1. Introduction
The field of cancer research was introduced to a new putative oncogene, TP53, relatively recently. However, later, TP53 was actively researched as a strong tumor suppressor and recognized for its ability to drive the apoptotic demise of cancer cells [
1]. TP53 is a tumor suppressor gene and represents the most commonly mutated locus in human cancers [
2]. The TP53 gene spans approximately 16–20 kb of cellular DNA and is located on the short arm of human chromosome 17 at position 17p13.1. It encodes p53, a 375-amino acid nuclear phosphoprotein that functions as a transcription factor, regulating the expression of a wide variety of genes involved in cell cycle arrest and apoptosis in response to genotoxic or cellular stress [
3,
4]. The p53 tumor suppressor belongs to a multi-protein family that also includes the transcription factors p63 and p73. These proteins recognize the same DNA consensus sequences and can activate overlapping target genes, indicating potential functional redundancy among them [
5]. The TP53 gene was initially thought to function as a dominant oncogene, since its overexpression led to immortalization of rodent cells and cooperated with activated genes to transform rat embryonic fibroblasts. Subsequent studies revealed, however, that many of the p53 clones examined were actually mutated forms of the gene, whereas the wild-type p53 serves as a tumor suppressor. Loss of functional p53 has since been linked to cellular transformation in vitro and tumor development in vivo [
6]. p53 functions as a tumor suppressor by regulating cell division and preventing cells from proliferating too rapidly or uncontrollably. Located in the nucleus, p53 binds directly to DNA to carry out its regulatory role. When cellular DNA is damaged by factors such as toxic chemicals, radiation, or ultraviolet (UV) light, p53 plays a pivotal role in deciding whether the DNA will be repaired or the cell will undergo apoptosis. If repair is possible, p53 activates genes that facilitate DNA repair; if not, it halts cell division and directs the damaged cell toward programmed cell death. By preventing the proliferation of cells with irreparable mutations, p53 acts as a safeguard against tumor development. Owing to its central role in maintaining genomic stability, p53 is often referred to as the ‘guardian of the genome [
7].
H2AX is a variant of the histone H2A protein, distinguished by a unique C-terminal tail that is 14 amino acids longer than that of H2A [
8]. The total length of H2AX is 143 amino acids, resulting in a theoretical molecular weight of approximately 15 kDa [
9]. H2AX is incorporated in place of H2A in the histones throughout the DNA in a fairly random distribution [
10,
11,
12]. Two copies of each histone protein (H2A, H2B, H3, and H4) form an octamer [
13]. The histone helps the chromatin maintain a stable and compact structure [
13]. H2AX has gained interest because it can be used as a biomarker for double-stranded breaks (DSBs) in DNA [
11,
14,
15]. In the event of a double-strand break (DSB)—caused by factors such as ionizing radiation, reactive oxygen species (ROS), or chemotherapeutic agents [
16]—H2AX undergoes phosphorylation at serine 139 (four residues from the carboxyl terminus) [
14] by the ATM kinase or occasionally ATR or DNA-PK, generating γH2AX foci [
17,
18]. Techniques for detecting and quantifying γH2AX foci (and consequently the number of DSBs) include confocal microscopy, flow cytometry, and Western blot, along with emerging high throughput methods such as laser-scanning cytometry, and image flow cytometry [
15]. The primary antibody used to stain γH2AX is typically anti-phospho-Histone H2AX (Ser139) which can be paired with an Alexa Fluor 488 conjugated to secondary antibodies [
15]. Dissociation-Enhanced Lanthanide Fluorescent Immunoassay (DELFI) can also be used as a more sensitive approach to detect and differentiate low levels of DNA DSBs [
15]. γH2AX also helps recruit DNA repair and cell cycle proteins that localize at these foci including MRE11/RAD50/NBS1(MRN complex), 53BP1, MDC1, BRCA1, and RAD51 to facilitate DNA repair through non-homologous end joining (NHEJ) and homologous recombination (HR) [
17].
Graphene quantum dots (GQDs) are nanoscale carbon-based particles with exceptional chemical, physical, and biological properties, making them highly versatile for applications in nanomedicine [
19]. As a recent addition to the graphene family, GQDs have gained attention as promising materials in diverse biomedical applications owing to their unique physicochemical characteristics and excellent biocompatibility [
20]. The distinctive electronic structure of GQDs endows them with valuable functional properties, including strong and tunable photoluminescence for fluorescence bioimaging and biosensing, high loading capacity for aromatic compounds in small-molecule drug delivery, and efficient absorption of incident radiation for photothermal and photodynamic cancer therapies [
19].
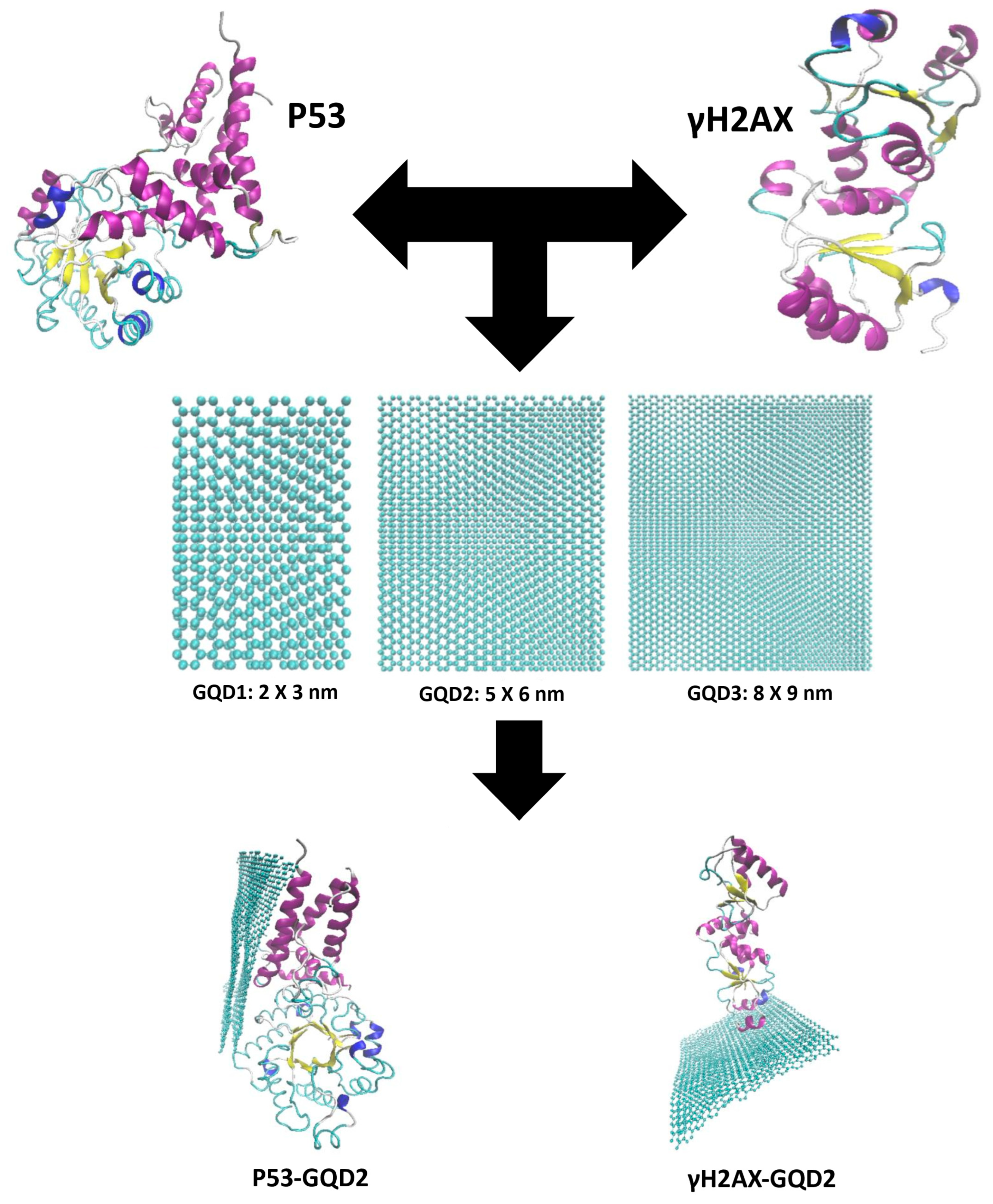
The purpose of this study is to investigate the effects of three different-sized two-layered GQDs, 2 × 3 nm (GQD1), 5 × 6 nm (GQD2), and 8 × 9 nm (GQD3), as they interact with γH2AX and p53 to form a molecular basis for isolating these two protein biomarkers based on their molecular weight (size) as shown in
Figure 1. The human proteome is estimated to contain over one million distinct protein types. Interactions between nanomaterials and biomolecules, such as DNA and proteins, can disrupt biological functions and potentially cause cytotoxicity. While graphene is known to disrupt the structure of polypeptides, protein fragments, and full proteins, GQDs offer advantages by generating intrinsic fluorescence, enhancing the aqueous stability of graphene oxide (GO), and retaining broad chemical versatility and high adsorption capacity. As the size of GQDs increases, more protein residues can adsorb onto their surface, resulting in greater protein adsorption. At the same time, larger GQDs tend to induce more pronounced alterations in the secondary structure of proteins [
21]. Our study is structured into two primary sections: the stability analysis and a more exhaustive analysis of the energies at the interface of the interacting GQDs and the two proteins, γH2AX and p53. For stability, we quantified several biophysical properties, including the root mean square deviation (RMSD), center of mass between the interacting molecules, analysis of the salt bridges and hydrogen bonds, conformational energy changes, secondary structure analysis, and the interaction energies between the adsorbed atoms (within 5 Å) and the surface of the GQDs. On the energetics front, we focus on the calculation of interaction energies, particularly van der Waals (VDW) and electrostatic energies, to understand the fundamental forces driving the protein-GQD interactions, with the exhaustive analysis further building upon this foundation by examining a range of additional parameters. These include the relationship between the number of atoms involved in the interaction and the distance, the variation in interaction energy with distance, and the ratio between these two, which we term the optimal distance, providing deeper insights into the effectiveness of protein-GQD binding. Moreover, we examine the role of interfacial water and its hydrogen bonds to evaluate their impact on the interactions at the interface. Finally, we conduct a thorough examination of the number of adsorbed residues for each case. This comprehensive analysis enables us to gain a detailed understanding of protein-GQD interactions, with critical implications for applications in nanobiotechnology, biosensing, and cancer research.
3. Results and Discussion
Trajectories from the 250 ns production runs were analyzed to assess the stability of the interacting molecules and to evaluate the interfacial energies. The trajectory screenshots at four different time intervals for the simulations between GQD1, GQD2, and GQD3 with γH2AX and p53 are provided in the
Supplementary Figures S1 and S2. This helped visualize the conformational evolution of protein–nanomaterial interactions over time, where a total of 24 trajectory images were captured—covering each protein in complex with each GQD—providing a comparative overview of the structural changes and binding behavior across the simulation timescale.
RMSD (Root Mean Square Deviation), which is a measure of the average deviation of atomic positions in a molecular system over time, is typically used to assess the structural stability and conformational changes in proteins, nucleic acids, or other biomolecules. A lower RMSD value suggests lesser structural deviation, whereas higher RMSD values indicate significant deviations. Furthermore, the fluctuations on the RMSD plot are indicative of the conformational changes in the molecule. Analyzing the RMSD of p53 and γH2AX proteins conjugated with GQDs of varying sizes provided valuable insights into how size affects the stability and conformational dynamics of these proteins. Towards understanding the stability of these proteins as they adsorb onto the GQDs,
Figure 2 shows the RMSD of the proteins p53 and γH2AX in the presence of GQD1, GQD2, and GQD3 (
Figure 2A), RMSD of the three GQDs in the presence of the two proteins (
Figure 2B,C), and the variation in the distance between the center of masses of the GQDs and the two proteins (
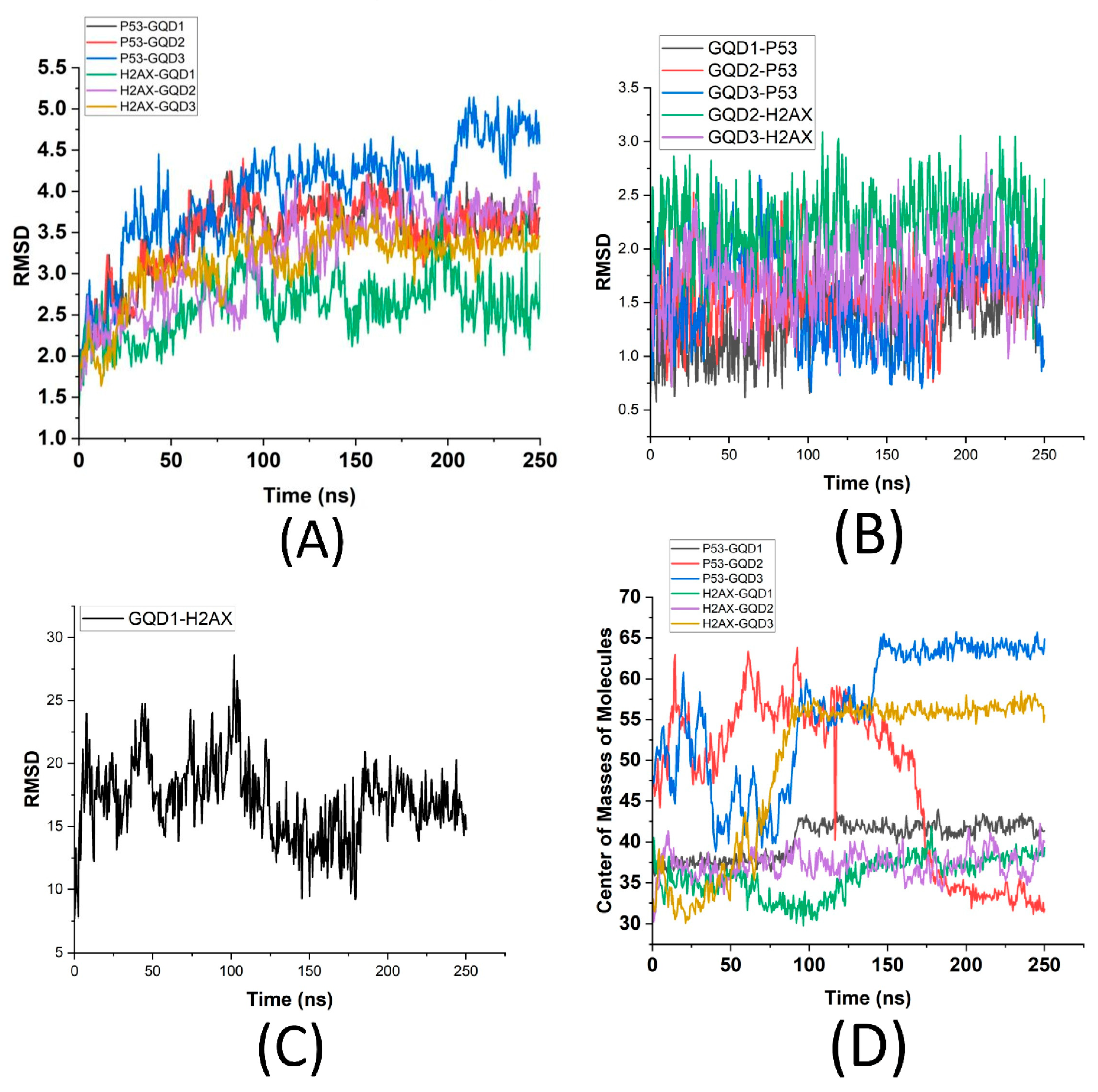
Figure 2D). From
Figure 2A, it is found that γH2AX shows the least deviation in the presence of GQD1 (~2.75 Å) but also has less stability (more fluctuations). In comparison, it has more deviation in the presence of GQD2 and GQD3 (~3.5 Å), but it is also more stable (less fluctuations) after ~125 ns.
This implies that γH2AX has a more stable interface with GQD2 and GQD3 but not with GQD1. In contrast, p53 in the presence of GQD3 shows significant deviation (~4 Å during the first 200 ns and afterwards up to 5 Å) with a highly unstable conformation (large fluctuations), whereas p53 in the presence of GQD1 and GQD2 shows a deviation of ~3.5 Å starting ~75 ns with a higher stability (minor fluctuations). This indicates that both GQD1 and GQD2 facilitate relatively stable interactions at the interface of p53. These results further suggest that while GQD1 and GQD2 are more suitable for adsorption of p53, GQD2 and GQD3 are better suited for γH2AX, owing to a high level of stability of their interfaces. To explore this further, the stability of GQDs in the presence of these proteins was also quantified, as shown in
Figure 2B,C, which clearly demonstrates significant variations in the RMSD of the GQDs depending on the type of protein present. Notably, GQD3 in the presence of p53 shows larger deviation and fluctuations (up to ~2.5 Å) in comparison to GQD1 and GQD2 (up to ~1.75 Å), supporting the RMSD of the p53 in
Figure 2A. Similarly, GQD1 in the presence of γH2AX (
Figure 2C) exhibits the largest deviation (~25 Å), indicating a high degree of structural instability and dynamic movement, further suggesting that γH2AX does not favor smaller GQDs. This can be attributed to γH2AX exhibiting the regions of structural flexibility or partial unfolding that influence its biological role in DNA damage signaling. While the histone core of γH2AX remains relatively stable within the nucleosome, the C-terminal tail—where phosphorylation at Ser139 occurs—is intrinsically disordered and highly flexible. This structural plasticity would allow the phosphorylated tail to extend outward from the nucleosome core, making it accessible to DNA repair factors. Partial unfolding or local fluctuations in the histone-DNA interface may further facilitate exposure of the γH2AX modification site and enhance recognition by repair proteins. In comparison, both GQD2 and GQD3 in the presence of γH2AX show lesser deviation, with GQD3 having an RMSD of ~1.5 Å, which is smaller than GQD2 (~2.5 Å), further supporting the RMSD quantification of γH2AX in
Figure 2A that larger GQDs are better able to maintain their integrity during interactions with γH2AX. In other words, γH2AX, which is smaller in size (~15 kDa) compared to p53 (~53 kDa), did not stabilize in the presence of the smallest GQD1 (2 × 3 nm), whereas p53 did not stabilize in the presence of the largest GQD3 (8 × 9 nm).
To further analyze the attractive interactions between these proteins and GQDs, the Center of Mass (CoM) analysis was conducted. CoM serves as a crucial indicator of the average position of all atoms within a molecule and plays a significant role in understanding molecular interactions. An increasing CoM value suggests the presence of repulsive forces and diminished interactions between the molecules, leading to an increased separation. Conversely, a decreasing CoM value indicates stronger attraction and potential adsorption, drawing the molecules closer together, forming a molecular complex. By examining the CoM trajectory of the p53 and γH2AX proteins when complexed with the GQDs of varying sizes (
Figure 2D), it is found that the p53-GQD3 complex exhibits the highest CoM (~65 Å), indicating significant repulsion and weakened binding, resulting in greater distances between the CoM of the two molecules. On similar lines, even though p53-GQD1 starts closer to each other (~33 Å), after ~100 ns they move further apart (~43 Å), whereas p53-GQD2 were found to move the closest (below 33 Å) after ~175 ns, implying a strong attraction. For γH2AX-bound GQDs, the γH2AX-GQD1 complex reveals a slight repulsive behavior where the two molecules moved away from each other (from 35 Å to 37 Å) after 100 ns. In comparison to this weak repulsive behavior of GQD1, γH2AX-GQD3 showed a very strong repulsive behavior with the CoM of the two molecules moving away by 55 Å, while γH2AX-GQD2 showed stable attraction at ~37 Å. Remarkably, the CoM values for γH2AX bound GQDs are consistently lower than those for their p53 counterparts, suggesting that γH2AX has a greater propensity to associate with smaller GQDs compared to p53. The findings also indicate that smaller GQD1 particles have a higher affinity for both types of proteins, while their larger counterparts (GQD3) display increased repulsive interactions. This phenomenon illustrates an inverse relationship between the strength of protein-GQD interactions and the size of the GQDs, with smaller variants forming more stable complexes. In summary, this data suggests that there is a size-dependent interaction at the interface of GQDs and these proteins, where stable interactions are observed in case of moderately sized p53 in the presence of moderately sized GQD2 and for a smaller sized γH2AX in the presence of a smaller GQD.
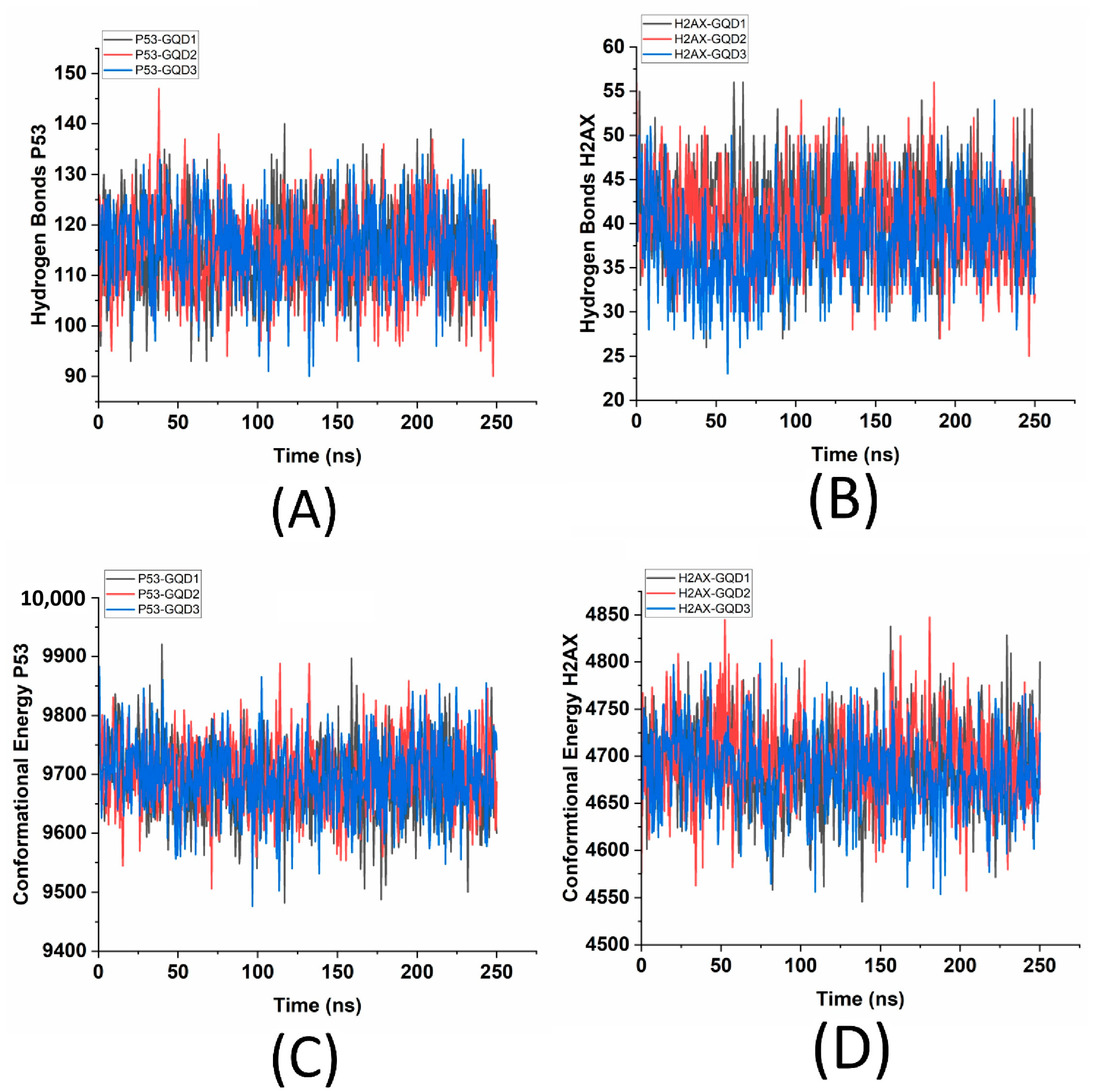
Another measure of stability is the hydrogen bonds, which play a crucial role in controlling the stability and specificity of molecular interactions, particularly in the complex structures formed between proteins and nanoparticles such as GQDs. These non-covalent bonds, which involve the sharing of hydrogen atoms between donors and acceptors, are essential for maintaining the integrity and functionality of biomolecules. In the case of interactions between proteins and GQDs, hydrogen bonding is vital for determining binding affinities, stability, and the overall strength of these interactions. A greater and consistent count of hydrogen bonds indicates a strong and stable interaction. Conversely, fluctuations in this count over time may signify transient bonding events or reorganizations within the complex structures. It is found that the sizes of the GQDs do not significantly affect the number of hydrogen bonds formed, with p53 having ~115 hydrogen bonds (
Figure 3A) and γH2AX having ~40 hydrogen bonds (
Figure 3B) for all combinations of GQDs.
For both γH2AX and p53, the number of hydrogen bonds remains consistent across the different sizes of GQDs, suggesting that the interactions between these proteins and GQDs are not governed by the hydrogen bonds. The RMSD and the interfacial stability as dictated by CoM are also further confirmed through the computed conformational energies for the p53-GQD and γH2AX-GQD complexes. Conformational energy refers to the energy associated with a specific 3D arrangement of atoms in a molecule and its stability. It encompasses a variety of factors, including bond stretching, which measures the energy involved in extending and compressing atomic bonds; bending angles, which indicate the variations in energy when the angles between bonds deviate from their ideal values; and torsional (dihedral) angles, which reflect the energy changes that occur during rotational movement around a bond. It is found that for p53, the mean conformational energy is ~9700 kcal/mol (
Figure 3C), whereas for γH2AX, it is ~4700 kcal/mol (
Figure 3D), which is much lower than the p53-GQD complex. The lower conformational energy values associated with γH2AX-GQD complexes are indicative of the smaller size of γH2AX compared to p53. A potential explanation for γH2AX’s relatively low conformational energy also stems from the inherent structural differences between the two proteins. γH2AX may adopt a relatively stable conformation when complexed with GQD, resulting in lower energy. In contrast, p53’s flexible and dynamic nature likely contributes to its higher conformational energy, attributed to increased fluctuations and strain. Similarly, the conformational energies of the three GQDs over 250 ns in the presence of p53 and γH2AX are also analyzed (
Supplementary Figure S3) and it is found that the three variants of GQD exhibit similar trends, with energies fluctuating within comparable ranges (~7975 kcal/mol for GQD3, ~3366 kcal/mol for GQD2 and ~680 kcal/mol for GQD1). This lends strong support to the assertion that all three variants exhibit comparable conformational dynamics. Despite minor fluctuations in energy levels, the conformational energy values for each variant show significant overlap, suggesting that the presence of these protein molecules does not affect the conformation of GQDs. The correlation between energy profiles and structural configurations reinforces the idea that all three variants exist within a similar energetic environment.
Another measure to quantify the stability of the proteins is their internal salt bridges.
Table 1 reports an investigation on the salt bridges within the proteins, p53 and γH2AX, in the presence of the three GQDs, providing important insights into the behavior of these proteins. Salt bridges, formed by electrostatic attractions between positively and negatively charged residues, play a crucial role in maintaining the structural integrity of proteins. From the six simulated trajectories, we assessed the dynamics of salt bridges to find that in the presence of GQD1, out of the 49 salt bridges, p53 had 10 salt bridges that were formed, 31 that were sustained, and 8 that were broken. In contrast, the interactions between GQD2 and p53 involved 54 salt bridges, with 17 initially formed, 30 sustained, and 7 broken. This higher proportion of formed salt bridges suggests a relatively more stable and robust interaction compared to GQD1. For GQD3 and p53, there were 44 salt bridges detected, with 20 initially formed, 14 sustained, and 10 broken. Although there were fewer sustained salt bridges compared to GQD2, the number of broken bridges was lower than that of GQD1, indicating an acceptable level of structural integrity for this interaction. For γH2AX and GQD1, only 22 salt bridges were identified, comprising 10 that were formed, 8 sustained, and 4 broken. This interaction appears weaker than that of p53, as it has fewer total salt bridges and a more balanced distribution of sustained and broken bridges. In the γH2AX-GQD2 interaction, there was a slight improvement, with 22 salt bridges identified, 6 formed, 9 sustained, and 7 broken. Despite a comparable number of sustained bridges to GQD1, the higher count of broken bridges suggests that γH2AX may be more sensitive to variations in GQD2’s structure. Finally, the γH2AX-GQD3 interaction demonstrated the least stability, with 21 salt bridges recorded, including 7 that were formed, 7 sustained, and 7 broken. The equal count of sustained and broken salt bridges indicates a relatively unstable interaction. Overall, the comparisons revealed that p53 tends to form a greater number of salt bridges with GQDs than γH2AX, and the stability of these salt bridges is higher in the case of p53.
The salt bridge analysis in the
Supplementary Table S1 of the p53 protein in complex with different sizes of GQDs reveals valuable insights into the structural behavior of the protein in response to nanoparticle interaction. Several salt bridges, such as ASP279-ARG299, ASP287-ARG290, and GLU246-LYS308, are consistently observed across all three GQD variants, indicating a set of stable electrostatic interactions that are preserved regardless of the size of the GQD. In addition, a subset of salt bridges, including GLU132-ARG167 and GLU115-LYS102, is shared between two of the three complexes, suggesting partial conservation of structural features that may shift depending on the GQD size. Notably, some interactions are unique to individual GQD systems, such as ASP563-LYS574 in GQD2 and GLU390-ARG104 in GQD2, reflecting specific conformational adaptations induced by a specific nanoparticle dimension. The presence of sustained, newly formed, and disrupted salt bridges throughout the simulation highlights the dynamic nature of the p53-GQD interface. These results collectively suggest that while core electrostatic interactions remain intact, the size of the GQD plays a significant role in modulating the local structural environment of p53, potentially influencing its stability, flexibility, and functional conformation. Furthermore, a relatively small number of sustained salt bridges in p53 in the presence of GQD3 indicate an outsized role of GQD3 to compromise p53’s structural integrity and functional stability. p53 is known for its conformational flexibility and partial disorder, particularly in regions such as the DNA-binding domain, which is highly sensitive to destabilizing mutations. A few persistent salt bridges in this domain act as crucial stabilizing interactions that help preserve the correct folding of β-sheets and loops required for DNA recognition. They also contribute to maintaining the balance between structural rigidity and the conformational plasticity needed for p53’s interactions with diverse DNA response elements and regulatory partners. Thus, GQD3 has the potential to interfere with these crucial electrostatic interactions serving as “structural keystones” that reinforce the fold and safeguard the protein’s tumor suppressor function.
Similarly, the salt bridge analysis in
Supplementary Table S2 of the γH2AX protein interacting with three different sizes of GQDs also revealed notable trends in the structural adaptation and stability of the protein complex. Several salt bridges, such as GLU1014-LYS1011 and GLU931-LYS885, are sustained across all three GQD variants, indicating the presence of conserved electrostatic interactions that remain stable regardless of GQD size. Additionally, certain interactions, like ASP1045-LYS1016 and ASP1045-LYS1067, are formed in the presence of GQD1 but are broken in GQD2 and GQD3, highlighting a size-dependent disruption of specific salt bridges. Notably, the GLU951-LYS907 and GLU951-ARG948 salt bridges are consistently formed in all GQD variants, suggesting they play a critical role in stabilizing the γH2AX-GQD interface. On the other hand, several bridges such as GLU938-LYS900 and GLU1038-LYS1016 are only observed in isolated cases, reflecting the dynamic and flexible nature of the γH2AX structure in response to different nanoparticle sizes. Overall, the data suggests that although certain key electrostatic interactions are preserved, the structural response of γH2AX varies with the size of GQD, indicating that nanoparticle dimensions influence not only interaction patterns but also the conformational stability of the complex.
An additional measure to quantify the stability is to examine the number of protein atoms within 5 Å (adsorption criteria) of the GQDs, which is crucial for understanding the interactions at the interface. This also directly impacts binding capacity, stability of the molecular complex, and reactivity of the GQD surface. For the number of atoms of p53/γH2AX within 5 Å of GQDs, the stability is analyzed through Van der Waals (VDW) energy to determine the GQD size that is the most favorable to these proteins.
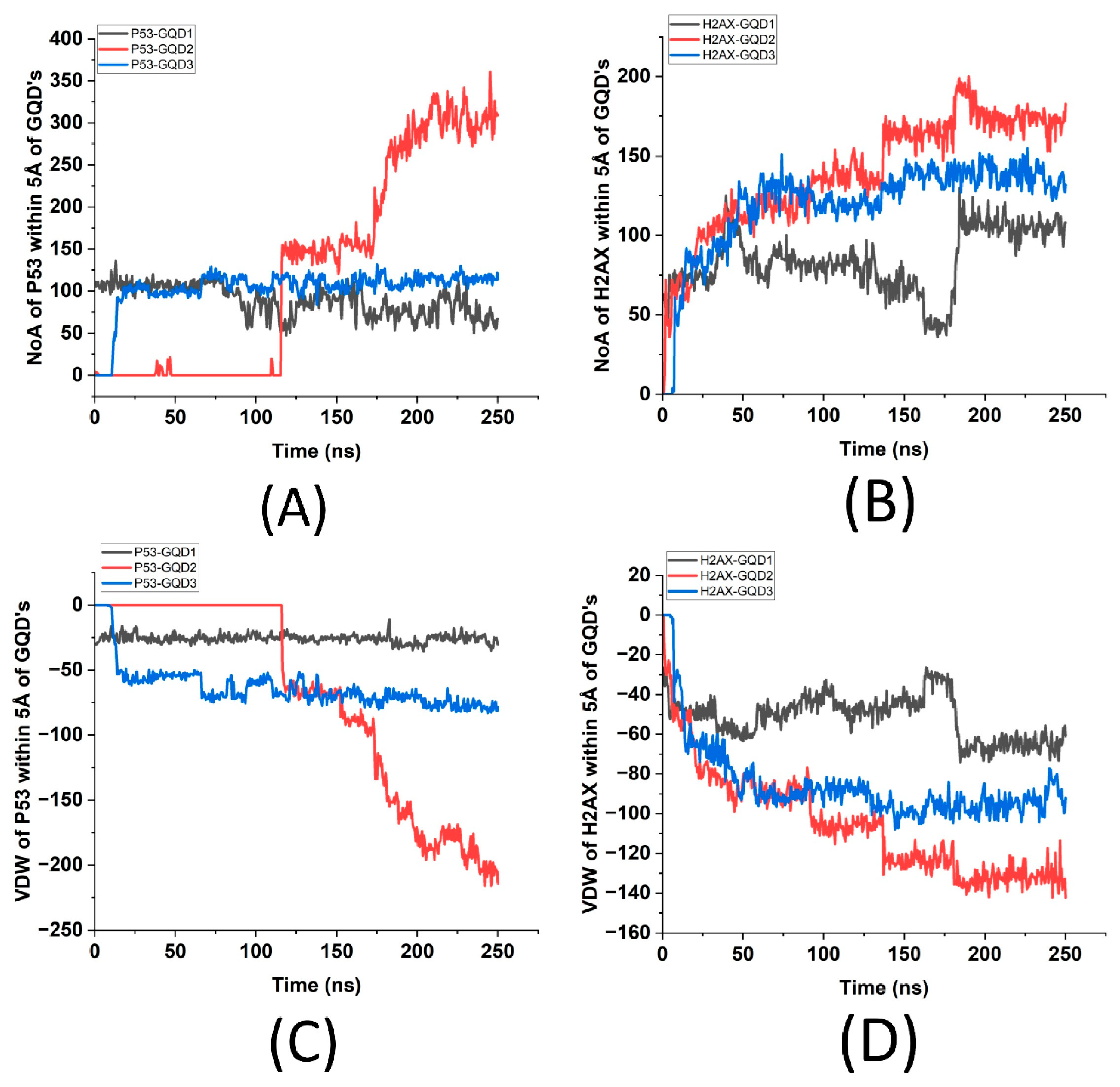
Figure 4 displays the number of atoms and their interaction energies (VDW) of p53 and γH2Ax within 5 Å of each GQD. It shows that the greatest number of atoms of p53 is ~300 in the presence of GQD2, followed by ~100 for GQD3 and ~75 for GQD1 (
Figure 4A). Similarly, γH2AX has the greatest number of atoms, ~175, for GQD2, followed by ~125 in the presence of GQD3 and ~100 in the presence of GQD1 (
Figure 4B). This graph further supports that γH2Ax stabilizes in four stages while interacting with GQD2, finally stabilizing at ~170 ns, and at ~130 ns while interacting with GQD3, while p53 stabilizes only after ~200 ns for GQD2, which is much later compared to GQD1 and GQD3. The VDW energies as seen from
Figure 4C imply that even though p53 stabilizes much later, it has a significant VDW energy (~200 kcal/mol) compared to GQD1 (~25 kcal/mol) and GQD3 (~60 kcal/mol), indicating that GQD2 is clearly the more suitable nanoparticle for p53. Similarly, from
Figure 4D, γH2AX interacts the most with GQD2 (~140 kcal/mol) compared to GQD1 (~70 kcal/mol) and GQD3 (~90 kcal/mol), indicating that GQD2 also may be a better option for adsorption of γH2AX. To resolve these differences between p53 and γH2AX in terms of their stability, it became important to analyze their secondary structures to understand localized conformational changes in the presence of the three GQDs.
Supplementary Figures S4 and S5 provide details on the secondary structure analysis of the two proteins from the simulated trajectories. It is found that γH2AX maintained its secondary structure while interacting with GQD2 compared to GQD1 and GQD3. While interacting with GQD1, the structure of reactive (phosphorylation) site on γH2AX, Serine 139 (resid 1065), changed from 3-10 helix to a turn after ~175 ns; the same occurred after 80 ns in the presence of GQD3. This indicates that γH2AX is not only the most stable but also retains its secondary structure in the presence of GQD2. This indicates that γH2AX is more stable and is more likely to maintain its functionality while interacting with GQD2. A similar analysis was performed on the secondary structure of p53 in the presence of three GQDs (
Figure S5), where it is found that the reactive site on p53 from SER162 to LYS333 undergoes a massive permanent conformational change in the presence of GQD3 and GQD1, where it loses most of its α-helices, but not in the presence of GQD2. Thus, both γH2AX and p53 retain their secondary structures in the presence of GQD2, thereby indicating that GQD2 is well suited for both proteins if they are to retain their function. On the other hand, if the goal is to denature the proteins, then GQD3 is the best choice out of the three GQDs, followed by GQD1.
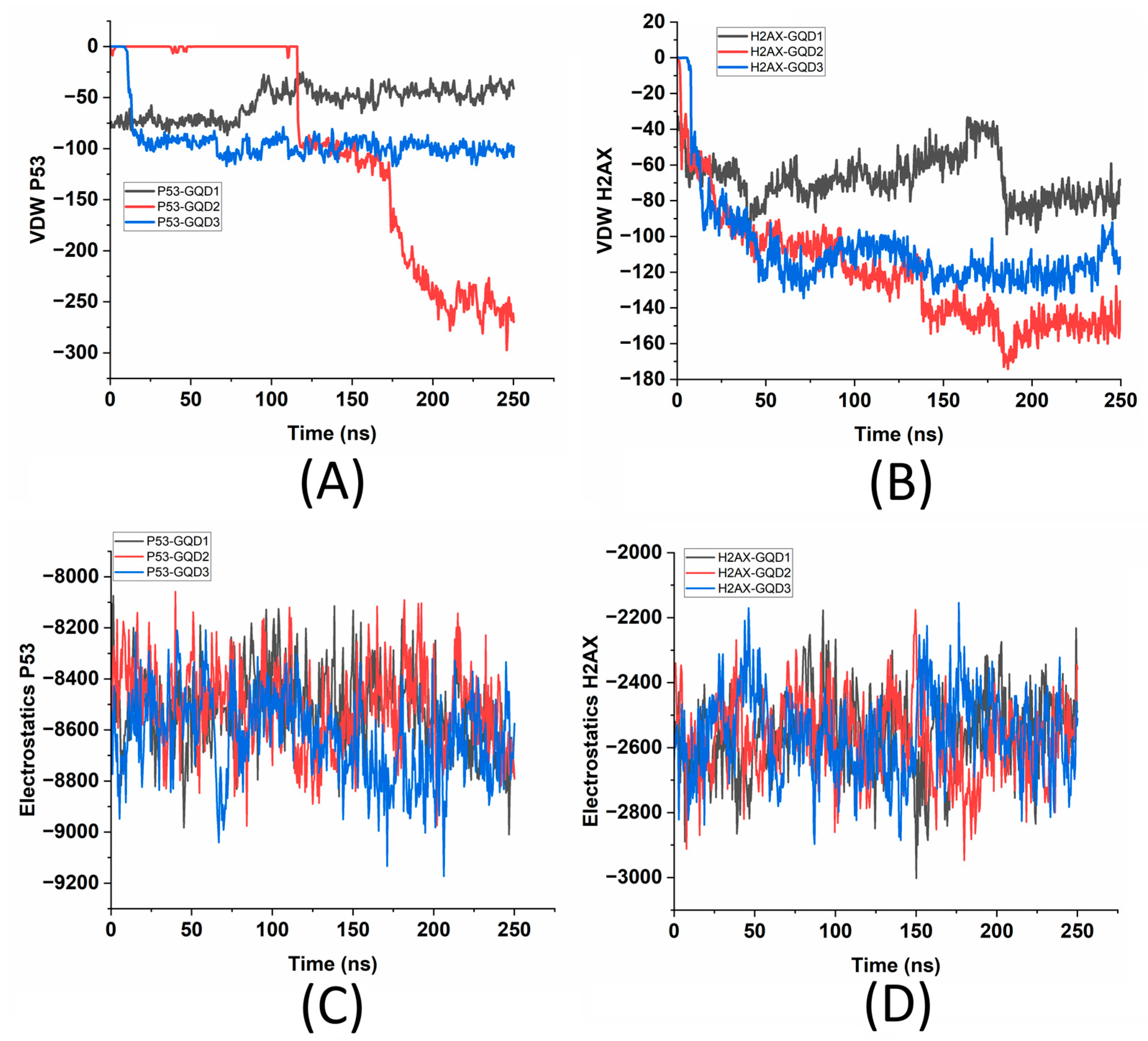
To compare the VDW energies of the two proteins within 5 Å of the GQDs to the overall energy landscape of the proteins, a thorough analysis of the overall VDW and electrostatic energies in the presence of the three GQDs was also performed.
Figure 5A and 5B illustrate the van der Waals (VDW) interaction energies of the six different systems involving the two proteins, p53 and γH2AX, and the three different GQDs (GQD1, GQD2, and GQD3). On similar lines,
Figure 5C,D shows the electrostatic energy analysis of the two proteins in the presence of the three GQDs. More negative values indicate stronger attractive interactions, where it is found that the p53-GQD2 combination exhibits the strongest interaction, with a notable VDW energy up to 275 kcal/mol. This suggests that GQD2 offers the most favorable binding surface for the p53 protein. In comparison, both GQD1 and GQD3 show moderate interactions with ~50 kcal/mol and ~100 kcal/mol (
Figure 5A). For γH2AX, interactions with GQD2 and GQD3 are relatively strong, with γH2AX-GQD2 displaying the highest attractive energy, indicating a stronger affinity (
Figure 5B). Conversely, γH2AX-GQD1 shows the weakest interaction among all combinations, highlighting a less favorable binding with the smallest GQD. Overall, the data indicate that GQD2 consistently exhibits the strongest binding interactions for both p53 and γH2AX, while the larger GQD (GQD3) generally facilitates stronger interactions than the smallest GQD (GQD1). Additionally, p53 appears to interact more intensively with the GQDs compared to γH2AX, particularly in the case of GQD2, which can be attributed to its larger size compared to γH2AX. These findings underscore the impact of GQD size on the strength of protein binding. To compare these interactive energies with the internal energy of the two proteins,
Figure 5C,D demonstrate that owing to their sizes, the electrostatic energies differ in both proteins, and the presence of different-sized GQDs does affect the internal energies of the proteins. It is known that electrostatic forces are fundamental in understanding the dynamics of biomolecular interactions, especially regarding nanomaterials and protein-ligand binding.
These electrostatic interactions occur due to the attraction and repulsion between molecular dipoles and charged atoms, which play a crucial role in determining binding affinity, molecular stability, and the overall functionality of biological systems. In analyzing the GQDs in relation to the p53 and γH2AX proteins, a comprehensive examination of electrostatic energy offers valuable insights into protein adsorption behavior, binding strength, and the impact of GQD size on binding efficacy. As seen from
Figure 5C,D, the electrostatic interaction energies of the p53 and γH2AX proteins reveal two clusters of energy values: the p53-GQD systems display significantly attractive electrostatic energies compared to the γH2AX-GQD systems. For the p53 complexes, all three GQD sizes exhibit similar trends, with electrostatic energy values converging around −9000 kcal/mol, reflecting strong and stable interactions. Slightly stronger interactions are noted for p53-GQD2 compared to GQD1 and GQD3, further supporting the conclusion that GQD2 provides a more favorable binding environment. In contrast, the electrostatic interactions between GQDs and γH2AX are noticeably weaker, stabilizing at around −3000 kcal/mol. The variations in the interaction strength among γH2AX complexes are also relatively small, suggesting a uniform interaction strength across the different GQD sizes. To further assess the interaction strength at the interface of these six systems, two additional analyses on the optimal distances of p53/γH2AX proteins from the GQDs and the role of interfacial water molecules and their hydrogen bonds were performed.
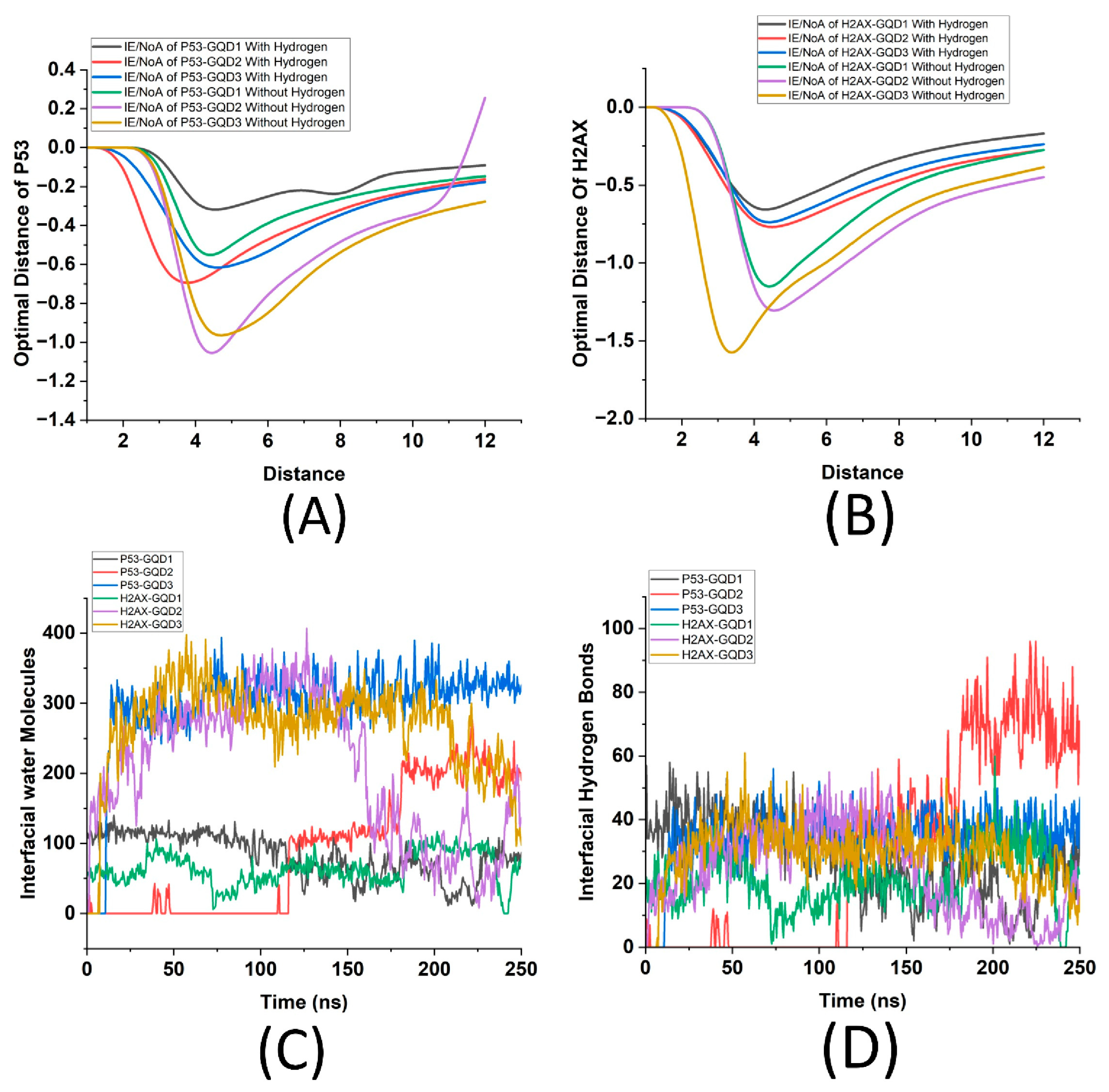
The optimal distance in molecular interactions is defined as the precise separation at which the interaction energy per atom (IE/NoA) reaches a maximum, reflecting the most stable configuration among interacting entities. This parameter is critically important for applications involving molecular docking and the interactions between nanomaterials and proteins, as it facilitates the identification of favorable binding distances where various forces—including VDW interactions, electrostatics, and hydrogen bonding—are effectively balanced. Analyzing this optimal distance is essential for elucidating the interactions between proteins and nanomaterials, which subsequently influence their adsorption characteristics, stability, and potential applications.
Figure 6A illustrates the relationship between interaction energy per atom (IE/NoA) and non-bonding cut-off distance for the interaction of the p53 protein with the GQDs, both in the presence and absence of hydrogen. Each curve corresponds to different sizes of GQDs, distinguished according to whether hydrogen is present in the system. Hydrogen atoms are considered because their presence can significantly alter a protein’s affinity for GQDs through hydrogen bonding and electrostatic interactions.
Analyzing protein interactions both with and without hydrogen offers insights into the structural and functional behavior of proteins, enhancing our understanding of how hydrogen affects protein conformation and binding capacity. Essentially, this analysis emphasizes the significance of GQD size and the presence of hydrogen in modulating protein-GQD interactions, providing valuable insights for designing more efficient protein-functionalized nanodevices. Similarly, by focusing on heavy atoms (excluding hydrogen), we can better understand the structural interactions between the proteins and GQDs, isolating the hydrogen-dependent interactions and eliminating the influence of hydrogen bonding and electrostatic interactions, which can significantly affect the stability and reactivity of proteins. Analysis of
Figure 6A indicates that interaction energy per atom has a peak for every combination of six systems and that p53 shows maximum energy per atom in the presence of GQD2 for both with and without hydrogen cases, reinforcing earlier findings that GQD2 provides an optimal interaction interface for p53. These peaks correspond to the ideal interaction distance where the binding affinity of p53 to GQD is maximized. The systems that include hydrogen atoms exhibit distinct behaviors compared to their hydrogen-free counterparts. Moreover, the differences between the interactions with and without hydrogen highlight the significant role hydrogen plays in modulating interaction energies. The presence of hydrogen appears to enhance interactions at their optimal distances, thereby increasing the stability of the system, which indicates that hydrogen bonding plays a crucial role in the binding process of p53 to GQDs. Similarly,
Figure 6B illustrates the interaction energy per number of atoms (IE/NoA) as a function of distance for the γH2AX protein interacting with different sizes of GQDs, both with and without hydrogen. Each curve corresponds to a specific GQD size (GQD1, GQD2, GQD3) and its interaction with γH2AX, distinguishing between systems with hydrogen from those without. The overall trend in the graph shows that for each system, the interaction energy initially decreases, reaching a minimum before stabilizing at larger distances. This minimum represents the optimal interaction distance, where the binding between γH2AX and the GQDs is strongest. As in case of p53, the presence of hydrogen significantly influences the interactions, demonstrating greater stabilization, suggesting that hydrogen enhances the interaction energy, likely through additional bonding interactions that strengthen the protein-GQD complex. Additionally, larger GQDs (GQD2 and GQD3) display more pronounced stabilization and a smoother trend compared to the smallest GQD1. This can be attributed to the increased surface area and more available interaction sites in the larger GQDs, facilitating stronger and more uniform interactions with γH2AX. Notably, GQD3 shows the deepest energy well, supporting indicating it provides the most favorable interaction surface for γH2AX.
To further explore the nature of interface between the proteins and the GQDs, the role of interfacial water molecules between the proteins and the GQDs, and their hydrogen bonds were quantified. These key parameters are crucial in studying molecular interactions, as they play a critical role in stabilizing interactions, mediating hydrogen bonds, and influencing the structural and dynamic properties of the system. Furthermore, this also offers insights into interaction strength, binding stability, and overall molecular behavior.
Figure 6C illustrates the number of interfacial water molecules over time for the six molecular systems, which involve the interactions of p53 and γH2AX proteins with the three GQDs. From this analysis, it is evident that larger GQDs (e.g., GQD3) exhibit a significantly higher number of interfacial water molecules compared to smaller GQDs (e.g., GQD1). This observation indicates a larger interaction surface area and more extensive hydration. Additionally, γH2AX showed higher counts of interfacial water molecules than p53 for the same GQD size, suggesting stronger hydration or a higher affinity for water at the γH2AX-GQD interface. Moreover, the number of interfacial water molecules fluctuates over time for each system, reflecting the dynamic nature of water molecule exchange at the interface. It is known that the hydrogen bonds formed by the interfacial water molecules also play a crucial role in stabilizing the interface, so studying the number of interfacial hydrogen bonds provides further insights into the strength and specificity of these interactions, which are essential for understanding binding mechanisms and designing functional biomolecular systems.
Figure 6D illustrates the number of interfacial hydrogen bonds over time for the six systems involving the interactions of p53 and γH2AX proteins with three GQDs, where it can be observed that the number of interfacial hydrogen bonds varies with both the type of protein and the size of the GQD. For p53 interactions, GQD2 exhibits the highest number of hydrogen bonds (~70), particularly during the later stages of the simulation, indicating stronger and more stable interactions for this size. In contrast, GQD3 shows fewer hydrogen bonds (~40), while GQD1 exhibits the lowest (~20). In case of γH2AX, the data demonstrate relatively consistent hydrogen bond numbers across all GQD sizes, with slight variations for GQD1 and GQD3 (~20 to ~30 bonds), indicating that both GQD1 and GQD3 facilitate better interactions compared to GQD2.
Finally, as a last measure of studying the interface between p53 and γH2AX proteins with the GQDs, the number and type of adsorbed residues (amino acids) of the proteins on the surface of the GQDs were analyzed. As shown in
Table 2, proteins are classified into different types based on their chemical properties. These residues can be categorized as polar (e.g., SER, TYR, ASN, GLN), non-polar (e.g., PRO, VAL, PHE, LEU, TRP), positively charged (e.g., ARG, LYS, HSD), or negatively charged (e.g., GLU). These properties significantly influence how residues interact with other molecules, such as GQDs. Studying the number of adsorbed residues during the interaction between proteins like p53 and γH2AX with GQDs provides insights into the binding mechanisms, stability, and specificity of these interactions. Adsorbed residues interact directly with the GQD surface, forming non-covalent bonds, including hydrogen bonds, π-π stacking, and hydrophobic interactions. Understanding these interactions further helps to comprehend the bio-nano interface. As seen from
Table 2, for p53, the number of adsorbed residues varies significantly across the three GQD sizes. Smaller GQD1 adsorbs residues such as SER (resid 189 and 190) and TYR (resid 225), while larger GQD2 and GQD3 adsorb additional residues, including GLN (resid 118, 396, 659) and ARG (resid 71, 634). This highlights the increased number of interaction sites available on larger GQDs due to their greater surface area. Residues like PRO, VAL, and HSD are more prominently observed in larger GQDs, indicating that larger GQDs create more opportunities for interactions with non-polar and positively charged residues. For γH2AX, the number of adsorbed residues consistently increases with the size of the GQD. Polar residues such as GLN (resid 1017) and charged residues like ARG (resid 1010, 1037, 1042) are consistently adsorbed across all GQD sizes. Notably, larger GQDs like GQD3 also adsorb additional residues, such as GLU (resid 1014, 1038) and LYS (resid 1016, 1067), indicating stronger or more extensive interactions with charged and polar residues.