SCCM: An Interpretable Enhanced Transfer Learning Model for Improved Skin Cancer Classification
Abstract
1. Introduction
2. Literature Review
3. Proposed System: Design and Implementation
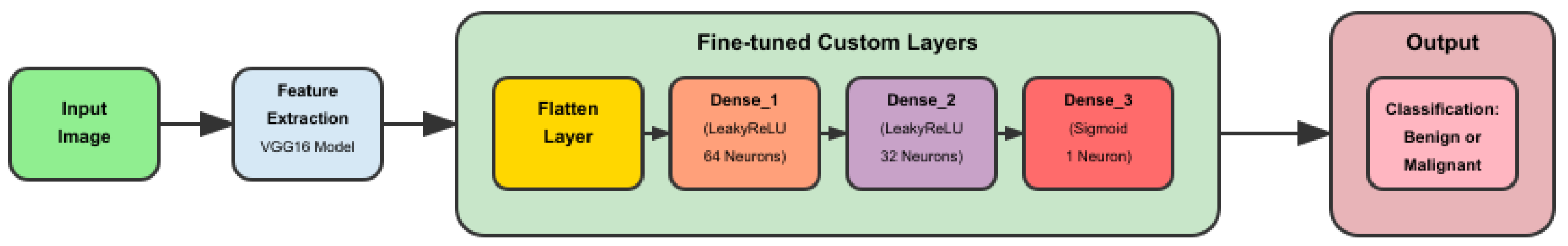
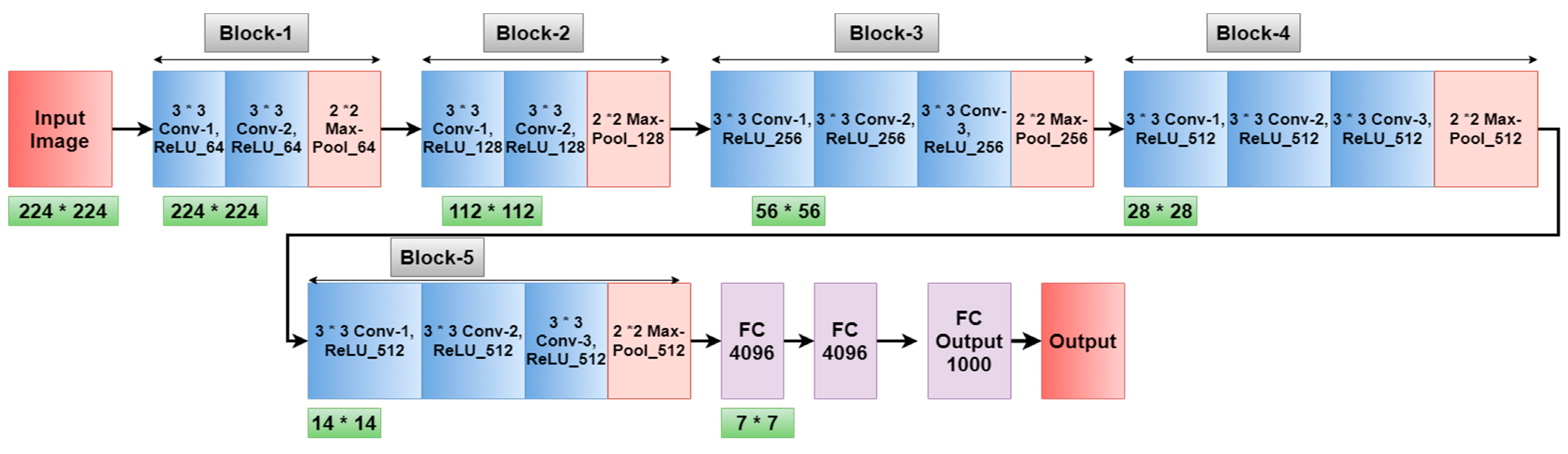
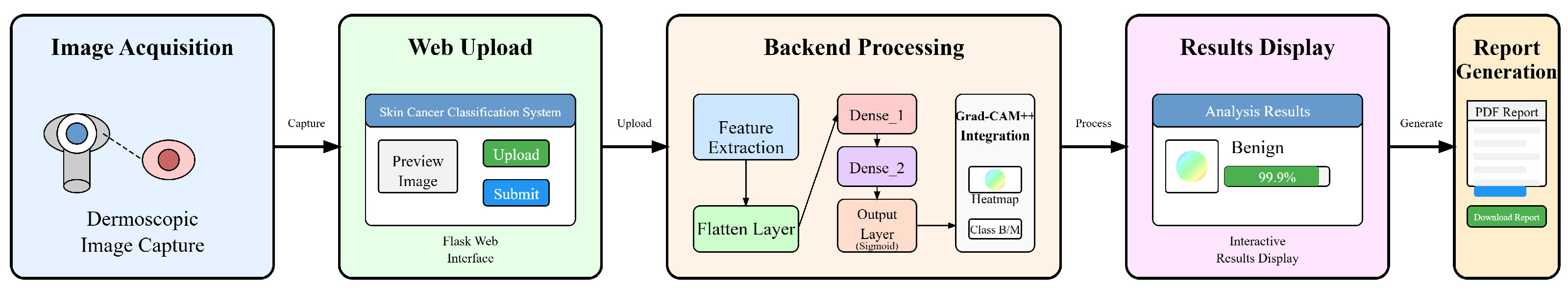
3.1. Model and System Architecture
3.2. Requirement Analysis
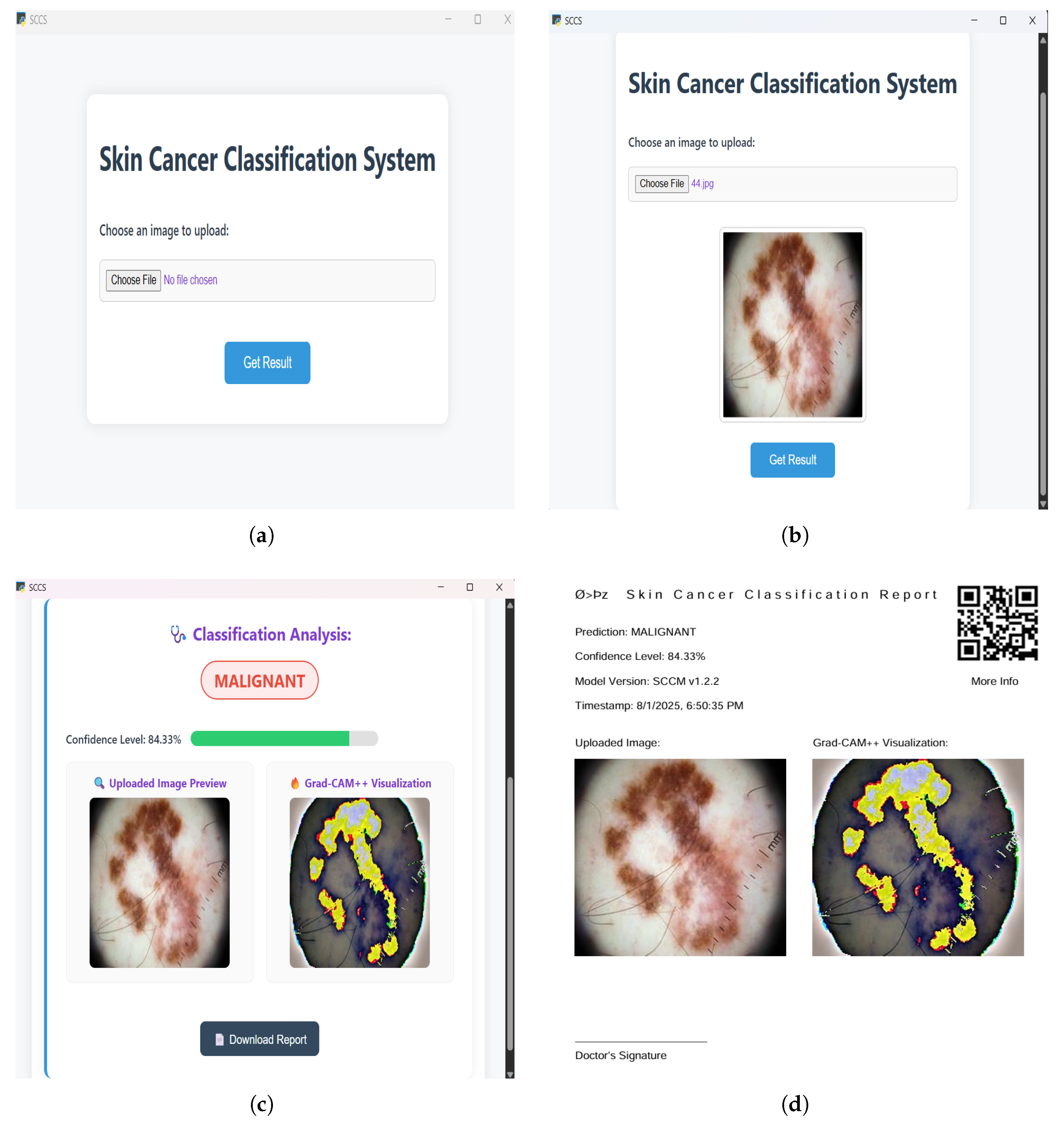
3.3. Implementation
4. Result Analysis and Discussion
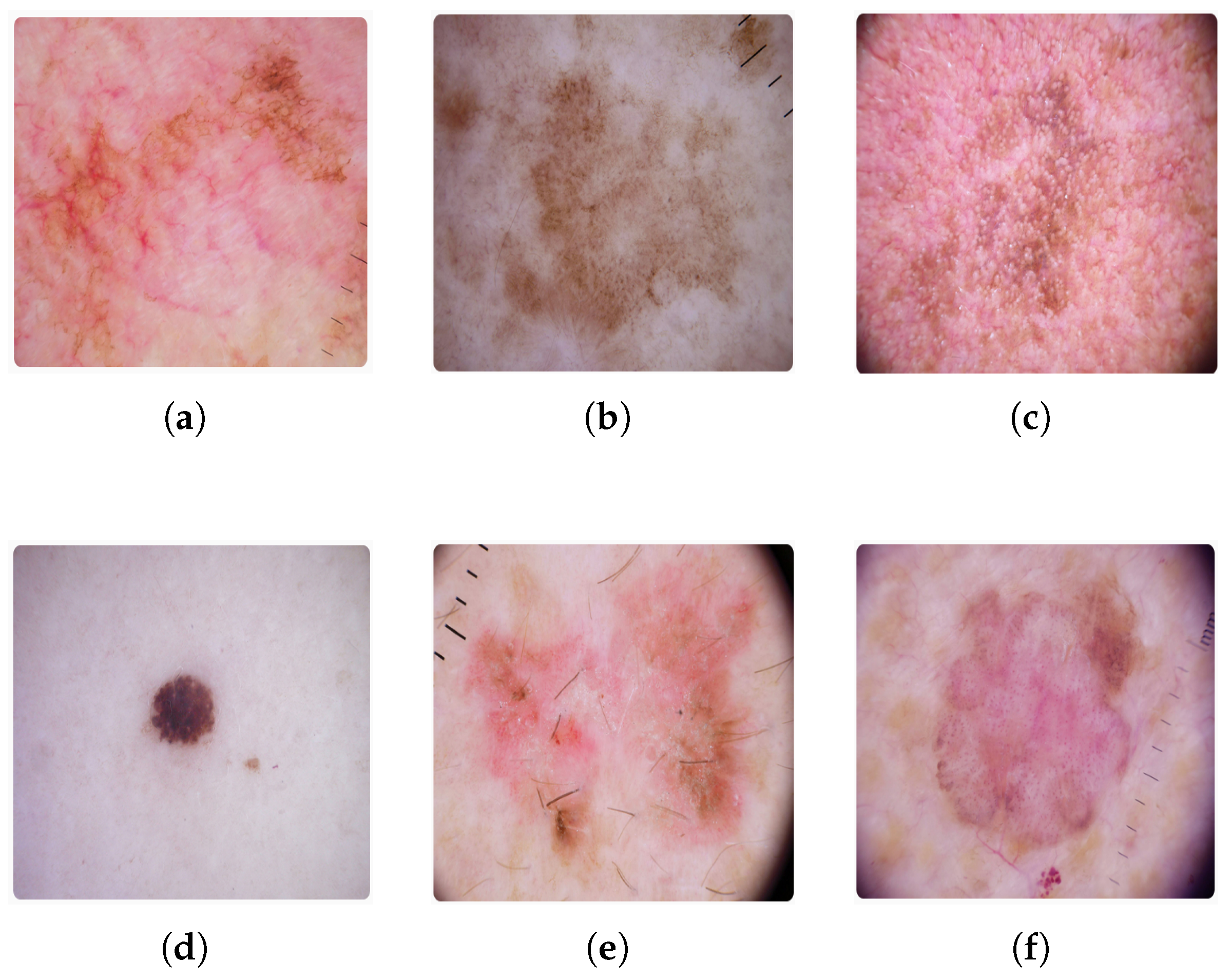
4.1. Dataset Specifications
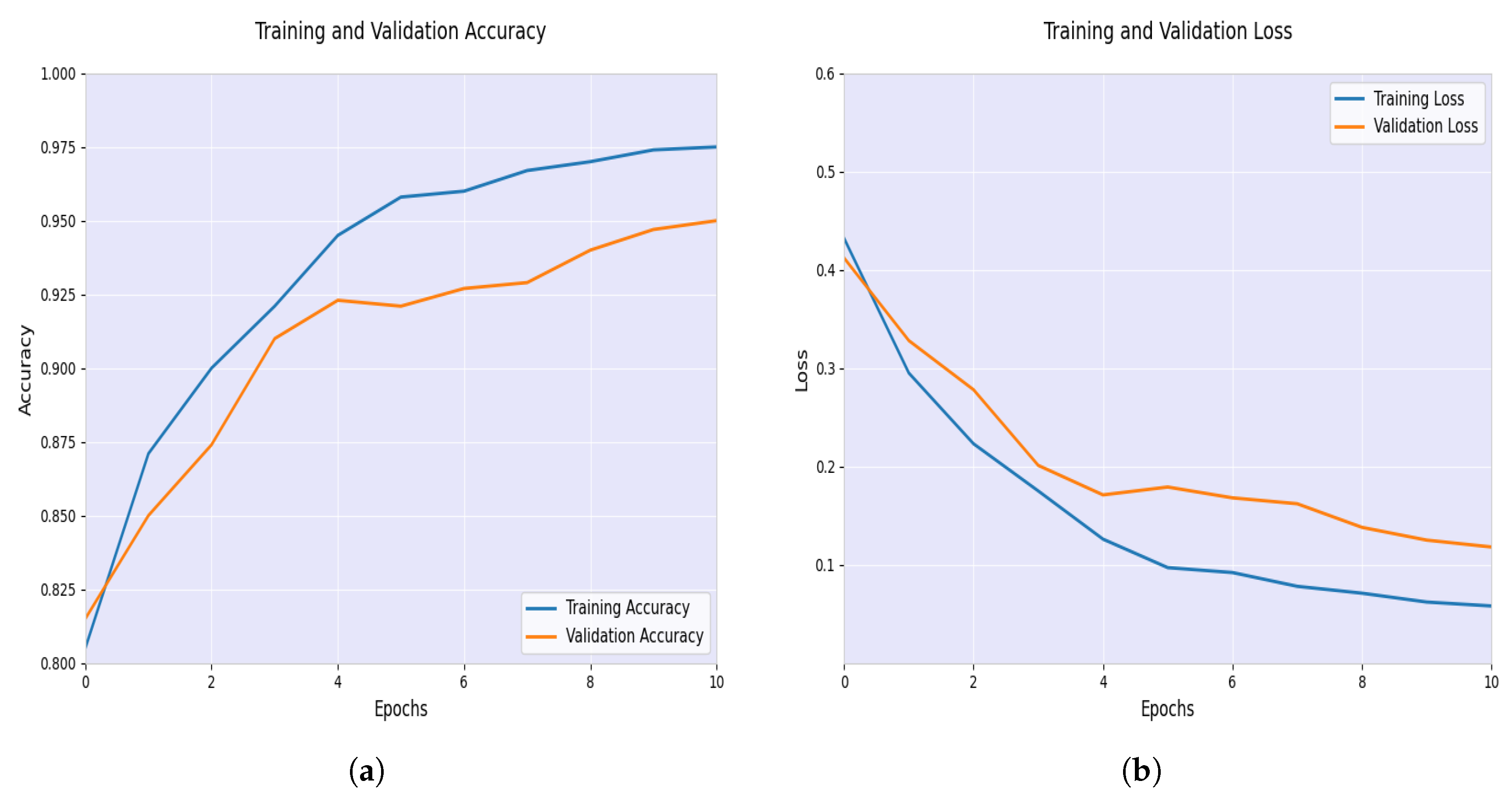
4.2. Training, Validation, and Test
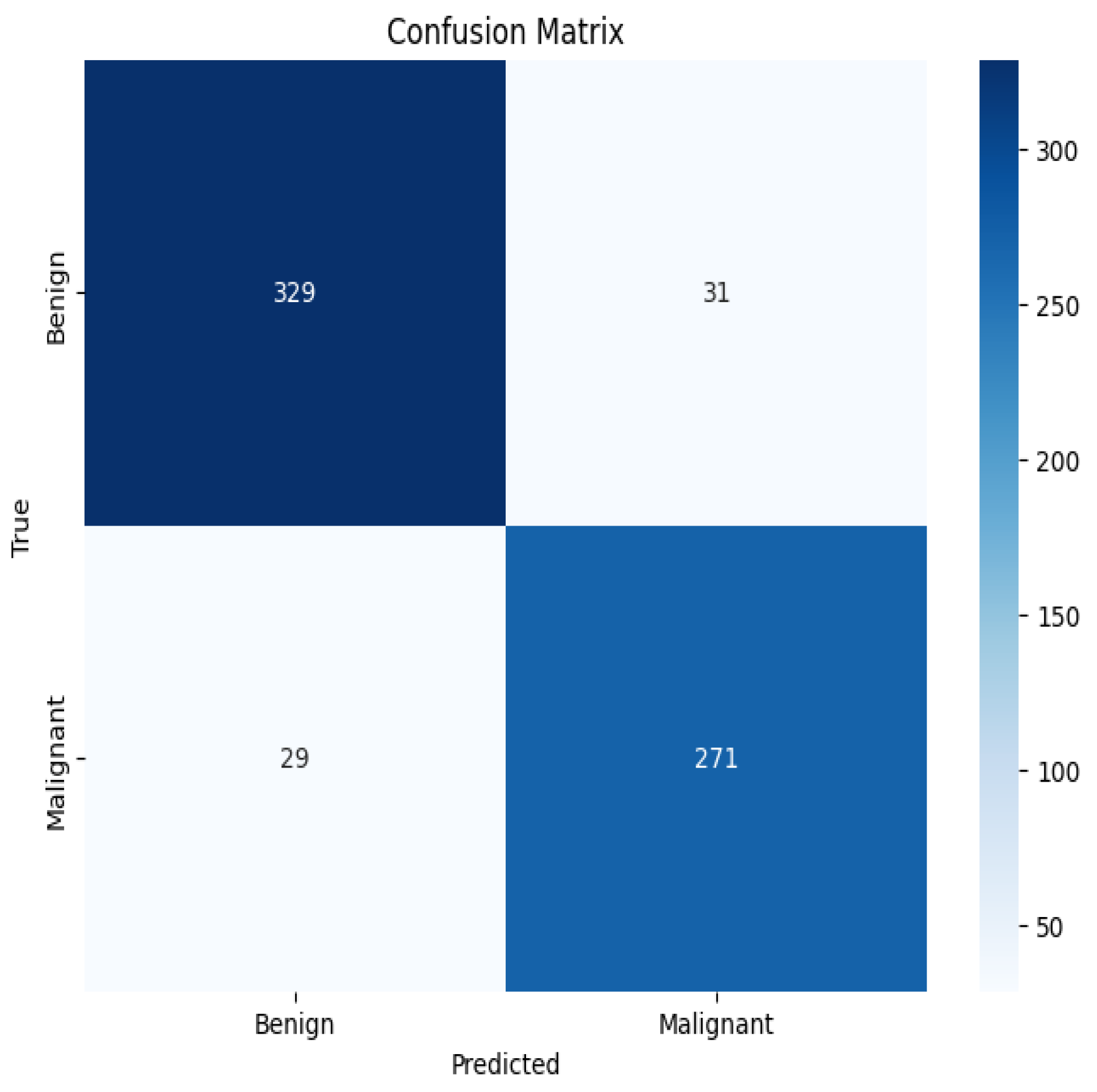
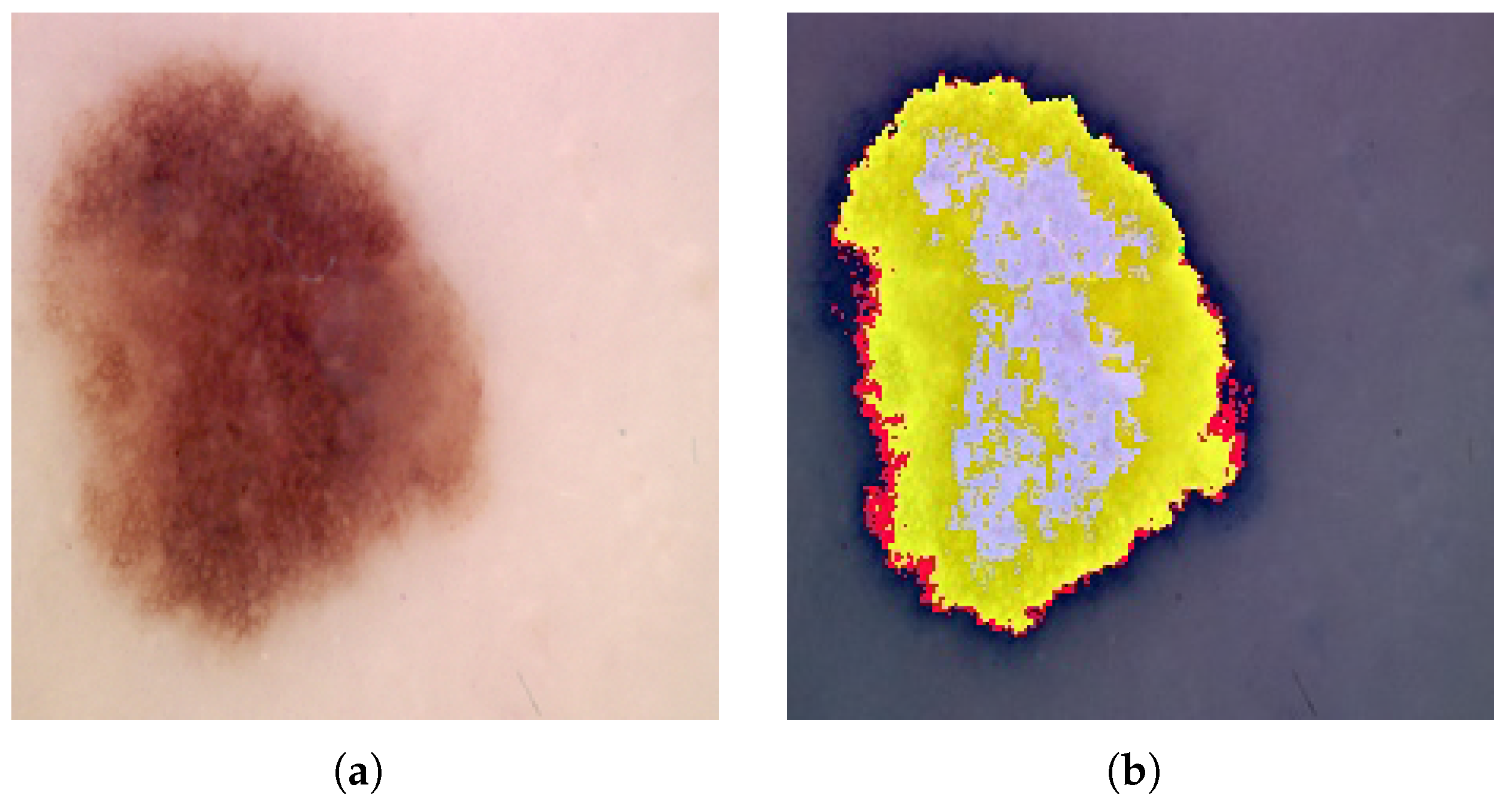
4.3. Performance Analysis
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
References
- Anand, V.; Gupta, S.; Altameem, A.; Nayak, S.R.; Poonia, R.C.; Saudagar, A.K.J. An enhanced transfer learning based classification for diagnosis of skin cancer. Diagnostics 2022, 12, 1628. [Google Scholar] [CrossRef] [PubMed]
- Cancer Research UK. Melanoma Skin Cancer Incidence Statistics. Available online: https://www.cancerresearchuk.org/health-professional/cancer-statistics/statistics-by-cancer-type/melanoma-skin-cancer/incidence (accessed on 18 June 2025).
- World Health Organization. Ultraviolet Radiation. Available online: https://www.who.int/news-room/fact-sheets/detail/ultraviolet-radiation (accessed on 18 June 2025).
- American Cancer Society. Survival Rates for Melanoma Skin Cancer by Stage. Available online: https://www.cancer.org/cancer/types/melanoma-skin-cancer/detection-diagnosis-staging/survival-rates-for-melanoma-skin-cancer-by-stage.html (accessed on 18 June 2025).
- Ghorbani, M.; Raahemifar, K.; Mahjani, F.; Moradi, F. Early detection of skin cancer using AI: Deciphering dermatology images for melanoma detection. AIP Adv. 2024, 14, 040701. [Google Scholar] [CrossRef]
- Reddy, S.; Shaheed, A.; Patel, R. Artificial intelligence in dermoscopy: Enhancing diagnosis to distinguish benign and malignant skin lesions. Cureus 2024, 16, e22547. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Chen, B.; Zeng, A.; Pan, D.; Wang, R.; Zhao, S. Skin Cancer Classification With Deep Learning: A Systematic Review. Front. Oncol. 2022, 12, 893972. [Google Scholar] [CrossRef] [PubMed]
- Fanconi, C. Skin Cancer: Malignant vs. Benign. Available online: https://www.kaggle.com/datasets/fanconic/skin-cancer-malignant-vs-benign (accessed on 18 June 2025).
- Ibrahim, A.M.; Elbasheir, M.; Badawi, S.; Mohammed, A.; Alalmin, A.F.M. Skin cancer classification using transfer learning by VGG16 architecture (case study on Kaggle dataset). J. Intell. Learn. Syst. Appl. 2023, 15, 67–75. [Google Scholar] [CrossRef]
- Agarwal, K.; Singh, T. Classification of skin cancer images using convolutional neural networks. arXiv 2022, arXiv:2202.00678. [Google Scholar] [CrossRef]
- Bazgir, E.; Haque, E.; Maniruzzaman, M.; Hoque, R. Skin cancer classification using Inception Network. World J. Adv. Res. Rev. 2024, 21, 839–849. [Google Scholar] [CrossRef]
- Hussein, H.; Magdy, A.; Abdel-Kader, R.F.; Ali, K.A.E. Binary Classification of Skin Cancer using Pretrained Deep Neural Networks. Suez Canal Eng. Energy Environ. Sci. 2023, 1, 10–14. [Google Scholar] [CrossRef]
- Alrabai, A.; Echtioui, A.; Kallel, F. Exploring Pre-Trained Models for Skin Cancer Classification. Appl. Syst. Innov. 2025, 8, 35. [Google Scholar] [CrossRef]
- Yildiz, A. A comparative analysis of skin cancer detection applications using histogram-based local descriptors. Diagnostics 2023, 13, 3142. [Google Scholar] [CrossRef] [PubMed]
- Djaroudib, K.; Lorenz, P.; Bouzida, R.B.; Merzougui, H. Skin cancer diagnosis using VGG16 and transfer learning: Analyzing the effects of data quality over quantity on model efficiency. Appl. Sci. 2024, 14, 7447. [Google Scholar] [CrossRef]
- Othman, S.; Mourad, H. Skin Cancer Detection Using Convolutional Neural Network (CNN). J. Acs Adv. Comput. Sci. 2022, 13, 42–48. [Google Scholar] [CrossRef]
- Nayan, A.A.; Mozumder, A.N.; Haque, M.R.; Sifat, F.H.; Mahmud, K.R.; Azad, A.K.A.; Kibria, M.G. A Deep Learning Approach for Brain Tumor Detection from MRI Images. Int. J. Electr. Comput. Eng. 2022, 13, 1039–1047. [Google Scholar] [CrossRef]
- Akther, J.; Harun-Or-Roshid, M.; Nayan, A.A.; Kibria, M.G. Transfer Learning on VGG16 for the Classification of Potato Leaves Infected by Blight Diseases. In Proceedings of the 2021 Emerging Technology in Computing, Communication and Electronics (ETCCE), Online, 21–23 December 2021. [Google Scholar] [CrossRef]
- Ibrahim, S. Skin Cancer ISIC 2019 & 2020 Malignant or Benign. Available online: https://www.kaggle.com/datasets/sallyibrahim/skin-cancer-isic-2019-2020-malignant-or-benign (accessed on 10 July 2025).
- Sathyanarayanan, S.; Tantri, B.R. Confusion matrix-based performance evaluation metrics. Afr. J. Biomed. Res. 2024, 27, 4023–4031. [Google Scholar] [CrossRef]
- Garbe, C.; Amaral, T.; Peris, K.; Hauschild, A.; Arenberger, P.; Basset-Seguin, N.; Bastholt, L.; Bataille, V.; Del Marmol, V.; Dréno, B.; et al. European consensus-based interdisciplinary guideline for melanoma. Part 1: Diagnostics: Update 2022. Eur. J. Cancer 2022, 170, 236–255. [Google Scholar] [CrossRef]
| Metric | VGG16 | Proposed Model | Difference |
|---|---|---|---|
| Accuracy | 85.37% | 90.91% | +5.54% |
| Misclassification Rate | 14.63% | 9.09% | −5.54% |
| Sensitivity (Recall) | 87.42% | 90.33% | +2.91% |
| Specificity | 83.61% | 91.39% | +7.78% |
| False Negative Rate | 12.58% | 9.67% | −2.91% |
| False Positive Rate | 16.39% | 8.61% | −7.78% |
| Precision | 82.12% | 89.74% | +7.62% |
| F1-Score | 84.67% | 90.03% | +5.36% |
| Metric | VGG19 | Proposed Model | Difference |
|---|---|---|---|
| Accuracy | 87.61% | 90.91% | +3.30% |
| Misclassification Rate | 12.39% | 9.09% | −3.30% |
| Sensitivity (Recall) | 80.00% | 90.33% | +10.33% |
| Specificity | 94.17% | 91.39% | −2.78% |
| False Negative Rate | 20.00% | 9.67% | −10.33% |
| False Positive Rate | 5.83% | 8.61% | +2.78% |
| Precision | 92.19% | 89.74% | −2.45% |
| F1-Score | 85.67% | 90.03% | +4.36% |
| Metric | ResNet-18 | Proposed Model | Difference |
|---|---|---|---|
| Accuracy | 88.51% | 90.91% | +2.40% |
| Misclassification Rate | 11.49% | 9.09% | −2.40% |
| Sensitivity (Recall) | 88.71% | 90.33% | +1.62% |
| Specificity | 88.33% | 91.39% | +3.06% |
| False Negative Rate | 11.29% | 9.67% | −1.62% |
| False Positive Rate | 11.67% | 8.61% | −3.06% |
| Precision | 86.75% | 89.74% | +2.99% |
| F1-Score | 87.73% | 90.03% | +2.30% |
| Metric | InceptionNet-v4 | Proposed Model | Difference |
|---|---|---|---|
| Accuracy | 87.31% | 90.91% | +3.60% |
| Misclassification Rate | 12.69% | 9.09% | −3.60% |
| Sensitivity (Recall) | 86.45% | 90.33% | +3.88% |
| Specificity | 88.06% | 91.39% | +3.33% |
| False Negative Rate | 13.55% | 9.67% | −3.88% |
| False Positive Rate | 11.94% | 8.61% | −3.33% |
| Precision | 86.17% | 89.74% | +3.57% |
| F1-Score | 86.31% | 90.03% | +3.72% |
| Metric | AlexNet | Proposed Model | Difference |
|---|---|---|---|
| Accuracy | 89.10% | 90.91% | +1.81% |
| Misclassification Rate | 10.90% | 9.09% | −1.81% |
| Sensitivity (Recall) | 91.29% | 90.33% | −0.96% |
| Specificity | 87.22% | 91.39% | +4.17% |
| False Negative Rate | 8.71% | 9.67% | +0.96% |
| False Positive Rate | 12.78% | 8.61% | −4.17% |
| Precision | 86.02% | 89.74% | +3.72% |
| F1-Score | 88.54% | 90.03% | +1.49% |
| Model Architecture | Dataset | Accuracy |
|---|---|---|
| Modified VGG16 with data augmentation | Skin cancer: Malignant vs. benign | 89.09% |
| Fine-tuned VGG16 with texture and shape features | Skin cancer: Malignant vs. benign | 84.24% |
| CNN with transfer learning | Skin cancer: Malignant vs. benign | 86.65% |
| Fine-tuned InceptionNet | Skin cancer: Malignant vs. benign | 85.94% |
| Pre-trained deep neural networks (AlexNet, ResNet-18, SqueezeNet, ShuffleNet) | Skin cancer: Malignant vs. benign | 89.00% (ResNet-18) |
| VGG19 with explainable AI | Skin cancer: Malignant vs. benign | 86.21% |
| Histogram-based local descriptors with XGBoost classifier | Skin cancer: Malignant vs. benign | 90.00% |
| VGG16 (last convolutional block) + custom dense classifier + explainable AI (This research) | Skin cancer: Malignant vs. benign | 90.91% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aknda, M.R.; Farid, F.A.; Uddin, J.; Mansor, S.; Kibria, M.G. SCCM: An Interpretable Enhanced Transfer Learning Model for Improved Skin Cancer Classification. BioMedInformatics 2025, 5, 43. https://doi.org/10.3390/biomedinformatics5030043
Aknda MR, Farid FA, Uddin J, Mansor S, Kibria MG. SCCM: An Interpretable Enhanced Transfer Learning Model for Improved Skin Cancer Classification. BioMedInformatics. 2025; 5(3):43. https://doi.org/10.3390/biomedinformatics5030043
Chicago/Turabian StyleAknda, Md. Rifat, Fahmid Al Farid, Jia Uddin, Sarina Mansor, and Muhammad Golam Kibria. 2025. "SCCM: An Interpretable Enhanced Transfer Learning Model for Improved Skin Cancer Classification" BioMedInformatics 5, no. 3: 43. https://doi.org/10.3390/biomedinformatics5030043
APA StyleAknda, M. R., Farid, F. A., Uddin, J., Mansor, S., & Kibria, M. G. (2025). SCCM: An Interpretable Enhanced Transfer Learning Model for Improved Skin Cancer Classification. BioMedInformatics, 5(3), 43. https://doi.org/10.3390/biomedinformatics5030043








