Abstract
Theodoxus fluviatilis (Linnaeus, 1758) (Gastropoda: Neritidae) is an oligohaline aquatic gastropod that inhabits most of Europe and adjacent areas of Asia. Two different ecotypes can be distinguished: One in freshwater (FW) and another along the Baltic Sea coast in brackish water habitats (BW). Individuals of either ecotype use free amino acids and urea as organic osmolytes to adjust body fluid osmolality to the external medium; however, the BW ecotype is able to accumulate them in larger quantities. The use of urea as an organic osmolyte in aquatic gastropods such as T. fluviatilis has only recently been initially described and raised the question of how urea transport between body fluids and the environment is balanced. Upon examining transcriptome and preliminary genome sequence data of T. fluviatilis, we identified putative homologues of DUR3 genes, which code for urea transporters (UTs) in other organisms. In this study, we provide evidence for the presence of four different subtypes of DUR3-like UTs that belong to two distinct families. Two of the UT subtypes were subject to qRT-PCR analyses to investigate differences in mRNA expression during the acclimation of individuals of both ecotypes to different salinities. Our results indicate that only BW animals regulate DUR3 gene expression in the context of osmoregulation.
1. Introduction
Urea is one of the terminal products of nitrogen metabolism in animals, e.g., the breakdown of proteins and nucleotides [1]. Compared with ammonia, it is a less toxic substance [2] that is readily excreted via the kidney or the gills in water-dwelling organisms. In certain organisms, however, urea serves as an organic osmolyte to adjust the osmolality of body fluids to those of adjacent organs or organ compartments or to the external medium. Prominent examples are the renal medulla in mammalian kidneys for the retention of water during urine formation [3] or the adjustment of the body fluids in elasmobranch fishes such as sharks or rays to the osmolality of their marine environment [4,5].
Urea is produced intracellularly, but in order to be excreted or to act as an organic osmolyte also in extracellular body fluids, it has to be transported across cell membranes. Since the passive diffusion of urea is very limited, several channels and transporters act to enhance the permeability of cell membranes for urea. Some aquaporine subtypes, namely the aquaglyceroporines Aqp3, 7, 9, and 10, are permeable for urea [6]. Specific transport of urea across biological membranes may occur by facilitated diffusion (along the concentration gradient) [7] or may be energized by a sodium concentration gradient that drives urea/Na+ cotransport [8,9] or antiport [10] depending on the animal species, the respective organ, or the subcellular localization [11].
In vertebrates, the specific transport of urea is mediated by the urea transporters UT-A and UT-B [12,13]. Both UTs facilitate the transport of urea down a concentration gradient [14]. In contrast, DUR3 is the urea transporter in yeast [15,16], fungi [17,18], plants [19], and the fluted giant clam, Tridacna squamosa (Lamarck, 1819) (Cardiida: Cardiidae) [20]. DUR3 is a urea/H+ symporter and hence also allows, in principle, for the transport of urea against a concentration gradient. Indeed, an active urea uptake from the environment has been described for yeast [21], plants [22,23,24], and molluscs [20]. The presence of the DUR3-like urea transporter encoding genes in invertebrates has so far been confirmed in mollusc species [20,25], but reports for other invertebrate groups are spare.
Aquatic snails can be classified according to their origin (marine, freshwater, and brackish water species) and their osmoregulatory capacity (stenohaline and eury- or oligohaline species). In general, all groups keep the osmolality of their body fluids close to the level of their environment (osmoconformers) to reduce the necessity of energy-consuming osmoregulatory processes. However, oligohaline snails are able to hyperregulate their internal osmolality under very low-salinity conditions [26,27,28,29].
Recently, the nerite snail Theodoxus fluviatilis (Linnaeus, 1758) (Gastropoda: Neritidae), a representative of the oligohaline gastropods, was reported to accumulate urea under hyperosmotic conditions. T. fluviatilis naturally occurs in both freshwater (FW, 0.5‰ salinity) and brackish water (BW, approximately 8‰ salinity) habitats, and the accumulation of urea in body fluids upon transfer of an animal to hyperosmotic conditions was observed in representatives of both ecotypes [30]. However, BW individuals displayed a much higher base level of urea already when they were kept under natural water conditions (8‰ salinity), and they were capable of increasing the concentration of urea to 500–700 mmol/kg fresh mass under hyperosmotic conditions (28‰ salinity). In contrast, FW individuals reached a maximal urea concentration under hyperosmotic conditions (21‰ salinity) of just 50 mmol/kg fresh mass. Further investigations indicated that FW individuals primarily rely on the accumulation of amino acids instead of urea to increase the osmolality of their body fluids [30].
Osmoregulation is an energy-demanding process, and a detailed look at both the expression levels and the activities of transport ATPases that are potentially relevant for a successful adaptation to changing external salinities revealed that the V-ATPase may not be important at all for ion homeostasis and that the basal activity of the Na+/K+-ATPase seems to be sufficient for both ion and volume balances in T. fluviatilis [31].
The main objective of the present study was to further evaluate the molecular mechanisms behind the osmoregulatory processes in T. fluviatilis and to elucidate whether or not differences in the expression of putative urea transporters occur between the two ecotypes and after exposure of respective individuals to either hyper- or hypoosmotic conditions. We used transcriptome sequence data (NCBI, accession numbers SRR15300633 and GKDH00000000.1) to identify relevant coding sequences and applied a qRT-PCR approach to detect differences in the expression levels of corresponding mRNAs. Our results indicate the presence of two families of putative DUR3-like urea transporters in T. fluviatilis and provide evidence for their contribution to the changes in body fluid urea concentrations during osmotic acclimation of the animals.
2. Materials and Methods
2.1. Animals
Adult snails were collected from late spring to early autumn in 2019–2021. Local water temperatures (approximately 12–20 °C) and salinity conditions (0.5‰ for FW and approximately 8‰ for BW sampling sites, respectively) were determined. BW snails were collected at the southern shore of the Baltic Sea in shallow water near the island of Riems (54°11′00″ N, 13°21′10.2″ E) and the island of Hiddensee (54°34′40” N, 13°06′48″ E). FW animals originated from the lake ‘Schmaler Luzin’ in northern Germany (53°18′01.4″ N, 13°25′53.2″ E).
T. fluviatilis individuals were kept along with stones and substrate from their natural habitat in aerated glass aquariums with a ground area of 30 × 60 cm at room temperature (20 °C) and a 16 to 8 h day–night cycle in addition to the natural light cycle. The salinity level of the water was adjusted to the level of their native environment (BW animals at 8‰ salinity, FW animals at 0.5‰ salinity), achieved by dissolving ‘Tropic Marine Classic’ sea salt (Dr. Biener GmbH, Wartenberg, Germany) in deionized water and monitored with a conductivity meter (CO 301, VWR International GmbH Germany, Darmstadt, Germany) in a daily routine.
2.2. Transfer Experiments
The transfer experiments followed the same routine as already described in great detail in Knobloch et al. [31]. The two control groups were maintained at salinity levels corresponding to the natural habitat (BW 8‰, FW 0.5‰). Two hyperosmotic groups with final salinities of 28‰ for BW and 18‰ for FW individuals, respectively, and one hypoosmotic group (final salinity of 0.5‰) for the BW snails were formed. The animals were stepwise transferred to the different salinity levels (see Supplementary Material Figure S1) and kept together at room temperature and a 16 to 8 h day–night cycle in small glass tanks with a ground area of 24 × 15 cm that were filled with 1 L of water and some pebbles. The duration of the experiment was limited to 10 days with 2 intermediate stages before the animals were kept in the final salinity for 5 days (see Figure S1). Survival was checked every 24 h. Sexes were not separated for the experiments. At the end, the animals were stored at −20 °C for 10 min to induce an anesthetic effect before the shells and the opercula were removed. The wet weight of the remaining tissue was measured for each animal, and the tissue samples were separately stored in liquid nitrogen until further use.
2.3. Total RNA Extraction and cDNA Synthesis
RNA extraction was performed using the Monarch Total RNA Miniprep Kit (New England Biolabs GmbH, Frankfurt am Main, Germany). Because of size variances, 400 µL of the 1X DNA/RNA Protection Reagent was used for the generally smaller BW animals and 800 µL for the larger FW snails. The total soft tissues of individual snails were weighed and homogenized in 2 mL tubes on ice with a T8-Ultra-Turrax (IKA-Werke, Staufen im Breisgau, Germany). After a centrifugation step for 2 min at 16,000× g, the supernatant was filled in a new tube. The recommended proteinase K treatment was skipped based on experiences in previous preparations. All further steps, however, followed the recommendations of the manufacturer, including the DNAse I treatment to remove remaining DNA contaminations. Total RNA was eluted using 100 µL of RNAse free water with an incubation step of 2 min prior to the centrifugation. Aliquots were used for the photometric determination of RNA concentration and purity. Furthermore, agarose gel electrophoresis (2% gels) and ethidium bromide staining were carried out to check the integrity of the RNA. Eluates were stored at −80 °C until further use.
RNA was transcribed into cDNA using the reverse transcriptase RevertAid (EP0441, Fisher Scientific, Schwerte, Germany). Each 20 µL reaction volume aliquot contained approximately 300 ng of total RNA, 2.5 µmol/L each of random hexamer- and oligodT-primer, and 200 U of the enzyme. The reactions were performed according to the manufacturer’s protocol.
2.4. Generation and Assembly of Transcriptome Data
A pool of foot muscle preparations of 24 individuals that were collected at freshwater and brackish water locations in Northern Germany and exposed to different salinities under laboratory conditions was homogenized in an adequate amount of TRIzol reagent (Thermo Fisher Scientific, Schwerte, Germany). Total RNA was extracted according to the manufacturer’s protocol and solubilized in 500 µL RNase free water. Sequencing and assembling were carried out by GATC Biotech AG (Konstanz, Germany) using an Illumina HiSeq2500 system with 300-bp paired-end reads. The obtained raw data consisted of 64,383,400 reads and were uploaded to Genbank (SRR15300633). The CLC mapper software (version: 4.22.107090) was used to assemble the raw data. A total of 442,667 de novo transcriptome contigs were generated and uploaded to GenBank (GKDH00000000.1).
2.5. Generation and Assembly of Genome Data
The tissue of the foot muscle from one individual of the brackish water ecotype was disrupted in liquid nitrogen using a mortar and pestle and an additional proteinase K digestion step. The DNA extraction was performed using the MagAttract HMW DNA Kit (Qiagen, Hilden, Germany) according to the manufacturer’s protocol. The DNA was stored at −80 °C until use. The sample and library preparation, as well as the sequencing procedure, were carried out in the Competence Centre for Genomic Analysis (CCGA Kiel, Kiel, Germany). A PacBio SMRTcell express template prep kit 2.0 (PacBio, Menlo Park, CA, USA) was used to prepare the library with a modification for low-input samples without size selection. The library was sequenced on 4 SMRTcells (SMRTCell 1M v3, sequencing chemistry 2.1) obtaining 14 Gb of data and a raw read length of approximately 5.7 Kb. PacBio subreads were assembled using Flye v2.8 [32] with the PacBio-raw mode. The resulting genome assembly of T. fluviatilis was 1.04 Gb in size with a fragment N50 of 55.4 Kb. The assembly contains 67.5% complete metazoan conserved orthologs, as analyzed using BUSCO v5.3.2 [33].
2.6. Identification and Characterization of DUR3-like Urea Transporter Homologues
Coding sequences for DUR3-like urea transporters of various mollusc species (e.g., T. squamosa [MF073181.1] and Biomphalaria glabrata (Say, 1818) (Basommatophora: Planorbidae) [XM_013209241.1]) were used to identify respective sequences in the T. fluviatilis transcriptome. Searches were performed using the Geneious software version 9.1.5. (Biomatters Ltd., Auckland, New Zealand). Repeatedly found BLAST hits with an E-value cut-off smaller than 10−10 were used for further investigation. To enhance the reliability of the outputs, the newly identified sequences were re-blasted against the NCBI database using both the blastn and tblastn functions, respectively. Final sequences of four different putative DUR3-like urea transporters of T. fluviatilis were deposited in GenBank under the accession numbers OP762683, OP762684, OP762685, and OP762686.
The putative DUR3-like urea transporters were analyzed for the presence of transmembrane (TM) regions/helices using the MEMSAT3 and MEMSAT_SVM algorithms provided by the PSIPRED protein structure prediction server (http://bioinf.cs.ucl.ac.uk/psipred, accessed on 17 March 2022). Further analyses to identify conserved domains or putative ATP binding sites were conducted using either the Conserved Domain Database (https://www.ncbi.nlm.nih.gov/Structure/cdd/docs/cdd_search.html, accessed on 17 March 2022) [34] or NsitePred (http://biomine.cs.vcu.edu/servers/NsitePred/, accessed on 17 March 2022) [35], respectively.
2.7. Bioinformatic Tools
Basic Local Alignment Search Tool (BLAST) searches were performed using the respective NCBI web portal and default settings for search algorithm parameters. Multiple sequence alignments were generated using the CLC Sequence Viewer software package v8.0 (CLC bio, QIAGEN Aarhus A/S, Aarhus, Denmark) with the following settings: Gap open cost: 5.0; gap extension cost: 2.0; end gap cost: as any other. Alignments were exported as msf files and further processed using GeneDoc v2.7 [36]. The neighbor-joining algorithm, Jukes–Cantor distance measurement, and bootstrapping with 10,000 replicates were used to create an unrooted phylogenetic tree of putative mollusc DUR3-like urea transporters.
2.8. Primer Design for qRT-PCR
Two of the four putative DUR3-like urea transporters coding sequences of T. fluviatilis were selected as targets for the deduction of primers for qRT-PCR analyses. The coding sequence of the glyceraldehyde 3-phosphate dehydrogenase (GAPDH) was used as the control target. Suitable corresponding primer pairs were derived and selected based on the localization in different exons. The sequences of all primers are listed in Table 1.

Table 1.
Sequences of forward and reverse primer pairs that were used for qRT-PCR amplification of target genes (TfUT-A3 and TfUT-B) and the reference gene (GAPDH).
2.9. qRT-PCR Reactions and Gene Expression Analyses
qRT-PCR reactions were performed on an Applied Biosystems 7900 HT Fast Real-Time PCR System (Thermo Fisher Scientific, Schwerte, Germany) with the Fast Evagreen qPCR master mix (Biotrend, Cologne, Germany). The master mix was used in a 0.75× concentration. Further components of the reaction mix were 1× ROX as the reference dye, 0.5 µmol/L of both the forward and the reverse primer, and 60 pg (for GAPDH and TfUT-A3) or 1.5 ng (for TfUT-B) of the template cDNA. The reaction volume of 10 µL was applied on an Applied Biosystems MicroAmp Optical 96-Well Reaction Plate (Thermo Fisher Scientific, Schwerte, Germany). The PCR program consisted of an initial denaturation step (95 °C for 2 min) followed by 45 cycles of 95 °C for 10 s, 60 °C for 10 sec, and 72 °C for 15 sec each. Fluorescence data were collected at the end of each cycle. A dissociation curve analysis was used after each qRT-PCR reaction to ensure the specificity of the PCR product and the quality of each reaction. Standard curve testing was performed using a cDNA mixture of all samples in a series of ten-fold dilutions and was repeated with a series of five-fold dilutions for all targets.
The qRT-PCR results were analyzed according to Pfaffl [37]. The Ct average of all animals in the experimental control conditions from one population was used as the reference (calibrator). The Ct values for each biological replicate were subtracted from the calibrator of the respective gene and snail population (ΔCt). The relative quantity (RQ) was calculated using the primer efficiency (E) as a base to the ΔCt as an exponent. The ratio of gene expression was obtained by dividing the RQ of the gene of interest (GOI) by the housekeeping gene GAPDH.
2.10. Statistical Analyses
The transcript expression fold changes of experimental groups compared to the control group were logarithmized and tested for their normal distribution using the Kolmogorov–Smirnov test followed by Levene’s test to evaluate the homogeneity of variance. Differences between the mean values of the experimental groups and the mean values of the corresponding control groups were tested for statistical significance. If variances were equal between groups, the two-sided t-test was used to test for significant differences in means, otherwise the Welch’s t-test was used. Significant differences in means were presented as p < 0.05 (*), p < 0.01 (**), or p < 0.001 (***).
3. Results
3.1. Identification of Putative DUR3-like Urea Transporter Sequences
As a basis for the current manuscript and also for future research projects, we generated both the first transcriptome sequence and the first preliminary genome sequence dataset of Theodoxus fluviatilis. Whereas the transcriptome sequence dataset was generated from a pool of 24 individuals of both the freshwater and the brackish water ecotypes of T. fluviatilis, the preliminary genome dataset was generated from a single individual of the brackish water ecotype. The transcriptome sequence data have already been uploaded to GenBank both as raw (SRR15300633) and as assembled data (GKDH00000000.1). The preliminary genome dataset is still under revision; a respective manuscript, however, is currently in preparation (Fuchs et al., in preparation)
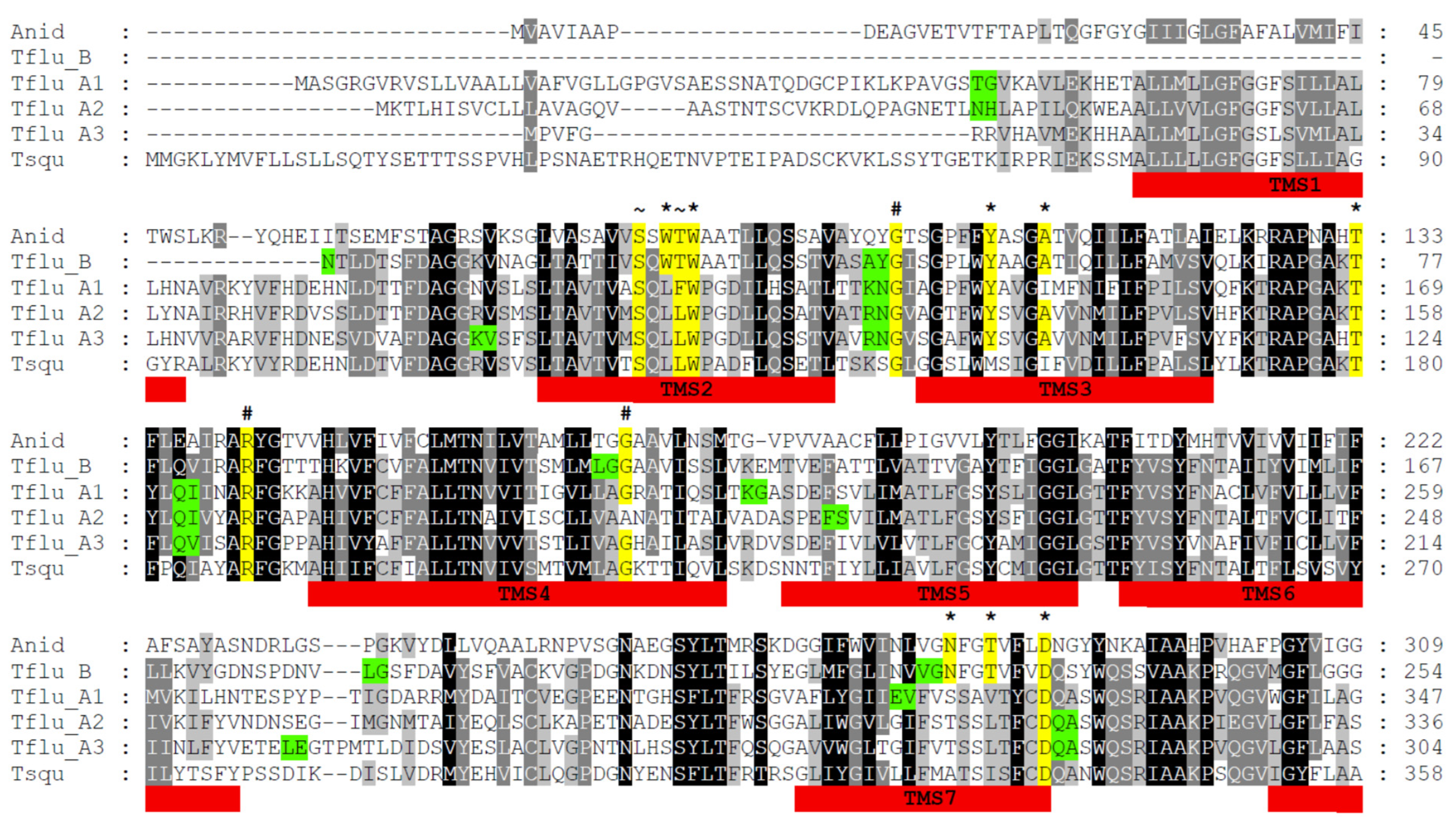
A thorough analysis of the transcriptome data revealed strong evidence for the expression of at least four different DUR3-like urea transporters in T. fluviatilis, namely TfUT-A1-A3 and TfUT-B. Where necessary, complete consensus sequences were manually generated and assembled from overlapping transcriptome contigs, whereas the genome contigs served as controls. All final sequences were uploaded to GenBank and are available under the accession numbers OP762683, OP762684, OP762685, and OP762686, respectively. A multiple-sequence alignment of all four putative DUR3-like urea transporters of T. fluviatilis together with respective sequences of T. squamosa and Aspergillus nidulans (Winter, 1884) (Eurotiales: Trichocomaceae) is provided in Figure 1.
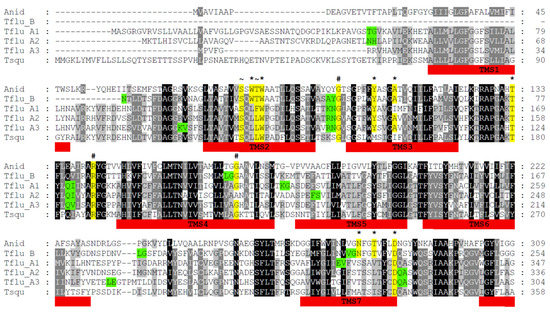

Figure 1.
Multiple-sequence alignment of DUR3-like urea transporters of Aspergillus nidulans (Anid), Tridacna squamosa (Tsqu), and Theodoxus fluviatilis (Tflu_A1, A2, A3, and B). The black background indicates conserved amino acid residues; the gray background indicates similar residues. The red bars mark the position of transmembrane segments (TMS). The yellow background indicates highly conserved residues that are involved in urea binding and translocation (*), protein folding and structure (#), and Na+/H+-binding (~). The green background indicates amino acid residues with coding triplets that comprise introns in the respective gene sequences.
The multiple-sequence alignment indicates that the four putative DUR3-like urea transporters of T. fluviatilis very likely belong to two different families, named type A (with the members A1, A2, and A3) and type B (with only one member). The overall degree of sequence identity/similarity among the TfUT-A family members is between 44–52% and 63–68% compared to 28–30% and 45–48% for TfUT-B. Interestingly, the DUR3-like urea transporter of T. squamosa displays higher degrees of sequence identity/similarity to the TfUT-A family members (37–41%/60–61%) compared to the TfUT-B family member (26%/41%). In contrast, the DUR3-like urea transporter of A. nidulans exhibits almost the same degrees of sequence identity/similarity to members of both families (18–24%/34–40%).
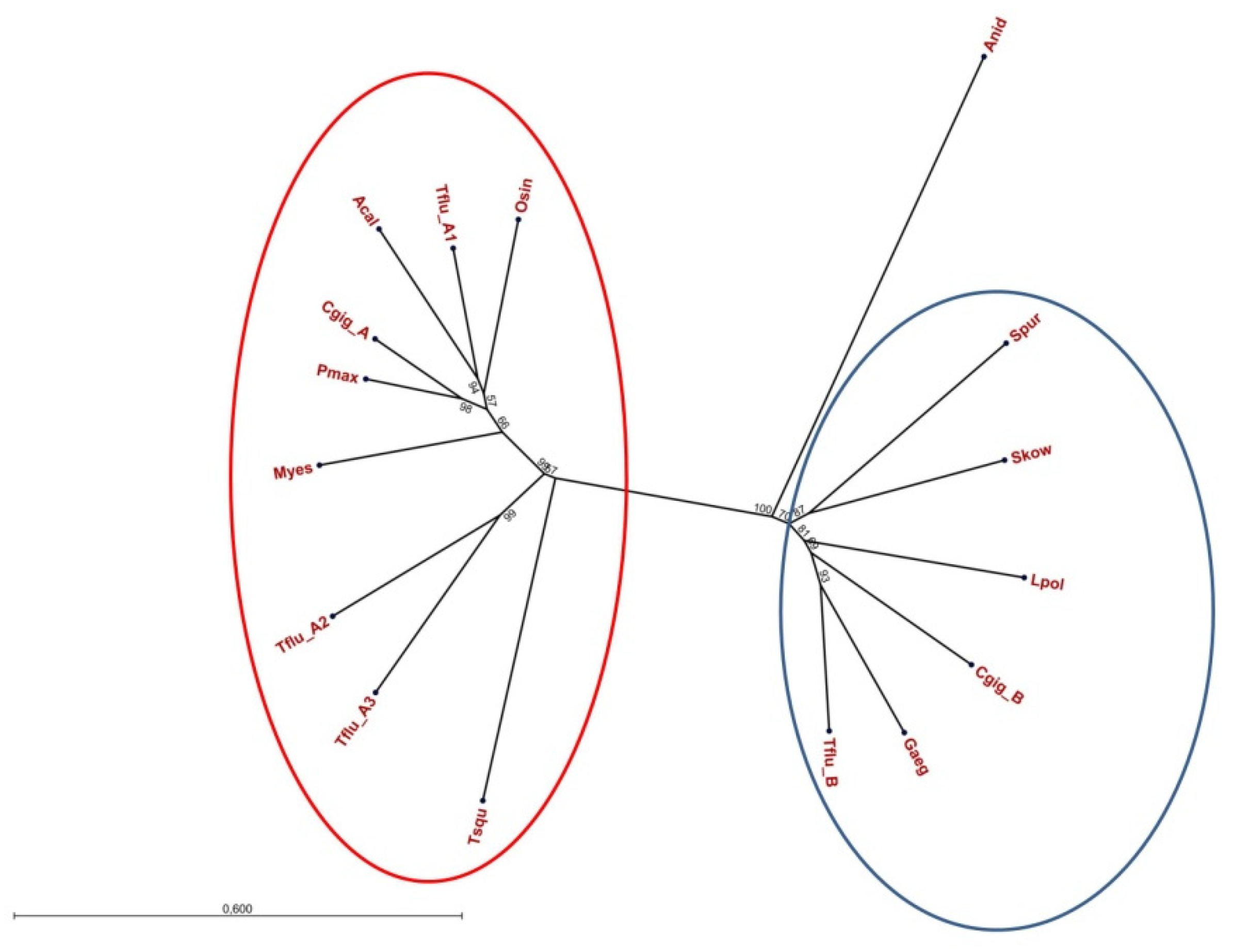
To evaluate whether or not the two families of DUR3-like urea transporters are a feature of the genus Theodoxus only or are present in other invertebrate genera as well, we collected respective protein sequences from various GenBank entries and performed a phylogenetic analysis. The resulting unrooted tree is shown in Figure 2 and supports the presence of two families of DUR3-like urea transporters in invertebrates. Interestingly, also for Crassostrea gigas (Thunberg, 1793) (Ostreoida: Ostreidae) (the Pacific oyster), members of both families have been identified (GenBank entries XP_034300843.1 for CgUT-A and XP_011441136.2 for CgUT-B).
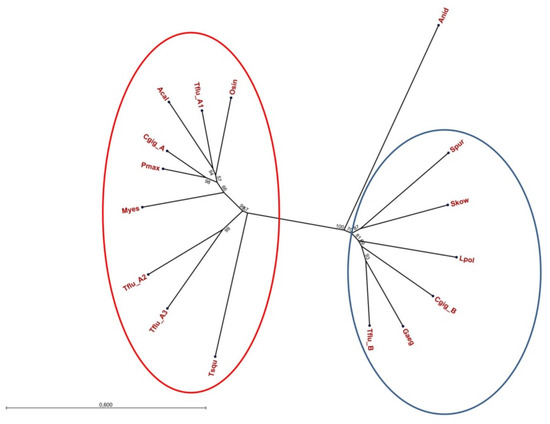
Figure 2.
Unrooted phylogenetic tree of invertebrate DUR3-like urea transporters. The red ellipse encloses UT-A type transporters and the blue ellipse encloses UT-B type transporters. The digits at the nodes represent the bootstrap values after 10,000 replications. Tflu = Theodoxus fluviatilis; Cgig-A = Crassostrea gigas XP_034300743.1; Cgig-B = Crassostrea gigas XP_01441136.2; Lpol = Limulus polyphemis XP_13776744.1; Skow = Saccoglossus kowalevskii XP_006823326.1; Spur = Strongylocentrotus purpuratus XP_030829061.1; Gaeg = Gigantopelta aegis XP_041377330.1; Tsqu = Tridacna squamosa ARQ84970.1; Osin = Octopus sinensis XP_029639211.1; Myes = Mizuhopecten yessoensis OWF45146.1; Pmax = Pecten maximus XP_033744244.1; Acal = Aplysia californica XP_035827281.1; Anid = Aspergillus nidulans ACZ62639.1.
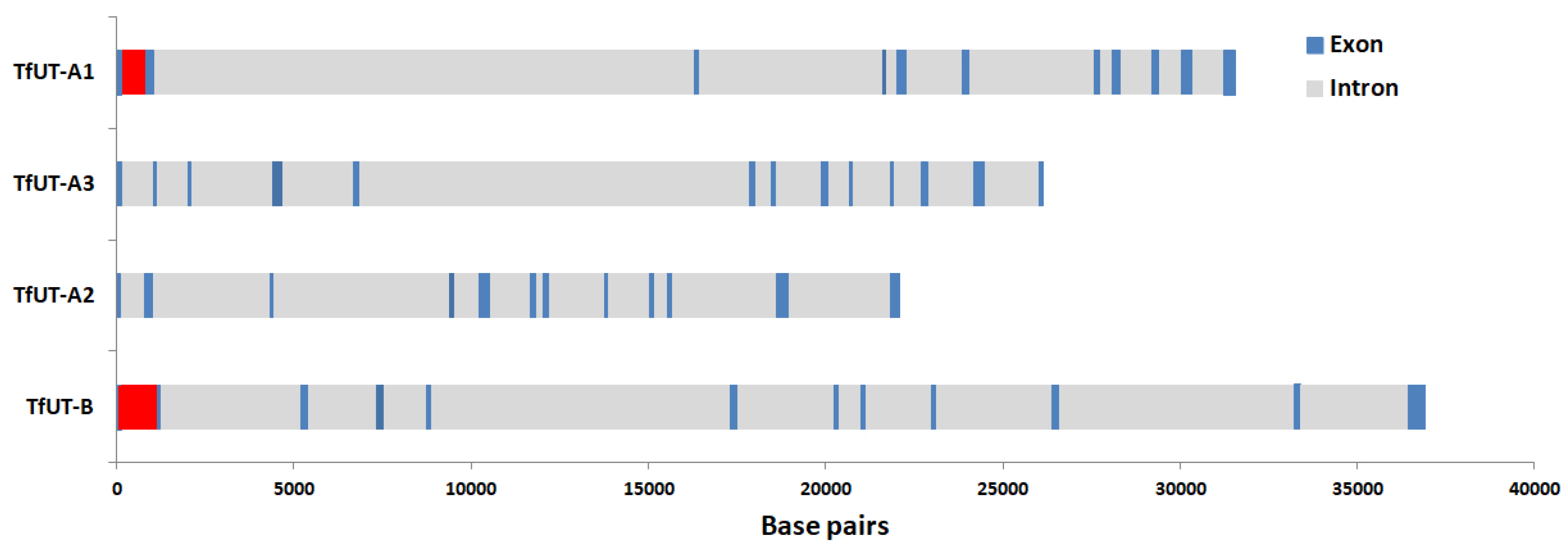
As already pointed out in the Introduction section, vertebrates express two types of urea transporters as well, namely UT-A and UT-B. Whereas six isoforms of UT-A exist, for UT-B, two (almost identical) isoforms have been described. However, both for UT-A and UT-B, the respective isoforms are encoded by the same gene and hence represent different splice variants [12,13]. In contrast, each of the three isoforms of TfUT-A and TfUT-B of T. fluviatilis is encoded by an individual gene. The structures of all four genes were determined and are given in Figure 3. Additionally, we have identified the exact localizations of all introns within the four gene sequences and marked them according to the positions of the respective amino acid residues (see Figure 1). The localizations of three introns are conserved among all TfUT genes, the localizations of two additional ones are conserved among the TfUT-A genes, and one is conserved among the TfUT-A1/A2, TfUT-A1/3, and TfUT-A2/3 genes each, respectively (see Figure 1 for details).
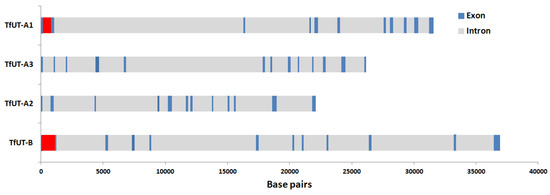
Figure 3.
Schematic representation of the gene structures of the DUR3-like urea transporters TfUT-A1–A3 and TfUT-B in Theodoxus fluviatilis. Size and position of exons (blue) and introns (grey) are marked. Red indicates uncertainty (extended repetitive sequences, TfUT-A1) or lack of data (TfUT-B).
3.2. Structural Features of Putative DUR3-like Urea Transporters of T. fluviatilis
DUR3-like urea transporters are membrane proteins and contain 14 to 15 transmembrane segments (TMSs) [18]. Computational structure analyses for all four putative DUR3-like urea transporters of T. fluviatilis using PSIPRED concordantly revealed the presence of 15 TMSs for TfUT-A1–A3 und 14 for TfUT-B (see Figure 1, TMSs are numbered and highlighted in red). The position and lengths of TMSs in TfUT-As and TfUT-B of T. fluviatilis are in agreement with the corresponding domains in UreA of A. nidulans [18].
In addition to the overall structure, several amino acid residues are of particular importance for the correct function of DUR3-like urea transporters including substrate binding and translocation and folding and membrane sorting [18]. An in-depth analysis revealed that most, but not all, of these functionally relevant residues are present in TfUT-As and TfUT-B of T. fluviatilis. In detail, this encompasses residues that are supposed to be involved in urea binding and translocation (W120, Y142, A/I146, T169, D324, Y429, and W549; marked with an * in Figure 1, numbers are according to UreA of A. nidulans), in protein folding and structure (G99, R141, G168, and P639, marked with a # in Figure 1), and in Na+/H+-binding (S80, L/T/F83, A/S370, S373, and T/A374, marked with a ~ in Figure 1), respectively. However, the residues W82, N279, T282, Y437, and S446 of UreA are missing in TfUT-A1-A3 and the DUR3-like urea transporter of T. squamosa, but are present in TfUT-B. For W82, N279, and T282, direct involvement in urea binding and translocation has recently been shown [38], whereas Y437 seems to be involved in substrate selectivity and S446 is necessary for the correct sorting of UreA to the membrane [39].
Although we have not yet performed any functional tests with the T. fluviatilis proteins, we are nevertheless confident that the putative DUR3-like urea transporters of T. fluviatilis are very likely functional proteins.
3.3. Effects of Osmotic Stress on Transcript Levels of TfUT-A3 and TfUT-B
In the course of physiological adaptation to higher salinities, individuals of T. fluviatilis accumulate organic osmolytes including larger quantities of urea. In addition, individuals of the brackish water ecotype exhibit a considerably higher base level of urea in their body fluids compared to individuals of the freshwater ecotype [30]. In order to obtain a better understanding of the regulatory mechanisms behind these observations, we conducted qRT-PCR experiments to quantify the relative mRNA expression of two types of putative DUR3-like urea transporters, namely TfUT-A3 and TfUT-B. Individuals of the brackish water ecotype were exposed to both hyper- (28‰ salinity) and hypoosmotic (0.5‰ salinity) conditions, whereas individuals of the freshwater ecotype were exposed to hyperosmotic (18‰ salinity) conditions only. Individuals that were kept under their natural conditions (8‰ salinity for BW and 0.5‰ salinity for FW individuals, respectively) served as the controls.
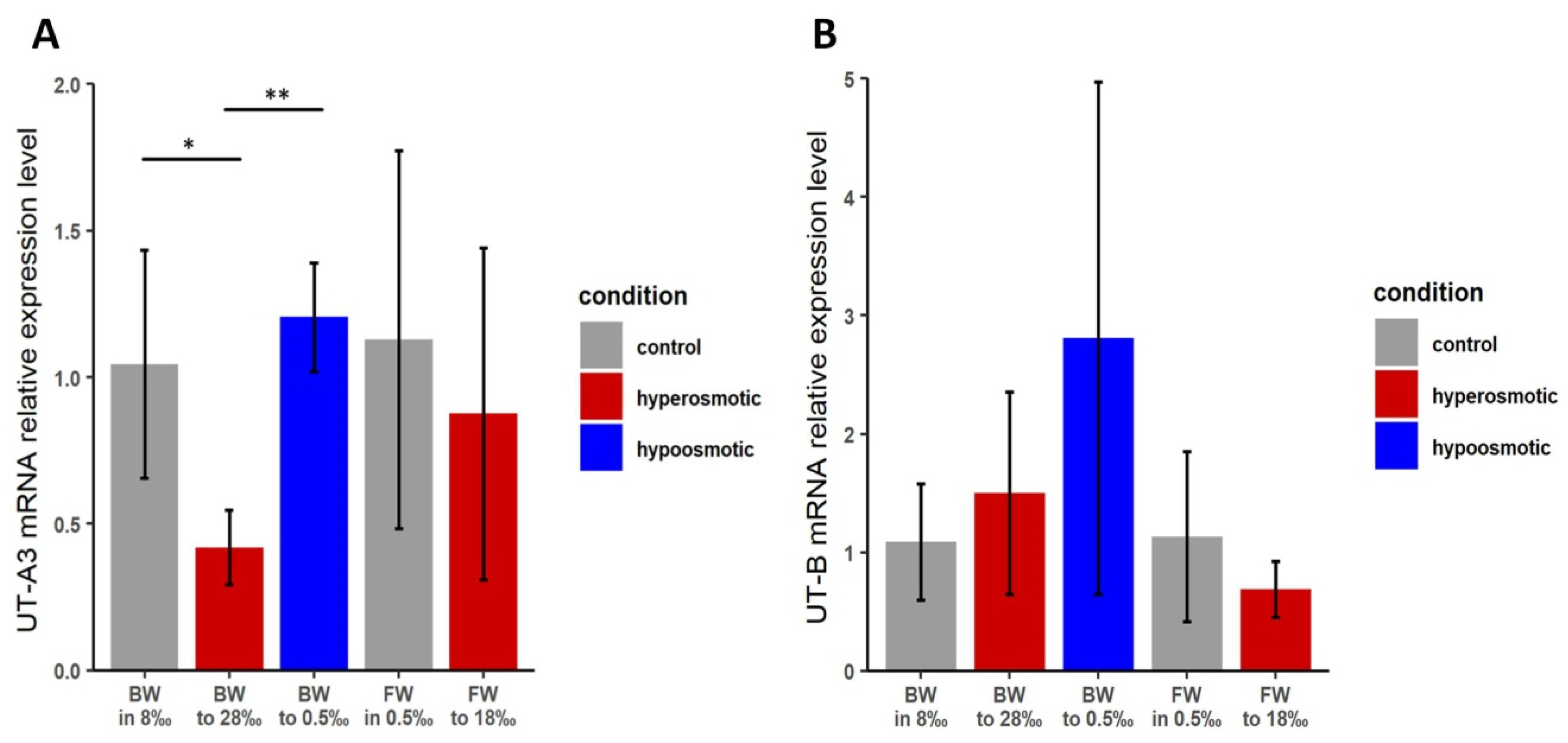
The qRT-PCR data for TfUT-A3 revealed a significant (p < 0.05) decrease in mRNA expression when brackish water individuals of T. fluviatilis were exposed to higher salinities. A slight but statistically insignificant increase in mRNA expression occurred upon the transfer to freshwater conditions (Figure 4A). For individuals of the freshwater ecotype, a slight but statistically non-significant increase in mRNA expression of TfUT-A3 could be observed. Strikingly, the normalized Ct-values for TfUT-A3 mRNA expression in freshwater individuals were approximately 1.5–2.2 cycles higher compared to brackish water individuals, indicating a generally lower base level of TfUT-A3 expression in freshwater individuals.
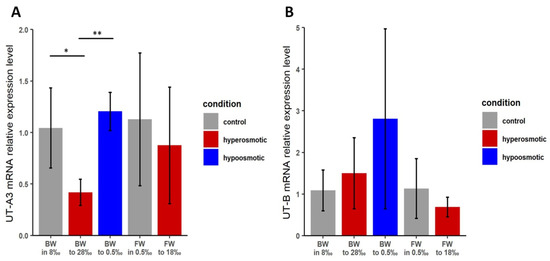
Figure 4.
Relative expression levels of urea transporter mRNAs in whole-body tissue preparations of brackish water and freshwater individuals of Theodoxus fluviatilis upon transfer to different salinities for TfUT-A3 (A) and TfUT-B (B). * p < 0.05; ** p < 0.005; error bars indicate standard deviation; n = 4 with 3 technical replicates each.
For the expression of TfUT-B mRNA, our analyses did not reveal any statistically significant changes in mRNA expression in either ecotype or kind of salinity change (Figure 4B). However, there is a certain increase in the mRNA expression of TfUT-B in brackish water individuals following a transfer to lower salinities, and it might be that the high degree of biological variability (as indicated by the large standard deviation bars) obscures any effect that may actually be present.
When both ecotypes were kept under control conditions (8‰ salinity for BW and 0.5‰ salinity for FW individuals), the normalized Ct-values for TfUT-A3 differed markedly between the BW (0.9 on average) and the FW (2.9 on average) individuals, whereas the respective value for TfUT-B was the same for both ecotypes (1.9).
4. Discussion
Urea has long been known as a key osmolyte in the body fluids of elasmobraches to maintain an osmotic balance with the external medium [5,40,41]. However, terrestrial snails also accumulate urea during aestivation and desiccation to prevent water loss [42,43,44]. For the first time, Wiesenthal et al. [30] described the use of urea as an organic osmolyte in the course of adaptation to higher salinities in aquatic snails, namely the euryhaline nerite snail T. fluviatilis. However, striking quantitative differences could be observed in the use of urea during osmotic adaptation between the two ecotypes of T. fluviatilis. It was hence very likely that distinct molecular mechanisms exist that take responsibility for the observed differences.
Urea uptake, release, or retention is influenced by membrane permeability and specific urea transporters [10,45,46]. UT-A and UT-B are the major urea transporters in vertebrates [12,13,47], whereas in plants [19], yeast/fungi [15,18], and some mollusc species [20,25], the DUR3 proteins fulfil the respective functions.
In the current manuscript, we describe the identification and molecular characterization of putative DUR3-like protein coding sequences in T. fluviatilis. Four different coding sequences (Figure 1) and the respective genes (Figure 3) representing two families of putative DUR3-like urea transporters could be identified in transcriptome and genome sequence datasets, named TfUT-A and TfUT-B, respectively. Phylogenetic analyses strongly supported the presence of two DUR3 families in invertebrates (see Figure 2). Members of both families have so far been identified both in T. fluviatilis and in C. gigas, but the presence of isoforms (TfUT-A1–A3) is currently a unique feature in T. fluviatilis. It is important to note in that context that both the families and the isoforms are present in each of the two ecotypes (FW and BW) of T. fluviatilis. In addition, the isoforms of TfUT-A do not represent splice variants of one gene as it is described for the vertebrate urea transporters but are encoded by separate genes. Similar observations for the presence of families and isoforms of DUR3-like urea transporters have been described for the fungus A. nidulans [18] and the marine macroalga Pyropia yezoensis [48].
DUR3-like urea transporters are transmembrane proteins, and a PSIRED MEMSAT-SVM analysis of TfUT-A1–A3 and TfUT-B revealed the presence of 15 (TfUT-A isoforms) or 14 (TfUT-B, 5′-incomplete sequence) TMSs (see Figure 1), as expected. The number of TMSs matches the respective values for DUR3-like urea transporters of T. squamosa [20], A. nidulans [18] and other yeast/fungi [39].
We have not yet functionally characterized the putative DUR3-like urea transporters of T. fluviatilis, but the presence of most of the amino acid residues that were reported to be essential for the function of UreA, the respective DUR3 protein in A. nidulans [18], and that are present in DUR3-like of T. squamosa [20] (see Figure 1) strongly indicates functionality. In a recent publication, however, Sanguinetti et al. [38] convincingly demonstrated that the amino acid residues W82, W84, N279, and T282 in UreA of A. nidulans are directly involved in urea binding and translocation. All four amino acid residues occur in TfUT-B, but only W84 is present in TfUT-A1-A3 and DUR3-like of T. squamosa (see Figure 1). Moreover, a multiple-sequence alignment of various invertebrate DUR3-like UTs revealed that the full W/W-N/T motif is present in all members of the UT-B type transporters, but not in members of the UT-A type (Supplementary Material Figure S2). Hence, the UT-A type transporters either do not transport urea (but a different solute) or work differentially compared to UreA. The first alternative is very unlikely since the DUR3-like UT of T. squamosa is clearly a UT [20]. The second alternative touches on the question of what monovalent cation gradient may actually drive the transport of urea. DUR3s can facilitate active uptake of urea, but in contrast to urea-binding ABC-type transporters found, e.g., in cyanobacteria [49], they lack an ATP-binding site. Instead, urea transport by DUR3s seems to be a secondary transport that is energized by gradients for monovalent cations across the membranes [15,22]. Whereas the urea transport by AtDUR3 of Arabidopsis thaliana (L.) Heynh. (Brassicales: Brassicaceae) requires a proton gradient for urea translocation [22], DUR3-like of T. squamosa comprises conserved domains that correspond to the sodium solute carrier 5- and 6-like families [20]. It might well be that the transport activities of DUR3-like UT-A type and UT-B type transporters rely on different ion gradients, namely on a proton gradient for UT-B type and on a sodium ion gradient for UT-A type transporters, but this hypothesis remains to be confirmed by experimental evidence.
The identification of putative DUR3-like coding sequences opened up the possibility to further evaluate the molecular mechanisms behind the remarkable capability of T. fluviatilis to acclimate to changing environmental salinities. Of special interest was the question of whether or not the accumulation of urea during a hyperosmotic challenge [30] is accompanied by equivalent changes in the expression of the respective urea transporters. Unfortunately, suitable antibodies to detect the proteins in vitro or in vivo [20] are not available. Thus, we decided to perform qRT-PCR experiments instead to obtain at least an impression of possible changes in mRNA abundances. TfUT-A3 and TfUT-B as representatives of both families of DUR3-like urea transporters were selected, and an appropriate experimental setup (including primer pairs that span at least one intron–exon boundary) was developed. Our data indicate that under control conditions, the expression of TfUT-A3 is more prominent in brackish water (BW) individuals compared to freshwater (FW) individuals, whereas the expression of TfUT-B is approximately the same in both ecotypes. In BW individuals, the expression of TfUT-A3 was significantly decreased upon transfer from normal conditions (8‰ salinity) to hyperosmotic conditions (28‰ salinity) and slightly increased upon transfer to hypoosmotic conditions (0.5‰ salinity) (Figure 4A). Only minor changes in TfUT-A3 mRNA expression could be observed in FW individuals.
The downregulation of TfUT-A3 expression in brackish water individuals under hyperosmotic conditions may be interpreted in different ways. If the expression of this transporter occurs in the gills or in the integument, its main function under basal environmental conditions might be the facilitation of urea excretion. It would make sense to downregulate such a transporter if urea accumulation in the body fluids is required under hyperosmotic stress conditions. The decrease in TfUT-A3 mRNA and subsequently also in protein abundances may prevent the loss of urea to the environment and hence facilitate the increase in body fluid urea concentration as was observed by Wiesenthal et al. [30]. Upregulation of TfUT-A3 may be beneficial under hypoosmotic conditions: An increase in TfUT-A3 expression allows for a more rapid release of urea to the environment to reduce internal osmolality. A trend that may point to such a mechanism could be seen when BW animals were subjected to hypoosmotic stress (Figure 4A); however, the observed changes did not reach statistical significance compared to the controls.
For FW individuals of T. fluviatilis, only a slight decrease in TfUT-A3 mRNA expression under hyperosmotic conditions could be observed. In light of the much less prominent role of urea as an organic osmolyte in FW individuals and the generally lower expression of TfUT-A3 in FW individuals compared to BW individuals, minor changes in the urea transporter expression might already be sufficient to adjust the internal urea concentration to the changing demands. For the expression of TfUT-B, the qRT-PCR data suffer from exceptionally high standard deviation errors (Figure 4B) that prevent any meaningful statistical analysis. The increase in TfUT-B mRNA expression in brackish water individuals upon transfer to hypoosmotic conditions, nevertheless, supports the assumption that higher abundances of urea transporters may help to quickly get rid of an excess of urea.
However, two limitations of our experimental approach have to be taken into account: (I) We used whole body tissue and (II) we did not test for the expression of TfUT-A1 and 2. It might well be that the expression of urea transporters differs across tissues and may be more prominent in gills that are exposed to the environment and are hence more likely to be involved in urea release/retention and uptake mechanisms. Light-regulated expression of a DUR3-like urea transporter has been detected in the ctenidium (gills) of the fluted giant clam, T. squamosa, which facilitates urea uptake from the environment [20]. The analysis of tissue-specific expression patterns of DUR3-like urea transporters in T. fluviatilis (e.g., by fluorescence in situ hybridization (FISH), immune fluorescence, or Western blotting experiments) would certainly help to obtain a better understanding of the complex regulatory processes during the acclimation of animals to changing environmental salinities.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/physiologia3020020/s1, Figure S1: Transfer regimes used to acclimate Theodoxus fluviatilis individuals to media with different salinities; Figure S2: Multiple sequence alignment of putative invertebrate DUR3-like urea transporters.
Author Contributions
J.K., C.M. and J.-P.H. conceived the ideas and designed the methodology; J.K. and S.G. performed the experiments; J.K., L.I.R.F., J.F., M.T.-O. and C.M. generated and analyzed the sequence data; J.K. and C.M. analyzed the data and drafted the manuscript; C.M. and J.-P.H. supervised the experimental work. All authors have read and agreed to the published version of the manuscript.
Funding
J.K. and L.I.R.F. received stipends and financial support from the German Research Foundation (DFG), RTG 2010 (RESPONSE). This work was supported by the DFG Research Infrastructure NGS_CC (project 407495230) as part of the Next-Generation Sequencing Competence Network (project 423957469). NGS analyses were carried out at the Competence Centre for Genomic Analysis (Kiel).
Institutional Review Board Statement
Ethical review and approval were waived for this study because it did not include the use of humans, vertebrates, cephalopods or crustaceans.
Data Availability Statement
Original data are deposited at BioStudies (EMBL-EBI) (https://www.ebi.ac.uk/biostudies/, accessed on 15 February 2023) under the accession number S-BSST1031.
Acknowledgments
The authors are thankful to all members of the Animal Physiology Working Group at the University of Greifswald for their help and support. Special thanks go to Kristina Kuprina and Alexander Wacker (Animal Ecology, University Greifswald) and to Mladen Tzvetkov (Dept. of General Pharmacology, University Medicine Greifswald) for technical support. Permissions for snail collections were given by the Nationalparkamt Vorpommersche Boddenlandschaft, Born, Germany (24–5303.3).
Conflicts of Interest
The authors declare that they have no conflict of interest.
Ethical Approval
We declare that the experiments described in this paper comply with the current laws in Germany. All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
References
- Wang, H.; Ran, J.; Jiang, T. Urea. In Urea Transporters in Subcellular Biochemistry; Yang, B., Sands, J.M., Eds.; Springer: Dordrecht, The Netherlands, 2014; pp. 7–29. [Google Scholar] [CrossRef]
- Oja, S.S.; Saransaari, P.; Korpi, E.R. Neurotoxicity of ammonia. Neurochem. Res. 2017, 42, 713–720. [Google Scholar] [CrossRef] [PubMed]
- Pallone, T.L.; Turner, M.R.; Edwards, A.; Jamison, R.L. Countercurrent exchange in the renal medulla. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2003, 284, R1153–R1175. [Google Scholar] [CrossRef] [PubMed]
- Browning, J. Urea levels in plasma and erythrocytes of the southern fiddler skate, Trygonorhina fasciata guanerius. J. Exp. Biol. 1978, 203, 325–329. [Google Scholar] [CrossRef]
- Hazon, N.; Wells, A.; Pillans, R.D.; Good, J.P.; Anderson, W.G.; Franklin, C.E. Urea based osmoregulation and endocrine control in elasmobranch fish with special reference to euryhalinity. Comp. Biochem. Physiol. B 2003, 136, 685–700. [Google Scholar] [CrossRef]
- Li, C.; Wang, W. Urea transport mediated by aquaporin water channel proteins. In Urea Transporters in Subcellular Biochemistry; Yang, B., Sands, J.M., Eds.; Springer: Dordrecht, The Netherlands, 2014; pp. 227–265. [Google Scholar] [CrossRef]
- McDonald, M.D.; Wood, C.M. Evidence for facilitated diffusion of urea across the gill basolateral membrane of the rainbow trout (Oncorhynchus mykiss). Biochim. Biophys. Acta 2004, 1663, 89–96. [Google Scholar] [CrossRef]
- Kato, A.; Sands, J.M. Active sodium-urea counter-transport is inducible in the basolateral membrane of rat renal initial inner medullary collecting ducts. J. Clin. Investig. 1998, 102, 1008–1015. [Google Scholar] [CrossRef] [PubMed]
- Morgan, R.L.; Wright, P.A.; Ballantyne, J.S. Urea transport in kidney brush-border membrane vesicles from an elasmobranch, Raja erinacea. J. Exp. Biol. 2003, 206, 3293–3302. [Google Scholar] [CrossRef] [PubMed]
- Fines, G.A.; Ballantyne, J.S.; Wright, P.A. Active urea transport and an unusual basolateral membrane composition in the gills of a marine elasmobranch. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2001, 280, R16–R24. [Google Scholar] [CrossRef]
- McDonald, M.D.; Smith, C.P.; Walsh, P.J. The physiology and evolution of urea transport in fishes. J. Membr. Biol. 2006, 212, 93–107. [Google Scholar] [CrossRef]
- Smith, C.P. Mammalian urea transporters. Exp. Physiol. 2009, 94, 180–185. [Google Scholar] [CrossRef]
- Sands, J.M.; Blount, M.A. Genes and proteins of urea transporters. In Urea Transporters in Subcellular Biochemistry; Yang, B., Sands, J.M., Eds.; Springer: Dordrecht, The Netherlands, 2014; pp. 45–63. [Google Scholar] [CrossRef]
- Levin, E.J.; Cao, J.; Enkavi, G.; Quick, M.; Pan, Y.; Tajkhorshid, E.; Zhou, M. Structure and permeation mechanism of a mammalian urea transporter. Proc. Natl. Acad. Sci. USA 2012, 109, 11194–11199. [Google Scholar] [CrossRef] [PubMed]
- Sumrada, R.; Gorski, M.; Cooper, T. Urea transport-defective strains of Saccharomyces cerevisiae. J. Bacteriol. 1976, 125, 1048–1056. [Google Scholar] [CrossRef] [PubMed]
- ElBerry, H.M.; Majumdar, M.L.; Cunningham, T.S.; Sumrada, R.A.; Cooper, T.G. Regulation of the urea active transporter gene (DUR3) in Saccharomyces cerevisiae. J. Bacteriol. 1993, 175, 4688–4698. [Google Scholar] [CrossRef] [PubMed]
- Navarathna, D.H.M.L.P.; Das, A.; Morschhäuser, J.; Nickerson, K.W.; Roberts, D.D. Dur3 is the major urea transporter in Candida albicans and is co-regulated with the urea amidolyase Dur1,2. Microbiology 2011, 157, 270–279. [Google Scholar] [CrossRef]
- Abreu, C.; Sanguinetti, M.; Amillis, S.; Ramon, A. UreA, the major urea/H+ symporter in Aspergillus nidulans. Fungal Genet. Biol. 2010, 47, 1023–1033. [Google Scholar] [CrossRef]
- Kojima, S.; Bohner, A.; von Wirén, N. Molecular mechanisms of urea transport in plants. J. Membr. Biol. 2006, 212, 83–91. [Google Scholar] [CrossRef]
- Chan, C.Y.L.; Hiong, K.C.; Boo, M.V.; Choo, C.Y.L.; Wong, W.P.; Chew, S.F.; Ip, Y.K. Light exposure enhances urea absorption in the fluted giant clam, Tridacna squamosa, and up-regulates the protein abundance of a light-dependent urea active transporter, DUR3-like, in its ctenidium. J. Exp. Biol. 2018, 221, jeb176313. [Google Scholar] [CrossRef]
- Cooper, T.G.; Sumrada, R. Urea transport in Saccharomyces cerevisiae. J. Bacteriol. 1978, 121, 571–576. [Google Scholar] [CrossRef]
- Liu, L.H.; Ludewig, U.; Frommer, W.B.; von Wirén, N. AtDUR3 encodes a new type of high-affinity urea/H+ symporter in Arabidopsis. Plant Cell 2003, 15, 790–800. [Google Scholar] [CrossRef]
- Kojima, S.; Bohner, A.; Gassert, B.; Yuan, L.X.; von Wiren, N. AtDUR3 represents the major transporter for high-affinity urea transport across the plasma membrane of nitrogen-deficient Arabidopsis roots. Plant J. 2007, 52, 30–40. [Google Scholar] [CrossRef]
- Wang, W.H.; Köhler, B.; Cao, F.Q.; Liu, G.W.; Gong, Y.Y.; Sheng, S.; Song, Q.C.; Cheng, X.Y.; Garnett, T.; Okamoto, M.; et al. Rice DUR3 mediates high-affinity urea transport and plays an effective role in improvement of urea acquisition and utilization when expressed in Arabidopsis. New Phytol. 2011, 193, 432–444. [Google Scholar] [CrossRef]
- Hermida, M.; Robledo, D.; Diaz, S.; Costas, D.; Bruzos, A.L.; Blanco, A.; Pardo, B.G.; Martinez, P. The first high density genetic map of common cockle (Cerastoderma edule) reveals a major QTL controlling shell color variation. Sci. Rep. 2022, 12, 16971. [Google Scholar] [CrossRef]
- Shumway, S.E.; Freeman, R.F.H. Osmotic balance in a marine pulmonate, Amphibola crenata. Mar. Freshw. Behav. Physiol. 1984, 11, 157–183. [Google Scholar] [CrossRef]
- Henry, R.P.; McBride, C.J.; Williams, A.H. Responses of the marsh periwinkle, Littoraria (Littorina) irrorata to temperature, salinity and desiccation, and the potential physiological relationship to climbing behavior. Mar. Freshw. Behav. Physiol. 1993, 24, 45–54. [Google Scholar] [CrossRef]
- Jordan, P.J.; Deaton, L.E. Osmotic regulation and salinity tolerance in the freshwater snail Pomacea bridgesi and the freshwater clam Lampsilis teres. Comp. Biochem. Physiol. Part A Mol. Integr. Physiol. 1999, 122, 199–205. [Google Scholar] [CrossRef]
- Symanowski, F.; Hildebrandt, J.-P. Differences in osmotolerance in freshwater and brackish water populations of Theodoxus fluviatilis (Gastropoda: Neritidae) are associated with differential protein expression. J. Comp. Physiol. B 2010, 180, 337–346. [Google Scholar] [CrossRef] [PubMed]
- Wiesenthal, A.A.; Müller, C.; Harder, K.; Hildebrandt, J.-P. Alanine, proline and urea are major organic osmolytes in the snail Theodoxus fluviatilis under hyperosmotic stress. J. Exp. Biol. 2019, 222, 193557. [Google Scholar] [CrossRef] [PubMed]
- Knobloch, J.; Müller, C.; Hildebrandt, J.-P. Expression levels and activities of energy-yielding ATPases in the oligohaline neritid snail Theodoxus fluviatilis under changing environmental salinities. Biol. Open 2022, 11, bio059190. [Google Scholar] [CrossRef]
- Kolmogorov, M.; Yuan, J.; Lin, Y.; Pevzner, P.A. Assembly of long, error-prone reads using repeat graphs. Nat. Biotechnol. 2019, 37, 540–546. [Google Scholar] [CrossRef] [PubMed]
- Manni, M.; Berkeley, M.R.; Seppey, M.; Simão, F.A.; Zdobnov, E.M. BUSCO update: Novel and streamlined workflows along with broader and deeper phylogenetic coverage for scoring of eukaryotic, prokaryotic, and viral genomes. Mol. Biol. Evol. 2021, 38, 4647–4654. [Google Scholar] [CrossRef]
- Marchler-Bauer, A.; Anderson, J.B.; Cherukuri, P.F.; DeWeese-Scott, C.; Geer, L.Y.; Gwadz, M.; He, S.; Hurwitz, D.I.; Jackson, J.D.; Ke, Z.; et al. CDD: A Conserved Domain Database for protein classification. Nucleic Acids Res. 2005, 33, D192–D196. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Mizianty, M.J.; Kurgan, L. Prediction and analysis of nucleotide-binding residues using sequence and sequence-derived structural descriptors. Bioinformatics 2012, 28, 331–341. [Google Scholar] [CrossRef] [PubMed]
- Nicholas, K.B.; Nicholas, H.B., Jr. GeneDoc: A Tool for Editing and Annotating Multiple Sequence Alignments. Distributed by the Author 1997. Available online: https://nrbsc.org/gfx/genedoc (accessed on 17 February 2023).
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef] [PubMed]
- Sanguinetti, M.; Santos, L.H.S.; Dourron, J.; Alamón, C.; Idiarte, J.; Amillis, S.; Pantano, S.; Ramón, A. Substrate recognition properties from an intermediate structural state of the UreA transporter. Int. J. Mol. Sci. 2022, 23, 16039. [Google Scholar] [CrossRef]
- Sanguinetti, M.; Amillis, S.; Pantano, S.; Scazzocchio, C.; Ramon, A. Modelling and mutational analysis of Aspergillus nidulans UreA, a member of the subfamily of urea/H+ transporters in fungi and plants. Open Biol. 2014, 4, 140070. [Google Scholar] [CrossRef]
- Smith, H.W. The retention and physiological role of urea in Elasmobranchii. Biol. Rev. 1936, 11, 49–82. [Google Scholar] [CrossRef]
- Trischitta, F.; Faggio, C.; Torre, A. Living with high concentrations of urea: They can! Open J. Anim. Sci. 2012, 2, 32–40. [Google Scholar] [CrossRef]
- Horne, F.R.; Barnes, G. Reevaluation of urea biosynthesis in prosobranch and pulmonate snails. J. Comp. Physiol. 1970, 69, 452–457. [Google Scholar] [CrossRef]
- Rees, B.B.; Hand, S.C. Biochemical correlates of estivation tolerance in the mountainsnail Oreohelix (Pulmonata: Oreohelicidae). Biol. Bull. 1993, 184, 230–242. [Google Scholar] [CrossRef]
- Hiong, K.C.; Loong, A.M.; Chew, S.F.; Ip, Y.K. Increases in urea synthesis and the ornithine-urea cycle capacity in the Giant African Snail, Achatina fulica, during fasting or aestivation, or after the injection with ammonium chloride. J. Exp. Zool. A 2005, 303, 1040–1053. [Google Scholar] [CrossRef]
- Lippe, C. Urea and thiourea permeabilities of phospholipid and cholesterol bilayer membranes. J. Mol. Biol. 1969, 39, 669–672. [Google Scholar] [CrossRef] [PubMed]
- Bankir, L. Active urea transport in lower vertebrates and mammals. In Urea Transporters in Subcellular Biochemistry; Yang, B., Sands, J.M., Eds.; Springer: Dordrecht, The Netherlands, 2014; pp. 193–226. [Google Scholar] [CrossRef]
- LeMoine, C.M.R.; Walsh, P.J. Evolution of urea transporters in vertebrates: Adaptation to urea’s multiple roles and metabolic sources. J. Exp. Biol. 2015, 218, 1936–1945. [Google Scholar] [CrossRef] [PubMed]
- Kakinuma, M.; Suzuki, K.; Iwata, S.; Coury, D.A.; Iwade, S.; Mikami, K. Isolation and characterization of a new DUR3-like gene, PyDUR3.3, from the marine macroalga Pyropia yezoensis (Rhodophyta). Fish. Sci. 2016, 82, 171–184. [Google Scholar] [CrossRef]
- Valladares, A.; Montesinos, M.L.; Herrero, A.; Flores, E. An ABC-type, high-affinity urea permease identified in cyanobacteria. Mol. Microbiol. 2002, 43, 703–715. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).