Physiological and Genomic Characterization of Two Novel Bacteroidota Strains Asinibacterium spp. OR43 and OR53
Abstract
:1. Introduction
2. Results and Discussion
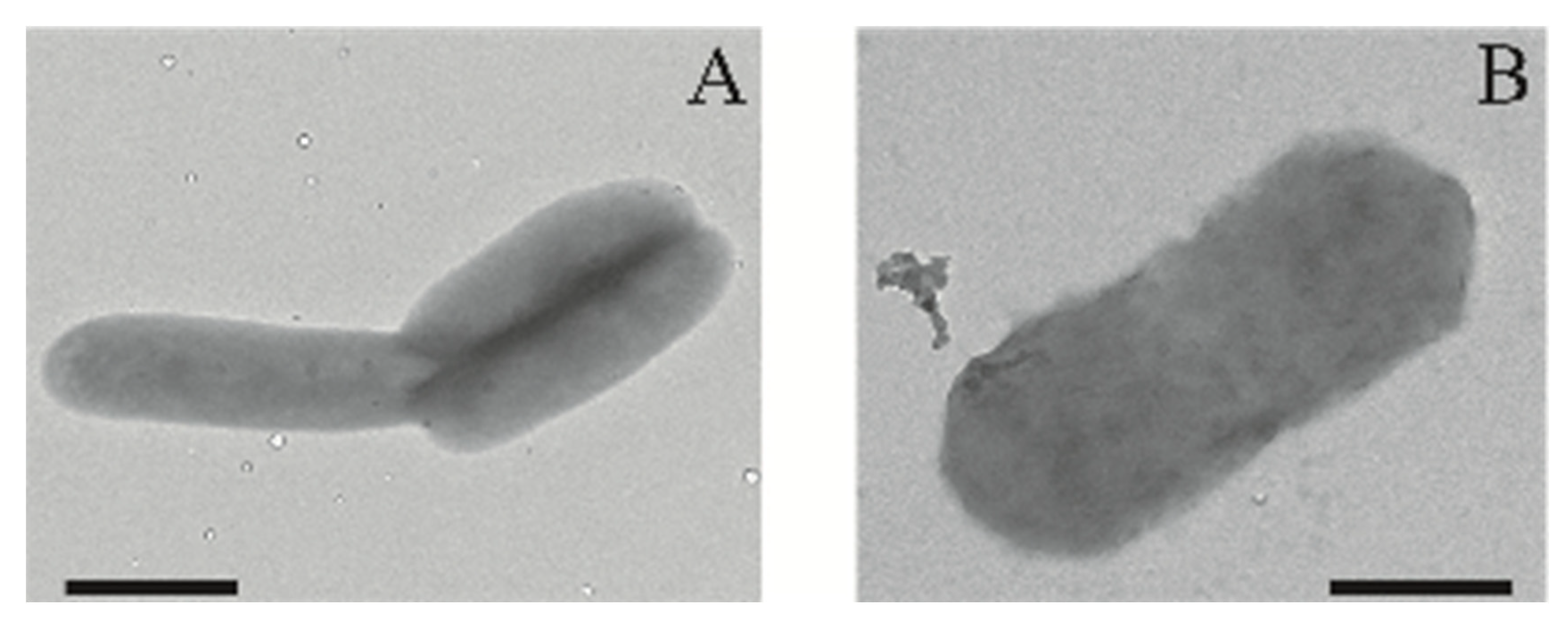
2.1. Physiological Characterization
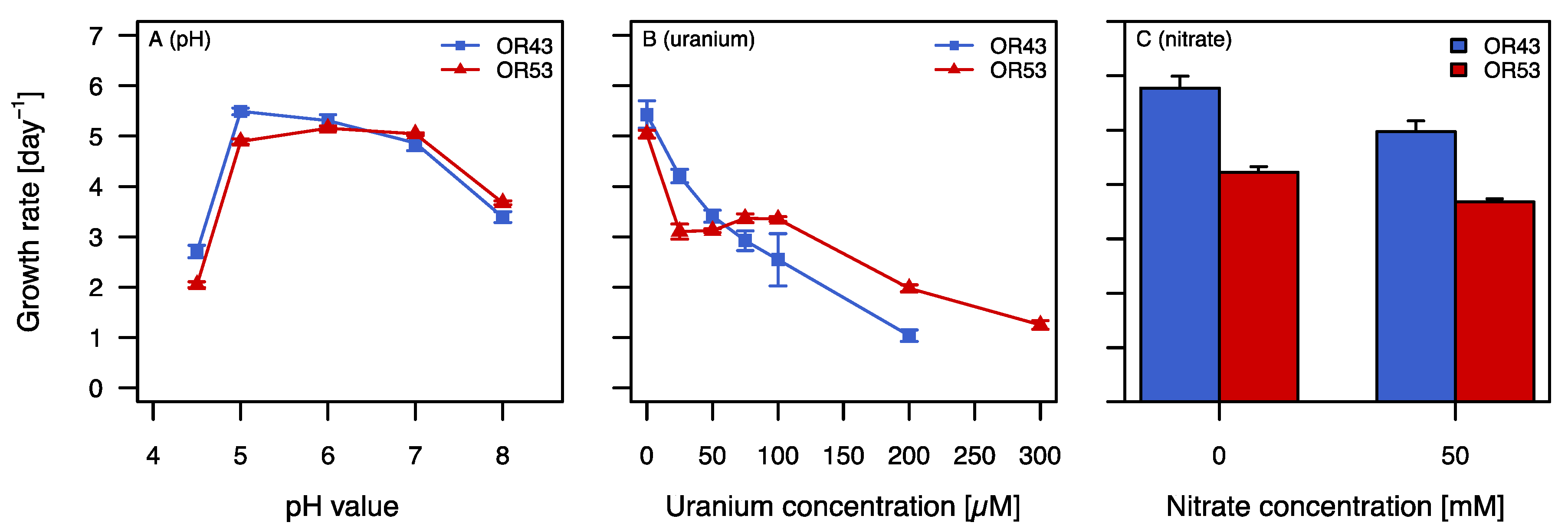
2.2. Growth in the Presence of Stress Factors
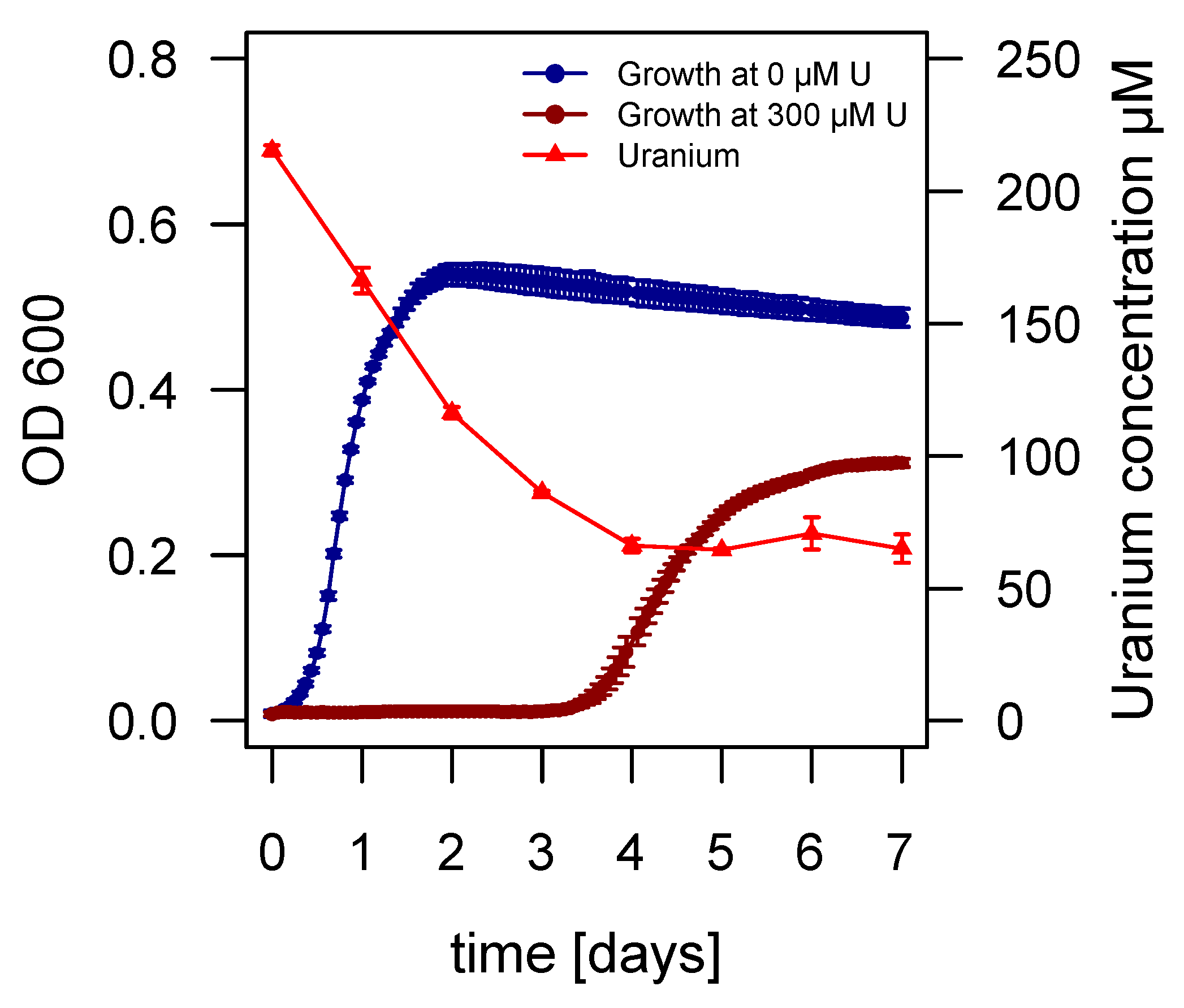
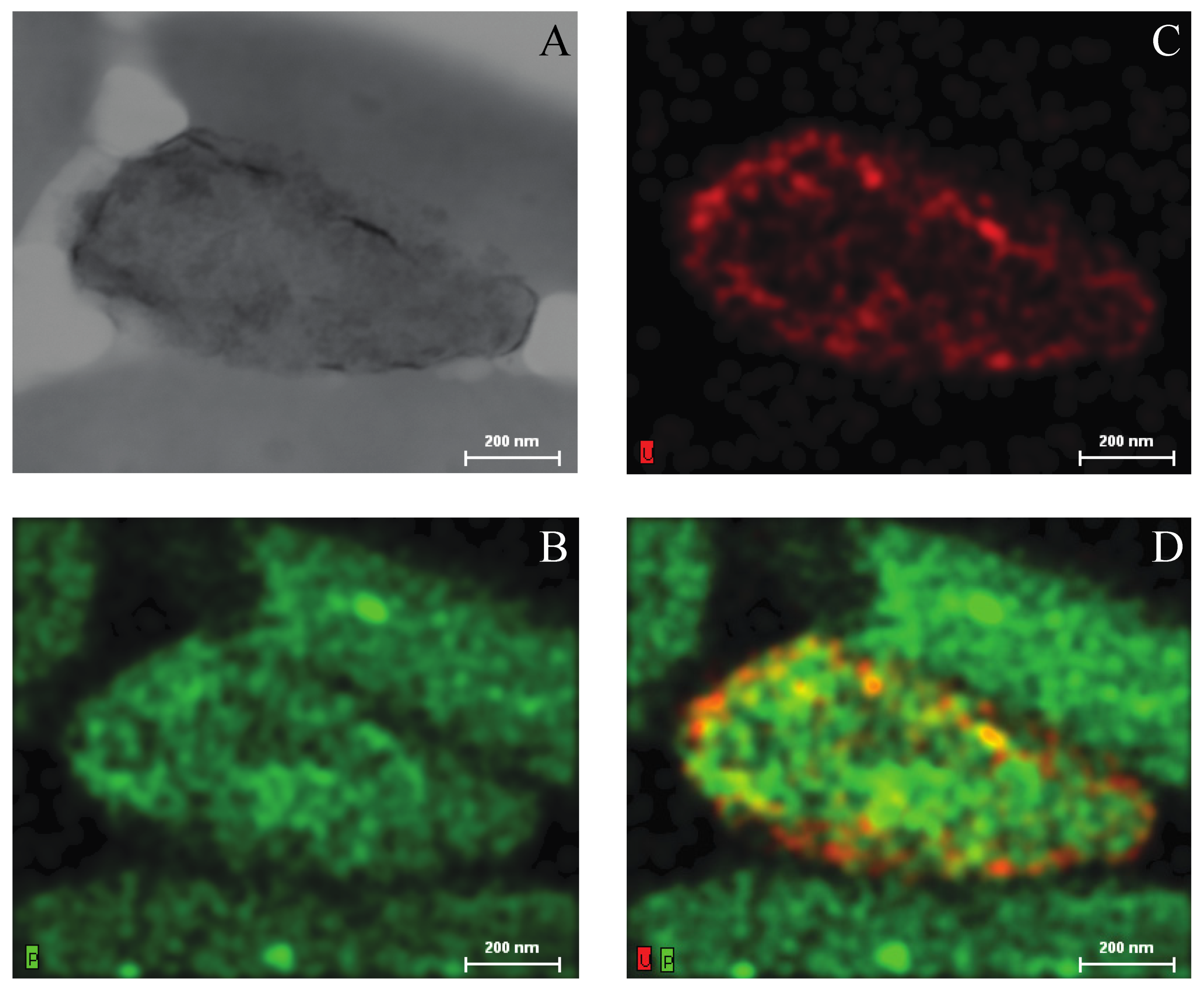
2.3. Growth of Asinibacterium sp. OR53 in the Presence of Uranium
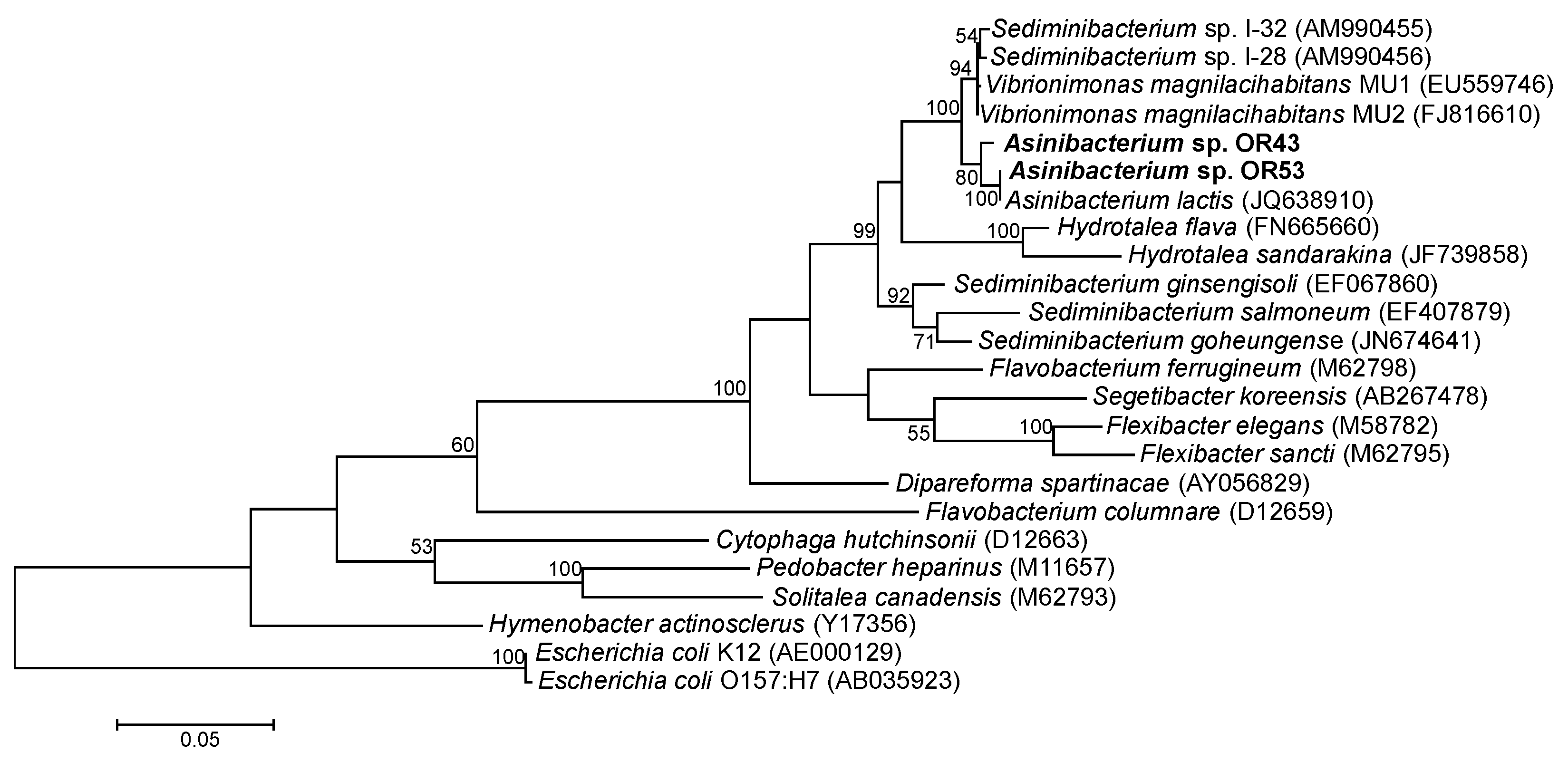
2.4. Genomes of Asinibacterium spp. OR43 and OR53
3. Material and Methods
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Kämpfer, P.; Lodders, N.; Falsen, E. Hydrotalea flava Gen. Nov., Sp. Nov., a New Member of the Phylum Bacteroidetes and Allocation of the Genera Chitinophaga, Sediminibacterium, Lacibacter, Flavihumibacter, Flavisolibacter, Niabella, Niastella, Segetibacter, Parasegetibacter, Terrimonas, Ferruginibacter, Filimonas and Hydrotalea to the Family Chitinophagaceae Fam. Nov. Int. J. Syst. Evol. Microbiol. 2011, 61, 518–523. [Google Scholar] [CrossRef]
- Lee, D.-G.; Park, J.-M.; Kang, H.; Hong, S.-Y.; Lee, K.R.; Chang, H.-B.; Trujillo, M.E. Asinibacterium lactis Gen. Nov., Sp. Nov., a Member of the Family Chitinophagaceae, Isolated from Donkey (Equus asinus) Milk. Int. J. Syst. Evol. Microbiol. 2013, 63, 3180–3185. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Albuquerque, L.; Rainey, F.A.; Nobre, M.F.; Costa, M.S.D. Hydrotalea sandarakina Sp. Nov., Isolated from a Hot Spring Runoff, and Emended Descriptions of the Genus Hydrotalea and the Species Hydrotalea flava. Int. J. Syst. Evol. Microbiol. 2012, 62, 1603–1608. [Google Scholar] [CrossRef] [Green Version]
- Kim, Y.-J.; Nguyen, N.-L.; Weon, H.-Y.; Yang, D.-C. Sediminibacterium ginsengisoli Sp. Nov., Isolated from Soil of a Ginseng Field, and Emended Descriptions of the Genus Sediminibacterium and of Sediminibacterium salmoneum. Int. J. Syst. Evol. Microbiol. 2013, 63, 905–912. [Google Scholar] [CrossRef]
- Kang, H.; Kim, H.; Lee, B.-I.; Joung, Y.; Joh, K. Sediminibacterium goheungense Sp. Nov., Isolated from a Freshwater Reservoir. Int. J. Syst. Evol. Microbiol. 2014, 64, 1328–1333. [Google Scholar] [CrossRef] [PubMed]
- Qu, J.-H.; Yuan, H.-L. Sediminibacterium salmoneum Gen. Nov., Sp. Nov., a Member of the Phylum Bacteroidetes Isolated from Sediment of a Eutrophic Reservoir. Int. J. Syst. Evol. Microbiol. 2008, 58, 2191–2194. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pisciotta, J.M.; Zou, Y.; Baskakov, I.V. Light-Dependent Electrogenic Activity of Cyanobacteria. PLoS ONE 2010, 5, e10821. [Google Scholar] [CrossRef] [PubMed]
- Dillon, J.G.; Carlin, M.; Gutierrez, A.; Nguyen, V.; McLain, N. Patterns of Microbial Diversity along a Salinity Gradient in the Guerrero Negro Solar Saltern, Baja CA Sur, Mexico. Front. Microbiol. 2013, 4, 399. [Google Scholar] [CrossRef] [Green Version]
- Bagatini, I.; Eiler, A.; Bertilsson, S.; Klaveness, D.; Tessarolli, L.P.; Vieira, A.A.H. Host-Specificity and Dynamics in Bacterial Communities Associated with Bloom-Forming Freshwater Phytoplankton. PLoS ONE 2014, 9, e85950. [Google Scholar] [CrossRef] [Green Version]
- Besemer, K.; Peter, H.; Logue, J.B.; Langenheder, S.; Lindström, E.; Tranvik, L.J.; Battin, T.J. Unraveling Assembly of Stream Biofilm Communities. ISME J. 2012, 6, 1459–1468. [Google Scholar] [CrossRef] [Green Version]
- Youngblut, N.D.; Shade, A.; Read, J.; McMahon, K.D.; Whitaker, R.J. Lineage-Specific Responses of Microbial Communities to Environmental Change. Appl. Environ. Microbiol. 2012, 79, 39–47. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cai, H.; Jiang, H.; Krumholz, L.R.; Yang, Z. Bacterial Community Composition of Size-Fractioned Aggregates within the Phycosphere of Cyanobacterial Blooms in a Eutrophic Freshwater Lake. PLoS ONE 2014, 9, e102879. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Medihala, P.; Lawrence, J.; Swerhone, G.; Korber, D. Transient Response of Microbial Communities in a Water Well Field to Application of an Impressed Current. Water Res. 2013, 47, 672–682. [Google Scholar] [CrossRef] [PubMed]
- Arias-Cordero, E.; Ping, L.; Reichwald, K.; Delb, H.; Platzer, M.; Boland, W. Comparative Evaluation of the Gut Microbiota Associated with the Below- and Above-Ground Life Stages (Larvae and Beetles) of the Forest Cockchafer, Melolontha hippocastani. PLoS ONE 2012, 7, e51557. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reti, K.L.; Thomas, M.C.; Yanke, L.J.; Selinger, L.B.; Inglis, G.D. Effect of Antimicrobial Growth Promoter Administration on the Intestinal Microbiota of Beef Cattle. Gut Pathog. 2013, 5, 8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Abulencia, C.B.; Wyborski, D.L.; Garcia, J.A.; Podar, M.; Chen, W.; Chang, S.H.; Chang, H.W.; Watson, D.; Brodie, E.L.; Hazen, T.; et al. Environmental Whole-Genome Amplification to Access Microbial Populations in Contaminated Sediments. Appl. Environ. Microbiol. 2006, 72, 3291–3301. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Singleton, D.R.; Richardson, S.D.; Aitken, M.D. Pyrosequence Analysis of Bacterial Communities in Aerobic Bioreactors Treating Polycyclic Aromatic Hydrocarbon-Contaminated Soil. Biodegradation 2011, 22, 1061–1073. [Google Scholar] [CrossRef] [Green Version]
- Al-Awadhi, H.; Dashti, N.; Khanafer, M.; Al-Mailem, D.; Ali, N.; Radwan, S. Bias Problems in Culture-Independent Analysis of Environmental Bacterial Communities: A Representative Study on Hydrocarbonoclastic Bacteria. SpringerPlus 2013, 2, 369. [Google Scholar] [CrossRef] [Green Version]
- Laplante, K.; Sébastien, B.; Derome, N. Parallel Changes of Taxonomic Interaction Networks in Lacustrine Bacterial Communities Induced by a Polymetallic Perturbation. Evol. Appl. 2013, 6, 643–659. [Google Scholar] [CrossRef]
- Callbeck, C.; Agrawal, A.; Voordouw, G.; Ishii, Y.; Matsuura, Y.; Kakizawa, S.; Nikoh, N.; Fukatsu, T. Acetate Production from Oil under Sulfate-Reducing Conditions in Bioreactors Injected with Sulfate and Nitrate. Appl. Environ. Microbiol. 2013, 79, 5059–5068. [Google Scholar] [CrossRef] [Green Version]
- Bollmann, A.; Palumbo, A.V.; Lewis, K.; Epstein, S.S. Isolation and Physiology of Bacteria from Contaminated Subsurface Sediments. Appl. Environ. Microbiol. 2010, 76, 7413–7419. [Google Scholar] [CrossRef] [Green Version]
- Nies, D.H. Microbial Heavy-Metal Resistance. Appl. Microbiol. Biotechnol. 1999, 51, 730–750. [Google Scholar] [CrossRef] [PubMed]
- Bruins, M.R.; Kapil, S.; Oehme, F.W. Microbial Resistance to Metals in the Environment. Ecotoxicol. Environ. Saf. 2000, 45, 198–207. [Google Scholar] [CrossRef]
- Nies, D.H. Efflux-Mediated Heavy Metal Resistance in Prokaryotes. FEMS Microbiol. Rev. 2003, 27, 313–339. [Google Scholar] [CrossRef]
- Newsome, L.; Morris, K.; Lloyd, J. The Biogeochemistry and Bioremediation of Uranium and other Priority Radionuclides. Chem. Geol. 2013, 363, 164–184. [Google Scholar] [CrossRef]
- Albert, R.A.; Zitomer, D.; Dollhopf, M.; Schauer-Gimenez, A.E.; Struble, C.; King, M.; Son, S.; Langer, S.; Busse, H.-J. Proposal of Vibrionimonas magnilacihabitans Gen. Nov., Sp. Nov., a Curved Gram-Stain-Negative Bacterium Isolated from Lake Water. Int. J. Syst. Evol. Microbiol. 2014, 64, 613–620. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ayarza, J.M.; Figuerola, E.L.M.; Erijman, L. Draft Genome Sequences of Type Strain Sediminibacterium salmoneum NJ-44 and Sediminibacterium sp. Strain C3, a Novel Strain Isolated from Activated Sludge. Genome Announc. 2014, 2, e01073-13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dhal, P.K.; Islam, E.; Kazy, S.K.; Sar, P. Culture-Independent Molecular Analysis of Bacterial Diversity in Uranium-Ore/-Mine Waste-Contaminated and Non-Contaminated Sites from Uranium Mines. 3 Biotech 2011, 1, 261–272. [Google Scholar] [CrossRef] [Green Version]
- Limbri, H.; Gunawan, C.; Thomas, T.; Smith, A.; Scott, J.; Rosche, B. Coal-Packed Methane Biofilter for Mitigation of Green House Gas Emissions from Coal Mine Ventilation Air. PLoS ONE 2014, 9, e94641. [Google Scholar] [CrossRef]
- Eberdugo-Clavijo, C.; Gieg, L.M. Conversion of Crude Oil to Methane by a Microbial Consortium Enriched from Oil Reservoir Production Waters. Front. Microbiol. 2014, 5, 197. [Google Scholar] [CrossRef] [Green Version]
- Brzoska, R.M.; Bollmann, A. The long-term effect of uranium and pH on the community composition of an artificial consortium. FEMS Microbiol. Ecol. 2015, 92, fiv158. [Google Scholar] [CrossRef] [PubMed]
- Barkleit, A.; Moll, H.; Bernhard, G. Interaction of Uranium(vi) with Lipopolysaccharide. Dalton Trans. 2008, 21, 2879–2886. [Google Scholar] [CrossRef] [PubMed]
- Brzoska, R.M.; Huntemann, M.; Clum, A.; Chen, A.; Kyrpides, N.; Palaniappan, K.; Ivanova, N.; Mikhailova, N.; Ovchinnikova, G.; Varghese, N.; et al. Complete Genome Sequence for Asinibacterium sp. Strain OR53 and Draft Genome Sequence for Asinibacterium sp. Strain OR43, Two Bacteria Tolerant to Uranium. Microbiol. Resour. Announc. 2019, 8, e01701-18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Goris, J.; Konstantinidis, K.T.; Klappenbach, J.A.; Coenye, T.; Vandamme, P.; Tiedje, J.M. DNA–DNA Hybridization Values and their Relationship to Whole-Genome Sequence Similarities. Int. J. Syst. Evol. Microbiol. 2007, 57, 81–91. [Google Scholar] [CrossRef] [Green Version]
- Varghese, N.J.; Mukherjee, S.; Ivanova, N.; Konstantinidis, K.T.; Mavrommatis, K.; Kyrpides, N.C.; Pati, A. Microbial Species Delineation Using Whole Genome Sequences. Nucleic Acids Res. 2015, 43, 6761–6771. [Google Scholar] [CrossRef]
- Zumft, W. Cell Biology and Molecular Basis of Denitrification. Microbiol. Mol. Biol. Rev. 1997, 61, 533–616. [Google Scholar]
- Hino, T.; Nagano, S.; Sugimoto, H.; Tosha, T.; Shiro, Y. Molecular Structure and Function of Bacterial Nitric Oxide Reductase. Biochim. Biophys. Acta 2011, 1817, 680–687. [Google Scholar] [CrossRef] [Green Version]
- Tavares, P.; Pereira, A.S.; Moura, J.; Moura, I. Metalloenzymes of the Denitrification Pathway. J. Inorg. Biochem. 2006, 100, 2087–2100. [Google Scholar] [CrossRef]
- Pauleta, S.R.; Dell’Acqua, S.; Moura, I. Nitrous Oxide Reductase. Co-ord. Chem. Rev. 2012, 257, 332–349. [Google Scholar] [CrossRef]
- Smith, J.L.; Liu, Y.; Paoli, G.C. How Does Listeria monocytogenes combat Acid Conditions? Can. J. Microbiol. 2013, 59, 141–152. [Google Scholar] [CrossRef]
- Phan-Thanh, L.; Mahouin, F. A Proteomic Approach to Study the Acid Response in Listeria monocytogenes. ELECTROPHORESIS Int. J. 1999, 20, 2214–2224. [Google Scholar] [CrossRef]
- Olson, E.R. Influence of pH on Bacterial Gene Expression. Mol. Microbiol. 1993, 8, 5–14. [Google Scholar] [CrossRef]
- Kobayashi, H.; Saito, H.; Kakegawa, T. Bacterial Strategies to Inhabit Acidic Environments. J. Gen. Appl. Microbiol. 2000, 46, 235–243. [Google Scholar] [CrossRef] [Green Version]
- Trchounian, A.; Kobayashi, H. Kup Is the Major K+uptake System in Escherichia coli upon Hyper-Osmotic Stress at a Low pH. FEBS Lett. 1999, 447, 144–148. [Google Scholar] [CrossRef] [Green Version]
- Foster, J.W. Escherichia coli Acid Resistance: Tales of an Amateur Acidophile. Nat. Rev. Genet. 2004, 2, 898–907. [Google Scholar] [CrossRef]
- Macaskie, L.E.; Bonthrone, K.M.; Yong, P.; Goddard, D.T. Enzymically Mediated Bioprecipitation of Uranium by a Citrobacter sp.: A Concerted Role for Exocellular Lipopolysaccharide and Associated Phosphatase in Biomineral Formation. Microbiology 2000, 146, 1855–1867. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Renninger, N.; Knopp, R.; Nitsche, H.; Clark, D.S.; Keasling, J.D. Uranyl Precipitation by Pseudomonas aeruginosa via Controlled Polyphosphate Metabolism. Appl. Environ. Microbiol. 2004, 70, 7404–7412. [Google Scholar] [CrossRef] [Green Version]
- Martinez, R.; Beazley, M.; Taillefert, M.; Arakaki, A.K.; Skolnick, J.; Sobecky, P.A. Aerobic Uranium (VI) Bioprecipitation by Metal-Resistant Bacteria Isolated from Radionuclide- and Metal-Contaminated Subsurface Soils. Environ. Microbiol. 2007, 9, 3122–3133. [Google Scholar] [CrossRef]
- Nilgiriwala, K.; Alahari, A.; Rao, A.S.; Apte, S.K. Cloning and Overexpression of Alkaline Phosphatase PhoK from Sphingomonas sp. Strain BSAR-1 for Bioprecipitation of Uranium from Alkaline Solutions. Appl. Environ. Microbiol. 2008, 74, 5516–5523. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kulkarni, S.; Misra, C.S.; Gupta, A.; Ballal, A.; Apte, S.K. Interaction of Uranium with Bacterial Cell Surfaces: Inferences from Phosphatase-Mediated Uranium Precipitation. Appl. Environ. Microbiol. 2016, 82, 4965–4974. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Choudhary, S.; Sar, P. Uranium Biomineralization by a Metal Resistant Pseudomonas aeruginosa Strain Isolated from Contaminated Mine Waste. J. Hazard. Mater. 2011, 186, 336–343. [Google Scholar] [CrossRef]
- Nedelkova, M.; Merroun, M.L.; Rossberg, A.; Hennig, C.; Selenska-Pobell, S. Microbacterium Isolates from the Vicinity of a Radioactive Waste Depository and their Interactions with Uranium. FEMS Microbiol. Ecol. 2007, 59, 694–705. [Google Scholar] [CrossRef] [Green Version]
- Merroun, M.; Nedelkova, M.; Rossberg, A.; Hennig, C.; Selenska-Pobell, S. Interaction Mechanisms of Bacterial Strains Isolated from Extreme Habitats with Uranium. Radiochim. Acta 2006, 94, 723–729. [Google Scholar] [CrossRef]
- Jroundi, F.; Merroun, M.L.; Arias, J.M.; Rossberg, A.; Selenska-Pobell, S.; Gonzalez-Muñoz, M.T. Spectroscopic and Microscopic Characterization of Uranium Biomineralization in Myxococcus Xanthus. Geomicrobiol. J. 2007, 24, 441–449. [Google Scholar] [CrossRef]
- Li, X.; Ding, C.; Liao, J.; Du, L.; Sun, Q.; Yang, J.; Yang, Y.; Zhang, D.; Tang, J.; Liu, N. Bioaccumulation Characterization of Uranium by a Novel Streptomyces sporoverrucosus dwc-3. J. Environ. Sci. 2016, 41, 162–171. [Google Scholar] [CrossRef]
- Gerber, U.; Zirnstein, I.; Krawczyk-Bärsch, E.; Lünsdorf, H.; Arnold, T.; Merroun, M. Combined Use of Flow Cytometry and Microscopy to Study the Interactions between the Gram-Negative Betaproteobacterium Acidovorax facilis and Uranium (VI). J. Hazard. Mater. 2016, 317, 127–134. [Google Scholar] [CrossRef] [PubMed]
- Chen, I.-M.A.; Chu, K.; Palaniappan, K.; Pillay, M.; Ratner, A.; Huang, J.; Huntemann, M.; Varghese, N.; White, J.R.; Seshadri, R.; et al. IMG/M v.5.0: An Integrated Data Management and Comparative Analysis System for Microbial Genomes and Microbiomes. Nucleic Acids Res. 2018, 47, D666–D677. [Google Scholar] [CrossRef] [PubMed]
- Pal, C.; Bengtsson-Palme, J.; Rensing, C.; Kristiansson, E.; Larsson, D.G.J. BacMet: Antibacterial Biocide and Metal Resistance Genes Database. Nucleic Acids Res. 2013, 42, D737–D743. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Silver, S.; Phung, L.T. Bacterial Heavy Metal Resistance: New Surprises. Annu. Rev. Microbiol. 1996, 50, 753–789. [Google Scholar] [CrossRef]
- Hobman, J.; Crossman, L.C. Bacterial Antimicrobial Metal Ion Resistance. J. Med Microbiol. 2015, 64, 471–497. [Google Scholar] [CrossRef]
- Ladomersky, E.; Petris, M.J. Copper Tolerance and Virulence in Bacteria. Metallomics 2015, 7, 957–964. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bondarczuk, K.; Piotrowska-Seget, Z. Molecular Basis of Active Copper Resistance Mechanisms in Gram-Negative Bacteria. Cell Biol. Toxicol. 2013, 29, 397–405. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rensing, C.; Grass, G. Escherichia coli Mechanisms of Copper Homeostasis in a Changing Environment. FEMS Microbiol. Rev. 2003, 27, 197–213. [Google Scholar] [CrossRef] [Green Version]
- Alvarez-Ortega, C.; Olivares, J.; Martínez, J.L. RND Multidrug Efflux Pumps: What Are they Good for? Front. Microbiol. 2013, 4, 7. [Google Scholar] [CrossRef] [Green Version]
- Goldberg, M.; Pribyl, T.; Juhnke, S.; Nies, D.H. Energetics and Topology of CzcA, a Cation/Proton Antiporter of the Resistance-Nodulation-Cell Division Protein Family. J. Biol. Chem. 1999, 274, 26065–26070. [Google Scholar] [CrossRef] [Green Version]
- Kolaj-Robin, O.; Russell, D.; Hayes, K.; Pembroke, T.; Soulimane, T. Cation Diffusion Facilitator Family: Structure and Function. FEBS Lett. 2015, 589, 1283–1295. [Google Scholar] [CrossRef]
- Eitinger, T.; Mandrand-Berthelot, M.-A. Nickel Transport Systems in Microorganisms. Arch. Microbiol. 2000, 173, 1–9. [Google Scholar] [CrossRef]
- Barkay, T.; Miller, S.M.; Summers, A.O. Bacterial Mercury Resistance from Atoms to Ecosystems. FEMS Microbiol. Rev. 2003, 27, 355–384. [Google Scholar] [CrossRef] [Green Version]
- Alvarez, A.H.; Moreno-Sánchez, R.; Cervantes, C. Chromate Efflux by Means of the ChrA Chromate Resistance Protein from Pseudomonas aeruginosa. J. Bacteriol. 1999, 181, 7398–7400. [Google Scholar] [CrossRef] [Green Version]
- Aguilar-Barajas, E.; Paluscio, E.; Cervantes, C.; Rensing, C. Expression of Chromate Resistance Genes from Shewanella sp. Strain ANA-3 in Escherichia Coli. FEMS Microbiol. Lett. 2008, 285, 97–100. [Google Scholar] [CrossRef] [Green Version]
- Bollmann, A.; French, E.; Laanbroek, H.J. Isolation, Cultivation, and Characterization of Ammonia-Oxidizing Bacteria and Archaea Adapted to Low Ammonium Concentrations. Methods Enzymol. 2011, 486, 55–88. [Google Scholar] [CrossRef] [PubMed]
- Reasoner, D.J.; Geldreich, E.E. A New Medium for the Enumeration and Subculture of Bacteria from Potable Water. Appl. Environ. Microbiol. 1985, 49, 1–7. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rashid, M.H.; Kornberg, A. Inorganic Polyphosphate Is Needed for Swimming, Swarming, and Twitching Motilities of Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 2000, 97, 4885–4890. [Google Scholar] [CrossRef] [Green Version]
- Hatchel, J.M.; Balish, R.S.; Duley, M.L.; Balish, M.F. Ultrastructure and Gliding Motility of Mycoplasma amphoriforme, a Possible Human respiratory Pathogen. Microbiology 2006, 152, 2181–2189. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miller, L.T. Single Derivatization Method for Routine Analysis of Bacterial Whole-Cell Fatty Acid Methyl Esters, Including Hydroxy Acids. J. Clin. Microbiol. 1982, 16, 584–586. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kuykendall, L.D.; Roy, M.A.; O’Neill, J.J.; Devine, T.E. Fatty Acids, Antibiotic Resistance, and Deoxyribonucleic Acid Homology Groups of Bradyrhizobium japonicum. Int. J. Syst. Bacteriol. 1988, 38, 358–361. [Google Scholar] [CrossRef] [Green Version]
- Kämpfer, P.; Kroppenstedt, R.M. Numerical Analysis of Fatty Acid Patterns of Coryneform Bacteria and Related Taxa. Can. J. Microbiol. 1996, 42, 989–1005. [Google Scholar] [CrossRef]
- Teixeira, L.S.G.; Costa, A.C.S.; Ferreira, S.L.C.; Freitas, M.D.L.; Carvalho, M.S.D. Spectrophotometric Determination of Uranium Using 2-(2-Thiazolylazo)-p-Cresol (TAC) in the Presence of Surfactants. J. Braz. Chem. Soc. 1999, 10, 519–522. [Google Scholar] [CrossRef] [Green Version]
- Weisburg, W.G.; Barns, S.M.; Pelletier, D.A.; Lane, D.J. 16S Ribosomal DNA Amplification for Phylogenetic Study. J. Bacteriol. 1991, 173, 697–703. [Google Scholar] [CrossRef] [Green Version]
- Muyzer, G.; de Waal, E.C.; Uitterlinden, A.G. Profiling of Complex Microbial Populations by Denaturing Gradient Gel Electrophoresis Analysis of Polymerase Chain Reaction-Amplified Genes Coding for 16S rRNA. Appl. Environ. Microbiol. 1993, 59, 695–700. [Google Scholar] [CrossRef] [Green Version]
- Ludwig, W.; Strunk, O.; Westram, R.; Richter, L.; Meier, H.; Yadhukumar; Buchner, A.; Lai, T.; Steppi, S.; Jobb, G.; et al. ARB: A Software Environment for Sequence Data. Nucleic Acids Res. 2004, 32, 1363–1371. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| Gene Product Name | Gene Symbol | OR43 | OR53 |
|---|---|---|---|
| Nitrate/nitrite transporter | |||
| Nitrite/nitrate transporter | narK | + | + |
| ABC transporter | nrtABC | + | - |
| Assimilatory nitrate reduction | |||
| Ferredoxin nitrate reductase | narB | + | + |
| Dissimilatory nitrate reduction | |||
| Nitrite reductase | nirBD | + | + |
| Denitrification | |||
| Nitrate reductase | napAB | - | - |
| Nitrite reductase (copper containing) | nirK | + | + |
| Nitric oxide reductase (qNOR) | norC | + | + |
| Nitrous oxide reductase | nosZLDFY | + | + |
| Gene Product Name | Gene Symbol | OR43 | OR53 |
|---|---|---|---|
| F-type proton translocating ATPase | atpG | 1 | 1 |
| atpCD | 1 | 1 | |
| atpBEFHA | 1 | 1 | |
| High affinity potassium transporter | kdpABCD | 1 | 1 |
| Low affinity potassium transporter | kup | 2 | 2 |
| Gene Product Name | Gene Symbol | OR43 | OR53 |
|---|---|---|---|
| Arsenic | |||
| Arsenic resistance operon | arsBCR | 0 | 1 |
| Arsenate reductase (protein-tyrosine phosphatase) | arsC | 1 | 1 |
| Copper and silver | |||
| Copper exporting P-type ATPase (Cu2+) | copAB | 1 | 1 |
| Copper exporting P-type ATPase (Cu2+) | copA | 2 | 2 |
| Copper exporting P-type ATPase (Cu2+) | copB | 2 | 2 |
| Copper chaperone | copZ | 1 | 1 |
| Copper homeostasis protein | cutC | 1 | 1 |
| Cu(I)/Ag(I) efflux system operon | cusAB, silAB | 1 | 3 |
| Copper two component regulatory systems | copRS, cusRS | 4 | 4 |
| Cobalt, zinc, cadmium | |||
| Cobalt–zinc–cadmium resistance operon | czcCBA | 2 | 1 |
| Heavy metal efflux pump | czcA | 3 | 3 |
| Membrane fusion protein | czcB | 2 | 2 |
| Cation diffusion facilitator | czcD | 2 | 2 |
| Cd2+/Zn2+ exporting ATPase | zntA | 1 | 1 |
| Mercury | |||
| Possible transcriptional regulator | merR | 2 | 2 |
| Putative mercury transport protein | merT-P | 2 | 2 |
| Mercury reductase | merA | 1 | 1 |
| Mercury resistance protein | merC | 1 | 1 |
| Zinc | |||
| Cd2+/Zn2+ exporting ATPase | zntA | 1 | 1 |
| Nickel | |||
| Nickel transport operon | nikABCE | 1 | 1 |
| Chromium | |||
| Chromate transporter | chrA | 1 | 1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Brzoska, R.M.; Edelmann, R.E.; Bollmann, A. Physiological and Genomic Characterization of Two Novel Bacteroidota Strains Asinibacterium spp. OR43 and OR53. Bacteria 2022, 1, 33-47. https://doi.org/10.3390/bacteria1010004
Brzoska RM, Edelmann RE, Bollmann A. Physiological and Genomic Characterization of Two Novel Bacteroidota Strains Asinibacterium spp. OR43 and OR53. Bacteria. 2022; 1(1):33-47. https://doi.org/10.3390/bacteria1010004
Chicago/Turabian StyleBrzoska, Ryann M., Richard E. Edelmann, and Annette Bollmann. 2022. "Physiological and Genomic Characterization of Two Novel Bacteroidota Strains Asinibacterium spp. OR43 and OR53" Bacteria 1, no. 1: 33-47. https://doi.org/10.3390/bacteria1010004





