Interconnection of Gut Microbiome and Efficacy of Immune Checkpoint Inhibitors in Inoperable Non-Small-Cell Lung Cancer
Abstract
1. Introduction
2. Results
2.1. Clinical Samples
2.2. Microbiome Analysis and Its Association with Clinical Characteristics
3. Discussion
4. Materials and Methods
4.1. Clinical Samples
4.2. 16S rRNA Gene Sequencing
4.3. Reads Processing
4.4. Statistical Analysis of α-Diversity
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ICI | Immune checkpoint inhibitor |
| NSCLC | Non-small-cell lung cancer |
| IO | Immune oncology |
| PFS | Progression-free survival |
| OS | Overall survival |
| BMI | Body mass index |
| NLR | Ratio of neutrophil to lymphocyte number |
| SIBA | Sum index of binary abundance |
| MDT | Thoracic multidisciplinary team |
| OTU | Operational taxonomic unit |
References
- Kim, M.J.; Ha, S.J. Differential Role of PD-1 Expressed by Various Immune and Tumor Cells in the Tumor Immune Microenvironment: Expression, Function, Therapeutic Efficacy, and Resistance to Cancer Immunotherapy. Front. Cell Dev. Biol. 2021, 9, 767466. [Google Scholar] [CrossRef]
- Hanahan, D. Hallmarks of Cancer: New Dimensions. Cancer Discov. 2022, 12, 31–46. [Google Scholar] [CrossRef] [PubMed]
- Johnson, D.B.; Nebhan, C.A.; Moslehi, J.J.; Balko, J.M. Immune-Checkpoint Inhibitors: Long-Term Implications of Toxicity. Nat. Rev. Clin. Oncol. 2022, 19, 254–267. [Google Scholar] [CrossRef]
- Alsaafeen, B.H.; Ali, B.R.; Elkord, E. Resistance Mechanisms to Immune Checkpoint Inhibitors: Updated Insights. Mol. Cancer 2025, 24, 20. [Google Scholar] [CrossRef]
- Thomas, A.M.; Fidelle, M.; Routy, B.; Kroemer, G.; Wargo, J.A.; Segata, N.; Zitvogel, L. Gut OncoMicrobiome Signatures (GOMS) as next-Generation Biomarkers for Cancer Immunotherapy. Nat. Rev. Clin. Oncol. 2023, 20, 583–603. [Google Scholar] [CrossRef] [PubMed]
- Routy, B.; Le Chatelier, E.; Derosa, L.; Duong, C.P.M.; Alou, M.T.; Daillère, R.; Fluckiger, A.; Messaoudene, M.; Rauber, C.; Roberti, M.P.; et al. Gut Microbiome Influences Efficacy of PD-1-Based Immunotherapy against Epithelial Tumors. Science 2018, 359, 91–97. [Google Scholar] [CrossRef]
- Peters, B.A.; Wilson, M.; Moran, U.; Pavlick, A.; Izsak, A.; Wechter, T.; Weber, J.S.; Osman, I.; Ahn, J. Relating the Gut Metagenome and Metatranscriptome to Immunotherapy Responses in Melanoma Patients. Genome Med. 2019, 11, 61. [Google Scholar] [CrossRef] [PubMed]
- Kalaora, S.; Nagler, A.; Nejman, D.; Alon, M.; Barbolin, C.; Barnea, E.; Ketelaars, S.L.C.; Cheng, K.; Vervier, K.; Shental, N.; et al. Identification of Bacteria-Derived HLA-Bound Peptides in Melanoma. Nature 2021, 592, 138–143. [Google Scholar] [CrossRef]
- Mager, L.F.; Burkhard, R.; Pett, N.; Cooke, N.C.A.; Brown, K.; Ramay, H.; Paik, S.; Stagg, J.; Groves, R.A.; Gallo, M.; et al. Microbiome-Derived Inosine Modulates Response to Checkpoint Inhibitor Immunotherapy. Science 2020, 369, 1481–1489. [Google Scholar] [CrossRef]
- Frankel, A.E.; Coughlin, L.A.; Kim, J.; Froehlich, T.W.; Xie, Y.; Frenkel, E.P.; Koh, A.Y. Metagenomic Shotgun Sequencing and Unbiased Metabolomic Profiling Identify Specific Human Gut Microbiota and Metabolites Associated with Immune Checkpoint Therapy Efficacy in Melanoma Patients. Neoplasia 2017, 19, 848–855. [Google Scholar] [CrossRef]
- Cascone, T.; William, W.N.; Weissferdt, A.; Leung, C.H.; Lin, H.Y.; Pataer, A.; Godoy, M.C.B.; Carter, B.W.; Federico, L.; Reuben, A.; et al. Neoadjuvant Nivolumab or Nivolumab plus Ipilimumab in Operable Non-Small Cell Lung Cancer: The Phase 2 Randomized NEOSTAR Trial. Nat. Med. 2021, 27, 504–514. [Google Scholar] [CrossRef]
- Derosa, L.; Routy, B.; Thomas, A.M.; Iebba, V.; Zalcman, G.; Friard, S.; Mazieres, J.; Audigier-Valette, C.; Moro-Sibilot, D.; Goldwasser, F.; et al. Intestinal Akkermansia Muciniphila Predicts Clinical Response to PD-1 Blockade in Patients with Advanced Non-Small-Cell Lung Cancer. Nat. Med. 2022, 28, 315–324. [Google Scholar] [CrossRef]
- Lee, K.A.; Thomas, A.M.; Bolte, L.A.; Björk, J.R.; de Ruijter, L.K.; Armanini, F.; Asnicar, F.; Blanco-Miguez, A.; Board, R.; Calbet-Llopart, N.; et al. Cross-Cohort Gut Microbiome Associations with Immune Checkpoint Inhibitor Response in Advanced Melanoma. Nat. Med. 2022, 28, 535–544. [Google Scholar] [CrossRef] [PubMed]
- Haberman, Y.; Kamer, I.; Amir, A.; Goldenberg, S.; Efroni, G.; Daniel-Meshulam, I.; Lobachov, A.; Daher, S.; Hadar, R.; Gantz-Sorotsky, H.; et al. Gut Microbial Signature in Lung Cancer Patients Highlights Specific Taxa as Predictors for Durable Clinical Benefit. Sci. Rep. 2023, 13, 2007. [Google Scholar] [CrossRef]
- Derosa, L.; Routy, B.; Fidelle, M.; Iebba, V.; Alla, L.; Pasolli, E.; Segata, N.; Desnoyer, A.; Pietrantonio, F.; Ferrere, G.; et al. Gut Bacteria Composition Drives Primary Resistance to Cancer Immunotherapy in Renal Cell Carcinoma Patients. Eur. Urol. 2020, 78, 195–206. [Google Scholar] [CrossRef] [PubMed]
- Peng, Z.; Cheng, S.; Kou, Y.; Wang, Z.; Jin, R.; Hu, H.; Zhang, X.; Gong, J.F.; Li, J.; Lu, M.; et al. The Gut Microbiome Is Associated with Clinical Response to Anti-PD-1/PD-L1 Immunotherapy in Gastrointestinal Cancer. Cancer Immunol. Res. 2020, 8, 1251–1261. [Google Scholar] [CrossRef] [PubMed]
- Sang, Y.; Zheng, K.; Zhao, Y.; Liu, Y.; Zhu, S.; Xie, X.; Shang, L.; Liu, J.; Li, L. Efficacy and Regulatory Strategies of Gut Microbiota in Immunotherapy: A Narrative Review. Transl. Cancer Res. 2024, 13, 2043–2063. [Google Scholar] [CrossRef]
- Porras, A.M.; Shi, Q.; Zhou, H.; Callahan, R.; Montenegro-Bethancourt, G.; Solomons, N.; Brito, I.L. Geographic Differences in Gut Microbiota Composition Impact Susceptibility to Enteric Infection. Cell Rep. 2021, 36, 4. [Google Scholar] [CrossRef]
- Dwiyanto, J.; Hussain, M.H.; Reidpath, D.; Ong, K.S.; Qasim, A.; Lee, S.W.H.; Lee, S.M.; Foo, S.C.; Chong, C.W.; Rahman, S. Ethnicity Influences the Gut Microbiota of Individuals Sharing a Geographical Location: A Cross-Sectional Study from a Middle-Income Country. Sci. Rep. 2021, 11, 2618. [Google Scholar] [CrossRef]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glöckner, F.O. Evaluation of General 16S Ribosomal RNA Gene PCR Primers for Classical and Next-Generation Sequencing-Based Diversity Studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, Interactive, Scalable and Extensible Microbiome Data Science Using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef] [PubMed]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-Resolution Sample Inference from Illumina Amplicon Data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef] [PubMed]
- Rognes, T.; Flouri, T.; Nichols, B.; Quince, C.; Mahé, F. VSEARCH: A Versatile Open Source Tool for Metagenomics. PeerJ 2016, 4, e2584. [Google Scholar] [CrossRef] [PubMed]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA Ribosomal RNA Gene Database Project: Improved Data Processing and Web-Based Tools. Nucleic Acids Res. 2013, 41, D590–D596. [Google Scholar] [CrossRef]
- Davidson-Pilon, C. Lifelines: Survival Analysis in Python. J. Open Source Softw. 2019, 4, 1317. [Google Scholar] [CrossRef]
| Benefit | |||||||
|---|---|---|---|---|---|---|---|
| No | Yes | Total | |||||
| N | % | N | % | N | % | ||
| Line of ICIs | First-line | 10 | 32.3% | 13 | 40.6% | 23 | 36.5% |
| Pembrolizumab | 6 | 60% | 9 | 69.2% | 15 | ||
| Atezolizumab | 4 | 40% | 4 | 30.8% | 8 | ||
| Second-line | 21 | 67.7% | 19 | 59.4% | 40 | 63.5% | |
| Pembrolizumab | 16 | 76.2% | 12 | 63.2% | 28 | ||
| Atezolizumab | 4 | 19.0% | 2 | 10.5% | 6 | ||
| Nivolumab Ipilimumab | 1 | 4.8% | 5 | 26.3% | 6 | ||
| Toxicity | No | 24 | 88.9% | 26 | 81.3% | 50 | 84.7% |
| Liver | 0 | 0.0% | 2 | 6.3% | 2 | 3.4% | |
| Skin | 0 | 0.0% | 1 | 3.1% | 1 | 1.7% | |
| Thyroid | 0 | 0.0% | 2 | 6.3% | 2 | 3.4% | |
| Response | N/A | 2 | 6.5% | 1 | 3.2% | 3 | 4.8% |
| Complete response | 0 | 0.0% | 1 | 3.2% | 1 | 1.6% | |
| Partial response | 3 | 9.7% | 9 | 29.0% | 12 | 19.4% | |
| Stable | 12 | 38.7% | 19 | 61.3% | 31 | 50.0% | |
| Disease | 14 | 45.2% | 1 | 3.2% | 15 | 24.2% | |
| Benefit | |||||||
|---|---|---|---|---|---|---|---|
| No | Yes | Total | |||||
| Characteristics | N | % | N | % | N | % | p-Value |
| Gender | 0.018 | ||||||
| Male | 28 | 90.3% | 21 | 65.6% | 49 | 77.8% | |
| Female | 3 | 9.7% | 11 | 34.4% | 14 | 22.2% | |
| Age | 0.916 | ||||||
| <65 | 13 | 41.9% | 13 | 40.6% | 26 | 41.3% | |
| ≥65 | 18 | 58.1% | 19 | 59.4% | 37 | 58.7% | |
| BMI ≥ 29 | 0.591 | ||||||
| No | 18 | 58.1% | 16 | 50.0% | 34 | 54.0% | |
| Yes | 11 | 35.5% | 15 | 46.9% | 26 | 41.3% | |
| NA | 2 | 6.5% | 1 | 3.1% | 3 | 4.8% | |
| NLR ≥ T3 | 0.124 | ||||||
| No | 24 | 77.4% | 19 | 59.4% | 43 | 68.3% | |
| Yes | 7 | 22.6% | 13 | 40.6% | 20 | 31.7% | |
| NLR ≥ T3 and BMI ≥ 29 | 0.758 | ||||||
| No | 27 | 87.1% | 27 | 84.4% | 54 | 85.7% | |
| Yes | 4 | 12.9% | 5 | 15.6% | 9 | 14.3% | |
| Histology | 0.259 | ||||||
| Adenocarcinoma | 14 | 45.2% | 19 | 59.4% | 33 | 52.4% | |
| Squamous cell | 17 | 54.8% | 13 | 40.6% | 30 | 47.6% | |
| Activating mutations | 0.721 | ||||||
| None | 28 | 90.3% | 29 | 90.6% | 57 | 90.5% | |
| EGFR | 1 | 3.2% | 1 | 3.1% | 2 | 3.2% | |
| ALK/ROS1 | 1 | 3.2% | 0 | 0.0% | 1 | 1.6% | |
| KRAS | 1 | 3.2% | 2 | 6.3% | 3 | 4.8% | |
| PD-L1 staining | 0.007 | ||||||
| <50% | 27 | 87.1% | 18 | 56.3% | 45 | 71.4% | |
| >50% | 4 | 12.9% | 14 | 43.8% | 18 | 28.6% | |
| Clinical Characteristic | Coverage Depth Threshold | Chao1 | Shannon | Simpson |
|---|---|---|---|---|
| Benefit | 1000–8000 | 207 vs. 147 (0.0078) | 5.69 vs. 5.44 (0.041) | 0.958 vs. 0.947 (0.039) |
| Age ≥ 72 | 3000–7000 | 136 vs. 186 (0.0285) | 5.44 vs. 5.57 (0.232) | 0.953 vs. 0.955 (0.282) |
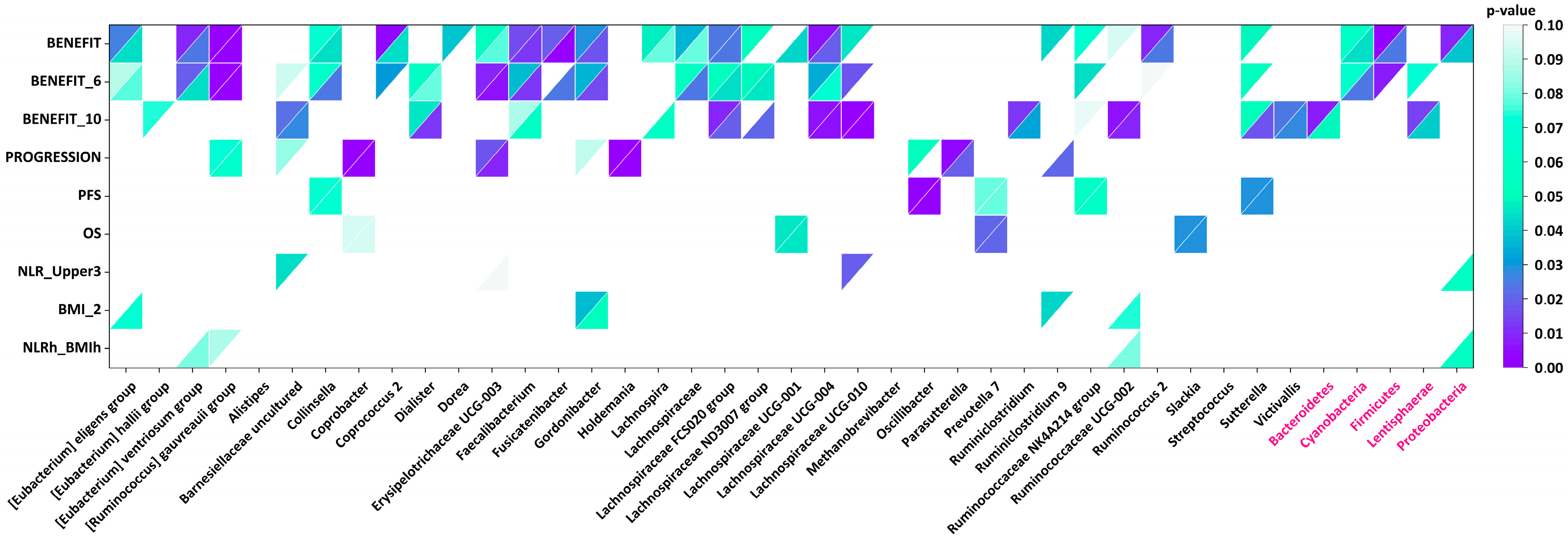
| Taxon | Bacterial Genus/Group/Phylum | Associated with | Abundance Threshold |
|---|---|---|---|
| Genus | [Eubacterium] eligens group | Benefit | 0.01 |
| Genus | [Eubacterium] hallii group | Progression | 0.002 |
| Genus | [Eubacterium] ventriosum group | Benefit | 0.0001 |
| Genus | [Ruminococcus] gauvreauii group | Benefit | 0.0001 |
| Genus | Alistipes | Benefit | 0.05 |
| Phylum | Bacteroidetes | Benefit | 0.36 |
| Genus | Uncultured genus from Barnesiellaceae family | Benefit | 0.00007 |
| Genus | Collinsella | Benefit | 0.009 |
| Genus | Coprobacter | Progression | 0.0002 |
| Genus | Coprococcus 2 | Benefit | 0.0001 |
| Phylum | Cyanobacteria | Benefit | 0.0001 |
| Genus | Dialister | Benefit | 0.01 |
| Genus | Dorea | Benefit | 0.0001 |
| Genus | Erysipelotrichaceae UCG-003 | Progression | 0.0008 |
| Genus | Faecalibacterium | Benefit | 0.03 |
| Phylum | Firmicutes | Benefit | 0.4 |
| Genus | Fusicatenibacter | Benefit | 0.002 |
| Genus | Gordonibacter | Benefit | 0.01 |
| Genus | Holdemania | Progression | 0.00006 |
| Genus | Lachnospira | Benefit | 0.0001 |
| Genus | Lachnospiraceae | Benefit | 0.006 |
| Genus | Lachnospiraceae FCS020 group | Benefit | 0.0001 |
| Genus | Lachnospiraceae ND3007 group | Benefit | 0.001 |
| Genus | Lachnospiraceae UCG-001 | Benefit | 0.001 |
| Genus | Lachnospiraceae UCG-004 | Benefit | 0.0001 |
| Genus | Lachnospiraceae UCG-010 | Benefit | 0.00004 |
| Phylum | Lentisphaerae | Benefit | 0.0003 |
| Genus | Methanobrevibacter | Benefit | 0.005 |
| Genus | Oscillibacter | Progression | 0.002 |
| Genus | Parasutterella | Progression | 0.0001 |
| Genus | Prevotella 7 | Progression | 0.001 |
| Phylum | Proteobacteria | Progression | 0.05 |
| Genus | Ruminiclostridium | Benefit | 0.00004 |
| Genus | Ruminiclostridium 9 | Benefit | 0.0001 |
| Genus | Ruminococcaceae NK4A214 group | Benefit | 0.002 |
| Genus | Ruminococcaceae UCG-002 | Benefit | 0.01 |
| Genus | Ruminococcus 2 | Benefit | 0.01 |
| Genus | Slackia | Progression | 0.002 |
| Genus | Streptococcus | Progression | 0.006 |
| Genus | Sutterella | Benefit | 0.002 |
| Genus | Victivallis | Benefit | 0.00004 |
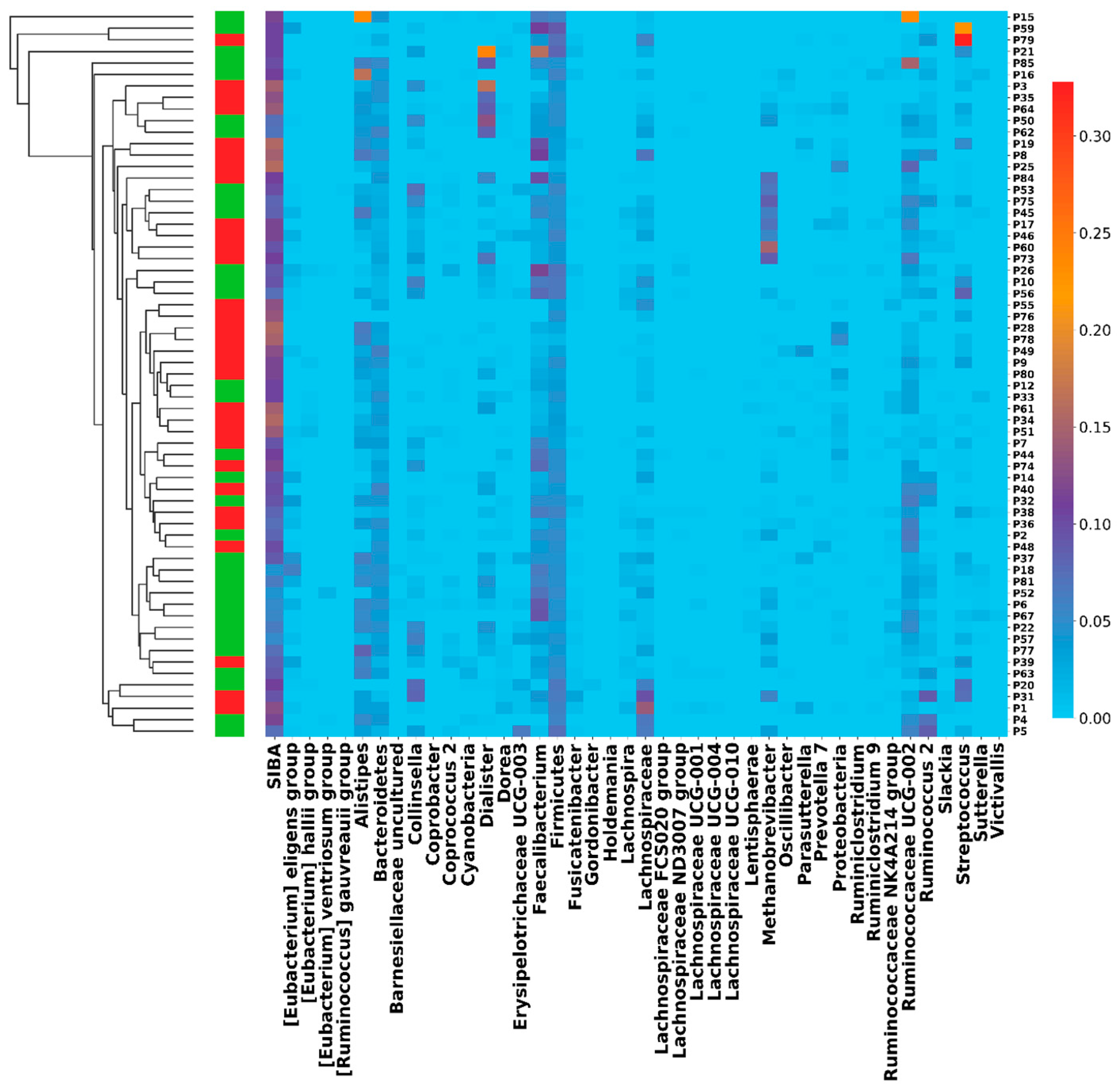
| Characteristics | Progression | Benefit | BMI ≥ 29 | NLR > T3 | BMIh_NLRh | PFS | OS |
|---|---|---|---|---|---|---|---|
| Median SIBA (Yes) | 21.0 | 18.0 | 19.5 | 19.5 | 19.0 | ||
| Median SIBA (No) | 19.5 | 24.0 | 21.0 | 21.0 | 21.0 | ||
| Mann–Whitney’s p-value | 0.70 | 6 × 10−7 | 0.87 | 0.40 | 0.84 | ||
| Threshold for the Fisher’s exact test/log-rank test | 20.25 | 21 | 20.25 | 20.25 | 20 | 23 | 23 |
| Fisher’ exact test/log-rank test p-value | 0.31 | 9 × 10−7 | 0.61 | 0.29 | 0.48 | 6 × 10−9 | 0.01 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Moiseenko, F.; Kechin, A.; Koryukov, M.; Boyarskikh, U.; Gabina, A.; Oganesian, A.; Belukhin, S.; Makarkina, M.; Elsakova, E.; Artemeva, E.; et al. Interconnection of Gut Microbiome and Efficacy of Immune Checkpoint Inhibitors in Inoperable Non-Small-Cell Lung Cancer. Int. J. Mol. Sci. 2025, 26, 7758. https://doi.org/10.3390/ijms26167758
Moiseenko F, Kechin A, Koryukov M, Boyarskikh U, Gabina A, Oganesian A, Belukhin S, Makarkina M, Elsakova E, Artemeva E, et al. Interconnection of Gut Microbiome and Efficacy of Immune Checkpoint Inhibitors in Inoperable Non-Small-Cell Lung Cancer. International Journal of Molecular Sciences. 2025; 26(16):7758. https://doi.org/10.3390/ijms26167758
Chicago/Turabian StyleMoiseenko, Fedor, Andrey Kechin, Maksim Koryukov, Ulyana Boyarskikh, Albina Gabina, Ani Oganesian, Sergey Belukhin, Maria Makarkina, Ekaterina Elsakova, Elizaveta Artemeva, and et al. 2025. "Interconnection of Gut Microbiome and Efficacy of Immune Checkpoint Inhibitors in Inoperable Non-Small-Cell Lung Cancer" International Journal of Molecular Sciences 26, no. 16: 7758. https://doi.org/10.3390/ijms26167758
APA StyleMoiseenko, F., Kechin, A., Koryukov, M., Boyarskikh, U., Gabina, A., Oganesian, A., Belukhin, S., Makarkina, M., Elsakova, E., Artemeva, E., Myslik, A., Volkov, N., Bogdanov, A., Kuligina, E., Aleksakhina, S., Iyevleva, A., Ivantsov, A., Bogdanov, A., Sidorenko, S., ... Moiseyenko, V. (2025). Interconnection of Gut Microbiome and Efficacy of Immune Checkpoint Inhibitors in Inoperable Non-Small-Cell Lung Cancer. International Journal of Molecular Sciences, 26(16), 7758. https://doi.org/10.3390/ijms26167758







