Characterization of Phosphate Solubilizing Bacteria in Sediments from a Shallow Eutrophic Lake and a Wetland: Isolation, Molecular Identification and Phosphorus Release Ability Determination
Abstract
:1. Introduction
2. Results
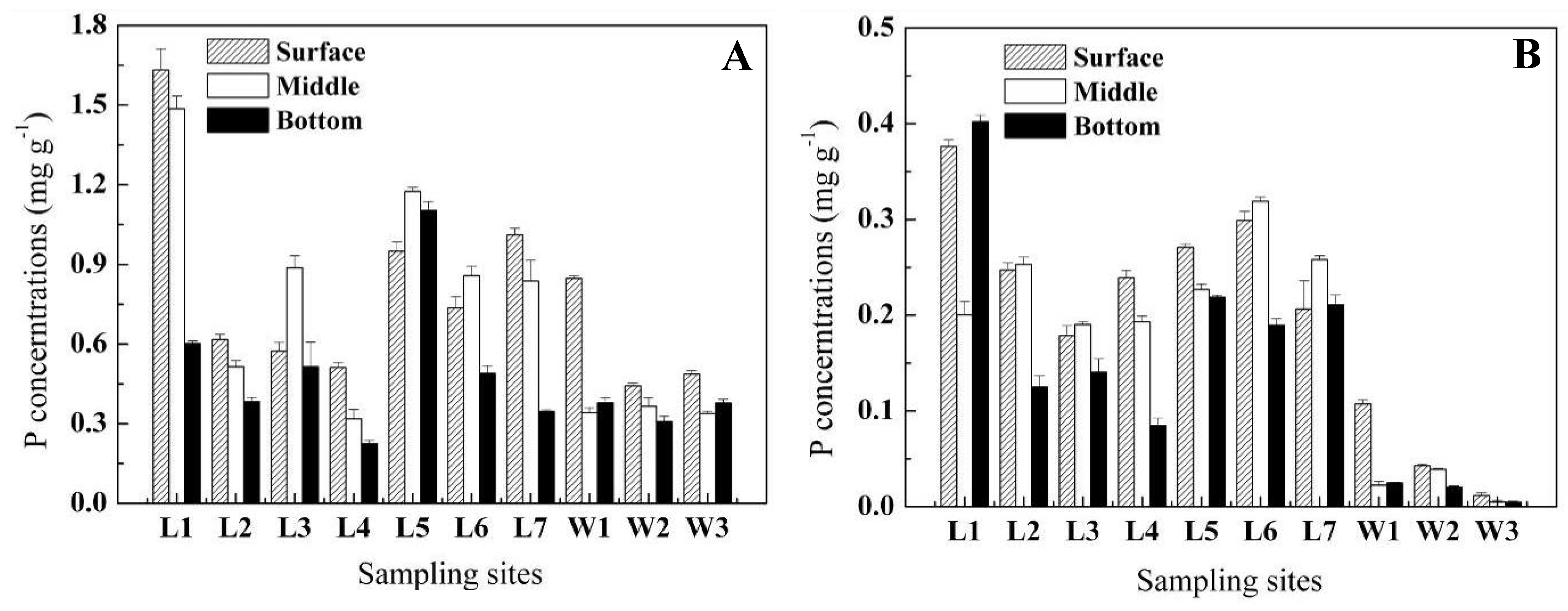
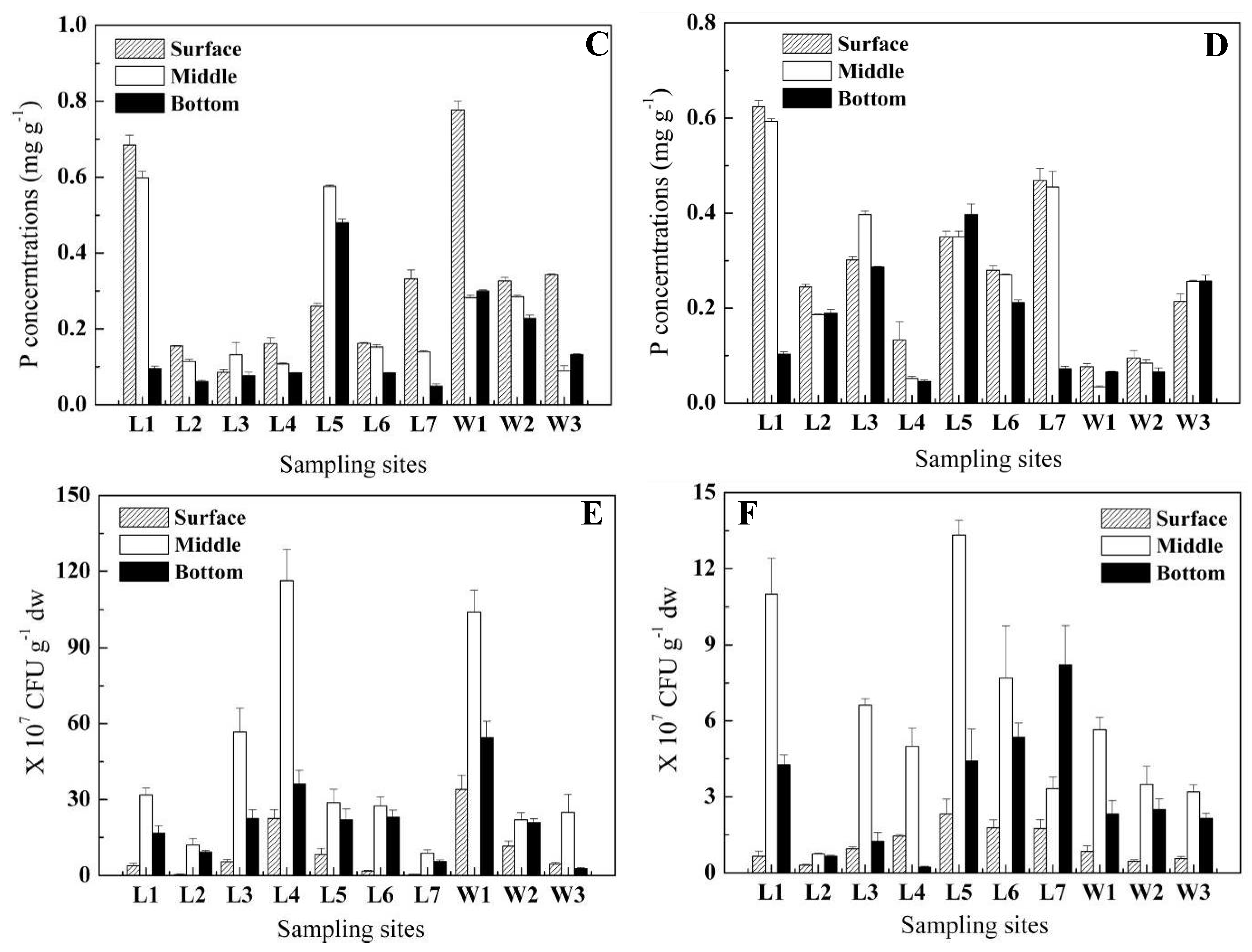
2.1. Vertical distribution of P in sediment
2.2. Abundance of IPB and OPB in sediment
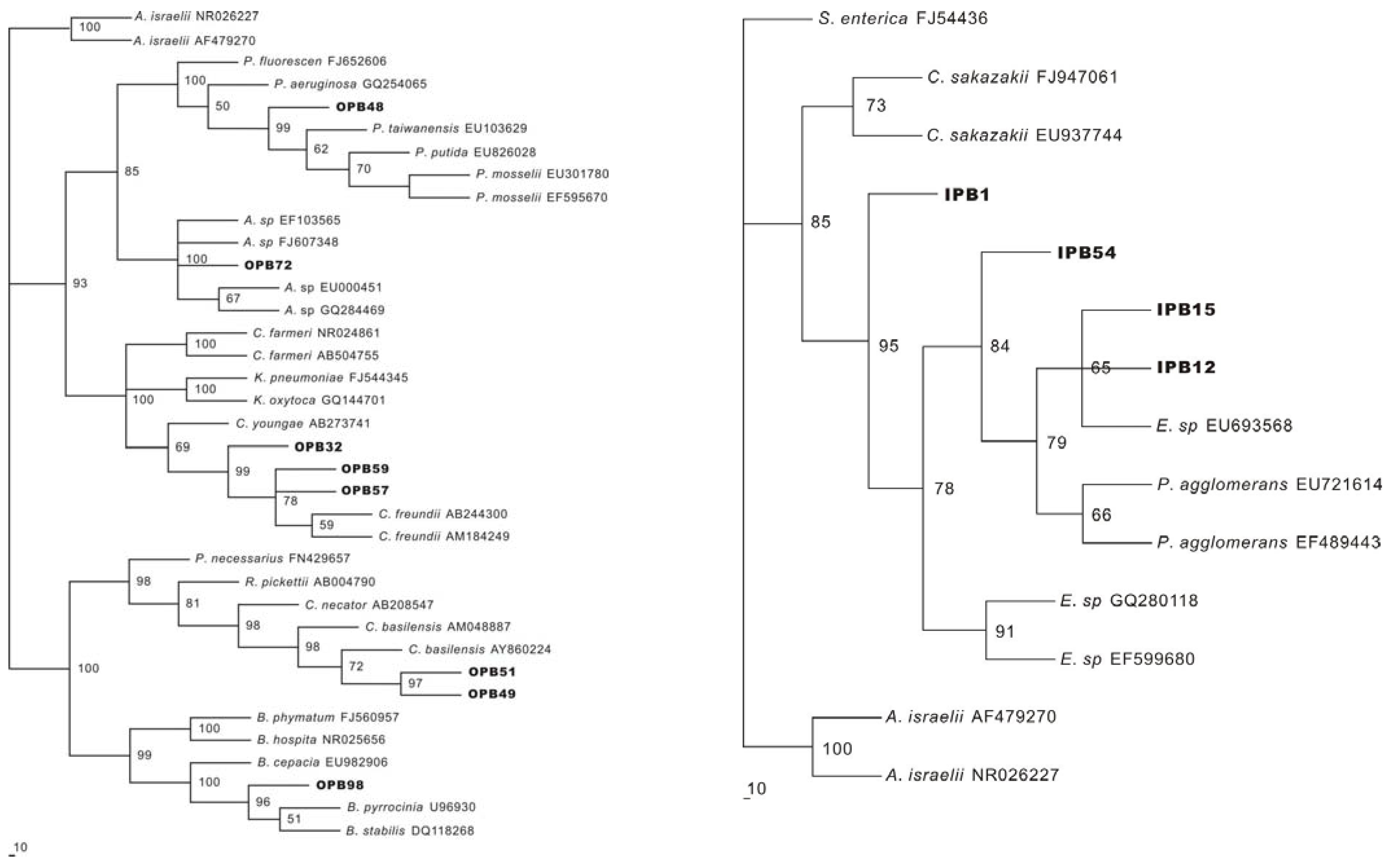
2.3. Identification of P releasing bacteria
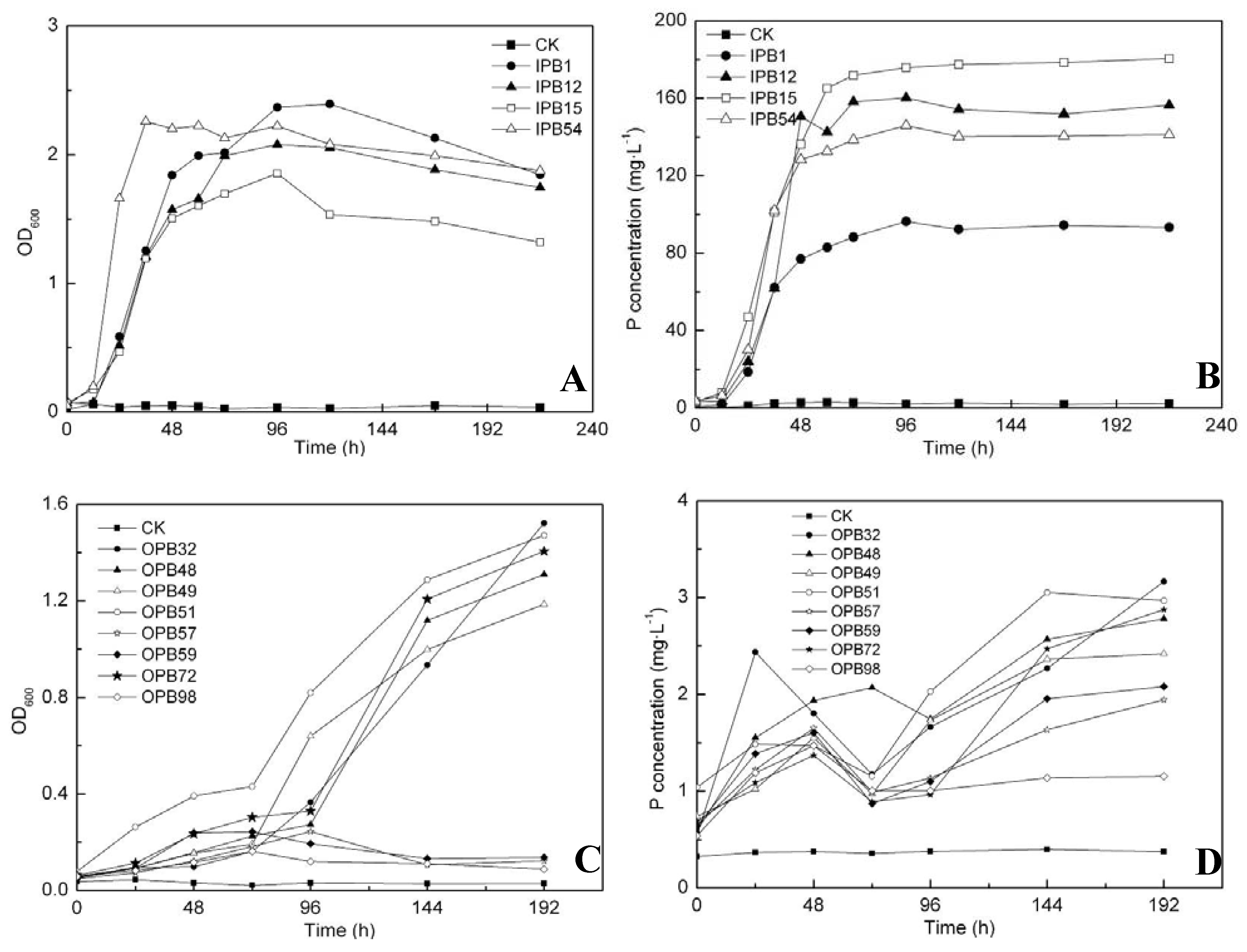
2.4. P release ability of predominant phosphate solubilizing bacteria
3. Discussion
4. Experimental
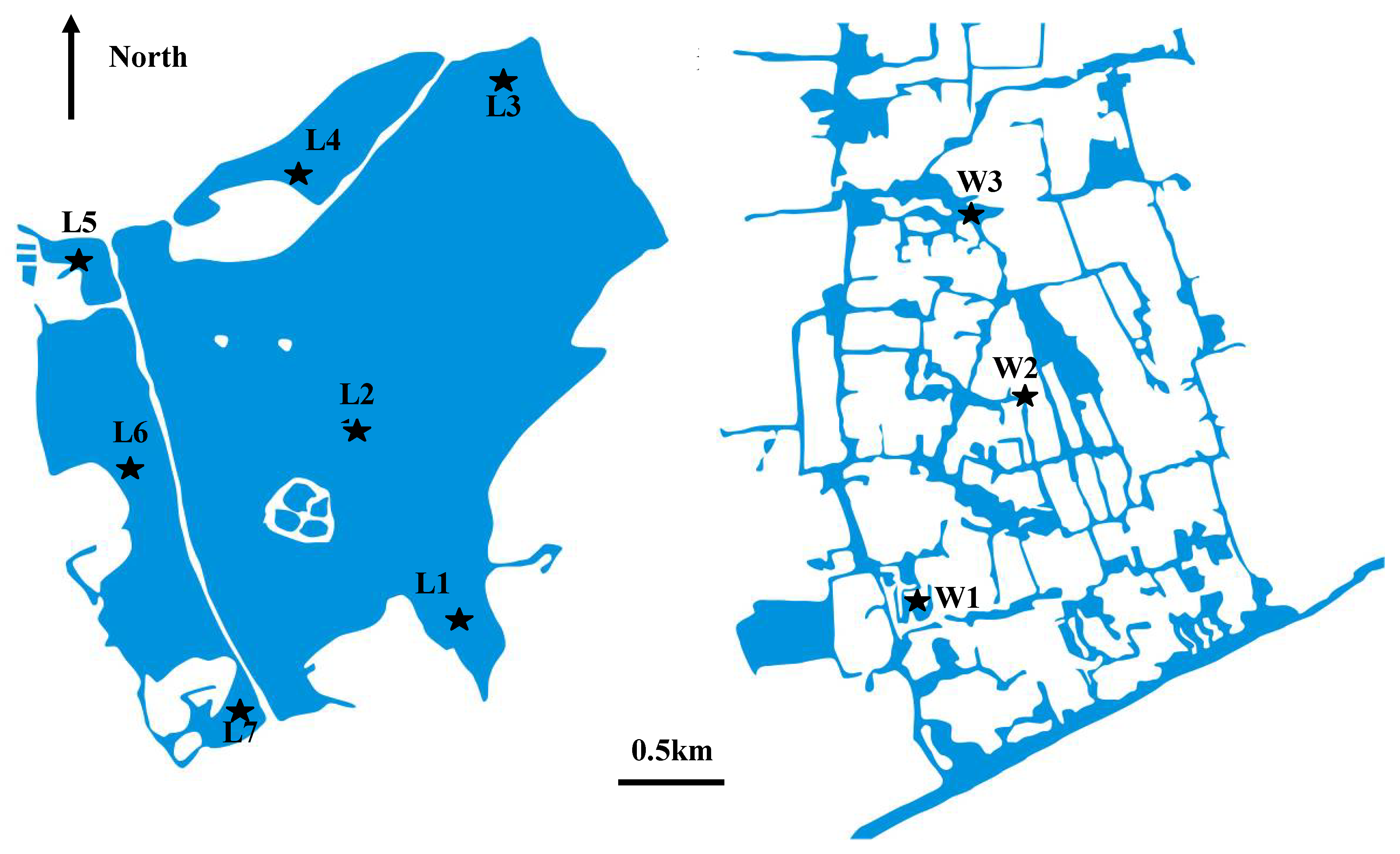
4.1. Description of the study area
4.2. Sediment sampling
4.3. Analysis of P fraction
4.4. Enumeration of phosphate solubilizing bacteria
4.5. Determination of P release ability
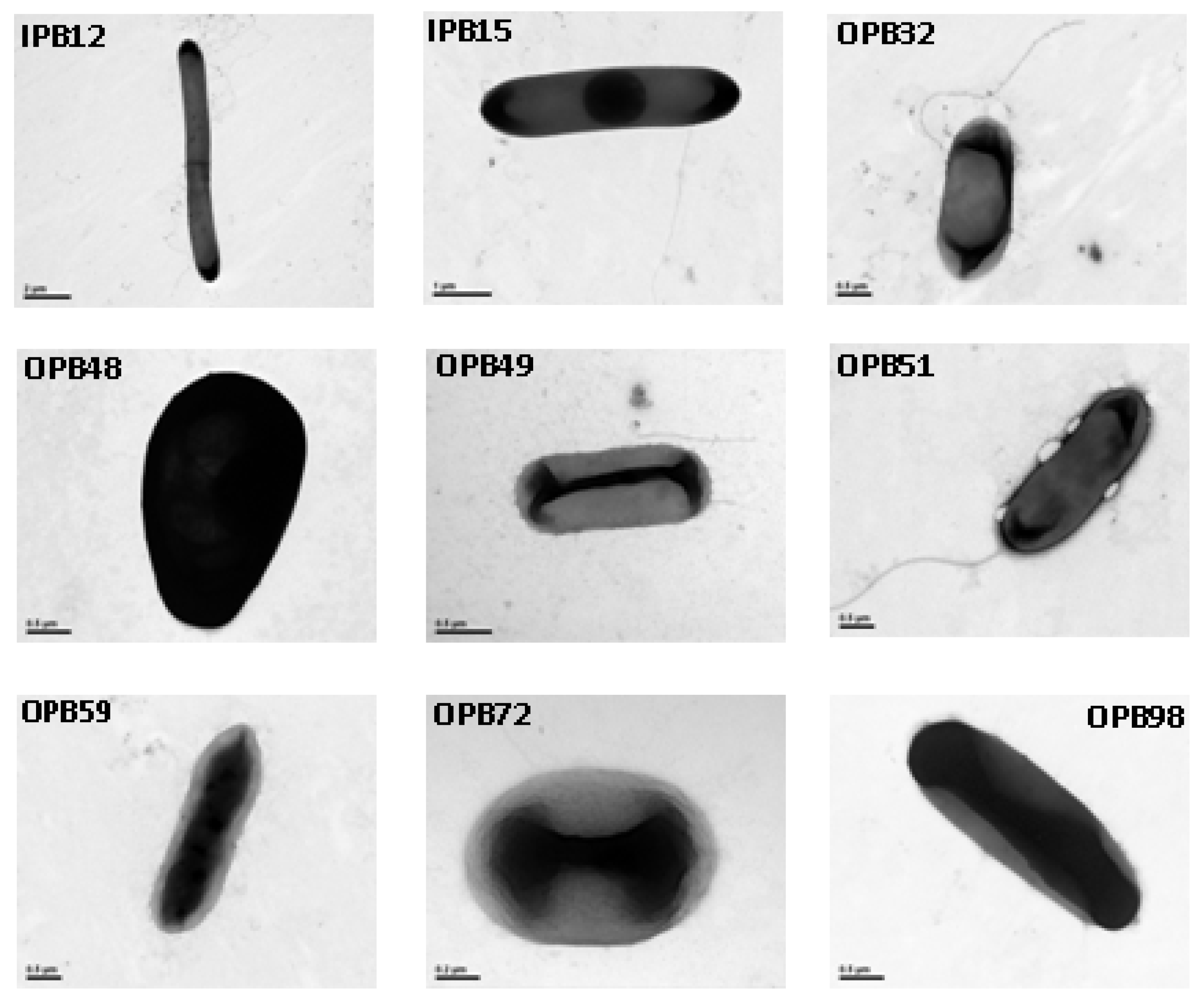
4.6. Morphological and molecular characterization of predominant phosphate solubilizing bacteria
4.7. Statistical analysis
5. Conclusions
Acknowledgements
References
- Sundareshwar, P.V.; Morris, J.T.; Koepfler, E.K.; Fornwalt, B. Phosphorus limitation of coastal ecosystem processes. Science 2003, 299, 563–565. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, D.C.; Martins, G.; Nogueira, R.; Cruz, J.V.; Brito, A.G. Phosphorus fractionation in volcanic lake sediments (Azores - Portugal). Chemosphere 2008, 70, 1256–1263. [Google Scholar] [CrossRef] [PubMed]
- Søndergaard, M.; Jensen, J.P.; Jeppesen, E. Internal phosphorus loading in shallow Danish lakes. Hydrobiologia 1999, 408/409, 145–152. [Google Scholar]
- Spears, M.B.; Carvalho, L.; Perkins, R.; Kirika, A.; Paterson, D.M. Spatial and historical variation in sediment phosphorus fractions and mobility in a large shallow lake. Water Res. 2006, 40, 383–391. [Google Scholar] [CrossRef] [PubMed]
- Golterman, H.L. Fractionation of sediment phosphate with chelating compounds: a simplification, and comparison with other methods. Hydrobiologia 1996, 335, 87–95. [Google Scholar] [CrossRef]
- Tian, J.R.; Zhou, P.J. Phosphorus fractions of floodplain sediments and phosphorus exchange on the sediment–water interface in the lower reaches of the Han River in China. Ecol. Eng. 2007, 30, 264–270. [Google Scholar] [CrossRef]
- Ruban, V.; Brigault, S.; Demare, D.; Philippe, A.M. An investigation of the origin and mobility of phosphorus in freshwater sediments from Bort-Les-Orgues Reservoir, France. J. Environmen. Monitor. 1999, 1, 403–407. [Google Scholar] [CrossRef]
- Wang, H.; Appan, A.; Gulliver, J.S. Modeling of phosphorus dynamics in aquatic sediments: II-examination of model performance. Water Res. 2003, 3, 3939–3953. [Google Scholar] [CrossRef]
- Van, H.E.; Auvray, F.; Deluchat, V.; Chazal, P.M.; Baudu, M. Phosphorus fractionation and short-term mobility in the surface sediment of a polymictic shallow lake treated with a low dose of alum (Courtille Lake, France). Water Air Soil Pollu. 2003, 146, 75–91. [Google Scholar]
- Edlund, G.; Carman, R. Distribution and digenesis of organic and inorganic phosphorus in sediments of the Baltic proper. Chemosphere 2001, 45, 1053–1061. [Google Scholar] [CrossRef]
- Wang, S.R.; Jin, X.C.; Zhao, H.C.; Wu, F.C. Phosphorus release characteristics of different trophic lake sediments under simulative disturbing conditions. J. Hazard. Mater. 2009, 161, 1551–1559. [Google Scholar] [CrossRef] [PubMed]
- Jacobs, H.; Boswell, G.P.; Ritz, K.; Davidson, F.A.; Gadd, G.M. Solubilization of calcium phosphate as a consequence of carbon translocation by Rhizoctonia solani. FEMS Microbiol. Ecol. 2002, 40, 65–71. [Google Scholar] [CrossRef] [PubMed]
- Khoshmanesh, A.; Hart, B.T.; Duncan, A.; Beckett, R. Luxury uptake of phosphorus by sediment bacteria. Water Res. 2002, 36, 774–778. [Google Scholar] [CrossRef]
- Anthony, C. The quinoprotein dehydrogenases for methanol and glucose. Arch. Biochem. Biophys. 2004, 428, 2–9. [Google Scholar] [CrossRef] [PubMed]
- Gautier, A.; Lopin, C.; Garipova, G.; Dubert, O.; Kalinina, I.; Salcedo, C.; Balieu, S.; Glatigny, S.; Valnot, J.; Gouhier, G.; Piettre, S.R. The preparation of new phosphorus-centered functional groups for modified oligonucleotides and other natural phosphates. Molecules 2005, 10, 1048–1073. [Google Scholar] [CrossRef] [PubMed]
- Mino, T.; van Loosdrecht, M.C.M.; Heijnen, J.J. Microbiology and biochemistry of the enhanced biological phosphate removal process. Water Res. 1999, 32, 3193–3207. [Google Scholar] [CrossRef]
- Saghir, K.M.; Zaidi, A.; Wani, P.A. Role of phosphate-solubilizing microorganisms in sustainable agriculture: a review. Agron. Sustain. Dev. 2007, 27, 29–43. [Google Scholar]
- Wu, G.F.; Zhou, X.P. Characterization of phosphorus-releasing bacteria in a small eutrophic shallow, Eastern Lake. Water Res. 2005, 39, 4623–4632. [Google Scholar]
- Zhou, Y.Y.; Li, J.Q.; Zhang, M. Temporal and spatial variations in kinetics of alkaline phosphatase in sediments of a shallow Chinese eutrophic lake (Lake Donghu). Water Res. 2002, 36, 2084–2090. [Google Scholar]
- Vassilev, N.; Toro, M.; Vassileva, M.; Azcon, R.; Barea, J.M. Rock phosphate solubilization by immobilized cells of Enterobacter sp. in fermentation and soil conditions. Bioresouc. Technol. 1999, 61, 29–32. [Google Scholar] [CrossRef]
- Goris, J.; De Vos, P.; Coenye, T.; Hoste, B.; Janssens, D.; Brim, H.; Diels, L.; Mergeay, M.; Kersters, K.; Vandamme, P. Classification of metal-resistant bacteria from industrial biotopes as Ralstonia campinensis sp. nov., Ralstonia metallidurans sp. nov. and Ralstonia basilensis Steinle et al. 1998 emend. Int. J. Syst. Evol. Micr. 2001, 51, 1773–1782. [Google Scholar] [CrossRef] [PubMed]
- Gerischer, U. Acinetobacter Molecular Biology, 1st ed.; Caister Academic Press: Hethersett, UK, 2008. [Google Scholar]
- Ruban, V.; López-Sánchez, J.F.; Pardo, P.; Rauret, G.; Muntau, H.; Quevauviller, Ph. Development of a harmonised phosphorus extraction procedure and certification of a sediment reference material. J. Environmen. Monitor. 2001, 3, 121–125. [Google Scholar] [CrossRef]
- Brigault, S.; Ruban, V. External Phosphorus Load Estimates and P-budget for the hydroelectric reservoir of Bort-les-Orgues, France. Water Air Soil Pollu. 2000, 119, 1–4. [Google Scholar] [CrossRef]
- Sánchez-Román, M.; Rivadeneyra, M.A.; Vasconcelos, C.; McKenzie, J.A. Biomineralization of carbonate and phosphate by moderately halophilic bacteria. FEMS Microbiol. Ecol. 2007, 61, 273–284. [Google Scholar] [CrossRef] [PubMed]
- Nautiyal, C.S.; Bhadauria, S.; Kumar, P.; Lal, H.; Mondal, R.; Verma, D. Stress induced phosphate solubilization inbacteria isolated from alkaline soils. FEMS Microbiol. Lett. 2000, 182, 291–296. [Google Scholar] [CrossRef] [PubMed]
- Jean, N.; Boge, G.; Jamet, J.L.; Richard, S.; Jamet, D. Seasonal changes in zooplanktonic alkaline phosphatase activity in Toulon Bay (France): the role of Cypris larvae. Mar. Pollut. Bull. 2003, 46, 346–352. [Google Scholar] [CrossRef]
- Chung, T.D.Y.; Sergienko, E.; Millán, J.L. Assay format as a critical success factor for identification of novel inhibitor chemotypes of tissue-nonspecific alkaline phosphatase from high-throughput screening. Molecules 2010, 15, 3010–3037. [Google Scholar] [CrossRef] [PubMed]
- Rossolini, G.M.; Shippa, S.; Riccio, M.L.; Berlutti, F.; Macaskie, L.E.; Thaller, M.C. Bacterial nonspecific acid phosphatases: physiology, evolution, and use as tools in microbial biotechnology. Cell. Mol. Life Sci. 1998, 54, 833–850. [Google Scholar] [CrossRef] [PubMed]
- Takano, Y.; Mori, H.; Kaneko, T.; Ishikawa, Y.; Marumo, K.; Kobayashi, K. Phosphatase and microbial activity with biochemical indicators in semi-permafrost active layer sediments over the past 10,000 years. Appl. Geochem. 2006, 21, 48–57. [Google Scholar] [CrossRef]
- Rodríguez, H.; Fraga, R. Phosphate solubilizing bacteria and their role in plant growth promotion. Biotechnol. Adv. 1999, 17, 319–339. [Google Scholar] [CrossRef]
- Thaller, M.C.; Berlutti, F.; Schippa, S.; Iori, P.; Passariello, C.; Rossolini, G.M. Heterogeneous patterns of acid phosphatases containing low-molecular-mass Polipeptides in members of the family Enterobacteriaceae. Int. J. Syst. Evol. Micr. 1995b, 4, 255–261. [Google Scholar]
- Lin, T.F.; Huang, H.I.; Shen, F.T.; Young, C.C. The protons of gluconic acid are the major factor responsible for the dissolution of tricalcium phosphate by Burkholderia cepacia CC-Al74. Bioresource Technol. 2006, 97, 957–960. [Google Scholar] [CrossRef] [PubMed]
- Tao, G.C.; Tian, S.J.; Cai, M.Y.; Xie, G.H. Phosphate-solubilizing and -mineralizing abilities of bacteria isolated from soils. Pedosphere 2008, 18, 515–523. [Google Scholar]
- Kim, K.Y.; Jordan, D.; McDonald, G.A. Enterobacter agglomerans, phosphate solubilizing bacteria, and microbial activity in soil: effect of carbon sources. Soil Biol. Biochem. 1998, 30, 995–1003. [Google Scholar] [CrossRef]
- Chen, W.Y.; Zheng, H.L. Current situation and countermeasures of water pollution of Xixi Wetland. Hangzhou Agr. Sci. Technol. 2008, 4, 25–26. (in Chinese). [Google Scholar]
- Holt, J.G.; Krieg, N.R.; Sneath, P.H.A.; Staley, J.T.; Williams, S.T. Bergey’s Manual of Determinative Bacteriology, 9th ed; Williams & Wilkins: Baltimore, MD, USA, 1994. [Google Scholar]
- Liu, W.; Lu, H.H.; Wu, W.X.; Wei, Q.K.; Chen, Y.X.; Thies, J.E. Transgenic Bt rice does not affect enzyme activities and microbial composition in the rhizosphere during crop development. Soil Biol. Biochem. 2008, 40, 475–486. [Google Scholar] [CrossRef]
| Strain | Number of nucleotides compared | Closest matcha and its accession code | Identity | Accession number |
|---|---|---|---|---|
| OPB32 | 730 | Citrobacter freundii (AB244300) | 99% | GQ465231 |
| OPB48 | 740 | Pseudomonas mosselii (EU301780) | 100% | GQ465232 |
| OPB49 | 1210 | Cupriavidus basilensis (AM048887) | 96% | GQ465222 |
| OPB51 | 720 | Cupriavidus basilensis (AY860244) | 98% | GQ465223 |
| OPB57 | 780 | Citrobacter freundii (AB184249) | 100% | GQ465233 |
| OPB59 | 700 | Citrobacter freundii (FJ494899) | 100% | GQ465224 |
| OPB72 | 1060 | Acinetobacter sp. (EF103565) | 99% | GQ465225 |
| OPB98 | 770 | Burkholderia pyrrocinia (U96930) Burkholderia cepacia (EF095217) Burkholderia ambifaria (CP000440) | 99% | GQ465226 |
| IPB1 | 1150 | Enterobacter sp. (GQ280118) | 99% | GQ465227 |
| IPB12 | 780 | Enterobacter sp. (EU693568) Pantoea agglomerans (EF489443) | 99% | GQ465228 |
| IPB15 | 680 | Enterobacter sp. (EU693568) Pantoea agglomerans (EF489443) | 99% | GQ465229 |
| IPB54 | 680 | Pantoea agglomerans (EU721614) | 100% | GQ465230 |
| Cupriavidus basilensis | OPB49 | OPB51 | |
|---|---|---|---|
| Gram | - | - | - |
| Flagella | + | + | + |
| Catalase | + | + | + |
| Oxidase | + | + | + |
| Nitrate Reduction | / | + | - |
| Indole test | - | - | - |
| Adipate | + | + | + |
| Caprate | + | + | + |
| Citrate | + | + | + |
| Glucose | - | - | - |
| Malate | + | + | + |
| Maltose | - | - | - |
| Mannitol | - | - | - |
| Mannose | - | - | - |
| Phenylacetate | + | + | + |
| Mobility | + | + | + |
© 2010 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Qian, Y.; Shi, J.; Chen, Y.; Lou, L.; Cui, X.; Cao, R.; Li, P.; Tang, J. Characterization of Phosphate Solubilizing Bacteria in Sediments from a Shallow Eutrophic Lake and a Wetland: Isolation, Molecular Identification and Phosphorus Release Ability Determination. Molecules 2010, 15, 8518-8533. https://doi.org/10.3390/molecules15118518
Qian Y, Shi J, Chen Y, Lou L, Cui X, Cao R, Li P, Tang J. Characterization of Phosphate Solubilizing Bacteria in Sediments from a Shallow Eutrophic Lake and a Wetland: Isolation, Molecular Identification and Phosphorus Release Ability Determination. Molecules. 2010; 15(11):8518-8533. https://doi.org/10.3390/molecules15118518
Chicago/Turabian StyleQian, Yichao, Jiyan Shi, Yingxu Chen, Liping Lou, Xinyi Cui, Rukun Cao, Pengfei Li, and Jie Tang. 2010. "Characterization of Phosphate Solubilizing Bacteria in Sediments from a Shallow Eutrophic Lake and a Wetland: Isolation, Molecular Identification and Phosphorus Release Ability Determination" Molecules 15, no. 11: 8518-8533. https://doi.org/10.3390/molecules15118518
APA StyleQian, Y., Shi, J., Chen, Y., Lou, L., Cui, X., Cao, R., Li, P., & Tang, J. (2010). Characterization of Phosphate Solubilizing Bacteria in Sediments from a Shallow Eutrophic Lake and a Wetland: Isolation, Molecular Identification and Phosphorus Release Ability Determination. Molecules, 15(11), 8518-8533. https://doi.org/10.3390/molecules15118518




