Structures and Dynamics of Dengue Virus Nonstructural Membrane Proteins
Abstract
1. Introduction
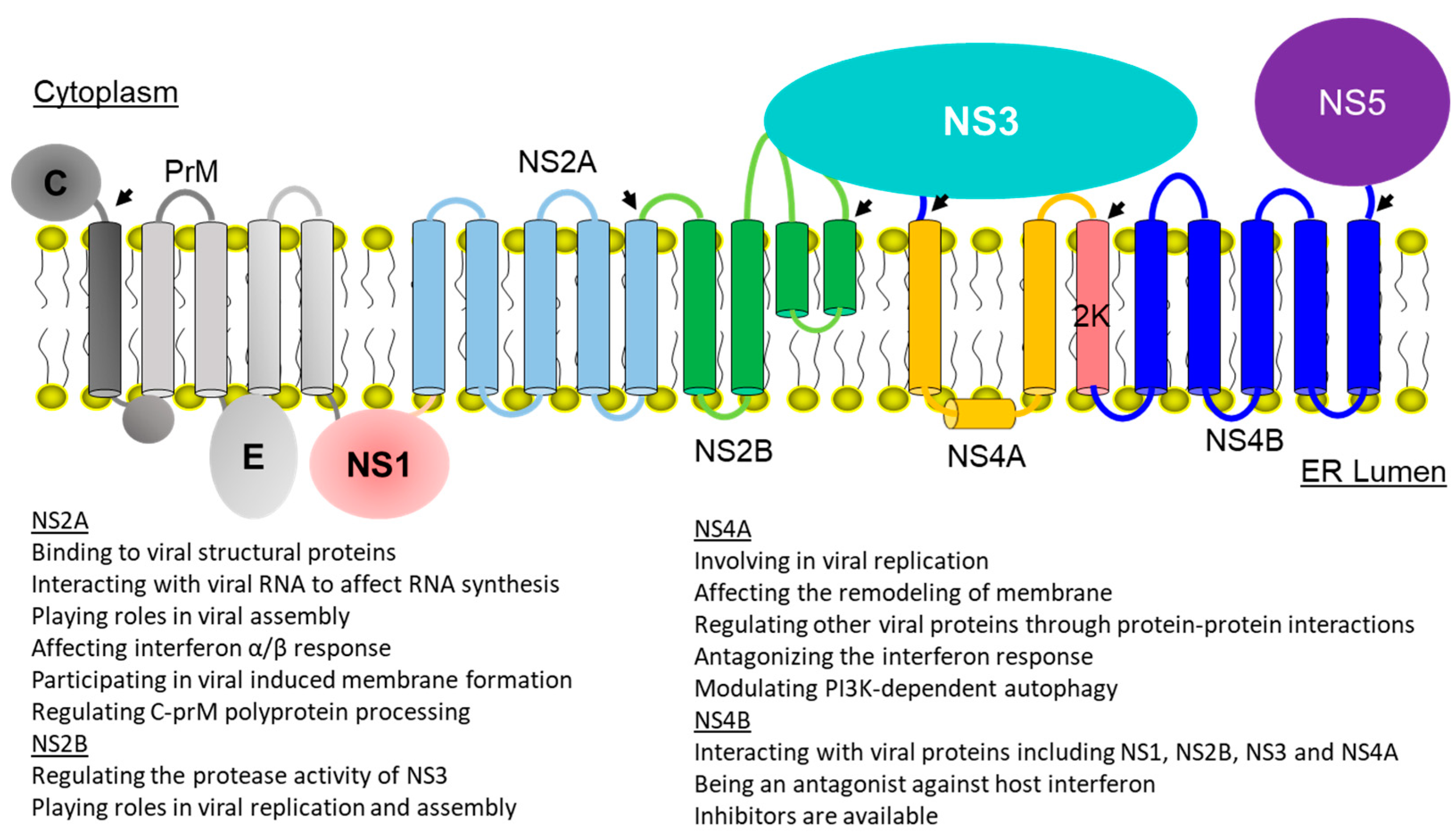
2. Membrane Topologies and Functions of Viral Membrane Proteins
2.1. Dengue NS2A
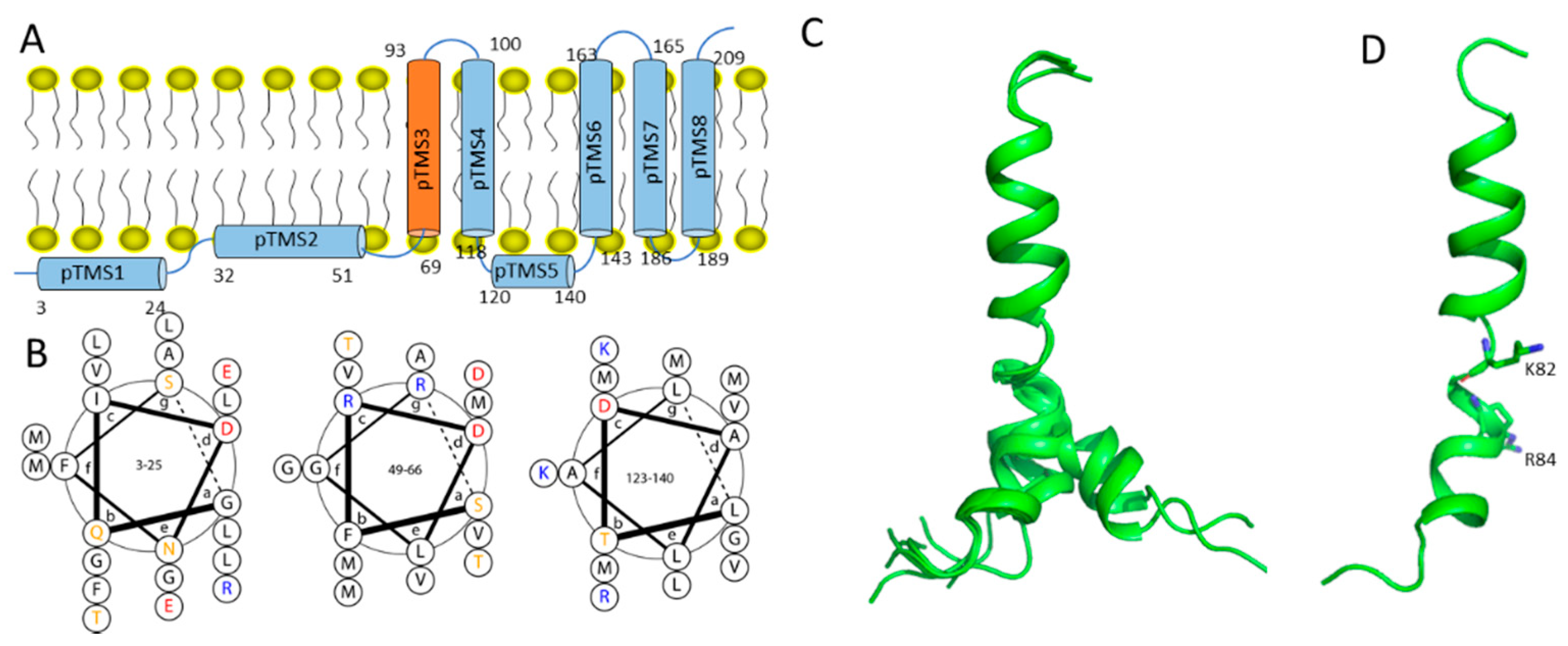
2.2. Dengue NS2B

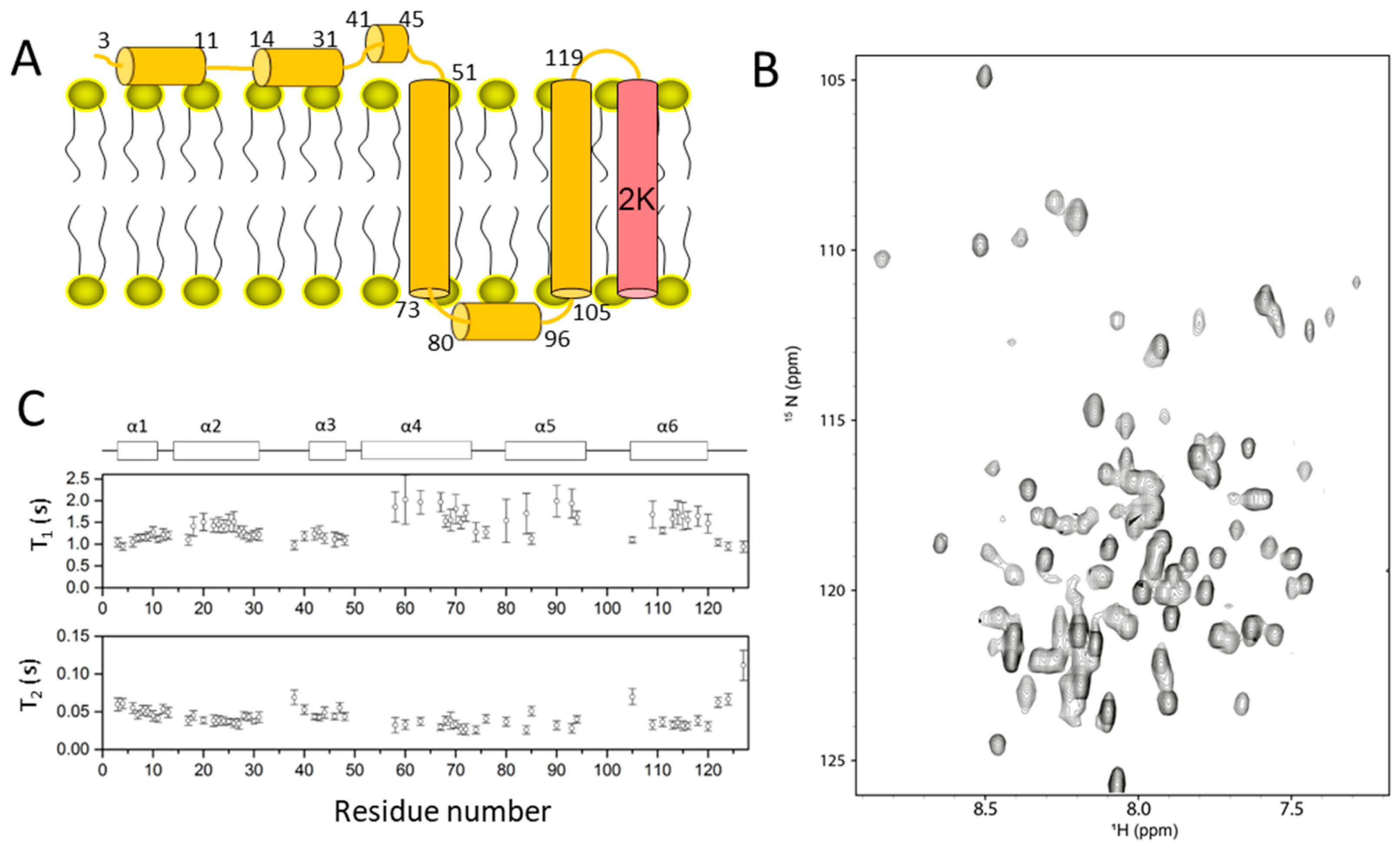
2.3. Dengue NS4A
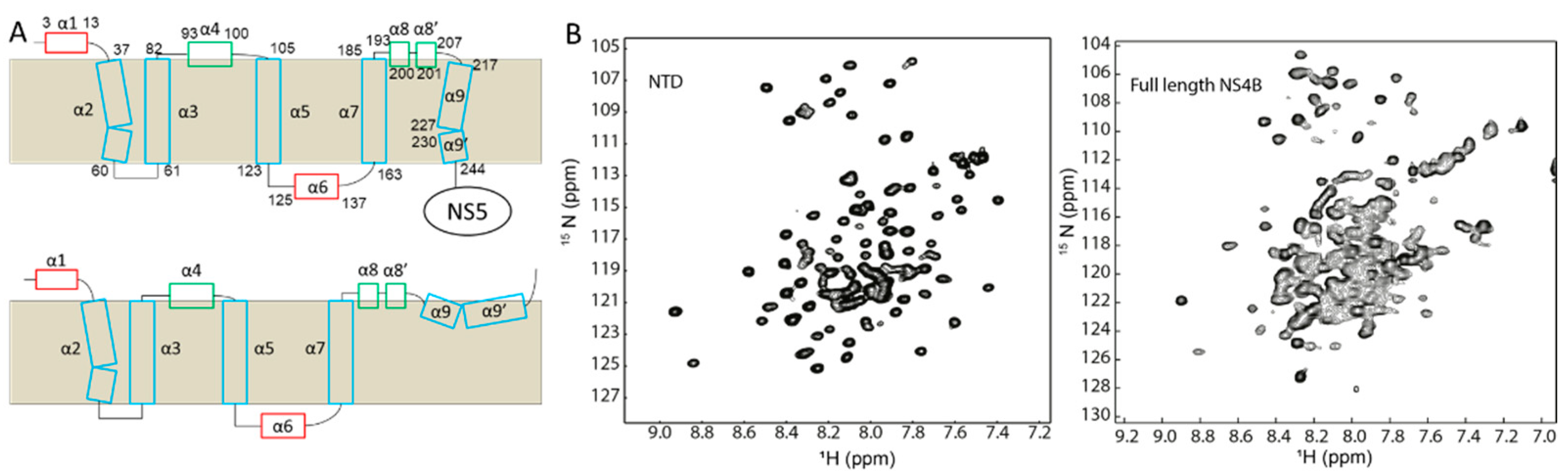
2.4. Dengue Virus NS4B
3. Dengue Membrane Proteins as a Drug Target
3.1. NS2A Inhibitors
3.2. NS2B Inhibitors
3.3. NS4A Inhibitors
3.4. NS4B Inhibitors
4. Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Bhatt, S.; Gething, P.W.; Brady, O.J.; Messina, J.P.; Farlow, A.W.; Moyes, C.L.; Drake, J.M.; Brownstein, J.S.; Hoen, A.G.; Sankoh, O.; et al. The global distribution and burden of dengue. Nature 2013, 496, 504–507. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.P.; Wang, Q.Y.; Noble, C.G.; Chen, Y.L.; Dong, H.; Zou, B.; Yokokawa, F.; Nilar, S.; Smith, P.; Beer, D.; et al. Ten years of dengue drug discovery: Progress and prospects. Antivir. Res. 2013, 100, 500–519. [Google Scholar] [CrossRef] [PubMed]
- Villar, L.; Dayan, G.H.; Arredondo-García, J.L.; Rivera, D.M.; Cunha, R.; Deseda, C.; Reynales, H.; Costa, M.S.; Morales-Ramírez, J.O.; Carrasquilla, G.; et al. Efficacy of a Tetravalent Dengue Vaccine in Children in Latin America. N. Engl. J. Med. 2015, 372, 113–123. [Google Scholar] [PubMed]
- Nitsche, C.; Holloway, S.; Schirmeister, T.; Klein, C.D. Biochemistry and Medicinal Chemistry of the Dengue Virus Protease. Chem. Rev. 2014, 114, 11348–11381. [Google Scholar] [CrossRef] [PubMed]
- Rawlinson, S.M.; Pryor, M.J.; Wright, P.J.; Jans, D.A. Dengue virus RNA polymerase NS5: A potential therapeutic target? Curr. Drug Targets 2006, 7, 1623–1638. [Google Scholar] [CrossRef]
- Behnam, M.A.M.; Nitsche, C.; Boldescu, V.; Klein, C.D. The Medicinal Chemistry of Dengue. Virus J. Med. Chem. 2016, 59, 5622–5649. [Google Scholar] [CrossRef]
- Xie, X.; Zou, J.; Wang, Q.Y.; Shi, P.Y. Targeting dengue virus NS4B protein for drug discovery. Antivir. Res. 2015, 118, 39–45. [Google Scholar] [CrossRef]
- Luo, D.; Vasudevan, S.G.; Lescar, J. The flavivirus NS2B-NS3 protease-helicase as a target for antiviral drug development. Antivir. Res. 2015, 118, 148–158. [Google Scholar] [CrossRef]
- Lim, S.P.; Noble, C.G.; Shi, P.Y. The dengue virus NS5 protein as a target for drug discovery. Antivir. Res. 2015, 119, 57–67. [Google Scholar]
- Lim, S.P.; Shi, P.Y. West Nile virus drug discovery. Viruses 2013, 5, 2977–3006. [Google Scholar] [CrossRef]
- Brecher, M.; Zhang, J.; Li, H. The flavivirus protease as a target for drug discovery. Virol. Sin. 2013, 28, 326–336. [Google Scholar] [CrossRef] [PubMed]
- Noble, C.G.; Shi, P.Y. Structural biology of dengue virus enzymes: Towards rational design of therapeutics. Antivir. Res. 2012, 96, 115–126. [Google Scholar] [CrossRef] [PubMed]
- Sampath, A.; Padmanabhan, R. Molecular targets for flavivirus drug discovery. Antivir. Res. 2009, 81, 6–15. [Google Scholar] [CrossRef] [PubMed]
- Perera, R.; Kuhn, R.J. Structural proteomics of dengue virus. Curr. Opin. Microbiol. 2008, 11, 369–377. [Google Scholar] [CrossRef] [PubMed]
- Malet, H.; Masse, N.; Selisko, B.; Romette, J.L.; Alvarez, K.; Guillemot, J.C.; Tolou, H.; Yap, T.L.; Vasudevan, S.; Lescar, J.; et al. The flavivirus polymerase as a target for drug discovery. Antivir. Res. 2008, 80, 23–35. [Google Scholar] [CrossRef]
- Melino, S.; Paci, M. Progress for dengue virus diseases. Towards the NS2B-NS3pro inhibition for a therapeutic-based approach. FEBS J. 2007, 274, 2986–3002. [Google Scholar] [CrossRef]
- Mass, N.; Selisko, B.; Malet, H.; Peyrane, F.; Debarnot, C.; Decroly, E.; Benarroch, D.; Egloff, M.P.; Guillernot, J.C.; Alvarez, K.; et al. Dengue virus: Viral targets and antiviral drugs. Virologie 2007, 11, 121–133. [Google Scholar]
- Aleshin, A.E.; Shiryaev, S.A.; Strongin, A.Y.; Liddington, R.C. Structural evidence for regulation and specificity of flaviviral proteases and evolution of the Flaviviridae fold. Protein Sci. 2007, 16, 795–806. [Google Scholar] [CrossRef]
- Zou, J.; Shi, P.-Y. Strategies for Zika drug discovery. Curr. Opin. Virol. 2019, 35, 19–26. [Google Scholar] [CrossRef]
- Pierson, T.C.; Kielian, M. Flaviviruses: Braking the entering. Curr. Opin. Virol. 2013, 3, 3–12. [Google Scholar] [CrossRef]
- Barnard, T.R.; Abram, Q.H.; Lin, Q.F.; Wang, A.B.; Sagan, S.M. Molecular Determinants of Flavivirus Virion Assembly. Trends Biochem. Sci. 2021, 46, 378–390. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.P.; Wen, D.; Yap, T.L.; Yan, C.K.; Lescar, J.; Vasudevan, S.G. A scintillation proximity assay for dengue virus NS5 2′-O-methyltransferase-kinetic and inhibition analyses. Antivir. Res. 2008, 80, 360–369. [Google Scholar] [PubMed]
- Yon, C.; Teramoto, T.; Mueller, N.; Phelan, J.; Ganesh, V.K.; Murthy, K.H.M.; Padmanabhan, R. Modulation of the nucleoside triphosphatase/RNA helicase and 5′ -RNA triphosphatase activities of dengue virus type 2 nonstructural protein 3 (NS3) by interaction with NS5, the RNA-dependent RNA polymerase. J. Biol. Chem. 2005, 280, 27412–27419. [Google Scholar] [PubMed]
- Nasar, S.; Rashid, N.; Iftikhar, S. Dengue proteins with their role in pathogenesis, and strategies for developing an effective anti-dengue treatment: A review. J. Med. Virol. 2020, 92, 941–955. [Google Scholar] [CrossRef]
- Vossmann, S.; Wieseler, J.; Kerber, R.; Kummerer, B.M. A basic cluster in the N terminus of yellow fever virus NS2A contributes to infectious particle production. J. Virol. 2015, 89, 4951–4965. [Google Scholar] [CrossRef]
- Leung, J.Y.; Pijlman, G.P.; Kondratieva, N.; Hyde, J.; Mackenzie, J.M.; Khromykh, A.A. Role of nonstructural protein NS2A in flavivirus assembly. J. Virol. 2008, 82, 4731–4741. [Google Scholar] [CrossRef]
- Swarbrick, C.M.D.; Basavannacharya, C.; Chan, K.W.K.; Chan, S.-A.; Singh, D.; Wei, N.; Phoo, W.W.; Luo, D.; Lescar, J.; Vasudevan, S.G. NS3 helicase from dengue virus specifically recognizes viral RNA sequence to ensure optimal replication. Nucleic Acids Res. 2017, 45, 12904–12920. [Google Scholar] [CrossRef]
- Luo, D.; Xu, T.; Watson, R.P.; Scherer-Becker, D.; Sampath, A.; Jahnke, W.; Yeong, S.S.; Wang, C.H.; Lim, S.P.; Strongin, A.; et al. Insights into RNA unwinding and ATP hydrolysis by the flavivirus NS3 protein. EMBO J. 2008, 27, 3209–3219. [Google Scholar] [CrossRef]
- Lescar, J.; Luo, D.; Xu, T.; Sampath, A.; Lim, S.P.; Canard, B.; Vasudevan, S.G. Towards the design of antiviral inhibitors against flaviviruses: The case for the multifunctional NS3 protein from Dengue virus as a target. Antivir. Res. 2008, 80, 94–101. [Google Scholar] [CrossRef]
- Xu, T.; Sampath, A.; Chao, A.; Wen, D.; Nanao, M.; Luo, D.; Chene, P.; Vasudevan, S.G.; Lescar, J. Towards the design of flavivirus helicase/NTPase inhibitors: Crystallographic and mutagenesis studies of the dengue virus NS3 helicase catalytic domain. Novartis Found. Symp. 2006, 277, 87–97; discussion 97–101, 103–251. [Google Scholar]
- Umareddy, I.; Chao, A.; Sampath, A.; Gu, F.; Vasudevan, S.G. Dengue virus NS4B interacts with NS3 and dissociates it from single-stranded RNA. J. Gen. Virol. 2006, 87, 2605–2614. [Google Scholar] [CrossRef] [PubMed]
- Prikhod’ko, G.G.; Prikhod’ko, E.A.; Pletnev, A.G.; Cohen, J.I. Langat flavivirus protease NS3 binds caspase-8 and induces apoptosis. J. Virol. 2002, 76, 5807–5812. [Google Scholar] [CrossRef] [PubMed]
- Zou, G.; Chen, Y.L.; Dong, H.; Lim, C.C.; Yap, L.J.; Yau, Y.H.; Shochat, S.G.; Lescar, J.; Shi, P.Y. Functional analysis of two cavities in flavivirus NS5 polymerase. J. Biol. Chem. 2011, 286, 14362–14372. [Google Scholar] [CrossRef] [PubMed]
- Luo, D.; Xu, T.; Hunke, C.; Gruber, G.; Vasudevan, S.G.; Lescar, J. Crystal structure of the NS3 protease-helicase from dengue virus. J. Virol. 2008, 82, 173–183. [Google Scholar] [CrossRef]
- Shiryaev, S.A.; Ratnikov, B.I.; Aleshin, A.E.; Kozlov, I.A.; Nelson, N.A.; Lebl, M.; Smith, J.W.; Liddington, R.C.; Strongin, A.Y. Switching the substrate specificity of the two-component NS2B-NS3 flavivirus proteinase by structure-based mutagenesis. J. Virol. 2007, 81, 4501–4509. [Google Scholar] [CrossRef]
- Shiryaev, S.A.; Aleshin, A.E.; Ratnikov, B.I.; Smith, J.W.; Liddington, R.C.; Strongin, A.Y. Expression and purification of a two-component flaviviral proteinase resistant to autocleavage at the NS2B-NS3 junction region. Protein Expr. Purif. 2007, 52, 334–339. [Google Scholar] [CrossRef][Green Version]
- Yin, Z.; Lim, S.P.; Patel, S.; Patel, V.; Beer, D.; Ma, N.L.; Vasudevan, S.; Keller, T. Targeting the protease activity of Dengue virus NS3. Acta Pharmacol. Sin. 2006, 27, 251. [Google Scholar]
- Kroschewski, H.; Lim, S.P.; Butcher, R.E.; Yap, T.L.; Lescar, J.; Wright, P.J.; Vasudevan, S.G.; Davidson, A.D. Mutagenesis of the dengue virus type 2 NS5 methyltransferase domain. J. Biol. Chem. 2008, 283, 19410–19421. [Google Scholar] [CrossRef]
- Poulsen, A.; Kang, C.; Keller, T.H. Drug design for flavivirus proteases: What are we missing? Curr. Pharm. Des. 2014, 20, 3422–3427. [Google Scholar] [CrossRef]
- Kang, C.; Gayen, S.; Wang, W.; Severin, R.; Chen, A.S.; Lim, H.A.; Chia, C.S.; Schuller, A.; Doan, D.N.; Poulsen, A.; et al. Exploring the binding of peptidic West Nile virus NS2B-NS3 protease inhibitors by NMR. Antivir. Res. 2013, 97, 137–144. [Google Scholar] [CrossRef]
- Lim, S.P. Dengue drug discovery: Progress, challenges and outlook. Antivir. Res. 2019, 163, 156–178. [Google Scholar] [CrossRef] [PubMed]
- Nitsche, C. Strategies Towards Protease Inhibitors for Emerging Flaviviruses. In Dengue and Zika: Control and Antiviral Treatment Strategies; Hilgenfeld, R., Vasudevan, S.G., Eds.; Springer: Singapore, 2018; pp. 175–186. [Google Scholar]
- Timiri, A.K.; Sinha, B.N.; Jayaprakash, V. Progress and prospects on DENV protease inhibitors. Eur. J. Med. Chem. 2016, 117, 125–143. [Google Scholar] [CrossRef] [PubMed]
- Moquin, S.A.; Simon, O.; Karuna, R.; Lakshminarayana, S.B.; Yokokawa, F.; Wang, F.; Saravanan, C.; Zhang, J.; Day, C.W.; Chan, K.; et al. NITD-688, a pan-serotype inhibitor of the dengue virus NS4B protein, shows favorable pharmacokinetics and efficacy in preclinical animal models. Sci. Transl. Med. 2021, 13, eabb2181. [Google Scholar] [CrossRef] [PubMed]
- Kaptein, S.J.; Goethals, O.; Kiemel, D.; Marchand, A.; Kesteleyn, B.; Bonfanti, J.-F.; Bardiot, D.; Stoops, B.; Jonckers, T.H.M.; Dallmeier, K.; et al. A pan-serotype dengue virus inhibitor targeting the NS3–NS4B interaction. Nature 2021, 598, 504–509. [Google Scholar] [CrossRef]
- Wang, Q.Y.; Dong, H.; Zou, B.; Karuna, R.; Wan, K.F.; Zou, J.; Susila, A.; Yip, A.; Shan, C.; Yeo, K.L.; et al. Discovery of Dengue Virus NS4B Inhibitors. J. Virol. 2015, 89, 8233–8244. [Google Scholar] [CrossRef]
- Li, Y.; Wong, Y.L.; Lee, M.Y.; Li, Q.; Wang, Q.Y.; Lescar, J.; Shi, P.Y.; Kang, C. Secondary Structure and Membrane Topology of the Full-Length Dengue Virus NS4B in Micelles. Angew. Chem. Int. Ed. 2016, 55, 12068–12072. [Google Scholar] [CrossRef]
- Li, Y.; Li, Q.; Wong, Y.L.; Liew, L.S.; Kang, C. Membrane topology of NS2B of dengue virus revealed by NMR spectroscopy. Biochim. Biophys. Acta 2015, 1848, 2244–2252. [Google Scholar] [CrossRef]
- Li, Y.; Kim, Y.M.; Zou, J.; Wang, Q.Y.; Gayen, S.; Wong, Y.L.; Lee, L.T.; Xie, X.; Huang, Q.; Lescar, J.; et al. Secondary structure and membrane topology of dengue virus NS4B N-terminal 125 amino acids. Biochim. Biophys. Acta 2015, 1848, 3150–3157. [Google Scholar] [CrossRef]
- Li, Y.; Lee, M.Y.; Loh, Y.R.; Kang, C. Secondary structure and membrane topology of dengue virus NS4A protein in micelles. Biochim. Biophys. Acta Biomembr. 2018, 1860, 442–450. [Google Scholar] [CrossRef]
- Reddy, S.B.G.; Chin, W.-X.; Shivananju, N.S. Dengue virus NS2 and NS4: Minor proteins, mammoth roles. Biochem. Pharmacol. 2018, 154, 54–63. [Google Scholar] [CrossRef]
- Huang, Q.; Li, Q.; Joy, J.; Chen, A.S.; Ruiz-Carrillo, D.; Hill, J.; Lescar, J.; Kang, C. Lyso-myristoyl phosphatidylcholine micelles sustain the activity of Dengue non-structural (NS) protein 3 protease domain fused with the full-length NS2B. Protein Expr. Purif. 2013, 92, 156–162. [Google Scholar] [CrossRef] [PubMed]
- Huang, Q.; Chen, A.S.; Li, Q.; Kang, C. Expression, purification, and initial structural characterization of nonstructural protein 2B, an integral membrane protein of Dengue-2 virus, in detergent micelles. Protein Expr. Purif. 2011, 80, 169–175. [Google Scholar] [CrossRef] [PubMed]
- Kang, C.; Li, Q. Solution NMR study of integral membrane proteins. Curr. Opin. Chem. Biol. 2011, 15, 560–569. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Ng, H.Q.; Kang, C. Secondary structure and topology of the transmembrane domain of Syndecan-2 in detergent micelles. FEBS Lett. 2019, 593, 554–561. [Google Scholar] [CrossRef]
- Ng, E.Y.; Loh, Y.R.; Li, Y.; Li, Q.; Kang, C. Expression, purification of Zika virus membrane protein-NS2B in detergent micelles for NMR studies. Protein Expr. Purif. 2019, 154, 1–6. [Google Scholar] [CrossRef]
- Xie, X.; Gayen, S.; Kang, C.; Yuan, Z.; Shi, P.Y. Membrane topology and function of dengue virus NS2A protein. J. Virol. 2013, 87, 4609–4622. [Google Scholar] [CrossRef]
- Mackenzie, J.M.; Khromykh, A.A.; Jones, M.K.; Westaway, E.G. Subcellular localization and some biochemical properties of the flavivirus Kunjin nonstructural proteins NS2A and NS4A. Virology 1998, 245, 203–215. [Google Scholar] [CrossRef]
- Taniguchi, T.; Takaoka, A. The interferon-α/β system in antiviral responses: A multimodal machinery of gene regulation by the IRF family of transcription factors. Curr. Opin. Immunol. 2002, 14, 111–116. [Google Scholar] [CrossRef]
- Muñoz-Jordán, J.L.; Sánchez-Burgos, G.G.; Laurent-Rolle, M.; García-Sastre, A. Inhibition of interferon signaling by dengue virus. Proc. Natl. Acad. Sci. USA 2003, 100, 14333–14338. [Google Scholar] [CrossRef]
- Xie, X.; Zou, J.; Zhang, X.; Zhou, Y.; Routh, A.L.; Kang, C.; Popov, V.L.; Chen, X.; Wang, Q.Y.; Dong, H.; et al. Dengue NS2A Protein Orchestrates Virus Assembly. Cell Host Microbe 2019, 26, 606–622.e608. [Google Scholar] [CrossRef]
- Bañó-Polo, M.; Baeza-Delgado, C.; Orzáez, M.; Marti-Renom, M.A.; Abad, C.; Mingarro, I. Polar/Ionizable Residues in Transmembrane Segments: Effects on Helix-Helix Packing. PLoS ONE 2012, 7, e44263. [Google Scholar] [CrossRef] [PubMed]
- Yusof, R.; Clum, S.; Wetzel, M.; Murthy, H.M.K.; Padmanabhan, R. Purified NS2B/NS3 serine protease of dengue virus type 2 exhibits cofactor NS2B dependence for cleavage of substrates with dibasic amino acids in vitro. J. Biol. Chem. 2000, 275, 9963–9969. [Google Scholar] [CrossRef] [PubMed]
- Clum, S.; Ebner, K.E. Padmanabhan, Cotranslational membrane insertion of the serine proteinase precursor NS2B-NS3(Pro) of dengue virus type 2 is required for efficient in vitro processing and is mediated through the hydrophobic regions of NS2B. J. Biol. Chem. 1997, 272, 30715–30723. [Google Scholar] [CrossRef] [PubMed]
- Robin, G.; Chappell, K.; Stoermer, M.J.; Hu, S.H.; Young, P.R.; Fairlie, D.P.; Martin, J.L. Structure of West Nile virus NS3 protease: Ligand stabilization of the catalytic conformation. J. Mol. Biol. 2009, 385, 1568–1577. [Google Scholar] [CrossRef]
- Noble, C.G.; Seh, C.C.; Chao, A.T.; Shi, P.Y. Ligand-bound structures of the dengue virus protease reveal the active conformation. J. Virol. 2011, 86, 438–446. [Google Scholar] [CrossRef]
- Radichev, I.; Shiryaev, S.A.; Aleshin, A.E.; Ratnikov, B.I.; Smith, J.W.; Liddington, R.C.; Strongin, A.Y. Structure-based mutagenesis identifies important novel determinants of the NS2B cofactor of the West Nile virus two-component NS2B-NS3 proteinase. J. Gen. Virol. 2008, 89, 636–641. [Google Scholar] [CrossRef]
- Erbel, P.; Schiering, N.; D’Arcy, A.; Renatus, M.; Kroemer, M.; Lim, S.P.; Yin, Z.; Keller, T.H.; Vasudevan, S.G.; Hommel, U. Structural basis for the activation of flaviviral NS3 proteases from dengue and West Nile virus. Nat. Struct. Mol. Biol. 2006, 13, 372–373. [Google Scholar] [CrossRef]
- de la Cruz, L.; Nguyen, T.H.; Ozawa, K.; Shin, J.; Graham, B.; Huber, T.; Otting, G. Binding of low molecular weight inhibitors promotes large conformational changes in the dengue virus NS2B-NS3 protease: Fold analysis by pseudocontact shifts. J. Am. Chem. Soc. 2011, 133, 19205–19215. [Google Scholar] [CrossRef]
- Kim, Y.M.; Gayen, S.; Kang, C.; Joy, J.; Huang, Q.; Chen, A.S.; Wee, J.L.; Ang, M.J.; Lim, H.A.; Hung, A.W.; et al. NMR analysis of a novel enzymatically active unlinked dengue NS2B-NS3 protease complex. J. Biol. Chem. 2013, 288, 12891–12900. [Google Scholar] [CrossRef]
- Zhang, Z.; Li, Y.; Loh, Y.R.; Phoo, W.W.; Hung, A.W.; Kang, C.; Luo, D. Crystal structure of unlinked NS2B-NS3 protease from Zika virus. Science 2016, 354, 1597–1600. [Google Scholar] [CrossRef]
- Phoo, W.W.; Li, Y.; Zhang, Z.; Lee, M.Y.; Loh, Y.R.; Tan, Y.B.; Ng, E.Y.; Lescar, J.; Kang, C.; Luo, D. Structure of the NS2B-NS3 protease from Zika virus after self-cleavage. Nat. Commun. 2016, 7, 13410. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Kang, C. Structure and Dynamics of Zika Virus Protease and Its Insights into Inhibitor Design. Biomedicines 2021, 9, 1044. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Kang, C. Mechanisms of Action for Small Molecules Revealed by Structural Biology in Drug Discovery. Int. J. Mol. Sci. 2020, 21, 5262. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Kang, C. Insights into Structures and Dynamics of Flavivirus Proteases from NMR Studies. Int. J. Mol. Sci. 2020, 21, 2527. [Google Scholar] [CrossRef] [PubMed]
- de la Cruz, L.; Chen, W.N.; Graham, B.; Otting, G. Binding mode of the activity-modulating C-terminal segment of NS2B to NS3 in the dengue virus NS2B-NS3 protease. FEBS J. 2014, 281, 1517–1533. [Google Scholar] [CrossRef]
- Chen, W.N.; Loscha, K.V.; Nitsche, C.; Graham, B.; Otting, G. The dengue virus NS2B-NS3 protease retains the closed conformation in the complex with BPTI. FEBS Lett. 2014, 588, 2206–2211. [Google Scholar] [CrossRef] [PubMed]
- Su, X.C.; Ozawa, K.; Qi, R.; Vasudevan, S.G.; Lim, S.P.; Otting, G. NMR analysis of the dynamic exchange of the NS2B cofactor between open and closed conformations of the West Nile virus NS2B-NS3 protease. PLOS Negl. Trop. Dis. 2009, 3, e561. [Google Scholar] [CrossRef]
- Choksupmanee, O.; Hodge, K.; Katzenmeier, G.; Chimnaronk, S. Structural platform for the autolytic activity of an intact NS2B-NS3 protease complex from dengue virus. Biochemistry 2012, 51, 2840–2851. [Google Scholar] [CrossRef]
- Wu, R.-H.; Tsai, M.-H.; Tsai, K.-N.; Tian, J.N.; Wu, J.-S.; Wu, S.-Y.; Chern, J.-H.; Chen, C.-H.; Yueh, A.; Diamond, M.S. Mutagenesis of Dengue Virus Protein NS2A Revealed a Novel Domain Responsible for Virus-Induced Cytopathic Effect and Interactions between NS2A and NS2B Transmembrane Segments. J. Virol. 2017, 91, e01836-16. [Google Scholar] [CrossRef]
- Cordero, J.G.; Juárez, M.L.; González-Y-Merchand, J.A.; Barrón, L.C.; Castañeda, B.G. Caveolin-1 in Lipid Rafts Interacts with Dengue Virus NS3 during Polyprotein Processing and Replication in HMEC-1 Cells. PLoS ONE 2014, 9, e90704. [Google Scholar]
- Li, X.D.; Deng, C.L.; Ye, H.Q.; Zhang, H.L.; Zhang, Q.Y.; Chen, D.D.; Zhang, P.T.; Shi, P.Y.; Yuan, Z.M.; Zhang, B. Transmembrane Domains of NS2B Contribute to both Viral RNA Replication and Particle Formation in Japanese Encephalitis Virus. J. Virol. 2016, 90, 5735–5749. [Google Scholar] [CrossRef] [PubMed]
- León-Juárez, M.; Martínez-Castillo, M.; Shrivastava, G.; García-Cordero, J.; Villegas-Sepulveda, N.; Mondragón-Castelán, M.; Mondragón-Flores, R.; Cedillo-Barrón, L. Recombinant Dengue virus protein NS2B alters membrane permeability in different membrane models. Virol. J. 2016, 13, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Aguirre, S.; Luthra, P.; Sanchez-Aparicio, M.T.; Maestre, A.M.; Patel, J.; Lamothe, F.; Fredericks, A.C.; Tripathi, S.; Zhu, T.; Pintado-Silva, J.; et al. Dengue virus NS2B protein targets cGAS for degradation and prevents mitochondrial DNA sensing during infection. Nat. Microbiol. 2017, 2, 17037. [Google Scholar] [CrossRef]
- Yu, L.; Liu, P. Cytosolic DNA sensing by cGAS: Regulation, function, and human diseases. Signal Transduct. Target. Ther. 2021, 6, 170. [Google Scholar] [CrossRef] [PubMed]
- Gayen, S.; Chen, A.S.; Huang, Q.; Kang, C. West Nile Virus (WNV) protease and membrane interactions revealed by NMR spectroscopy. Biochem. Biophys. Res. Commun. 2012, 423, 799–804. [Google Scholar] [CrossRef] [PubMed]
- Su, X.C.; Ozawa, K.; Yagi, H.; Lim, S.P.; Wen, D.; Ekonomiuk, D.; Huang, D.; Keller, T.H.; Sonntag, S.; Caflisch, A.; et al. NMR study of complexes between low molecular mass inhibitors and the West Nile virus NS2B-NS3 protease. FEBS J. 2009, 276, 4244–4255. [Google Scholar] [CrossRef]
- Luo, D.; Wei, N.; Doan, D.N.; Paradkar, P.N.; Chong, Y.; Davidson, A.D.; Kotaka, M.; Lescar, J.; Vasudevan, S.G. Flexibility between the protease and helicase domains of the dengue virus NS3 protein conferred by the linker region and its functional implications. J. Biol. Chem. 2010, 285, 18817–18827. [Google Scholar] [CrossRef]
- Huang, Q.; Li, Q.; Chen, A.S.; Kang, C. West Nile virus protease activity in detergent solutions and application for affinity tag removal. Anal. Biochem. 2013, 435, 44–46. [Google Scholar] [CrossRef]
- Liew, L.S.; Lee, M.Y.; Wong, Y.L.; Cheng, J.; Li, Q.; Kang, C. Selection of suitable detergents for obtaining an active dengue protease in its natural form from E. coli. Protein Expr. Purif. 2016, 121, 141–148. [Google Scholar] [CrossRef]
- Miller, S.; Kastner, S.; Krijnse-Locker, J.; Bühler, S.; Bartenschlager, R. The Non-structural Protein 4A of Dengue Virus Is an Integral Membrane Protein Inducing Membrane Alterations in a 2K-regulated Manner. J. Biol. Chem. 2007, 282, 8873–8882. [Google Scholar] [CrossRef]
- Lin, C.; Amberg, S.M.; Chambers, T.J.; Rice, C.M. Cleavage at a novel site in the NS4A region by the yellow fever virus NS2B-3 proteinase is a prerequisite for processing at the downstream 4A/4B signalase site. J. Virol. 1993, 67, 2327–2335. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Kumar, P.; Giri, R. Zika virus NS4A cytosolic region (residues 1–48) is an intrinsically disordered domain and folds upon binding to lipids. Virology 2020, 550, 27–36. [Google Scholar] [CrossRef] [PubMed]
- Hung, Y.F.; Schwarten, M.; Hoffmann, S.; Willbold, D.; Sklan, E.H.; Koenig, B. Amino Terminal Region of Dengue Virus NS4A Cytosolic Domain Binds to Highly Curved Liposomes. Viruses 2015, 7, 4119–4130. [Google Scholar] [CrossRef] [PubMed]
- Hung, Y.F.; Schwarten, M.; Schunke, S.; Thiagarajan-Rosenkranz, P.; Hoffmann, S.; Sklan, E.H.; Willbold, D.; Koenig, B.W. Dengue virus NS4A cytoplasmic domain binding to liposomes is sensitive to membrane curvature. Biochim. Biophys. Acta 2015, 1848, 1119–1126. [Google Scholar] [CrossRef] [PubMed]
- To, J.; Torres, J. Trimerization of the N-Terminal Tail of Zika Virus NS4A Protein: A Potential In Vitro Antiviral Screening Assay. Membranes 2021, 11, 335. [Google Scholar] [CrossRef]
- Tan, M.J.A.; Brown, N.G.; Chan, K.W.K.; Jin, J.Y.; Kong, S.Y.Z.; Vasudevan, S.G. Mutations in the cytoplasmic domain of dengue virus NS4A affect virus fitness and interactions with other non-structural proteins. J. Gen. Virol. 2020, 101, 941–953. [Google Scholar] [CrossRef]
- Klaitong, P.; Smith, D.R. Roles of Non-Structural Protein 4A in Flavivirus Infection. Viruses 2021, 13, 2077. [Google Scholar] [CrossRef]
- Mikulasova, A.; Gillespie, L.K.; Ambrose, R.L.; Aktepe, T.E.; Trenerry, A.M.; Liebscher, S.; Mackenzie, J.M. A Putative Lipid-Associating Motif in the West Nile Virus NS4A Protein Is Required for Efficient Virus Replication. Front. Cell Dev. Biol. 2021, 9, 655606. [Google Scholar] [CrossRef]
- Ambrose, R.L.; Mackenzie, J.M. A Conserved Peptide in West Nile Virus NS4A Protein Contributes to Proteolytic Processing and Is Essential for Replication. J. Virol. 2011, 85, 11274–11282. [Google Scholar] [CrossRef]
- Ambrose, R.L.; Mackenzie, J.M. Conserved amino acids within the N-terminus of the West Nile virus NS4A protein contribute to virus replication, protein stability and membrane proliferation. Virology 2015, 481, 95–106. [Google Scholar] [CrossRef]
- Teo, C.S.H.; Chu, J.J.H. Cellular Vimentin Regulates Construction of Dengue Virus Replication Complexes through Interaction with NS4A Protein. J. Virol. 2014, 88, 1897–1913. [Google Scholar] [CrossRef] [PubMed]
- Gillespie, L.K.; Hoenen, A.; Morgan, G.; Mackenzie, J.M. The Endoplasmic Reticulum Provides the Membrane Platform for Biogenesis of the Flavivirus Replication Complex. J. Virol. 2010, 84, 10438–10447. [Google Scholar] [CrossRef] [PubMed]
- Junjhon, J.; Pennington, J.G.; Edwards, T.J.; Perera, R.; Lanman, J.; Kuhn, R.J.; Doms, R.W. Ultrastructural Characterization and Three-Dimensional Architecture of Replication Sites in Dengue Virus-Infected Mosquito Cells. J. Virol. 2014, 88, 4687–4697. [Google Scholar] [CrossRef] [PubMed]
- Welsch, S.; Miller, S.; Romero-Brey, I.; Merz, A.; Bleck, C.K.E.; Walther, P.; Fuller, S.D.; Antony, C.; Krijnse-Locker, J.; Bartenschlager, R. Composition and Three-Dimensional Architecture of the Dengue Virus Replication and Assembly Sites. Cell Host Microbe 2009, 5, 365–375. [Google Scholar] [CrossRef] [PubMed]
- Ngo, A.M.; Shurtleff, M.J.; Popova, K.D.; Kulsuptrakul, J.; Weissman, J.S.; Puschnik, A.S. The ER membrane protein complex is required to ensure correct topology and stable expression of flavivirus polyproteins. eLife 2019, 8, 48469. [Google Scholar] [CrossRef]
- Lin, M.H.; Hsu, H.J.; Bartenschlager, R.; Fischer, W.B. Membrane undulation induced by NS4A of Dengue virus: A molecular dynamics simulation study. J. Biomol. Struct. Dyn. 2013, 32, 1552–1562. [Google Scholar] [CrossRef]
- Stern, O.; Hung, Y.-F.; Valdau, O.; Yaffe, Y.; Harris, E.; Hoffmann, S.; Willbold, D.; Sklan, E.H. An N-Terminal Amphipathic Helix in Dengue Virus Nonstructural Protein 4A Mediates Oligomerization and Is Essential for Replication. J. Virol. 2013, 87, 4080–4085. [Google Scholar] [CrossRef]
- Lee, C.M.; Xie, X.; Zou, J.; Li, S.H.; Lee, M.Y.; Dong, H.; Qin, C.F.; Kang, C.; Shi, P.Y. Determinants of Dengue Virus NS4A Protein Oligomerization. J. Virol. 2015, 89, 6171–6183. [Google Scholar] [CrossRef]
- Riva, L.; Goellner, S.; Biering, S.B.; Huang, C.-T.; Rubanov, A.N.; Haselmann, U.; Warnes, C.M.; Jesus, P.D.D.; Martin-Sancho, L.; Terskikh, A.V.; et al. The Compound SBI-0090799 Inhibits Zika Virus Infection by Blocking De Novo Formation of the Membranous Replication Compartment. J. Virol. 2021, 95, e00996-21. [Google Scholar] [CrossRef]
- Preugschat, F.; Yao, C.W.; Strauss, J.H. In vitro processing of dengue virus type 2 nonstructural proteins NS2A, NS2B, and NS3. J. Virol. 1990, 64, 4364–4374. [Google Scholar] [CrossRef]
- Zhang, L.; Mohan, P.M.; Padmanabhan, R. Processing and localization of Dengue virus type 2 polyprotein precursor NS3-NS4A-NS4B-NS5. J. Virol. 1992, 66, 7549–7554. [Google Scholar] [CrossRef] [PubMed]
- Shiryaev, S.A.; Chernov, A.V.; Aleshin, A.E.; Shiryaeva, T.N.; Strongin, A.Y. NS4A regulates the ATPase activity of the NS3 helicase: A novel cofactor role of the non-structural protein NS4A from West Nile virus. J. Gen. Virol. 2009, 90, 2081–2085. [Google Scholar] [CrossRef] [PubMed]
- Lindenbach, B.D.; Rice, C.M. Genetic Interaction of Flavivirus Nonstructural Proteins NS1 and NS4A as a Determinant of Replicase Function. J. Virol. 1999, 73, 4611–4621. [Google Scholar] [CrossRef] [PubMed]
- Płaszczyca, A.; Scaturro, P.; Neufeldt, C.J.; Cortese, M.; Cerikan, B.; Ferla, S.; Brancale, A.; Pichlmair, A.; Bartenschlager, R. A novel interaction between dengue virus nonstructural protein 1 and the NS4A-2K-4B precursor is required for viral RNA replication but not for formation of the membranous replication organelle. PLOS Pathog. 2019, 15, e1007736. [Google Scholar] [CrossRef]
- Zou, J.; Xie, X.; Wang, Q.Y.; Dong, H.; Lee, M.Y.; Kang, C.; Yuan, Z.; Shi, P.Y. Characterization of dengue virus NS4A and NS4B protein interaction. J. Virol. 2015, 89, 3455–3470. [Google Scholar] [CrossRef]
- Liang, Q.; Luo, Z.; Zeng, J.; Chen, W.; Foo, S.-S.; Lee, S.-A.; Ge, J.; Wang, S.; Goldman, S.-A.; Zlokovic, B.-V.; et al. Zika Virus NS4A and NS4B Proteins Deregulate Akt-mTOR Signaling in Human Fetal Neural Stem Cells to Inhibit Neurogenesis and Induce Autophagy. Cell Stem Cell 2016, 19, 663–671. [Google Scholar] [CrossRef]
- Ma, J.; Ketkar, H.; Geng, T.; Lo, E.; Wang, L.; Xi, J.; Sun, Q.; Zhu, Z.; Cui, Y.; Yang, L.; et al. Zika Virus Non-structural Protein 4A Blocks the RLR-MAVS Signaling. Front. Microbiol. 2018, 9, 1350. [Google Scholar] [CrossRef]
- Ngueyen, T.T.N.; Kim, S.J.; Lee, J.Y.; Myoung, J. Zika Virus Proteins NS2A and NS4A Are Major Antagonists that Reduce IFN-β Promoter Activity Induced by the MDA5/RIG-I Signaling Pathway. J. Microbiol. Biotechnol. 2019, 29, 1665–1674. [Google Scholar]
- McLean, J.E.; Wudzinska, A.; Datan, E.; Quaglino, D.; Zakeri, Z. Flavivirus NS4A-induced Autophagy Protects Cells against Death and Enhances Virus Replication. J. Biol. Chem. 2011, 286, 22147–22159. [Google Scholar] [CrossRef]
- Aktepe, T.E.; Liebscher, S.; Prier, J.E.; Simmons, C.P.; Mackenzie, J.M. The Host Protein Reticulon 3.1A Is Utilized by Flaviviruses to Facilitate Membrane Remodelling. Cell Rep. 2017, 21, 1639–1654. [Google Scholar] [CrossRef]
- Hung, Y.F.; Valdau, O.; Schunke, S.; Stern, O.; Koenig, B.W.; Willbold, D.; Hoffmann, S. Recombinant production of the amino terminal cytoplasmic region of dengue virus non-structural protein 4A for structural studies. PLoS ONE 2014, 9, e86482. [Google Scholar] [CrossRef]
- Zou, J.; Xie, X.; Le, T.L.; Chandrasekaran, R.; Reynaud, A.; Yap, L.; Wang, Q.Y.; Dong, H.; Kang, C.; Yuan, Z.; et al. Dimerization of flavivirus NS4B protein. J. Virol. 2014, 88, 3379–3391. [Google Scholar] [CrossRef]
- Westaway, E.G.; Khromykh, A.A.; Kenney, M.T.; Mackenzie, J.M.; Jones, M.K. Proteins C and NS4B of the flavivirus Kunjin translocate independently into the nucleus. Virology 1997, 234, 31–41. [Google Scholar] [CrossRef]
- Roosendaal, J.; Westaway, E.G.; Khromykh, A.; Mackenzie, J.M. Regulated Cleavages at the West Nile Virus NS4A-2K-NS4B Junctions Play a Major Role in Rearranging Cytoplasmic Membranes and Golgi Trafficking of the NS4A Protein. J. Virol. 2006, 80, 4623–4632. [Google Scholar] [CrossRef]
- Westaway, E.G.; Mackenzie, J.M.; Kenney, M.T.; Jones, M.K.; Khromykh, A.A. Ultrastructure of Kunjin virus-infected cells: Colocalization of NS1 and NS3 with double-stranded RNA, and of NS2B with NS3, in virus-induced membrane structures. J. Virol. 1997, 71, 6650–6661. [Google Scholar] [CrossRef]
- Lu, H.; Zhan, Y.; Li, X.; Bai, X.; Yuan, F.; Ma, L.; Wang, X.; Xie, M.; Wu, W.; Chen, Z. Novel insights into the function of an N-terminal region of DENV2 NS4B for the optimal helicase activity of NS3. Virus Res. 2021, 295, 198318. [Google Scholar] [CrossRef]
- Fanunza, E.; Grandi, N.; Quartu, M.; Carletti, F.; Ermellino, L.; Milia, J.; Corona, A.; Capobianchi, M.R.; Ippolito, G.; Tramontano, E. INMI1 Zika Virus NS4B Antagonizes the Interferon Signaling by Suppressing STAT1 Phosphorylation. Viruses 2021, 13, 2448. [Google Scholar] [CrossRef]
- Tian, J.-N.; Yang, C.-C.; Chuang, C.-K.; Tsai, M.-H.; Wu, R.-H.; Chen, C.-T.; Yueh, A. A Dengue Virus Type 2 (DENV-2) NS4B-Interacting Host Factor, SERP1, Reduces DENV-2 Production by Suppressing Viral RNA Replication. Viruses 2019, 11, 787. [Google Scholar] [CrossRef]
- Youn, S.; Li, T.; McCune, B.T.; Edeling, M.A.; Fremont, D.H.; Cristea, I.M.; Diamond, M.S. Evidence for a Genetic and Physical Interaction between Nonstructural Proteins NS1 and NS4B That Modulates Replication of West Nile Virus. J. Virol. 2012, 86, 7360–7371. [Google Scholar] [CrossRef]
- Barrett, P.J.; Chen, J.; Cho, M.-K.; Kim, J.-H.; Lu, Z.; Mathew, S.; Peng, D.; Song, Y.; van Horn, W.D.; Zhuang, T.; et al. The Quiet Renaissance of Protein Nuclear Magnetic Resonance. Biochemistry 2013, 52, 1303–1320. [Google Scholar] [CrossRef]
- Zou, J.; Lee, T.L.; Wang, Q.Y.; Xie, X.; Lu, S.; Yau, Y.H.; Yuan, Z.; Shochat, S.G.; Kang, C.; Lescar, J.; et al. Mapping the Interactions between the NS4B and NS3 proteins of dengue virus. J. Virol. 2015, 89, 3471–3483. [Google Scholar] [CrossRef] [PubMed]
- Munoz-Jordan, J.L.; Laurent-Rolle, M.; Ashour, J.; Martinez-Sobrido, L.; Ashok, M.; Lipkin, W.I.; Garcia-Sastre, A. Inhibition of alpha/beta interferon signaling by the NS4B protein of flaviviruses. J. Virol. 2005, 79, 8004–8013. [Google Scholar] [CrossRef]
- Miller, S.; Sparacio, S.; Bartenschlager, R. Subcellular localization and membrane topology of the Dengue virus type 2 Non-structural protein 4B. J. Biol. Chem. 2006, 281, 8854–8863. [Google Scholar] [CrossRef] [PubMed]
- Kang, C.B. 19F-NMR in target-based drug discovery. Curr. Med. Chem. 2019, 26, 4964–4983. [Google Scholar] [CrossRef] [PubMed]
- Bautista-Carbajal, P.; Soto-Acosta, R.; Angel-Ambrocio, A.H.; Cervantes-Salazar, M.; Loranca-Vega, C.I.; Herrera-Martínez, M.; del Angel, R.M. The calmodulin antagonist W-7 (N-(6-aminohexyl)-5-chloro-1-naphthalenesulfonamide hydrochloride) inhibits DENV infection in Huh-7 cells. Virology 2017, 501, 188–198. [Google Scholar] [CrossRef] [PubMed]
- Nemésio, H.; Villalaín, J. Membrane Interacting Regions of Dengue Virus NS2A Protein. J. Phys. Chem. B 2014, 118, 10142–10155. [Google Scholar] [CrossRef] [PubMed]
- Fajardo-Sánchez, E.; Galiano, V.; Villalaín, J. Spontaneous membrane insertion of a dengue virus NS2A peptide. Arch. Biochem. Biophys. 2017, 627, 56–66. [Google Scholar] [CrossRef]
- Soto-Acosta, R.; Bautista-Carbajal, P.; Syed, G.H.; Siddiqui, A.; del Angel, R.M. Nordihydroguaiaretic acid (NDGA) inhibits replication and viral morphogenesis of dengue virus. Antivir. Res. 2014, 109, 132–140. [Google Scholar] [CrossRef]
- Li, Y.; Loh, Y.R.; Hung, A.W.; Kang, C. Characterization of molecular interactions between Zika virus protease and peptides derived from the C-terminus of NS2B. Biochem. Biophys. Res. Commun. 2018, 503, 691–696. [Google Scholar] [CrossRef]
- Yin, Z.; Patel, S.J.; Wang, W.L.; Wang, G.; Chan, W.L.; Rao, K.R.; Alam, J.; Jeyaraj, D.A.; Ngew, X.; Patel, V.; et al. Peptide inhibitors of Dengue virus NS3 protease. Part 1: Warhead. Bioorganic Med. Chem. Lett. 2006, 16, 36–39. [Google Scholar] [CrossRef]
- Tomlinson, S.M.; Malmstrom, R.D.; Russo, A.; Mueller, N.; Pang, Y.P.; Watowich, S.J. Structure-based discovery of dengue virus protease inhibitors. Antivir. Res. 2009, 82, 110–114. [Google Scholar] [CrossRef] [PubMed]
- Leung, D.; Abbenante, G.; Fairlie, D.P. Protease inhibitors: Current status and future prospects. J. Med. Chem. 2000, 43, 305–341. [Google Scholar] [CrossRef] [PubMed]
- Yildiz, M.; Ghosh, S.; Bell, J.A.; Sherman, W.; Hardy, J.A. Allosteric Inhibition of the NS2B-NS3 Protease from Dengue Virus. ACS Chem. Biol. 2013, 8, 2744–2752. [Google Scholar] [CrossRef] [PubMed]
- Lim, L.; Dang, M.; Roy, A.; Kang, J.; Song, J. Curcumin Allosterically Inhibits the Dengue NS2B-NS3 Protease by Disrupting Its Active Conformation. ACS Omega 2020, 5, 25677–25686. [Google Scholar] [CrossRef]
- Shiryaev, S.A.; Farhy, C.; Pinto, A.; Huang, C.-T.; Simonetti, N.; Ngono, A.E.; Dewing, A.; Shresta, S.; Pinkerton, A.B.; Cieplak, P.; et al. Characterization of the Zika virus two-component NS2B-NS3 protease and structure-assisted identification of allosteric small-molecule antagonists. Antivir. Res. 2017, 143, 218–229. [Google Scholar] [CrossRef]
- Yao, Y.; Huo, T.; Lin, Y.L.; Nie, S.; Wu, F.; Hua, Y.; Wu, J.; Kneubehl, A.R.; Vogt, M.B.; Rico-Hesse, R.; et al. Discovery, X-ray Crystallography and Antiviral Activity of Allosteric Inhibitors of Flavivirus NS2B-NS3 Protease. J. Am. Chem. Soc. 2019, 141, 6832–6836. [Google Scholar] [CrossRef]
- Li, Q.; Kang, C. Perspectives on Fragment-based Drug Discovery: A Strategy Applicable to Diverse Targets. Curr. Top. Med. Chem. 2021, 21, 1099–1112. [Google Scholar]
- Quek, J.P.; Liu, S.; Zhang, Z.; Li, Y.; Ng, E.Y.; Loh, Y.R.; Hung, A.W.; Luo, D.; Kang, C. Identification and structural characterization of small molecule fragments targeting Zika virus NS2B-NS3 protease. Antivir. Res. 2020, 175, 104707. [Google Scholar] [CrossRef]
- Chang, Y.S.; Graves, B.; Guerlavais, V.; Tovar, C.; Packman, K.; To, K.-H.; Olson, K.A.; Kesavan, K.; Gangurde, P.; Mukherjee, A.; et al. Stapled α−helical peptide drug development: A potent dual inhibitor of MDM2 and MDMX for p53-dependent cancer therapy. Proc. Natl. Acad. Sci. USA 2013, 110, E3445–E3454. [Google Scholar] [CrossRef]
- Mabonga, L.; Kappo, A.P. Peptidomimetics: A Synthetic Tool for Inhibiting Protein–Protein Interactions in Cancer. Int. J. Pept. Res. Ther. 2020, 26, 225–241. [Google Scholar] [CrossRef]
- Zou, B.; Chan, W.L.; Ding, M.; Leong, S.Y.; Nilar, S.; Seah, P.G.; Liu, W.; Karuna, R.; Blasco, F.; Yip, A.; et al. Lead Optimization of Spiropyrazolopyridones: A New and Potent Class of Dengue Virus Inhibitors. ACS Med. Chem. Lett. 2015, 6, 344–348. [Google Scholar] [CrossRef] [PubMed]
- Behnam, M.A.M.; Klein, C.D. On track to tackle dengue: History and future of NS4B ligands. Cell Host Microbe 2021, 29, 1735–1737. [Google Scholar] [CrossRef] [PubMed]
- Low, J.G.; Gatsinga, R.; Vasudevan, S.G.; Sampath, A. Dengue Antiviral Development: A Continuing Journey. In Dengue and Zika: Control and Antiviral Treatment Strategies; Hilgenfeld, R., Vasudevan, S.G., Eds.; Springer: Singapore, 2018; pp. 319–332. [Google Scholar]
- Knyazhanskaya, E.; Morais, M.C.; Choi, K.H. Chapter Ten—Flavivirus enzymes and their inhibitors. In The Enzymes; Cameron, C.E., Arnold, J.J., Kaguni, L.S., Eds.; Academic Press: Cambridge, MA, USA, 2021; pp. 265–303. [Google Scholar]
- Kang, C.; Keller, T.H.; Luo, D. Zika Virus Protease: An Antiviral Drug Target. Trends Microbiol. 2017, 25, 797–808. [Google Scholar] [CrossRef]
- Obi, J.O.; Gutiérrez-Barbosa, H.; Chua, J.V.; Deredge, D.J. Current Trends and Limitations in Dengue Antiviral Research. Trop. Med. Infect. Dis. 2021, 6, 180. [Google Scholar] [CrossRef]
- Lescar, J.; Soh, S.; Lee, L.T.; Vasudevan, S.G.; Kang, C.; Lim, S.P. The Dengue Virus Replication Complex: From RNA Replication to Protein-Protein Interactions to Evasion of Innate Immunity. Adv. Exp. Med. Biol. 2018, 1062, 115–129. [Google Scholar] [PubMed]
- Biering, S.B.; Harris, E. A step towards therapeutics for dengue. Nature 2021, 598, 420–421. [Google Scholar] [CrossRef]
- Li, Y.; Kang, C. Solution NMR Spectroscopy in Target-Based Drug Discovery. Molecules 2017, 22, 1399. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, Q.; Kang, C. Structures and Dynamics of Dengue Virus Nonstructural Membrane Proteins. Membranes 2022, 12, 231. https://doi.org/10.3390/membranes12020231
Li Q, Kang C. Structures and Dynamics of Dengue Virus Nonstructural Membrane Proteins. Membranes. 2022; 12(2):231. https://doi.org/10.3390/membranes12020231
Chicago/Turabian StyleLi, Qingxin, and Congbao Kang. 2022. "Structures and Dynamics of Dengue Virus Nonstructural Membrane Proteins" Membranes 12, no. 2: 231. https://doi.org/10.3390/membranes12020231
APA StyleLi, Q., & Kang, C. (2022). Structures and Dynamics of Dengue Virus Nonstructural Membrane Proteins. Membranes, 12(2), 231. https://doi.org/10.3390/membranes12020231





