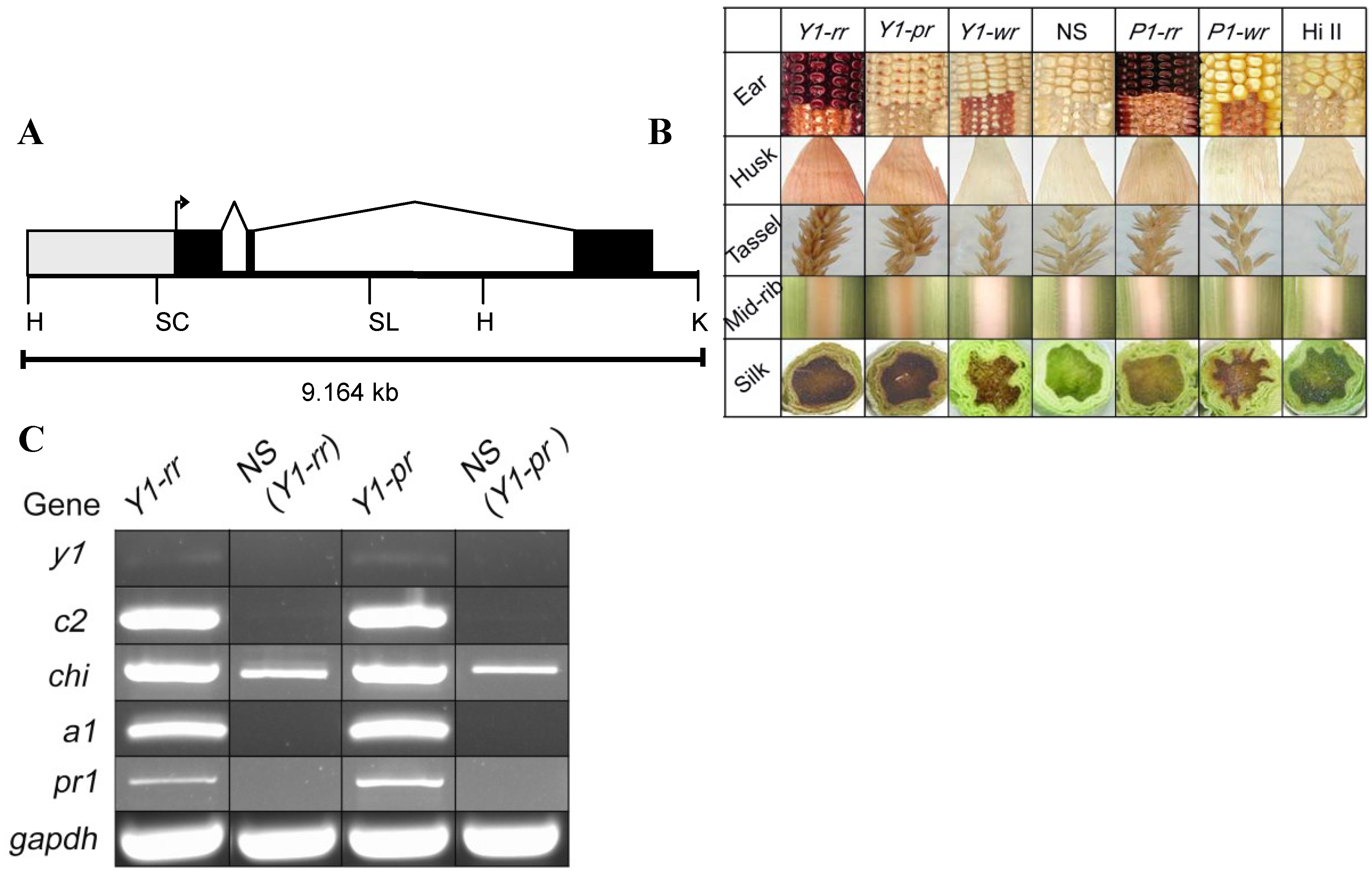
2.2. y1 Regulates Accumulation of 3-Deoxyflavonoids (flavan-4-ols) in Maize
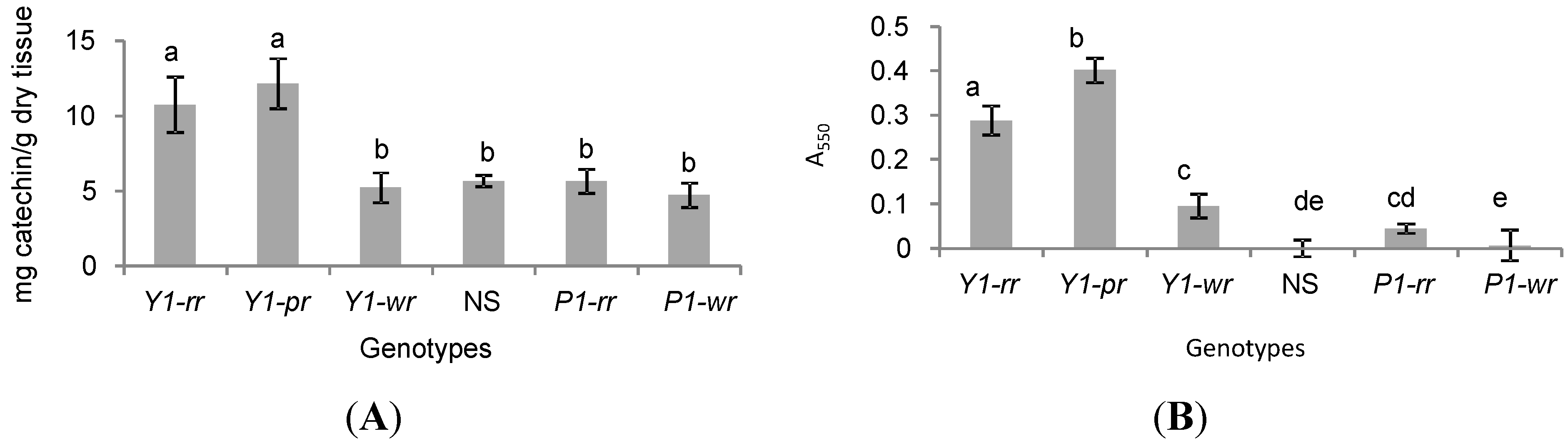
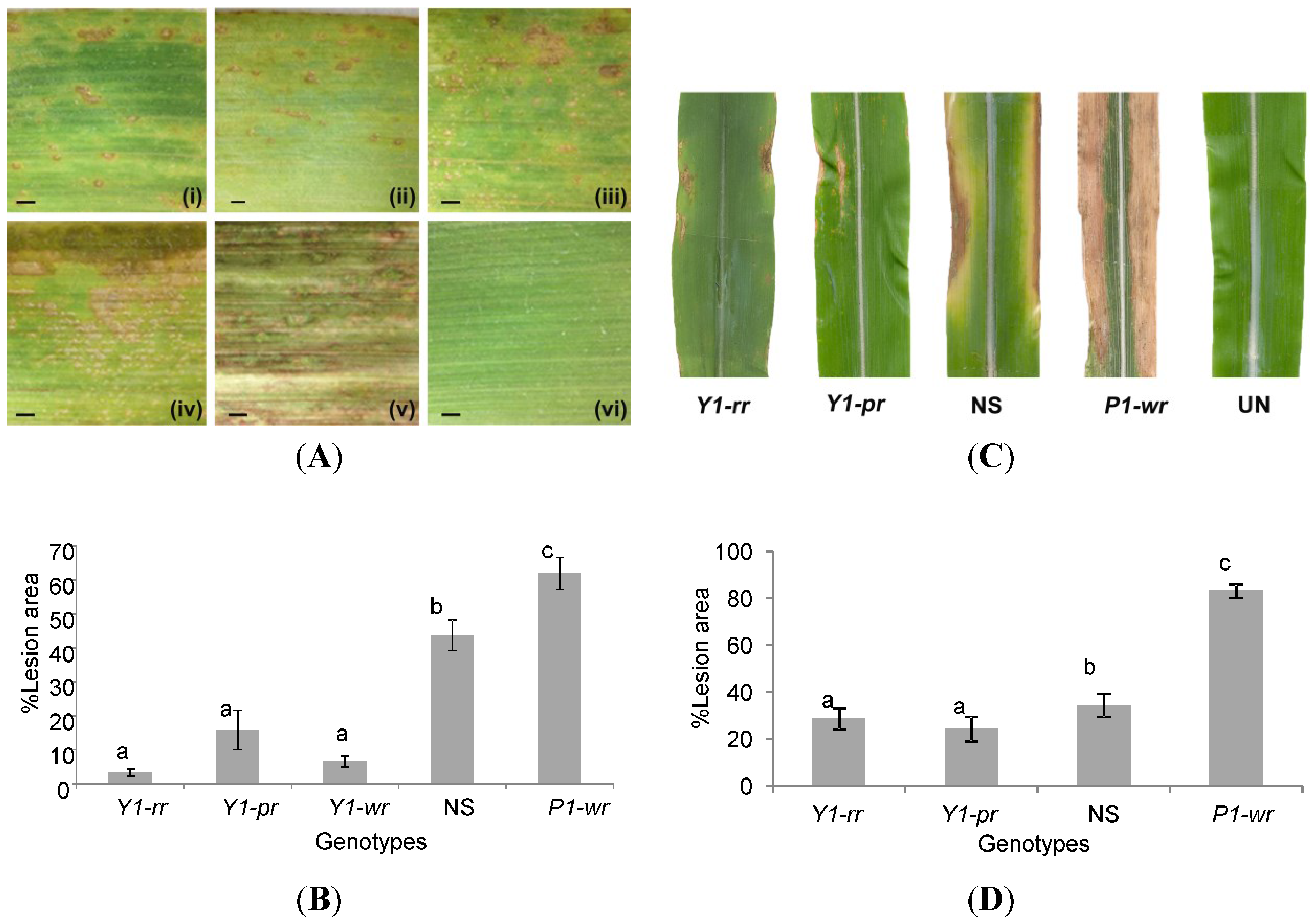
2.3. Y1-Maize Exhibits Enhanced Resistance to C. heterostrophus and C. graminicola
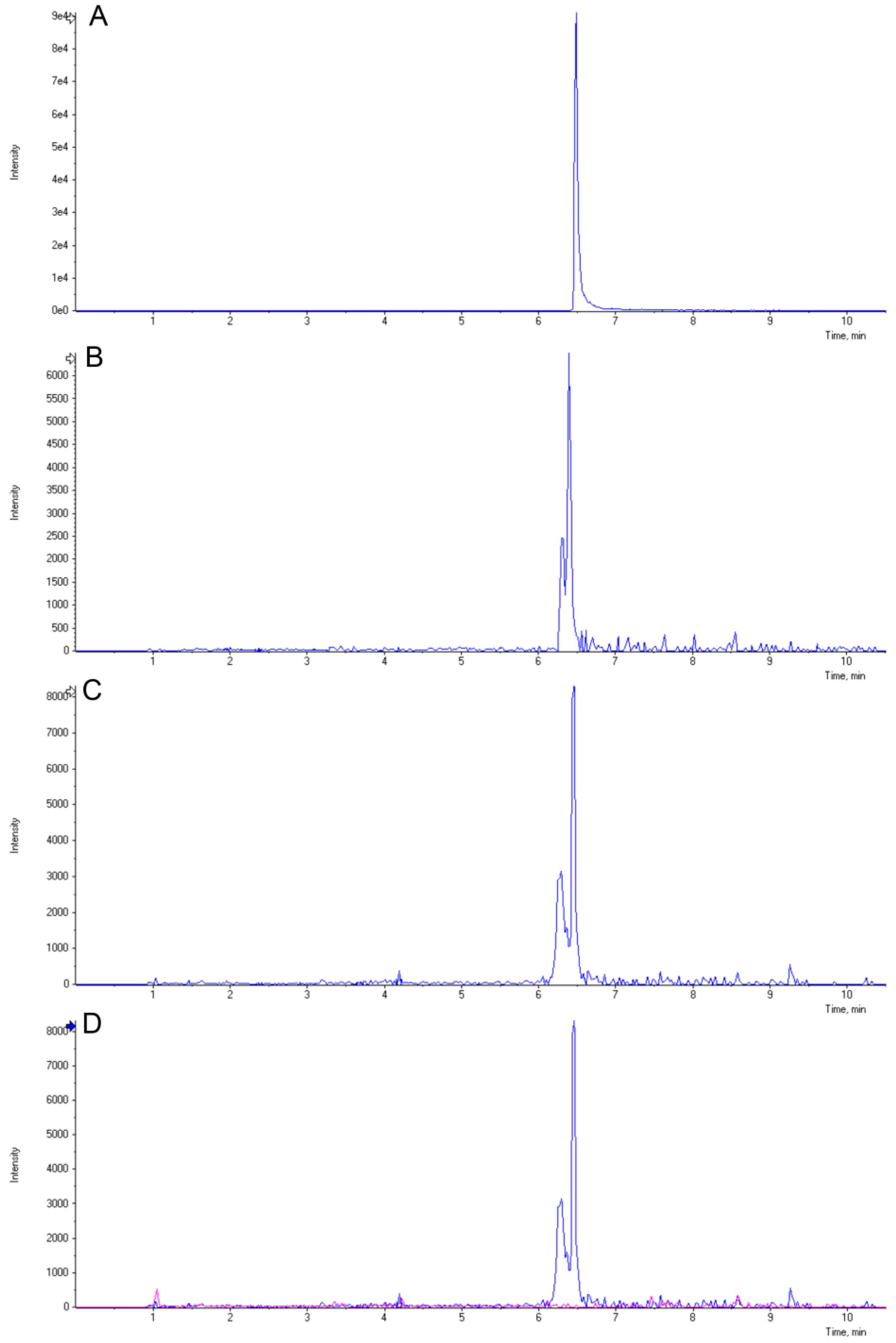
2.4. Induction of 3-Deoxyanthocyanidins during Y1-Maize–C. graminicola Interaction
2.5. Discussion
3. Experimental Section
3.1. Maize Genetic Stocks
3.2. Transgene Constructs
3.3. Tissue Culture, Transformation, and Regeneration of Transgenic Maize Plants
3.4. Expression Analysis of Genes Induced by y1
3.5. Tissue Collection for Chemical Analyses
3.6. Quantification of Total Flavonoids and Flavan-4-ols
3.7. Evaluation of Y1-Transgenic Plants for Resistance to Cochliobolus heterostrophus and Colletotrichum graminicola
3.8. Image Analysis for Evaluation of Disease Response
3.9. LC-MS Analysis of 3-Deoxyanthocyanidins
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Winkel-Shirley, B. Flavonoid biosynthesis. A colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiol. 2001, 126, 485–493. [Google Scholar] [CrossRef] [PubMed]
- Dixon, R.A.; Liu, C.; Jun, J.H. Metabolic engineering of anthocyanins and condensed tannins in plants. Curr. Opin. Biotechnol. 2013, 24, 329–335. [Google Scholar] [CrossRef] [PubMed]
- Verpoorte, R.; Memelink, J. Engineering secondary metabolite production in plants. Curr. Opin. Biotechnol. 2002, 13, 181–187. [Google Scholar] [CrossRef] [PubMed]
- Kristensen, C.; Morant, M.; Olsen, C.E.; Ekstrøm, C.T.; Galbraith, D.W.; Møller, B.L.; Bak, S. Metabolic engineering of dhurrin in transgenic arabidopsis plants with marginal inadvertent effects on the metabolome and transcriptome. Proc. Natl. Acad. Sci. USA 2005, 102, 1779–1784. [Google Scholar] [CrossRef] [PubMed]
- Tattersall, D.B.; Bak, S.; Jones, P.R.; Olsen, C.E.; Nielsen, J.K.; Hansen, M.L.; Høj, P.B.; Møller, B.L. Resistance to an herbivore through engineered cyanogenic glucoside synthesis. Science 2001, 293, 1826–1828. [Google Scholar] [CrossRef] [PubMed]
- Styles, E.D.; Ceska, O. Pericarp flavonoids in genetic strains of zea mays. Maydica 1989, 34, 227–237. [Google Scholar]
- Grotewold, E.; Drummond, B.J.; Bowen, B.; Peterson, T. The myb-homologous p gene controls phlobaphene pigmentation in maize floral organs by directly activating a flavonoid biosynthetic gene subset. Cell 1994, 76, 543–553. [Google Scholar] [CrossRef] [PubMed]
- Morohashi, K.; Casas, M.I.; Ferreyra, L.F.; Mejía-Guerra, M.K.; Pourcel, L.; Yilmaz, A.; Feller, A.; Carvalho, B.; Emiliani, J.; Rodriguez, E.; et al. A genome-wide regulatory framework identifies maize pericarp color1 controlled genes. Plant Cell 2012, 24, 2745–2764. [Google Scholar] [CrossRef] [PubMed]
- Boddu, J.; Jiang, C.H.; Sangar, V.; Olson, T.; Peterson, T.; Chopra, S. Comparative structural and functional characterization of sorghum and maize duplications containing orthologous myb transcription regulators of 3-deoxyflavonoid biosynthesis. Plant Mol. Biol. 2006, 60, 185–199. [Google Scholar] [CrossRef] [PubMed]
- Bennetzen, J.L.; Freeling, M. The unified grass genome: Synergy in synteny. Genome Res. 1997, 7, 301–306. [Google Scholar] [PubMed]
- Gaut, B.S.; le Thierry d’Ennequin, M.; Peek, A.S.; Sawkins, M.C. Maize as a model for the evolution of plant nuclear genomes. Proc. Natl. Acad. Sci. USA 2000, 97, 7008–7015. [Google Scholar] [CrossRef] [PubMed]
- Devos, K.M.; Gale, M. Genome relationships: The grass model in current research. Plant Cell 2000, 12, 637–646. [Google Scholar] [CrossRef] [PubMed]
- Melake-Berhan, A.; Hurber, S.H.; Butler, L.G.; Bennetzen, J.L. Structure and evolution of the genome of sorghum bicolor and zea mays. Theor. Appl. Genet. 1993, 86, 598–604. [Google Scholar] [CrossRef] [PubMed]
- Zanta, C.A.; Yang, X.; Axtell, J.D.; Bennetzen, J.L. The candystripe locus, y-cs, determines mutable pigmentation of the sorghum leaf, flower, and pericarp. J. Hered. 1994, 85, 23–29. [Google Scholar]
- Chopra, S.; Brendel, V.; Zhang, J.B.; Axtell, J.D.; Peterson, T. Molecular characterization of a mutable pigmentation phenotype and isolation of the first active transposable element from sorghum bicolor. Proc. Natl. Acad. Sci. USA 1999, 96, 15330–15335. [Google Scholar] [CrossRef] [PubMed]
- Ibraheem, F.; Gaffoor, I.; Chopra, S. Flavonoid phytoalexin dependent resistance to anthracnose leaf blight requires a functional yellow seed1 in sorghum bicolor. Genetics 2010, 184, 915–926. [Google Scholar] [CrossRef] [PubMed]
- Chopra, S.; Gevens, A.; Svabek, C.; Wood, K.V.; Peterson, T.; Nicholson, R.L. Excision of the candystripe1 transposon from a hyper-mutable y1-cs allele shows that the sorghum y1 gene controls the biosynthesis of both 3-deoxyanthocyanidin phytoalexins and phlobaphene pigments. Physiol. Mol. Plant Pathol. 2002, 60, 321–330. [Google Scholar] [CrossRef]
- Snyder, B.A.; Nicholson, R.L. Synthesis of phytoalexins in sorghum as a site-specific response to fungal ingress. Science 1990, 248, 1637–1639. [Google Scholar] [CrossRef] [PubMed]
- Hipskind, J.D.; Nicholson, R.L.; Goldsbrough, P.B. Isolation of a cdna encoding a novel leucine-rich repeat motif from Sorghum bicolor inoculated with fungi. Mol. Plant-Microbe Interact. 1996, 9, 819–825. [Google Scholar] [CrossRef] [PubMed]
- McMullen, M.D.; Snook, M.; Lee, E.A.; Byrne, P.F.; Kross, H.; Musket, T.A.; Houchins, K.; Coe, E.H., Jr. The biological basis of epistasis between quantitative trait loci for flavone and 3-deoxyanthocyanin synthesis in maize (Zea mays L.). Genome 2001, 44, 667–676. [Google Scholar] [CrossRef] [PubMed]
- Grotewold, E.; Chamberlin, M.; Snook, M.; Siame, B.; Butler, L.; Swenson, J.; Maddock, S.; Clair, G.S.; Bowen, B. Engineering secondary metabolism in maize cells by ectopic expression of transcription factors. Plant Cell 1998, 10, 721–740. [Google Scholar] [PubMed]
- McMullen, M.D.; Kross, H.; Snook, M.E.; Cortes-Cruz, M.; Houchins, K.E.; Musket, T.A.; Coe, E.H. Salmon silk genes contribute to the elucidation of the flavone pathway in maize (Zea mays L.). J. Hered. 2004, 95, 225–233. [Google Scholar] [CrossRef] [PubMed]
- Sharma, M.; Chai, C.; Morohashi, K.; Grotewold, E.; Snook, M.E.; Chopra, S. Expression of flavonoid 3′-hydroxylase is controlled by p1, the regulator of 3-deoxyflavonoid biosynthesis in maize. BMC Plant Biol. 2012, 12, 196. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Wang, Y.; Zhang, J.; Maddock, S.; Snook, M.; Peterson, T. A maize qtl for silk maysin levels contains duplicated myb-homologous genes which jointly regulate flavone biosynthesis. Plant Mol. Biol. 2003, 52, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Nicholson, R.L.; Kollipara, S.S.; Vincent, J.R.; Lyons, P.C.; Cadena-Gomez, G. Phytoalexin synthesis by the sorghum mesocotyl in response to infection by pathogenic and nonpathogenic fungi. Proc. Natl. Acad. Sci. USA 1987, 84, 5520–5524. [Google Scholar] [CrossRef] [PubMed]
- Bate-Smith, E.C. Luteoforol (3',4,4',5,7-pentahydroxyflavan) in Sorghum vulgare L. Phytochemistry 1969, 8, 1803–1810. [Google Scholar] [CrossRef]
- Chopra, S.; Athma, P.; Peterson, T. Alleles of the maize p gene with distinct tissue specificities encode myb-homologous proteins with c-terminal replacements. Plant Cell 1996, 8, 1149–1158. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Chopra, S.; Peterson, T. A segmental gene duplication generated differentially expressed myb-homologous genes in maize. Plant Cell 2000, 12, 2311–2322. [Google Scholar] [CrossRef] [PubMed]
- Boddu, J.; Svabek, C.; Ibraheem, F.; Jones, A.D.; Chopra, S. Characterization of a deletion allele of a sorghum myb gene, yellow seed1 showing loss of 3-deoxyflavonoids. Plant Sci. 2005, 169, 542–552. [Google Scholar] [CrossRef]
- Cocciolone, S.M.; Nettleton, D.; Snook, M.; Peterson, T. Transformation of maize with the p1 transcription factor directs production of silk maysin, a corn earworm resistance factor, in concordance with a hierarchy of floral organ pigmentation. Plant Biotechnol. 2005, 3, 225–235. [Google Scholar] [CrossRef]
- Chopra, S.; Cocciolone, S.M.; Bushman, S.; Sangar, V.; McMullen, M.D.; Peterson, T. The maize unstable factor for orange1 is a dominant epigenetic modifier of a tissue specifically silent allele of pericarp color1. Genetics 2003, 163, 1135–1146. [Google Scholar] [PubMed]
- Robbins, M.L.; Roy, A.; Wang, P.-H.; Gaffoor, I.; Sekhon, R.S.; de O. Buanafina, M.M.; Rohila, J.S.; Chopra, S. Comparative proteomics analysis by DIGE and iTRAQ provides insight into the regulation of phenylpropanoids in maize. J. Proteomics 2013, 93, 254–275. [Google Scholar] [CrossRef] [PubMed]
- Hammerschmidt, R. Phenols and plant-pathogen interactions: The saga continues. Physiol. Mol. Plant Pathol. 2005, 66, 77–78. [Google Scholar] [CrossRef]
- Kambal, A.E.; Bate-Smith, E.C. A genetic and biochemical study on pericarp pigmentation between two cultivars of grain sorghum, sorghum bicolor. Heredity 1976, 37, 417–421. [Google Scholar] [CrossRef]
- Schutt, C.; Netzly, D. Effect of apiforol and apigeninidin on growth of selected fungi. J. Chem. Ecol. 1991, 17, 2261–2266. [Google Scholar] [CrossRef] [PubMed]
- Stich, K.; Forkmann, G. Biosynthesis of 3-deoxyanthocyanins with flower extracts from Sinningia cardinalis. Phytochemistry 1988, 27, 785–789. [Google Scholar] [CrossRef]
- Winefield, C.S.; Lewis, D.H.; Swinny, E.E.; Zhang, H.B.; Arathoon, H.S.; Fischer, T.C.; Halbwirth, H.; Stich, K.; Gosch, C.; Forkmann, G.; et al. Investigation of the biosynthesis of 3-deoxyanthocyanins in Sinningia cardinalis. Physiol. Plant. 2005, 124, 419–430. [Google Scholar] [CrossRef]
- Jambunathan, R.; Butler, L.G.; Bandyopadhyay, R.; Lewisk, N. Polyphenol concentration in grain leaf and callus of mold susceptible and mold resistant sorghum cultivars. J. Agric. Food Chem. 1986, 34, 425–420. [Google Scholar] [CrossRef]
- Jambunathan, R.; Kherdekar, M.S.; Bandyopadhyay, R. Flavan-4-ols concentration in mold-susceptible and mold resistant sorghum at different stages of grain development. J. Agric. Food Chem. 1990, 38, 545–548. [Google Scholar] [CrossRef]
- Menkir, A.; Ejeta, G.; Butler, L.; Melakeberhan, A. Physical and chemical kernel properties associated with resistance to grain mold in sorghum. Cereal Chem. 1996, 73, 613–617. [Google Scholar]
- Spinelli, F.; Speakman, J.B.; Rademacher, W.; Halbwirth, H.; Stich, K.; Costa, G. Luteoforol, a flavan 4-ol, is induced in pome fruits by prohexadione-calciumand shows phytoalexin-like properties against erwinia amylovora and other plant pathogens. Eur. J. Plant Pathol. 2005, 112, 133–142. [Google Scholar] [CrossRef]
- Boddu, J.; Svabek, C.; Sekhon, R.; Gevens, A.; Nicholson, R.L.; Jones, A.D.; Pedersen, J.F.; Gustine, D.L.; Chopra, S. Expression of a putative flavonoid 3'-hydroxylase in sorghum mesocotyls synthesizing 3-deoxyanthocyanidin phytoalexins. Physiol. Mol. Plant Pathol. 2004, 65, 101–113. [Google Scholar] [CrossRef]
- Bruce, W.; Folkerts, O.; Garnaat, C.; Crasta, O.; Roth, B.; Bowen, B. Expression profiling of the maize flavonoid pathway genes controlled by estradiol-inducible transcription factors crc and p. Plant Cell 2000, 12, 65–80. [Google Scholar] [CrossRef] [PubMed]
- Bushman, B.S.; Snookc, M.E.; Gerkeb, J.B.; Szalmaa, S.J.; Berhowd, M.A.; Houchinse, K.E.; McMullen, M.D. Two loci exert major effects on chlorogenic acid synthesis in maize. Crop Sci. 2002, 42, 1669–1678. [Google Scholar]
- Dixon, R.A.; Achnine, L.; Kota, P.; Liu, C.; Reddy, M.S.; Wang, L. The phenylpropanoid pathway and plant defence—A genomics perspective. Mol. Plant Pathol. 2002, 3, 371–390. [Google Scholar] [CrossRef] [PubMed]
- Blount, J.W.; Korth, K.L.; Masoud, S.A.; Rasmussen, S.; Lamb, C.; Dixon, R.A. Altering expression of cinnamic acid 4-hydroxylase in transgenic plants provides evidence for a feedback loop at the entry point into the phenylpropanoid pathway. Plant Physiol. 2000, 122, 107–116. [Google Scholar] [CrossRef] [PubMed]
- Jin, H.; Martin, C. Multifunctionality and diversity within the plant myb-gene family. Plant Mol. Biol. 1999, 41, 577–585. [Google Scholar] [CrossRef] [PubMed]
- Jirschitzka, J.; Mattern, D.J.; Gershenzon, J.; D’Auria, J.C. Learning from nature: New approaches to the metabolic engineering of plant defense pathways. Curr. Opin. Biotechnol. 2013, 24, 320–328. [Google Scholar] [CrossRef] [PubMed]
- Mengiste, T.; Chen, X.; Salmeron, J.; Dietrich, R. The Botrytis susceptible1 gene encodes an r2r3myb transcription factor protein that is required for biotic and abiotic stress responses in arabidopsis. Plant Cell 2003, 15, 2551–2565. [Google Scholar] [CrossRef] [PubMed]
- Stracke, R.; Werber, M.; Weisshaar, B. The r2r3-myb gene family in arabidopsis thaliana. Curr. Opin. Plant Biol. 2001, 4, 447–456. [Google Scholar] [CrossRef] [PubMed]
- Armstrong, C.L.; Romero-Severson, J.; Hodges, T.K. Improved tissue culture response of an elite maize inbred through backcross breeding, and identification of chromosomal regions important for regeneration by rflp analysis. Theor. Appl. Genet. 1992, 84, 755–762. [Google Scholar] [PubMed]
- Armstrong, C.L.; Green, C.E. Establishment and maintenance of friable, embryogenic maize callus and the involvement of l-proline. Planta 1985, 164, 207–214. [Google Scholar] [CrossRef] [PubMed]
- Frame, B.R.; Zhang, H.; Cocciolone, S.M.; Sidorenko, L.V.; Dietrich, C.R.; Pegg, S.E.; Zhen, S.; Schnable, P.S.; Wang, K. Production of transgenic maize from bombarded type ii callus: Effect of gold particle size and callus morphology on transformation efficiency. Vitr. Cell. Dev. Biol. 2000, 36, 21–29. [Google Scholar] [CrossRef]
- Wolfe, K.; Wu, X.Z.; Liu, R.H. Antioxidant activity of apple peels. J. Agric. Food Chem. 2003, 51, 609–614. [Google Scholar] [CrossRef] [PubMed]
- Coca, M.; Bortolotti, C.; Rufat, M.; Penas, G.; Eritja, R.; Tharreau, D.; del Pozo, A.M.; Messeguer, J.; San Segundo, B. Transgenic rice plants expressing the antifungal afp protein from Aspergillus giganteus show enhanced resistance to the rice blast fungus Magnaporthe grisea. Plant Mol. Biol. 2004, 54, 245–259. [Google Scholar] [CrossRef] [PubMed]
- Harborne, J.B. Phytochemical Methods. A Guide to Modern Techniques of Plant Analysis, 3rd ed.; Champman and Hall: New York, NY, USA, 1998. [Google Scholar]
- Sample Availability: Not available.
© 2015 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ibraheem, F.; Gaffoor, I.; Tan, Q.; Shyu, C.-R.; Chopra, S. A Sorghum MYB Transcription Factor Induces 3-Deoxyanthocyanidins and Enhances Resistance against Leaf Blights in Maize. Molecules 2015, 20, 2388-2404. https://doi.org/10.3390/molecules20022388
Ibraheem F, Gaffoor I, Tan Q, Shyu C-R, Chopra S. A Sorghum MYB Transcription Factor Induces 3-Deoxyanthocyanidins and Enhances Resistance against Leaf Blights in Maize. Molecules. 2015; 20(2):2388-2404. https://doi.org/10.3390/molecules20022388
Chicago/Turabian StyleIbraheem, Farag, Iffa Gaffoor, Qixian Tan, Chi-Ren Shyu, and Surinder Chopra. 2015. "A Sorghum MYB Transcription Factor Induces 3-Deoxyanthocyanidins and Enhances Resistance against Leaf Blights in Maize" Molecules 20, no. 2: 2388-2404. https://doi.org/10.3390/molecules20022388
APA StyleIbraheem, F., Gaffoor, I., Tan, Q., Shyu, C.-R., & Chopra, S. (2015). A Sorghum MYB Transcription Factor Induces 3-Deoxyanthocyanidins and Enhances Resistance against Leaf Blights in Maize. Molecules, 20(2), 2388-2404. https://doi.org/10.3390/molecules20022388






