A Novel Method for Colorectal Cancer Screening Based on Circulating Tumor Cells and Machine Learning
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Sample Preparation
2.3. Antibodies and Staining Procedure
2.4. Sample Blinding
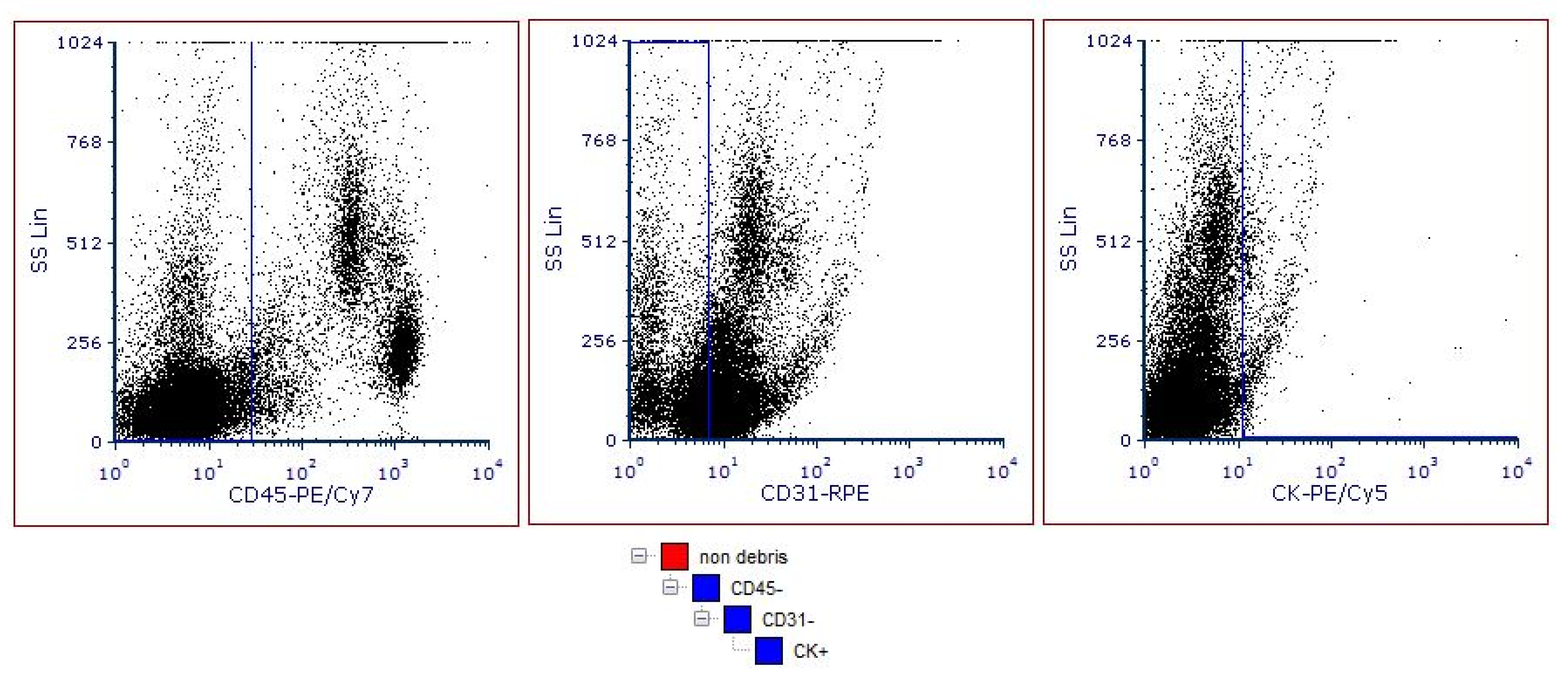
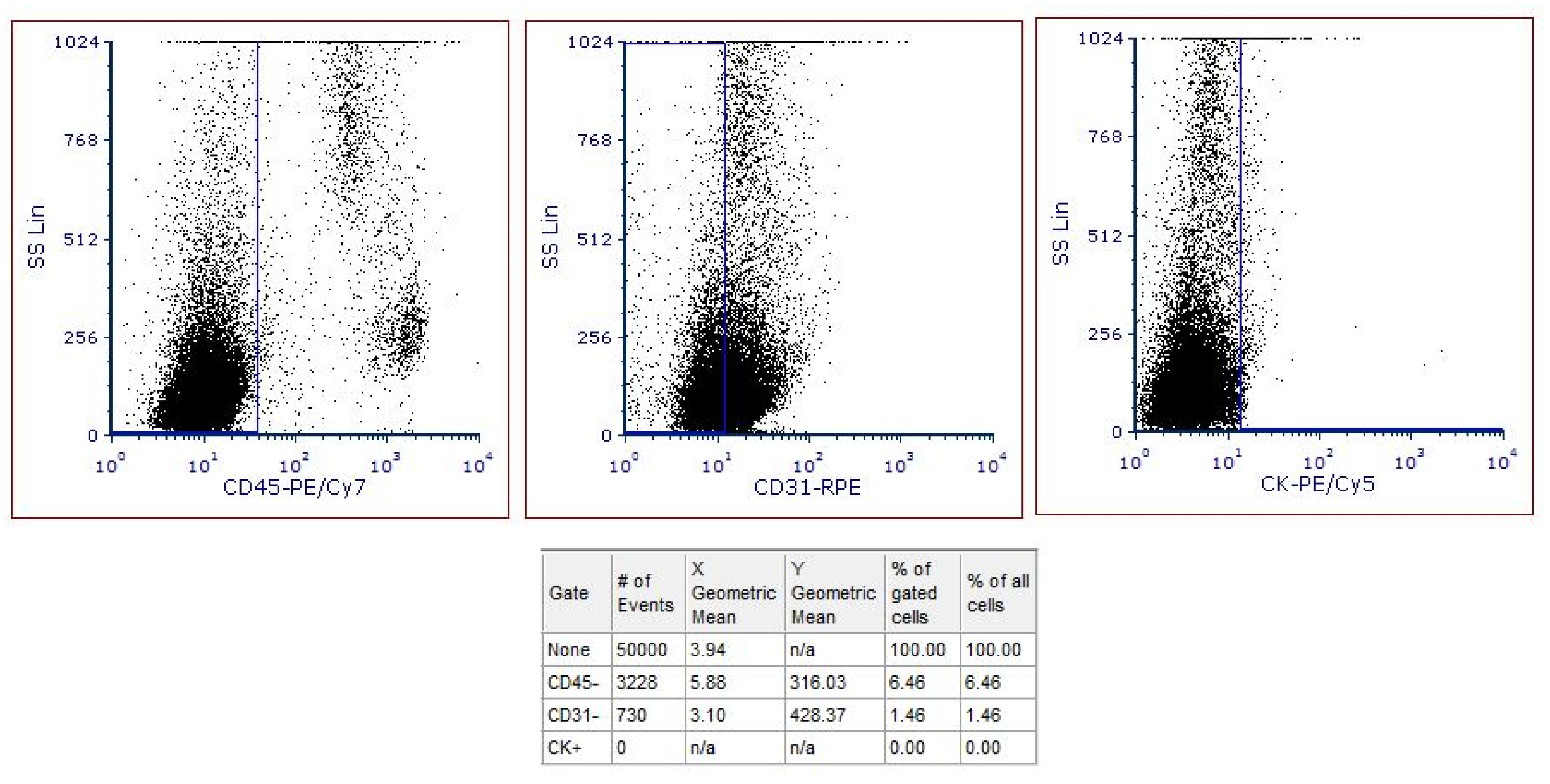
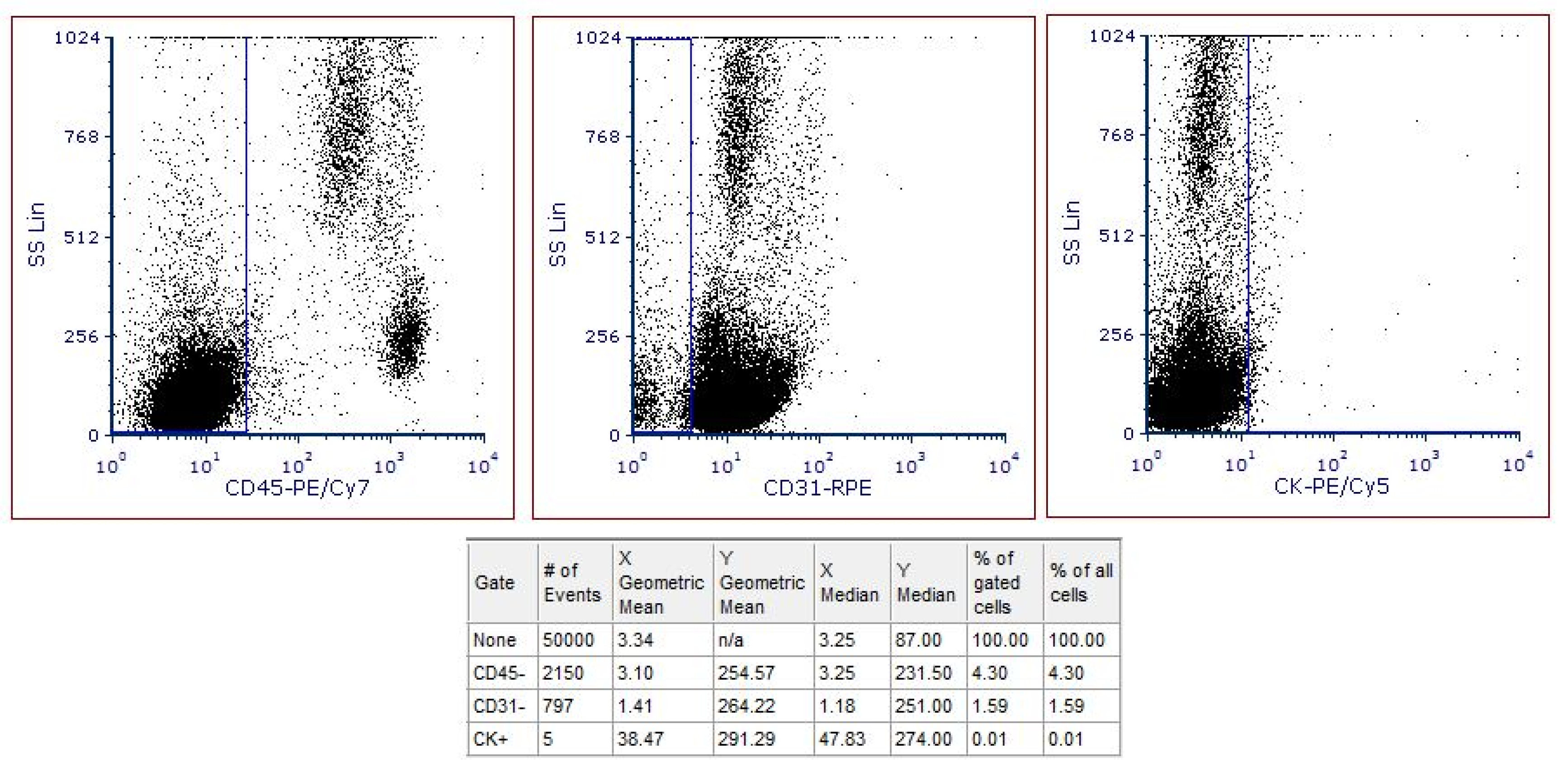
2.5. Sample Acquisition and FCS Data Analysis
2.6. Mathematical Analysis
2.6.1. Two-Sample Kolmogorov–Smirnov Test
2.6.2. Wilcoxon Rank Sum Test
2.6.3. Support Vector Machines
2.6.4. Comparison between Different Classifiers
2.6.5. Performance Measures for Binary Classifiers
3. Results
3.1. Statistical Tests
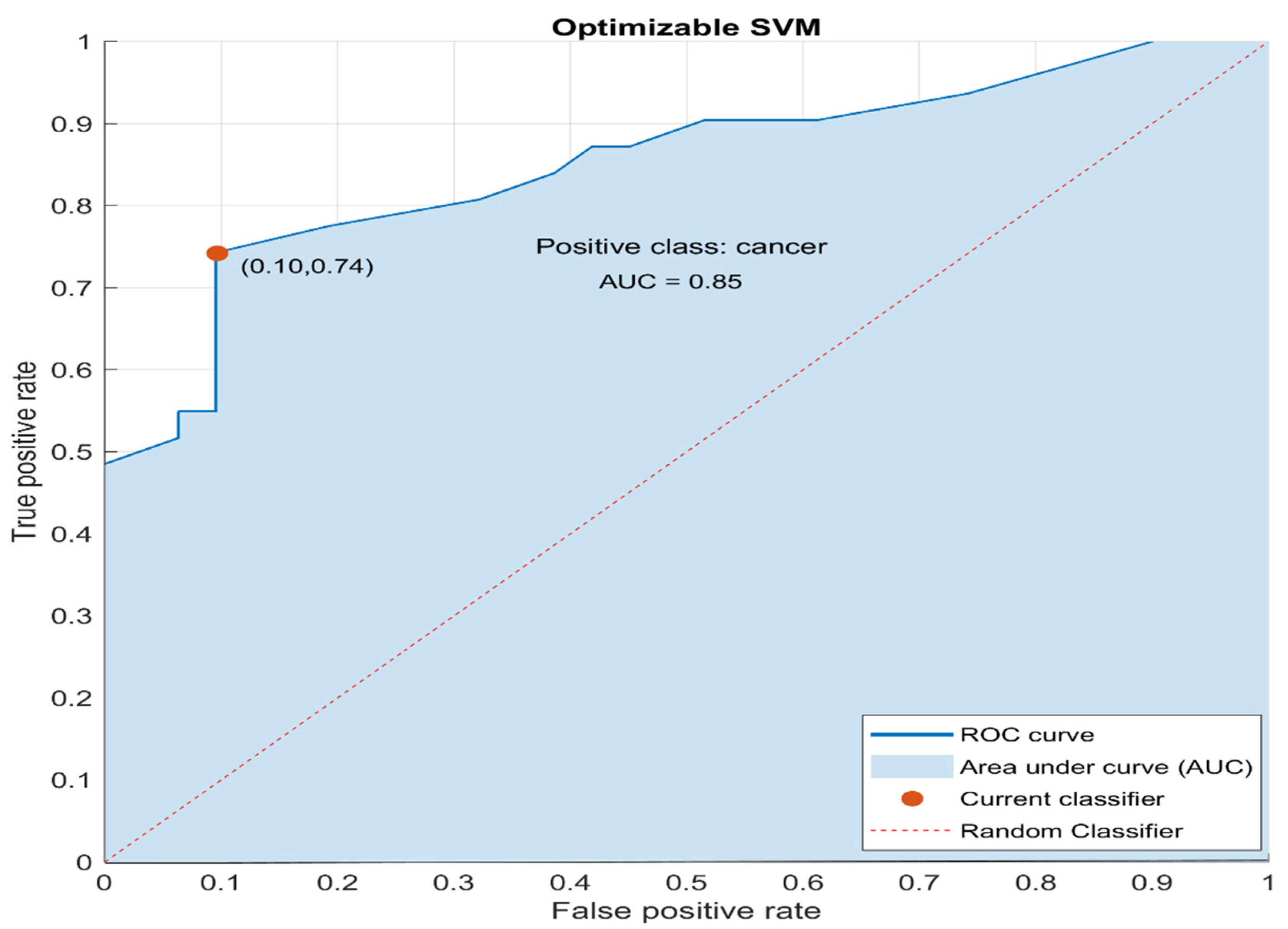
3.2. SVM Classifier
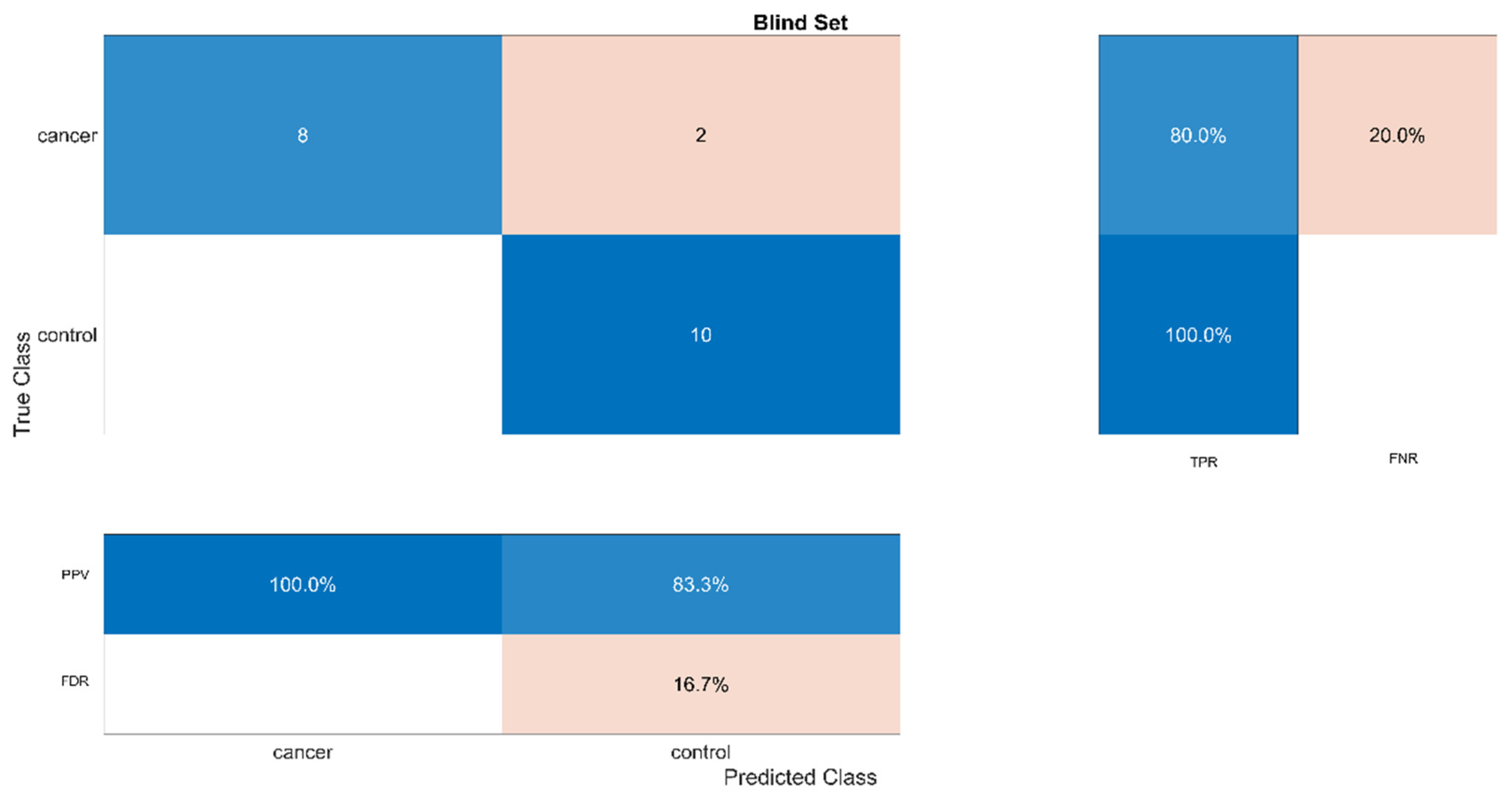
3.3. Blind Set
3.4. Comparison between Different Classifiers
4. Discussion
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Hanahan, D.; Weinberg, R.A. Hallmarks of Cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Butcher, E.C.; Berg, E.; Kunkel, E.J. Systems biology in drug discovery. Nat. Biotechnol. 2004, 22, 1253–1259. [Google Scholar] [CrossRef]
- Hornberg, J.J.; Bruggeman, F.; Westerhoff, H.V.; Lankelma, J. Cancer: A Systems Biology disease. Biosystems 2006, 83, 81–90. [Google Scholar] [CrossRef]
- Grizzi, F.; Chiriva-Internati, M. Cancer: Looking for simplicity and finding complexity. Cancer Cell Int. 2006, 6, 4. [Google Scholar] [CrossRef] [PubMed]
- Moore, N.M.; Kuhn, N.Z.; Hanlon, S.E.; Lee, J.S.H.; Nagahara, L.A. De-convoluting cancer’s complexity: Using a ‘physical sciences lens’ to provide a different (clearer) perspective of cancer. Phys. Biol. 2011, 8, 010302. [Google Scholar] [CrossRef] [PubMed]
- Bray, F.; Me, J.F.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Dekker, E.; Tanis, P.J.; Vleugels, J.L.A.; Kasi, P.M.; Wallace, M.B. Colorectal cancer. Lancet 2019, 394, 1467–1480. [Google Scholar] [CrossRef]
- Henrikson, N.B.; Webber, E.M.; Goddard, K.A.; Scrol, A.; Piper, M.; Williams, M.S.; Zallen, D.T.; Calonge, N.; Ganiats, T.G.; Msc, A.C.J.J.; et al. Family history and the natural history of colorectal cancer: Systematic review. Genet. Med. 2015, 17, 702–712. [Google Scholar] [CrossRef]
- Qaseem, A.; Crandall, C.J.; Mustafa, R.A.; Hicks, L.A.; Wilt, T.J. Clinical Guidelines Committee of the American College of Physicians. Screening for Colorectal Cancer in Asymptomatic Average-Risk Adults: A Guidance Statement from the American College of Physicians. Ann. Intern. Med. 2019, 171, 643–654. [Google Scholar] [CrossRef]
- Gentles, A.J.; Gallahan, D. Systems Biology: Confronting the Complexity of Cancer. Cancer Res. 2011, 71, 5961–5964. [Google Scholar] [CrossRef]
- Biemar, F.; Foti, M. Global progress against cancer—Challenges and opportunities. Cancer Biol. Med. 2013, 10, 183–186. [Google Scholar]
- Cagan, R.; Meyer, P. Rethinking cancer: Current challenges and opportunities in cancer research. Dis. Model. Mech. 2017, 10, 349–352. [Google Scholar] [CrossRef]
- Iliopoulos, A.; Beis, G.; Apostolou, P.; Papasotiriou, I. Complex Networks, Gene Expression and Cancer Complexity: A Brief Review of Methodology and Applications. Curr. Bioinform. 2020, 15, 629–655. [Google Scholar] [CrossRef]
- Karakatsanis, L.P.; Pavlos, E.G.; Tsoulouhas, G.; Stamokostas, G.L.; Mosbruger, T.; Duke, J.L.; Pavlos, G.P.; Monos, D.S. Spatial constrains and information content of sub-genomic regions of the human genome. iScience 2021, 24, 102048. [Google Scholar] [CrossRef]
- Cruz, J.A.; Wishart, D.S. Applications of Machine Learning in Cancer Prediction and Prognosis. Cancer Inform. 2006, 2, 59–77. [Google Scholar] [CrossRef]
- Kourou, K.; Exarchos, T.P.; Exarchos, K.P.; Karamouzis, M.V.; Fotiadis, D.I. Machine learning applications in cancer prognosis and prediction. Comput. Struct. Biotechnol. J. 2015, 13, 8–17. [Google Scholar] [CrossRef] [PubMed]
- Munir, K.; Elahi, H.; Ayub, A.; Frezza, F.; Rizzi, A. Cancer Diagnosis Using Deep Learning: A Bibliographic Review. Cancers 2019, 11, 1235. [Google Scholar] [CrossRef] [PubMed]
- Apostolou, P.; Iliopoulos, A.C.; Parsonidis, P.; Papasotiriou, I. Gene expression profiling as a potential predictor between normal and cancer samples in gastrointestinal carcinoma. Oncotarget 2019, 10, 3328–3338. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Iqbal, M.J.; Javed, Z.; Sadia, H.; Qureshi, I.A.; Irshad, A.; Ahmed, R.; Malik, K.; Raza, S.; Abbas, A.; Pezzani, R.; et al. Clinical applications of artificial intelligence and machine learning in cancer diagnosis: Looking into the future. Cancer Cell Int. 2021, 21, 1–11. [Google Scholar] [CrossRef]
- Menden, M.P.; Iorio, F.; Garnett, M.; McDermott, U.; Benes, C.H.; Ballester, P.J.; Saez-Rodriguez, J. Machine Learning Prediction of Cancer Cell Sensitivity to Drugs Based on Genomic and Chemical Properties. PLoS ONE 2013, 8, e61318. [Google Scholar] [CrossRef]
- Bashiri, A.; Ghazisaeedi, M.; Safdari, R.; Shahmoradi, L.; Ehtesham, H. Improving the Prediction of Survival in Cancer Patients by Using Machine Learning Techniques: Experience of Gene Expression Data: A Narrative Review. Iran. J. Public Health 2017, 46, 165–172. [Google Scholar]
- De Silva, D.; Ranasinghe, W.; Bandaragoda, T.; Adikari, A.; Mills, N.; Iddamalgoda, L.; Alahakoon, D.; Lawrentschuk, N.; Persad, R.; Osipov, E.; et al. Machine learning to support social media empowered patients in cancer care and cancer treatment decisions. PLoS ONE 2018, 13, e0205855. [Google Scholar] [CrossRef]
- Levine, A.B.; Schlosser, C.; Grewal, J.; Coope, R.; Jones, S.; Yip, S. Rise of the Machines: Advances in Deep Learning for Cancer Diagnosis. Trends Cancer 2019, 5, 157–169. [Google Scholar] [CrossRef] [PubMed]
- Ronen, J.; Hayat, S.; Akalin, A. Evaluation of colorectal cancer subtypes and cell lines using deep learning. Life Sci. Alliance 2019, 2, e201900517. [Google Scholar] [CrossRef] [PubMed]
- Nartowt, B.J.; Hart, G.R.; Roffman, D.A.; Llor, X.; Ali, I.; Muhammad, W.; Liang, Y.; Deng, J. Scoring colorectal cancer risk with an artificial neural network based on self-reportable personal health data. PLoS ONE 2019, 14, e0221421. [Google Scholar] [CrossRef] [PubMed]
- Nartowt, B.J.; Hart, G.R.; Muhammad, W.; Liang, Y.; Stark, G.F.; Deng, J. Robust Machine Learning for Colorectal Cancer Risk Prediction and Stratification. Front. Big Data 2020, 3, 6. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.S.; Yu, G.; Xu, C.; Meng, X.H.; Zhou, J.; Zheng, C.; Deng, Z.; Shang, L.; Liu, R.; Su, S.; et al. Accurate diagnosis of colorectal cancer based on histopathology images using artificial intelligence. BMC Med. 2021, 19, 76. [Google Scholar] [CrossRef]
- Mitsala, A.; Tsalikidis, C.; Pitiakoudis, M.; Simopoulos, C.; Tsaroucha, A. Artificial Intelligence in Colorectal Cancer Screening, Diagnosis and Treatment. A New Era. Curr. Oncol. 2021, 28, 1581–1607. [Google Scholar] [CrossRef] [PubMed]
- Chu, F.; Wang, L. Applications of support vector machines to cancer classification with microarray data. Int. J. Neural Syst. 2005, 15, 475–484. [Google Scholar] [CrossRef]
- Zhang, B.; Liang, X.; Gao, H.; Ye, L.; Wang, Y. Models of logistic regression analysis, support vector machine, and back-propagation neural network based on serum tumor markers in colorectal cancer diagnosis. Genet. Mol. Res. 2016, 15. [Google Scholar] [CrossRef]
- Aziz, M.; Hussein, M.A.; Gabere, M.N. Filtered selection coupled with support vector machines generate a functionally relevant prediction model for colorectal cancer. OncoTargets Ther. 2016, 9, 3313–3325. [Google Scholar] [CrossRef] [PubMed]
- Gao, L.; Ye, M.; Wu, C. Cancer Classification Based on Support Vector Machine Optimized by Particle Swarm Optimization and Artificial Bee Colony. Molecules 2017, 22, 2086. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Cai, N.; Pacheco, P.P.; Narandes, S.; Wang, Y.; Xu, W. Applications of Support Vector Machine (SVM) Learning in Cancer Genomics. Cancer Genom. Proteom. 2018, 15, 41–51. [Google Scholar] [CrossRef]
- Chawla, N.V.; Bowyer, K.; Hall, L.O.; Kegelmeyer, W.P. SMOTE: Synthetic Minority Over-sampling Technique. J. Artif. Intell. Res. 2002, 16, 321–357. [Google Scholar] [CrossRef]
- Wang, W.-C.; Zhang, X.-F.; Peng, J.; Li, X.-F.; Wang, A.-L.; Bie, Y.-Q.; Shi, L.-H.; Lin, M.-B. Survival Mechanisms and Influence Factors of Circulating Tumor Cells. BioMed Res. Int. 2018, 2018, 6304701. [Google Scholar] [CrossRef]
- Veyrune, L.; Naumann, D.; Christou, N. Circulating Tumour Cells as Prognostic Biomarkers in Colorectal Cancer: A Systematic Review. Int. J. Mol. Sci. 2021, 22, 3437. [Google Scholar] [CrossRef] [PubMed]
- Ribatti, D.; Tamma, R.; Annese, T. Epithelial-Mesenchymal Transition in Cancer: A Historical Overview. Transl. Oncol. 2020, 13, 100773. [Google Scholar] [CrossRef]
- Cabel, L.; Proudhon, C.; Gortais, H.; Loirat, D.; Coussy, F.; Pierga, J.-Y.; Bidard, F.-C. Circulating tumor cells: Clinical validity and utility. Int. J. Clin. Oncol. 2017, 22, 421–430. [Google Scholar] [CrossRef]
- Gorges, T.M.; Tinhofer, I.; Drosch, M.; Röse, L.; Zollner, T.M.; Krahn, T.; von Ahsen, O. Circulating tumour cells escape from EpCAM-based detection due to epithelial-to-mesenchymal transition. BMC Cancer 2012, 16, 178. [Google Scholar] [CrossRef]
- Agarwal, A.; Balic, M.; El-Ashry, D.; Cote, R.J. Circulating Tumor Cells: Strategies for Capture, Analyses, and Propagation. Cancer J. 2018, 24, 70–77. [Google Scholar] [CrossRef]
- Papasotiriou, I.; Chatziioannou, M.; Pessiou, K.; Retsas, I.; Dafouli, G.; Kyriazopoulou, A.; Toloudi, M.; Kaliara, I.; Vlachou, I.; Kourtidou, E.; et al. Detection of Circulating Tumor Cells in Patients with Breast, Prostate, Pancreatic, Colon and Melanoma Cancer: A Blinded Comparative Study Using Healthy Donors. J. Cancer Ther. 2015, 6, 543–553. [Google Scholar] [CrossRef][Green Version]
- Marsaglia, G.; Tsang, W.W.; Wang, J. Evaluating Kolmogorov’s Distribution. J. Stat. Softw. 2003, 8, 1–4. [Google Scholar] [CrossRef]
- Whitley, E.; Ball, J. Statistics review 6: Nonparametric methods. Crit. Care 2002, 6, 509–513. [Google Scholar] [CrossRef]
- Vapnik, V. Pattern recognition using generalized portrait method. Autom. Remote Control 1963, 24, 774–780. [Google Scholar]
- Noble, W.S. What is a support vector machine? Nat. Biotechnol. 2006, 24, 1565–1567. [Google Scholar] [CrossRef]
- Mitchell, T. Machine Learning; McGraw Hill: New York, NY, USA, 1997. [Google Scholar]
- Krzywinski, M.; Altman, N. Classification and regression trees. Nat. Meth. 2017, 14, 757–758. [Google Scholar] [CrossRef]
- Hardle, W.; Simar, L. Applied Multivariate Statistical Analysis; Springer: Berlin, Germany, 2015. [Google Scholar]
- LaValley, M.P. Logistic Regression. Circulation 2008, 117, 2395–2399. [Google Scholar] [CrossRef]
- Hastie, T.; Tibshirani, R.; Friendman, J. The Elements of Statistical Learning—Data Mining, Inference, and Prediction; Springer: Berlin, Germany, 2013. [Google Scholar]
- Abu Alfeilat, H.A.; Hassanat, A.; Lasassmeh, O.; Altarawneh, A.S.A.; Alhasanat, M.B.; Salman, H.S.E.; Prasath, S. Effects of Distance Measure Choice on K-Nearest Neighbor Classifier Performance: A Review. Big Data 2019, 7, 221–248. [Google Scholar] [CrossRef] [PubMed]
- Opitz, D.; Maclin, R. Popular Ensemble Methods: An empirical Study. J. Artif. Intell. Res. 1999, 11, 169–198. [Google Scholar] [CrossRef]
- Yang, P.; Yang, Y.H.; Zhou, B.B.; Zomaya, A.Y. A Review of Ensemble Methods in Bioinformatics. Curr. Bioinform. 2010, 5, 296–308. [Google Scholar] [CrossRef]
- Blagus, R.; Lusa, L. SMOTE for high-dimensional class-imbalanced data. BMC Bioinform. 2013, 14, 106. [Google Scholar] [CrossRef] [PubMed]
- Fernández, A.; García, S.; Herrera, F.; Chawla, N.V. SMOTE for Learning from Imbalanced Data: Progress and Challenges, Marking the 15-year Anniversary. J. Artif. Intell. Res. 2018, 61, 863–905. [Google Scholar] [CrossRef]
- Sokolova, M.; Lapalme, G. A systematic analysis of performance measures for classification tasks. Inf. Process. Manag. 2009, 45, 427–437. [Google Scholar] [CrossRef]
- Streiner, D.L.; Cairney, J. What’s Under the ROC? An Introduction to Receiver Operating Characteristics Curves. Can. J. Psychiatry 2007, 52, 121–128. [Google Scholar] [CrossRef] [PubMed]
- Fawcett, T. An introduction to ROC analysis. Pattern Recognit. Lett. 2006, 27, 861–874. [Google Scholar] [CrossRef]
- MATLAB. Statistics and Machine Learning Toolbox; The MathWorks, Inc.: Natick, MA, USA, 2021. [Google Scholar]
- Larsen, B.S. Synthetic Minority Over-Sampling Technique (SMOTE). 2021. Available online: https://github.com/dkbsl/matlab_smote/releases/tag/1.0 (accessed on 1 September 2021).
- Friedman, J.; Hastie, T.; Tibshirani, R. Additive logistic regression: A statistical view of boosting. Ann. Stat. 2000, 28, 337–407. [Google Scholar] [CrossRef]
- Breiman, L. Random Forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]

| D1 | D2 | D3 | D4 | D5 | D6 | |
|---|---|---|---|---|---|---|
| N = 1 | N = 1 | N = 3 | N = 3 | N = 10 | N = 10 | |
| K = 5 | K = 10 | K = 10 | K = 20 | K = 20 | K = 30 | |
| Trees | 86.0 | 88.4 | 88.4 | 85.1 | 87.6 | 88.1 |
| Discriminant | 84.1 | 87.2 | 85.7 | 86.0 | 86.7 | 88.0 |
| Logistic Regression | 84.1 | 86.0 | 86.0 | 86.6 | 87.1 | 87.8 |
| Naïve Bayes | 84.1 | 86.6 | 85.7 | 86.0 | 86.9 | 88.1 |
| SVM | 86.0 | 89.0 | 89.6 | 87.2 | 87.3 | 88.0 |
| KNN | 85.4 | 87.8 | 89.6 | 85.7 | 84.4 | 87.9 |
| Ensemble | 86.0 | 88.4 | 88.4 | 87.2 | 86.9 | 88.2 |
| D1 | D2 | D3 | D4 | D5 | D6 | |
|---|---|---|---|---|---|---|
| N = 1 | N = 1 | N = 3 | N = 3 | N = 10 | N = 10 | |
| K = 5 | K = 10 | K = 10 | K = 20 | K = 20 | K = 30 | |
| Trees | 0.89 | 0.88 | 0.92 | 0.88 | 0.94 | 0.86 |
| Discriminant | 0.89 | 0.88 | 0.91 | 0.92 | 0.94 | 0.93 |
| Logistic Regression | 0.89 | 0.88 | 0.91 | 0.92 | 0.94 | 0.95 |
| Naïve Bayes | 0.88 | 0.88 | 0.89 | 0.92 | 0.94 | 0.94 |
| SVM | 0.84 | 0.89 | 0.88 | 0.89 | 0.94 | 0.95 |
| KNN | 0.89 | 0.88 | 0.91 | 0.92 | 0.94 | 0.92 |
| Ensemble | 0.89 | 0.89 | 0.92 | 0.92 | 0.94 | 0.94 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hatzidaki, E.; Iliopoulos, A.; Papasotiriou, I. A Novel Method for Colorectal Cancer Screening Based on Circulating Tumor Cells and Machine Learning. Entropy 2021, 23, 1248. https://doi.org/10.3390/e23101248
Hatzidaki E, Iliopoulos A, Papasotiriou I. A Novel Method for Colorectal Cancer Screening Based on Circulating Tumor Cells and Machine Learning. Entropy. 2021; 23(10):1248. https://doi.org/10.3390/e23101248
Chicago/Turabian StyleHatzidaki, Eleana, Aggelos Iliopoulos, and Ioannis Papasotiriou. 2021. "A Novel Method for Colorectal Cancer Screening Based on Circulating Tumor Cells and Machine Learning" Entropy 23, no. 10: 1248. https://doi.org/10.3390/e23101248
APA StyleHatzidaki, E., Iliopoulos, A., & Papasotiriou, I. (2021). A Novel Method for Colorectal Cancer Screening Based on Circulating Tumor Cells and Machine Learning. Entropy, 23(10), 1248. https://doi.org/10.3390/e23101248





