Visualizing the Machine Learning Process in Multichannel Time Series Classification
Abstract
1. Introduction
2. Related Work
3. Materials and Methods
3.1. Classification Algorithms
3.1.1. The Random Convolutional Kernel Transform (ROCKET)
3.1.2. Residual Network (ResNet)
3.1.3. InceptionTime
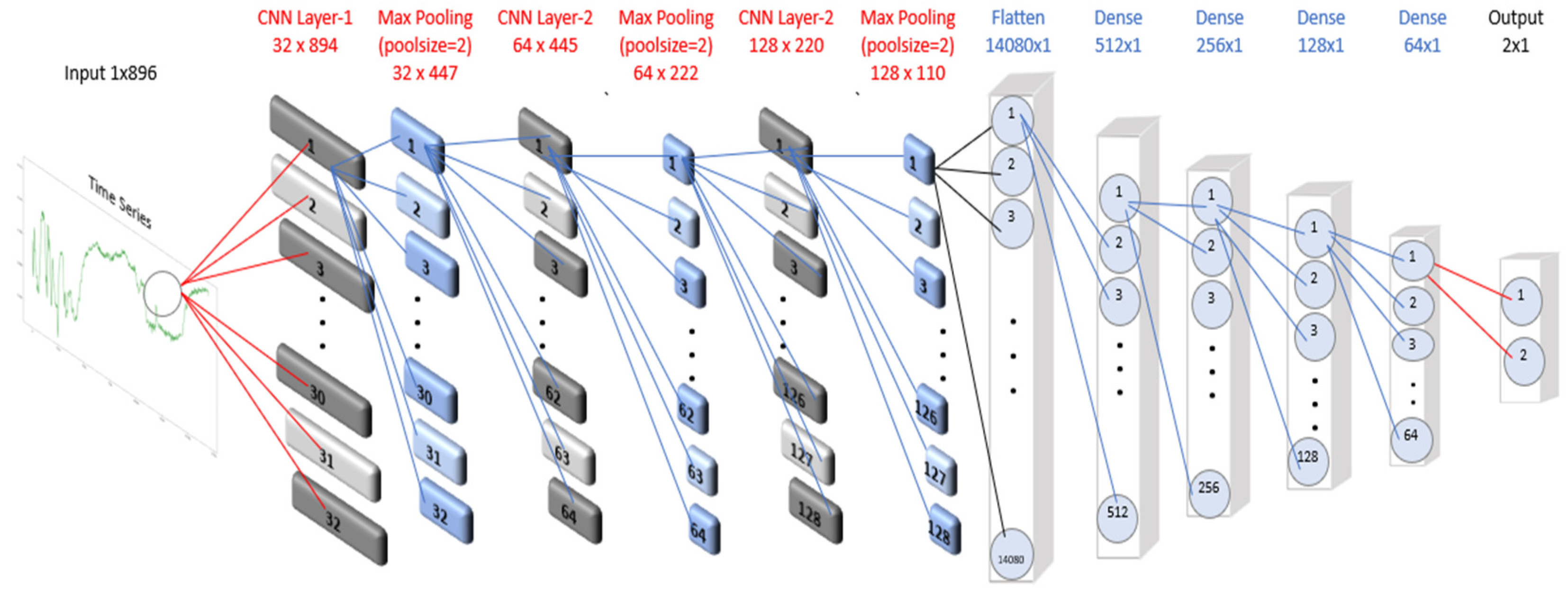
3.1.4. One Dimensional Convolutional Neural Network (CNN-1D)
3.1.5. LSTM
3.1.6. Transformers
3.2. Datasets
3.3. Research Workflow
4. Results
4.1. Classifiers Accuracy Results
4.2. Discussion of Individual Dataset Results
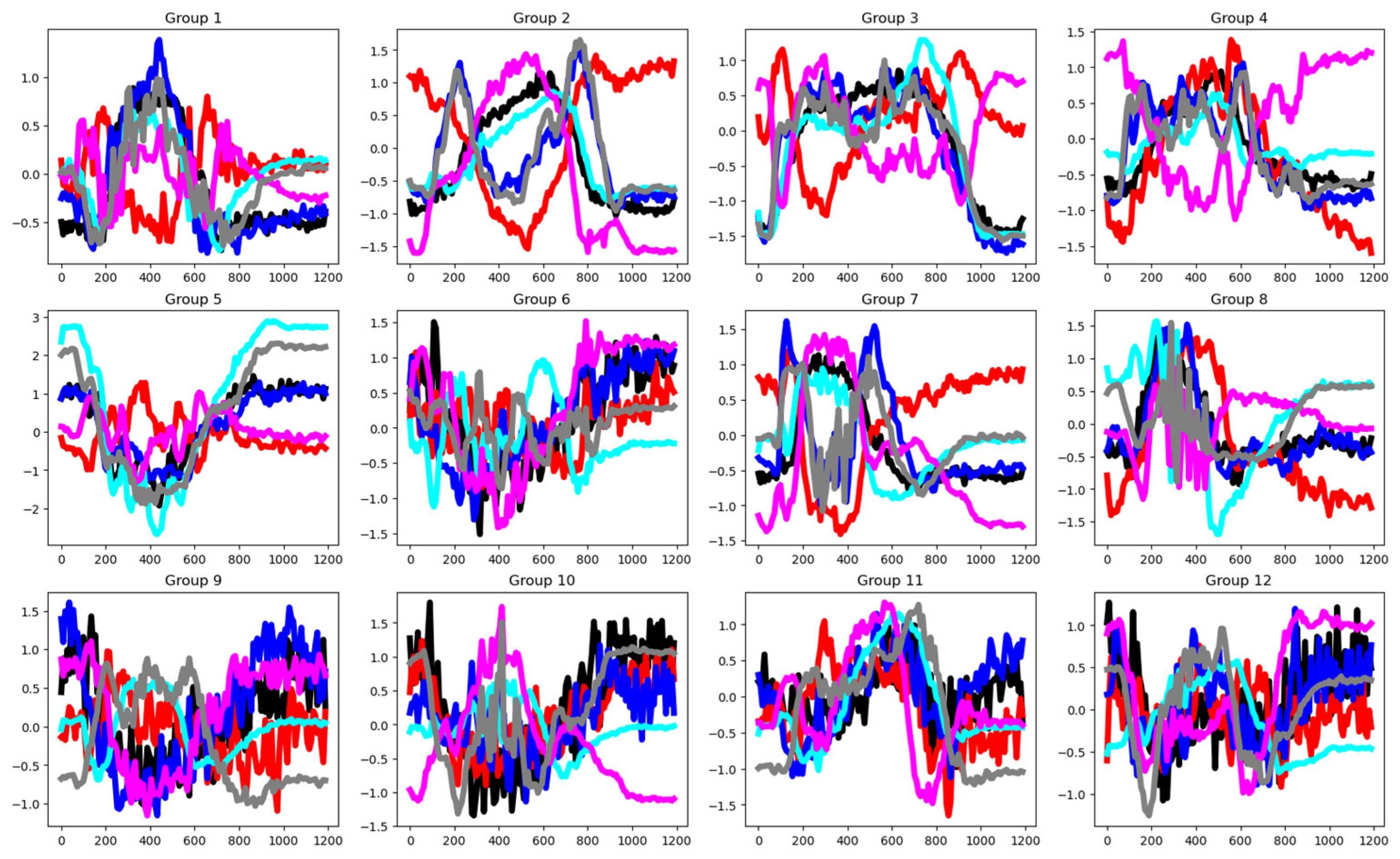
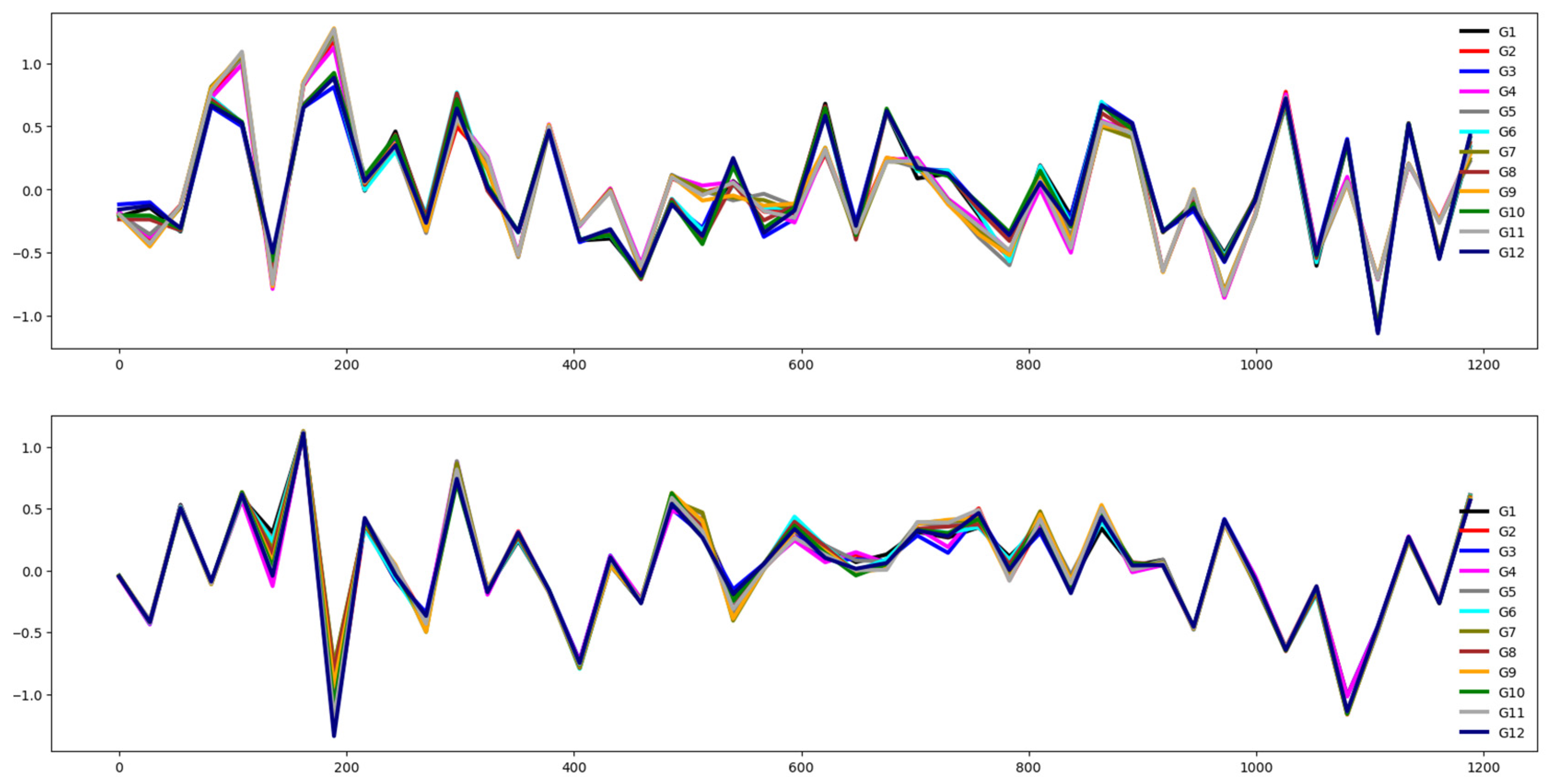
4.2.1. Cricket (CRI)
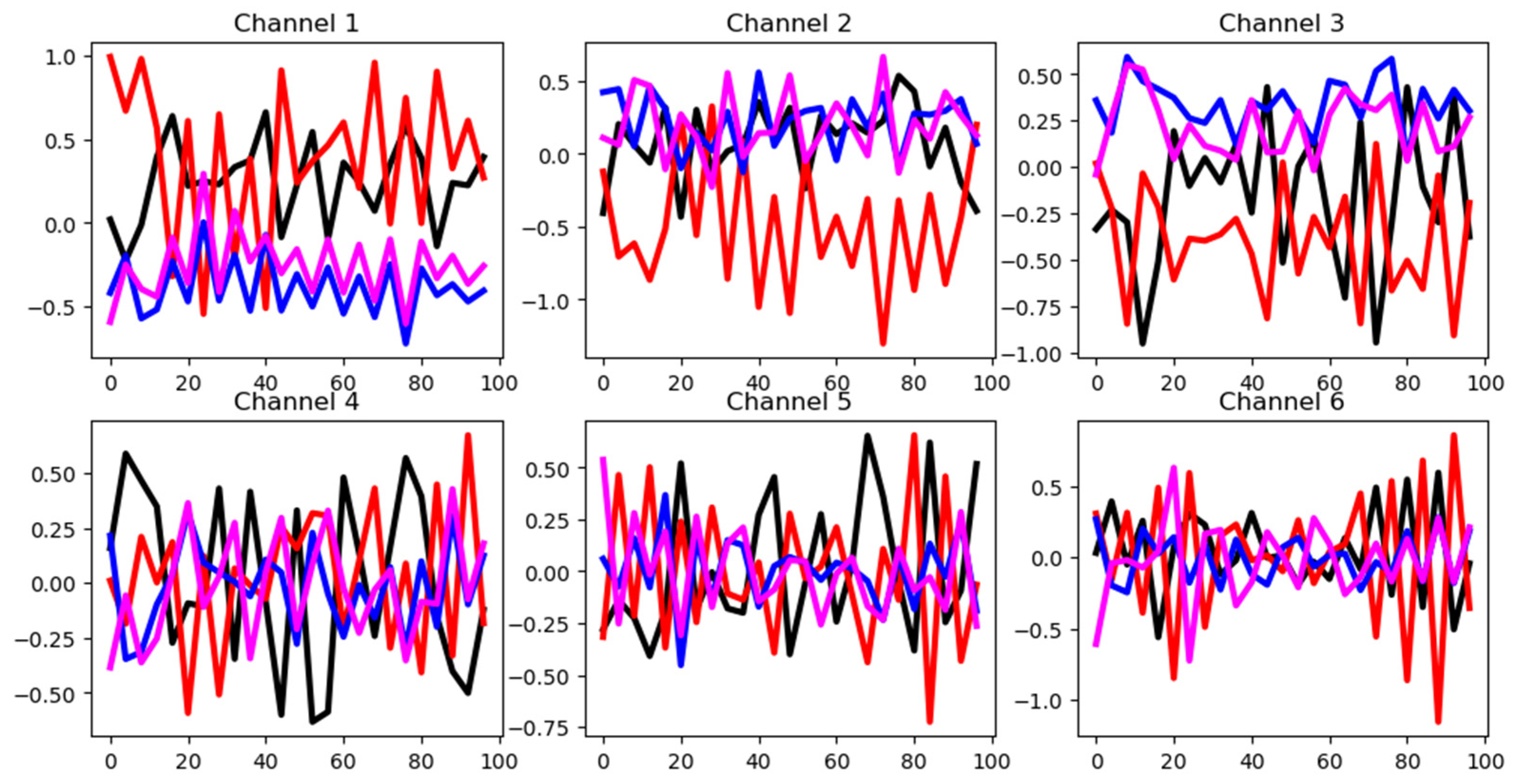
4.2.2. Basic Motions (BM)
4.2.3. Epilepsy (EPI)
4.2.4. NATOPS
4.2.5. Articulacy Word Recognition (AWR)
4.2.6. Racket Sports (RS)
4.2.7. PEMS-SF
4.2.8. SelfRegulationSCP1 (SCP1)
4.2.9. Libras (LIB)
4.2.10. Heartbeat (HB)
4.2.11. Face Detection (FD)
4.2.12. Duck Duck Geese (DDG)
4.2.13. SelfRegulationSCP2 (SCP2)
4.2.14. Finger Movements (FM)
4.2.15. Hand Movement Direction (HMD)
4.2.16. Ethanol Concentration (EC)
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Appendix A
| Dataset Name | Abbreviation | Brief Description |
|---|---|---|
| Cricket | CRI | Wrist accelerometer data capturing umpire hand signals in cricket. |
| Basic Motions | BM | Smartwatch sensor data of basic physical activities (walk, run, rest, badminton). |
| Epilepsy | EPI | Wrist accelerometer recordings of simulated epileptic seizures and activities. |
| NATOPS | NATOPS | Kinect-based 3D motion data of standardized aircraft handling gestures. |
| Articulacy Word Recognition | AWR | Electromagnetic articulography data of tongue and lip movements during speech. |
| Racket Sports | RS | Smartwatch sensor data of badminton and squash strokes. |
| PEMS-SF | PEMS-SF | Freeway sensor time series for classifying day-of-week traffic patterns. |
| Self-Regulation SCP1 | SCP | EEG slow cortical potentials from motor imagery cursor control tasks. |
| Libras | LIB | Hand movement trajectories from Brazilian sign language gestures. |
| Heartbeat | HB | Heart sound recordings classified as normal or abnormal. |
| Face Detection | FD | MEG signals distinguishing face versus scrambled visual stimuli. |
| Duck Duck Geese | DDG | Audio spectrograms of bird species vocalizations. |
| Self-Regulation SCP2 | SCP2 | EEG slow cortical potentials from cursor control by an ALS patient. |
| Finger Movements | FM | Wrist accelerometer data capturing umpire hand signals in cricket. |
| Hand Movement Direction | HMD | Smartwatch sensor data of basic physical activities (walk, run, rest, badminton). |
| Ethanol Concentration. | EC | Spectral data classifying ethanol concentration in whisky samples. |
References
- Bagnall, A.; Lines, J.; Bostrom, A.; Large, J.; Keogh, E. The great time series classification bake off: A review and experimental evaluation of recent algorithmic advances. Data Min. Knowl. Discov. 2017, 31, 606–660. [Google Scholar] [CrossRef]
- Bagnall, A.; Dau, H.A.; Lines, J.; Flynn, M.; Large, J.; Bostrom, A.; Southam, P.; Keogh, E. The UEA multivariate time series classification archive. arXiv 2018, arXiv:1811.00075. [Google Scholar] [CrossRef]
- Fawaz, H.; Forestier, G.; Weber, J.; Idoumghar, L.; Muller, P.A. Deep learning for time series classification: A review. Data Min. Knowl. Discov. 2019, 33, 917–963. [Google Scholar] [CrossRef]
- Pasos Ruiz, A.; Flynn, M.; Large, J.; Middlehurst, M.; Bagnall, A. The great multivariate time series classification bake off: A review and experimental evaluation of recent algorithmic advances. Data Min. Knowl. Discov. 2021, 35, 401–449. [Google Scholar] [CrossRef]
- Baldan, F.; Benítez, J. Multivariable time series classification through an interpretable representation. arXiv 2020, arXiv:2009.03614. [Google Scholar] [CrossRef]
- Pasos Ruiz, A.; Bagnall, A. Dimension selection strategies for multivariate time series classification with HIVE-COTE v2.0. In Advanced Analytics and Learning on Temporal Data; Lecture Notes in Computer Science; Springer: Berlin/Heidelberg, Germany, 2023; pp. 133–147. [Google Scholar]
- Ilbert, R.; Huang, T.V.; Zhang, Z. Data augmentation for multivariate time series classification: An experimental study. In IEEE 40th International Conference on Data Engineering Workshops (ICDEW), Utrecht, The Netherlands, 13–16 May 2024; IEEE: New York, NY, USA, 2024. [Google Scholar]
- Dempster, A.; Petitjean, F.; Webb, G. ROCKET: Exceptionally fast and accurate time series classification using random convolutional kernels. Data Min. Knowl. Discov. 2020, 34, 1454–1495. [Google Scholar] [CrossRef]
- Löning, M.; Bagnall, A.; Ganesh, S.; Kazakov, V.; Király, F.J. sktime: A unified interface for machine learning with time series. arXiv 2019, arXiv:1909.07872. [Google Scholar] [CrossRef]
- Wang, J.; Balasubramanian, A.; de La Vega, L.; Green, J.R.; Samal, A.; Prabhakaran, B. Word recognition from continuous articulatory movement time-series data using symbolic representations. In Proceedings of the Fourth Workshop on Speech and Language Processing for Assistive Technologies, Grenoble, France, 21–22 August 2013; Association for Computational Linguistics: Stroudsburg, PA, USA, 2013; pp. 119–127. [Google Scholar]
- Szegedy, C.; Liu, W.; Jia, Y.; Sermanet, P.; Reed, S.; Anguelov, D.; Erhan, D.; Vanhoucke, V.; Rabinovich, A. Going deeper with convolutions. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 1–9. [Google Scholar] [CrossRef]
- Fawaz, H.; Lucas, B.; Forestier, G.; Pelletier, C.; Schmidt, D.F.; Weber, J.; Webb, G.; Idoumghar, L.; Muller, P.A.; Petitjean, F. InceptionTime: Finding AlexNet for time series classification. Data Min. Knowl. Discov. 2020, 34, 1936–1962. [Google Scholar] [CrossRef]
- Khan, A.; Sohail, A.; Zahoora, U.; Qureshi, A.Q. A survey of the recent architectures of deep convolutional neural networks. Artif. Intell. Rev. 2020, 53, 5455–5516. [Google Scholar] [CrossRef]
- Kiranyaz, S.; Avci, O.; Abdeljaber, O.; Ince, T.; Gabbouj, M.; Inman, D.J. 1D convolutional neural networks and applications: A survey. Mech. Syst. Signal Process. 2021, 151, 107398. [Google Scholar] [CrossRef]
- Acuna, E.; Kendziora, C.; Fustenberg, R.; Breshike, C.J.; Kendziora, D. Machine learning algorithms for analytes classification based on simulated spectra. In Proceedings of the SPIE 13031, Algorithms, Technologies, and Applications for Multispectral and Hyperspectral Imaging XXX, National Harbor, MA, USA, 21–25 April 2024. [Google Scholar] [CrossRef]
- Sak, H.; Senior, A.; Beaufays, F. Long short-term memory recurrent neural network architectures for large scale acoustic modeling. In Proceedings of the 15th Annual Conference of the International Speech Communication Association, INTERSPEECH-2014, Singapore, 14–18 September 2014; ISCA: Singapore, 2014; pp. 338–342. [Google Scholar]
- Cura, A.; Kucuk, H.; Ergen, E.; Oksuzoglu, I.B. Driver profiling using long short term memory (LSTM) and convolutional neural network (CNN) methods. IEEE Trans. Intell. Transp. Syst. 2020, 22, 6572–6582. [Google Scholar] [CrossRef]
- Xu, G.; Ren, T.; Chen, Y.; Che, W. A one-dimensional CNN-LSTM model for epileptic seizure recognition using EEG. Front. Neurosci. 2020, 14, 578126. [Google Scholar] [CrossRef]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, Ł.; Polosukhin, I. Attention is all you need. In Advances in Neural Information Processing Systems, Proceedings of the 31st International Conference on Neural Information Processing Systems, Red Hook, NY, USA, 4–9 December 2017; Curran Associates: Red Hook, NY, USA, 2017; Volume 30. [Google Scholar]
- Qingsong, W.; Tian, Z.; Chaoli, Z.; Weiqi, C.; Ziqing, M.; Junchi, Y.; Liang, S. Transformers in time series: A survey. In Proceedings of the Thirty-Second International Joint Conference on Artificial Intelligence (IJCAI-23), Macao, China, 19–25 August 2023; Available online: https://github.com/qingsongedu/time-series-transformers-review (accessed on 1 June 2025).
- Zerveas, G.; Jayaraman, S.; Patel, D.; Bhamidipaty, A.; Eickhoff, C. A transformer-based framework for multivariate time series representation learning. In Proceedings of the 27th ACM SIGKDD Conference on Knowledge Discovery & Data Mining, Singapore, 14–18 August 2021; ACM: New York, NY, USA, 2021; pp. 2114–2124. [Google Scholar] [CrossRef]
- Wang, Z.; Zhang, J.; Zhang, X.; Chen, P.; Wang, B. Transformer model for functional near-infrared spectroscopy classification. IEEE J. Biomed. Health Inform. 2022, 26, 2609–2619. [Google Scholar] [CrossRef] [PubMed]
- Ko, M.H.; West, G.; Venkatesh, S.; Kumar, M. Context recognition in multisensor systems using dynamic time warping. In International Conference on Intelligent Sensors 2005, Sensor Networks and Information Processing, Melbourne, VIC, Australia, 5–8 December 2005; IEEE: New York, NY, USA, 2005; pp. 283–288. [Google Scholar] [CrossRef]
- Villar, J.R.; Vergara, P.; Menendez, M.; de la Cal, E.; Gonzalez, V.; Sedano, J. Generalized models for the classification of abnormal movements in daily life and its applicability to epilepsy convulsion recognition. Int. J. Neural Syst. 2016, 26, 1650037. [Google Scholar] [CrossRef] [PubMed]
- Ghouaiel, N.; Marteau, P.; Dupont, M. Continuous pattern detection and recognition in stream—A benchmark for online gesture recognition. Int. J. Appl. Pattern Recognit. 2017, 4, 146–160. [Google Scholar] [CrossRef]
- Cuturi, M. Fast global alignment kernels. In Proceedings of the 28th International Conference on Machine Learning (ICML-11), Washington, DC, USA, 28 June–2 July 2011; pp. 929–936. [Google Scholar]
- Birbaumer, N.; Ghanayim, N.; Hinterberger, T.; Iversen, I.; Kotchoubey, B.; Kubler, A.; Perelmouter, J.; Taub, E.; Flor, H. A spelling device for the paralysed. Nature 1999, 398, 297. [Google Scholar] [CrossRef] [PubMed]
- Dias, D.; Peres, S. Algoritmos bio-inspirados aplicados ao reconhecimento de padroes da libras: Enfoque no parâmetro movimento. In Proceedings of the 16 Simpósio Internacional de Iniciaçao Cientıfica da Universidade de Sao Paulo, São Paulo, Brasil, 26–31 October 2008. [Google Scholar]
- Goldberger, A.; Amaral, L.; Glass, L.; Hausdorff, J.; Ivanov, P.; Mark, R.; Mietus, J.; Moody, G.; Peng, C.K.; Stanley, E. Physiobank, physiotoolkit, and physionet: Components of a new research resource for complex physiologic signals. Circulation 2000, 101, e215–e220. [Google Scholar] [CrossRef] [PubMed]
- Olivetti, R.; Kia, M.; Avesani, P. DecMeg2014—Decoding the Human Brain. Kaggle. 2014. Available online: https://www.kaggle.com/competitions/decoding-the-human-brain (accessed on 1 June 2025).
- Xeno-Canto. Sharing Wildlife Sounds from Around the World. Repository. Available online: https://xeno-canto.org/ (accessed on 1 June 2025).
- Blankertz, B.; Curio, G.; Müller, K.R. Classifying single trial EEG: Towards brain computer interfacing. In Advances in Neural Information Processing Systems; Curran Associates, Inc.: Red Hook, NY, USA, 2002; pp. 157–164. [Google Scholar]
- Large, J.; Kemsley, E.K.; Wellner, N.; Goodall, I.; Bagnall, A. Detecting forged alcohol non-invasively through vibrational spectroscopy and machine learning. In Proceedings of the Pacific-Asia Conference on Knowledge Discovery and Data Mining, Melbourne, VIC, Australia, 3–6 June 2018; Springer: Berlin/Heidelberg, Germany, 2025; pp. 298–309. [Google Scholar] [CrossRef]
- Dhariyal, B.; Nguyen, T.L.; Ifrim, G. Fast channel selection for scalable multivariate time series classification. In Advanced Analytics and Learning on Temporal Data (AALTD 2021); Lemaire, V., Malinowski, S., Bagnall, A., Guyet, T., Tavenard, R., Ifrim, G., Eds.; Lecture Notes in Computer Science; Springer: Berlin/Heidelberg, Germany, 2021; Volume 13114. [Google Scholar] [CrossRef]
- Meert, W.; Hendrickx, K.; Van Craenendonck, T.; Robberechts, P.; Blockeel, H.; Davis, J. DTAIDistance, Version v2; Zenodo: Geneva, Switzerland, 2025. [CrossRef]












| Dataset | Train | Test | Channels (Ch) | Length | Groups | Train × Ch | Train × Ch/L | Acc. Default % |
|---|---|---|---|---|---|---|---|---|
| CRI [23] | 108 | 72 | 6 | 1197 | 12 | 648 | 0.54 | 8.33 |
| BM [2] | 40 | 40 | 6 | 100 | 4 | 240 | 2.40 | 25.00 |
| EPI [24] | 137 | 138 | 3 | 206 | 4 | 411 | 1.99 | 26.80 |
| NATOPS [25] | 180 | 180 | 24 | 51 | 6 | 4320 | 84.70 | 26.81 |
| AWR [10] | 275 | 300 | 9 | 144 | 25 | 2475 | 17.18 | 4.00 |
| RS [2] | 151 | 152 | 6 | 30 | 4 | 906 | 30.20 | 28.30 |
| PEMS-SF [26] | 267 | 173 | 963 | 144 | 7 | 257,121 | 1786.00 | 17.34 |
| SCP1 [27] | 268 | 293 | 6 | 896 | 2 | 1608 | 1.79 | 50.20 |
| LIB [28] | 180 | 180 | 2 | 45 | 15 | 360 | 8.00 | 6.70 |
| HB [29] | 204 | 205 | 61 | 405 | 2 | 12,444 | 30.70 | 72.19 |
| FD [30] | 5890 | 3524 | 144 | 62 | 2 | 848,160 | 13,680.00 | 50.00 |
| DDG [31] | 50 | 50 | 1345 | 270 | 5 | 67,250 | 249.00 | 20.00 |
| SCP2 [27] | 200 | 180 | 7 | 1152 | 2 | 1400 | 1.22 | 50.00 |
| FM [32] | 316 | 100 | 28 | 50 | 2 | 8848 | 177.00 | 51.00 |
| HMD [30] | 160 | 74 | 10 | 400 | 4 | 1600 | 4.00 | 40.54 |
| EC [33] | 261 | 263 | 3 | 1751 | 4 | 783 | 0.44 | 25.09 |
| Dataset | LSTM | CNN-1D | Transformer | ResNet | Inception Time | ROCKET |
|---|---|---|---|---|---|---|
| CRI | 93.20 ± 4.96 | 94.10 ± 2.18 | 69.00 ± 7.15 | 98.84 ± 0.57 | 98.41 ± 0.96 | 100.00 ± 0.00 |
| BM | 97.00 ± 1.82 | 100.00 ± 0.00 | 82.85 ± 6.98 | 97.00 ± 6.70 | 55.20 ± 2.82 | 100.00 ± 0.00 |
| EPI | 82.53 ± 1.79 | 83.48 ± 1.44 | 76.45 ± 3.33 | 94.05 ± 4.30 | 93.62 ± 5.11 | 99.20 ± 0.79 |
| NATOPS | 83.99 ± 1.69 | 91.67 ± 1.94 | 77.07 ± 5.09 | 92.88 ± 2.68 | 94.27 ± 1.14 | 88.32 ± 0.64 |
| AWR | 87.92 ± 2.09 | 92.93 ± 1.26 | 48.78 ± 3.19 | 93.36 ± 5.70 | 97.56 ± 2.22 | 99.29 ± 0.18 |
| RS | 75.80 ± 1.98 | 82.00 ± 2.89 | 66.00 ± 4.26 | 88.68 ± 2.0 | 88.28 ± 1.35 | 91.17 ± 0.59 |
| PEMS-SF | 89.70 ± 2.22 | 88.55 ± 4.27 | 78.09 ± 2.78 | 73.80 ± 6.95 | 73.86 ± 6.21 | 81.26 ± 1.44 |
| SCP1 | 77.74 ± 1.51 | 85.60 ± 2.11 | 75.63 ± 2.91 | 73.03 ± 6.40 | 76.79 ± 8.80 | 84.88 ± 1.02 |
| LIB | 74.60 ± 6.27 | 79.50 ± 2.27 | 54.50 ± 3.17 | 81.66 ± 10.51 | 87.44 ± 0.63 | 90.55 ± 0.39 |
| HB | 71.70 ± 1.21 | 73.56 ± 1.30 | 73.72 ± 2.13 | 57.56 ± 9.77 | 67.36 ± 9.45 | 74.48 ± 0.94 |
| FD | 63.62 ± 0.79 | 61.14 ± 0.46 | 61.38 ± 1.14 | 55.26 ± 1.14 | 65.92 ± 0.94 | 58.67 ± 0.55 |
| DDG | 51.80 ± 5.62 | 62.00 ± 4.42 | 42.60 ± 6.80 | 59.80 ± 4.46 | 61.20 ± 2.35 | 49.40 ± 3.27 |
| SCP2 | 53.93 ± 2.67 | 53.60 ± 2.01 | 51.94 ± 2.80 | 51.10 ± 1.75 | 52.77 ± 2.79 | 55.16 ± 2.06 |
| FM | 51.79 ± 3.18 | 51.80 ± 2.09 | 50.40 ± 3.02 | 53.60 ± 4.14 | 55.20 ± 2.82 | 55.10 ± 1.28 |
| HMD | 38.11 ± 8.14 | 52.30 ± 2.40 | 36.10 ± 4.35 | 30.40 ± 3.89 | 40.80 ± 1.89 | 50.89 ± 3.57 |
| EC | 26.92 ± 1.90 | 41.50 ± 10.36 | 27.97 ± 2.35 | 27.63 ± 2.18 | 28.09 ± 2.79 | 40.83 ± 1.88 |
| Dataset | % of Relevant Channels | % of Relevant Features | Min Eucl. Dist_Group | Min DTW Dist_Group | Mean DTW Dist Train_Test | |
|---|---|---|---|---|---|---|
| F-Test | MI | |||||
| CRI | 100.00 | 2.59 | 7.60 | 1.43 | 1.40 | 7.90 |
| BM | 50.00 | 9.00 | 9.00 | 1.51 | 0.92 | 0.76 |
| EPI | 33.33 | 0.48 | 7.28 | 1.60 | 1.46 | 3.12 |
| NATOPS | 70.83 | 5.88 | 17.64 | 0.37 | 0.36 | 0.37 |
| AWR | 100.00 | 0.69 | 11.81 | 0.66 | 0.52 | 0.57 |
| RS | 66.67 | 3.33 | 46.66 | 0.44 | 0.31 | 0.30 |
| PEMS-SF | 33.02 | 4.86 | 14.58 | 0.05 | 0.05 | 1.45 |
| SCP1 | 50.00 | 0.55 | 1.34 | 2.47 | 2.45 | 22.15 |
| LIB | 100.00 | 20.00 | 37.77 | 1.59 | 1.37 | 3.51 |
| HB | 14.75 | 0.49 | 11.85 | 0.37 | 0.37 | 12.92 |
| FD | 8.33 | 14.52 | 19.35 | 0.02 | 0.01 | 0.05 |
| DDG | 28.77 | 1.11 | 10.00 | 0.20 | 0.20 | 8.76 |
| SCP2 | 71.40 | 0.17 | 0.00 | 2.19 | 2.18 | 18.14 |
| FM | 21.42 | 12.00 | 14.00 | 0.06 | 0.06 | 3.06 |
| HMD | 20.00 | 8.25 | 13.00 | 4.34 | 3.75 | 5.05 |
| EC | 100.00 | 2.85 | 21.01 | 3.60 | 3.58 | 21.91 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Acuña, E.; Aparicio, R. Visualizing the Machine Learning Process in Multichannel Time Series Classification. Analytics 2026, 5, 15. https://doi.org/10.3390/analytics5010015
Acuña E, Aparicio R. Visualizing the Machine Learning Process in Multichannel Time Series Classification. Analytics. 2026; 5(1):15. https://doi.org/10.3390/analytics5010015
Chicago/Turabian StyleAcuña, Edgar, and Roxana Aparicio. 2026. "Visualizing the Machine Learning Process in Multichannel Time Series Classification" Analytics 5, no. 1: 15. https://doi.org/10.3390/analytics5010015
APA StyleAcuña, E., & Aparicio, R. (2026). Visualizing the Machine Learning Process in Multichannel Time Series Classification. Analytics, 5(1), 15. https://doi.org/10.3390/analytics5010015





