1. Introduction
Breast cancer is one of the most prevalent malignancies among women and remains a leading cause of cancer-related mortality worldwide. According to GLOBOCAN 2022 [
1], breast cancer accounts for the highest female mortality in over 100 countries, underscoring the urgent need for accurate and efficient diagnostic systems that support early detection and improved survival.
In recent years, machine learning (ML) and deep learning (DL) techniques have been increasingly applied to assist in medical image classification, pattern recognition, and diagnostic decision-making. DL approaches such as CNNs have shown remarkable performance in biomedical applications due to their ability to automatically extract hierarchical features from raw data.
In theory, a three-layer CNN can be trained to function as a perfect classifier. This is consistent with the universal approximation theorem, which states that a CNN with at least one sufficiently large hidden layer can approximate any continuous function with arbitrary precision. However, it is not necessarily the optimal choice for BCS classification, as a three-layer network may not adequately capture the underlying structure of the BCS classification problem, which involves complex features that may be better modeled by alternative techniques. Simpler models such as RF, SVM, NB, and KNN, which rely on different learning approaches, may be more suitable given the complexity of the multiclassification task and the size and quality of the data.
However, these models often require large datasets, high computational power, and complex tuning procedures, which can limit their efficiency and interpretability in clinical environments. TMLA models, in contrast, are less computationally demanding and perform robustly on smaller or structured datasets.
To overcome the limitations of both paradigms, hybrid systems combining CNN-based feature extraction with TMLA have emerged as promising alternatives. By leveraging CNNs to capture intricate spatial features and conventional algorithms—such as RF, SVM, NB, and KNN—for classification, these hybrid models aim to balance accuracy, provide an easy-to-understand interpretation, and benefit from the efficiency of feature extractors.
This study focuses on evaluating the performance of such hybrid architectures for breast cancer classification. Specifically, we investigate how combining CNN feature extraction with TMLA has the potential to serve as a viable alternative capable of achieving accuracy levels comparable to state-of-the-art approaches and methodologies.
2. Related Work
In 2020, the study entitled
Comparative Study of Machine Learning Algorithms for Breast Cancer Prediction [
2] compared the performance of various ML algorithms for breast cancer prognosis. The authors quantitatively evaluated decision tree (DT) and logistic regression (LR) classifiers, concluding that ML-based early detection plays a vital role in improving clinical outcomes. The models achieved high accuracies of 94.4% (LR) and 95.1% (DT), highlighting that algorithmic selection strongly influences predictive performance even when applied to the same dataset. During the same year, in
Conventional Machine Learning and Deep Learning Approach for Multi-Classification of Breast Cancer Histopathology Images [
3], the authors addressed the automation of multiclass breast cancer diagnosis using the BreakHis image set. The study compared conventional feature-based classifiers with transfer learning models (VGG16, VGG19, and ResNet50). Results indicated that pretrained networks as feature extractors significantly outperformed handcrafted feature approaches. The hybrid combination of the VGG16 network with a linear SVM achieved the best performance, reaching 91.79% accuracy at 400× magnification.
In 2021, the study entitled
Malignant and Benign Breast Cancer Classification using Machine Learning Algorithms [
4] focused on early detection of benign and malignant tumors, using the
Wisconsin breast cancer dataset. Several classifiers were tested, including SVM, LR, KNN, DT, NB, and RF. RF and SVM demonstrated superior results, both achieving accuracies around 96.5%, supporting their potential for integration into automated diagnostic systems. In 2022, the authors of
An Improvised Random Forest Model for Breast Cancer Classification [
5] proposed a cost-sensitive enhancement to the standard RF algorithm. The model incorporated a penalty matrix assigning higher costs to false negatives, thereby improving minority class detection. This adjustment yielded a notable accuracy of 97.51%, outperforming conventional RF models and reinforcing the algorithm’s adaptability to clinical data imbalance.
More recently, the 2023 study entitled
Breast cancer diagnosis through knowledge distillation of Swin transformer-based teacher–student models [
6] explored transformer-based networks for histopathological image analysis. The authors proposed a teacher–student framework where a compact learner model was trained via knowledge distillation from a Swin Transformer teacher model. Despite being significantly lighter, the learner achieved 98.71% accuracy, only marginally lower than the teacher’s 98.91%, suggesting its practical value for real-time diagnostic support. Also, in the same year, the authors of
Deep learning- and expert knowledge-based feature extraction and performance evaluation in breast histopathology images [
7] examined how DL-derived and expert-defined features affect classification performance. The study compared CNN-based, VGG16 transfer learning, and knowledge-driven feature extraction, tested across seven classifiers. The knowledge-based approach achieved the highest accuracy of up to 98% when combined with neural networks, RF, and multilayer perceptron classifiers.
By reviewing these works, it becomes evident that ML and DL models substantially contribute to breast cancer diagnosis. However, differences in accuracy, computational demand, and interpretability highlight the need for hybrid frameworks that combine the representational strength of deep networks with the efficiency and explainability of TMLA.
3. Proposed Methodology
The overall workflow of the proposed methodology is illustrated in
Figure 1. The process begins with data augmentation, where oversampling techniques and digital image processing (DIP) operations are applied to expand the dataset and mitigate errors inherent to limited data, characterized by low representativeness, low diversity, and class imbalance [
8].
The augmented image set is then fed into a VGG16 CNN configured as a feature extractor to obtain high-level representations of the input data. These extracted features are subsequently used to train four TMLAs: RF, SVM, NB, and KNN. Finally, k-fold cross-validation is performed on all trained models to evaluate performance metrics, including accuracy (Acc), precision (P), recall (R), specificity (Sp), and F1-score (F1). The hybrid model achieving the highest overall performance is selected for its potential in the breast cancer classification task.
3.1. Implementation Steps
The methodology was implemented in four main stages, as described below.
3.1.1. Data Augmentation
The BreakHis image set consists of 7909 histopathological biopsy images collected from 82 patients by P&D Laboratory in Brazil in 2014 [
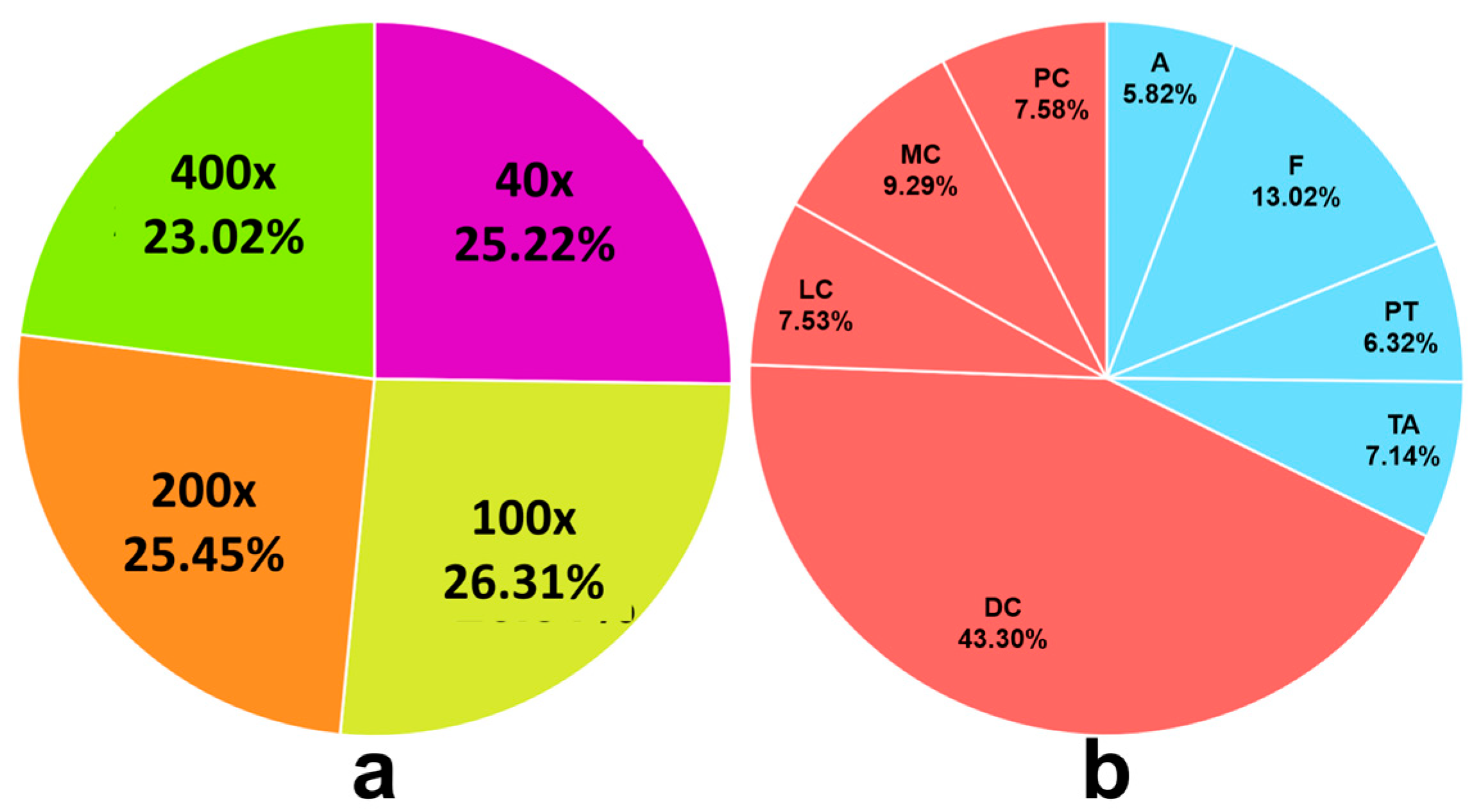
9]. These images are organized into four magnifications (40×, 100×, 200×, and 400×), as shown in
Figure 2a. Each magnification level contains an unbalanced distribution of eight BCS. For the BreakHis
400× subset, there are 1820 images distributed across the following categories (
Figure 2b): Adenosis (A), Fibroadenoma (F), Phyllodes tumor (PT), Tubular adenoma (TA), Ductal carcinoma (DC), Lobular carcinoma (LC), Mucinous carcinoma (MC), and Papillary carcinoma (PC). As a first approximation, each image in the BreakHis dataset is treated as an independent sample, omitting any existing correlations among images.
In ML model training, a limited dataset can lead to overfitting, poor generalization, and inaccurate predictions [
10], since the model does not have enough information to learn representative patterns. This constraint is particularly critical in exploratory research, where insufficient training data can compromise model performance.
To mitigate data scarcity, data augmentation techniques [
11] are applied to the BreakHis
400× subset. Synthetic samples were generated using DIP operations without altering the integrity or biological relevance of the original images. The following transformations were applied: rotation of an image, mirroring an image, and contrast limited adaptive histogram equalization (CLAHE). Applied sequentially, they generate eight images shown in
Figure 3.
In this way, synthetic images are generated without empty pixels or null values that complicate classification, while increasing the number of images and allowing analysis from different angles and perspectives without affecting image quality or content.
3.1.2. VGG16 Feature Extractor
A feature extractor [
13] is an algorithm that identifies and encodes relevant visual patterns such as shape, texture, and color into numerical feature vectors. A popular method for feature extraction is the use of autoencoders, which are deep neural networks capable of learning compressed representations (encodings) of the input data. Autoencoders can reduce data dimensionality, perform noise reduction, and extract salient features that may improve downstream classification performance. However, they may also lead to information loss due to forced compression in the latent space, and they often suffer from training instabilities such as vanishing gradients, which can affect convergence and overall performance. For multiclass tasks, autoencoders focus on learning latent representations and reconstruction, rather than optimizing directly for classification objectives, which may reduce their accuracy and effectiveness.
In contrast, specialized architectures such as VGG16 are explicitly designed for efficient feature extraction, providing richer hierarchical and discriminative representations. This structure supports robust and clinically relevant recognition of complex breast tumor samples.
The VGG16 network, a pretrained CNN, was employed as a feature extractor. This architecture consists of 16 layers in total: 13 convolutional layers and 3 fully connected layers. The convolutional layers are organized into five blocks, each containing multiple 3 × 3 convolutional filters, interleaved with pooling layers to progressively reduce spatial dimensionality.
To analyze the potential of TMLM integrated into hybrid systems, VGG16 was used exclusively as the sole feature extractor. This choice is based on its well-established ability to extract high-quality features and serves as a starting point for the present study, allowing us to establish an initial foundation upon which future methodological improvements can be developed and evaluated.
To use VGG16 as a feature extractor, the convolutional layers are frozen, meaning that their weights remain unchanged during the feature extraction process. Feature extraction is performed using the final output of the convolutional blocks, just before the fully connected layers. This produces a convolutional feature map with dimensions 7 × 7 × 512, assuming the input is an RGB image of size 224 × 224 pixels. A global average pooling or global max pooling strategy is then applied to transform the 7 × 7 × 512 tensor into a 512-dimensional vector, resulting in 25,088 high-quality attributes per image. These vectors capture essential spatial information and serve as inputs for subsequent TMLA models, leveraging transfer learning to improve efficiency and performance (
Figure 4).
When training hybrid models, it is possible that they inherit certain disadvantages from the CNN-based feature extractors, such as the need for large amounts of labeled data, long training periods, and substantial hardware and energy requirements.
However, these limitations are offset by the ability of CNN feature extractors to generate high-level representations that capture complex details and spatial hierarchies in histopathological micrographs, thereby improving the quality of model training and enhancing generalization ability.
Although CNNs typically require extensive data and computational resources for training, the proposed methodology leverages their pretrained capabilities to extract high-quality features that strengthen the predictive power of TMLM classifiers. This hybrid approach combines the best of both worlds: powerful automated feature extraction with faster and more cost-efficient classification. Therefore, despite the computational demands and data requirements, this hybrid strategy improves model accuracy and allows us to examine the potential of hybrid models for multiclass breast cancer classification.
3.1.3. ML Models
This study compared four hybrid multiclassifier models that combine the VGG16 feature extractor with TMLA methods—specifically, RF, SVM, KNN, and NB—to determine the most potentially effective hybrid model for the BCS. Although TMLAs are less complex than DL architectures, integrating them with CNN-based feature extraction can yield comparable results in settings where data availability and computational resources are limiting factors.
Since traditional classifiers cannot process raw images directly, the VGG16 converts images into a structured feature vector. These vectors are compiled into a dataset to train the four TMLA models described below.
RF: An ensemble algorithm that combines multiple decision trees to enhance predictive accuracy and reduce overfitting [
15]. For a dataset with
N samples and
M features, bootstrap sampling generates
B subsets, each used to train a decision tree. For each bootstrap set
b, a decision tree
is built.
Mathematically, the prediction of an RF with
B trees can be expressed as
SVM: is a supervised maximum margin model [
16], which makes it resistant to noise and misclassified data. It is used to analyze simple or high-dimensional data and to solve classifications. This algorithm involves finding the hyperplane that best separates two different classes of data points and maximizes the margin between them. The margin is defined as the maximum width of the region parallel to the hyperplane that contains no interior data points.
Given a training dataset , where are feature vectors and are class labels, the objective of SVMs is to find the hyperplane that maximizes the margin between the two classes.
Mathematically, the maximal margin classifier principle behind SVM is formulated as a quadratic optimization problem that minimizes the norm of the normal vector to the hyperplane
subject to the constraint that all data points lie on the correct side of the hyperplane with a minimum margin of 1.
where
w is a normal vector to the hyperplane,
b is the independent term of the hyperplane, and
is the Euclidean norm of
w, which is minimized to maximize margin.
KNN: This model is based on the mathematical principle of assigning a new instance the most frequent class among its k nearest neighbors in the feature space [
17].
Given a training dataset with instances
, where
is a vector in
, and
is the class label, the distance
between a new point
and each training point is computed as a distance metric, such as the Euclidean distance. The
k training points with the smallest distances
are selected. The predicted class for
is then determined by the most frequent class among these
k neighbors:
where
is the set of indices corresponding to the
k nearest neighbors of
, and 1(⋅) is the indicator function.
NB: this ML algorithm is a probabilistic classifier that utilizes Bayes’ theorem for prediction [
18]. It operates under the assumption that predictor variables are mutually independent, significantly simplifying the required calculations. Given a set of predictor variables
and a class variable
C, NB computes the posterior probability
P(C|X) using the formula.
where
P(
C) is the prior probability of the class,
is the conditional probability of each predictor variable given the class, and
P(X) is the probability of the predictor variables.
3.1.4. Model Training and Performance
Supervised ML models automatically adjust their internal parameters during the training process to optimize their performance in BCS classification, learning directly from the data. However, to ensure reproducibility and to maintain control over the learning process, training hyperparameters must be defined in advance. These hyperparameters are external configurations set prior to model training and govern key aspects of the learning process.
Table 1 presents the training hyperparameters for the RF, SVM, KNN, and NB models. In addition to listing the primary hyperparameters for each model, the table includes the specific values used in this study as well as suggested search ranges for future reference.
The primary goal is to individually evaluate the performance of the proposed hybrid models and to identify the most suitable model for BCS multiclass classification using the Acc, R, Sp, P, and F1 metrics, which are standard and deterministic criteria for assessing ML model performance. A final ensemble of all models is not necessarily required, as the comparative analysis aims to determine the optimal model.
The train–test split is one of the most fundamental techniques for training and validating ML models. It divides the dataset into two subsets: the training set and the validation set, typically allocating 70–80% of the data for training and 20–30% for validation.
However, because this method relies on a single random data partition, it may produce a less reliable estimate of model performance and be influenced by sampling variability. In some cases, the selected partition may not accurately represent the true distribution of the data, potentially biasing the results and leading to misleading conclusions.
K-fold cross-validation is a more rigorous and reliable technique for training and validating ML models compared to the simple train–test split. In this approach, the dataset is systematically divided into
k equally sized subsets (folds). The model is iteratively trained on
k–1 fold and validated on the remaining fold, ensuring that every sample is used for both training and validation across the
k iterations (
Figure 5).
This process provides a more stable and unbiased estimate of model performance by maximizing data utilization and mitigating the randomness inherent in single-split validation. As a result, k-fold cross-validation offers a robust and comprehensive assessment that effectively compensates for the limitations of the traditional train–test split.
Furthermore, k-fold cross-validation is particularly effective for comparative model analysis, as it evaluates all models under identical data partitions and experimental conditions. This consistency ensures a fair and reliable comparison of performance results across different classifiers.
During each training and validation iteration, a confusion matrix [
19] is generated to summarize the model’s correct and incorrect predictions for each class. The matrix is organized into four categories that describe classification outcomes:
True Positives (TP): Malignant cases correctly identified as malignant.
False Positives (FP): Benign cases incorrectly identified as malignant.
True Negatives (TN): Benign cases correctly identified as benign.
False Negatives (FN): Malignant cases incorrectly identified as benign
Evaluation metrics
The elements of the confusion matrix (TP, FP, TN, and FN) are used to compute several quantitative measures that assess the accuracy and effectiveness of a classification model. In this case, because the task involves an eight-category multiclass classification, the resulting confusion matrix has dimensions of 8 × 8.
These metrics, Acc, P, R, Sp, and F1, evaluate a model’s performance in terms of its ability to generalize and avoid overfitting to training data patterns.
Micro Acc represents the proportion of correctly predicted instances among all predictions. It provides an overall measure of model performance, and for multiclass classification, it is calculated as follows:
Micro P measures the proportion of
TP predictions among all instances predicted as positive. In other words, it quantifies how many of the samples classified as positive are actually positive.
Micro R measures the model’s ability to correctly identify positive cases. It represents the proportion of actual positive instances that are correctly detected by the model. Mathematically, it is expressed as follows:
Micro Sp measures the model’s ability to correctly identify negative cases. It represents the proportion of actual negative instances that are correctly classified as negative by the model. Mathematically, it is defined as follows:
Micro F1 represents the harmonic mean of micro
P and micro
R, providing a single metric that balances both. It evaluates the overall quality of a classification model by considering the impact of both false positives and false negatives. Mathematically, it is expressed as follows:
These evaluation metrics are fundamental for assessing the effectiveness and performance of ML models in predictive tasks.
3.2. Performance Considerations
1: For each hybrid model, two sets of evaluation tables are presented: the first corresponds to the performance obtained with the original image set, and the second shows the metrics derived from the synthetically augmented image set. This comparison highlights the impact of data augmentation on the hybrid’s training performance.
2: The comparison between models trained on the original and augmented datasets is not entirely equivalent because the validation subset of the augmented dataset is larger. This discrepancy affects the absolute counts of TP, TN, FP, and FN, potentially introducing bias in the performance evaluation metrics.
For this reason, the evaluation of model performance was extended to include a testing stage using an independent image set (BreakHis Version 2). This image set is distinct from the BreakHis 400× image set for training but was documented following the same acquisition and annotation methodology. The dataset is distributed as follows: A—30 images, F—74 images, PT—39 images, TA—33 images, DC—248 images, LC—27 images, MC—50 images, and PC—44 images, for a total of 545 images. Each image in this dataset is treated as an independent sample, omitting the patient-wise split and any correlations that may exist between images.
To ensure a fair comparison, the metrics obtained during the testing stage are used to determine the best-performing hybrid model, while those from the validation stage serve as internal references for assessing model behavior during training.
3: In accordance with the above considerations, each results table is organized into two sections: the first section reports the validation metrics (highlighted in blue), while the second section presents the testing metrics (highlighted in green). This structure provides a clear distinction between internal model performance and external generalization capability.
4. Results
As described in the methodology section, the values of TP, FP, TN, and FN were quantified to calculate the evaluation metrics for each trained model. Two sets of performance tables were constructed for each hybrid classifier: one based on the original image set and another using the synthetically augmented dataset. In addition to the tables, confusion matrices, and ROC-AUC curve plots are provided, which complement and strengthen the evaluation of the hybrid model by offering both a visual and quantitative measure of its ability to distinguish among the eight BCS classes.
5. Discussion
The RF model (
Table 2 and
Table 3 and
Figure 6,
Figure 7,
Figure 8 and
Figure 9) showed that data augmentation produced a moderate but consistent improvement in performance. While the validation
Acc remained around 0.51, the testing
Acc increased to 0.94, demonstrating that the inclusion of synthetic data enhanced the model’s generalization ability.
For the SVM models (
Table 4 and
Table 5 and
Figure 10,
Figure 11,
Figure 12 and
Figure 13), the improvement was more pronounced. After training with augmented data, the average
Acc rose from 0.82 to 0.91 and the
F1 from 0.77 to 0.89. These results confirm that the hybrid integration of VGG16 feature extraction with SVM enables superior discrimination of BCS compared to models trained exclusively on the original dataset.
In contrast, the NB-based model (
Table 6 and
Table 7 and
Figure 14,
Figure 15,
Figure 16 and
Figure 17) experienced a decline in
Acc and
F1 when trained with synthetic data. This suggests that the proposed methodology is not well suited for hybrid frameworks relying on NB, likely due to its strong independence assumptions and reduced adaptability to high-dimensional features.
The KNN-based hybrid model (
Table 8 and
Table 9 and
Figure 18,
Figure 19,
Figure 20 and
Figure 21) exhibited the most significant improvement. Training with augmented data increased
Acc from 0.92 to 0.97 and
F1 from 0.90 to 0.96, demonstrating excellent stability and generalization. These findings position the KNN-based hybrid model as the most promising approach for multi-class BCS classification.
Table 10 and
Table 11 summarize the average, SD, and CI
95 values for all models, clearly showing that the KNN hybrid model trained with synthetically augmented data consistently outperformed the others across all evaluation metrics. In turn,
Table 12 presents the mean values, SD, and CI
95 of the ROC-AUC scores for each hybrid model, confirming the generalization capability of each model in relation to the calculated metrics.
Overall, these findings demonstrate that hybrid systems combining feature extraction with classical classifiers have the potential to achieve high performance, while also revealing the possibility of data leakage and the need for a feature-selection process to reduce the dimensionality of the attributes extracted by VGG16. Together, these insights enable specification of a rigorous experimental design that incorporates selective sampling, which groups images from the same patient within the same fold of cross-validation to ensure better control over training data and to prevent data leakage that would inflate the performance metrics of ML models.
6. Conclusions
This study presented a methodology for breast cancer classification using hybrid models that combine VGG16 feature extraction with traditional ML classifiers. The proposed approach demonstrated that high performance can be achieved in terms of Acc, P, R, Sp, and F1, highlighting the viability of hybrid models for BCS classification.
Overall, the RF and SVM hybrids achieved computationally satisfactory levels of accuracy; however, these levels remain insufficient from a clinical standpoint. Therefore, further optimization and methodological refinement are needed to enhance their applicability to multi-class classification of BCS.
In contrast, the NB-hybrid model showed limited adaptability and failed to achieve significant performance gains, reflecting its inherent limitations when applied to high-dimensional data. This restricts its suitability for contexts where precision and reliability are paramount.
The KNN-hybrid model, on the other hand, demonstrated the most consistent and accurate performance, reaching an average Acc of 0.97. These results underscore the potential of hybrid architectures integrating TMLA-based systems, demonstrating that the hybrid approach is a viable alternative.
One area for improvement is addressing a key limitation in the proposed methodology related to potential information leakage during the training of hybrid models, which poses a critical risk to the model’s reliability and practical utility. Possible causes include insufficient rigor and oversight in the construction of the training set, as well as artificially inflated model performance resulting from placing highly correlated histopathological micrographs in both the training and validation sets. This issue may arise from the incorporation of synthetic data during the preprocessing stage, leading to performance estimates that do not accurately reflect the model’s true predictive capacity in real-world scenarios.
Other disadvantages of the proposed methodology include the limited generalization of the model due to working exclusively with 400× magnification micrographs and the reliance on a single feature extractor (VGG16), which could potentially lead to model overfitting due to its pretrained biases.
In future work, efforts to mitigate information leakage in the training of hybrid models will focus on implementing rigorous safeguards to ensure data integrity and security without compromising the scientific validity of the model. In addition, a feature-selection process will be implemented to reduce data dimensionality and improve the generalization capability of the hybrid models, along with a statistical analysis that supports the robustness of the experimental design.
Author Contributions
F.J.R.-P.: Conceptualization, methodology, software, validation, formal analysis, investigation, resources, data curation, writing—review and editing, visualization, supervision, project; J.R.C.-S.: Conceptualization, validation, formal analysis, writing—review and editing, project administration; A.M.-P.: Validation, investigation, formal analysis, writing—review and editing; F.M.-B.: Conceptualization, methodology, formal analysis, writing—review and editing; J.C.R.-F.: Conceptualization, software, writing—review and editing; J.L.-M.: Conceptualization, validation, investigation, writing—review and editing; G.V.-H.: Conceptualization, validation, investigation, writing—review and editing; B.E.J.-L.: Conceptualization, validation, investigation, writing—review and editing; J.M.X.-P.: Conceptualization, software, writing—review and editing; E.G.P.-P.: Conceptualization, methodology, software, formal analysis, data curation, writing—original draft, writing—review and editing, supervision, project administration. All authors have read and agreed to the published version of the manuscript.
Funding
The first author received a grant from the Consejo Nacional de Humanidades, Ciencia y Tecnología (CONACYT), CVU: 1078268.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The BreakHis
Version 2 breast cancer histopathology micrograph set is publicly available at the following URL:
https://web.inf.ufpr.br/vri/databases/breast-cancer-histopathological-database-breakhis (accessed on 9 January 2025) or
https://www.kaggle.com/datasets/ambarish/breakhis (accessed on 9 January 2025). This dataset contains 7909 microscopic images of breast tumor tissues, classified as benign and malignant, obtained from 82 patients with different magnification factors (40×, 100×, 200×, and 400×). This dataset is widely used for classification tasks in ML and allows for comparative evaluation of models on histopathological images of breast cancer. The BreakHis breast cancer histopathology micrograph set is publicly available at the following URL:
https://www.kaggle.com/datasets/forderation/breakhis-400x (accessed on 9 January 2025). This dataset contains 545 microscopic images of breast tumor tissues, classified as benign and malignant, with a 400× magnification factor. The dataset is distributed as follows: The dataset is distributed as follows: A—30 images, F—74 images, PT—39 images, TA—33 images, DC—248 images, LC—27 images, MC—50 images, and PC—44 images. The purpose of this dataset is to serve as a resource for deep learning training and for the training or testing of machine learning models.
Acknowledgments
The first author received a grant from the Consejo Nacional de Humanidades, Ciencia y Tecnología (CONACYT), for which we are grateful for the support provided by this institution. The authors have reviewed and edited the output and take full responsibility for the content of this publication.
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| TMLA | Traditional Machine Learning Algorithms |
| BCS | Breast Cancer Subtypes |
| RF | Random Forest |
| SVM | Support Vector Machine |
| KNN | K-Nearest Neighbors |
| NB | Naive Bayes |
| AI | Artificial Intelligence |
| DT | Decision Tree |
| LR | Logistic Regression |
| DL | Deep Learning |
| CNN | Convolutional Neural Network |
| ML | Machine Learning |
| DIP | Digital Image Processing |
| Acc | Accuracy |
| P | Precision |
| R | Recall |
| Sp | Specificity |
| F1 | F1-Score |
| CLAHE | Contrast Limited Adaptive Histogram Equalization |
| TP | True Positive |
| FP | False Positive |
| TN | True Negative |
| FN | False Negative |
| SD | Standard deviation |
| CI95 | 95% Confidence Interval |
| ROC-AUC | Receiver Operating Characteristic–Area Under the Curve |
References
- Bray, F.; Laversanne, M.; Sung, H.; Ferlay, J.; Siegel, R.L.; Soerjomataram, I.; Jemal, A. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA A Cancer J. Clin. 2024, 74, 229–263. [Google Scholar] [CrossRef] [PubMed]
- Sengar, P.P.; Gaikwad, M.J.; Nagdive, A.S. Comparative study of machine learning algorithms for breast cancer prediction. In Proceedings of the 2020 Third International Conference on Smart Systems and Inventive Technology (ICSSIT), Tirunelveli, India, 20–22 August 2020; IEEE: New York, NY, USA, 2020; pp. 796–801. [Google Scholar]
- Sharma, S.; Mehra, R. Conventional machine learning and deep learning approach for multi-classification of breast cancer histopathology images—A comparative insight. J. Digit. Imaging 2020, 33, 632–654. [Google Scholar] [CrossRef] [PubMed]
- Ara, S.; Das, A.; Dey, A. Malignant and benign breast cancer classification using machine learning algorithms. In Proceedings of the 2021 International Conference on Artificial Intelligence (ICAI), Lucknow, India, 22–23 May 2021; IEEE: New York, NY, USA, 2021; pp. 97–101. [Google Scholar]
- Mathew, T.E. An improvised random forest model for breast cancer classification. NeuroQuantology 2022, 20, 713. [Google Scholar] [CrossRef]
- Kolla, B.; Venugopal, P. Breast cancer diagnosis through knowledge distillation of Swin transformer-based teacher–student models. Mach. Learn. Sci. Technol. 2023, 4, 045047. [Google Scholar] [CrossRef]
- Kode, H.; Barkana, B.D. Deep learning- and expert knowledge-based feature extraction and performance evaluation in breast histopathology images. Cancers 2023, 15, 3075. [Google Scholar] [CrossRef] [PubMed]
- Ebrahimy, H.; Mirbagheri, B.; Matkan, A.A.; Azadbakht, M. Effectiveness of the integration of data balancing techniques and tree-based ensemble machine learning algorithms for spatially-explicit land cover accuracy prediction. Remote Sens. Appl. Soc. Environ. 2022, 27, 100785. [Google Scholar] [CrossRef]
- Spanhol, F.A.; Oliveira, L.S.; Petitjean, C.; Heutte, L. A dataset for breast cancer histopathological image classification. IEEE Trans. Biomed. Eng. 2015, 63, 1455–1462. [Google Scholar] [CrossRef] [PubMed]
- Barkah, A.S.; Selamat, S.R.; Abidin, Z.Z.; Wahyudi, R. Impact of data balancing and feature selection on machine learning-based network intrusion detection. JOIV Int. J. Inform. Vis. 2023, 7, 241–248. [Google Scholar] [CrossRef]
- Tarawneh, A.S.; Hassanat, A.B.; Altarawneh, G.A.; Almuhaimeed, A. Stop oversampling for class imbalance learning: A review. IEEE Access 2022, 10, 47643–47660. [Google Scholar] [CrossRef]
- Krawczyk, B.; Jeleń, Ł.; Krzyżak, A.; Fevens, T. Oversampling methods for classification of imbalanced breast cancer malignancy data. In Proceedings of the Computer Vision and Graphics: International Conference, ICCVG 2012, Warsaw, Poland, 24–26 September 2012; Springer: Berlin/Heidelberg, Germany, 2012; pp. 483–490. [Google Scholar]
- Dara, S.; Tumma, P. Feature extraction by using deep learning: A survey. In Proceedings of the 2018 Second International Conference on Electronics, Communication and Aerospace Technology (ICECA), Coimbatore, India, 29–31 March 2018; IEEE: New York, NY, USA, 2018; pp. 1795–1801. [Google Scholar]
- Bakasa, W.; Viriri, S. Vgg16 feature extractor with extreme gradient boost classifier for pancreas cancer prediction. J. Imaging 2023, 9, 138. [Google Scholar] [CrossRef] [PubMed]
- Salman, H.A.; Kalakech, A.; Steiti, A. Random forest algorithm overview. Babylon. J. Mach. Learn. 2024, 2024, 69–79. [Google Scholar] [CrossRef] [PubMed]
- Guido, R.; Ferrisi, S.; Lofaro, D.; Conforti, D. An overview on the advancements of support vector machine models in healthcare applications: A review. Information 2024, 15, 235. [Google Scholar] [CrossRef]
- Kramer, O. K-nearest neighbors. In Dimensionality Reduction with Unsupervised Nearest Neighbors; Springer: Berlin/Heidelberg, Germany, 2013; pp. 13–23. [Google Scholar]
- Wickramasinghe, I.; Kalutarage, H. Naive Bayes: Applications, variations and vulnerabilities: A review of literature with code snippets for implementation. Soft Comput. 2021, 25, 2277–2293. [Google Scholar] [CrossRef]
- Liang, J. Confusion matrix: Machine learning. POGIL Act. Clgh. 2022, 3, 4. [Google Scholar]
Figure 1.
Proposed training methodology diagram for hybrid ML models.
Figure 1.
Proposed training methodology diagram for hybrid ML models.
Figure 2.
Distribution of images in the BreakHis training set: (a) BreakHis image distribution per magnification level; (b) BreakHis 400×—BCS distribution.
Figure 2.
Distribution of images in the BreakHis training set: (a) BreakHis image distribution per magnification level; (b) BreakHis 400×—BCS distribution.
Figure 3.
Oversampling process [
12]: (
a) original image; (
b) original image with rotation of 180° (in range −360° to 360°); (
c) original image with mirror effect (horizontal axis); (
d) original image with rotation of 180° and horizontal mirror effect; (
e) CLAHE image (with clipLimit = 2.0 and tileGridSize = (8,8)); (
f) CLAHE image with rotation of 180°; (
g) CLAHE image with horizontal mirror effect; (
h) CLAHE image with rotation of 180° and horizontal mirror effect.
Figure 3.
Oversampling process [
12]: (
a) original image; (
b) original image with rotation of 180° (in range −360° to 360°); (
c) original image with mirror effect (horizontal axis); (
d) original image with rotation of 180° and horizontal mirror effect; (
e) CLAHE image (with clipLimit = 2.0 and tileGridSize = (8,8)); (
f) CLAHE image with rotation of 180°; (
g) CLAHE image with horizontal mirror effect; (
h) CLAHE image with rotation of 180° and horizontal mirror effect.
Figure 4.
The VGG16 CNN [
14] is modified into a feature extractor by removing its fully connected layers and its last classification layer (red box). Instead, an output of an intermediate layer is taken (green box), which represents a vector of high-resolution features. The vectors are then stacked to create a dataset to train TMLM.
Figure 4.
The VGG16 CNN [
14] is modified into a feature extractor by removing its fully connected layers and its last classification layer (red box). Instead, an output of an intermediate layer is taken (green box), which represents a vector of high-resolution features. The vectors are then stacked to create a dataset to train TMLM.
Figure 5.
Schematic illustration of k-fold cross-validation with K = 10 (folds = 10). The dataset is divided into 10 subsets, each representing 10% of the total training set. In each cycle, the training subsets (shown in blue) and the validation subset (shown in red) is alternated, such that 90 percent of the data are used to train the model and the remaining 10 percent are used to validate the same model.
Figure 5.
Schematic illustration of k-fold cross-validation with K = 10 (folds = 10). The dataset is divided into 10 subsets, each representing 10% of the total training set. In each cycle, the training subsets (shown in blue) and the validation subset (shown in red) is alternated, such that 90 percent of the data are used to train the model and the remaining 10 percent are used to validate the same model.
Figure 6.
Confusion matrices of the RF-based hybrid model trained with the original image set (TP indicated in red): (a) confusion matrix for the validation stage (182 images); (b) confusion matrix for the test stage (545 images).
Figure 6.
Confusion matrices of the RF-based hybrid model trained with the original image set (TP indicated in red): (a) confusion matrix for the validation stage (182 images); (b) confusion matrix for the test stage (545 images).
Figure 7.
ROC-AUC curves of the RF model trained with the original dataset (reference line equal to 0.5 in black): (a) ROC-AUC curve of the validation stage; (b) ROC-AUC curve of the test stage.
Figure 7.
ROC-AUC curves of the RF model trained with the original dataset (reference line equal to 0.5 in black): (a) ROC-AUC curve of the validation stage; (b) ROC-AUC curve of the test stage.
Figure 8.
Confusion matrices of the RF-based hybrid model trained with the synthetically augmented image set (TP indicated in red): (a) confusion matrix for the validation stage (1456 images); (b) confusion matrix for the test stage (545 images).
Figure 8.
Confusion matrices of the RF-based hybrid model trained with the synthetically augmented image set (TP indicated in red): (a) confusion matrix for the validation stage (1456 images); (b) confusion matrix for the test stage (545 images).
Figure 9.
ROC-AUC curves of the RF model trained with the synthetically augmented dataset (reference line equal to 0.5 in black): (a) ROC-AUC curve of the validation stage; (b) ROC-AUC curve of the test stage.
Figure 9.
ROC-AUC curves of the RF model trained with the synthetically augmented dataset (reference line equal to 0.5 in black): (a) ROC-AUC curve of the validation stage; (b) ROC-AUC curve of the test stage.
Figure 10.
Confusion matrices of the SVM-based hybrid model trained with the original image set (TP indicated in yellow): (a) confusion matrix for the validation stage (182 images); (b) confusion matrix for the test stage (545 images).
Figure 10.
Confusion matrices of the SVM-based hybrid model trained with the original image set (TP indicated in yellow): (a) confusion matrix for the validation stage (182 images); (b) confusion matrix for the test stage (545 images).
Figure 11.
ROC-AUC curves of the SVM model trained with the original dataset (reference line equal to 0.5 in black): (a) ROC-AUC curve of the validation stage; (b) ROC-AUC curve of the test stage.
Figure 11.
ROC-AUC curves of the SVM model trained with the original dataset (reference line equal to 0.5 in black): (a) ROC-AUC curve of the validation stage; (b) ROC-AUC curve of the test stage.
Figure 12.
Confusion matrices of the SVM-based hybrid model trained with the synthetically augmented image set (TP indicated in yellow): (a) confusion matrix for the validation stage (1456 images); (b) confusion matrix for the test stage (545 images).
Figure 12.
Confusion matrices of the SVM-based hybrid model trained with the synthetically augmented image set (TP indicated in yellow): (a) confusion matrix for the validation stage (1456 images); (b) confusion matrix for the test stage (545 images).
Figure 13.
ROC-AUC curves of the SVM model trained with the synthetically augmented dataset (reference line equal to 0.5 in black): (a) ROC-AUC curve of the validation stage; (b) ROC-AUC curve of the test stage.
Figure 13.
ROC-AUC curves of the SVM model trained with the synthetically augmented dataset (reference line equal to 0.5 in black): (a) ROC-AUC curve of the validation stage; (b) ROC-AUC curve of the test stage.
Figure 14.
Confusion matrices of the NB-based hybrid model trained with the original image set (TP indicated in purple): (a) confusion matrix for the validation stage (182 images); (b) confusion matrix for the test stage (545 images).
Figure 14.
Confusion matrices of the NB-based hybrid model trained with the original image set (TP indicated in purple): (a) confusion matrix for the validation stage (182 images); (b) confusion matrix for the test stage (545 images).
Figure 15.
ROC-AUC curves of the NB model trained with the original dataset (reference line equal to 0.5 in black): (a) ROC-AUC curve of the validation stage; (b) ROC-AUC curve of the test stage.
Figure 15.
ROC-AUC curves of the NB model trained with the original dataset (reference line equal to 0.5 in black): (a) ROC-AUC curve of the validation stage; (b) ROC-AUC curve of the test stage.
Figure 16.
Confusion matrices of the NB-based hybrid model trained with the synthetically augmented image set (TP indicated in purple): (a) confusion matrix for the validation stage (1456 images); (b) confusion matrix for the test stage (545 images).
Figure 16.
Confusion matrices of the NB-based hybrid model trained with the synthetically augmented image set (TP indicated in purple): (a) confusion matrix for the validation stage (1456 images); (b) confusion matrix for the test stage (545 images).
Figure 17.
ROC-AUC curves of the NB model trained with the synthetically augmented dataset (reference line equal to 0.5 in black): (a) ROC-AUC curve of the validation stage; (b) ROC-AUC curve of the test stage.
Figure 17.
ROC-AUC curves of the NB model trained with the synthetically augmented dataset (reference line equal to 0.5 in black): (a) ROC-AUC curve of the validation stage; (b) ROC-AUC curve of the test stage.
Figure 18.
Confusion matrices of the KNN-based hybrid model trained with the original image set (TP indicated in orange): (a) confusion matrix for the validation stage (182 images); (b) confusion matrix for the test stage (545 images).
Figure 18.
Confusion matrices of the KNN-based hybrid model trained with the original image set (TP indicated in orange): (a) confusion matrix for the validation stage (182 images); (b) confusion matrix for the test stage (545 images).
Figure 19.
ROC-AUC curves of the KNN model trained with the original dataset (reference line equal to 0.5 in black): (a) ROC-AUC curve of the validation stage; (b) ROC-AUC curve of the test stage.
Figure 19.
ROC-AUC curves of the KNN model trained with the original dataset (reference line equal to 0.5 in black): (a) ROC-AUC curve of the validation stage; (b) ROC-AUC curve of the test stage.
Figure 20.
Confusion matrices of the KNN-based hybrid model trained with the synthetically augmented image set (TP indicated in orange): (a) confusion matrix for the validation stage (1456 images); (b) confusion matrix for the test stage (545 images).
Figure 20.
Confusion matrices of the KNN-based hybrid model trained with the synthetically augmented image set (TP indicated in orange): (a) confusion matrix for the validation stage (1456 images); (b) confusion matrix for the test stage (545 images).
Figure 21.
ROC-AUC curves of the KNN model trained with the synthetically augmented dataset (reference line equal to 0.5 in black); (a) ROC-AUC curve of the validation stage; (b) ROC-AUC curve of the test stage.
Figure 21.
ROC-AUC curves of the KNN model trained with the synthetically augmented dataset (reference line equal to 0.5 in black); (a) ROC-AUC curve of the validation stage; (b) ROC-AUC curve of the test stage.
Table 1.
Hyperparameters and suggested search ranges for the RF, SVM, KNN, and NB models.
Table 1.
Hyperparameters and suggested search ranges for the RF, SVM, KNN, and NB models.
| Model | Main Hyperparameters | Suggested Search Ranges |
|---|
| RF | n_estimators = 100; random_state = 42; max_depth = None; min_samples_split = 2; max_features = 1.0; bootstrap = True | n_estimators: [50, 100, 200, 500]; random_state = Constant value for reproducibility; max_depth: [None, 10, 20, 30]; min_samples_split: [2, 5, 10, 20]; max_features: [‘sqrt’, ‘log2’, None]; bootstrap: [True, False] |
| SVM | kernel = ‘rbf’, probability = True, C = 1.0, gamma = ‘scale’, degree = 3 (only for “poly,” irrelevant in rbf) | Kernel: [‘rbf’, ‘poly’]; probability: True (fixed); C: [0.1, 1.0, 10, 100]; gamma = [‘scale’, ‘auto’, 0.001, 0.01, 0.1]; degree: [3] (fixed) |
| KNN | n_neighbors = 1, metric = “minkowski”, p = 2 | N_neighbors: [1, 3, 5, 7, 9, 15]; metric: [minkowski, euclidean, manhattan]; p: [1,2] (only for minkowski) |
| NB | var_smoothing = 1 × 10−6 | var_smoothing: [1 × 10−9, 1 × 10−8, 1 × 10−7, 1 × 10−6, 1 × 10−5] |
Table 2.
Performance of the RF model trained with the original dataset.
Table 2.
Performance of the RF model trained with the original dataset.
| Model Id | Validation | Test |
|---|
| Acc | P | R | Sp | F1 | Acc | P | R | Sp | F1 |
|---|
| 1 | 0.4341 | 0.1065 | 0.1761 | 0.8875 | 0.1293 | 0.9321 | 0.9671 | 0.8884 | 0.9851 | 0.9242 |
| 2 | 0.4890 | 0.1309 | 0.1913 | 0.8906 | 0.1521 | 0.9284 | 0.9591 | 0.8751 | 0.9847 | 0.9114 |
| 3 | 0.5549 | 0.2687 | 0.2046 | 0.8963 | 0.1794 | 0.9431 | 0.9724 | 0.8988 | 0.9876 | 0.9321 |
| 4 | 0.5330 | 0.1794 | 0.1984 | 0.9000 | 0.1730 | 0.9303 | 0.9660 | 0.8820 | 0.9847 | 0.9201 |
| 5 | 0.4890 | 0.3774 | 0.2063 | 0.8967 | 0.1777 | 0.9413 | 0.9543 | 0.9014 | 0.9884 | 0.9260 |
| 6 | 0.4835 | 0.1474 | 0.2034 | 0.8970 | 0.1630 | 0.9486 | 0.9651 | 0.9125 | 0.9894 | 0.9370 |
| 7 | 0.4615 | 0.1238 | 0.1857 | 0.8938 | 0.1422 | 0.9486 | 0.9667 | 0.9134 | 0.9891 | 0.9378 |
| 8 | 0.5165 | 0.1362 | 0.1656 | 0.8874 | 0.1387 | 0.9450 | 0.9642 | 0.9195 | 0.9887 | 0.9403 |
| 9 | 0.4341 | 0.2318 | 0.1903 | 0.8885 | 0.1498 | 0.9358 | 0.9693 | 0.8901 | 0.9863 | 0.9260 |
| 10 | 0.5220 | 0.1276 | 0.1838 | 0.8915 | 0.1489 | 0.9413 | 0.9672 | 0.9048 | 0.9877 | 0.9333 |
| Mean | 0.4918 | 0.1830 | 0.1906 | 0.8929 | 0.1554 | 0.9395 | 0.9651 | 0.8986 | 0.9872 | 0.9288 |
Table 3.
Performance of the RF model trained with synthetically augmented data.
Table 3.
Performance of the RF model trained with synthetically augmented data.
| Model Id | Validation | Test |
|---|
| Acc | P | R | Sp | F1 | Acc | P | R | Sp | F1 |
|---|
| 1 | 0.5034 | 0.7277 | 0.2279 | 0.8962 | 0.2267 | 0.9376 | 0.9590 | 0.8945 | 0.9869 | 0.9232 |
| 2 | 0.5460 | 0.7661 | 0.2416 | 0.9004 | 0.2488 | 0.9450 | 0.9667 | 0.9094 | 0.9887 | 0.9358 |
| 3 | 0.5041 | 0.7194 | 0.2273 | 0.8955 | 0.2228 | 0.9505 | 0.9789 | 0.9089 | 0.9891 | 0.9398 |
| 4 | 0.5144 | 0.7104 | 0.2237 | 0.8974 | 0.2179 | 0.9248 | 0.9627 | 0.8755 | 0.9836 | 0.9143 |
| 5 | 0.5082 | 0.7793 | 0.2327 | 0.8957 | 0.2368 | 0.9505 | 0.9780 | 0.9074 | 0.9893 | 0.9388 |
| 6 | 0.5275 | 0.7710 | 0.2412 | 0.8982 | 0.2421 | 0.9394 | 0.9635 | 0.8968 | 0.9876 | 0.9274 |
| 7 | 0.5110 | 0.7549 | 0.2256 | 0.8959 | 0.2344 | 0.9394 | 0.9646 | 0.9002 | 0.9870 | 0.9298 |
| 8 | 0.5151 | 0.7673 | 0.2414 | 0.8986 | 0.2493 | 0.9413 | 0.9676 | 0.8944 | 0.9872 | 0.9273 |
| 9 | 0.5446 | 0.7749 | 0.2512 | 0.9006 | 0.2629 | 0.9468 | 0.9710 | 0.9077 | 0.9883 | 0.9367 |
| 10 | 0.5103 | 0.7200 | 0.2484 | 0.8977 | 0.2605 | 0.9486 | 0.9677 | 0.9157 | 0.9891 | 0.9396 |
| Mean | 0.5185 | 0.7491 | 0.2361 | 0.8976 | 0.2402 | 0.9424 | 0.9680 | 0.9011 | 0.9877 | 0.9313 |
Table 4.
Performance of the SVM model trained on the original dataset.
Table 4.
Performance of the SVM model trained on the original dataset.
| Model Id | Validation | Test |
|---|
| Acc | P | R | Sp | F1 | Acc | P | R | Sp | F1 |
|---|
| 1 | 0.4890 | 0.3205 | 0.2359 | 0.9017 | 0.1991 | 0.8128 | 0.9316 | 0.6817 | 0.9607 | 0.7572 |
| 2 | 0.5330 | 0.4400 | 0.2706 | 0.9041 | 0.2576 | 0.8239 | 0.9239 | 0.6993 | 0.9631 | 0.7700 |
| 3 | 0.6044 | 0.5064 | 0.2684 | 0.9140 | 0.2609 | 0.8294 | 0.9357 | 0.7034 | 0.9645 | 0.7778 |
| 4 | 0.5659 | 0.4000 | 0.2362 | 0.9119 | 0.2287 | 0.8202 | 0.9247 | 0.6878 | 0.9619 | 0.7616 |
| 5 | 0.5220 | 0.3161 | 0.2441 | 0.9071 | 0.2099 | 0.8220 | 0.9312 | 0.6944 | 0.9632 | 0.7654 |
| 6 | 0.5549 | 0.5305 | 0.3082 | 0.9153 | 0.2821 | 0.8275 | 0.9210 | 0.7010 | 0.9639 | 0.7720 |
| 7 | 0.5330 | 0.3394 | 0.2512 | 0.9138 | 0.2080 | 0.8330 | 0.9361 | 0.7112 | 0.9653 | 0.7831 |
| 8 | 0.5934 | 0.6330 | 0.2793 | 0.9099 | 0.3039 | 0.8257 | 0.9364 | 0.7074 | 0.9631 | 0.7786 |
| 9 | 0.5110 | 0.4889 | 0.2767 | 0.9076 | 0.2507 | 0.8165 | 0.9305 | 0.6902 | 0.9618 | 0.7696 |
| 10 | 0.5879 | 0.3867 | 0.2473 | 0.9095 | 0.2252 | 0.8257 | 0.9302 | 0.7019 | 0.9636 | 0.7768 |
| Mean | 0.5495 | 0.4362 | 0.2618 | 0.9095 | 0.2426 | 0.8237 | 0.9301 | 0.6978 | 0.9631 | 0.7712 |
Table 5.
Performance of the SVM model trained with synthetically augmented data.
Table 5.
Performance of the SVM model trained with synthetically augmented data.
| Model Id | Validation | Test |
|---|
| Acc | P | R | Sp | F1 | Acc | P | R | Sp | F1 |
|---|
| 1 | 0.8029 | 0.8513 | 0.7228 | 0.9623 | 0.7718 | 0.9174 | 0.9468 | 0.8633 | 0.9833 | 0.8991 |
| 2 | 0.8180 | 0.8659 | 0.7096 | 0.9633 | 0.7700 | 0.9028 | 0.9302 | 0.8444 | 0.9807 | 0.8816 |
| 3 | 0.8015 | 0.8567 | 0.7006 | 0.9619 | 0.7559 | 0.9083 | 0.9394 | 0.8440 | 0.9816 | 0.8841 |
| 4 | 0.8001 | 0.8754 | 0.6970 | 0.9604 | 0.7633 | 0.9119 | 0.9451 | 0.8558 | 0.9819 | 0.8946 |
| 5 | 0.8036 | 0.8676 | 0.7126 | 0.9613 | 0.7708 | 0.9156 | 0.9523 | 0.8574 | 0.9824 | 0.8971 |
| 6 | 0.8036 | 0.8668 | 0.7068 | 0.9606 | 0.7669 | 0.9046 | 0.9350 | 0.8426 | 0.9809 | 0.8811 |
| 7 | 0.7720 | 0.8360 | 0.6610 | 0.9551 | 0.7243 | 0.9083 | 0.9436 | 0.8478 | 0.9811 | 0.8881 |
| 8 | 0.7953 | 0.8529 | 0.6989 | 0.9608 | 0.7576 | 0.9083 | 0.9386 | 0.8441 | 0.9813 | 0.8836 |
| 9 | 0.8235 | 0.8763 | 0.7291 | 0.9643 | 0.7837 | 0.9064 | 0.9368 | 0.8421 | 0.9808 | 0.8806 |
| 10 | 0.7988 | 0.8562 | 0.7150 | 0.9613 | 0.7681 | 0.9119 | 0.9376 | 0.8559 | 0.9824 | 0.8909 |
| Mean | 0.8019 | 0.8605 | 0.7053 | 0.9611 | 0.7632 | 0.9096 | 0.9405 | 0.8497 | 0.9816 | 0.8881 |
Table 6.
Performance of the NB model trained on the original dataset.
Table 6.
Performance of the NB model trained on the original dataset.
| Model Id | Validation | Test |
|---|
| Acc | P | R | Sp | F1 | Acc | P | R | Sp | F1 |
|---|
| 1 | 0.3132 | 0.0948 | 0.1169 | 0.0910 | 0.8694 | 0.8569 | 0.8700 | 0.8643 | 0.8644 | 0.9752 |
| 2 | 0.4121 | 0.2414 | 0.1663 | 0.1553 | 0.8816 | 0.8550 | 0.8686 | 0.8651 | 0.8618 | 0.9754 |
| 3 | 0.4725 | 0.2699 | 0.1940 | 0.1935 | 0.8837 | 0.8752 | 0.8833 | 0.8946 | 0.8849 | 0.9791 |
| 4 | 0.3846 | 0.1873 | 0.1525 | 0.1434 | 0.8803 | 0.8532 | 0.8555 | 0.8710 | 0.8604 | 0.9750 |
| 5 | 0.3791 | 0.1511 | 0.1530 | 0.1342 | 0.8784 | 0.8716 | 0.8762 | 0.8787 | 0.8747 | 0.9780 |
| 6 | 0.3681 | 0.1045 | 0.1353 | 0.1041 | 0.8785 | 0.8679 | 0.8748 | 0.8796 | 0.8725 | 0.9780 |
| 7 | 0.3791 | 0.1812 | 0.1629 | 0.1473 | 0.8820 | 0.8716 | 0.8764 | 0.8940 | 0.8805 | 0.9791 |
| 8 | 0.4176 | 0.0982 | 0.1280 | 0.1066 | 0.8743 | 0.8752 | 0.8847 | 0.8964 | 0.8859 | 0.9791 |
| 9 | 0.3516 | 0.3958 | 0.1589 | 0.1506 | 0.8761 | 0.8624 | 0.8772 | 0.8677 | 0.8686 | 0.9760 |
| 10 | 0.4066 | 0.1320 | 0.1305 | 0.1156 | 0.8722 | 0.8624 | 0.8652 | 0.8749 | 0.8625 | 0.9772 |
| Mean | 0.3885 | 0.1856 | 0.1498 | 0.1342 | 0.8777 | 0.8651 | 0.8732 | 0.8786 | 0.8716 | 0.9772 |
Table 7.
Performance of the NB model trained with synthetically augmented data.
Table 7.
Performance of the NB model trained with synthetically augmented data.
| Model Id | Validation | Test |
|---|
| Acc | P | R | Sp | F1 | Acc | P | R | Sp | F1 |
|---|
| 1 | 0.4251 | 0.4277 | 0.4402 | 0.9116 | 0.4136 | 0.5211 | 0.6466 | 0.6283 | 0.9341 | 0.5549 |
| 2 | 0.4210 | 0.4155 | 0.4219 | 0.9119 | 0.3941 | 0.5138 | 0.6427 | 0.6313 | 0.9333 | 0.5481 |
| 3 | 0.4265 | 0.4403 | 0.4297 | 0.9122 | 0.4088 | 0.5064 | 0.6291 | 0.6173 | 0.9321 | 0.5388 |
| 4 | 0.4190 | 0.4330 | 0.4340 | 0.9109 | 0.4095 | 0.5101 | 0.6255 | 0.6202 | 0.9325 | 0.5428 |
| 5 | 0.4245 | 0.4376 | 0.4509 | 0.9128 | 0.4201 | 0.5174 | 0.6438 | 0.6370 | 0.9333 | 0.5586 |
| 6 | 0.4025 | 0.4204 | 0.4143 | 0.9084 | 0.3901 | 0.5174 | 0.6440 | 0.6348 | 0.9334 | 0.5570 |
| 7 | 0.4093 | 0.4131 | 0.4246 | 0.9099 | 0.3951 | 0.5174 | 0.6291 | 0.6306 | 0.9330 | 0.5500 |
| 8 | 0.4121 | 0.4320 | 0.4278 | 0.9105 | 0.4080 | 0.5028 | 0.6265 | 0.6173 | 0.9318 | 0.5387 |
| 9 | 0.4196 | 0.4209 | 0.4522 | 0.9125 | 0.4118 | 0.5174 | 0.6466 | 0.6238 | 0.9334 | 0.5504 |
| 10 | 0.4588 | 0.4634 | 0.4711 | 0.9178 | 0.4479 | 0.5119 | 0.6304 | 0.6223 | 0.9322 | 0.5473 |
| Mean | 0.4218 | 0.4304 | 0.4367 | 0.9119 | 0.4099 | 0.5136 | 0.6364 | 0.6263 | 0.9329 | 0.5487 |
Table 8.
Performance of the KNN model trained on the original dataset.
Table 8.
Performance of the KNN model trained on the original dataset.
| Model Id | Validation | Test |
|---|
| Acc | P | R | Sp | F1 | Acc | P | R | Sp | F1 |
|---|
| 1 | 0.3791 | 0.2630 | 0.3017 | 0.9063 | 0.2661 | 0.9229 | 0.8912 | 0.9027 | 0.9880 | 0.8964 |
| 2 | 0.3736 | 0.2325 | 0.2497 | 0.9052 | 0.2285 | 0.9138 | 0.8862 | 0.9053 | 0.9867 | 0.8946 |
| 3 | 0.3462 | 0.1978 | 0.1898 | 0.8986 | 0.1874 | 0.9138 | 0.8857 | 0.9087 | 0.9872 | 0.8947 |
| 4 | 0.3516 | 0.2605 | 0.2365 | 0.8998 | 0.2213 | 0.9248 | 0.9071 | 0.9031 | 0.9877 | 0.9038 |
| 5 | 0.3242 | 0.2039 | 0.2128 | 0.8990 | 0.1911 | 0.9266 | 0.8931 | 0.9072 | 0.9891 | 0.898 |
| 6 | 0.3407 | 0.3314 | 0.2909 | 0.9036 | 0.2865 | 0.9229 | 0.8937 | 0.9244 | 0.9884 | 0.9063 |
| 7 | 0.3791 | 0.3794 | 0.2863 | 0.9048 | 0.2596 | 0.9339 | 0.9109 | 0.9259 | 0.9897 | 0.915 |
| 8 | 0.3297 | 0.2048 | 0.2278 | 0.8941 | 0.1970 | 0.9101 | 0.8745 | 0.9281 | 0.9865 | 0.8986 |
| 9 | 0.3297 | 0.2251 | 0.2130 | 0.8983 | 0.2111 | 0.9156 | 0.8841 | 0.8875 | 0.9869 | 0.8826 |
| 10 | 0.3846 | 0.3395 | 0.2699 | 0.9081 | 0.2512 | 0.9266 | 0.9014 | 0.925 | 0.9891 | 0.9088 |
| Mean | 0.3539 | 0.2638 | 0.2478 | 0.9018 | 0.2300 | 0.9211 | 0.8928 | 0.9118 | 0.9879 | 0.8999 |
Table 9.
Performance of the KNN model trained with synthetically augmented data.
Table 9.
Performance of the KNN model trained with synthetically augmented data.
| Model Id | Validation | Test |
|---|
| Acc | P | R | Sp | F1 | Acc | P | R | Sp | F1 |
|---|
| 1 | 0.8585 | 0.8241 | 0.8512 | 0.9785 | 0.8352 | 0.9725 | 0.9644 | 0.9623 | 0.9954 | 0.9631 |
| 2 | 0.8592 | 0.8284 | 0.8457 | 0.978 | 0.8334 | 0.9596 | 0.9492 | 0.9566 | 0.9933 | 0.9526 |
| 3 | 0.8571 | 0.8381 | 0.8248 | 0.9774 | 0.8254 | 0.9743 | 0.9737 | 0.9630 | 0.9952 | 0.9676 |
| 4 | 0.8688 | 0.8519 | 0.8448 | 0.9794 | 0.8404 | 0.9615 | 0.9487 | 0.9558 | 0.9938 | 0.9509 |
| 5 | 0.8757 | 0.8551 | 0.8662 | 0.9804 | 0.8581 | 0.9761 | 0.9656 | 0.9694 | 0.9961 | 0.9675 |
| 6 | 0.8606 | 0.8332 | 0.8383 | 0.9783 | 0.8297 | 0.9725 | 0.9584 | 0.9673 | 0.9959 | 0.9619 |
| 7 | 0.851 | 0.8211 | 0.8236 | 0.9763 | 0.8155 | 0.9706 | 0.9593 | 0.9678 | 0.9953 | 0.9633 |
| 8 | 0.8661 | 0.8458 | 0.8462 | 0.9791 | 0.8409 | 0.9725 | 0.9640 | 0.9595 | 0.9956 | 0.9613 |
| 9 | 0.8736 | 0.8515 | 0.8517 | 0.9792 | 0.8485 | 0.9798 | 0.9700 | 0.9694 | 0.9966 | 0.9695 |
| 10 | 0.8468 | 0.8263 | 0.8259 | 0.976 | 0.8222 | 0.9761 | 0.9685 | 0.9658 | 0.9959 | 0.9669 |
| Mean | 0.8617 | 0.8376 | 0.8418 | 0.9783 | 0.8349 | 0.9716 | 0.9622 | 0.9637 | 0.9953 | 0.9625 |
Table 10.
Average, standard deviation (SD), and 95% confidence interval (CI95) values of evaluation metrics (validation stage) for the overall and comparative interpretation of the performance of hybrid ML models.
Table 10.
Average, standard deviation (SD), and 95% confidence interval (CI95) values of evaluation metrics (validation stage) for the overall and comparative interpretation of the performance of hybrid ML models.
| Model Id | Validation |
|---|
| Acc | P | R | Sp | F1 |
|---|
RF (Original data)
| Mean ± SD | 0.49176 ± 0.0406 | 0.18297 ± 0.0857 | 0.19055 ± 0.0132 | 0.89293 ± 0.0045 | 0.15541 ± 0.0172 |
| CI95 | [0.4627, 0.5208] | [0.1217, 0.2443] | [0.1811, 0.2] | [0.8897, 0.8961] | [0.1431, 0.1677] |
RF (Synthetic data)
| Mean ± SD | 0.51846 ± 0.0157 | 0.7491 ± 0.0267 | 0.2361 ± 0.0099 | 0.89762 ± 0.0019 | 0.24022 ± 0.0153 |
| CI95 | [0.5072, 0.5297] | [0.73, 0.7682] | [0.229, 0.2432] | [0.8963, 0.899] | [0.2293, 0.2512] |
SVM (Original data)
| Mean ± SD | 0.54945 ± 0.0382 | 0.43615 ± 0.1034 | 0.26179 ± 0.023 | 0.90949 ± 0.0044 | 0.24261 ± 0.0343 |
| CI95 | [0.5221, 0.5768] | [0.3622, 0.5101] | [0.2453, 0.2782] | [0.9063, 0.9127] | [0.2181, 0.2672] |
SVM (Synthetic data)
| Mean ± SD | 0.80193 ± 0.0137 | 0.86051 ± 0.0123 | 0.70534 ± 0.0187 | 0.96113 ± 0.0025 | 0.76324 ± 0.0158 |
| CI95 | [0.7921, 0.8117] | [0.8517, 0.8693] | [0.692, 0.7187] | [0.9594, 0.9629] | [0.752, 0.7745] |
NB (Original data)
| Mean ± SD | 0.38845 ± 0.0427 | 0.18562 ± 0.0948 | 0.14983 ± 0.0227 | 0.13416 ± 0.0305 | 0.87765 ± 0.0046 |
| CI95 | [0.3579, 0.419] | [0.1178, 0.2535] | [0.1336, 0.1661] | [0.1123, 0.156] | [0.8744, 0.8809] |
NB (Synthetic data)
| Mean ± SD | 0.42184 ± 0.0151 | 0.43039 ± 0.0148 | 0.43667 ± 0.0171 | 0.91185 ± 0.0025 | 0.4099 ± 0.0164 |
| CI95 | [0.4111, 0.4326] | [0.4198, 0.441] | [0.4244, 0.4489] | [0.9101, 0.9136] | [0.3981, 0.4217] |
KNN (Original data)
| Mean ± SD | 0.35385 ± 0.0233 | 0.26379 ± 0.0646 | 0.24784 ± 0.0381 | 0.90178 ± 0.0044 | 0.22998 ± 0.0344 |
| CI95 | [0.3372, 0.3705] | [0.2176, 0.31] | [0.2206, 0.2751] | [0.8986, 0.905] | [0.2053, 0.2546] |
KNN (Synthetic data)
| Mean ± SD | 0.862 ± 0.009 | 0.838 ± 0.013 | 0.842 ± 0.014 | 0.978 ± 0.001 | 0.835 ± 0.013 |
| CI95 | [0.8551, 0.8684] | [0.8284, 0.8467] | [0.832, 0.8517] | [0.9773, 0.9793] | [0.8259, 0.844] |
Table 11.
Average, standard deviation (SD), and 95% confidence interval (CI95) values of evaluation metrics (test stage) for the overall and comparative interpretation of the performance of hybrid ML models.
Table 11.
Average, standard deviation (SD), and 95% confidence interval (CI95) values of evaluation metrics (test stage) for the overall and comparative interpretation of the performance of hybrid ML models.
| Model Id | Test |
|---|
| Acc | P | R | Sp | F1 |
|---|
RF (Original data)
| Mean ± SD | 0.93945 ± 0.0074 | 0.96514 ± 0.0051 | 0.8986 ± 0.0145 | 0.98717 ± 0.0018 | 0.92882 ± 0.009 |
| CI95 | [0.9342, 0.9447] | [0.9615, 0.9688] | [0.8882, 0.909] | [0.9859, 0.9885] | [0.9224, 0.9352] |
RF (Synthetic data)
| Mean ± SD | 0.94239 ± 0.0078 | 0.96797 ± 0.0064 | 0.90105 ± 0.0115 | 0.98768 ± 0.0017 | 0.93127 ± 0.0084 |
| CI95 | [0.9368, 0.948] | [0.9634, 0.9726] | [0.8928, 0.9093] | [0.9865, 0.9889] | [0.9253, 0.9373] |
SVM (Original data)
| Mean ± SD | 0.82367 ± 0.006 | 0.93013 ± 0.0054 | 0.69783 ± 0.0092 | 0.96311 ± 0.0014 | 0.77121 ± 0.0081 |
| CI95 | [0.8194, 0.828] | [0.9263, 0.934] | [0.6913, 0.7044] | [0.9621, 0.9641] | [0.7654, 0.777] |
SVM (Synthetic data)
| Mean ± SD | 0.90955 ± 0.0046 | 0.94054 ± 0.0064 | 0.84974 ± 0.0076 | 0.98164 ± 0.0009 | 0.88808 ± 0.007 |
| CI95 | [0.9062, 0.9129] | [0.9359, 0.9451] | [0.8443, 0.8552] | [0.981, 0.9822] | [0.8831, 0.8931] |
NB (Original data)
| Mean ± SD | 0.86514 ± 0.0083 | 0.87319 ± 0.0087 | 0.87863 ± 0.0124 | 0.87162 ± 0.0096 | 0.97721 ± 0.0017 |
| CI95 | [0.8592, 0.8711] | [0.867, 0.8794] | [0.8697, 0.8875] | [0.8647, 0.8785] | [0.976, 0.9784] |
NB (Synthetic data)
| Mean ± SD | 0.51357 ± 0.0058 | 0.63643 ± 0.0089 | 0.62629 ± 0.0071 | 0.93291 ± 0.0007 | 0.54866 ± 0.007 |
| CI95 | [0.5095, 0.5177] | [0.63, 0.6428] | [0.6212, 0.6314] | [0.9324, 0.9334] | [0.5436, 0.5537] |
KNN (Original data)
| Mean ± SD | 0.9211 ± 0.0075 | 0.89279 ± 0.0111 | 0.91179 ± 0.0134 | 0.98793 ± 0.0011 | 0.89988 ± 0.009 |
| CI95 | [0.9158, 0.9264] | [0.8848, 0.9007] | [0.9022, 0.9214] | [0.9871, 0.9887] | [0.8934, 0.9063] |
KNN (Synthetic data)
| Mean ± SD | 0.972 ± 0.006 | 0.962 ± 0.008 | 0.964 ± 0.005 | 0.995 ± 0.001 | 0.962 ± 0.006 |
| CI95 | [0.967, 0.9761] | [0.9562, 0.9681] | [0.9601, 0.9673] | [0.9946, 0.996] | [0.958, 0.967] |
Table 12.
Average, standard deviation (SD), and 95% confidence interval (CI95) values of ROC-AUC.
Table 12.
Average, standard deviation (SD), and 95% confidence interval (CI95) values of ROC-AUC.
| Model Id | Validation | Test |
|---|
RF (Original data)
| Mean ± SD | 0.77745 ± 0.01688 | 0.9948 ± 0.0013 |
| CI95 | [0.7654, 0.7895] | [0.9939, 0.9957] |
RF (Synthetic data)
| Mean ± SD | 0.9017 ± 0.00518 | 0.99672 ± 0.00211 |
| CI95 | [0.898, 0.9054] | [0.9952, 0.9982] |
SVM (Original data)
| Mean ± SD | 0.88203 ± 0.01085 | 0.99016 ± 0.0014 |
| CI95 | [0.8743, 0.8898] | [0.9892, 0.9912] |
SVM (Synthetic data)
| Mean ± SD | 0.97967 ± 0.0015 | 0.995325 ± 0.00098 |
| CI95 | [0.9786, 0.9807] | [0.9946, 0.996] |
NB (Original data)
| Mean ± SD | 0.51393 ± 0.01398 | 0.93027 ± 0.00719 |
| CI95 | [0.5039, 0.5239] | [0.9251, 0.9354] |
NB (Synthetic data)
| Mean ± SD | 0.69413 ± 0.01001 | 0.80661 ± 0.00263 |
| CI95 | [0.687, 0.7013] | [0.8047, 0.8085] |
KNN (Original data)
| Mean ± SD | 0.5748 ± 0.02081 | 0.94984 ± 0.00695 |
| CI95 | [0.5599, 0.5897] | [0.9449, 0.9548] |
KNN (Synthetic data)
| Mean ± SD | 0.91004 ± 0.0075 | 0.97951 ± 0.00301 |
| CI95 | [0.9047, 0.9154] | [0.9774, 0.9817] |
| Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |