Possible Roles of Plant Sulfurtransferases in Detoxification of Cyanide, Reactive Oxygen Species, Selected Heavy Metals and Arsenate
Abstract
:1. Introduction
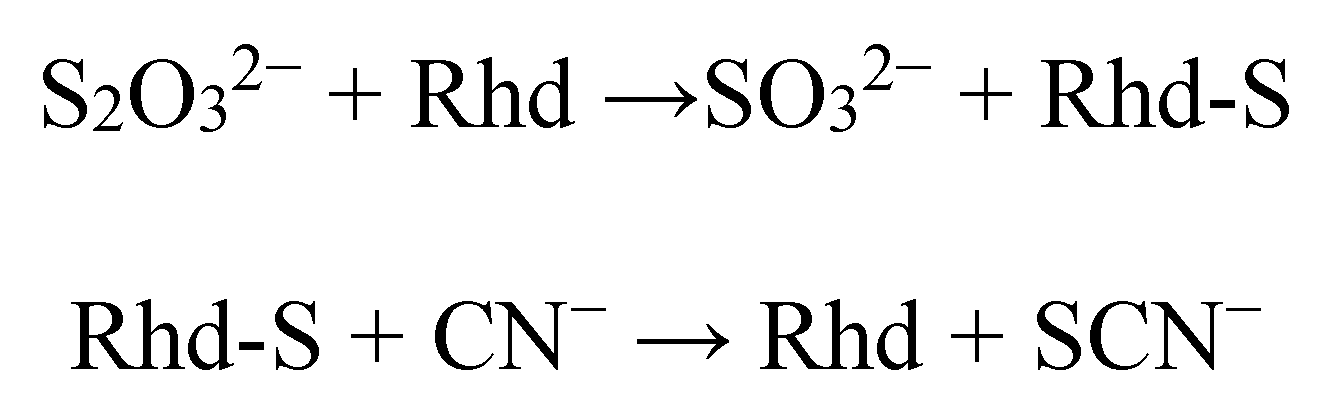
2. Detoxification of Cyanide
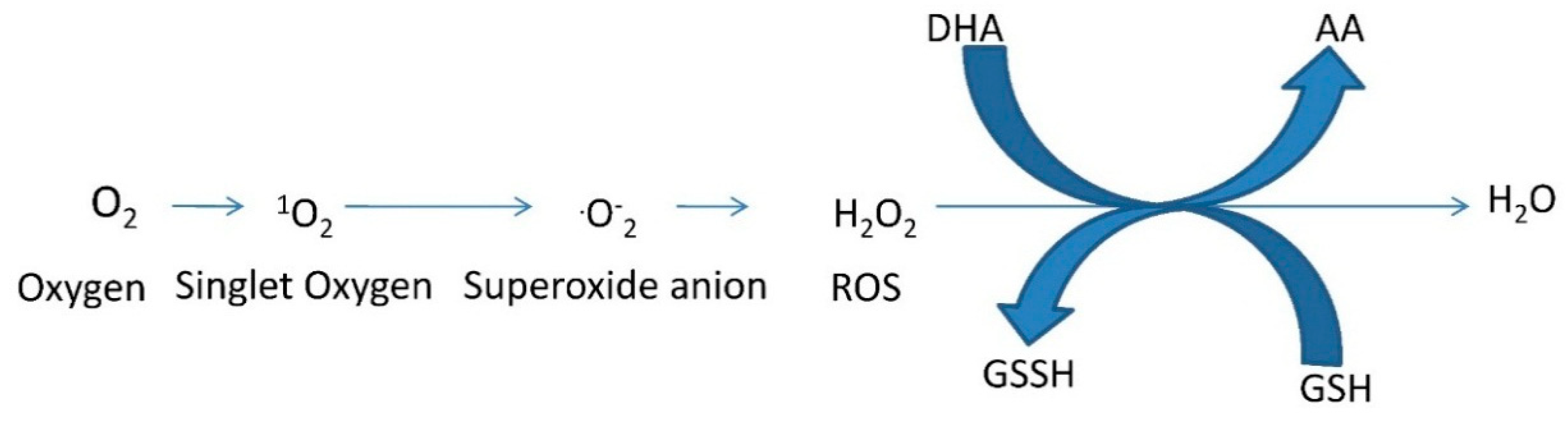
3. Detoxification of Reactive Oxygen Species
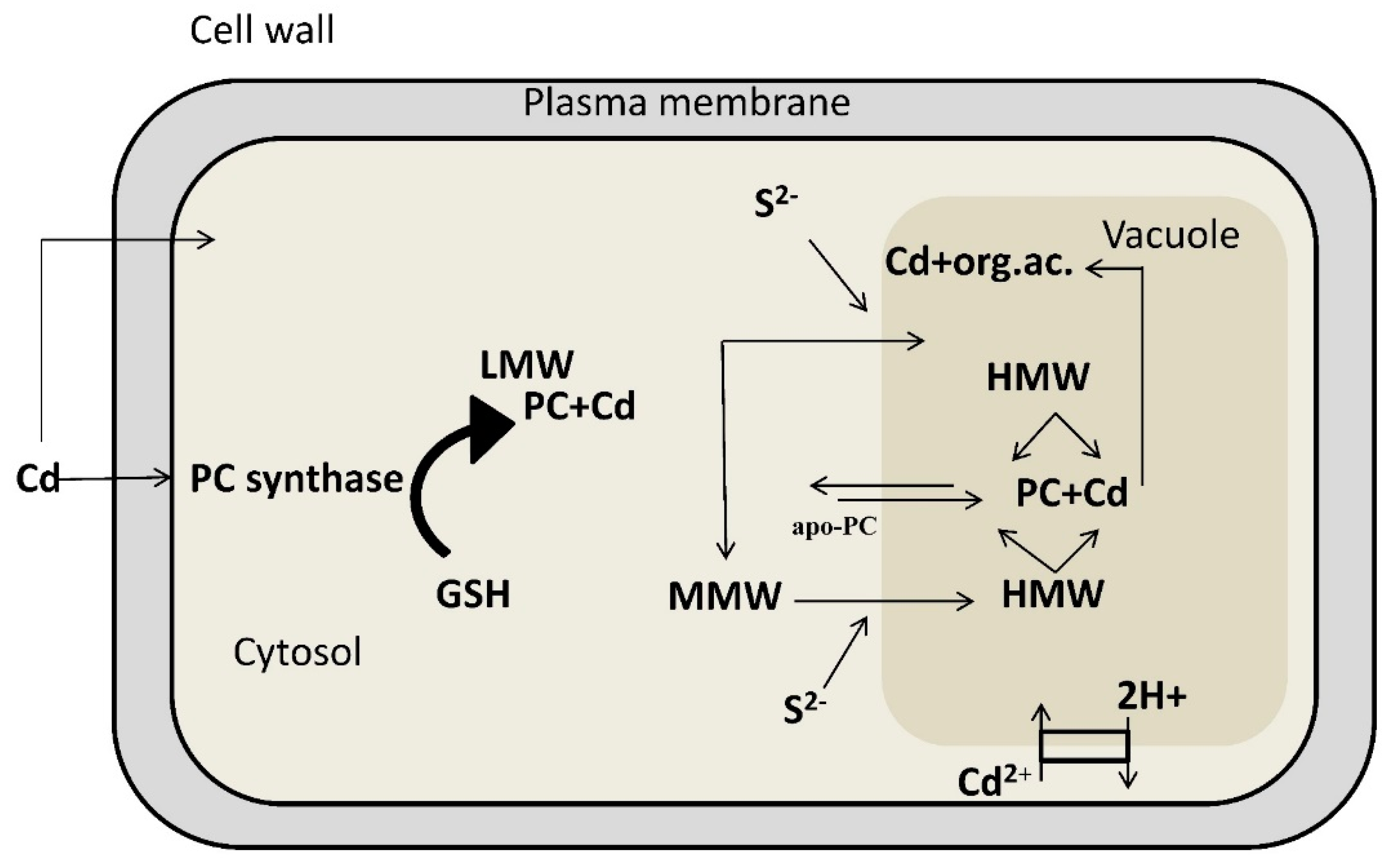
4. Detoxification of Heavy Metals
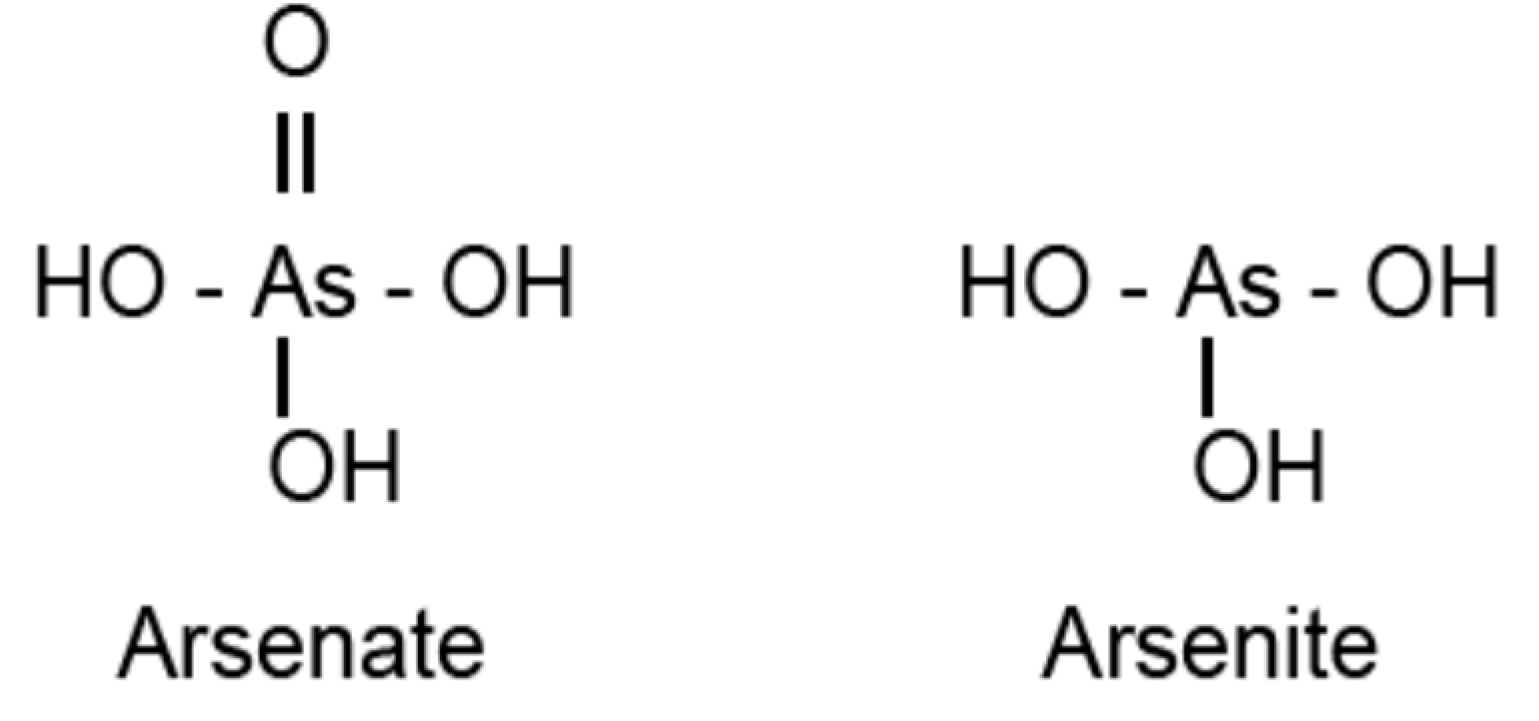
4.1. Sulfurtransferases with Arsenate Reductase Activity
4.2. AtStr5 as Arsenate Reductase: Possible Role in Arsenic Phytoremediation
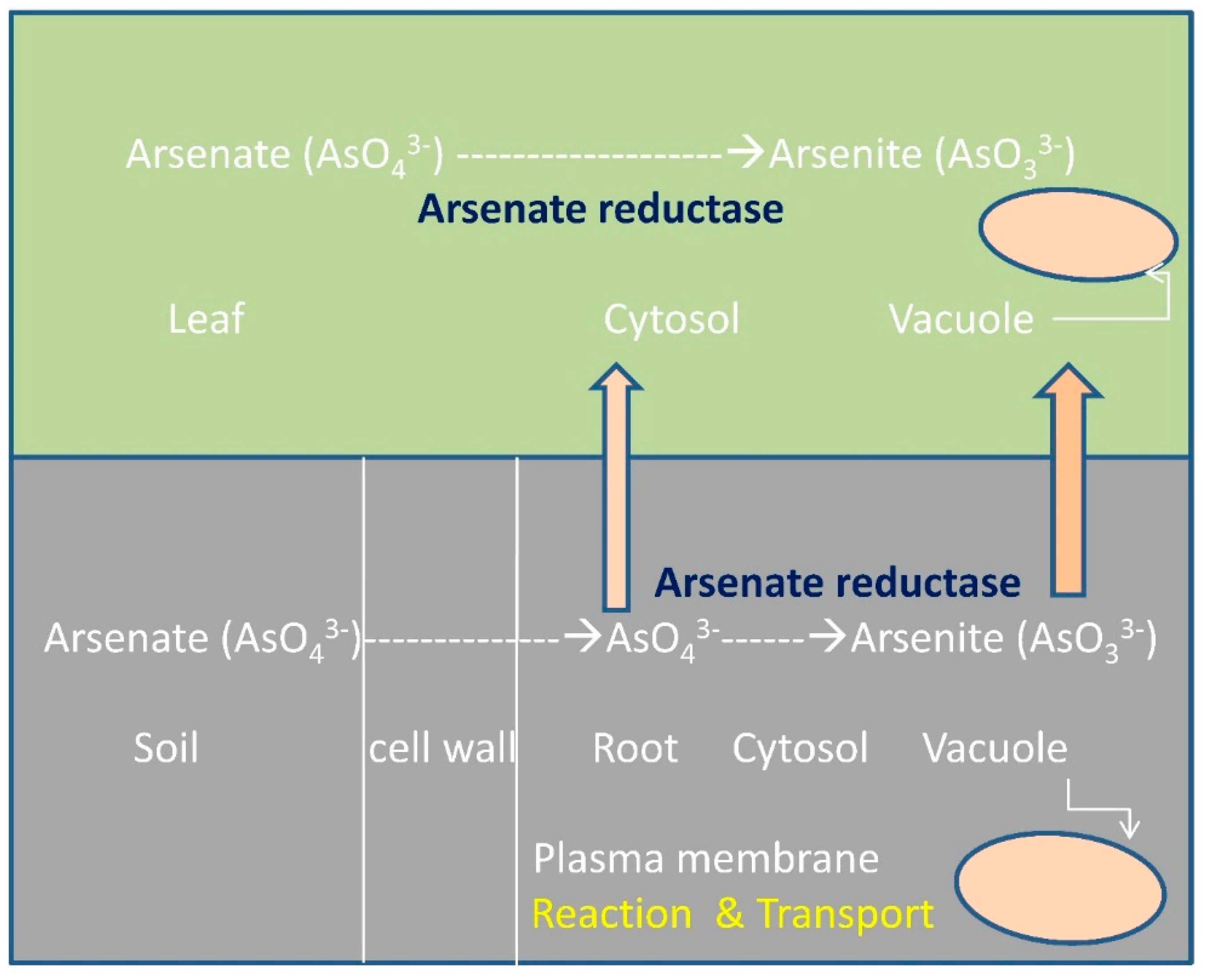
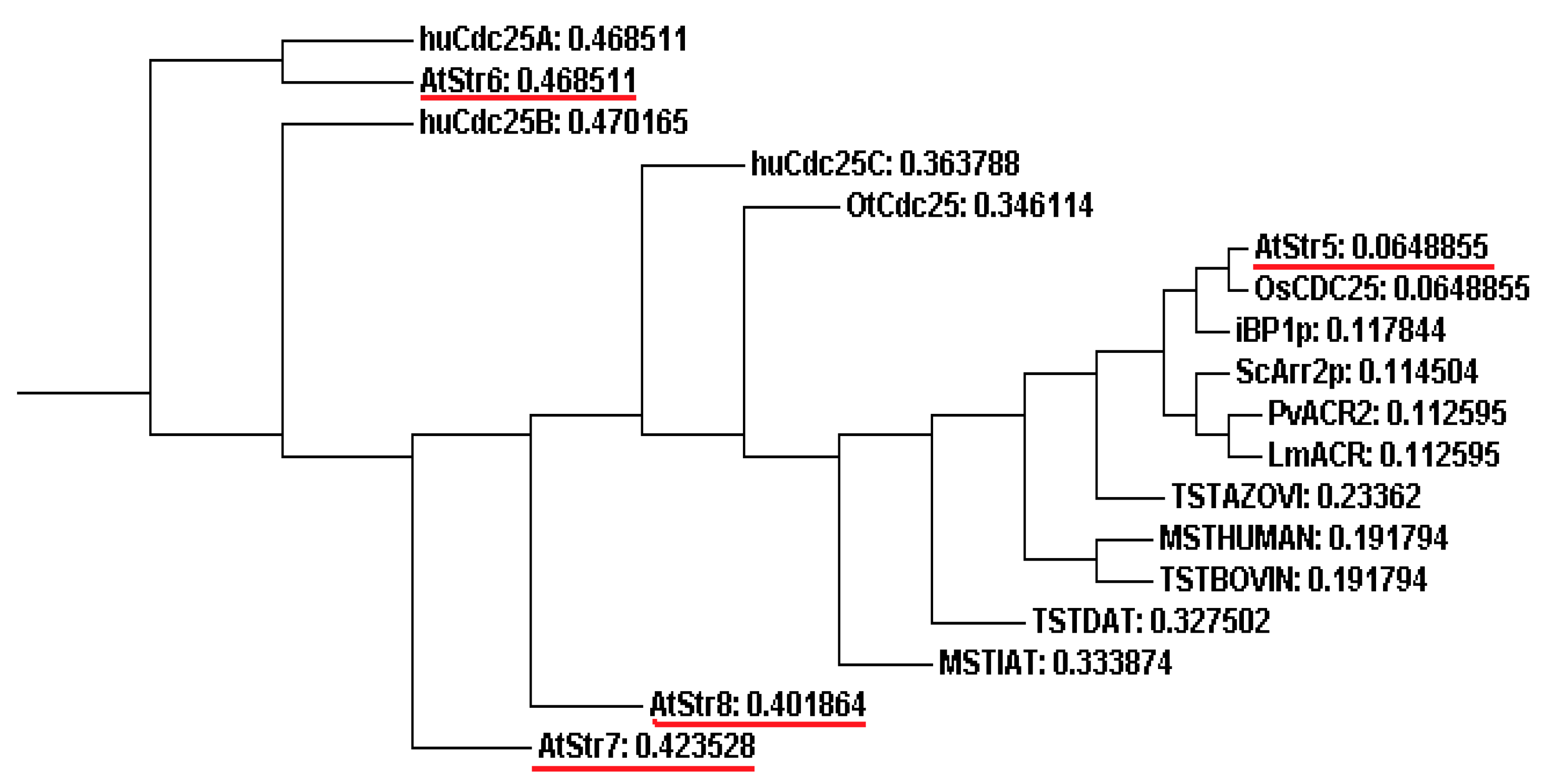
4.3. Phytoremediation
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Papenbrock, J.; Guretzki, S.; Henne, M. Latest news about the sulfurtransferases of higher plants. Amino Acids 2010, 41, 53–57. [Google Scholar]
- Gliubich, F.; Gazerro, M.; Zanotti, G.; Delbono, S.; Bombieri, G.; Berni, R. Active site structural features for chemically modified forms of rhodanase. J. Biol. Chem. 1996, 27, 21054–21061. [Google Scholar]
- Westley, J. Rhodanese. Adv. Enzymol. Relat. Areas Mol. Biol. 1973, 39, 327–368. [Google Scholar]
- Ray, W.K.; Zeng, G.; Potters, M.B.; Mansuri, A.M.; Larson, T.J. Characterization of a 12-kilodalton rhodanese encoded by glpE of Escherichia coli and its interaction with thioredoxin. J. Bacteriol. 2000, 182, 2277–2284. [Google Scholar] [CrossRef] [PubMed]
- Spallarossa, A.; Donahue, J.L.; Larson, T.J.; Bolognesi, M.; Bordo, D. Escherichia coli GlpE is a prototype sulfurtransferase for the single-domain rhodanese homology superfamily. Structure 2001, 9, 1117–1125. [Google Scholar] [CrossRef] [PubMed]
- Adams, H.; Teertstra, W.; Koster, M.; Tommassen, J. PspE (phage shock protein E) of Escherichia coli is a rhodanese. FEBS Lett. 2002, 518, 173–176. [Google Scholar] [CrossRef] [PubMed]
- Heidelberg, J.F.; Eisen, J.A.; Nelson, W.C.; Clayton, R.A.; Gwinn, M.L.; Dodson, R.J.; Haft, D.H.; Hickey, E.K.; Peterson, J.D.; Umayam, L.A.; et al. DNA sequence of both chromosomes of the cholera pathogen Vibrio cholerae. Nature 2000, 406, 477–483. [Google Scholar] [CrossRef] [PubMed]
- Azumi, Y.; Watanabe, A. Evidence for a senescence-associated gene induced by darkness. Plant Physiol. 1991, 95, 577–583. [Google Scholar] [CrossRef] [PubMed]
- Oh, S.A.; Lee, S.Y.; Chung, I.K.; Lee, C.H.; Nam, H.G. A senescence-associated gene of Arabidopsis thaliana is distinctively regulated during natural and artificially induced leaf senescence. Plant Mol. Biol. 1996, 30, 739–754. [Google Scholar] [CrossRef] [PubMed]
- Ploegman, J.H.; Drent, G.; Kalk, K.H.; Hol, W.G. Structure of bovine liver rhodanese. I. Structure determination at 2.5 A resolution and a comparison of the conformation and sequence of its two domains. J. Mol. Biol. 1978, 123, 557–594. [Google Scholar] [CrossRef] [PubMed]
- Bordo, D.; Deriu, D.; Colnaghi, R.; Carpen, A.; Pagani, S.; Bolognesi, M. The crystal structure of a sulfurtransferase from Azotobacter vinelandii highlights the evolutionary relationship between the rhodanese and phosphatase enzyme families. J. Mol. Biol. 2000, 298, 691–704. [Google Scholar] [CrossRef] [PubMed]
- Bordo, D.; Bork, P. The rhodanese/Cdc25 phosphatase super family: Sequence structure and functions relations. EMBO Rep. 2002, 3, 741–746. [Google Scholar] [CrossRef] [PubMed]
- Forlani, F.; Carpen, A.; Pagani, S. Evidence that elongation of the catalytic loop of the Azotobacter vinelandii rhodanese changed selectivity from sulfur-to phosphate-containing substrates. Protein Eng. 2003, 16, 515–519. [Google Scholar] [CrossRef] [PubMed]
- Hofmann, K.; Bucher, P.; Kajava, A.V. A model of Cdc25 phosphatase activity catalytic domain and Cdk-interaction surface based on the presence of a rhodanese homology domain. J. Mol. Biol. 1998, 282, 195–208. [Google Scholar] [CrossRef] [PubMed]
- Farooq, A.; Chaturvedi, G.; Mujtaba, S.; Plotnikova, O.; Zeng, L.; Dhalluin, C.; Ashton, R.; Zhou, M.M. Solution structure of ERK2 binding domain of MAPK phosphatase MKP-3: Structural insights into MKP-3 activation by ERK2. Mol. Cell 2001, 7, 387–399. [Google Scholar] [CrossRef] [PubMed]
- Fauman, E.B.; Cogswell, J.P.; Lovejoy, B.; Rocque, W.J.; Holmes, W.; Montana, V.G.; Rink, M.J.; Piwnica-Worms, H.; Saper, M.A. Crystal structure of the catalytic domain of the human cell cycle control phosphatase, Cdc25A. Cell 1998, 93, 617–625. [Google Scholar] [CrossRef] [PubMed]
- Nagahara, N.; Ito, T.; Minami, M. Mercaptopyruvate sulfurtransferase as a defense against cyanide toxification: Molecular properties and mode of detoxification. Histol. Hispathol. 1999, 14, 1277–1286. [Google Scholar]
- Bartels, A.; Mock, H.P.; Papenbrock, J. Differential expression of Arabidopsis sulfurtransferases under various growth conditions. Plant Physiol. Biochem. 2007, 45, 178–187. [Google Scholar] [CrossRef] [PubMed]
- Bauer, M.; Papenbrock, J. Identification and characterization of single-domain thiosulfate sulfurtransferases from Arabidopsis thaliana. FEBS Lett. 2002, 532, 427–431. [Google Scholar] [CrossRef] [PubMed]
- Landrieu, I.; da Costa, M.; de Veylder, L.; Dewitte, F.; Vandepoele, K.; Hassan, S.; Wieruszeski, J.M.; Corellou, F.; Faure, J.D.; van Montagu, M.; et al. A small CDC25 dual-specificity tyrosine-phosphatase isoform in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2004, 101, 13380–13385. [Google Scholar] [CrossRef] [PubMed]
- Niu, J.S.; Yu, L.; Ma, Z.-Q.; Chen, P.-D.; Liu, D.-J. Molecular cloning, characterization and mapping of a rhodanese like gene in wheat. Acta Genet. Sin. 2002, 29, 266–272. [Google Scholar] [PubMed]
- Louie, M.; Kondor, N.; Dewitt, J.G. Gene expression in cadmium-tolerant Datura innoxia: Detection and characterization of cDNAs induced in response to Cd2+. Plant Mol. Biol. 2003, 52, 81–89. [Google Scholar] [CrossRef] [PubMed]
- Henny, C.J.; Hallock, R.J.; Hill, E.F. Cyanide and migratory birds at gold-mines in Nevada, USA. Ecotoxicology 1994, 3, 45–58. [Google Scholar] [CrossRef] [PubMed]
- Yip, W.-K.; Yang, S.F. Cyanide metabolism in relation to ethylene production in plant tissues. Plant Physiol. 1988, 88, 473–476. [Google Scholar] [CrossRef] [PubMed]
- Doucleff, M.; Terry, N. Pumping out the arsenic. Nat. Biotechnol. 2002, 20, 1094–1096. [Google Scholar] [CrossRef] [PubMed]
- Oracz, K.; El-Maarouf-Bouteau, H.; Kranner, I.; Bogatek, R.; Corbineau, F.; Bailly, C. The mechanisms involved in seed dormancy alleviation by hydrogen cyanide unravel the role of reactive oxygen species as key factors of cellular signaling during germination. Plant Physiol. 2009, 150, 494–505. [Google Scholar] [CrossRef] [PubMed]
- Ressler, C.; Tatake, J.G. Vicianin, prunasin, and β-cyanoalanine in common vetch seed as sources of urinary thiocyanate in the rat. J. Agric. Food Chem. 2001, 49, 5075–5080. [Google Scholar] [CrossRef] [PubMed]
- Cipollone, R.P.; Ascenzi, P.; Tomao, F.; Imperi, P.; Visca, P. Enzymatic detoxification of cyanide: Clues from Pseudomonas aeruginosa Rhodanese. J. Mol. Microbiol. Biotechnol. 2008, 15, 199–211. [Google Scholar] [CrossRef] [PubMed]
- Meyer, T.; Burow, M.; Bauer, M.; Papenbrock, J. Arabidopsis sulfurtransferases: Investigation of their function during senescence and in cyanide detoxification. Planta 2003, 217, 1–10. [Google Scholar] [PubMed]
- Machingura, M.; Ebbs, S.D. Functional redundancies in cyanide tolerance provided by β-cyanoalanine pathway genes in Arabidopsis thaliana. Int. J. Plant Sci. 2014, 175, 346–358. [Google Scholar] [CrossRef]
- Hatzfeld, Y.; Maruyama, A.; Schmidt, A.; Noji, M.; Ishizawa, K.; Saito, K. β-cyanoalanine synthase is a mitochondrial cysteine synthase-like protein in spinach and Arabidopsis. Plant Physiol. 2000, 123, 1163–1172. [Google Scholar] [CrossRef] [PubMed]
- Piotrowski, M.; Schonfelder, S.; Weiler, E.W. The Arabidopsis thaliana isogene NIT4 and its orthologs in tobacco encode β-cyano-l-alanine hydratase/nitrilase. J. Biol. Chem. 2001, 276, 2616–2621. [Google Scholar] [CrossRef] [PubMed]
- Moller, I.M. Plant mitochondria and oxidative stress: Electron transport, NADPH turnover, and metabolism of reactive oxygen species. Annu. Rev. Plant Physiol. Mol. Biol. 2001, 52, 561–591. [Google Scholar] [CrossRef]
- Uhteg, L.C.; Westley, J. Purification and steady-state kinetic analysis of yeast thiosulfate reductase. Arch. Biochem. Biophys. 1979, 195, 211–222. [Google Scholar] [CrossRef] [PubMed]
- Slater, A.; Scott, N.W.; Fowler, M.R. Plant Biotechnology: The Genetic Manipulation of Plants, 2nd ed.; Oxford University Press: Oxford, UK, 2008; p. 231. [Google Scholar]
- Mittler, R.; Vanderauwera, S.; Gollery, M.; van Breusegem, F. Reactive oxygen gene network of plants. Trends Plant Sci. 2004, 9, 490–498. [Google Scholar] [CrossRef] [PubMed]
- Chalapathi Rao, A.S.V.; Reddy, A.R. Glutathione Reductase: A Putative Redox Regulatory System in Plant Cells. Sulfur Assimilation and Abiotic Stresses in Plants; Khan, N.A., Singh, S., Umar, S., Eds.; Springer: Heidelberg, Berlin, 2008; pp. 111–147. [Google Scholar]
- Ding, S.; Lu, Q.; Zhang, Y.; Yang, Z.; Wen, X.; Zhang, L.; Lu, C. Enhanced sensitivity to oxidative stress in transgenic tobacco plants with decreased glutathione reductase activity leads to a decrease in ascorbate pool and ascorbate redox state. Plant Mol. Biol. 2009, 69, 577–592. [Google Scholar] [CrossRef] [PubMed]
- Foyer, C.H.; Noctor, G. Redox homeostasis and antioxidant signaling: A metabolic interface between stress perception and physiological responses. Plant Cell 2005, 17, 1866–1875. [Google Scholar] [CrossRef] [PubMed]
- Dixon, D.P.; Davis, B.G.; Edwards, E. Functional divergence in the glutathione transferase super-family in plants: Identification of two classes with putative functions in redox homeostasis in Arabidopsis thaliana. J. Biol. Chem. 2002, 277, 30859–30869. [Google Scholar] [CrossRef] [PubMed]
- Toohey, T.I. Sulphane sulfur in biological systems: A possible regulatory role. Biochemistry 1989, 264, 625–632. [Google Scholar]
- Kaczor-Kamińska, M.; Sura, P.; Wróbel, M. Changes in activity of three sulfurtransferases in response to exposure to cadmium, lead and mercury ions. J. Environ. Prot. 2013, 4, 19–28. [Google Scholar] [CrossRef]
- Tomsett, A.B.; Thurman, D.A. Molecular biology of metal tolerances of plants. Plant Cell Environ. 1988, 11, 383–394. [Google Scholar] [CrossRef]
- Nocito, F.F.; Pirovano, L.; Cocucci, M.; Sacchi, G.A. Cadmium-induced sulfate uptake in maize roots. Plant Physiol. 2002, 129, 1872–1879. [Google Scholar] [CrossRef] [PubMed]
- Gutiérez-Alcalá, G.; Gotor, C.; Meyer, A.J.; Fricker, M.; Vega, J.M.; Romeo, L.C. Glutathione biosynthesis in Arabidopsis trichome cells. Proc. Natl. Acad. Sci. USA 2000, 97, 11108–11113. [Google Scholar] [CrossRef] [PubMed]
- Bick, J.A.; Setterdahl, A.T.; Knaff, D.B.; Chen, Y.; Pitcher, L.H.; Zilinskas, B.A.; Leustek, T. Regulation of the plant-type 5-adenylsulfate reductase by oxidative stress. Biochemistry 2001, 40, 9040–9048. [Google Scholar] [CrossRef] [PubMed]
- Kazuki, S. Sulfur assimilatory metabolism: The long and smelling road. Plant Physiol. 2004, 136, 2443–2450. [Google Scholar] [CrossRef] [PubMed]
- Peer, W.A.; Baxter, I.R.; Richards, E.L.; Freeman, J.L.; Murphy, A.S. Phytoremediation and hyperaccumulator plants. In Molecular Biology of Metal Homeostasis and Detoxification; Tamás, M.J., Martinoia, E., Eds.; Springer: Heidelberg, Berlin, Germany, 2005; Volume 14, pp. 299–330. [Google Scholar]
- Fernandes, A.P.; Holmgren, A. Glutaredoxin: Glutathione-dependent redox enzymes with functions far beyond a simple thioredoxin backup system. Antioxid. Redox Signal. 2004, 6, 63–74. [Google Scholar] [CrossRef] [PubMed]
- Duan, G.L.; Zhu, Y.G.; Tong, Y.P.; Cai, C.; Kneer, R. Characterization of arsenate reductase in the extract of roots and fronds of Chinese brake fern, an arsenic hyperaccumulator. Plant Physiol. 2005, 138, 461–469. [Google Scholar] [CrossRef] [PubMed]
- Sorrell, D.A.; Chrimes, D.; Dickinson, J.R.; Rogers, H.J.; Francis, D. The Arabidopsis CDC25 induces a short cell length when overexpressed in fission yeast: Evidence for cell cycle functions. New Phytol. 2005, 165, 425–428. [Google Scholar] [CrossRef] [PubMed]
- Landrieu, I.; Hassan, S.; Sauty, M.; Dewitte, F.; Wieruszeski, J.M.; Inzé, D.; de Veylder, L.; Lippens, G. Characterization of the Arabidopsis thaliana Arath; CDC25 dual-specificity tyrosine phosphatase. Biochem. Biophys. Res. Commun. 2004, 322, 734–739. [Google Scholar] [CrossRef] [PubMed]
- Ogasawara, Y.; Lacourciere, G.; Stadtman, T.C. Formation of a selenium-substituted rhodanese by reaction with selenite and glutathione: Possible role of a protein perselenide in a selenium delivery system. Proc. Natl. Acad. Sci. USA 2001, 98, 9494–9498. [Google Scholar] [CrossRef] [PubMed]
- Mukhopadhyay, R.; Shi, J.; Rosen, B.P. Purification and characterization of ACR2p, the Saccharomyces cerevisiae arsenate reductase. J. Biol. Chem. 2000, 275, 21149–21157. [Google Scholar] [CrossRef] [PubMed]
- Mukhopadhyay, R.; Rosen, B.P. The phosphatase C(X)5R motif is required for catalytic activity of the Saccharomyces cerevisiae Acr2p arsenate reductase. J. Biol. Chem. 2001, 276, 34738–34742. [Google Scholar] [CrossRef] [PubMed]
- Mukhopadhyay, R.; Zhou, Y.; Rosen, B.P. Directed evolution of a yeast arsenate reductase into a protein-tyrosine phosphatase. J. Biol. Chem. 2003, 278, 24476–24480. [Google Scholar] [CrossRef] [PubMed]
- Bleeker, P.M.; Hakvoort, H.W.; Bliek, M.; Souer, E.; Schat, H. Enhanced arsenate reduction by a Cdc25-like tyrosine phosphatase explains increased phytochelatin accumulation in arsenate-tolerant Holcus lanatus. Plant J. 2006, 45, 917–929. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Bhattacharjee, H.; Mukhopadhyay, R. Bifunctional role of the leishmanial antimonate reductase LmACR2 as a protein tyrosine phosphatase. Mol. Biochem. Parasitol. 2006, 148, 161–168. [Google Scholar] [CrossRef] [PubMed]
- Dhankher, O.P.; Rosen, B.P.; McKinney, E.C.; Meagher, R.B. Hyper-accumulation of arsenic in the shoots of Arabidopsis silenced for arsenate reductase, ACR2. Proc. Natl. Acad. Sci. USA 2006, 103, 5413–5418. [Google Scholar] [CrossRef] [PubMed]
- Leonard, A.; Lauwerys, R. Carcinogenicity, teratogenicity, and mutagenicity of arsenic. Mutat. Res. 1980, 75, 49–62. [Google Scholar] [CrossRef] [PubMed]
- Helleday, T.; Nilsson, R.; Jenssen, D. Arsenic (III) and heavy metal ions induce intrachromosomal homologous recombination in the hprt gene of V79 Chinese hamster cells. Environ. Mol. Mutagen. 2000, 35, 114–122. [Google Scholar] [CrossRef] [PubMed]
- Chou, W.C.; Jie, C.; Kenedy, A.A.; Jones, R.J.; Trush, M.A.; Dang, C.V. Role of NADPH oxidase in arsenic-induced reactive oxygen species formation and cytotoxicity in myeloid leukemia cells. Proc. Natl. Acad. Sci. USA 2004, 101, 4578–4583. [Google Scholar] [CrossRef] [PubMed]
- Carbonell, A.A.; Aarabi, M.A.; Delaune, R.D.; Grambrell, R.P.; Patrick, W.H. Arsenic in wetland vegetation: Availability, phytotoxicity, uptake and effects on plants growth and nutrition. Sci. Total Environ. 1998, 217, 189–199. [Google Scholar] [CrossRef]
- Shah, K.; Nongkynrih, J.M. Metal hyperaccumulation and bioremediation. Biol. Plant. 2007, 51, 618–634. [Google Scholar] [CrossRef]
- Li, R.; Haile, J.D.; Kennelly, P.J. An arsenate reductase from Synechocystis sp. strain PCC 6803 exhibits a novel combination of catalytic characteristics. J. Bacteriol. 2003, 185, 6780–6789. [Google Scholar] [CrossRef] [PubMed]
- Shipley, S.; Nordin, A.B.; Tang, C.G.; Kim, S.K. Phytoremediation for arsenic contamination: Arsenate reductase. The Pulse 2008, 6, 1–12. [Google Scholar]
© 2015 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Most, P.; Papenbrock, J. Possible Roles of Plant Sulfurtransferases in Detoxification of Cyanide, Reactive Oxygen Species, Selected Heavy Metals and Arsenate. Molecules 2015, 20, 1410-1423. https://doi.org/10.3390/molecules20011410
Most P, Papenbrock J. Possible Roles of Plant Sulfurtransferases in Detoxification of Cyanide, Reactive Oxygen Species, Selected Heavy Metals and Arsenate. Molecules. 2015; 20(1):1410-1423. https://doi.org/10.3390/molecules20011410
Chicago/Turabian StyleMost, Parvin, and Jutta Papenbrock. 2015. "Possible Roles of Plant Sulfurtransferases in Detoxification of Cyanide, Reactive Oxygen Species, Selected Heavy Metals and Arsenate" Molecules 20, no. 1: 1410-1423. https://doi.org/10.3390/molecules20011410
APA StyleMost, P., & Papenbrock, J. (2015). Possible Roles of Plant Sulfurtransferases in Detoxification of Cyanide, Reactive Oxygen Species, Selected Heavy Metals and Arsenate. Molecules, 20(1), 1410-1423. https://doi.org/10.3390/molecules20011410






