Classification of Covid-19 Coronavirus, Pneumonia and Healthy Lungs in CT Scans Using Q-Deformed Entropy and Deep Learning Features
Abstract
1. Introduction
2. Related Work
- (1)
- By achieving efficient classification results under limited computational resources with the use of fewer parameters on the collected 321 chest CT scans, we have shown that the proposed approach could effectively improve the performance of classifying lungs in CT scans.
- (2)
- The new proposed Q-deformed entropy features which are used as new texture extracted features for image classification tasks.
- (3)
- The proposed nine layers fully convolutional network architecture which is used to extract the deep features from lungs’ CT scans.
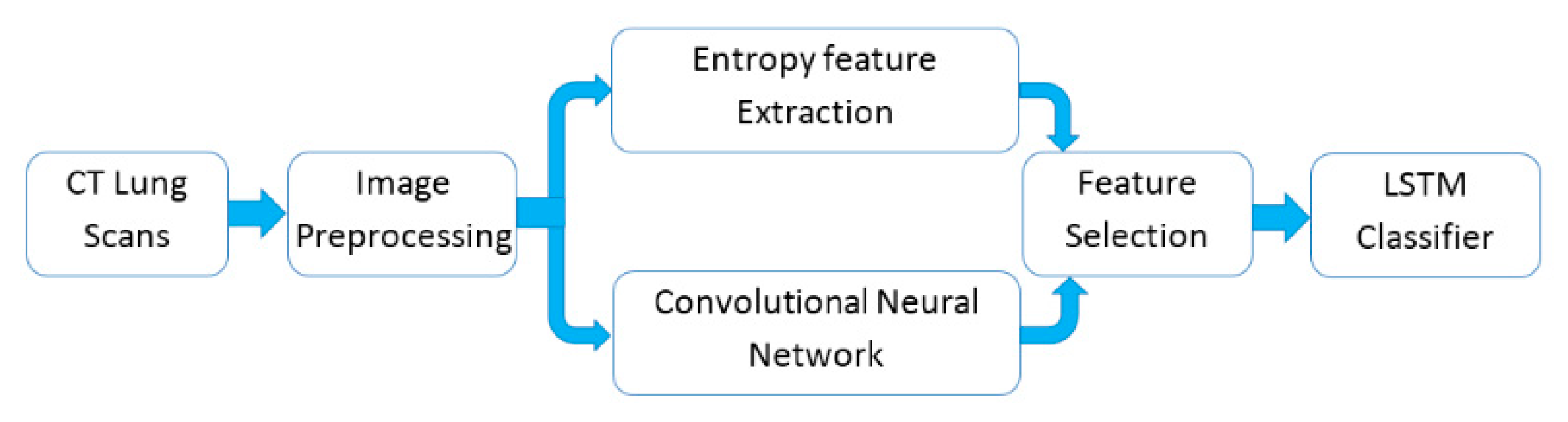
3. Materials and Methods
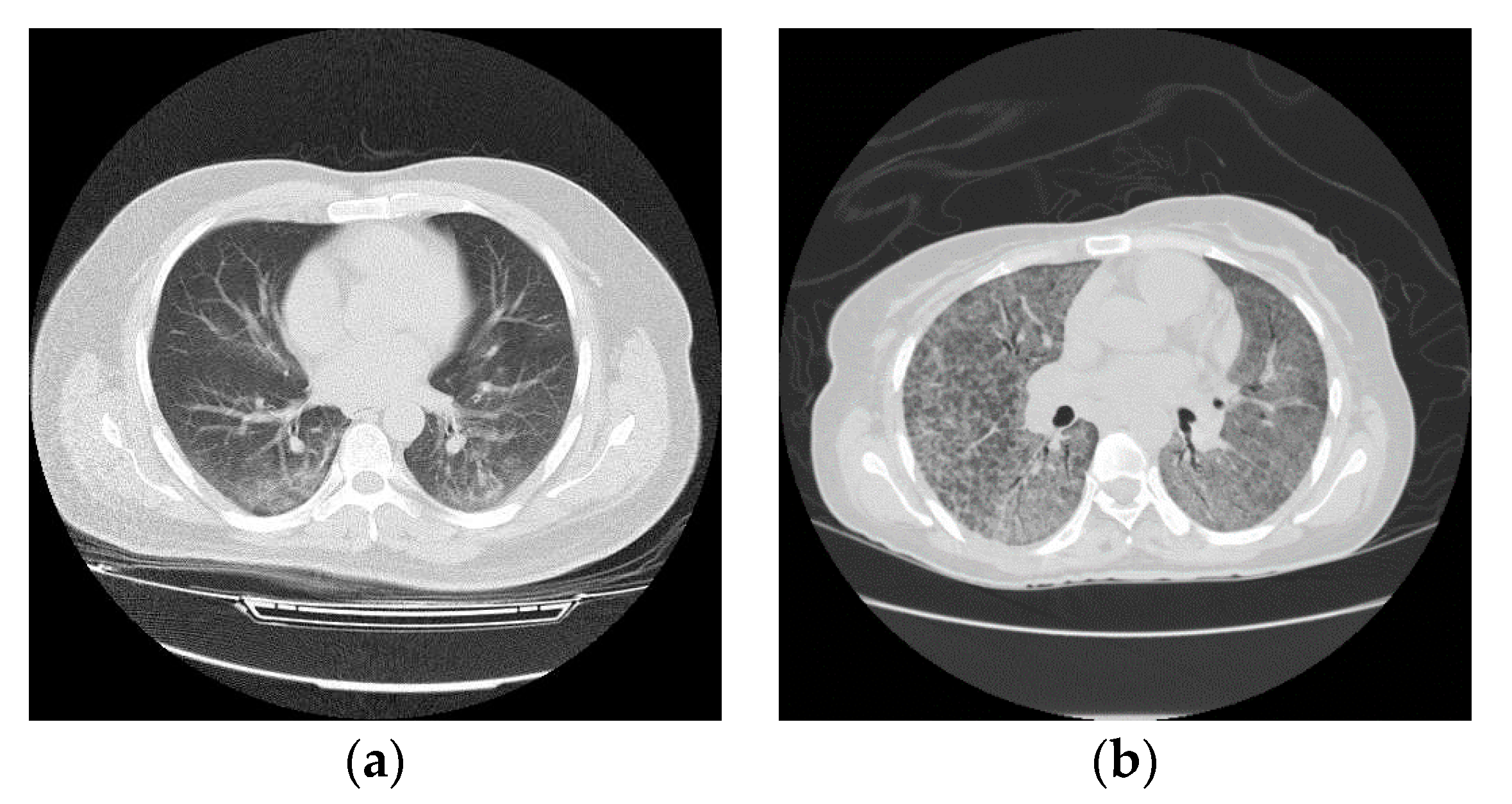
3.1. Data Collection
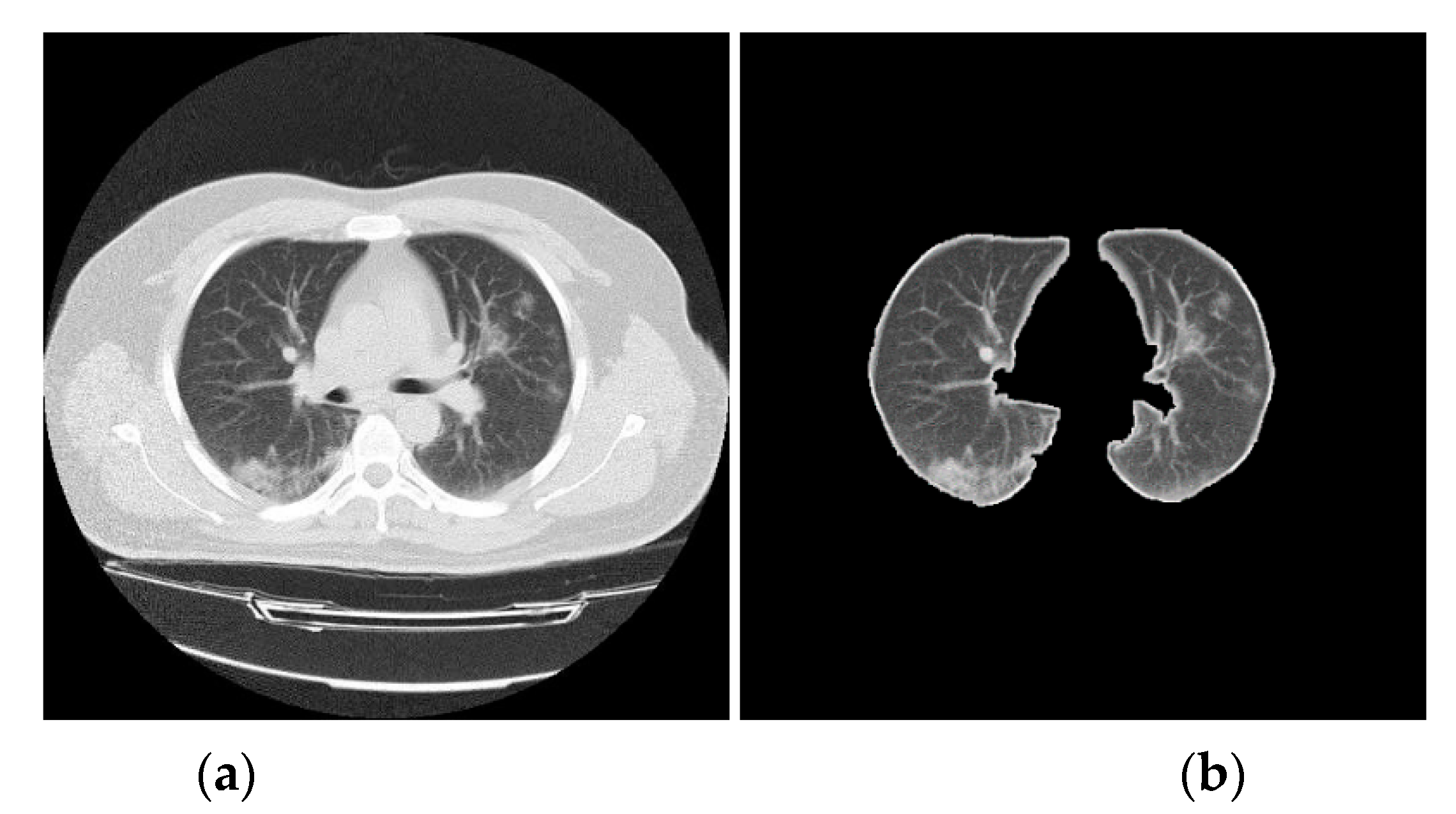
3.2. CT Lung Scan Preprocessing
| Algorithm 1: Pseudo-code for CT lung scans preprocessing. |
| Input: Input image I(n,m) |
| Output: Output image K(n,m) |
| begin |
| Adjust image intensity values |
| Convert the image into a binary image(B) |
| For all I pixels: |
| IF the grayscale value < the image Mean, |
| THEN, the pixel value = 0 |
| ELSE the pixel grayscale value = 255 |
| End IF |
| End For |
| Remove small objects from binary image, and Fill image regions and holes |
| Produce the output image(K) |
| For each Input image I do |
| For i = 1 to n do |
| For j = 1 to m do |
| Multiply each element in I(i,j) by the corresponding element |
| in B(i,j) and return the output image(K) |
| End For |
| End For |
| End For |
3.3. Q-Deformed Entropy Feature Extraction (QDE)
| Algorithm 2: Pseudo-code for the proposed Q-deformed entropy feature extraction (QDE) algorithm. |
| Initialization: I = Input image, 0 < q < 3 |
| For each Input image I do |
| (b1, b2, …, bn) divide I into n blocks of size m x m pixels |
| For i = 1 to n do |
| QDE in Equation (5), where i denotes the ith block of m x m |
| dimension |
| End For |
| QDE ← I = (1, 2, …n) // QDE Features of all (n) blocks |
| End For |
3.4. Deep Learning for Feature Extraction
- (filters of size 3 × 3, stride of 1, padding of 1, and kernels of 16) are applied:For the feature maps, we have 256 × 256 × 16 = 1,048,576 neurons.
- is equal to the previous feature maps divided by the stride number:For the feature maps, we have 128 × 128 × 16 = 262,144 neurons in the feature map of the first max pooling layer.
- (filters of size 5 × 5, a stride of 1, padding of 2 and kernels of 32) are applied:For the feature maps, there are 128 × 128 × 32 = 524,288 neurons.
- is equal to the previous feature maps divided by the stride number:For the feature maps, we have 64 × 64 × 32 = 131,072 neurons.
- (filters of size 5 × 5, a stride of 1, padding of 2 and kernels of 64) are applied:For the feature maps, we have 64 × 64 × 64 = 262,144 neurons in the feature map of the third convolution layer.
- is equal to the previous feature maps divided by the stride number:For the feature maps, we have 32 × 32 × 64 = 65,536 neurons.
- (convolutional filters of size 7 × 7, a stride of 1, padding of 3 and kernels of 128) are applied:For the feature maps, we have 32 × 32 × 128 = 131,072 neurons.
- The fully connected (FC) layer determines the class scores by combining all features which are produced and learned by the previous layers to produce a feature map of size 1 × 1 × 3, that is equal to the number of classes in the dataset. The input size of the FC layer is equal to 131,072 that is produced by .
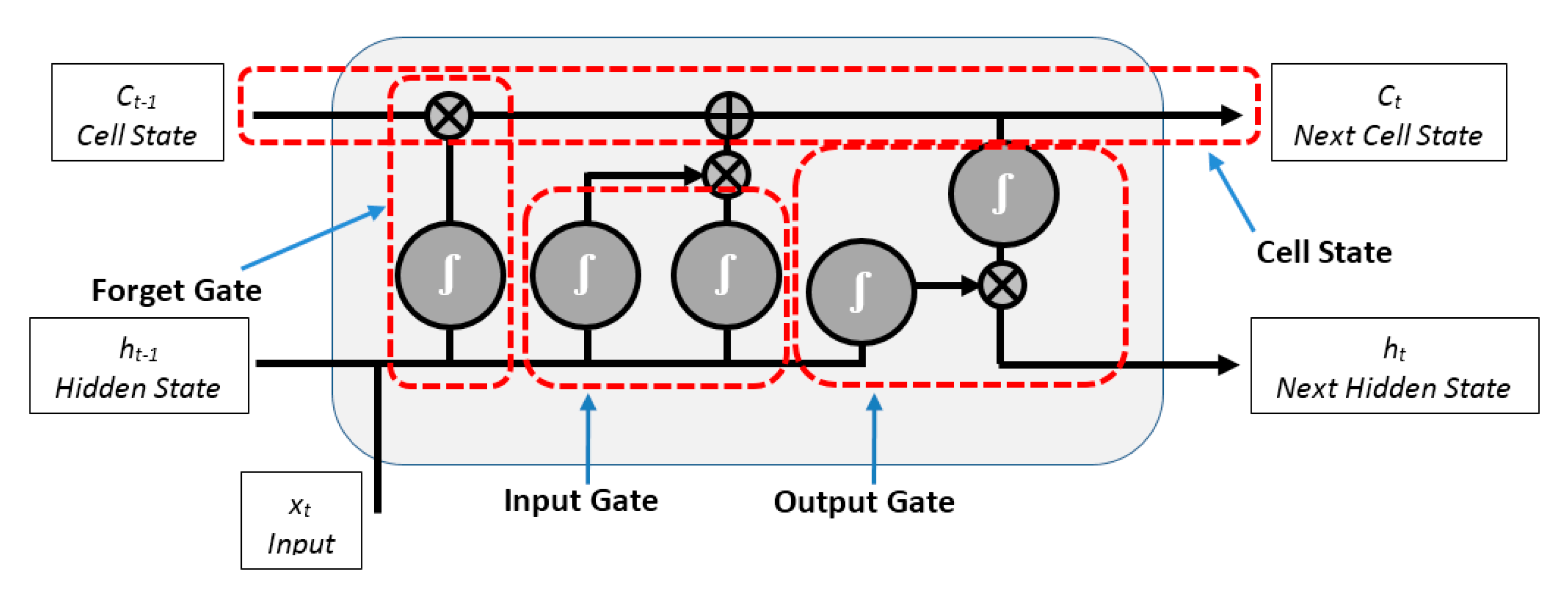
3.5. LSTM Neural Network Classifier
4. Experimental Results
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Fan, L.; Li, D.; Xue, H.; Zhang, L.; Liu, Z.; Zhang, B.; Zhang, L.; Yang, W.; Xie, B.; Duan, X.; et al. Progress and prospect on imaging diagnosis of COVID-19. Chin. J. Acad. Radiol. 2020, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Hu, Z.; Song, C.; Xu, C.; Jin, G.; Chen, Y.; Xu, X.; Ma, H.; Chen, W.; Lin, Y.; Zheng, Y.; et al. Clinical Characteristics of 24 Asymptomatic Infections with COVID-19 Screened among Close Contacts in Nanjing, China; Cold Spring Harbor Laboratory: Cold Spring Harbor, NY, USA, 2020; pp. 1–6. [Google Scholar]
- Hermanek, P.; Carmignato, S. Porosity measurements by X-ray computed tomography: Accuracy evaluation using a calibrated object. Precis. Eng. 2017, 49, 377–387. [Google Scholar] [CrossRef]
- Zonneveld, F.W.; Hanafee, W.N. Computed Tomography of the Temporal Bone and Orbit. J. Comput. Assist. Tomogr. 1988, 12, 540. [Google Scholar] [CrossRef]
- Samei, E. Computed Tomography: Approaches, Applications, and Operations; Springer Nature: Berlin/Heidelberg, Germany, 2020. [Google Scholar]
- Abdullayev, C.-P. COVID-19 Pneumonia. Available online: https://radiopaedia.org/cases/covid-19-pneumonia-45?lang=us (accessed on 17 April 2020).
- Depeursinge, A.; Al-Kadi, O.S.; Mitchell, J.R. Biomedical Texture Analysis: Fundamentals, Tools and Challenges; Academic Press: Cambridge, MA, USA, 2017. [Google Scholar]
- Zhang, J.; Xie, Y.; Li, Y.; Shen, C.; Xia, Y. COVID-19 Screening on Chest X-ray Images Using Deep Learning based Anomaly Detection. arXiv 2020, arXiv:2003.12338. [Google Scholar]
- Wang, L.; Wong, A. COVID-Net: A tailored deep convolutional neural network design for detection of COVID-19 cases from chest radiography images. arXiv 2020, arXiv:2003.09871. [Google Scholar]
- Li, L.; Qin, L.; Xu, Z.; Yin, Y.; Wang, X.; Kong, B.; Bai, J.; Lu, Y.; Fang, Z.; Song, Q. Artificial Intelligence Distinguishes COVID-19 from Community Acquired Pneumonia on Chest CT. Radiology 2020, 200905. [Google Scholar] [CrossRef]
- Wang, S.; Kang, B.; Ma, J.; Zeng, X.; Xiao, M.; Guo, J.; Cai, M.; Yang, J.; Li, Y.; Meng, X. A deep learning algorithm using CT images to screen for Corona Virus Disease (COVID-19). MedRxiv 2020. [Google Scholar] [CrossRef]
- Xu, X.; Jiang, X.; Ma, C.; Du, P.; Li, X.; Lv, S.; Yu, L.; Chen, Y.; Su, J.; Lang, G. Deep Learning System to Screen Coronavirus Disease 2019 Pneumonia. arXiv 2020, arXiv:2002.09334. [Google Scholar]
- Song, Y.; Zheng, S.; Li, L.; Zhang, X.; Zhang, X.; Huang, Z.; Chen, J.; Zhao, H.; Jie, Y.; Wang, R. Deep learning Enables Accurate Diagnosis of Novel Coronavirus (COVID-19) with CT images. MedRxiv 2020. [Google Scholar] [CrossRef]
- Yang, X.-J.; Gao, F.; Ju, Y. General Fractional Derivatives with Applications in Viscoelasticity; Academic Press: Cambridge, MA, USA, 2020. [Google Scholar]
- Yang, X.-J. General Fractional Derivatives: Theory, Methods and Applications; CRC Press: Boca Raton, FL, USA, 2019. [Google Scholar]
- Al-Shamasneh, A.R.; Jalab, H.A.; Shivakumara, P.; Ibrahim, R.W.; Obaidellah, U.H. Kidney segmentation in MR images using active contour model driven by fractional-based energy minimization. SignalImage Video Process. 2020, 1–8. [Google Scholar] [CrossRef]
- Radiopaedia. COVID-19 CT Cases. 2020. Available online: www.radiopaedia.org (accessed on 31 March 2020).
- Archive, C.I. SPIE-AAPM-NCI Lung Nodule Classification Challenge Dataset. 2019. Available online: www.cancerimagingarchive.net (accessed on 1 April 2020).
- Barrett, J.F.; Keat, N. Artifacts in CT: Recognition and avoidance. Radiographics 2004, 24, 1679–1691. [Google Scholar] [CrossRef] [PubMed]
- Tantisatirapong, S. Texture Analysis of Multimodal Magnetic Resonance Images in Support of Diagnostic Classification of Childhood Brain Tumours; University of Birmingham: Birmingham, UK, 2015. [Google Scholar]
- Nabizadeh, N.; Kubat, M. Brain tumors detection and segmentation in MR images: Gabor wavelet vs. statistical features. Comput. Electr. Eng. 2015, 45, 286–301. [Google Scholar] [CrossRef]
- Ibrahim, R.W.; Hasan, A.; Jalab, H.A. A new deformable model based on fractional Wright energy function for tumor segmentation of volumetric brain MRI scans. Comput. Methods Programs Biomed. 2018, 163, 21–28. [Google Scholar] [CrossRef] [PubMed]
- Mansoor, A.; Bagci, U.; Foster, B.; Xu, Z.; Papadakis, G.Z.; Folio, L.R.; Udupa, J.K.; Mollura, D.J. Segmentation and image analysis of abnormal lungs at CT: Current approaches, challenges, and future trends. RadioGraphics 2015, 35, 1056–1076. [Google Scholar] [CrossRef]
- Gonzalez, R.; Woods, R. Digital Image Processing, 3rd ed.; Prentice-Hall, Inc.: Upper Saddle River, NJ, USA, 2002. [Google Scholar]
- Hasan, A.M.; Jalab, H.A.; Meziane, F.; Kahtan, H.; Al-Ahmad, A.S. Combining deep and handcrafted image features for MRI brain scan classification. IEEE Access 2019, 7, 79959–79967. [Google Scholar] [CrossRef]
- Jalab, H.A.; Subramaniam, T.; Ibrahim, R.W.; Kahtan, H.; Noor, N.F.M. New Texture Descriptor Based on Modified Fractional Entropy for Digital Image Splicing Forgery Detection. Entropy 2019, 21, 371. [Google Scholar] [CrossRef]
- Al-Shamasneh, A.a.R.; Jalab, H.A.; Palaiahnakote, S.; Obaidellah, U.H.; Ibrahim, R.W.; El-Melegy, M.T. A new local fractional entropy-based model for kidney MRI image enhancement. Entropy 2018, 20, 344. [Google Scholar] [CrossRef]
- Umarov, S.; Tsallis, C.; Steinberg, S. On a q-central limit theorem consistent with nonextensive statistical mechanics. Milan J. Math. 2008, 76, 307–328. [Google Scholar] [CrossRef]
- Callen, H.B. Thermodynamics and an Introduction to Thermostatistics; American Association of Physics Teachers: College Park, MD, USA, 1998. [Google Scholar]
- Chang, P.; Grinband, J.; Weinberg, B.; Bardis, M.; Khy, M.; Cadena, G.; Su, M.-Y.; Cha, S.; Filippi, C.; Bota, D. Deep-learning convolutional neural networks accurately classify genetic mutations in gliomas. Am. J. Neuroradiol. 2018, 39, 1201–1207. [Google Scholar] [CrossRef]
- Gu, J.; Wang, Z.; Kuen, J.; Ma, L.; Shahroudy, A.; Shuai, B.; Liu, T.; Wang, X.; Wang, G.; Cai, J. Recent advances in convolutional neural networks. Pattern Recognit. 2018, 77, 354–377. [Google Scholar] [CrossRef]
- Kutlu, H.; Avcı, E. A Novel Method for Classifying Liver and Brain Tumors Using Convolutional Neural Networks, Discrete Wavelet Transform and Long Short-Term Memory Networks. Sensors 2019, 19, 1992. [Google Scholar] [CrossRef] [PubMed]
- Lundervold, A.S.; Lundervold, A. An overview of deep learning in medical imaging focusing on MRI. Zeitschrift für Medizinische Physik 2019, 29, 102–127. [Google Scholar] [CrossRef] [PubMed]
- Alom, M.Z.; Taha, T.M.; Yakopcic, C.; Westberg, S.; Sidike, P.; Nasrin, M.S.; Hasan, M.; Van Essen, B.C.; Awwal, A.A.; Asari, V.K. A State-of-the-Art Survey on Deep Learning Theory and Architectures. Electronics 2019, 8, 292. [Google Scholar] [CrossRef]
- Duan, M.; Li, K.; Yang, C.; Li, K. A hybrid deep learning CNN–ELM for age and gender classification. Neurocomputing 2018, 275, 448–461. [Google Scholar] [CrossRef]
- Dubitzky, W.; Granzow, M.; Berrar, D.P. Fundamentals of Data Mining in Genomics and Proteomics; Springer Science & Business Media: Berlin/Heidelberg, Germany, 2007. [Google Scholar]
- Le, X.-H.; Ho, H.V.; Lee, G.; Jung, S. Application of long short-term memory (LSTM) neural network for flood forecasting. Water 2019, 11, 1387. [Google Scholar] [CrossRef]
- Chen, G. A gentle tutorial of recurrent neural network with error backpropagation. arXiv 2016, arXiv:1610.02583. [Google Scholar]
- Tsiouris, Κ.Μ.; Pezoulas, V.C.; Zervakis, M.; Konitsiotis, S.; Koutsouris, D.D.; Fotiadis, D.I. A Long Short-Term Memory deep learning network for the prediction of epileptic seizures using EEG signals. Comput. Biol. Med. 2018, 99, 24–37. [Google Scholar] [CrossRef]
- Sak, H.; Senior, A.; Beaufays, F. Long Short-Term Memory Recurrent Neural Network Architectures for Large Scale Acoustic Modeling. In Proceedings of the Fifteenth Annual Conference of the International Speech Communication Association, Dresden, Germany, 6–10 September 2015. [Google Scholar]
- Babatunde, O.; Armstrong, L.; Leng, J.; Diepeveen, D. A genetic algorithm-based feature selection. Br. J. Math. Comput. Sci. 2014, 4, 889–905. [Google Scholar]




| Layer Name | Kernel Size | Feature Map |
|---|---|---|
| Input layer | (256 × 256) | |
| Conv1 | (3 × 3) | (256 × 256 × 16) |
| Max. Pooling1 | (2 × 2) | (128 × 128 × 16) |
| Conv2 | (5 × 5) | (128 × 128 × 32) |
| Max. Pooling2 | (2 × 2) | (64 × 64 × 32) |
| Conv3 | (5 × 5) | (64 × 64 × 64) |
| Max. Pooling3 | (2 × 2) | (32 × 32 × 64) |
| Conv4 | (7 × 7) | (32 × 32 × 128) |
| FC | (1 × 3) | (1 × 3) |
| Method | Accuracy 100% | TP 100% COVID-19 | TP 100% Healthy | TP 100% Pneumonia |
|---|---|---|---|---|
| QDE | 97.50 | 95.70 | 100 | 96.80 |
| DF | 98 | 97.40 | 100 | 96.80 |
| QDE–DF | 99.68 | 100 | 100 | 98.90 |
| Method | Accuracy 100% | TP 100% COVID-19 | TP 100% Healthy | TP 100% Pneumonia |
|---|---|---|---|---|
| Linear SVM | 96.20 | 94.90 | 98.10 | 95.80 |
| KNN | 95.30 | 93.20 | 97.20 | 95.80 |
| Logistic Regression | 97.20 | 96.60 | 98.10 | 96.80 |
| LSTM | 99.68 | 100 | 100 | 98.90 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hasan, A.M.; AL-Jawad, M.M.; Jalab, H.A.; Shaiba, H.; Ibrahim, R.W.; AL-Shamasneh, A.R. Classification of Covid-19 Coronavirus, Pneumonia and Healthy Lungs in CT Scans Using Q-Deformed Entropy and Deep Learning Features. Entropy 2020, 22, 517. https://doi.org/10.3390/e22050517
Hasan AM, AL-Jawad MM, Jalab HA, Shaiba H, Ibrahim RW, AL-Shamasneh AR. Classification of Covid-19 Coronavirus, Pneumonia and Healthy Lungs in CT Scans Using Q-Deformed Entropy and Deep Learning Features. Entropy. 2020; 22(5):517. https://doi.org/10.3390/e22050517
Chicago/Turabian StyleHasan, Ali M., Mohammed M. AL-Jawad, Hamid A. Jalab, Hadil Shaiba, Rabha W. Ibrahim, and Ala’a R. AL-Shamasneh. 2020. "Classification of Covid-19 Coronavirus, Pneumonia and Healthy Lungs in CT Scans Using Q-Deformed Entropy and Deep Learning Features" Entropy 22, no. 5: 517. https://doi.org/10.3390/e22050517
APA StyleHasan, A. M., AL-Jawad, M. M., Jalab, H. A., Shaiba, H., Ibrahim, R. W., & AL-Shamasneh, A. R. (2020). Classification of Covid-19 Coronavirus, Pneumonia and Healthy Lungs in CT Scans Using Q-Deformed Entropy and Deep Learning Features. Entropy, 22(5), 517. https://doi.org/10.3390/e22050517








