Probing the Immune System Dynamics of the COVID-19 Disease for Vaccine Designing and Drug Repurposing Using Bioinformatics Tools
Abstract
1. Introduction
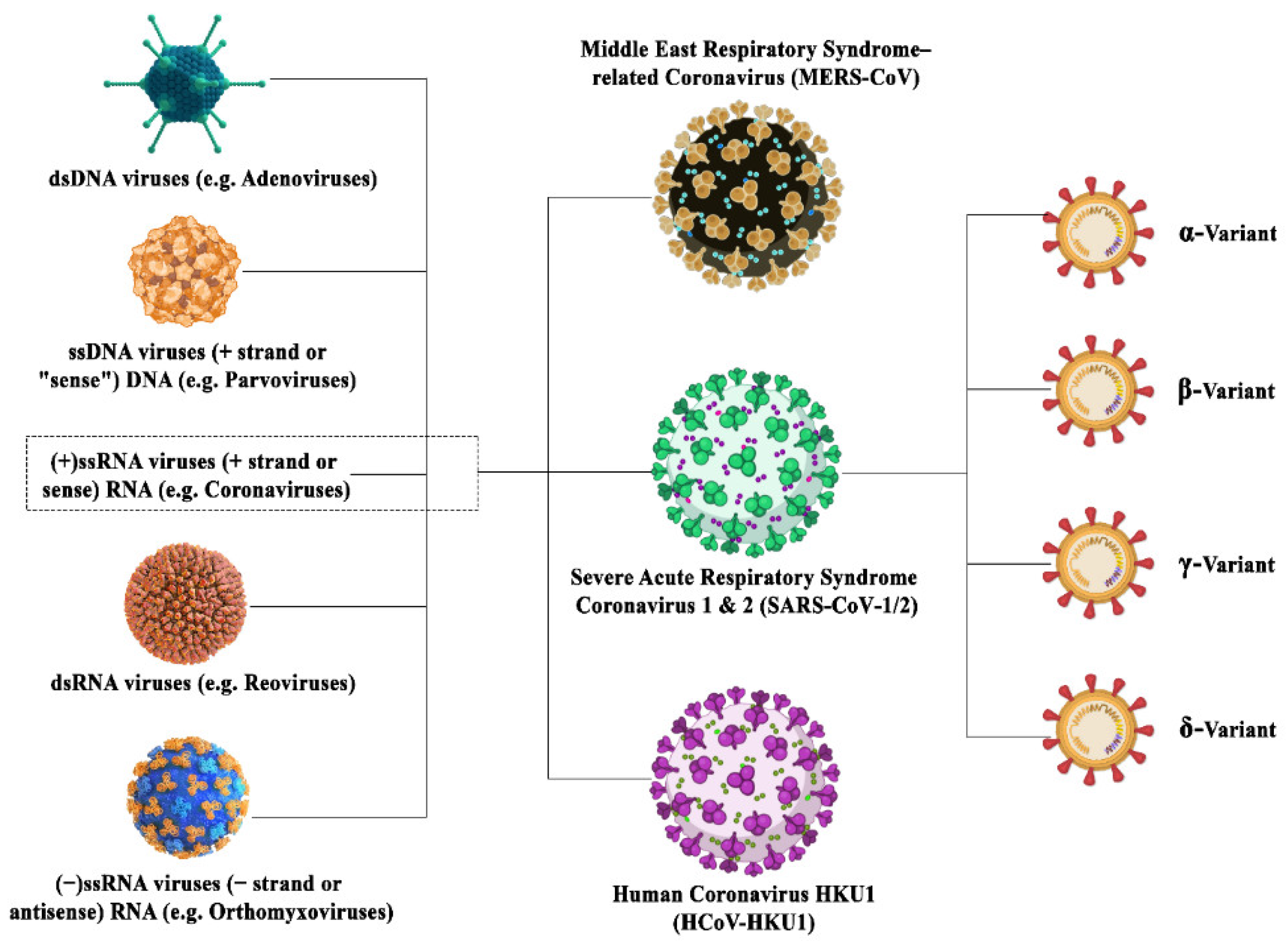
2. Viral Mutations
3. Coronavirus Adaptations
4. Diagnosis of Coronavirus
- Nucleic acid amplification test (NAAT): The most reliable and accurate methodology for the detection of SARS-CoV-2 is the nucleic acid amplification test (NAAT) [39,40]. The most popular laboratory-based NAAT is the quantitative reverse transcription-polymerase chain reaction [41,42]. The samples that are used for diagnosis include sputum, saliva, nasal, pharyngeal and tracheal swabs, broncho-alveolar lavage, pleural effusion fluid, blood, faeces and sometimes urine and semen [6].According to the World Health Organization (WHO), the detection of a single RNA sequence of coronavirus by RT-PCR is enough for the confirmation of the disease. This test is used for the real-time qualitative detection of nucleic acids from suspected viral pathogens [43]. In case of detection of SARC-CoV-2, the nucleic acid (RNA) isolated from the specific sample is first reverse transcribed into cDNA and then amplified. During the amplification process, the probe anneals to a specific target sequence located between the forward and reverse primers. During the extension phase of the PCR cycle, the 5′ nuclease activity of Taq polymerase degrades the bound probe, causing the reporter dye to separate from the quencher dye, generating a fluorescent signal. Fluorescence intensity is monitored at each PCR cycle [43]. According to the WHO, based on the first sequences of SARS-CoV-2 made available on the GISAID database on 11 January 2020, primers and probes (nCoV_IP2 and nCoV_IP4) were designed to target the RdRp gene spanning nt 12,621–12,727 and 14,010–14,116 [44]. The current test kits may lead to false-negative results because the detection of COVID-19 in the early stages of the infection is challenging due to the improper isolation of RNA or inadequate methods for detection.
- Chest computerized tomography (CT): CT scans are used for the diagnosis and imaging of viral pneumonia. It has been extensively used for the timely diagnosis and treatment of other coronavirus outbreaks caused by SARS-CoV and MERS-CoV. It is a confirmatory scan that is used to detect any false negatives as robust detection of COVID-19 is imperative for avoiding these false negatives because a CT scan can detect the infection even before the manifestation of symptoms resulting in timely treatment [39].
- Serological immunoassays: A plethora of serological immunoassays exists which detect SARS-CoV-2 viral proteins and antibodies against those proteins in the plasma or serum. The most popular biomolecules detected by commercial immunological tests such as rapid lateral flow immunoassay (LFIA) tests, automated chemiluminescence immunoassay (CLIA) and manual ELISA and other formats are IgM and IgG antibodies. The antibodies are released into the bloodstream in the second week of viral infection. IgM and IgG may be detected within 10–30 days and 20 days post-infection, respectively [40]. The IgM antibody unveils a drop in concentration, whereas IgG persists in the systemic circulation for prolonged periods and may play a role in adaptive immunity against SARS-CoV-2 in a possible second encounter. ELISA kits against nucleocapsid (NP) and spike (SP) viral proteins exist in the market but they are mainly used for rsearch and development purposes only [45]. The details of all the diagnostic techniques used for SARS-CoV-2 detection are summarized in the Table 2 below:
4.1. Molecular Targets for Diagnostic Kits
D-Dimer-Based Detection
4.2. Effect of Viral Mutations on the Accuracy of Diagnostic Tests
5. Transmission of Coronavirus
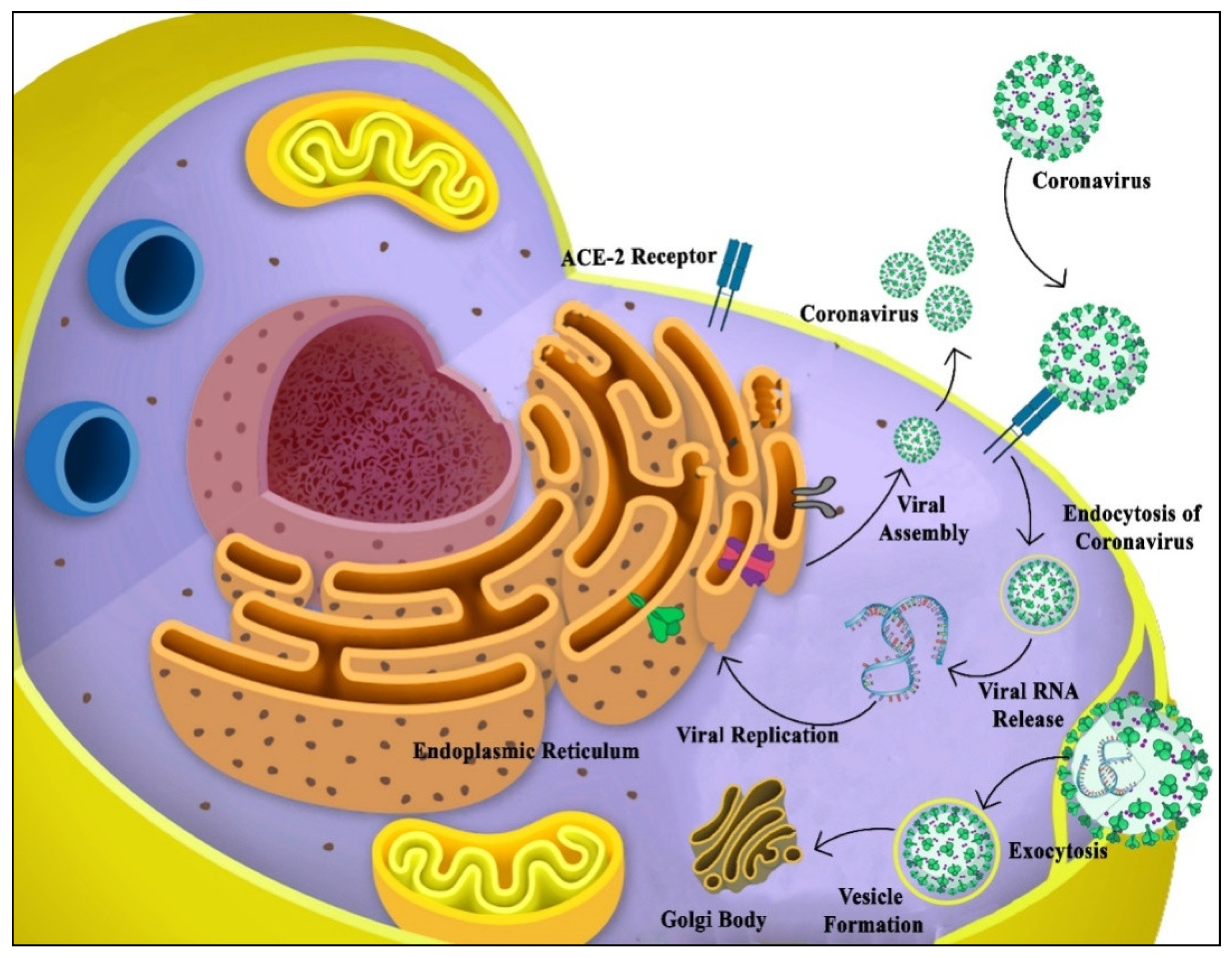
6. Mechanism of Viral Entry into the Host
7. Immunogenic Trigger in Response to Coronavirus
7.1. Hallmarks of the Immune Response to COVID-19
7.2. The Interplay between the Production of Interferons and Inflammatory Cytokines
Mechanism of Action of Interferons
7.3. Role of the Inflammasome in Causing Inflammation
7.3.1. Histopathological Markers of Inflammation
7.3.2. Role of Damage-Associated Molecular Patterns (DAMPs) Inflammation Trigger
7.4. Role of the Adaptive Immune System against COVID-19
7.5. Damage to the T-Lymphocytes Inflicted by COVID-19
7.6. Gender-Based Differences in the Immune Response against COVID-19
7.7. Contribution of Bioinformatics Tools for Designing Theranostics Approaches
7.8. Epitope Designing: The Process
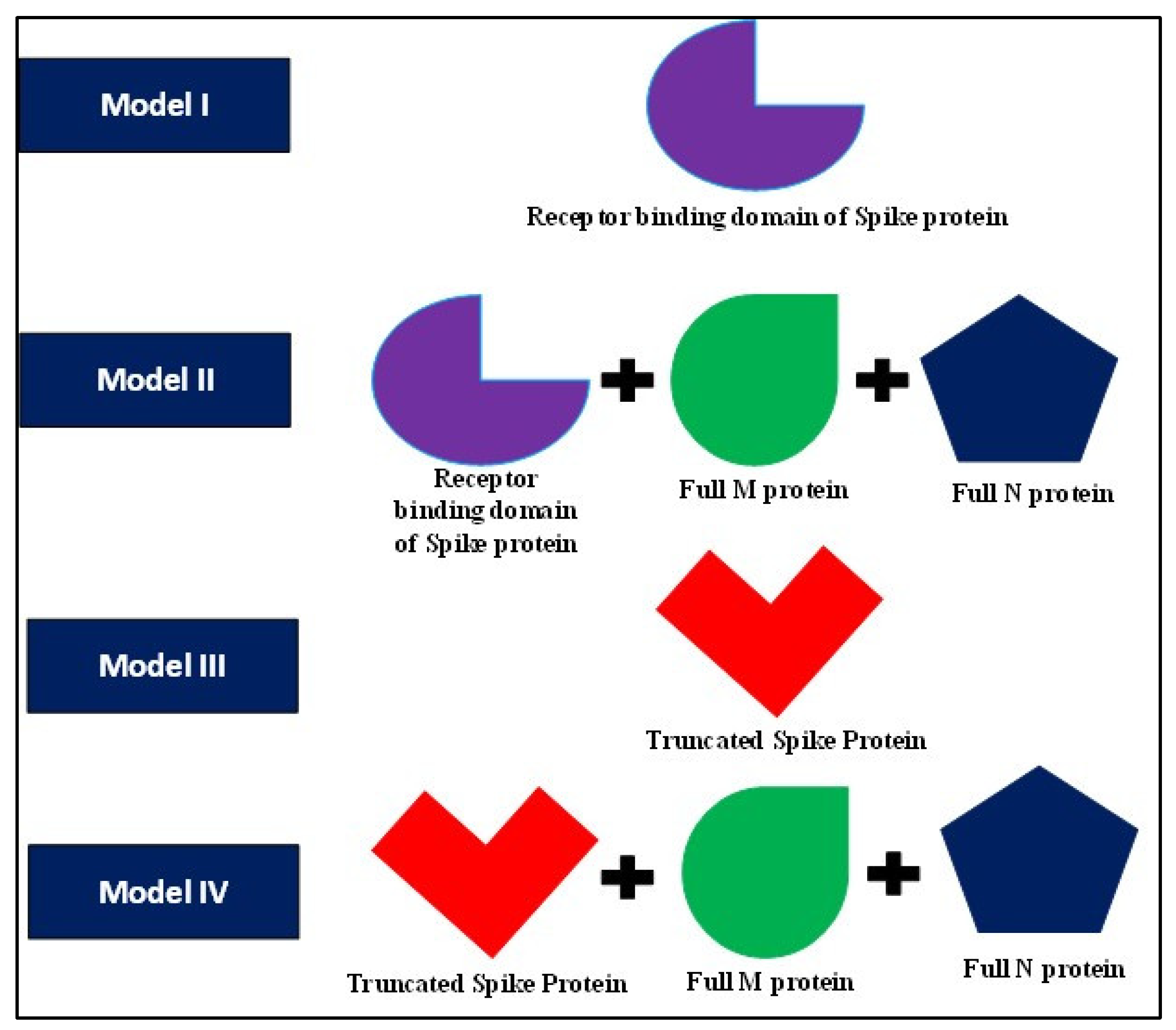
7.9. Approaches for Vaccine Development
- Targeting the cellular immune system: For effective vaccine development against intracellular pathogens, the identification of protective antigens from thousands of candidates is a prerequisite. However, the relevant properties of the antigens which render protection to the endogenous pathogens against the cellular immune system are poorly characterized [138,139], delaying the process of vaccine development. Antigen abundance is an important property that can be exploited for this purpose [138,140]. The antigen expression profile varies from a few to millions of molecules per cell and the antigens which are highly expressed can be selectively tested for their immunogenicity [141,142].In one such study by Rollenhagen et al., the selective recognition of a few highly expressing antigens by CD4+ cells led to the induction of a potent immune response which was shown by the in vivo selection of abundantly expressed antigens. Therefore, the selection of such abundant antigens may facilitate the development of effective vaccines against infectious diseases such as influenza, typhoid fever, COVID-19, etc. This process can be improved by studying the transcriptomic data from viral–host interactions [138].
- Pooling of highly immunogenic epitopes: The human serum consists of a range of polyclonal antibodies that may or may not correspond to immunogenic antigens. This is because a pathogen consists of several antigenic particles/epitopes which have originated from regions of the pathogens that were genetically evolved to be less immunogenic This deceives the immune system into recognizing the non-immunogenic epitopes which undergo constant mutations. Even after having a memory of the antigens, the immune system fails on a second encounter because of the accumulation of mutations in the epitopes. Selecting such antigens will ensure that the immune system can induce the production of antibodies specific to the pathogen Therefore, pooling out the highly conserved epitopes of the virus which are seemingly non-immunogenic, and raising an antibody response against them in absence of the whole virus would enhance their immunogenicity and promise a robust immune response against the virus [90,143].
- Computational Docking: Designing vaccines can also be done by docking studies. The first step in the process of vaccine designing is the generation of crystal structures of the antigens and their respective antibodies. Once this task is accomplished, the next step is the in silico docking of both the structures in a bid to discover useful epitopes which are immunogenic.This approach was attempted by Kuntz et al. in 1982 [144]. In their study, they analyzed the binding geometries of different ligands and their respective receptors for steric overlapping by devising a docking program DOCK. This aided in identifying binding sites on the macromolecular surface. They studied the binding interactions between heme-myoglobin and thyroid hormone and pre-albumin. Since then, this field has had major reformations and is still flourishing at a rapid rate [144,145]. Some of the studies also involved the utilization of Ayurvedic plants Piper longum and Ocimum sanctum for the treatment of coronavirus by targeting ACE2 and TMPRSS2 receptor proteins [146].
- Multi-subunit-based vaccines:In a study by Dar et al., in silico studies were conducted on the spike protein of COVID-19 and multiple epitopes were identified for their ability to elicit a strong immune response. All of these epitopes were then combined via flexible GPGPG linkers along with a cholera toxin β subunit (CTB) sequence at the N-terminal joined by an EAAAK linker, as an adjuvant to form a multi-subunit vaccine candidate. This vaccine was then subjected to molecular docking studies with toll-like receptors to understand their interaction dynamics and to check for the immunogenicity of the multi-subunit vaccine against COVID-19. Quality assessment by the proSA web server showed a good match with experimentally resolved similar PDB structures. Ramachandran plot results revealed that 95% of residues were in favourable regions. Discotope and Ellipro servers revealed the competency of the vaccine in terms of possessing 27% and 52% epitopes recognized by the B-cells. The HADDOCK server showed that the binding between the protein clusters exhibited good stability and flexibility. However, there was an improvement in the HADDOCK scores when molecular refinements were applied to the docked protein complex. Moreover, the interactions between the protein clusters were also energetically favourable. Hence, in vitro studies must be performed to validate the docking results to further prove the effectiveness of the vaccine against COVID-19 [147].
8. Drug Repurposing
Targeting Viral Replication Proteins for Drug Repurposing
- (a)
- Enriching the drug candidates using glide software;
- (b)
- Evaluation of the docking hits using the MM-PBSA-WSAS method;
- (c)
- Selection of the drug candidates based on their binding energies.
9. Phage Display Techniques for Understanding the Molecular Dynamics of Viral Pathogenesis
10. Artificial Intelligence (AI) Based Vaccine Designing and Drug Repurposing against SARS-CoV-2 Virus
- Machine learning (ML): ML allows for the generation of models which analyze and learn from the available data to identify patterns and derive inferences from previously unseen data [203]. Some of the popular ML algorithms include support vector machines, random forest classifiers, k-means, hierarchical clustering, etc., and recently artificial neural networks (ANN) [204].
11. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Zheng, J. SARS-CoV-2: An Emerging Coronavirus that Causes a Global Threat. Int. J. Biol. Sci. 2020, 16, 1678–1685. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.X.; Liang, J.Q.; Fung, T.S. Human Coronavirus-229E, -OC43, -NL63, and -HKU1 (Coronaviridae). Encycl. Virol. 2021, 428–440. [Google Scholar]
- Malik, Y.A. Properties of Coronavirus and SARS-CoV-2. Malays. J. Pathol. 2020, 42, 3–11. [Google Scholar]
- Schoeman, D.; Gordon, B.; Fielding, B.C. Pathogenic Human Coronaviruses. In Reference Module in Biomedical Sciences; Elsevier: Amsterdam, The Netherlands, 2021. [Google Scholar]
- Beck, B.R.; Shin, B.; Choi, Y.; Park, S.; Kang, K. Predicting commercially available antiviral drugs that may act on the novel coronavirus (SARS-CoV-2) through a drug-target interaction deep learning model. Comput. Struct. Biotechnol. J. 2020, 18, 784–790. [Google Scholar] [CrossRef] [PubMed]
- Kowalik, M.M.; Trzonkowski, P.; Łasińska-Kowara, M.; Mital, A.; Smiatacz, T.; Jaguszewski, M. COVID-19—Toward a comprehensive understanding of the disease. Cardiol. J. 2020, 27, 99–114. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharya, S.; Sharma, P.; Mathur, H.; Rasheed, H.; Singh, S.; Rajput, G.; Agnihotri, S.; Nirmal, P.; Kaur, S. Recent apprise on coronavirus and its terrible insinuations. Virusdisease 2020, 31, 121–127. [Google Scholar] [CrossRef][Green Version]
- Bhattacharya, M.; Sharma, A.R.; Patra, P.; Ghosh, P.; Sharma, G.; Patra, B.C.; Saha, R.P.; Lee, S.S.; Chakraborty, C. A SARS-CoV-2 vaccine candidate: In-silico cloning and validation. Inform. Med. Unlocked 2020, 20, 100394. [Google Scholar] [CrossRef]
- Woo, P.C.; Lau, S.K.; Lam, C.S.; Lai, K.K.; Huang, Y.; Lee, P.; Luk, G.S.; Dyrting, K.C.; Chan, K.H.; Yuen, K.Y. Comparative analysis of complete genome sequences of three avian coronaviruses reveals a novel group 3c coronavirus. J. Virol. 2009, 83, 908–917. [Google Scholar] [CrossRef]
- Woo, P.C.; Wang, M.; Lau, S.K.; Xu, H.; Poon, R.W.; Guo, R.; Wong, B.H.; Gao, K.; Tsoi, H.W.; Huang, Y.; et al. Comparative analysis of twelve genomes of three novel group 2c and group 2d coronaviruses reveals unique group and subgroup features. J. Virol. 2007, 81, 1574–1585. [Google Scholar] [CrossRef]
- Woo, P.C.; Lau, S.K.; Chu, C.M.; Chan, K.H.; Tsoi, H.W.; Huang, Y.; Wong, B.H.; Poon, R.W.; Cai, J.J.; Luk, W.K.; et al. Characterization and complete genome sequence of a novel coronavirus, coronavirus HKU1, from patients with pneumonia. J. Virol. 2005, 79, 884–895. [Google Scholar] [CrossRef]
- V’kovski, P.; Kratzel, A.; Steiner, S.; Stalder, H.; Thiel, V. Coronavirus biology and replication: Implications for SARS-CoV-2. Nat. Rev. Microbiol. 2021, 19, 155–170. [Google Scholar] [CrossRef] [PubMed]
- Alluwaimi, A.M.; Alshubaith, I.H.; Al-Ali, A.M.; Abohelaika, S. The Coronaviruses of Animals and Birds: Their Zoonosis, Vaccines, and Models for SARS-CoV and SARS-CoV2. Front. Vet. Sci. 2020, 7, 582287. [Google Scholar] [CrossRef] [PubMed]
- Woo, P.C.; Huang, Y.; Lau, S.K.; Yuen, K.Y. Coronavirus genomics and bioinformatics analysis. Viruses 2010, 2, 1804–1820. [Google Scholar] [CrossRef] [PubMed]
- Harvey, W.T.; Carabelli, A.M.; Jackson, B.; Gupta, R.K.; Thomson, E.C.; Harrison, E.M.; Ludden, C.; Reeve, R.; Rambaut, A.; Peacock, S.J.; et al. SARS-CoV-2 variants, spike mutations and immune escape. Nat. Rev. Microbiol. 2021, 19, 409–424. [Google Scholar] [CrossRef]
- Duchene, S.; Featherstone, L.; Haritopoulou-Sinanidou, M.; Rambaut, A.; Lemey, P.; Baele, G. Temporal signal and the phylodynamic threshold of SARS-CoV-2. Virus Evol. 2020, 6, veaa061. [Google Scholar] [CrossRef]
- Worobey, M.; Pekar, J.; Larsen, B.B.; Nelson, M.I.; Hill, V.; Joy, J.B.; Rambaut, A.; Suchard, M.A.; Wertheim, J.O.; Lemey, P. The emergence of SARS-CoV-2 in Europe and North America. Science 2020, 370, 564–570. [Google Scholar] [CrossRef]
- Banoun, H. Evolution of SARS-CoV-2: Review of Mutations, Role of the Host Immune System. Nephron 2021, 145, 392–403. [Google Scholar] [CrossRef]
- Muth, D.; Corman, V.M.; Roth, H.; Binger, T.; Dijkman, R.; Gottula, L.T.; Gloza-Rausch, F.; Balboni, A.; Battilani, M.; Rihtarič, D.; et al. Attenuation of replication by a 29 nucleotide deletion in SARS-coronavirus acquired during the early stages of human-to-human transmission. Sci. Rep. 2018, 8, 15177. [Google Scholar] [CrossRef]
- Cosar, B.; Karagulleoglu, Z.Y.; Unal, S.; Ince, A.T.; Uncuoglu, D.B.; Tuncer, G.; Kilinc, B.R.; Ozkan, Y.E.; Ozkoc, H.C.; Demir, I.N.; et al. SARS-CoV-2 Mutations and their Viral Variants. Cytokine Growth Factor Rev. 2022, 63, 10–22. [Google Scholar] [CrossRef]
- Viana, R.; Moyo, S.; Amoako, D.G.; Tegally, H.; Scheepers, C.; Althaus, C.L.; Anyaneji, U.J.; Bester, P.A.; Boni, M.F.; Chand, M.; et al. Rapid epidemic expansion of the SARS-CoV-2 Omicron variant in southern Africa. Nature 2022, 603, 679–686. [Google Scholar] [CrossRef]
- WHO. COVID-19 New Variants: Knowledge Gaps and Research; WHO R&D Blueprint: Geneva, Switzerland, 2021. [Google Scholar]
- Naveca, F.; Nascimento, V.; Souza, V.; Corado, A.; Nascimento, F.; Silva, G.; Costa, Á.; Duarte, D.; Pessoa, K.; Gonçalves, L.; et al. Phylogenetic Relationship of SARS-CoV-2 Sequences from Amazonas with Emerging Brazilian Variants Harboring Mutations E484K and N501Y in the Spike Protein. 2021. Available online: https://virological.org/t/phylogenetic-relationship-of-sars-cov-2-sequences-from-amazonas-with-emerging-brazilian-variants-harboring-mutations-e484k-and-n501y-in-the-spike-protein/585 (accessed on 26 April 2022).
- Zhang, W.; Davis, B.D.; Chen, S.S.; Sincuir Martinez, J.M.; Plummer, J.T.; Vail, E. Emergence of a Novel SARS-CoV-2 Variant in Southern California. JAMA 2021, 325, 1324–1326. [Google Scholar] [CrossRef] [PubMed]
- Tian, D.; Sun, Y.; Xu, H.; Ye, Q. The emergence and epidemic characteristics of the highly mutated SARS-CoV-2 Omicron variant. J. Med. Virol. 2022, 94, 2376–2383. [Google Scholar] [CrossRef] [PubMed]
- Kar, S. COVID-19 Alert: WHO Says New Virus Strain ‘XE’ Could be Most Contagious So Far. 2022. Available online: https://www.hindustantimes.com/world-news/covid19-alert-who-says-new-virus-strain-xe-could-be-most-contagious-so-far-101648894695415.html (accessed on 26 April 2022).
- Zhang, J.; Zhang, Y.; Kang, J.-Y.; Chen, S.; He, Y.; Han, B.; Liu, M.-F.; Lu, L.; Li, L.; Yi, Z.; et al. Potential transmission chains of variant B.1.1.7 and co-mutations of SARS-CoV-2. Cell Discov. 2021, 7, 44. [Google Scholar] [CrossRef] [PubMed]
- Tegally, H.; Wilkinson, E.; Giovanetti, M.; Iranzadeh, A.; Fonseca, V.; Giandhari, J.; Doolabh, D.; Pillay, S.; San, E.J.; Msomi, N.; et al. Detection of a SARS-CoV-2 variant of concern in South Africa. Nature 2021, 592, 438–443. [Google Scholar] [CrossRef] [PubMed]
- Negi, S.S.; Schein, C.H.; Braun, W. Regional and temporal coordinated mutation patterns in SARS-CoV-2 spike protein revealed by a clustering and network analysis. Sci. Rep. 2022, 12, 1128. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.-J. SARS-COV-2 γ variant acquires spike P681H or P681R for improved viral fitness. bioRxiv 2021. [Google Scholar] [CrossRef]
- Kemp, S.A.; Collier, D.A.; Datir, R.P.; Ferreira, I.A.T.M.; Gayed, S.; Jahun, A.; Hosmillo, M.; Rees-Spear, C.; Mlcochova, P.; Lumb, I.U.; et al. SARS-CoV-2 evolution during treatment of chronic infection. Nature 2021, 592, 277–282. [Google Scholar] [CrossRef]
- Bal, A.; Destras, G.; Gaymard, A.; Bouscambert-Duchamp, M.; Valette, M.; Escuret, V.; Frobert, E.; Billaud, G.; Trouillet-Assant, S.; Cheynet, V.; et al. Molecular characterization of SARS-CoV-2 in the first COVID-19 cluster in France reveals an amino acid deletion in nsp2 (Asp268del). CMI 2020, 26, 960–962. [Google Scholar] [CrossRef]
- Robson, F.; Khan, K.S.; Le, T.K.; Paris, C.; Demirbag, S.; Barfuss, P.; Rocchi, P.; Ng, W.-L. Coronavirus RNA Proofreading: Molecular Basis and Therapeutic Targeting. Mol. Cell 2020, 79, 710–727. [Google Scholar] [CrossRef]
- Fehr, A.R.; Perlman, S. Coronaviruses: An overview of their replication and pathogenesis. Methods Mol. Biol. 2015, 1282, 1–23. [Google Scholar]
- Jacob, C.O. On the genetics and immunopathogenesis of COVID-19. J. Clin. Immunol. 2020, 220, 108591. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, A.; Kulcsar, K.; Misra, V.; Frieman, M.; Mossman, K. Bats and Coronaviruses. Viruses 2019, 11, 41. [Google Scholar] [CrossRef] [PubMed]
- Astuti, I.; Ysrafil. Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2): An overview of viral structure and host response. Diabetes Metab. Syndr. 2020, 14, 407–412. [Google Scholar] [CrossRef] [PubMed]
- Harapan, H.; Itoh, N.; Yufika, A.; Winardi, W.; Keam, S.; Te, H.; Megawati, D.; Hayati, Z.; Wagner, A.L.; Mudatsir, M. Coronavirus disease 2019 (COVID-19): A literature review. J. Infect. Public Health 2020, 13, 667–673. [Google Scholar] [CrossRef]
- Mathuria, J.P.; Yadav, R. Rajkumar, Laboratory diagnosis of SARS-CoV-2—A review of current methods. J. Infect. Public Health 2020, 13, 901–905. [Google Scholar] [CrossRef]
- Deng, H.; Jayawardena, A.; Chan, J.; Tan, S.M.; Alan, T.; Kwan, P. An ultra-portable, self-contained point-of-care nucleic acid amplification test for diagnosis of active COVID-19 infection. Sci. Rep. 2021, 11, 15176. [Google Scholar] [CrossRef]
- FDA. OPTI SARS-CoV-2 RT-PCR Test. 2020. Available online: https://www.fda.gov/media/137739/download (accessed on 26 April 2022).
- WHO. Protocol: Real-Time RT-PCR Assays for the Detection of SARS-CoV-2; Institut Pasteur: Paris, France, 2019. [Google Scholar]
- He, J.-L.; Luo, L.; Luo, Z.-D.; Lyu, J.-X.; Ng, M.-Y.; Shen, X.-P.; Wen, Z. Diagnostic performance between CT and initial real-time RT-PCR for clinically suspected 2019 coronavirus disease (COVID-19) patients outside Wuhan, China. Respir. Med. 2020, 168, 105980. [Google Scholar] [CrossRef]
- Dzimianski, J.V.; Lorig-Roach, N.; O’Rourke, S.M.; Alexander, D.L.; Kimmey, J.M.; DuBois, R.M. Rapid and sensitive detection of SARS-CoV-2 antibodies by biolayer interferometry. Sci. Rep. 2020, 10, 21738. [Google Scholar] [CrossRef]
- FDA. Megna Health Rapid COVID-19 IgM/IgG Combo Test Kit. 2020. Available online: https://www.fda.gov/media/140297/download (accessed on 26 April 2022).
- Corman, V.M.; Landt, O.; Kaiser, M.; Molenkamp, R.; Meijer, A.; Chu, D.K.W.; Bleicker, T.; Brünink, S.; Schneider, J.; Schmidt, M.L.; et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro Surveill. 2020, 25, 2000045. [Google Scholar] [CrossRef]
- Ai, T.; Yang, Z.; Hou, H.; Zhan, C.; Chen, C.; Lv, W.; Tao, Q.; Sun, Z.; Xia, L. Correlation of Chest CT and RT-PCR Testing for Coronavirus Disease 2019 (COVID-19) in China: A Report of 1014 Cases. Radiology 2020, 296, E32–E40. [Google Scholar] [CrossRef]
- Gheblawi, M.; Wang, K.; Viveiros, A.; Nguyen, Q.; Zhong, J.-C.; Turner, A.J.; Raizada, M.K.; Grant, M.B.; Oudit, G.Y. Angiotensin-Converting Enzyme 2: SARS-CoV-2 Receptor and Regulator of the Renin-Angiotensin System: Celebrating the 20th Anniversary of the Discovery of ACE2. Circ. Res. 2020, 126, 1456–1474. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Yang, C.; Xu, X.-F.; Xu, W.; Liu, S.-W. Structural and functional properties of SARS-CoV-2 spike protein: Potential antivirus drug development for COVID-19. Acta Pharmacol. Sin. 2020, 41, 1141–1149. [Google Scholar] [CrossRef] [PubMed]
- Samavati, L.; Uhal, B.D. ACE2, Much More Than Just a Receptor for SARS-COV-2. Front. Cell. Infect. Microbiol. 2020, 10, 317. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; He, S.; Wang, X.; Yan, Y.; Liu, J.; Wu, S.; Liu, S.; Lei, Y.; Chen, M.; Li, L.; et al. Rapid lateral flow immunoassay for the fluorescence detection of SARS-CoV-2 RNA. Nat. Biomed. Eng. 2020, 4, 1150–1158. [Google Scholar] [CrossRef]
- Soleimani, R.; Khourssaji, M.; Gruson, D.; Rodriguez-Villalobos, H.; Berghmans, M.; Belkhir, L.; Yombi, J.C.; Kabamba-Mukadi, B. Clinical usefulness of fully automated chemiluminescent immunoassay for quantitative antibody measurements in COVID-19 patients. J. Med. Virol. 2021, 93, 1465–1477. [Google Scholar] [CrossRef]
- MacMullan, M.A.; Ibrayeva, A.; Trettner, K.; Deming, L.; Das, S.; Tran, F.; Moreno, J.R.; Casian, J.G.; Chellamuthu, P.; Kraft, J.; et al. ELISA detection of SARS-CoV-2 antibodies in saliva. Sci. Rep. 2020, 10, 20818. [Google Scholar] [CrossRef]
- FDA. Accelerated Emergency Use Authorization (EUA) Summary COVID-19 Elisa IGG Antibody Test (Mount Sinai Laboratory, New York). 2020. Available online: https://www.fda.gov/media/137029/download (accessed on 26 April 2022).
- Guglielmi, G. Fast coronavirus tests: What they can and can’t do. Nature 2020, 585, 496–498. [Google Scholar] [CrossRef]
- Poudel, A.; Poudel, Y.; Adhikari, A.; Aryal, B.B.; Dangol, D.; Bajracharya, T.; Maharjan, A.; Gautam, R. D-dimer as a biomarker for assessment of COVID-19 prognosis: D-dimer levels on admission and its role in predicting disease outcome in hospitalized patients with COVID-19. PLoS ONE 2021, 16, e0256744. [Google Scholar] [CrossRef]
- Yamamoto, K.; Ohmagari, N. Microbiological Testing for Coronavirus Disease. JMA J. 2021, 4, 67–75. [Google Scholar]
- Liu, Y.; Zhan, L.; Qin, Z.; Sackrison, J.; Bischof, J.C. Ultrasensitive and Highly Specific Lateral Flow Assays for Point-of-Care Diagnosis. ACS Nano 2021, 15, 3593–3611. [Google Scholar] [CrossRef]
- Khoury, D.S.; Cromer, D.; Reynaldi, A.; Schlub, T.E.; Wheatley, A.K.; Juno, J.A.; Subbarao, K.; Kent, S.J.; Triccas, J.A.; Davenport, M.P. Neutralizing antibody levels are highly predictive of immune protection from symptomatic SARS-CoV-2 infection. Nat. Med. 2021, 27, 1205–1211. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Petitjean, S.J.L.; Koehler, M.; Zhang, Q.; Dumitru, A.C.; Chen, W.; Derclaye, S.; Vincent, S.P.; Soumillion, P.; Alsteens, D. Molecular interaction and inhibition of SARS-CoV-2 binding to the ACE2 receptor. Nat. Commun. 2020, 11, 4541. [Google Scholar] [CrossRef] [PubMed]
- Choudhury, A.; Mukherjee, S. In silico studies on the comparative characterization of the interactions of SARS-CoV-2 spike glycoprotein with ACE-2 receptor homologs and human TLRs. J. Med. Virol. 2020, 92, 2105–2113. [Google Scholar] [CrossRef] [PubMed]
- McBride, R.; van Zyl, M.; Fielding, B.C. The coronavirus nucleocapsid is a multifunctional protein. Viruses 2014, 6, 2991–3018. [Google Scholar] [CrossRef] [PubMed]
- Surjit, M.; Lal, S.K. The SARS-CoV nucleocapsid protein: A protein with multifarious activities. Infect. Genet. Evol. 2008, 8, 397–405. [Google Scholar] [CrossRef] [PubMed]
- Thomas, S. The Structure of the Membrane Protein of SARS-CoV-2 Resembles the Sugar Transporter SemiSWEET. Pathog. Immun. 2020, 5, 342–363. [Google Scholar] [CrossRef] [PubMed]
- Mandala, V.S.; McKay, M.J.; Shcherbakov, A.A.; Dregni, A.J.; Kolocouris, A.; Hong, M. Structure and drug binding of the SARS-CoV-2 envelope protein transmembrane domain in lipid bilayers. Nat. Struct. Mol. Biol. 2020, 27, 1202–1208. [Google Scholar] [CrossRef]
- Schoeman, D.; Fielding, B.C. Coronavirus envelope protein: Current knowledge. Virol. J. 2019, 16, 69. [Google Scholar] [CrossRef]
- Garcia-Olivé, I.; Sintes, H.; Radua, J.; Abad Capa, J.; Rosell, A. D-dimer in patients infected with COVID-19 and suspected pulmonary embolism. Respir. Med. 2020, 169, 106023. [Google Scholar] [CrossRef]
- Salvatore, P.P.; Shah, M.M.; Ford, L.; Delaney, A.; Hsu, C.H.; Tate, J.E.; Kirking, H.L. Quantitative comparison of SARS-CoV-2 nucleic acid amplification test and antigen testing algorithms: A decision analysis simulation model. BMC Public Health 2022, 22, 82. [Google Scholar] [CrossRef]
- Chau, C.H.; Strope, J.D.; Figg, W.D. COVID-19 Clinical Diagnostics and Testing Technology. Pharmacotherapy 2020, 40, 857–868. [Google Scholar] [CrossRef] [PubMed]
- Zoabi, Y.; Deri-Rozov, S.; Shomron, N. Machine learning-based prediction of COVID-19 diagnosis based on symptoms. npj Digit. Med. 2021, 4, 3. [Google Scholar] [CrossRef] [PubMed]
- Ahsan, M.M.; Nazim, R.; Siddique, Z.; Huebner, P. Detection of COVID-19 Patients from CT Scan and Chest X-ray Data Using Modified MobileNetV2 and LIME. Healthcare 2021, 9, 1099. [Google Scholar] [CrossRef] [PubMed]
- Sidiq, Z.; Hanif, M.; Dwivedi, K.K.; Chopra, K.K. Benefits and limitations of serological assays in COVID-19 infection. Indian J. Tuberc. 2020, 67 (Suppl. S4), S163–S166. [Google Scholar] [CrossRef]
- Chantal Babb de Villiers, L.B.; Cook, S.; Janus, J. SARS-CoV-2 Variants. 2021. Available online: https://www.finddx.org/wp-content/uploads/2021/03/COVID-variants-report-FINAL-12MAR2021.pdf (accessed on 26 April 2022).
- MacKay, M.J.; Hooker, A.C.; Afshinnekoo, E.; Salit, M.; Kelly, J.; Feldstein, J.V.; Haft, N.; Schenkel, D.; Nambi, S.; Cai, Y.; et al. The COVID-19 XPRIZE and the need for scalable, fast, and widespread testing. Nat. Biotechnol. 2020, 38, 1021–1024. [Google Scholar] [CrossRef]
- Artesi, M.; Bontems, S.; Göbbels, P.; Franckh, M.; Maes, P.; Boreux, R.; Meex, C.; Melin, P.; Hayette, M.P.; Bours, V.; et al. A Recurrent Mutation at Position 26340 of SARS-CoV-2 Is Associated with Failure of the E Gene Quantitative Reverse Transcription-PCR Utilized in a Commercial Dual-Target Diagnostic Assay. J. Clin. Microbiol. 2020, 58, e01598-20. [Google Scholar] [CrossRef]
- WHO. Transmission of SARS-CoV-2: Implications for Infection Prevention Precautions. 2020. Available online: https://www.who.int/news-room/commentaries/detail/transmission-of-sars-cov-2-implications-for-infection-prevention-precautions (accessed on 26 April 2022).
- Hamming, I.; Timens, W.; Bulthuis, M.L.; Lely, A.T.; Navis, G.; van Goor, H. Tissue distribution of ACE2 protein, the functional receptor for SARS coronavirus. A first step in understanding SARS pathogenesis. J. Pathol. 2004, 203, 631–637. [Google Scholar] [CrossRef]
- Lodish, H.; Berk, A.; Zipursky, L.; Matsudaira, P.; Baltimore, D.; Darnell, J. Molecular Cell Biology; W. H. Freeman: New York, NY, USA, 2000; Volume 4. [Google Scholar]
- Janeway, C.A. Immunobiology: The Immune System in Health and Disease; Garland Science: New York, NY, USA, 2001; Volume 5. [Google Scholar]
- Dotiwala, F.; Mulik, S.; Polidoro, R.B.; Ansara, J.A.; Burleigh, B.A.; Walch, M.; Gazzinelli, R.T.; Lieberman, J. Killer lymphocytes use granulysin, perforin and granzymes to kill intracellular parasites. Nat. Med. 2016, 22, 210–216. [Google Scholar] [CrossRef]
- Le Page, C.; Génin, P.; Baines, M.G.; Hiscott, J. Interferon activation and innate immunity. Rev. Immunogenet. 2000, 2, 374–386. [Google Scholar]
- Alberts, B.; Johnson, A.; Lewis, J.; Raff, M.; Roberts, K.; Walter, P. Molecular Biology of the Cell; Garland Science: New York, NY, USA, 2002; Volume 4. [Google Scholar]
- Hosseini, A.; Hashemi, V.; Shomali, N.; Asghari, F.; Gharibi, T.; Akbari, M.; Gholizadeh, S.; Jafari, A. Innate and adaptive immune responses against coronavirus. Biomed Pharm. 2020, 132, 110859. [Google Scholar] [CrossRef]
- Catanzaro, M.; Fagiani, F.; Racchi, M.; Corsini, E.; Govoni, S.; Lanni, C. Immune response in COVID-19: Addressing a pharmacological challenge by targeting pathways triggered by SARS-CoV-2. Signal Transduct. Taregt. Ther. 2020, 5, 84. [Google Scholar] [CrossRef] [PubMed]
- Tay, M.Z.; Poh, C.M.; Rénia, L.; MacAry, P.A.; Ng, L.F.P. The trinity of COVID-19: Immunity, inflammation and intervention. Nat. Rev. Immunol. 2020, 20, 363–374. [Google Scholar] [CrossRef] [PubMed]
- Coperchini, F.; Chiovato, L.; Croce, L.; Magri, F.; Rotondi, M. The cytokine storm in COVID-19: An overview of the involvement of the chemokine/chemokine-receptor system. Cytokine Growth Factor Rev. 2020, 53, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Costela-Ruiz, V.J.; Illescas-Montes, R.; Puerta-Puerta, J.M.; Ruiz, C.; Melguizo-Rodríguez, L. SARS-CoV-2 infection: The role of cytokines in COVID-19 disease. Cytokine Growth Factor Rev. 2020, 54, 62–75. [Google Scholar] [CrossRef]
- Subbarao, K.; Mahanty, S. Respiratory Virus Infections: Understanding COVID-19. Immunity 2020, 52, 905–909. [Google Scholar] [CrossRef]
- Verdecchia, P.; Cavallini, C.; Spanevello, A.; Angeli, F. The pivotal link between ACE2 deficiency and SARS-CoV-2 infection. Eur. J. Intern. Med. 2020, 76, 14–20. [Google Scholar] [CrossRef]
- García, L.F. Immune Response, Inflammation, and the Clinical Spectrum of COVID-19. Front. Immunol. 2020, 11, 1441. [Google Scholar] [CrossRef]
- Shah, V.K.; Firmal, P.; Alam, A.; Ganguly, D.; Chattopadhyay, S. Overview of Immune Response During SARS-CoV-2 Infection: Lessons From the Past. Front. Immunol. 2020, 11, 1949. [Google Scholar] [CrossRef]
- Felsenstein, S.; Herbert, J.A.; McNamara, P.S.; Hedrich, C.M. COVID-19: Immunology and treatment options. Clin. Immunol. 2020, 215, 108448. [Google Scholar] [CrossRef]
- Fung, T.S.; Liu, D.X. Human Coronavirus: Host-Pathogen Interaction. Annu. Rev. Microbiol. 2019, 73, 529–557. [Google Scholar] [CrossRef]
- Ramasamy, S.; Subbian, S. Critical Determinants of Cytokine Storm and Type I Interferon Response in COVID-19 Pathogenesis. Clin. Miocrobiol. Rev. 2021, 34, e00299-20. [Google Scholar] [CrossRef] [PubMed]
- Del Valle, D.M.; Kim-Schulze, S.; Huang, H.-H.; Beckmann, N.D.; Nirenberg, S.; Wang, B.; Lavin, Y.; Swartz, T.H.; Madduri, D.; Stock, A.; et al. An inflammatory cytokine signature predicts COVID-19 severity and survival. Nat. Med. 2020, 26, 1636–1643. [Google Scholar] [CrossRef] [PubMed]
- Carty, M.; Guy, C.; Bowie, A.G. Detection of Viral Infections by Innate Immunity. Biochem. Pharmacol. 2021, 183, 114316. [Google Scholar] [CrossRef] [PubMed]
- Ong, E.Z.; Chan, Y.F.Z.; Leong, W.Y.; Lee, N.M.Y.; Kalimuddin, S.; Haja Mohideen, S.M.; Chan, K.S.; Tan, A.T.; Bertoletti, A.; Ooi, E.E.; et al. A Dynamic Immune Response Shapes COVID-19 Progression. Cell Host Microbe 2020, 27, 879–882.e2. [Google Scholar] [CrossRef]
- Mogensen, T.H. Pathogen recognition and inflammatory signaling in innate immune defenses. Clin. Microiol. Rev. 2009, 22, 240–273. [Google Scholar] [CrossRef] [PubMed]
- Fullam, A.; Schröder, M. DExD/H-box RNA helicases as mediators of anti-viral innate immunity and essential host factors for viral replication. Biochim. Biophys. Acta 2013, 1829, 854–865. [Google Scholar] [CrossRef]
- Weber, F. Antiviral Innate Immunity: Introduction. Encycl. Virol. 2021, 577–583. [Google Scholar] [CrossRef]
- Paul, S.; Lal, G. The Molecular Mechanism of Natural Killer Cells Function and Its Importance in Cancer Immunotherapy. Front. Immunol. 2017, 8, 1124. [Google Scholar] [CrossRef]
- Zhao, N.; Di, B.; Xu, L.-L. The NLRP3 inflammasome and COVID-19: Activation, pathogenesis and therapeutic strategies. Cytokine Growth Factor Rev. 2021, 61, 2–15. [Google Scholar] [CrossRef]
- de Rivero Vaccari, J.C.; Dietrich, W.D.; Keane, R.W.; de Rivero Vaccari, J.P. The Inflammasome in Times of COVID-19. Front. Immunol. 2020, 11, 583373. [Google Scholar] [CrossRef]
- Kawasaki, T.; Kawai, T. Toll-Like Receptor Signaling Pathways. Front. Immunol. 2014, 5, 461. [Google Scholar] [CrossRef] [PubMed]
- Lester, S.N.; Li, K. Toll-like receptors in antiviral innate immunity. J. Mol. Biol. 2014, 426, 1246–1264. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Fan, Y.; Lai, Y.; Han, T.; Li, Z.; Zhou, P.; Pan, P.; Wang, W.; Hu, D.; Liu, X.; et al. Coronavirus infections and immune responses. J. Med. Virol. 2020, 92, 424–432. [Google Scholar] [CrossRef] [PubMed]
- Lauxmann, M.; Santucci, N.; Autran-Gomez, A. The SARS-CoV-2 Coronavirus and the COVID-19 Outbreak. Int. Braz. J. Urol. Off. J. Braz. Soc. Urol. 2020, 46, 6–18. [Google Scholar] [CrossRef] [PubMed]
- Bolívar, B.E.; Vogel, T.P.; Bouchier-Hayes, L. Inflammatory caspase regulation: Maintaining balance between inflammation and cell death in health and disease. FEBS J. 2019, 286, 2628–2644. [Google Scholar] [CrossRef]
- Zahid, A.; Li, B.; Kombe, A.J.K.; Jin, T.; Tao, J. Pharmacological Inhibitors of the NLRP3 Inflammasome. Front. Immunol. 2019, 10, 2538. [Google Scholar] [CrossRef] [PubMed]
- Conti, P.; Younes, A. Coronavirus COV-19/SARS-CoV-2 affects women less than men: Clinical response to viral infection. J. Biol. Regul. Homeost 2020, 34, 339–343. [Google Scholar]
- Clay, C.; Donart, N.; Fomukong, N.; Knight, J.B.; Lei, W.; Price, L.; Hahn, F.; Van Westrienen, J.; Harrod, K.S. Primary severe acute respiratory syndrome coronavirus infection limits replication but not lung inflammation upon homologous rechallenge. J. Virol. 2012, 86, 4234–4244. [Google Scholar] [CrossRef]
- Yewdell, J.W.; Bennink, J.R. Mechanisms of Viral Interference with MHC Class I Antigen Processing and Presentation. Annu. Rev. Cell Dev. Biol. 1999, 15, 579–606. [Google Scholar] [CrossRef]
- Gaebler, C.; Wang, Z.; Lorenzi, J.C.C.; Muecksch, F.; Finkin, S.; Tokuyama, M.; Cho, A.; Jankovic, M.; Schaefer-Babajew, D.; Oliveira, T.Y.; et al. Evolution of antibody immunity to SARS-CoV-2. Nature 2021, 591, 639–644. [Google Scholar] [CrossRef]
- Callow, K.A.; Parry, H.F.; Sergeant, M.; Tyrrell, D.A. The time course of the immune response to experimental coronavirus infection of man. Epidemiol. Infect. 1990, 105, 435–446. [Google Scholar] [CrossRef] [PubMed]
- Punt, J.; Owen, J.A.; Stranford, S.A.; Jones, P.P.; Kuby, J. Kuby Immunology, 8th ed.; W.H. Freeman/Macmillan Learning: New York, NY, USA, 2019. [Google Scholar]
- Qin, C.; Zhou, L.; Hu, Z.; Zhang, S.; Yang, S.; Tao, Y.; Xie, C.; Ma, K.; Shang, K.; Wang, W.; et al. Dysregulation of immune response in patients with COVID-19 in Wuhan, China. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2020, 71, 762–768. [Google Scholar] [CrossRef] [PubMed]
- Yang, A.-P.; Liu, J.-P.; Tao, W.-Q.; Li, H.-M. The diagnostic and predictive role of NLR, d-NLR and PLR in COVID-19 patients. Int. Immunopharmacol. 2020, 84, 106504. [Google Scholar] [CrossRef] [PubMed]
- Gadi, N.; Wu, S.C.; Spihlman, A.P.; Moulton, V.R. What’s Sex Got to Do With COVID-19? Gender-Based Differences in the Host Immune Response to Coronaviruses. Front. Immunol. 2020, 11, 2147. [Google Scholar] [CrossRef] [PubMed]
- Pivonello, R.; Auriemma, R.S.; Pivonello, C.; Isidori, A.M.; Corona, G.; Colao, A.; Millar, R.P. Sex Disparities in COVID-19 Severity and Outcome: Are Men Weaker or Women Stronger? Neuroendocrinology 2020, 111, 1066–1085. [Google Scholar] [CrossRef]
- Paterni, I.; Granchi, C.; Katzenellenbogen, J.A.; Minutolo, F. Estrogen receptors alpha (ERα) and beta (ERβ): Subtype-selective ligands and clinical potential. Steroids 2014, 90, 13–29. [Google Scholar] [CrossRef]
- Kovats, S. Estrogen receptors regulate innate immune cells and signaling pathways. Cell. Immunol. 2015, 294, 63–69. [Google Scholar] [CrossRef]
- Koenig, A.; Buskiewicz, I.; Huber, S.A. Age-Associated Changes in Estrogen Receptor Ratios Correlate with Increased Female Susceptibility to Coxsackievirus B3-Induced Myocarditis. Front. Immunol. 2017, 8, 1585. [Google Scholar] [CrossRef]
- Stilhano, R.S.; Costa, A.J.; Nishino, M.S.; Shams, S.; Bartolomeo, C.S.; Breithaupt-Faloppa, A.C.; Silva, E.A.; Ramirez, A.L.; Prado, C.M.; Ureshino, R.P. SARS-CoV-2 and the possible connection to ERs, ACE2, and RAGE: Focus on susceptibility factors. FASEB J. 2020, 34, 14103–14119. [Google Scholar] [CrossRef]
- Ghosh, M.; Rodriguez-Garcia, M.; Wira, C.R. The immune system in menopause: Pros and cons of hormone therapy. J. Steroid. Biochem. Mol. Biol. 2014, 142, 171–175. [Google Scholar] [CrossRef]
- Rubtsova, K.; Marrack, P.; Rubtsov, A.V. TLR7, IFNγ, and T-bet: Their roles in the development of ABCs in female-biased autoimmunity. Cell. Immunol. 2015, 294, 80–83. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, T.; Ellingson, M.K.; Wong, P.; Israelow, B.; Lucas, C.; Klein, J.; Silva, J.; Mao, T.; Oh, J.E.; Tokuyama, M.; et al. Sex differences in immune responses that underlie COVID-19 disease outcomes. Nature 2020, 588, 315–320. [Google Scholar] [CrossRef] [PubMed]
- Souyris, M.; Mejía, J.E.; Chaumeil, J.; Guéry, J.C. Female predisposition to TLR7-driven autoimmunity: Gene dosage and the escape from X chromosome inactivation. Semin. Immunopathol. 2019, 41, 153–164. [Google Scholar] [CrossRef] [PubMed]
- Ji, Z.; Yan, K.; Li, W.; Hu, H.; Zhu, X. Mathematical and Computational Modeling in Complex Biological Systems. Biomed Res. Int. 2017, 2017, 5958321. [Google Scholar] [PubMed]
- Hufsky, F.; Lamkiewicz, K.; Almeida, A.; Aouacheria, A.; Arighi, C.; Bateman, A.; Baumbach, J.; Beerenwinkel, N.; Brandt, C.; Cacciabue, M.; et al. Computational strategies to combat COVID-19: Useful tools to accelerate SARS-CoV-2 and coronavirus research. Brief. Bioinform. 2021, 22, 642–663. [Google Scholar] [CrossRef] [PubMed]
- Abdellatiif, M.H.; Ali, A.; Ali, A.; Hussien, M.A. Computational studies by molecular docking of some antiviral drugs with COVID-19 receptors are an approach to medication for COVID-19. Open Chem. 2021, 19, 245–264. [Google Scholar] [CrossRef]
- Abel, R.; Paredes Ramos, M.; Chen, Q.; Pérez-Sánchez, H.; Coluzzi, F.; Rocco, M.; Marchetti, P.; Mura, C.; Simmaco, M.; Bourne, P.E.; et al. Computational Prediction of Potential Inhibitors of the Main Protease of SARS-CoV-2. Front. Chem. 2020, 8, 590263. [Google Scholar] [CrossRef]
- Sikora, M.; von Bülow, S.; Blanc, F.E.C.; Gecht, M.; Covino, R.; Hummer, G. Computational epitope map of SARS-CoV-2 spike protein. PLoS Comput. Biol. 2021, 17, e1008790. [Google Scholar] [CrossRef]
- Sadat, S.M.; Aghadadeghi, M.R.; Yousefi, M.; Khodaei, A.; Sadat Larijani, M.; Bahramali, G. Bioinformatics Analysis of SARS-CoV-2 to Approach an Effective Vaccine Candidate Against COVID-19. Mol. Biotechnol. 2021, 63, 389–409. [Google Scholar] [CrossRef]
- Hwang, W.; Lei, W.; Katritsis, N.M.; MacMahon, M.; Chapman, K.; Han, N. Current and prospective computational approaches and challenges for developing COVID-19 vaccines. Adv. Drug Deliv. Rev. 2021, 172, 249–274. [Google Scholar] [CrossRef]
- Al-Qaaneh, A.M.; Alshammari, T.; Aldahhan, R.; Aldossary, H.; Alkhalifah, Z.A.; Borgio, J.F. Genome composition and genetic characterization of SARS-CoV-2. Saudi J. Biol. Sci. 2021, 28, 1978–1989. [Google Scholar] [PubMed]
- Bhattacharya, M.; Sharma, A.R.; Patra, P.; Ghosh, P.; Sharma, G.; Patra, B.C.; Lee, S.S.; Chakraborty, C. Development of epitope-based peptide vaccine against novel coronavirus 2019 (SARS-COV-2): Immunoinformatics approach. J. Med. Virol. 2020, 92, 618–631. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.H.; Koohy, H. In silico identification of vaccine targets for 2019-nCoV. F1000Res 2020, 9, 145. [Google Scholar] [CrossRef] [PubMed]
- Rollenhagen, C.; Sörensen, M.; Rizos, K.; Hurvitz, R.; Bumann, D. Antigen selection based on expression levels during infection facilitates vaccine development for an intracellular pathogen. Proc. Natl. Acad. Sci. USA 2004, 101, 8739–8744. [Google Scholar] [PubMed]
- Adu-Bobie, J.; Capecchi, B.; Serruto, D.; Rappuoli, R.; Pizza, M. Two years into reverse vaccinology. Vaccine 2003, 21, 605–610. [Google Scholar] [CrossRef]
- Zinkernagel, R.M.; Ehl, S.; Aichele, P.; Oehen, S.; Kündig, T.; Hengartner, H. Antigen localisation regulates immune responses in a dose- and time-dependent fashion: A geographical view of immune reactivity. Immunol. Rev. 1997, 156, 199–209. [Google Scholar] [CrossRef]
- Pedersen, S.; Bloch, P.L.; Reeh, S.; Neidhardt, F.C. Patterns of protein synthesis in E. coli: A catalog of the amount of 140 individual proteins at different growth rates. Cell 1978, 14, 179–190. [Google Scholar]
- Ghaemmaghami, S.; Huh, W.K.; Bower, K.; Howson, R.W.; Belle, A.; Dephoure, N.; O’Shea, E.K.; Weissman, J.S. Global analysis of protein expression in yeast. Nature 2003, 425, 737–741. [Google Scholar] [CrossRef]
- Kushwaha, S.K.; Kesarwani, V.; Choudhury, S.; Gandhi, S.; Sharma, S. SARS-CoV-2 transcriptome analysis and molecular cataloguing of immunodominant epitopes for multi-epitope based vaccine design. Genomics 2020, 112, 5044–5054. [Google Scholar]
- Kuntz, I.D.; Blaney, J.M.; Oatley, S.J.; Langridge, R.; Ferrin, T.E. A geometric approach to macromolecule-ligand interactions. J. Mol. Biol. 1982, 161, 269–288. [Google Scholar] [CrossRef]
- DesJarlais, R.L.; Sheridan, R.P.; Dixon, J.S.; Kuntz, I.D.; Venkataraghavan, R. Docking flexible ligands to macromolecular receptors by molecular shape. J. Mol. Chem. 1986, 29, 2149–2153. [Google Scholar] [CrossRef] [PubMed]
- Jindal, D.; Rani, V. In Silico Studies of Phytoconstituents from Piper longum and Ocimum sanctum as ACE2 and TMRSS2 Inhibitors: Strategies to Combat COVID-19. Appl. Biochem. Biotechnol. 2022, 194, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Dar, H.A.; Waheed, Y.; Najmi, M.H.; Ismail, S.; Hetta, H.F.; Ali, A.; Muhammad, K. Multiepitope Subunit Vaccine Design against COVID-19 Based on the Spike Protein of SARS-CoV-2: An In Silico Analysis. J. Immunol. 2020, 2020, 8893483. [Google Scholar] [CrossRef]
- Singh, A.; Thakur, M.; Sharma, L.K.; Chandra, K. Designing a multi-epitope peptide-based vaccine against SARS-CoV-2. Sci. Rep. 2020, 10, 16219. [Google Scholar] [CrossRef]
- Mitra, D.; Pandey, J.; Jain, A.; Swaroop, S. In silico design of multi-epitope-based peptide vaccine against SARS-CoV-2 using its spike protein. J. Biomol. Struct. Dyn. 2021, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Grifoni, A.; Sidney, J.; Zhang, Y.; Scheuermann, R.H.; Peters, B.; Sette, A. A Sequence Homology and Bioinformatic Approach Can Predict Candidate Targets for Immune Responses to SARS-CoV-2. Cell Host Microbe 2020, 27, 671–680.e2. [Google Scholar] [CrossRef] [PubMed]
- Greenbaum, J.; Sidney, J.; Chung, J.; Brander, C.; Peters, B.; Sette, A. Functional classification of class II human leukocyte antigen (HLA) molecules reveals seven different supertypes and a surprising degree of repertoire sharing across supertypes. Immunogenetics 2011, 63, 325–335. [Google Scholar] [CrossRef]
- Weiskopf, D.; Angelo, M.A.; de Azeredo, E.L.; Sidney, J.; Greenbaum, J.A.; Fernando, A.N.; Broadwater, A.; Kolla, R.V.; De Silva, A.D.; de Silva, A.M.; et al. Comprehensive analysis of dengue virus-specific responses supports an HLA-linked protective role for CD8+ T cells. Proc. Natl. Acad. Sci. USA 2013, 110, E2046–E2053. [Google Scholar] [CrossRef]
- Pushpakom, S.; Iorio, F.; Eyers, P.A.; Escott, K.J.; Hopper, S.; Wells, A.; Doig, A.; Guilliams, T.; Latimer, J.; McNamee, C.; et al. Drug repurposing: Progress, challenges and recommendations. Nat. Rev. Drug Discov. 2019, 18, 41–58. [Google Scholar] [CrossRef]
- Wang, J. Fast Identification of Possible Drug Treatment of Coronavirus Disease-19 (COVID-19) through Computational Drug Repurposing Study. J. Chem. Inf. Model. 2020, 60, 3277–3286. [Google Scholar] [CrossRef]
- Ashburn, T.T.; Thor, K.B. Drug repositioning: Identifying and developing new uses for existing drugs. Nat. Rev. Drug Discov. 2004, 3, 673–683. [Google Scholar] [CrossRef] [PubMed]
- Breckenridge, A.; Jacob, R. Overcoming the legal and regulatory barriers to drug repurposing. Nat. Rev. Drug Discov. 2019, 18, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Galindez, G.; Matschinske, J.; Rose, T.D.; Sadegh, S.; Salgado-Albarrán, M.; Späth, J.; Baumbach, J.; Pauling, J.K. Lessons from the COVID-19 pandemic for advancing computational drug repurposing strategies. Nat. Comput. Sci. 2021, 1, 33–41. [Google Scholar] [CrossRef]
- The Genotype-Tissue Expression (GTEx) project. Nat. Gen. 2013, 45, 580–585. [CrossRef]
- Musa, A.; Tripathi, S.; Dehmer, M.; Emmert-Streib, F. L1000 Viewer: A Search Engine and Web Interface for the LINCS Data Repository. Front. Gen. 2019, 10, 557. [Google Scholar] [CrossRef]
- Duan, Q.; Flynn, C.; Niepel, M.; Hafner, M.; Muhlich, J.L.; Fernandez, N.F.; Rouillard, A.D.; Tan, C.M.; Chen, E.Y.; Golub, T.R.; et al. LINCS Canvas Browser: Interactive web app to query, browse and interrogate LINCS L1000 gene expression signatures. Nucleic Acids Res. 2014, 42, W449–W460. [Google Scholar] [CrossRef]
- Wishart, D.S.; Feunang, Y.D.; Guo, A.C.; Lo, E.J.; Marcu, A.; Grant, J.R.; Sajed, T.; Johnson, D.; Li, C.; Sayeeda, Z.; et al. DrugBank 5.0: A major update to the DrugBank database for 2018. Nucleic Acids Res. 2018, 46, D1074–D1082. [Google Scholar] [CrossRef]
- Sterling, T.; Irwin, J.J. ZINC 15--Ligand Discovery for Everyone. J. Chem. Inf. Model. 2015, 55, 2324–2337. [Google Scholar] [CrossRef]
- Reddy, M.R.; Reddy, C.R.; Rathore, R.S.; Erion, M.D.; Aparoy, P.; Reddy, R.N.; Reddanna, P. Free energy calculations to estimate ligand-binding affinities in structure-based drug design. Curr. Pharm. Des. 2014, 20, 3323–3337. [Google Scholar] [CrossRef]
- Friesner, R.A.; Banks, J.L.; Murphy, R.B.; Halgren, T.A.; Klicic, J.J.; Mainz, D.T.; Repasky, M.P.; Knoll, E.H.; Shelley, M.; Perry, J.K.; et al. Glide: A new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 2004, 47, 1739–1749. [Google Scholar] [CrossRef]
- Wang, J.; Morin, P.; Wang, W.; Kollman, P.A. Use of MM-PBSA in reproducing the binding free energies to HIV-1 RT of TIBO derivatives and predicting the binding mode to HIV-1 RT of efavirenz by docking and MM-PBSA. J. Am. Chem. Soc. 2001, 123, 5221–5230. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Kang, X.; Kuntz, I.D.; Kollman, P.A. Hierarchical database screenings for HIV-1 reverse transcriptase using a pharmacophore model, rigid docking, solvation docking, and MM-PB/SA. J. Med. Chem. 2005, 48, 2432–2444. [Google Scholar] [CrossRef] [PubMed]
- Bang, S.; Son, S.; Kim, S.; Shin, H. Disease Pathway Cut for Multi-Target drugs. BMC Bioinform. 2019, 20, 74. [Google Scholar] [CrossRef] [PubMed]
- Behera, S.K.; Vhora, N.; Contractor, D.; Shard, A.; Kumar, D.; Kalia, K.; Jain, A. Computational drug repurposing study elucidating simultaneous inhibition of entry and replication of novel corona virus by Grazoprevir. Sci. Rep. 2021, 11, 7307. [Google Scholar] [CrossRef]
- Bobrowski, T.; Chen, L.; Eastman, R.T.; Itkin, Z.; Shinn, P.; Chen, C.; Guo, H.; Zheng, W.; Michael, S.; Simeonov, A.; et al. Discovery of Synergistic and Antagonistic Drug Combinations against SARS-CoV-2 In Vitro. bioRxiv 2020. [Google Scholar] [CrossRef]
- Abatematteo, F.S.; Niso, M.; Contino, M.; Leopoldo, M.; Abate, C. Multi-Target Directed Ligands (MTDLs) Binding the σ(1) Receptor as Promising Therapeutics: State of the Art and Perspectives. Int. J. Mol. Sci. 2021, 22, 6359. [Google Scholar] [CrossRef]
- Agis-Torres, A.; Sölhuber, M.; Fernandez, M.; Sanchez-Montero, J.M. Multi-Target-Directed Ligands and other Therapeutic Strategies in the Search of a Real Solution for Alzheimer’s Disease. Curr. Neuropharmacol. 2014, 12, 2–36. [Google Scholar] [CrossRef]
- Balasubramanian, S.; Rao, N.M.; Goenka, A.; Roderick, M.; Ramanan, A.V. Coronavirus Disease 2019 (COVID-19) in Children—What We Know So Far and What We Do Not. Indian Pediatr. 2020, 57, 435–442. [Google Scholar] [CrossRef]
- Agostini, M.L.; Andres, E.L.; Sims, A.C.; Graham, R.L.; Sheahan, T.P.; Lu, X.; Smith, E.C.; Case, J.B.; Feng, J.Y.; Jordan, R.; et al. Coronavirus Susceptibility to the Antiviral Remdesivir (GS-5734) Is Mediated by the Viral Polymerase and the Proofreading Exoribonuclease. mBio 2018, 9, e00221-18. [Google Scholar] [CrossRef]
- Balasubramaniam, M.; Reis, R.J.S. Computational target-based drug repurposing of elbasvir, an antiviral drug predicted to bind multiple SARS-CoV-2 proteins. ChemRxiv Prepr. Serv. Chem. 2020. [Google Scholar] [CrossRef]
- De Clercq, E.; Li, G. Approved Antiviral Drugs over the Past 50 Years. Clin. Microbiol. Rev. 2016, 29, 695–747. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Yang, Y.; Li, J.; Wang, M.; Saravanan, K.M.; Wei, J.; Ng, J.T.-Y.; Hossain, T.; Liu, M.; Zhang, H.; et al. A novel virtual screening procedure identifies Pralatrexate as inhibitor of SARS-CoV-2 RdRp and it reduces viral replication in vitro. PLoS Comput. Biol. 2020, 16, e1008489. [Google Scholar] [CrossRef] [PubMed]
- Chaudhari, R.; Tan, Z.; Huang, B.; Zhang, S. Computational polypharmacology: A new paradigm for drug discovery. Expert Opin. Drug Discov. 2017, 12, 279–291. [Google Scholar] [CrossRef] [PubMed]
- Jacobson, I.M.; Lawitz, E.; Kwo, P.Y.; Hézode, C.; Peng, C.-Y.; Howe, A.Y.M.; Hwang, P.; Wahl, J.; Robertson, M.; Barr, E.; et al. Safety and Efficacy of Elbasvir/Grazoprevir in Patients With Hepatitis C Virus Infection and Compensated Cirrhosis: An Integrated Analysis. Gastroenterology 2017, 152, 1372–1382.e2. [Google Scholar] [CrossRef] [PubMed]
- Smith, G.P.; Petrenko, V.A. Phage Display. Chem. Rev. 1997, 97, 391–410. [Google Scholar] [CrossRef] [PubMed]
- Burton, D.R. Phage display. Immunotechnology 1995, 1, 87–94. [Google Scholar] [CrossRef]
- Valdes-Balbin, Y.; Santana-Mederos, D.; Paquet, F.; Fernandez, S.; Climent, Y.; Chiodo, F.; Rodríguez, L.; Sanchez Ramirez, B.; Leon, K.; Hernandez, T.; et al. Molecular Aspects Concerning the Use of the SARS-CoV-2 Receptor Binding Domain as a Target for Preventive Vaccines. ACS Cent. Sci. 2021, 7, 757–767. [Google Scholar] [CrossRef]
- Devarakonda, C.K.V.; Meredith, E.; Ghosh, M.; Shapiro, L.H. Coronavirus Receptors as Immune Modulators. J. Immunol. 2021, 206, 923–929. [Google Scholar] [CrossRef]
- Costa, L.B.; Perez, L.G.; Palmeira, V.A.; Macedo e Cordeiro, T.; Ribeiro, V.T.; Lanza, K.; Simões e Silva, A.C. Insights on SARS-CoV-2 Molecular Interactions With the Renin-Angiotensin System. Front. Cell. Dev. Biol. 2020, 8, 559841. [Google Scholar] [CrossRef]
- Li, F. Structure, Function, and Evolution of Coronavirus Spike Proteins. Annu. Rev. Virol. 2016, 3, 237–261. [Google Scholar] [CrossRef]
- Wang, N.; Shi, X.; Jiang, L.; Zhang, S.; Wang, D.; Tong, P.; Guo, D.; Fu, L.; Cui, Y.; Liu, X.; et al. Structure of MERS-CoV spike receptor-binding domain complexed with human receptor DPP4. Cell Res. 2013, 23, 986–993. [Google Scholar] [CrossRef] [PubMed]
- Carvacho, I.; Piesche, M. RGD-binding integrins and TGF-β in SARS-CoV-2 infections—Novel targets to treat COVID-19 patients? Clin. Transl. Immunol. 2021, 10, e1240. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Mi, L.; Xu, J.; Yu, J.; Wang, X.; Jiang, J.; Xing, J.; Shang, P.; Qian, A.; Li, Y.; et al. Function of HAb18G/CD147 in Invasion of Host Cells by Severe Acute Respiratory Syndrome Coronavirus. J. Infect. Dis. 2005, 191, 755–760. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Chen, W.; Zhang, Z.; Deng, Y.; Lian, J.-Q.; Du, P.; Wei, D.; Zhang, Y.; Sun, X.-X.; Gong, L.; et al. CD147-spike protein is a novel route for SARS-CoV-2 infection to host cells. Signal Transduct. Target. Ther. 2020, 5, 283. [Google Scholar] [CrossRef]
- Kuzmicheva, G.A.; Jayanna, P.K.; Eroshkin, A.M.; Grishina, M.A.; Pereyaslavskaya, E.S.; Potemkin, V.A.; Petrenko, V.A. Mutations in fd phage major coat protein modulate affinity of the displayed peptide. Protein Eng. Des. Sel. 2009, 22, 631–639. [Google Scholar] [CrossRef]
- Petrenko, V.A.; Smith, G.P.; Gong, X.; Quinn, T. A library of organic landscapes on filamentous phage. Protein Eng. Des. Sel. 1996, 9, 797–801. [Google Scholar] [CrossRef]
- Petrenko, V.A.; Gillespie, J.W.; De Plano, L.M.; Shokhen, M.A. Phage-Displayed Mimotopes of SARS-CoV-2 Spike Protein Targeted to Authentic and Alternative Cellular Receptors. Viruses 2022, 14, 384. [Google Scholar] [CrossRef]
- Thaper, D.; Prabha, V. Molecular mimicry: An explanation for autoimmune diseases and infertility. Scand. J. Immunol. 2018, 88, e12697. [Google Scholar] [CrossRef]
- Wildner, G.; Diedrichs-Möhring, M. Molecular Mimicry and Uveitis. Front. Immunol. 2020, 11, 580636. [Google Scholar] [CrossRef]
- Zhou, P.; Yang, X.-L.; Wang, X.-G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.-R.; Zhu, Y.; Li, B.; Huang, C.-L.; et al. Addendum: A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 588, E6. [Google Scholar] [CrossRef]
- Li, Q.; Wu, J.; Nie, J.; Zhang, L.; Hao, H.; Liu, S.; Zhao, C.; Zhang, Q.; Liu, H.; Nie, L.; et al. The Impact of Mutations in SARS-CoV-2 Spike on Viral Infectivity and Antigenicity. Cell 2020, 182, 1284–1294.e9. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Sarma, P.; Kaur, H.; Prajapat, M.; Bhattacharyya, A.; Avti, P.; Sehkhar, N.; Kaur, H.; Bansal, S.; Mahendiratta, S.; et al. Clinically relevant cell culture models and their significance in isolation, pathogenesis, vaccine development, repurposing and screening of new drugs for SARS-CoV-2: A systematic review. Tissue Cell 2021, 70, 101497. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Boorla, V.S.; Banerjee, D.; Chowdhury, R.; Cavener, V.S.; Nissly, R.H.; Gontu, A.; Boyle, N.R.; Vandegrift, K.; Nair, M.S.; et al. Computational prediction of the effect of amino acid changes on the binding affinity between SARS-CoV-2 spike RBD and human ACE2. Proc. Natl. Acad. Sci. USA 2021, 118, e2106480118. [Google Scholar] [CrossRef] [PubMed]
- Millet, J.K.; Jaimes, J.A.; Whittaker, G.R. Molecular diversity of coronavirus host cell entry receptors. FEMS Microbiol. Rev. 2020, 45, fuaa057. [Google Scholar] [CrossRef] [PubMed]
- Petukh, M.; Dai, L.; Alexov, E. SAAMBE: Webserver to Predict the Charge of Binding Free Energy Caused by Amino Acids Mutations. Int. J. Mol. Sci. 2016, 17, 547. [Google Scholar] [CrossRef] [PubMed]
- Xiong, P.; Zhang, C.; Zheng, W.; Zhang, Y. BindProfX: Assessing Mutation-Induced Binding Affinity Change by Protein Interface Profiles with Pseudo-Counts. J. Mol. Biol. 2017, 429, 426–434. [Google Scholar] [CrossRef]
- Shah, P.; Kendall, F.; Khozin, S.; Goosen, R.; Hu, J.; Laramie, J.; Ringel, M.; Schork, N. Artificial intelligence and machine learning in clinical development: A translational perspective. Digit. Med. 2019, 2, 69. [Google Scholar] [CrossRef]
- Dias, R.; Torkamani, A. Artificial intelligence in clinical and genomic diagnostics. Genome Med. 2019, 11, 70. [Google Scholar] [CrossRef]
- Wiens, J.; Shenoy, E.S. Machine Learning for Healthcare: On the Verge of a Major Shift in Healthcare Epidemiology. Clin. Infect. Dis. 2018, 66, 149–153. [Google Scholar] [CrossRef]
- Bansal, A.; Padappayil, R.P.; Garg, C.; Singal, A.; Gupta, M.; Klein, A. Utility of Artificial Intelligence Amidst the COVID 19 Pandemic: A Review. J. Med. Syst. 2020, 44, 156. [Google Scholar] [CrossRef]
- LeCun, Y.; Bengio, Y.; Hinton, G. Deep learning. Nature 2015, 521, 436–444. [Google Scholar] [CrossRef] [PubMed]
- Cao, C.; Liu, F.; Tan, H.; Song, D.; Shu, W.; Li, W.; Zhou, Y.; Bo, X.; Xie, Z. Deep Learning and Its Applications in Biomedicine. Gen. Prot. Bioinform. 2018, 16, 17–32. [Google Scholar] [CrossRef] [PubMed]
- Pham, T.-H.; Qiu, Y.; Zeng, J.; Xie, L.; Zhang, P. A deep learning framework for high-throughput mechanism-driven phenotype compound screening and its application to COVID-19 drug repurposing. Nat. Mach. Intell. 2021, 3, 247–257. [Google Scholar] [CrossRef]
- Wallach, I.; Dzamba, M.; Heifets, A. AtomNet: A Deep Convolutional Neural Network for Bioactivity Prediction in Structure-based Drug Discovery. arXiv 2015, arXiv:1510.02855. [Google Scholar]
- Ton, A.T.; Gentile, F.; Hsing, M.; Ban, F.; Cherkasov, A. Rapid Identification of Potential Inhibitors of SARS-CoV-2 Main Protease by Deep Docking of 1.3 Billion Compounds. Mol. Inform. 2020, 39, e2000028. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.; Khodadoust, M.S.; Olsson, N.; Wagar, L.E.; Fast, E.; Liu, C.L.; Muftuoglu, Y.; Sworder, B.J.; Diehn, M.; Levy, R.; et al. Predicting HLA class II antigen presentation through integrated deep learning. Nat. Biotechnol. 2019, 37, 1332–1343. [Google Scholar] [CrossRef]
- Moore, T.V.; Nishimura, M.I. Improved MHC II epitope prediction—A step towards personalized medicine. Nat. Rev. Clin. Oncol. 2020, 17, 71–72. [Google Scholar] [CrossRef]
- Racle, J.; Michaux, J.; Rockinger, G.A.; Arnaud, M.; Bobisse, S.; Chong, C.; Guillaume, P.; Coukos, G.; Harari, A.; Jandus, C.; et al. Robust prediction of HLA class II epitopes by deep motif deconvolution of immunopeptidomes. Nat. Biotechnol. 2019, 37, 1283–1286. [Google Scholar] [CrossRef]
- Ong, E.; Wong, M.U.; Huffman, A.; He, Y. COVID-19 Coronavirus Vaccine Design Using Reverse Vaccinology and Machine Learning. Front. Immunol. 2020, 11, 1581. [Google Scholar] [CrossRef]

| Type of Variant | Region of Viral Mutations | Types of Mutations | References |
|---|---|---|---|
| α-variant | Spike protein | 60–70 del, 145 del, N501Y, A570D, D614G, P681H, T7161, S982A, D1118H | [27] |
| β-variant | Spike protein | K417N, E484K and N501Y | [28] |
| Epsilon variant | Spike protein | S13I, W152C and L452R | [29] |
| γ-variant | Spike protein | K417T, E484K, N501Y | [30] |
| Eta-variant | Spike protein | E484K ΔH69/ΔV70 deletion | [31] |
| Cluster 5 | Spike protein | ΔH69/ΔV70 deletion | [32] |
| S. No. | Diagnostic Method | Principle of Method | Sample | Time Duration | Detected Component | References |
|---|---|---|---|---|---|---|
| 1. | RT-PCR | Polymerase chain reaction | Sputum, saliva, nasal, phalangeal and tracheal swabs, broncho-alveolar lavage, pleural effusion fluid, blood, faeces, urine and semen | 6–8 h | Viral RNA | [6,46,47] |
| 2. | PCR with fluorescently labelled probes | Polymerase chain reaction | RNA | 6–8 h | Viral RNA | [41] |
| 3. | Chest computerized tomography (CT) | X-ray | Chest X-ray | 30–60 min | Small nodules in the chest | [48,49,50] |
| 4. | Rapid lateral Flow Immunoassay (LFIA) | A liquid sample consisting of analyte transports without the help of capillary action through 3 zones of polymeric strips, upon which molecules that can interact with the analyte are attached | Throat swab or sputum | IgG | [51] | |
| 5. | Automated chemiluminescence immunoassay (CLIA) | Chemiluminescent methods utilizing luminophore markers | serum | IgM and IgG antibodies | [52] | |
| 6. | Manual ELISA | Specific antibody-antigen interactions | Saliva, serum, plasma | 5–6 h | IgG antibodies | [53,54] |
| 7. | Rapid antigen test | Rapid membrane-based lateral flow immunoassay | Saliva | 15–30 min | nucleocapsid protein antigen of the coronavirus SARS-CoV-2 | [55] |
| 8. | Detection of D-dimer levels | Coagulation | Blood | 1–2 days | D-dimer concentrations | [56] |
| 9. | Microbial culture test | NAAT (nucleic acid amplification test) | Nucleic acids | 2–3 days | Viral growth | [57] |
| 10. | Lateral flow antigen test | Immunochromatographic assay | Throat swab or sputum | 30–60 min | Nucleocapsid protein antigen | [58] |
| 11. | Neutralizing antibody test | Rapid membrane-based lateral flow immunoassay | Serum, throat swab or sputum | 30–60 min | Neutralizing antibodies | [59] |
| Diagnostic Technique | Advantages | Disadvantages | References |
|---|---|---|---|
| NAAT |
|
| [68,69] |
| CT scan |
|
| [70,71] |
| Serological immunoassays |
|
| [72] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yadav, D.; Agarwal, S.; Pancham, P.; Jindal, D.; Agarwal, V.; Dubey, P.K.; Jha, S.K.; Mani, S.; Rachana; Dey, A.; et al. Probing the Immune System Dynamics of the COVID-19 Disease for Vaccine Designing and Drug Repurposing Using Bioinformatics Tools. Immuno 2022, 2, 344-371. https://doi.org/10.3390/immuno2020022
Yadav D, Agarwal S, Pancham P, Jindal D, Agarwal V, Dubey PK, Jha SK, Mani S, Rachana, Dey A, et al. Probing the Immune System Dynamics of the COVID-19 Disease for Vaccine Designing and Drug Repurposing Using Bioinformatics Tools. Immuno. 2022; 2(2):344-371. https://doi.org/10.3390/immuno2020022
Chicago/Turabian StyleYadav, Deepshikha, Shriya Agarwal, Pranav Pancham, Divya Jindal, Vinayak Agarwal, Premshankar Kumar Dubey, Saurabh K. Jha, Shalini Mani, Rachana, Abhijit Dey, and et al. 2022. "Probing the Immune System Dynamics of the COVID-19 Disease for Vaccine Designing and Drug Repurposing Using Bioinformatics Tools" Immuno 2, no. 2: 344-371. https://doi.org/10.3390/immuno2020022
APA StyleYadav, D., Agarwal, S., Pancham, P., Jindal, D., Agarwal, V., Dubey, P. K., Jha, S. K., Mani, S., Rachana, Dey, A., Jha, N. K., Kesari, K. K., & Singh, M. (2022). Probing the Immune System Dynamics of the COVID-19 Disease for Vaccine Designing and Drug Repurposing Using Bioinformatics Tools. Immuno, 2(2), 344-371. https://doi.org/10.3390/immuno2020022









