Abstract
The intestinal microbiota is closely related to liver diseases via the intestinal barrier and bile secretion to the gut. Impairment of the barrier can translocate microbes or their components to the liver where they can contribute to liver damage and fibrosis. The components of the barrier are discussed in this review along with the other elements of the so-called gut–liver axis. This bidirectional relation has been widely studied in alcoholic and non-alcoholic liver disease. However, the involvement of microbiota in the pathogenesis and treatment of viral liver diseases have not been extensively studied, and controversial data have been published. Therefore, we reviewed data regarding the integrity and function of the intestinal barrier and the changes of the intestinal microbioma that contribute to progression of Hepatitis B (HBV) and Hepatitis C (HCV) infection. Their consequences, such as cirrhosis and hepatic encephalopathy, were also discussed in connection with therapeutic interventions such as the effects of antiviral eradication and the use of probiotics that may influence the outcome of liver disease. Profound alterations of the microbioma with significant reduction in microbial diversity and changes in the abundance of both beneficial and pathogenic bacteria were found.
1. Introduction
A connection between the intestine and the liver was already postulated approximately two thousand years ago when the Greek-Roman doctor Galen suggested a connection between the gut and the liver [1].
In modern medicine, the clustering of microorganisms living in the same environment has been defined as microbiota, while the term microbiome applies to the collective genomes of the microbes [2,3].Microbiota are composed of bacteria, archaea, protozoans, fungi, and viruses [4]. Interestingly, every human being has their unique composition of gut microbiota properly defined as the “microbial fingerprint” [5].
The bacterial component of the microbiota is classified into 12 different phyla and 93.5% of the total belongs to Proteobacteria, the Gram positive Firmicutes, Actinobacteria, and the Gram negative Bacteroidetes. Bacteroides and Prevotella are the main genera of Bacteriodetes. Clostridium, Blautia, Enterococcus, Faecalibacterium, Eubacterium, Roseburium, Ruminococcus, Streptococcus, and Lactobacillus are the most prevalent genera of Firmicutes. Actinobacteria include Bifidobacteria, Atopobium, and Collinsella, while Proteobacteria are mainly composed of Enterobacteriaceae such as Escherichia and Klebsiella. Akkermansia muciniphila is the only species of Verrucomicrobia found in the human gut [6,7,8]. Archaea are predominated by Methanobrevibacter species. Viruses and bacteriophages are also colonizing the gut in considerable quantities [9].
Firmicutes and Actinobacteria predominate among luminal bacteria populations, while Proteobacteria are abundant among mucosal populations [10]. Early in the life of humans, there is a restricted diversity of the microbiota which is mostly composed of Actinobacteria and Proteobacteria. Diversity and variability are increasing with age and the species of Bacteroides, Clostridium, and Escherichia coli predominate in the intestinal flora in individuals over 65 years of age [11,12,13]. The gut microbiome is also variable among different ethnic groups [14,15,16] and between rural and urbanized populations of the same ethnicity [15,16,17,18]. Microbiota also differ between countries and continents [14,15,19].
Fungal species are also found in the gut including Candida, Saccharomyces, Aspergillus, Penicillium, Rhodotorula, Trametes, Pleospora, Sclerotinia, Bullera, and Galactomyces [20].
The human intestinal microbiota is now considered as a significant superorganism [21], colonized by approximately one-hundred trillion bacteria comprising nearly 40,000 types of microbes [22,23,24,25] most of which cannot be cultured, and 200–300 fungal species [26,27,28]. Microbial cells in the body are 10- to 100-fold higher than human cells [29,30]. In all, the microbiota weights approximately 1–2 kg in the adult, while the genetic material exceeds that of the human by about 100 times indicating its significance in human homeostasis [31,32].
There are several pathways of communication between microbes and the human host. This is achieved through different microbial components and products such as lipopolysaccharides (LPS), bacterial DNA, flagellin, short-chain fatty acids (SCFAs), tryptophan (Trp), and secondary bile acids (BAs) [33]. All these are recognized by pattern recognition receptors, mainly the Toll-like receptors (TLRs) family.
These pathways have been recently studied in connection with alcoholic or metabolic liver disease, but viral hepatitis and its consequences have not studied in depth. The purpose of this review, therefore, is to present the current data on the interplay between the intestinal microflora and chronic viral disease and the therapeutic implications of the manipulation of the gut microbiota.
2. The Gut–Liver Axis
The gut–liver axis is the mutual interaction between the gut microbiota and the liver. The portal vein transports gut produced components directly to the liver, and the liver provides the intestine with bile components including a wealth of antibodies [34]. A critical element of this mutual communication is the intestinal permeability. Bacterial products cross the intestinal barrier and modify the gut-associated lymphatic tissue (GALT) to release cytokines and chemokines along with other bacterial metabolites such as trimethylamine, and ethanol. A second barrier, the gut–vascular barrier (GVB), is the molecular sieve of components entering the portal-venous circulation to directly reach the liver [35,36,37].
The intestinal mucosal barrier is a complex functional structure consisting of three elements (a) the physical element comprised of several types of cells sealed by the tight intercellular junctions, (b) the gut-associated lymphoid tissue comprised of several immune cells in concert with the Peyer’s cells and mesenteric lymph nodes, and (c) the mucus layer secreted by the goblet cells that also contains immunoglobulin A (IgA) and antimicrobial products [38,39]. The function of the intestinal barrier is to maintain the tolerance to commensal organisms and food antigens and to mount a protective immune reaction to microbial pathogens and/or to microbial components defined as pathogen-associated molecular patterns (PAMPs) [40].
2.1. Physical Elements of the Intestinal Barrier
The intestinal barrier is a single layer of cells comprised of enterocytes, goblet cells, Tuft cells and enterochromaffin cells [41]. A physical barrier is provided by the cellular monolayer and the tight junctions, and an electrical barrier is operational as the negatively charged brush border repels the negative charge of the microbiota. In addition, many hydrophilic molecules are denied passage by the hydrophobic nature of cell membranes of intestinal cells. The integrity of the epithelial barrier is further accomplished by an intercellular seal comprised—from the apex of the cell to the base—of tight junctions (TJs, zonula occludens), the adherens junction (AJ, zonula adherens), and the desmosome (macula adherens) [41,42]. TJs consist of more than 50 proteins. Membrane-associated scaffolding proteins such as zonula occludens 1, 2, and 3 anchor TJ to the actin cytoskeleton [43,44]. They include tight junction-associated MARVEL proteins (TAMPs), claudins, and junctional adhesion molecules (JAMs) [43,45]. The function of TAMPs, such as occludin, is not completely clarified as TJs are formed even in the absence of occludin [45]. More than 27 claudin family members have been described [46]. JAMs, on the other hand, are related to signaling pathways of cell polarity and regulate permeability via non-selective pathways [47,48,49].
As mentioned before, there is a second physical barrier, the gut–vascular barrier that also contains TJs and AJs. This barrier prevents bacterial and antigen translocation from entering the portal circulation and reaching the liver due to the presence of the plasmalemma vesicle-associated protein-1. Its function is dependent on the Wnt/β-catenin signaling pathway. Salmonella typhimurium can cross this barrier by interfering with Wnt/β-catenin that controls AJ functionality via E-cadherin/β-catenin [50,51]. It should be noted that Hepatitis B virus also affects the Wnt/β-catenin signaling [52].
2.2. Control of the Microbiota by the Gut-Associated Lymphoid Tissue
Several immune cells contribute to the intestinal barrier. Dendritic cells, various subgroups of lymphoid cells, and macrophages protect from pathogens and provide the necessary tolerance to ingested food antigens and commensal bacteria [53]. Immune cells are located either in the lamina propria or within the epithelium. Intraepithelial cells include αβ and γδ T lymphocytes and mononuclear phagocytes [54,55,56]. Intraepithelial lymphocytes are cytolytic and are activated by epithelial cell cytokines [57]. Intraepithelial phagocytes are critical for tolerance development. Their luminar protrusions sense bacterial and food components and present their peptides into the lamina propria dendritic cells [58]. Immunocytes of the lamina propria are the next line of protection. CD4+ T lymphocytes, innate immunity associated NKT cells, and mucosal associated invariant T cells (MAIT cells) are highly specialized for particular antigens. NKT cells recognize lipids [59], while MAIT cells recognize metabolites of vitamin B2 [60,61]. CD4+ T cells are mostly Th17 cells that are induced after adhesion of filamentous bacteria to the intestinal epithelium [62,63], and T regulatory cells [64]. The production of IL-22 by Th17 cells improves the integrity of the tight junctions. Conversely, production of IL-17 is pro-inflammatory and pro-fibrotic [65]. Interactions between the host and the microbiota are mediated by soluble factors called postbiotics [66,67]. Intestinal microbes produce during the degradation of dietary fibers short chain fatty acids (SCFAs), such as acetate, propionate, and butyrate, which are the main postbiotics. SCFAs are consumed by other butyrate-producing microorganisms, such as Roseburia, Faecalibacterium, and Eubacterium [68]. SCFAs directly strengthen tight junctions [69,70]. They also stimulate mucin production and intestinal motility [66]. SCFAs also sensitize intestinal epithelial cells (IEC) to bacterial products [71]. They also regulate immunity in the GALT as they inhibit macrophage and dendritic cell activation and shape the T helper cell repertoire [72,73,74] controlling the differentiation of T regulatory cells [75]. Bifidobacteria are examples of protective bacteria that produce SCFAs leading to decreased production of TNF-a, IL-1b, and IL-6 by macrophages while reinforcing production of IL-10 [76,77]. Conversely, bacteria of the Enterobacteriaceae family such as Klebsiella, Escherichia coli, Proteus, and Enterobacter produce ethanol and promote liver damage [78]. They also release large amounts of lipopolysaccharide (LPS) in the intestinal lumen that favor the production of pro-inflammatory cytokines [79], and increase intestinal permeability and the translocation of bacteria [80].
Commensal bacteria also strengthen the barrier through their interaction with Toll-like receptor (TLRs) [81] and the production of mediators that can affect the binding proteins [82,83]. Isoforms of protein kinase C are phosphorylated after activation of TLR2 leading to up-regulated expression of zona occludens and the sealing of tight junctions [82]. On the other hand, the expression of occludin is down-regulated after activation of TLR4 increasing intestinal permeability [83]. Escherichia coli and Clostridia difficile are examples of bacteria that can affect the binding proteins and open the paracellular routes [84]. Humans express ten TLRs [85] responding to viral and bacterial proteins or endogenous ligands without infection [86]. Kupffer cells, the main cells responding to TLRs ligands, express all TLRs except TLR5 [87]. Hepatocytes, biliary endothelial cells, hepatic stellate cells (HSCs), and sinusoidal epithelial cells also express TLRs, but only HSCs express all nine TLRs [88,89]. The production of inflammatory cytokines, chemokines, and reactive oxygen species by the Kupffer cells after TLRs activation, lead to liver damage [90] and the activation of both the innate and adaptive immune immunity [91,92]. Ligands bind to all TLRs except TLR3 and activate a signal pathway operated by the myeloid differentiation factor 88 (MyD88) [93,94] that in turn activates nuclear factor-kappa B (NF-κB) and promotes the transcription of pro-inflammatory cytokines, such as TNF-α, IL-1β, and IL-6 [85]. More specifically, activation of TLR4 and TLR9 releases IL-12 and pro-inflammatory cytokines [95], while TLR4 activation may also induce the release of IL-23 that favors the survival of pro-inflammatory Th 17 lymphocytes [96]. On the other hand, activation of TLR2 releases the anti-inflammatory IL-10 and IL-13 [97]. LPS from Gram-negative bacteria activates TLR4 [85,98], while unmethylated CpG bacterial and viral sequences activate TLR9 [99]. TLR2 is activated by ligands of the HCV virus [86,100,101]. Endotoxin-mediated activation of TLR4, TLR9, and macrophage TLR2 activation by endotoxin is the primary driver of the development of liver fibrosis [102,103,104,105]. TLR4 signaling induces fibrosis by reducing the BMP and activin membrane-bound inhibitor homologue (BAMBI) which is a decoy receptor for TGF-β in hepatic stellate cells [106]. TLRs activation also increase the expression of the major histocompatibility complex on antigen presenting cells [107].
Intestinal epithelial cells (IECs) are also involved in intestinal immunity as they are exposed to a wealth of antigens acting as sensors of the microbiome through the pattern recognition receptor families such as the nucleotide-binding oligomerization domain-like receptors (NODs), TLRs, and the aryl hydrocarbon receptor (AhR). IECs, unlike immune cells, produce through TLR4 and AhR signaling interleukin-10 (IL-10), leading to tolerance to commensal bacteria. Commensal stimulation of IECs also promotes tolerance of dendritic cells and macrophages through the production of transforming growth factor beta (TGF-β), IL-10, and retinoic acid (RA) [40,108]. Interestingly, only limited species such as Peptostreptococcus russellii and Lactobacillus produce AhR ligands [109]. Lactobacilli species can convert Trp into indole-3-aldehyde, a ligand for AhR leading to the production of IL-22 [110].
Detailed description of the interaction of HCV, HBV, and the TRLs have been published [111,112].
2.3. Stratification of the Microbiota by Mucus
The mucus separates the microbiota from the intestinal cells and inhibits an excessive inflammatory reaction. Very few species, such as the filamentous bacteria, present in early life [113], can cross the mucus directly interacting with the epithelial cells [62]. All other bacteria indirectly interact with the host through their metabolic products [66,114,115]. Intestinal mucus has two layers: the almost sterile inner layer attached to the epithelium, and the outer layer colonized by bacteria. Mucus is thicker in the terminal ileum and large bowel [116,117]. Secreted mucins (MUCs), such as MUC2 and transmembrane MUCs, comprise the inner and outer mucous layers [118]. Bacteria can attach to mucus through the mucin–immunoglobulin A interactions [119]. The composition of the mucus is determined by the diversity of microbiota [120] because goblet cells sense the presence of different bacterial products. This is probably due to the capacity of the so-called sentinel goblet cells to act as sensors for the presence of different bacterial products and to modify MUC2 mucin secretion after activation of the Nlrp6 inflammasome [121]. It should be noted that several bacteria, such as Akkermansia municiphila, can degrade mucins and use them as a source of nutrients [122].
The intestinal mucus layers increase their defenses by antibacterial lectins, such as the regenerating islet-derived protein IIIγ (REG3G), which are secreted by the Paneth cells to fight bacteria present in the mucosal lining [123,124,125]. In addition, plasma cells can secrete immunoglobulins A (sIgAs) which is transported to the lumen and neutralize microbial pathogens [126]. IgA may be used by commensal microbes, such as Bacteroides fragilis, to facilitate mucus attachment [127]. Additionally, sIgAs neutralize bacterial toxins as well [128,129]. The large diversity of antimicrobial peptides inhibits bacteria to develop resistance to these proteins [130].
Apart from intestinal inflammation, the most important driver of dysregulated barrier permeability is intestinal dysbiosis [131,132]. The major unsolved problem with dysbiosis is whether it is the cause or the effect of the disease. The main promoter of dysbiosis is the use of antibiotics leading to their association with several autoimmune diseases [133,134,135,136]. Dysbiosis may result from either a reduction of beneficial microorganisms or the predominance of pathogens and the alteration of the total microbial diversity [137]. Intestinal dysbiosis usually leads to tight junctions weakening and increased translocation of microbes or microbial elements such as LPS, DNA, and β-glucan from fungi. They are collectively designated as microbial-associated molecular patterns (MAMPs) or pathogen-associated molecular patterns (PAMPs). MAPS and PAMPS can activate Kupffer and hepatic stellate cells that ultimately lead to liver damage and fibrosis [106,138,139,140].
The communication between the gut and the liver is achieved through the biliary tract and the portal vein. Liver produced mediators such as bile acids influence the gut microbiota and intestinal permeability, while intestinal products are involved in bile acid synthesis and glucose and lipid metabolism in the liver [141,142]. Translocation of gut bacteria or bacterial components to the liver, mesenteric lymph nodes, and other extra-intestinal sites is the result of tight junction abnormalities [106,143,144]. Lactate, which is produced by bacterial carbohydrate fermentation, reduces barrier permeability before its own fermentation to butyrate by intestinal flora [145]. Harmful bacterial products, such as LPS and unmethylated CpG, are then delivered to the liver and activate TLRs as mentioned before [85].
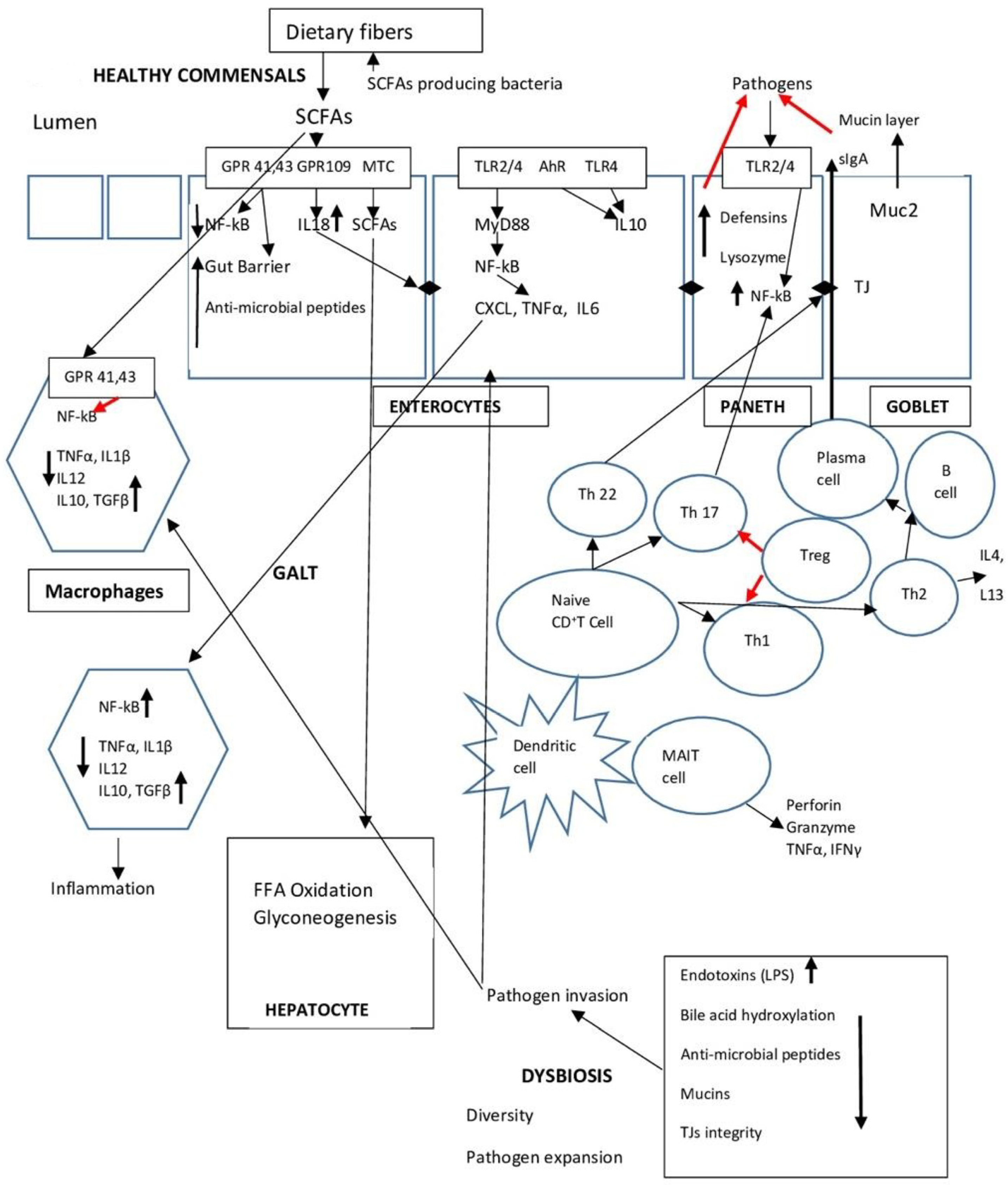
An important mechanism in the bidirectional communication between the liver and the intestine is the enterohepatic circulation of bile acids (BAs) [146]. Primary Bas, such as cholic acid (CA) and chenodeoxycholic acid (CDCA), are involved in enterohepatic circulation after their secretion by the hepatocytes. They are transported to the intestinal lumen as glycine or taurine conjugates [147]. In the lumen, the bacterial enzyme bile salt hydrolase (BSH) acts on BSH is produced by both Gram-positive and Gram-negative bacteria in the gut such as Bacteroides, Clostridium, Lactobacillus, Bifidobacterium, and Listeria [148]. Luminal bile acids are re-absorbed in the terminal ileum by the apical sodium-dependent bile acid transporter (ASBT) or passively dross the epithelium in the colon [36]. Secondary BAs may increase intestinal permeability affecting the stability of cellular membranes [149]. Gut bacteria also control the synthesis of BAs in the liver through the farnesoid X receptor (FXR) and G protein-coupled bile acid receptor 1 (GPBAR1, also known as TGR5) [150]. FXR is a transcriptional nuclear receptor mostly expressed in the liver, ileum, and kidneys. It is involved in lipid and glucose metabolism [151,152]. BAs bind to FXR of the enterocytes, inducing the transcription the fibroblast growth factor 19 (FGF19). FGF19 is consequently transported to the liver via the portal vein and binds to FGF receptor 4 on hepatocytes. This binding inhibits the enzyme cholesterol 7α-monooxygenase (CYP7A1) and reduces the de novo BA synthesis in hepatocytes [153,154,155]. In addition, binding of BA to FXR induces the production of antimicrobial peptides, such as angiogenin 1 and RNase family member 4, which restrict gut microbial overgrowth and intestinal barrier dysfunction [156,157]. FXR engagement can therefore preserve the epithelial barrier [158] and repair damage of the gut vascular barrier [159]. Additionally, BAs binding to TGR5 on the enterocyte membrane mediates host energy expenditure [160,161] and glucose homeostasis [162]. Figure 1 graphically depicts the complex interrelations as described above.
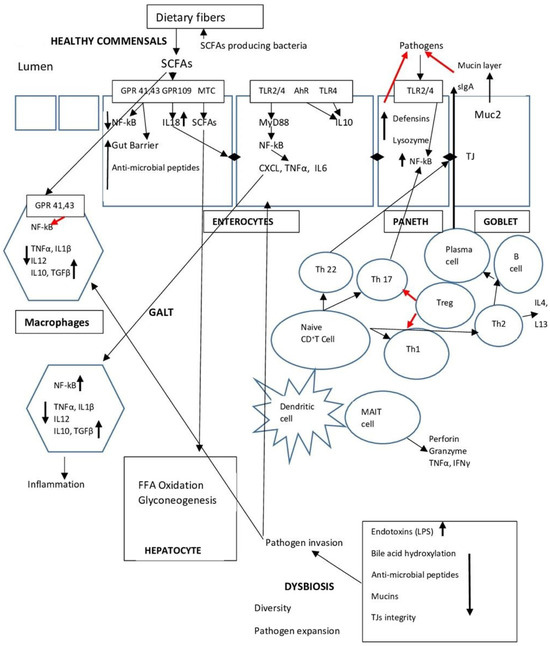
Figure 1.
Summarizes the main points of the interplay between microbiota, the intestinal barrier, and the gut immune system (GALT).
3. HBV Infection and Intestinal Microbiota
There are approximately 296 million people with chronic HBV infection worldwide, while 887,000 people die each year from complications of chronic HBV infection [163,164].
3.1. HBV and Intestinal Dysbiosis
HBV infection may be associated with intestinal dysbiosis [165] as demonstrated from animal experiments and clinical data. Thus, the ratio of Bacteroidetes and Firmicutes was stable in control mice, but it was significantly different in mice with HBV infection. Interestingly, differences were observed in Lactobacillus and Bifidobacterium between acute or chronic HBV infection [166]. In another experiment, decreased Blautia and Clostridium in HBV-infected mice were negatively correlated and increased Butyricicoccus, and Prevotellaceae were positively correlated with HBsAg and HBeAg levels. On the contrary, Akkermansia, which is considered a gut barrier protector, was reduced in HBV mice and was negatively correlated with HBV DNA in both serum and the liver [167].
Extensive changes in the gut microbiota composition have been reported in patients with chronic HBV infection [168,169]. Decreased genera of bacteria that metabolize bile acids have been described in association with changes in serum and fecal bile acids in chronic hepatitis B (CHB) patients with moderate/advanced fibrosis. Bacteroides and Ruminococcus were significantly lower in CHB patients compared to healthy controls. It was proposed that CHB fibrosis was in fact a modifier of the intestinal microbiota. Fibrosis limited the conversion of primary to secondary bile acids, activating the FXR and subsequently the FGF19 [170,171].
Microbiota changes already occur in early stage CHB patients. Operational taxonomic units (OTUs) belonging to Actinomyces, Clostridium, Lachnospiraceae, and Megamonas increased, while several OTUs decreased, including those belonging to Alistipes, Asaccharobacter, Bacteroides, and Butyricimonas [168]. The gut microbiota is also variable according to viral load. HBV patients with a low viral load have high diversity and taxa associated with fatty acid and lipid metabolism predominate [172]. LPS produced by Gram-negative intestinal bacteria was related to liver inflammation and cirrhosis. LPS levels were an independent predictor towards end-stage liver disease in patients with HBV infection [173]. Controversial results on the composition of microbiota have been reported. There was no difference in the intestinal microbiome between chronic HBV patients with normal ALT and normal volunteers. Megasphaera showed positive correlations, and Acidaminococcus exhibited a negative correlation with high ALT levels [174]. However, in another report, abundance of Lactobacillus, Clostridium, and Bifidobacterium were reduced in CHB patients with normal ALT compared to healthy controls [171]. In acute on chronic liver failure associated with HBV infection, the microbiota was enriched with Moraxellaceae, Sulfurovum, Comamonas, and Burkholderiaceae, but Actinobacteria, Deinococcus-Thermus, Alphaproteobacteria, Xanthomonadaceae, and Enterobacteriaceae were significantly reduced. Moreover, an increase of Prevotellaceae was a predictor of mortality [175].
In recent extensive studies, patients with all stages of HBV–related liver disease were examined and compared to healthy people. Firmicutes, Bacteroidetes, Proteobacteria, Actinobacteria, Verrucomicrobia, Cyanobacteria, and Fusobacteria accounted for almost 100% of the total sequences. Decreased Firmicutes and increased Bacteroidetes were found in all disease groups (Chronic Hepatitis, cirrhosis, Hepatocellular carcinoma) compared to healthy controls. Bifidobacterium and butyrate-producing bacteria families such as Clostridia and Ruminococcus were also decreased in all disease groups [176], but no difference was observed among patients with resolved HBV infection [176,177]. These findings may have pathogenetic implications as Bacteroidetes are Gram-negative bacteria which produce LPS, while Firmicutes are Gram-positive bacteria without LPS synthesis. Therefore, the higher Bacteroidetes/Firmicutes ratio means increased burden of LPS to the liver cells and increased liver damage [178]. On the other hand, the Enterobacteriaceae family bacteria comprising many pathogenic bacteria such as Klebsiella, Escherichia coli, Proteus, and Enterobacter were increased in all HBV groups [176,179]. The Enterobacteriaceae family were also increased in liver cirrhosis and were positively correlated to Child–Pugh (CP) score [180,181]. In detail, a negative correlation was found between the CP score and Bacteroidetes, while a positive correlation was demonstrated between CP score and Enterobacteriaceae or Veillonella [182]. Apart from increased LPS secretion, the Enterobacteriaceae produce endogenous ethanol that may be detrimental to the liver [79]. In addition, high Enterobacteriaceae release endotoxin that may cause inhibition of enterocyte protein synthesis leading to increased intestinal barrier permeability with further bacterial translocation to the liver [183]. In fact, two studies reported on barrier permeability in CHB patients. In the first, serum zonulin and copeptin were reduced in CHB patients and were negatively correlated with serum HBV DNA [184]. This was in disagreement with another study where serum zonulin was higher in HBV-related HCC, but no difference was observed in patients with CHB, cirrhosis or healthy controls [185].
A repeatedly confirmed finding of gut dysbiosis during progression of chronic HBV is the decrease of SCFAs-producing bacteria, such as Lachnospiraceae and Ruminococcaceae and their replacement by LPS-producing bacteria such as Enterobacteriaceae, Haemophilus, and Enterococcus [165,186]. The microbiota of HBV carriers contains more SCFA producers and less pro-inflammatory bacteria than patients with CHB, cirrhosis, and acute-on-chronic liver failure or hepatocellular carcinoma [187,188]. Another consistent finding of dysbiosis in HBV patients is that Bifidobacteria decrease with the increase of Enterobacteriaceae as the disease progresses. The ratio of Bifidobacteria/Enterobacteriaceae is reduced as disease severity progresses from CHB to cirrhosis and HCC [168,176,183,189].
Microbiota changes are difficult to be studied in human acute HBV. Results from animal studies have shown that the ratio of Firmicutes/Bacteroides increased early in the disease at day 14, and decreased in late disease at day 49 [166].
The above controversial reports indicate that interpretation and comparisons of results should be done with great caution as many studies are performed in populations with particular diet habits which influence the composition of the intestinal microbiome. Moreover, most studies are cross-sectional with samples representing an individual time point, and only a few were performed at different periods of HBV infection [165,187,190].
Detailed descriptions of the microbiome in the different stages of HBV infection have been recently published [191,192,193,194].
3.2. Microbiota and Immune Responses in HBV
Microbiota affects the immune response in HBV. Apart from the effects that LPS has on the immunological response through the activation of TLR4, an additional pathway is implicated in the immune response of patients with HBV. The unmethylated CpG DNA-TLR9 pathway can activate TLR9 that produces protective cytokines, such as Interferons. Unmethylated CpG DNAs is mainly produced by Lactobacilli, Bifidobacteria, Proteobacteria, and Bacteroidetes [195]. As mentioned above, Lactobacillus and Bifidobacteria are reduced in the gut microbiota of chronic HBV patients. Therefore, beneficial cytokines are reduced and the immune effects are defective in HBV [196,197].
Gut microbiota is implicated in the clearance of the HBV infection. When the gut microbiota is deregulated by antibiotics, the intestinal barrier function is probably impaired and the ability of immunity to clear HBV may be compromised [198]. Thus, adult mice with an intact intestinal microbiota clear HBV after 6 weeks of infection, while infection is not cleared in young mice or after antibiotic use [199,200]. Young mice with a TLR4 mutation achieved prompt HBV clearance. It therefore seems that a TLR4-dependent pathway of tolerance is operative in young animals and prevents HBV clearance. Development of intestinal microbiota stimulated the immune mechanisms and HBV clearance was feasible [201]. Additionally, impairment of intestinal microbiota was shown to affect the systemic adaptive immunity leading to delayed HBV antigen clearance. Gene analysis of Peyer’s patches (PPs) demonstrated that adaptive immunity was downregulated in intestinal microbiota-deficient mice, while the depletion of PPs led to higher HBsAg levels in serum [202]. Dysbiosis in mice and the resulting endotoxemia induced IL-10 production by the Kupffer cells and increased Kupffer cell-mediated T cell suppression. The immediate result was the protracted persistence of HBV infection [203]. However, in a mouse model of CHB, intestinal bacteria reduction by antibiotics had no effect on HBV replication in immune tolerant mice [204].
The immune response in HBV infection is also regulated by metabolic products produced by intestinal microbes, such as tryptophan, which interferes with the immune response of HBV through its metabolic product kynurenine [205]. Indoleamine-2,3-dioxygenase (IDO) is an enzyme induced by interferon that catalyzes tryptophan into kynurenine [206] acting as a suppressor of intracellular pathogens and as an immune regulator [207]. Inducible IDO was shown to suppress HBV replication in HepG2 cells with the HBV genome [208]. The effect of IDO in HBV clearance was investigated in HBV infected patients. In acute hepatitis patients who finally cleared the virus, IDO activity was high at the peak of ALT. In patients with hepatic flare, on the other hand, IDO activity remained low irrespective of ALT levels indicating that IDO is an anti-HBV factor only during the early phase of HBV infection [209].
Integrated studies of microbiome and metabolome showed an extensive shift of intestinal microbiota and metabolites in chronic HBV patients attributed to either disease evolution and/or antiviral treatment. Peripheral mononuclear cells incubated with bacterial extracts (BE) from non-cirrhotic patients promoted the expansion of Th17 lymphocytes, while BE from cirrhotics reduced Th1 cell count [210]. This is a particularly important findings that may explain some of the findings during liver fibrogenesis. Th17 immunity is an important factor in all stages of fibrogenesis in chronic HBV patients [211] including hepatic stellate cell activation [212,213], increased TGF-β production [214], the secretion of matrix metalloproteases (MMPs), and collagen synthesis [212,214].
3.3. Microbiota and HBV Treatment
Based on the above findings, it was only logical to suggest that manipulation of the microbiome might be beneficial for the evolution of HBV. Fecal microbiota transplantation (FMT) was tested, but the data are still restricted [215,216]. In an interesting experiment, the gut microbiome in BALB/c mice was abolished by antibiotics and replaced with FMT from naïve mice to investigate the effect of FMT on the immune response to HBV infection. HBV clearance differed considerably depending on the origin of FMT. The fecal microbiota from C57BL/6 but not from BALB/c mice induced tolerance and prolonged HBV infection [217].
Gut microbiota changes, induced via FMT, resulted in promising results in HBeAg-positive patients. A study on HBeAg-positive CHB patients under treatment with oral antivirals showed that FMT induces HBeAg clearance in some cases who had failed to clear HBeAg despite long-term antiviral treatment. The problem with this study is that only five patients were studied in the FMT group [218]. In a similarly designed recent larger study of 14 patients in the FMT arm, 16.7% of patients cleared and none in the antiviral only arm. It should be noted, however, that all patients retained the HBsAg in either arm. However, after six months, serum HBV DNA was reduced in the FMT arm but not in the controls [219].
An informative review on all aspects of FMT has been recently published [215].
The effects of oral antiviral treatment on gut microbiota have also been examined in HBV. In a persistent HBV mouse model, Akkermansia was significantly reduced in HBV-infected mice, while Entecavir therapy restored levels back to those of the normal controls. Akkermansia levels showed a negative correlation with HBV DNA levels in serum and liver [167]. On the contrary, Akkermansia was increased in patients with CHB and liver cirrhosis [176]. Therefore, additional studies are required on the actual role of Akkermansia in HBV. In the treatment of naïve patients, E. hallii group and Blautia were greatly reduced and were restored to normal levels after 5 years of entecavir treatment. Turicibacter with 4-hydroxyretinoic acid were negatively associated with AST [210,220].
The manipulation of intestinal microbiota with probiotics (Clostridium and Bifidobacterium) was tested in the treatment of minimal hepatic encephalopathy (MHE) in patients with HBV cirrhosis. Probiotics improved serum ALT and AST and albumin levels. Absolute fecal bacterial load of genera Fecal Clostridia and Bifidobacteria were increased, and Enterobacteriaceae were decreased. More importantly probiotics improved psychometric tests and cognition. Ammonia levels were reduced possibly due to the observed improvement of the intestinal microflora [221]. A recent study administered a mixture of lactulose, Clostridium butyricum, and Bifidobacterium longum infantis in a population of patients with HBV-related cirrhosis. The clinical response was insignificant, but intestinal dysbiosis and the metabolome of the patients improved compared to patients treated with placebo [222]. Obviously, more extensive studies are required, particularly when the above expressed reservations are considered.
4. HCV Infection and Intestinal Microbiota
Globally, approximately 58.5 million people are infected with HCV worldwide, while 1.75 million new cases are identified each year. Hepatocellular carcinoma (HCV-related) causes approximately 150,000 deaths and more than 350,000 deaths are HCV-related other complications. These figures are probably an underestimation of the real problem [223].
Gut microbiota has been connected to the various stages of HCV infection. A common finding of all studies performed so far is the lower bacterial diversity in HCV patients compared to healthy controls [180,224,225,226]. Diversity abnormalities are proportional to the stage of the disease [225]. Two hypotheses have been proposed that can explain how HCV infection can interfere with the gut–liver axis and the progression to fibrosis and cirrhosis. The first is that the gut microbiota is indirectly affected as a result of the liver damage. This is not compatible with changes in microbiota observed in early disease. The second hypothesis proposes a direct effect of HCV infection on B-lymphocytes and the consequent reduction of IgA production [224,227]. Reduced IgA secretion favors the abundance of Prevotella. Prevotella contains enzymes that may degrade mucin and increases the intestinal permeability leading to higher bacterial translocation [8]. A further indication of an impaired intestinal barrier in HCV-infected patients is also the finding of increased serum LPS levels [225,228].
Impairment of BAs metabolism is an additional explanation for the reduced microbial diversity in HCV. BAs profiles are different in chronic HCV compared with normal people. Fecal deoxycholic acid (DCA) was decreased and lithocholic or ursodeoxycholic acid predominated. The decrease in fecal DCA reduction was associated with Clostridiales reduction, while impaired synthesis of cholic acid (CA) was associated with a reduction in the transcription of CYP8B1, a key enzyme in CA synthesis [229]. This BAs disturbance results from overgrowth of pro-inflammatory bacteria, such as Porphyromonadaceae, Enterobacteriaceae, and reduction of Firmicutes the main producers of secondary bile acids [180,230,231,232].
The lower bacterial diversity is also associated with a reduction of the SCFAs producing Clostridiales, Lachnospiraceae, Ruminococcaceae, and an increase in Streptococcus and Lactobacillus, Prevotella and Faecaliberium [227,230]. SCFAs are critical for the differentiation of bowel regulatory T (Treg) cells that are the main suppressors of inflammation [233,234] as mentioned before. Apart from Clostridiales, the phylum of Firmicutes is also decreased in patients with chronic CHC. By contrast, the phylum of Bacterioidetes, the family of Enterobacteriaceae, Viridans streptococci, and the genera Bacteroides, Blautia, and Collinsella, are increased [216,235]. A recent study also demonstrated a decreased diversity and found that Lactic acid bacteria, and Lactobacillus acidophilus were higher in early stage of fibrosis compared to patients with advanced fibrosis [236].
Low diversity is already evident even in patients with normal transaminases and minimal disease with a transient increase in Bacteroides and Enterobacteriaceae. Metagenomics have shown an increase in the urease gene encoded by viridans streptococci that may account for the hyperammonemia present in the later stages of the disease [232]. Similarly, bacterial translocation due to intestinal barrier dysfunction was reported in the absence of fibrosis, indicating that impairment of the gut barrier occurs even at the early stages of chronic HCV [173,237].
In contrast to all other reports, a recent study showed an increased microbiota diversity in patients with HCV infection compared to healthy individuals. A higher abundance of Prevotella, Collinsella, Faecalibacterium, Megasphera, Mitsuokella multacida, and Ruminococcaceae, and a lower abundance of Bacteroides, Alistipes, Streptococcus, and Enterobacteriaceae was observed. Possible explanations for the discrepancy may be the stages of disease analyzed, the effect of HCV genotypes, and, most importantly, the demographic characteristics of the study groups [238].
An important finding was recently reported. The use of Proton pump inhibitors (PPIs) was related to significant alterations of the microbiota in patients with chronic HCV infection which were more pronounced in patients with liver cirrhosis. Streptococcus species, Enterobacter species, and Haemophilus species were significantly increased in patients with PPI use irrespective of the stage of liver disease [239].
Detailed descriptions of microbiota alterations in the different stages of progressive severity have been recently published [240,241].
Effect of HCV Treatment on Intestinal Microbiota
The initial treatment of HCV infection with interferon showed that the microbes before and after treatment were not different [242].
The use of effective direct acting antivirals (DAAS) in the HCV elimination prompted a series of studies of the potential effects of treatment on intestinal bacteria. The use of DAAs in patients with chronic HCV infection could only rectify the intestinal bacterial abnormalities only in with initial degrees of fibrosis [243]. A later study verified these results. Bacterial diversity was restored in patients without cirrhosis after sustained viral response (SVR) within 24 weeks after the end of treatment. No diversity improvement was found in SVR patients with cirrhosis. The abundances of Collinsella and Bifidobacter genera were increased between baseline and SVR only in non-cirrhotic patients [244]. However, in patients with genotypes 1,2,3 4 treated with glecaprevir/pibrentasvir, no significant differences in microbiota diversity, or microbial pattern were found before and after treatment at week 12 [245]. The same negative results were also very recently reported [246]. Two further reports also produced negative results. No significant alterations in the overall composition of gut microbiome or alpha diversity were observed after viral eradication. Some differences in abundance of certain bacteria, such as Coriobacteriaceae, Peptostreptococcaceae, Staphylococcaceae, and Morganellaceae, were identified but the overall compositions was not different after HCV eradication [247]. The diversity of the gut microbiota did not significantly alter before and after DAAs, even though the relative abundances of Faecalibacterium and Bacillus increased after eradication [248]. The reason for this discrepancy is not clear but the question is open to more detailed and larger studies.
The impact of DAAs on intestinal microbiota when cirrhosis is present also remains controversial as both favorable and negative studies have appeared and will be presented in the relevant section below [230,242].
Sustained viral response (SVR) seems to be a decisive factor, as alleviation of intestinal dysbiosis and microbial translocation were observed in responders but not in non-responders. Viral elimination increased the abundance of SCFAs-producing bacteria such as Blautia and Bifidobacterium [249]. However, successful response to DAAs eradication did not affect the intestinal barrier function. It is therefore likely that bacterial translocation is connected to abnormal composition of gut microbiota rather than to gut barrier dysfunction after DAAs therapy [230,249]. These reports are not consistent with findings demonstrating that microbial translocation markers, such as the lipopolysaccharide binding protein (LBP), were reduced after HCV elimination [250].
An interesting approach for restoration of gut dysbiosis is the use of Bacteriophages. In reality, phages are viruses that attack and eliminate bacteria [251]. The gut dysbiosis observed in HCV could potentially be corrected by using bacteriophages that target the chronic HCV associated bacteria [252], but this remains to be tested.
The current evidence on the effects of the gut microbiota in the evolution of HCV infection and the impact of DAAs elimination has been recently reviewed [31].
5. Other Hepatitis Viruses (A, D, and E)
Very limited information is available for these viruses, regarding mostly patients with acute hepatitis E (HEV) infection. The gut microbiota of these patients was enriched in Proteobacteria, and Enterobacteriaceae compared to normal controls. The presence of Gamma proteobacteria was positively related to ALT and total bilirubin levels and may be used as a predictor of the acute infection [253]. Significant changes were observed between acute uncomplicated HEV and HEV-associated acute liver failure (HEV-ALF). The HEV-ALF subgroup of patients had higher levels of Gamma proteobacteria, Proteobacteria, Xanthomonadceae, and Stenotrophomonas, and decreased abundance of Firmicutes, Streptococcus, and Lactobacillus, when compared with the acute HEV group. The relative abundances of Lactobacillaceae and Gammaproteobacteria were positively correlated with Th lymphocytes, and degree of hepatic encephalopathy. Survival was associated with higher levels of Lactobacillus mucosae compared to deceased patients [254]. The administration of the probiotic bacterium Enterococcus faecium in pigs led to the reduction of enteric HEV viruses and accelerated viral clearance. However, no human trials have been performed [255].
The sporadic nature of acute Hepatitis A (HAV) prevented an extensive investigation of the intestinal microbiota. A Japanese study of an HAV outbreak among HIV positive patients showed significant microbe abnormalities and persistence of dysbiosis persisted for a long time after recovery [256].
No data about gut microbiota changes during hepatitis D virus (HDV) infection exist as of yet. This is almost impossible to be clarified since HDV infection always co-exists with HBV, meaning separate data are difficult to obtain [257].
6. Cirrhosis and Intestinal Microbiota
Chronic liver disease is associated with several abnormalities of the intestinal microbiome leading to reduced commensal diversity, expansion of pathogenic species and disruption of the intestinal defensive barriers [40]. Interestingly, microbial abnormalities in cirrhosis are independent of etiology [169,258,259]. Therefore, they are also applicable in viral cirrhosis as well.
Microbiome alterations were reported in alcohol-associated and hepatitis-associated cirrhosis [180]. Increased levels of Veillonella, Megasphaera, Dialister, Atopobium, and Prevotella genera were described in the duodenum of patients with cirrhosis. Interestingly, Neisseria and Gemella genera could differentiate between HBV and PBC cirrhosis [260]. The role of intestinal microbiota in non-alcoholic liver disease is possibly the most extensively investigated, but analysis is beyond the scope of the present review [261,262].
Reduced diversity, increased abundance of pathogenic species, such as Staphylococcaceae and Enterobacteriaceae, and decreased colonization by beneficial commensals such as Lachnospiraceae and Ruminococcaceae are all characteristics of cirrhosis. Enterobacteriaceae increase with progression of liver disease and decompensation [169,180,263].
Gut barrier disruption is well recognized in cirrhosis. It is due to reduced expression of the tight-junction proteins occludin and claudin [264]. In addition, the impairment of antimicrobial host defense, as demonstrated in experimental cirrhosis, allows for bacterial invasion of the inner mucous layer of the gut and increased bacterial translocation [265].
BAs are important regulators of the intestinal microbiome. Abnormalities in either the quantity or the composition of BAs in the intestinal lumen as observed in cirrhosis, will lead to a reduction of beneficial bacteria and an increase in pathogenic bacteria [153,266,267]. De novo suppression of BAs may be obtained by activation of the intestinal FXR-FGF19 pathway. As a consequence, the levels of Firmicutes and Actinomycetes with BSH activity will increase in association with up-regulated excretion of BAs from feces to protect from liver injury and fibrosis that otherwise would result from the toxicity of hepatic bile acids [268,269].
6.1. Involvement of Microbiota in the Pathogenesis of Cirrhosis
Portal hypertension, inflammation, and oxidative stress damage the gut barrier and participate in the complications of cirrhosis The degree of the barrier damage and bacterial translocation parallels the severity of cirrhosis and the appearance of ascites [34,270]. A damaged barrier in cirrhosis is associated with restricted secretion of antibacterial peptides such as α- Defensins by Paneth cells [271]. LPS overproduction by gut microbes activate Paneth cells to secrete angiogenic molecules that promote mesenteric angiogenesis and the development of portal hypertension [272]. LPS overproduction may also participate in the progression of liver fibrosis through interaction with the TLR4. Bacterial translocation promotes liver fibrosis and inflammation via activation of hepatic stellate cells and Kupffer cells [37,102]. LPS also activates the pattern-recognition receptor (PRR) mostly on macrophages, leading to the activation of quiescent HSC into myofibroblasts [273,274,275,276] and progression of fibrosis.
An additional pathway through which microbiota are engaged in the pathogenesis of cirrhosis is the activation of inflammasomes, the protein complexes found in most cells including Kupffer cells, hepatocytes, and HSCs [277,278]. They release pro-inflammatory cytokines such as IL-1β and IL-18 and promote inflammation and fibrosis in the liver [279,280]. TLRs and inflammasomes have different routes of activation [277], but their role is complementary in the communication between the gut microbiota and the systemic immune response [86]. Interestingly, TLRs may counteract the inflammatory activity of inflammasomes. Thus, chronic stimulation of the TLRs by LPS induces IL-10 production restricting inflammasome activation [281]. Moreover, the activation of TLR2 or TLR4 can upregulate the autophagy of hepatocytes that leads to the degradation of inflammasomes attenuating inflammation [282]. It should be noted that the interplay between intestinal microbiota portal hypertension and fibrosis resembles a mutual relationship, similar to that between the chicken and the egg, as they affect each other [283].
6.2. Microbiota and HE
The gut microbiota is a critical mediator in the interrelation between the liver and the brain. Gut dysbiosis influences the cognitive behavior of cirrhotic patients mediated by the gut–liver–brain axis [284,285,286]. The already described change of reduction of beneficial and overgrowth of pathogenic bacteria in cirrhosis is augmented as hepatic encephalopathy (HE) appears indicating a bidirectional relation between gut microbiota and the nervous systems of the body including the enteric nervous system, the autonomic nervous system, and the neuroendocrine and neuro-immunity systems of the central nervous system [287]. This microbial alteration further compromises the production of SCFAs in concert with increased barrier permeability and bacterial translocation [287,288]. Derangements of bile acids are also implicated in the pathogenesis of HE and the role of microbiota. Alterations in intestinal microbiota are associated with a reduced conversion of primary to secondary fecal BAs [259]. It has been shown that the serum conjugated BAs are increased in cirrhosis. During HE, there is a further increase in serum and brain BAs that can act as detergents increasing the permeability of blood–brain barrier and brain damage [289]. Enterobacteriaceae and fecal CDCA were correlated with the degree of HE, while Ruminococcaceae positively correlated with DCA. DCA is an effective bactericidal for gut microbes, and a lower DCA/CA ratio improves the cognitive function in HE [290].
Patients with acute HE were found to have decreased Bacteroidetes and an increase in the relative abundance of Firmicutes, Proteobacteria, Actinobacteria, and Veillonella parvula increased [291]. Streptococcus salivarius was also increased even in minimal HE with sleep disturbances and had a positive correlation with ammonia levels [292,293]. A positive correlation between cognitive impairment and the overgrowth of Alcaligeneceae and Porphyromonadaceae has been demonstrated. This is particularly important as Alcaligenaceae produce ammonia by decomposing urea [294,295]. Other Gram-negative bacteria containing urease such as Streptococcus salivarius and Proteobacteria also metabolize urea to ammonia and are implicated in the pathogenesis of HE [296].
The extensive variety of microbial species and their dependence on exogenous factors not related to cirrhosis itself may cause difficulties in comparisons among different studies. An example is the presence of minimal HE. Microbiota results may differ in various studies depending on the methods used for the diagnosis of minimal HE. Thus, the abundances of Enterococcus and Streptococcus were higher in minimal HE diagnosed by the psychometric encephalopathy score, while Prevotella, Eggerthela, and Alistipes species were higher when minimal HE was diagnosed by the inhibitory control test [297]. Only Lactobacillaceae estimations were not dependent on the method used for minimal HE diagnosis [298,299]. A way to partly overcome this problem in clinical studies is the use of gut dysbiosis indices. One such index is the Cirrhosis Dysbiosis Ratio, which is the ratio of the abundance of Lachnospiraceae, Ruminococcaceae, Veillonellaceae, and Clostridiales Incertae sedis XIV over this of Bacteroidaceae and Enterobacteriaceae [263]. Another index is the Gut Dysbiosis Index [168] where a high index corresponds with severe dysbiosis.
6.3. HBV Cirrhosis
The prevalence of HBV-related cirrhosis is lower in Europe and the Americas, compared to Africa and Asia while HCV-related cirrhosis has very heterogeneous prevalence Globally, 42% of patient with cirrhosis had HBV infection and 21% HCV infection [300].
Specific intestinal microbiota alterations were described in HBV patients with cirrhosis. Prevalent phyla were Firmicutes (57%), Bacteroidetes (28.6%), with less abundant Proteobacteria, Actinobacteria, Fusobacteria, and Verrucomicrobia, adding up to almost 93% of the total sequences. Bacteroidetes were increased and Firmicutes were reduced HBV cirrhosis compared to the healthy individuals [194,301,302]. Patients with HBV cirrhosis had lower levels of beneficial bacterial taxa, such as Dialister and Alistipes, and higher levels of pathogenic species within Actinobacteria [165]. The lower Firmicutes/Bacteroidetes ratio may be pathogenetically associated with the progression of cirrhosis and inflammation. The Bifidobacteria/Enterobacteriaceae was decreased significantly in patients with decompensated HBV cirrhosis [183,303] while a reduced Megamonas genus level and increased Veillonella genus were risk factors for HBV-related liver cirrhosis [304]. In accordance with this scenario, Akkermansia, which is a protector of the intestinal barrier [305], was reduced in fecal samples of HBV cirrhosis with or without HCC [306,307].
Differences also exist between compensated and decompensated cirrhosis. Pathogenic bacteria, such as especially Alcaligenaceae, Porphyromonadaceae, Veillonellaceae, and Enterobacteriaceae, significantly increased in the decompensation stage [194,215]. Interestingly, there are differences in the composition of gut microbiota when diabetes mellitus co-exists with HBV cirrhosis (LCDM). Lactobacillus, Roseburia, and Veillonella increased in the LCDM compared to HBV cirrhosis. Moreover, Escherichia/Shigella, Veillonella, and Lactobacillus showed a positive correlation with liver damage and fasting blood glucose [308]. HBV patients with decompensated cirrhosis have increased sIgAs content in blood and stool compared to asymptomatic HBV groups and controls. This is consistent with increased bacterial migration. Enterobacteriaceae were positively correlated with sIgAs [183]. Zonulin, a regulator of tight junctions and a marker of intestinal permeability [309] was significantly increased in HBV cirrhosis and HCC patients, correlating with the stages of cirrhosis [185]. However, in HBV-associated HCC patients, unexpected up-regulation of anti-inflammatory bacteria, such as Prevotella and down-regulation of pro-inflammatory bacteria, like Escherichia, were reported in comparison to non-hepatitis- related HCC [179].
6.4. HBV-Related HCC
It is clearly established that the gut microbiome may influence the induction and progression of HCC by interfering with immune and metabolic pathways related to HCC. Data, both experimental [310] and clinical, mostly exists for non-viral HCC [311,312,313,314,315]. Recent findings have demonstrated that this is also true for HBV related HCC. Overgrowth of pathogenic bacteria of Gram-negative species and a significant increase in the fecal count of Escherichia coli are characteristic in HBV-related HCC [316]. Butyrate-producing bacteria, such as Ruminococcus, Oscillibacter, Faecalibacterium, Clostridium IV, and Coprococcus, were limited, while the LPS-producing bacteria Klebsiella and Haemophilus were augmented compared to cirrhosis patients [188,307].
Increased Prevotella abundance was also described in HBV-HCC compared to non-viral HCC [179]. Finally, changes in BA metabolism may contribute to the pathogenesis of HCC. Modifications of BAs metabolism by intestinal microbiota have already been described in HBV infection. Therefore, their implication in HBV-HCC induction is highly probable [317].
6.5. HCV Cirrhosis
As in HBV related cirrhosis, a reduced microbial diversity was reported in HCV cirrhotics compared to healthy individuals. Thus, higher levels of Prevotella and Faecalibacterium and lower levels of Acinetobacter, Veillonella, and Phascolarctobacterium were observed in the intestinal microbioma of Egyptian patients. Moreover, the ratio Prevotella/Bifidobacterium was proposed as a marker of fibrosis development [224]. Detailed descriptions of the diversity of microbial species in HCV cirrhosis have been recently presented. Thus, higher levels were found for Veillonella, Lactobacillus, Streptococcus, Alloprevotella, Bifidobacterium, Escherichia/Shigella, Haemophilus, Micrococcus, and Weissella species. On the other hands Bilophila, Clostridium, Mitsuokella, and Vampirovibrio species were significantly decreased. Interestingly, the beneficial Akkermansia series were also significantly increased [225,240,318].
Table 1 graphically depicts the main microbiome changes in chronic viral liver disease. It should be stressed however, that there are many discrepancies as the results are dependent on a variety of external factors.

Table 1.
Main Microbiome alterations in chronic viral liver disease. See text for details. Bidirectional light arrows indicate controversial results. Upward arrows indicate increase. Downward arrows indicate decrease.
The effect of treatment on gut microbiota has been examined in HCV-related cirrhosis. Treatment with pegylated interferon and ribavirin did not improve the composition of intestinal microbiota, even in those achieving SVR [242]. The effects of treatment with direct acting antivirals (DAAs) are controversial. DAAs administration modified the composition of the gut microbiota and reduced dysbiosis after achievement of SVR. The levels of pathogenic Enterobacteriaceae, Enterococcus, and Staphylococcus were decreased after treatment. However, intestinal barrier permeability was not affected [230]. A recent study reported that modifications of the gut microbiota after DAAs treatment was only observed in the absence of cirrhosis. No significant differences were observed in cirrhotic patients [244]. Recently, a small longitudinal study of patients with HCV-related cirrhosis and clinically significant portal hypertension was reported. Treatment with DAAs modified significantly the gut microbiome only in those with a significant reduction of portal pressure [319].
Fecal microbiota transplant may also improve gut dysbiosis and the intestinal microbiota in minimal HE as shown in two small studies that included a number of HCV patients [320,321]. Contrary to expectations, lactulose administration in cirrhotic patients did not affect intestinal microbiota. The study population included cirrhotic patients of mixed etiology, half of them patients with HBV and HCV cirrhosis. Phyla such as Tenericutes, Cyanobacteria, Spirochaetes, Elusimicrobia, and Lentisphaerae were lower in cirrhosis and did not change after lactulose [322]. This was not the case in a study of minimal HE patients, with half of them being HCV-related [323].This is also in disagreement with a more recent study of patients with viral cirrhosis, where administration of lactulose improved the cognitive abilities of cirrhotic patients and alleviated microbiota dysbiosis in minimal HE. In addition, lactulose responders had significantly different Actinobacteria, Bacteroidetes, Firmicutes, and Proteobacteria, compared to non-responders [324].
Table 2 summarizes the results of therapeutic modulations in the microbiota of chronic viral liver disease.

Table 2.
Therapeutic Manipulation of the microbioma in chronic viral liver disease. Upward arrows indicate increase. Downward arrows indicate decrease.
7. Conclusions
Gut microbiota is in constant communication with the liver microenvironment, affecting both hepatocytes and sinusoidal cells through the gut–liver axis. As in other chronic liver diseases, all the components of this communication are seriously affected in HBV and HCV infections. Intestinal barrier abnormalities lead to increased translocation of bacteria or their components that activate both innate and adaptive immunity in the intestinal lamina propria with subsequent activation of TLRs and various signaling pathways. A constant finding is the reduction of microbial diversity. Beneficial bacteria are reduced, and potential pathogens are increased. Thus, decreased Firmicutes and increased Bacteroidetes are found in all viral disease groups compared to healthy controls. Moreover, Bifidobacterium and SCFAs-producing bacteria families, such as Clostridia and Ruminococcus, also decreased in all disease groups. The changes are usually more pronounced as viral hepatitis progresses to cirrhosis and hepatocellular carcinoma. Based on these microbial alterations, specific treatments are tested. Fecal microbiota transplantation is tried with satisfactory results, mostly as an adjunct therapy in antiviral treatment of HBV and HCV or in patients with cirrhosis and hepatic encephalopathy. The same groups of patients are also treated with various combinations of probiotics with promising results. Attempts to strengthen the intestinal barrier by drugs or modulation of TLRs responses have not yet been tried in viral liver disease.
Author Contributions
Conceptualization, E.K. and I.T.; methodology, E.K.; software, I.T.; validation, E.K., I.T. and A.V.; formal analysis, I.T.; investigation, A.V.; resources, E.K.; data curation, I.T.; writing—original draft preparation, E.K. and A.V.; writing—review and editing, I.T.; visualization, E.K. and A.V.; supervision, E.K.; project administration, E.K.; funding acquisition, Funding not available. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Ethical review and approval were waived for this study, due to this is a Review.
Informed Consent Statement
Not applicable.
Data Availability Statement
No new data were generated.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Cohn, R. A brief history of the portal circulation. AMA Arch. Intern. Med. 1957, 100, 848–852. [Google Scholar] [CrossRef]
- Ursell, L.K.; Metcalf, J.L.; Parfrey, L.W.; Knight, R. Defining the human microbiome. Nutr. Rev. 2012, 70, S38–S44. [Google Scholar] [CrossRef]
- Turnbaugh, P.J.; Ley, R.E.; Hamady, M.; Fraser-Liggett, C.M.; Knight, R.; Gordon, J.I. The human microbiome project. Nature 2007, 449, 804–810. [Google Scholar] [CrossRef]
- De Sordi, L.; Lourenço, M.; Debarbieux, L. The Battle Within: Interactions of Bacteriophages and Bacteria in the Gastrointestinal Tract. Cell Host Microbe. 2019, 25, 210–218. [Google Scholar] [CrossRef]
- Bhattarai, Y.; Muniz Pedrogo, D.A.; Kashyap, P.C. Irritable bowel syndrome: A gut microbiota-related disorder? Am. J. Physiol. Gastrointest. Liver Physiol. 2017, 312, G52–G62. [Google Scholar] [CrossRef]
- Catinean, A.; Neag, M.A.; Muntean, D.M.; Bocsan, I.C.; Buzoianu, A.D. An overview on the interplay between nutraceuticals and gut microbiota. PeerJ 2018, 6, e4465. [Google Scholar] [CrossRef]
- Philips, C.A.; Augustine, P.; Yerol, P.K.; Ramesh, G.N.; Ahamed, R.; Rajesh, S.; George, T.; Kumbar, S. Modulating the Intestinal Microbiota: Therapeutic Opportunities in Liver Disease. J. Clin. Transl. Hepatol. 2020, 8, 87–99. [Google Scholar] [CrossRef]
- Milosevic, I.; Vujovic, A.; Barac, A.; Djelic, M.; Korac, M.; Radovanovic Spurnic, A.; Gmizic, I.; Stevanovic, O.; Djordjevic, V.; Lekic, N.; et al. Gut-Liver Axis, Gut Microbiota, and Its Modulation in the Management of Liver Diseases: A Review of the Literature. Int. J. Mol. Sci. 2019, 20, 395. [Google Scholar] [CrossRef]
- Reyes, A.; Semenkovich, N.P.; Whiteson, K.; Rohwer, F.; Gordon, J.I. Going viral: Next-generation sequencing applied to phage populations in the human gut. Nat. Rev. Microbiol. 2012, 10, 607–617. [Google Scholar] [CrossRef]
- Ringel, Y.; Maharshak, N.; Ringel-Kulka, T.; Wolber, E.A.; Sartor, R.B.; Carroll, I.M. High throughput sequencing reveals distinct microbial populations within the mucosal and luminal niches in healthy individuals. Gut Microbes 2015, 6, 173–181. [Google Scholar] [CrossRef]
- Mariat, D.; Firmesse, O.; Levenez, F.; Guimarăes, V.; Sokol, H.; Doré, J.; Corthier, G.; Furet, J.P. The Firmicutes/Bacteroidetes ratio of the human microbiota changes with age. BMC Microbiol. 2009, 9, 123. [Google Scholar] [CrossRef]
- Claesson, M.J.; Cusack, S.; O’Sullivan, O.; Greene-Diniz, R.; de Weerd, H.; Flannery, E.; Marchesi, J.R.; Falush, D.; Dinan, T.; Fitzgerald, G.; et al. Composition, variability, and temporal stability of the intestinal microbiota of the elderly. Proc. Natl. Acad. Sci. USA 2011, 108, 4586–4591. [Google Scholar] [CrossRef]
- Jalanka-Tuovinen, J.; Salonen, A.; Nikkilä, J.; Immonen, O.; Kekkonen, R.; Lahti, L.; Palva, A.; de Vos, W.M. Intestinal microbiota in healthy adults: Temporal analysis reveals individual and common core and relation to intestinal symptoms. PLoS ONE 2011, 6, e23035. [Google Scholar] [CrossRef]
- Lin, A.; Bik, E.M.; Costello, E.K.; Dethlefsen, L.; Haque, R.; Relman, D.A.; Singh, U. Distinct distal gut microbiome diversity and composition in healthy children from Bangladesh and the United States. PLoS ONE 2013, 8, e53838. [Google Scholar] [CrossRef]
- Chong, C.W.; Ahmad, A.F.; Lim, Y.A.; Teh, C.S.; Yap, I.K.; Lee, S.C.; Chin, Y.T.; Loke, P.; Chua, K.H. Effect of ethnicity and socioeconomic variation to the gut microbiota composition among pre-adolescent in Malaysia. Sci. Rep. 2015, 5, 13338. [Google Scholar] [CrossRef]
- Zhang, J.; Guo, Z.; Xue, Z.; Sun, Z.; Zhang, M.; Wang, L.; Wang, G.; Wang, F.; Xu, J.; Cao, H.; et al. A phylo-functional core of gut microbiota in healthy young Chinese cohorts across lifestyles, geography and ethnicities. ISME J. 2015, 9, 1979–1990. [Google Scholar] [CrossRef]
- Rampelli, S.; Schnorr, S.L.; Consolandi, C.; Turroni, S.; Severgnini, M.; Peano, C.; Brigidi, P.; Crittenden, A.N.; Henry, A.G.; Candela, M. Metagenome Sequencing of the Hadza Hunter-Gatherer Gut Microbiota. Curr. Biol. 2015, 25, 1682–1693. [Google Scholar] [CrossRef]
- Segata, N. Gut Microbiome: Westernization and the Disappearance of Intestinal Diversity. Curr. Biol. 2015, 25, R611–R613. [Google Scholar] [CrossRef]
- De Filippo, C.; Cavalieri, D.; Di Paola, M.; Ramazzotti, M.; Poullet, J.B.; Massart, S.; Collini, S.; Pieraccini, G.; Lionetti, P. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc. Natl. Acad. Sci. USA 2010, 107, 14691–14696. [Google Scholar] [CrossRef]
- Raimondi, S.; Amaretti, A.; Gozzoli, C.; Simone, M.; Righini, L.; Candeliere, F.; Brun, P.; Ardizzoni, A.; Colombari, B.; Paulone, S.; et al. Longitudinal Survey of Fungi in the Human Gut: ITS Profiling, Phenotyping, and Colonization. Front. Microbiol. 2019, 10, 1575. [Google Scholar] [CrossRef]
- Biedermann, L.; Rogler, G. The intestinal microbiota: Its role in health and disease. Eur. J. Pediatr. 2015, 174, 151–167. [Google Scholar] [CrossRef]
- Hiippala, K.; Jouhten, H.; Ronkainen, A.; Hartikainen, A.; Kainulainen, V.; Jalanka, J.; Satokari, R. The Potential of Gut Commensals in Reinforcing Intestinal Barrier Function and Alleviating Inflammation. Nutrients 2018, 10, 988. [Google Scholar] [CrossRef]
- Cervantes-Barragan, L.; Chai, J.N.; Tianero, M.D.; Di Luccia, B.; Ahern, P.P.; Merriman, J.; Cortez, V.S.; Caparon, M.G.; Donia, M.S.; Gilfillan, S.; et al. Lactobacillus reuteri induces gut intraepithelial CD4+CD8αα+ T cells. Science 2017, 357, 806–810. [Google Scholar] [CrossRef]
- Sender, R.; Fuchs, S.; Milo, R. Revised Estimates for the Number of Human and Bacteria Cells in the Body. PLoS Biol. 2016, 14, e1002533. [Google Scholar] [CrossRef]
- Hollister, E.B.; Gao, C.; Versalovic, J. Compositional and functional features of the gastrointestinal microbiome and their effects on human health. Gastroenterology 2014, 146, 1449–1458. [Google Scholar] [CrossRef]
- Doré, J.; Simrén, M.; Buttle, L.; Guarner, F. Hot topics in gut microbiota. United Eur. Gastroenterol. J. 2013, 1, 311–318. [Google Scholar] [CrossRef]
- Hillman, E.T.; Lu, H.; Yao, T.; Nakatsu, C.H. Microbial Ecology along the Gastrointestinal Tract. Microbes Environ. 2017, 32, 300–313. [Google Scholar] [CrossRef]
- Adak, A.; Khan, M.R. An insight into gut microbiota and its functionalities. Cell Mol. Life Sci. 2019, 76, 473–493. [Google Scholar] [CrossRef]
- Conlon, M.A.; Bird, A.R. The impact of diet and lifestyle on gut microbiota and human health. Nutrients 2014, 7, 17–44. [Google Scholar] [CrossRef]
- Gilbert, J.A.; Blaser, M.J.; Caporaso, J.G.; Jansson, J.K.; Lynch, S.V.; Knight, R. Current understanding of the human microbiome. Nat. Med. 2018, 24, 392–400. [Google Scholar] [CrossRef]
- Pinchera, B.; Moriello, N.S.; Buonomo, A.R.; Zappulo, E.; Viceconte, G.; Villari, R.; Gentile, I. Microbiota and hepatitis C virus in the era of direct-acting antiviral agents. Microb. Pathog. 2023, 175, 105968. [Google Scholar] [CrossRef]
- Ohtani, N.; Kawada, N. Role of the Gut-Liver Axis in Liver Inflammation, Fibrosis, and Cancer: A Special Focus on the Gut Microbiota Relationship. Hepatol. Commun. 2019, 3, 456–470. [Google Scholar] [CrossRef]
- Oliphant, K.; Allen-Vercoe, E. Macronutrient metabolism by the human gut microbiome: Major fermentation by-products and their impact on host health. Microbiome 2019, 7, 91. [Google Scholar] [CrossRef]
- Albillos, A.; de Gottardi, A.; Rescigno, M. The gut-liver axis in liver disease: Pathophysiological basis for therapy. J. Hepatol. 2020, 72, 558–577. [Google Scholar] [CrossRef]
- Round, J.L.; Mazmanian, S.K. The gut microbiota shapes intestinal immune responses during health and disease. Nat. Rev. Immunol. 2009, 9, 313–323. [Google Scholar] [CrossRef]
- Wiest, R.; Albillos, A.; Trauner, M.; Bajaj, J.S.; Jalan, R. Targeting the gut-liver axis in liver disease. J. Hepatol. 2017, 67, 1084–1103. [Google Scholar] [CrossRef]
- Simbrunner, B.; Mandorfer, M.; Trauner, M.; Reiberger, T. Gut-liver axis signaling in portal hypertension. World J. Gastroenterol. 2019, 25, 5897–5917. [Google Scholar] [CrossRef]
- Guo, X.; Okpara, E.S.; Hu, W.; Yan, C.; Wang, Y.; Liang, Q.; Chiang, J.Y.L.; Han, S. Interactive Relationships between Intestinal Flora and Bile Acids. Int. J. Mol. Sci. 2022, 23, 8343. [Google Scholar] [CrossRef]
- Li, S.; Han, W.; He, Q.; Zhang, W.; Zhang, Y. Relationship between Intestinal Microflora and Hepatocellular Cancer Based on Gut-Liver Axis Theory. Contrast Media Mol. Imaging 2022, 2022, 6533628. [Google Scholar] [CrossRef]
- Tranah, T.H.; Edwards, L.A.; Schnabl, B.; Shawcross, D.L. Targeting the gut-liver-immune axis to treat cirrhosis. Gut 2021, 70, 982–994. [Google Scholar] [CrossRef]
- Odenwald, M.A.; Turner, J.R. The intestinal epithelial barrier: A therapeutic target? Nat. Rev. Gastroenterol. Hepatol. 2017, 14, 9–21. [Google Scholar] [CrossRef]
- Marchiando, A.M.; Graham, W.V.; Turner, J.R. Epithelial barriers in homeostasis and disease. Annu. Rev. Pathol. 2010, 5, 119–144. [Google Scholar] [CrossRef]
- Van Itallie, C.M.; Anderson, J.M. Architecture of tight junctions and principles of molecular composition. Semin. Cell Dev. Biol. 2014, 36, 157–165. [Google Scholar] [CrossRef]
- Luissint, A.C.; Parkos, C.A.; Nusrat, A. Inflammation and the Intestinal Barrier: Leukocyte-Epithelial Cell Interactions, Cell Junction Remodeling, and Mucosal Repair. Gastroenterology 2016, 151, 616–632. [Google Scholar] [CrossRef]
- Raleigh, D.R.; Marchiando, A.M.; Zhang, Y.; Shen, L.; Sasaki, H.; Wang, Y.; Long, M.; Turner, J.R. Tight junction-associated MARVEL proteins marveld3, tricellulin, and occludin have distinct but overlapping functions. Mol. Biol. Cell. 2010, 21, 1200–1213. [Google Scholar] [CrossRef]
- Mineta, K.; Yamamoto, Y.; Yamazaki, Y.; Tanaka, H.; Tada, Y.; Saito, K.; Tamura, A.; Igarashi, M.; Endo, T.; Takeuchi, K.; et al. Predicted expansion of the claudin multigene family. FEBS Lett. 2011, 585, 606–612. [Google Scholar] [CrossRef]
- Monteiro, A.C.; Sumagin, R.; Rankin, C.R.; Leoni, G.; Mina, M.J.; Reiter, D.M.; Stehle, T.; Dermody, T.S.; Schaefer, S.A.; Hall, R.A.; et al. JAM-A associates with ZO-2, afadin, and PDZ-GEF1 to activate Rap2c and regulate epithelial barrier function. Mol. Biol. Cell. 2013, 24, 2849–2860. [Google Scholar] [CrossRef]
- Severson, E.A.; Parkos, C.A. Mechanisms of outside-in signaling at the tight junction by junctional adhesion molecule A. Ann. N. Y. Acad. Sci. 2009, 1165, 10–18. [Google Scholar] [CrossRef]
- Mandell, K.J.; Babbin, B.A.; Nusrat, A.; Parkos, C.A. Junctional adhesion molecule 1 regulates epithelial cell morphology through effects on beta1 integrins and Rap1 activity. J. Biol. Chem. 2005, 280, 11665–11674. [Google Scholar] [CrossRef]
- Spadoni, I.; Fornasa, G.; Rescigno, M. Organ-specific protection mediated by cooperation between vascular and epithelial barriers. Nat. Rev. Immunol. 2017, 17, 761–773. [Google Scholar] [CrossRef]
- Spadoni, I.; Zagato, E.; Bertocchi, A.; Paolinelli, R.; Hot, E.; Di Sabatino, A.; Caprioli, F.; Bottiglieri, L.; Oldani, A.; Viale, G.; et al. A gut-vascular barrier controls the systemic dissemination of bacteria. Science 2015, 350, 830–834. [Google Scholar] [CrossRef]
- von Olshausen, G.; Quasdorff, M.; Bester, R.; Arzberger, S.; Ko, C.; van de Klundert, M.; Zhang, K.; Odenthal, M.; Ringelhan, M.; Niessen, C.M.; et al. Hepatitis B virus promotes β-catenin-signalling and disassembly of adherens junctions in a Src kinase dependent fashion. Oncotarget 2018, 9, 33947–33960. [Google Scholar] [CrossRef]
- Mowat, A.M.; Agace, W.W. Regional specialization within the intestinal immune system. Nat. Rev. Immunol. 2014, 14, 667–685. [Google Scholar] [CrossRef]
- Chieppa, M.; Rescigno, M.; Huang, A.Y.; Germain, R.N. Dynamic imaging of dendritic cell extension into the small bowel lumen in response to epithelial cell TLR engagement. J. Exp. Med. 2006, 203, 2841–2852. [Google Scholar] [CrossRef]
- Niess, J.H.; Brand, S.; Gu, X.; Landsman, L.; Jung, S.; McCormick, B.A.; Vyas, J.M.; Boes, M.; Ploegh, H.L.; Fox, J.G.; et al. CX3CR1-mediated dendritic cell access to the intestinal lumen and bacterial clearance. Science 2005, 307, 254–258. [Google Scholar] [CrossRef]
- Ismail, A.S.; Severson, K.M.; Vaishnava, S.; Behrendt, C.L.; Yu, X.; Benjamin, J.L.; Ruhn, K.A.; Hou, B.; DeFranco, A.L.; Yarovinsky, F.; et al. Gammadelta intraepithelial lymphocytes are essential mediators of host-microbial homeostasis at the intestinal mucosal surface. Proc. Natl. Acad. Sci. USA 2011, 108, 8743–8748. [Google Scholar] [CrossRef]
- McDonald, B.D.; Jabri, B.; Bendelac, A. Diverse developmental pathways of intestinal intraepithelial lymphocytes. Nat. Rev. Immunol. 2018, 18, 514–525. [Google Scholar] [CrossRef]
- Mazzini, E.; Massimiliano, L.; Penna, G.; Rescigno, M. Oral tolerance can be established via gap junction transfer of fed antigens from CX3CR1+ macrophages to CD103+ dendritic cells. Immunity 2014, 40, 248–261. [Google Scholar] [CrossRef]
- Brennan, P.J.; Brigl, M.; Brenner, M.B. Invariant natural killer T cells: An innate activation scheme linked to diverse effector functions. Nat. Rev. Immunol. 2013, 13, 101–117. [Google Scholar] [CrossRef]
- Kjer-Nielsen, L.; Patel, O.; Corbett, A.J.; Le Nours, J.; Meehan, B.; Liu, L.; Bhati, M.; Chen, Z.; Kostenko, L.; Reantragoon, R.; et al. MR1 presents microbial vitamin B metabolites to MAIT cells. Nature 2012, 491, 717–723. [Google Scholar] [CrossRef]
- Dias, J.; Leeansyah, E.; Sandberg, J.K. Multiple layers of heterogeneity and subset diversity in human MAIT cell responses to distinct microorganisms and to innate cytokines. Proc. Natl. Acad. Sci. USA 2017, 114, E5434–E5443. [Google Scholar] [CrossRef]
- Atarashi, K.; Tanoue, T.; Ando, M.; Kamada, N.; Nagano, Y.; Narushima, S.; Suda, W.; Imaoka, A.; Setoyama, H.; Nagamori, T.; et al. Th17 Cell Induction by Adhesion of Microbes to Intestinal Epithelial Cells. Cell 2015, 163, 367–380. [Google Scholar] [CrossRef]
- Ivanov, I.I.; Atarashi, K.; Manel, N.; Brodie, E.L.; Shima, T.; Karaoz, U.; Wei, D.; Goldfarb, K.C.; Santee, C.A.; Lynch, S.V.; et al. Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell 2009, 139, 485–498. [Google Scholar] [CrossRef]
- Sharma, A.; Rudra, D. Emerging Functions of Regulatory T Cells in Tissue Homeostasis. Front. Immunol. 2018, 9, 883. [Google Scholar] [CrossRef]
- Sandquist, I.; Kolls, J. Update on regulation and effector functions of Th17 cells. F1000Research 2018, 7, 205. [Google Scholar] [CrossRef]
- Tsilingiri, K.; Rescigno, M. Postbiotics: What else? Benef. Microbes. 2013, 4, 101–107. [Google Scholar] [CrossRef]
- Mosca, F.; Gianni, M.L.; Rescigno, M. Can Postbiotics Represent a New Strategy for NEC? Adv. Exp. Med. Biol. 2019, 1125, 37–45. [Google Scholar]
- Rivière, A.; Selak, M.; Lantin, D.; Leroy, F.; De Vuyst, L. Bifidobacteria and Butyrate-Producing Colon Bacteria: Importance and Strategies for Their Stimulation in the Human Gut. Front. Microbiol. 2016, 7, 979. [Google Scholar] [CrossRef]
- Yaku, K.; Enami, Y.; Kurajyo, C.; Matsui-Yuasa, I.; Konishi, Y.; Kojima-Yuasa, A. The enhancement of phase 2 enzyme activities by sodium butyrate in normal intestinal epithelial cells is associated with Nrf2 and p53. Mol. Cell Biochem. 2012, 370, 7–14. [Google Scholar] [CrossRef]
- Ziegler, K.; Kerimi, A.; Poquet, L.; Williamson, G. Butyric acid increases transepithelial transport of ferulic acid through upregulation of the monocarboxylate transporters SLC16A1 (MCT1) and SLC16A3 (MCT4). Arch. Biochem. Biophys. 2016, 599, 3–12. [Google Scholar] [CrossRef]
- Morrison, D.J.; Preston, T. Formation of short chain fatty acids by the gut microbiota and their impact on human metabolism. Gut Microbes 2016, 7, 189–200. [Google Scholar] [CrossRef]
- Schulthess, J.; Pandey, S.; Capitani, M.; Rue-Albrecht, K.C.; Arnold, I.; Franchini, F.; Chomka, A.; Ilott, N.E.; Johnston, D.G.W.; Pires, E.; et al. The Short Chain Fatty Acid Butyrate Imprints an Antimicrobial Program in Macrophages. Immunity 2019, 50, 432–445.e7. [Google Scholar] [CrossRef]
- Park, J.; Kim, M.; Kang, S.G.; Jannasch, A.H.; Cooper, B.; Patterson, J.; Kim, C.H. Short-chain fatty acids induce both effector and regulatory T cells by suppression of histone deacetylases and regulation of the mTOR-S6K pathway. Mucosal Immunol. 2015, 8, 80–93. [Google Scholar] [CrossRef]
- Bachem, A.; Makhlouf, C.; Binger, K.J.; de Souza, D.P.; Tull, D.; Hochheiser, K.; Whitney, P.G.; Fernandez-Ruiz, D.; Dähling, S.; Kastenmüller, W.; et al. Microbiota-Derived Short-Chain Fatty Acids Promote the Memory Potential of Antigen-Activated CD8+ T Cells. Immunity 2019, 51, 285–297.e5. [Google Scholar] [CrossRef]
- Park, J.H.; Eberl, G. Type 3 regulatory T cells at the interface of symbiosis. J. Microbiol. 2018, 56, 163–171. [Google Scholar] [CrossRef]
- Sun, S.; Luo, L.; Liang, W.; Yin, Q.; Guo, J.; Rush, A.M.; Lv, Z.; Liang, Q.; Fischbach, M.A.; Sonnenburg, J.L.; et al. Bifidobacterium alters the gut microbiota and modulates the functional metabolism of T regulatory cells in the context of immune checkpoint blockade. Proc. Natl. Acad. Sci. USA 2020, 117, 27509–27515. [Google Scholar] [CrossRef]
- Alessandri, G.; Ossiprandi, M.C.; MacSharry, J.; van Sinderen, D.; Ventura, M. Bifidobacterial Dialogue With Its Human Host and Consequent Modulation of the Immune System. Front. Immunol. 2019, 10, 2348. [Google Scholar] [CrossRef]
- Elshaghabee, F.M.; Bockelmann, W.; Meske, D.; de Vrese, M.; Walte, H.G.; Schrezenmeir, J.; Heller, K.J. Ethanol Production by Selected Intestinal Microorganisms and Lactic Acid Bacteria Growing under Different Nutritional Conditions. Front. Microbiol. 2016, 7, 47. [Google Scholar] [CrossRef]
- Zhu, L.; Baker, S.S.; Gill, C.; Liu, W.; Alkhouri, R.; Baker, R.D.; Gill, S.R. Characterization of gut microbiomes in nonalcoholic steatohepatitis (NASH) patients: A connection between endogenous alcohol and NASH. Hepatology 2013, 57, 601–609. [Google Scholar] [CrossRef]
- Barreau, F.; Hugot, J.P. Intestinal barrier dysfunction triggered by invasive bacteria. Curr. Opin. Microbiol. 2014, 17, 91–98. [Google Scholar] [CrossRef]
- Rakoff-Nahoum, S.; Paglino, J.; Eslami-Varzaneh, F.; Edberg, S.; Medzhitov, R. Recognition of commensal microflora by toll-like receptors is required for intestinal homeostasis. Cell 2004, 118, 229–241. [Google Scholar] [CrossRef]
- Cario, E.; Gerken, G.; Podolsky, D.K. Toll-like receptor 2 enhances ZO-1-associated intestinal epithelial barrier integrity via protein kinase C. Gastroenterology 2004, 127, 224–238. [Google Scholar] [CrossRef]
- Li, X.; Wang, C.; Nie, J.; Lv, D.; Wang, T.; Xu, Y. Toll-like receptor 4 increases intestinal permeability through up-regulation of membrane PKC activity in alcoholic steatohepatitis. Alcohol 2013, 47, 459–465. [Google Scholar] [CrossRef]
- Sharma, R.; Young, C.; Neu, J. Molecular modulation of intestinal epithelial barrier: Contribution of microbiota. J. Biomed. Biotechnol. 2010, 2010, 305879. [Google Scholar] [CrossRef]
- Takeuchi, O.; Akira, S. Pattern recognition receptors and inflammation. Cell 2010, 140, 805–820. [Google Scholar] [CrossRef]
- Ignacio, A.; Morales, C.I.; Câmara, N.O.; Almeida, R.R. Innate Sensing of the Gut Microbiota: Modulation of Inflammatory and Autoimmune Diseases. Front. Immunol. 2016, 7, 54. [Google Scholar] [CrossRef]
- Wu, J.; Meng, Z.; Jiang, M.; Zhang, E.; Trippler, M.; Broering, R.; Bucchi, A.; Krux, F.; Dittmer, U.; Yang, D.; et al. Toll-like receptor-induced innate immune responses in non-parenchymal liver cells are cell type-specific. Immunology 2010, 129, 363–374. [Google Scholar] [CrossRef]
- Seki, E.; Brenner, D.A. Toll-like receptors and adaptor molecules in liver disease: Update. Hepatology 2008, 48, 322–335. [Google Scholar] [CrossRef]
- Miyake, Y.; Yamamoto, K. Role of gut microbiota in liver diseases. Hepatol. Res. 2013, 43, 139–146. [Google Scholar] [CrossRef]
- Seki, E.; Tsutsui, H.; Nakano, H.; Tsuji, N.; Hoshino, K.; Adachi, O.; Adachi, K.; Futatsugi, S.; Kuida, K.; Takeuchi, O.; et al. Lipopolysaccharide-induced IL-18 secretion from murine Kupffer cells independently of myeloid differentiation factor 88 that is critically involved in induction of production of IL-12 and IL-1beta. J. Immunol. 2001, 166, 2651–2657. [Google Scholar] [CrossRef]
- Rogier, R.; Koenders, M.I.; Abdollahi-Roodsaz, S. Toll-like receptor mediated modulation of T cell response by commensal intestinal microbiota as a trigger for autoimmune arthritis. J. Immunol. Res. 2015, 2015, 527696. [Google Scholar] [CrossRef]
- Pasare, C.; Medzhitov, R. Toll-like receptors and acquired immunity. Semin. Immunol. 2004, 16, 23–26. [Google Scholar] [CrossRef]
- Bonnert, T.P.; Garka, K.E.; Parnet, P.; Sonoda, G.; Testa, J.R.; Sims, J.E. The cloning and characterization of human MyD88: A member of an IL-1 receptor related family. FEBS Lett. 1997, 402, 81–84. [Google Scholar] [CrossRef]
- Yamamoto, M.; Takeda, K. Current views of toll-like receptor signaling pathways. Gastroenterol. Res. Pract. 2010, 2010, 240365. [Google Scholar] [CrossRef]
- Agrawal, S.; Agrawal, A.; Doughty, B.; Gerwitz, A.; Blenis, J.; Van Dyke, T.; Pulendran, B. Cutting edge: Different Toll-like receptor agonists instruct dendritic cells to induce distinct Th responses via differential modulation of extracellular signal-regulated kinase-mitogen-activated protein kinase and c-Fos. J. Immunol. 2003, 171, 4984–4989. [Google Scholar] [CrossRef]
- Kattah, M.G.; Wong, M.T.; Yocum, M.D.; Utz, P.J. Cytokines secreted in response to Toll-like receptor ligand stimulation modulate differentiation of human Th17 cells. Arthritis Rheum. 2008, 58, 1619–1629. [Google Scholar] [CrossRef]
- Dillon, S.; Agrawal, A.; Van Dyke, T.; Landreth, G.; McCauley, L.; Koh, A.; Maliszewski, C.; Akira, S.; Pulendran, B. A Toll-like receptor 2 ligand stimulates Th2 responses in vivo, via induction of extracellular signal-regulated kinase mitogen-activated protein kinase and c-Fos in dendritic cells. J. Immunol. 2004, 172, 4733–4743. [Google Scholar] [CrossRef]
- Chow, J.C.; Young, D.W.; Golenbock, D.T.; Christ, W.J.; Gusovsky, F. Toll-like receptor-4 mediates lipopolysaccharide-induced signal transduction. J. Biol. Chem. 1999, 274, 10689–10692. [Google Scholar] [CrossRef]
- Bauer, S.; Kirschning, C.J.; Häcker, H.; Redecke, V.; Hausmann, S.; Akira, S.; Wagner, H.; Lipford, G.B. Human TLR9 confers responsiveness to bacterial DNA via species-specific CpG motif recognition. Proc. Natl. Acad. Sci. USA 2001, 98, 9237–9242. [Google Scholar] [CrossRef]
- Dolganiuc, A.; Oak, S.; Kodys, K.; Golenbock, D.T.; Finberg, R.W.; Kurt-Jones, E.; Szabo, G. Hepatitis C core and nonstructural 3 proteins trigger toll-like receptor 2-mediated pathways and inflammatory activation. Gastroenterology 2004, 127, 1513–1524. [Google Scholar] [CrossRef]
- Wang, B.; Trippler, M.; Pei, R.; Lu, M.; Broering, R.; Gerken, G.; Schlaak, J.F. Toll-like receptor activated human and murine hepatic stellate cells are potent regulators of hepatitis C virus replication. J. Hepatol. 2009, 51, 1037–1045. [Google Scholar] [CrossRef]
- Seki, E.; De Minicis, S.; Osterreicher, C.H.; Kluwe, J.; Osawa, Y.; Brenner, D.A.; Schwabe, R.F. TLR4 enhances TGF-beta signaling and hepatic fibrosis. Nat. Med. 2007, 13, 1324–1332. [Google Scholar] [CrossRef]
- Isayama, F.; Hines, I.N.; Kremer, M.; Milton, R.J.; Byrd, C.L.; Perry, A.W.; McKim, S.E.; Parsons, C.; Rippe, R.A.; Wheeler, M.D. LPS signaling enhances hepatic fibrogenesis caused by experimental cholestasis in mice. Am. J. Physiol. Gastrointest. Liver Physiol. 2006, 290, G1318–G1328. [Google Scholar] [CrossRef]
- Gäbele, E.; Mühlbauer, M.; Dorn, C.; Weiss, T.S.; Froh, M.; Schnabl, B.; Wiest, R.; Schölmerich, J.; Obermeier, F.; Hellerbrand, C. Role of TLR9 in hepatic stellate cells and experimental liver fibrosis. Biochem. Biophys. Res. Commun. 2008, 376, 271–276. [Google Scholar] [CrossRef]
- Hartmann, P.; Haimerl, M.; Mazagova, M.; Brenner, D.A.; Schnabl, B. Toll-like receptor 2-mediated intestinal injury and enteric tumor necrosis factor receptor I contribute to liver fibrosis in mice. Gastroenterology 2012, 143, 1330–1340.e1. [Google Scholar] [CrossRef]
- Seki, E.; Schnabl, B. Role of innate immunity and the microbiota in liver fibrosis: Crosstalk between the liver and gut. J. Physiol. 2012, 590, 447–458. [Google Scholar] [CrossRef]
- Czaja, A.J. Factoring the intestinal microbiome into the pathogenesis of autoimmune hepatitis. World J. Gastroenterol. 2016, 22, 9257–9278. [Google Scholar] [CrossRef]
- Chopyk, D.M.; Grakoui, A. Contribution of the Intestinal Microbiome and Gut Barrier to Hepatic Disorders. Gastroenterology 2020, 159, 849–863. [Google Scholar] [CrossRef]
- Agus, A.; Planchais, J.; Sokol, H. Gut Microbiota Regulation of Tryptophan Metabolism in Health and Disease. Cell Host Microbe. 2018, 23, 716–724. [Google Scholar] [CrossRef]
- Zelante, T.; Iannitti, R.G.; Cunha, C.; De Luca, A.; Giovannini, G.; Pieraccini, G.; Zecchi, R.; D’Angelo, C.; Massi-Benedetti, C.; Fallarino, F.; et al. Tryptophan catabolites from microbiota engage aryl hydrocarbon receptor and balance mucosal reactivity via interleukin-22. Immunity 2013, 39, 372–385. [Google Scholar] [CrossRef]
- Ma, Z.; Cao, Q.; Xiong, Y.; Zhang, E.; Lu, M. Interaction between Hepatitis B Virus and Toll-Like Receptors: Current Status and Potential Therapeutic Use for Chronic Hepatitis B. Vaccines 2018, 6, 6. [Google Scholar] [CrossRef]
- Ashfaq, U.A.; Iqbal, M.S.; Khaliq, S. Role of Toll-Like Receptors in Hepatitis C Virus Pathogenesis and Treatment. Crit. Rev. Eukaryot. Gene Expr. 2016, 26, 353–362. [Google Scholar] [CrossRef]
- Chen, B.; Chen, H.; Shu, X.; Yin, Y.; Li, J.; Qin, J.; Chen, L.; Peng, K.; Xu, F.; Gu, W.; et al. Presence of Segmented Filamentous Bacteria in Human Children and Its Potential Role in the Modulation of Human Gut Immunity. Front. Microbiol. 2018, 9, 1403. [Google Scholar] [CrossRef]
- Blacher, E.; Levy, M.; Tatirovsky, E.; Elinav, E. Microbiome-Modulated Metabolites at the Interface of Host Immunity. J. Immunol. 2017, 198, 572–580. [Google Scholar] [CrossRef]
- Levy, M.; Blacher, E.; Elinav, E. Microbiome, metabolites and host immunity. Curr. Opin. Microbiol. 2017, 35, 8–15. [Google Scholar] [CrossRef]
- Johansson, M.E. Fast renewal of the distal colonic mucus layers by the surface goblet cells as measured by in vivo labeling of mucin glycoproteins. PLoS ONE 2012, 7, e41009. [Google Scholar] [CrossRef]
- Bergström, J.H.; Birchenough, G.M.; Katona, G.; Schroeder, B.O.; Schütte, A.; Ermund, A.; Johansson, M.E.; Hansson, G.C. Gram-positive bacteria are held at a distance in the colon mucus by the lectin-like protein ZG16. Proc. Natl. Acad. Sci. USA 2016, 113, 13833–13838. [Google Scholar] [CrossRef]
- Johansson, M.E.; Ambort, D.; Pelaseyed, T.; Schütte, A.; Gustafsson, J.K.; Ermund, A.; Subramani, D.B.; Holmén-Larsson, J.M.; Thomsson, K.A.; Bergström, J.H.; et al. Composition and functional role of the mucus layers in the intestine. Cell Mol. Life Sci. 2011, 68, 3635–3641. [Google Scholar] [CrossRef]
- Gibbins, H.L.; Proctor, G.B.; Yakubov, G.E.; Wilson, S.; Carpenter, G.H. SIgA binding to mucosal surfaces is mediated by mucin-mucin interactions. PLoS ONE 2015, 10, e0119677. [Google Scholar] [CrossRef]
- Jakobsson, H.E.; Rodríguez-Piñeiro, A.M.; Schütte, A.; Ermund, A.; Boysen, P.; Bemark, M.; Sommer, F.; Bäckhed, F.; Hansson, G.C.; Johansson, M.E. The composition of the gut microbiota shapes the colon mucus barrier. EMBO Rep. 2015, 16, 164–177. [Google Scholar] [CrossRef]
- Birchenough, G.M.; Nyström, E.E.; Johansson, M.E.; Hansson, G.C. A sentinel goblet cell guards the colonic crypt by triggering Nlrp6-dependent Muc2 secretion. Science 2016, 352, 1535–1542. [Google Scholar] [CrossRef] [PubMed]
- Derrien, M.; Van Baarlen, P.; Hooiveld, G.; Norin, E.; Müller, M.; de Vos, W.M. Modulation of Mucosal Immune Response, Tolerance, and Proliferation in Mice Colonized by the Mucin-Degrader Akkermansia muciniphila. Front. Microbiol. 2011, 2, 166. [Google Scholar] [CrossRef] [PubMed]
- Abreu, M.T. Toll-like receptor signalling in the intestinal epithelium: How bacterial recognition shapes intestinal function. Nat. Rev. Immunol. 2010, 10, 131–144. [Google Scholar] [CrossRef] [PubMed]
- Gallo, R.L.; Hooper, L.V. Epithelial antimicrobial defence of the skin and intestine. Nat. Rev. Immunol. 2012, 12, 503–516. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, K.; Sakuragi, N.; Takakuwa, A.; Ayabe, T. Paneth cell α-defensins and enteric microbiota in health and disease. Biosci. Microbiota Food Health 2016, 35, 57–67. [Google Scholar] [CrossRef]
- Mantis, N.J.; Rol, N.; Corthésy, B. Secretory IgA’s complex roles in immunity and mucosal homeostasis in the gut. Mucosal Immunol. 2011, 4, 603–611. [Google Scholar] [CrossRef]
- Donaldson, G.P.; Ladinsky, M.S.; Yu, K.B.; Sanders, J.G.; Yoo, B.B.; Chou, W.C.; Conner, M.E.; Earl, A.M.; Knight, R.; Bjorkman, P.J.; et al. Gut microbiota utilize immunoglobulin A for mucosal colonization. Science 2018, 360, 795–800. [Google Scholar] [CrossRef]
- Macpherson, A.J.; Geuking, M.B.; McCoy, K.D. Homeland security: IgA immunity at the frontiers of the body. Trends Immunol. 2012, 33, 160–167. [Google Scholar] [CrossRef]
- Chairatana, P.; Nolan, E.M. Defensins, lectins, mucins, and secretory immunoglobulin A: Microbe-binding biomolecules that contribute to mucosal immunity in the human gut. Crit. Rev. Biochem. Mol. Biol. 2017, 52, 45–56. [Google Scholar] [CrossRef]
- Mukherjee, S.; Hooper, L.V. Antimicrobial defense of the intestine. Immunity 2015, 42, 28–39. [Google Scholar] [CrossRef]
- Leclercq, S.; Cani, P.D.; Neyrinck, A.M.; Stärkel, P.; Jamar, F.; Mikolajczak, M.; Delzenne, N.M.; de Timary, P. Role of intestinal permeability and inflammation in the biological and behavioral control of alcohol-dependent subjects. Brain Behav. Immun. 2012, 26, 911–918. [Google Scholar] [CrossRef] [PubMed]
- Cresci, G.A.; Glueck, B.; McMullen, M.R.; Xin, W.; Allende, D.; Nagy, L.E. Prophylactic tributyrin treatment mitigates chronic-binge ethanol-induced intestinal barrier and liver injury. J. Gastroenterol. Hepatol. 2017, 32, 1587–1597. [Google Scholar] [CrossRef]
- Boursi, B.; Mamtani, R.; Haynes, K.; Yang, Y.X. The effect of past antibiotic exposure on diabetes risk. Eur. J. Endocrinol. 2015, 172, 639–648. [Google Scholar] [CrossRef] [PubMed]
- Kozyrskyj, A.L.; Ernst, P.; Becker, A.B. Increased risk of childhood asthma from antibiotic use in early life. Chest 2007, 131, 1753–1759. [Google Scholar] [CrossRef] [PubMed]
- Francino, M.P. Antibiotics and the Human Gut Microbiome: Dysbioses and Accumulation of Resistances. Front. Microbiol. 2016, 6, 1543. [Google Scholar] [CrossRef] [PubMed]
- Mårild, K.; Ye, W.; Lebwohl, B.; Green, P.H.; Blaser, M.J.; Card, T.; Ludvigsson, J.F. Antibiotic exposure and the development of coeliac disease: A nationwide case-control study. BMC Gastroenterol. 2013, 13, 109. [Google Scholar] [CrossRef] [PubMed]
- Petersen, C.; Round, J.L. Defining dysbiosis and its influence on host immunity and disease. Cell Microbiol. 2014, 16, 1024–1033. [Google Scholar] [CrossRef]
- Csak, T.; Ganz, M.; Pespisa, J.; Kodys, K.; Dolganiuc, A.; Szabo, G. Fatty acid and endotoxin activate inflammasomes in mouse hepatocytes that release danger signals to stimulate immune cells. Hepatology 2011, 54, 133–144. [Google Scholar] [CrossRef]
- Uesugi, T.; Froh, M.; Arteel, G.E.; Bradford, B.U.; Thurman, R.G. Toll-like receptor 4 is involved in the mechanism of early alcohol-induced liver injury in mice. Hepatology 2001, 34, 101–108. [Google Scholar] [CrossRef]
- Anand, G.; Zarrinpar, A.; Loomba, R. Targeting Dysbiosis for the Treatment of Liver Disease. Semin. Liver Dis. 2016, 36, 37–47. [Google Scholar] [CrossRef]
- Stärkel, P.; Schnabl, B. Bidirectional Communication between Liver and Gut during Alcoholic Liver Disease. Semin. Liver Dis. 2016, 36, 331–339. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, A.; Debelius, J.; Brenner, D.A.; Karin, M.; Loomba, R.; Schnabl, B.; Knight, R. The gut-liver axis and the intersection with the microbiome. Nat. Rev. Gastroenterol. Hepatol. 2018, 15, 397–411. [Google Scholar] [CrossRef] [PubMed]
- Wiest, R.; Garcia-Tsao, G. Bacterial translocation (BT) in cirrhosis. Hepatology 2005, 41, 422–433. [Google Scholar] [CrossRef] [PubMed]
- Mehal, W.Z. The Gordian Knot of dysbiosis, obesity and NAFLD. Nat. Rev. Gastroenterol. Hepatol. 2013, 10, 637–644. [Google Scholar] [CrossRef] [PubMed]
- Bourriaud, C.; Robins, R.J.; Martin, L.; Kozlowski, F.; Tenailleau, E.; Cherbut, C.; Michel, C. Lactate is mainly fermented to butyrate by human intestinal microfloras but inter-individual variation is evident. J. Appl. Microbiol. 2005, 99, 201–212. [Google Scholar] [CrossRef] [PubMed]
- Wahlström, A.; Sayin, S.I.; Marschall, H.U.; Bäckhed, F. Intestinal Crosstalk between Bile Acids and Microbiota and Its Impact on Host Metabolism. Cell Metab. 2016, 24, 41–50. [Google Scholar] [CrossRef]
- de Aguiar Vallim, T.Q.; Tarling, E.J.; Edwards, P.A. Pleiotropic roles of bile acids in metabolism. Cell Metab. 2013, 17, 657–669. [Google Scholar] [CrossRef]
- Staley, C.; Weingarden, A.R.; Khoruts, A.; Sadowsky, M.J. Interaction of gut microbiota with bile acid metabolism and its influence on disease states. Appl. Microbiol. Biotechnol. 2017, 101, 47–64. [Google Scholar] [CrossRef]
- Lachar, J.; Bajaj, J.S. Changes in the Microbiome in Cirrhosis and Relationship to Complications: Hepatic Encephalopathy, Spontaneous Bacterial Peritonitis, and Sepsis. Semin. Liver Dis. 2016, 36, 327–330. [Google Scholar] [CrossRef]
- Chiang, J.Y.L.; Ferrell, J.M. Bile acid receptors FXR and TGR5 signaling in fatty liver diseases and therapy. Am. J. Physiol. Gastrointest. Liver Physiol. 2020, 318, G554–G573. [Google Scholar] [CrossRef]
- Copple, B.L.; Li, T. Pharmacology of bile acid receptors: Evolution of bile acids from simple detergents to complex signaling molecules. Pharmacol. Res. 2016, 104, 9–21. [Google Scholar] [CrossRef] [PubMed]
- Stofan, M.; Guo, G.L. Bile Acids and FXR: Novel Targets for Liver Diseases. Front. Med. 2020, 7, 544. [Google Scholar] [CrossRef]
- Zhu, F.; Zheng, S.; Zhao, M.; Shi, F.; Zheng, L.; Wang, H. The regulatory role of bile acid microbiota in the progression of liver cirrhosis. Front. Pharmacol. 2023, 14, 1214685. [Google Scholar] [CrossRef]
- Zarrinpar, A.; Loomba, R. Review article: The emerging interplay among the gastrointestinal tract, bile acids and incretins in the pathogenesis of diabetes and non-alcoholic fatty liver disease. Aliment. Pharmacol. Ther. 2012, 36, 909–921. [Google Scholar] [CrossRef]
- Sayin, S.I.; Wahlström, A.; Felin, J.; Jäntti, S.; Marschall, H.U.; Bamberg, K.; Angelin, B.; Hyötyläinen, T.; Orešič, M.; Bäckhed, F. Gut microbiota regulates bile acid metabolism by reducing the levels of tauro-beta-muricholic acid, a naturally occurring FXR antagonist. Cell Metab. 2013, 17, 225–235. [Google Scholar] [CrossRef]
- Inagaki, T.; Moschetta, A.; Lee, Y.K.; Peng, L.; Zhao, G.; Downes, M.; Yu, R.T.; Shelton, J.M.; Richardson, J.A.; Repa, J.J.; et al. Regulation of antibacterial defense in the small intestine by the nuclear bile acid receptor. Proc. Natl. Acad. Sci. USA 2006, 103, 3920–3925. [Google Scholar] [CrossRef]
- Parséus, A.; Sommer, N.; Sommer, F.; Caesar, R.; Molinaro, A.; Ståhlman, M.; Greiner, T.U.; Perkins, R.; Bäckhed, F. Microbiota-induced obesity requires farnesoid X receptor. Gut 2017, 66, 429–437. [Google Scholar] [CrossRef] [PubMed]
- Gadaleta, R.M.; van Erpecum, K.J.; Oldenburg, B.; Willemsen, E.C.; Renooij, W.; Murzilli, S.; Klomp, L.W.; Siersema, P.D.; Schipper, M.E.; Danese, S.; et al. Farnesoid X receptor activation inhibits inflammation and preserves the intestinal barrier in inflammatory bowel disease. Gut 2011, 60, 463–472. [Google Scholar] [CrossRef]
- Mouries, J.; Brescia, P.; Silvestri, A.; Spadoni, I.; Sorribas, M.; Wiest, R.; Mileti, E.; Galbiati, M.; Invernizzi, P.; Adorini, L.; et al. Microbiota-driven gut vascular barrier disruption is a prerequisite for non-alcoholic steatohepatitis development. J. Hepatol. 2019, 71, 1216–1228. [Google Scholar] [CrossRef]
- Pols, T.W.; Noriega, L.G.; Nomura, M.; Auwerx, J.; Schoonjans, K. The bile acid membrane receptor TGR5 as an emerging target in metabolism and inflammation. J. Hepatol. 2011, 54, 1263–1272. [Google Scholar] [CrossRef] [PubMed]
- Broeders, E.P.; Nascimento, E.B.; Havekes, B.; Brans, B.; Roumans, K.H.; Tailleux, A.; Schaart, G.; Kouach, M.; Charton, J.; Deprez, B.; et al. The Bile Acid Chenodeoxycholic Acid Increases Human Brown Adipose Tissue Activity. Cell Metab. 2015, 22, 418–426. [Google Scholar] [CrossRef]
- Thomas, C.; Gioiello, A.; Noriega, L.; Strehle, A.; Oury, J.; Rizzo, G.; Macchiarulo, A.; Yamamoto, H.; Mataki, C.; Pruzanski, M.; et al. TGR5-mediated bile acid sensing controls glucose homeostasis. Cell Metab. 2009, 10, 167–177. [Google Scholar] [CrossRef]
- Hsu, Y.C.; Huang, D.Q.; Nguyen, M.H. Global burden of hepatitis B virus: Current status, missed opportunities and a call for action. Nat. Rev. Gastroenterol. Hepatol. 2023, 20, 524–537. [Google Scholar] [CrossRef]
- Iannacone, M.; Guidotti, L.G. Immunobiology and pathogenesis of hepatitis B virus infection. Nat. Rev. Immunol. 2022, 22, 19–32. [Google Scholar] [CrossRef]
- Chen, Z.; Xie, Y.; Zhou, F.; Zhang, B.; Wu, J.; Yang, L.; Xu, S.; Stedtfeld, R.; Chen, Q.; Liu, J.; et al. Featured Gut Microbiomes Associated With the Progression of Chronic Hepatitis B Disease. Front. Microbiol. 2020, 11, 383. [Google Scholar] [CrossRef]
- Zhu, Q.; Xia, P.; Zhou, X.; Li, X.; Guo, W.; Zhu, B.; Zheng, X.; Wang, B.; Yang, D.; Wang, J. Hepatitis B Virus Infection Alters Gut Microbiota Composition in Mice. Front. Cell Infect. Microbiol. 2019, 9, 377, Erratum in Front. Cell Infect. Microbiol. 2020, 10, 490. [Google Scholar] [CrossRef]
- Li, X.; Wu, S.; Du, Y.; Yang, L.; Li, Y.; Hong, B. Entecavir therapy reverses gut microbiota dysbiosis induced by hepatitis B virus infection in a mouse model. Int. J. Antimicrob. Agents. 2020, 56, 106000. [Google Scholar] [CrossRef]
- Wang, J.; Wang, Y.; Zhang, X.; Liu, J.; Zhang, Q.; Zhao, Y.; Peng, J.; Feng, Q.; Dai, J.; Sun, S.; et al. Gut Microbial Dysbiosis Is Associated with Altered Hepatic Functions and Serum Metabolites in Chronic Hepatitis B Patients. Front. Microbiol. 2017, 8, 2222. [Google Scholar] [CrossRef]
- Chen, Y.; Yang, F.; Lu, H.; Wang, B.; Chen, Y.; Lei, D.; Wang, Y.; Zhu, B.; Li, L. Characterization of fecal microbial communities in patients with liver cirrhosis. Hepatology 2011, 54, 562–572. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Chen, L.; Wang, H.; Cai, W.; Xie, Q. Modulation of bile acid profile by gut microbiota in chronic hepatitis B. J. Cell Mol. Med. 2020, 24, 2573–2581. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Huang, C.; Shi, Y.; Wang, R.; Fan, J.; Yu, Y.; Zhang, Z.; Zhu, K.; Li, M.; Ni, Q.; et al. Distinct Bile Acid Profiles in Patients With Chronic Hepatitis B Virus Infection Reveal Metabolic Interplay Between Host, Virus and Gut Microbiome. Front. Med. 2021, 8, 708495. [Google Scholar] [CrossRef] [PubMed]
- Joo, E.J.; Cheong, H.S.; Kwon, M.J.; Sohn, W.; Kim, H.N.; Cho, Y.K. Relationship between gut microbiome diversity and hepatitis B viral load in patients with chronic hepatitis B. Gut Pathog. 2021, 13, 65. [Google Scholar] [CrossRef] [PubMed]
- Sandler, N.G.; Koh, C.; Roque, A.; Eccleston, J.L.; Siegel, R.B.; Demino, M.; Kleiner, D.E.; Deeks, S.G.; Liang, T.J.; Heller, T.; et al. Host response to translocated microbial products predicts outcomes of patients with HBV or HCV infection. Gastroenterology 2011, 141, 1220–1230.e1–3. [Google Scholar] [CrossRef] [PubMed]
- Yun, Y.; Chang, Y.; Kim, H.N.; Ryu, S.; Kwon, M.J.; Cho, Y.K.; Kim, H.L.; Cheong, H.S.; Joo, E.J. Alterations of the Gut Microbiome in Chronic Hepatitis B Virus Infection Associated with Alanine Aminotransferase Level. J. Clin. Med. 2019, 8, 173. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhao, R.; Shi, D.; Sun, S.; Ren, H.; Zhao, H.; Wu, W.; Jin, L.; Sheng, J.; Shi, Y. Characterization of the circulating microbiome in acute-on-chronic liver failure associated with hepatitis B. Liver Int. 2019, 39, 1207–1216. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Y.; Chen, S.; Fu, Y.; Wu, W.; Chen, T.; Chen, J.; Yang, B.; Ou, Q. Gut microbiota dysbiosis in patients with hepatitis B virus-induced chronic liver disease covering chronic hepatitis, liver cirrhosis and hepatocellular carcinoma. J. Viral Hepat. 2020, 27, 143–155. [Google Scholar] [CrossRef] [PubMed]
- Lin, M.J.; Su, T.H.; Chen, C.C.; Wu, W.K.; Hsu, S.J.; Tseng, T.C.; Liao, S.H.; Hong, C.M.; Yang, H.C.; Liu, C.J.; et al. Diversity and composition of gut microbiota in healthy individuals and patients at different stages of hepatitis B virus-related liver disease. Gut Pathog. 2023, 15, 24. [Google Scholar] [CrossRef] [PubMed]
- Bailey, M.A.; Holscher, H.D. Microbiome-Mediated Effects of the Mediterranean Diet on Inflammation. Adv. Nutr. 2018, 9, 193–206. [Google Scholar] [CrossRef]
- Liu, Q.; Li, F.; Zhuang, Y.; Xu, J.; Wang, J.; Mao, X.; Zhang, Y.; Liu, X. Alteration in gut microbiota associated with hepatitis B and non-hepatitis virus related hepatocellular carcinoma. Gut Pathog. 2019, 11, 1. [Google Scholar] [CrossRef]
- Qin, N.; Yang, F.; Li, A.; Prifti, E.; Chen, Y.; Shao, L.; Guo, J.; Le Chatelier, E.; Yao, J.; Wu, L.; et al. Alterations of the human gut microbiome in liver cirrhosis. Nature 2014, 513, 59–64. [Google Scholar] [CrossRef]
- Liu, Y.; Li, J.; Jin, Y.; Zhao, L.; Zhao, F.; Feng, J.; Li, A.; Wei, Y. Splenectomy Leads to Amelioration of Altered Gut Microbiota and Metabolome in Liver Cirrhosis Patients. Front. Microbiol. 2018, 9, 963. [Google Scholar] [CrossRef]
- Wei, X.; Yan, X.; Zou, D.; Yang, Z.; Wang, X.; Liu, W.; Wang, S.; Li, X.; Han, J.; Huang, L.; et al. Abnormal fecal microbiota community and functions in patients with hepatitis B liver cirrhosis as revealed by a metagenomic approach. BMC Gastroenterol. 2013, 13, 175. [Google Scholar] [CrossRef]
- Lu, H.; Wu, Z.; Xu, W.; Yang, J.; Chen, Y.; Li, L. Intestinal microbiota was assessed in cirrhotic patients with hepatitis B virus infection. Intestinal microbiota of HBV cirrhotic patients. Microb. Ecol. 2011, 61, 693–703. [Google Scholar] [CrossRef]
- Calgin, M.K.; Cetinkol, Y. Decreased levels of serum zonulin and copeptin in chronic Hepatitis-B patients. Pak. J. Med. Sci. 2019, 35, 847–851. [Google Scholar] [CrossRef]
- Wang, X.; Li, M.M.; Niu, Y.; Zhang, X.; Yin, J.B.; Zhao, C.J.; Wang, R.T. Serum Zonulin in HBV-Associated Chronic Hepatitis, Liver Cirrhosis, and Hepatocellular Carcinoma. Dis. Markers 2019, 2019, 5945721. [Google Scholar] [CrossRef]
- Zheng, D.; Liwinski, T.; Elinav, E. Interaction between microbiota and immunity in health and disease. Cell Res. 2020, 30, 492–506. [Google Scholar] [CrossRef]
- Yang, X.A.; Lv, F.; Wang, R.; Chang, Y.; Zhao, Y.; Cui, X.; Li, H.; Yang, S.; Li, S.; Zhao, X.; et al. Potential role of intestinal microflora in disease progression among patients with different stages of Hepatitis B. Gut Pathog. 2020, 12, 50. [Google Scholar] [CrossRef]
- Ren, Z.; Li, A.; Jiang, J.; Zhou, L.; Yu, Z.; Lu, H.; Xie, H.; Chen, X.; Shao, L.; Zhang, R.; et al. Gut microbiome analysis as a tool towards targeted non-invasive biomarkers for early hepatocellular carcinoma. Gut 2019, 68, 1014–1023. [Google Scholar] [CrossRef]
- Zhao, Y.; Mao, Y.F.; Tang, Y.S.; Ni, M.Z.; Liu, Q.H.; Wang, Y.; Feng, Q.; Peng, J.H.; Hu, Y.Y. Altered oral microbiota in chronic hepatitis B patients with different tongue coatings. World J. Gastroenterol. 2018, 24, 3448–3461. [Google Scholar] [CrossRef]
- Li, R.; Yi, X.; Yang, J.; Zhu, Z.; Wang, Y.; Liu, X.; Huang, X.; Wan, Y.; Fu, X.; Shu, W.; et al. Gut Microbiome Signatures in the Progression of Hepatitis B Virus-Induced Liver Disease. Front. Microbiol. 2022, 13, 916061. [Google Scholar] [CrossRef]
- Lu, Y.X.; He, C.Z.; Wang, Y.X.; Ai, Z.S.; Liang, P.; Yang, C.Q. Effect of Entecavir on the Intestinal Microflora in Patients with Chronic Hepatitis B: A Controlled Cross-Sectional and Longitudinal Real-World Study. Infect. Dis. Ther. 2021, 10, 241–252. [Google Scholar] [CrossRef]
- Chen, B.; Huang, H.; Pan, C.Q. The role of gut microbiota in hepatitis B disease progression and treatment. J. Viral Hepat. 2022, 29, 94–106. [Google Scholar] [CrossRef]
- Li, Y.N.; Kang, N.L.; Jiang, J.J.; Zhu, Y.Y.; Liu, Y.R.; Zeng, D.W.; Wang, F. Gut microbiota of hepatitis B virus-infected patients in the immune-tolerant and immune-active phases and their implications in metabolite changes. World J. Gastroenterol. 2022, 28, 5188–5202. [Google Scholar] [CrossRef]
- Shu, W.; Shanjian, C.; Jinpiao, L.; Qishui, O. Gut microbiota dysbiosis in patients with hepatitis B virus-related cirrhosis. Ann. Hepatol. 2022, 27, 100676. [Google Scholar] [CrossRef]
- Gao, K.; Liu, L.; Wang, H. Advances in immunomodulation of microbial unmethylated CpG DNA on animal intestinal tract A review. Wei Sheng Wu Xue Bao 2015, 55, 543–550. [Google Scholar]
- Yang, R.; Xu, Y.; Dai, Z.; Lin, X.; Wang, H. The Immunologic Role of Gut Microbiota in Patients with Chronic HBV Infection. J. Immunol. Res. 2018, 2018, 2361963. [Google Scholar] [CrossRef]
- Yan, F.; Zhang, Q.; Shi, K.; Zhang, Y.; Zhu, B.; Bi, Y.; Wang, X. Gut microbiota dysbiosis with hepatitis B virus liver disease and association with immune response. Front. Cell Infect. Microbiol. 2023, 13, 1152987. [Google Scholar] [CrossRef]
- Xu, D.; Huang, Y.; Wang, J. Gut microbiota modulate the immune effect against hepatitis B virus infection. Eur. J. Clin. Microbiol. Infect. Dis. 2015, 34, 2139–2147. [Google Scholar] [CrossRef]
- Chou, H.H.; Chien, W.H.; Wu, L.L.; Cheng, C.H.; Chung, C.H.; Horng, J.H.; Ni, Y.H.; Tseng, H.T.; Wu, D.; Lu, X.; et al. Age-related immune clearance of hepatitis B virus infection requires the establishment of gut microbiota. Proc. Natl. Acad. Sci. USA 2015, 112, 2175–2180. [Google Scholar] [CrossRef]
- Guo, W.; Zhou, X.; Li, X.; Zhu, Q.; Peng, J.; Zhu, B.; Zheng, X.; Lu, Y.; Yang, D.; Wang, B.; et al. Depletion of Gut Microbiota Impairs Gut Barrier Function and Antiviral Immune Defense in the Liver. Front. Immunol. 2021, 12, 636803. [Google Scholar] [CrossRef]
- Wu, T.; Li, F.; Chen, Y.; Wei, H.; Tian, Z.; Sun, C.; Sun, R. CD4+ T Cells Play a Critical Role in Microbiota-Maintained Anti-HBV Immunity in a Mouse Model. Front. Immunol. 2019, 10, 927. [Google Scholar] [CrossRef]
- Li, Y.; Zhong, S.; Jin, Z.; Ye, G.; Zhang, T.; Liu, Z.; Liu, Z.; Zeng, Z.; Li, Q.; Wang, Y.; et al. Peyer’s patch-involved gut microbiota facilitates anti-HBV immunity in mice. Virus Res. 2023, 331, 199129. [Google Scholar] [CrossRef]
- Zhou, W.; Luo, J.; Xie, X.; Yang, S.; Zhu, D.; Huang, H.; Yang, D.; Liu, J. Gut Microbiota Dysbiosis Strengthens Kupffer Cell-mediated Hepatitis B Virus Persistence through Inducing Endotoxemia in Mice. J. Clin. Transl. Hepatol. 2022, 10, 17–25. [Google Scholar] [CrossRef]
- Bu, Y.; Zhao, K.; Xu, Z.; Zheng, Y.; Hua, R.; Wu, C.; Zhu, C.; Xia, Y.; Cheng, X. Antibiotic-induced gut bacteria depletion has no effect on HBV replication in HBV immune tolerance mouse model. Virol. Sin. 2023, 38, 335–343. [Google Scholar] [CrossRef]
- Sun, X.; Pan, C.Q.; Xing, H. Effect of microbiota metabolites on the progression of chronic hepatitis B virus infection. Hepatol. Int. 2021, 15, 1053–1067. [Google Scholar] [CrossRef]
- Munn, D.H.; Mellor, A.L. Indoleamine 2,3 dioxygenase and metabolic control of immune responses. Trends Immunol. 2013, 34, 137–143. [Google Scholar] [CrossRef]
- Schmidt, S.V.; Schultze, J.L. New Insights into IDO Biology in Bacterial and Viral Infections. Front. Immunol. 2014, 5, 384. [Google Scholar] [CrossRef]
- Mao, R.; Zhang, J.; Jiang, D.; Cai, D.; Levy, J.M.; Cuconati, A.; Block, T.M.; Guo, J.T.; Guo, H. Indoleamine 2,3-dioxygenase mediates the antiviral effect of gamma interferon against hepatitis B virus in human hepatocyte-derived cells. J. Virol. 2011, 85, 1048–1057. [Google Scholar] [CrossRef]
- Yoshio, S.; Sugiyama, M.; Shoji, H.; Mano, Y.; Mita, E.; Okamoto, T.; Matsuura, Y.; Okuno, A.; Takikawa, O.; Mizokami, M.; et al. Indoleamine-2,3-dioxygenase as an effector and an indicator of protective immune responses in patients with acute hepatitis B. Hepatology 2016, 63, 83–94. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.; Wu, S.D.; Chen, Y.; Li, X.Y.; Zhu, Q.; Nakayama, K.; Zhang, W.Q.; Weng, C.Z.; Zhang, J.; Wang, H.K.; et al. Alterations in gut microbiome and metabolomics in chronic hepatitis B infection-associated liver disease and their impact on peripheral immune response. Gut Microbes 2023, 15, 2155018. [Google Scholar] [CrossRef]
- Li, J.; Qiu, S.J.; She, W.M.; Wang, F.P.; Gao, H.; Li, L.; Tu, C.T.; Wang, J.Y.; Shen, X.Z.; Jiang, W. Significance of the balance between regulatory T (Treg) and T helper 17 (Th17) cells during hepatitis B virus related liver fibrosis. PLoS ONE 2012, 7, e39307. [Google Scholar] [CrossRef] [PubMed]
- Meng, F.; Wang, K.; Aoyama, T.; Grivennikov, S.I.; Paik, Y.; Scholten, D.; Cong, M.; Iwaisako, K.; Liu, X.; Zhang, M.; et al. Interleukin-17 signaling in inflammatory, Kupffer cells, and hepatic stellate cells exacerbates liver fibrosis in mice. Gastroenterology 2012, 143, 765–776.e3. [Google Scholar] [CrossRef]
- Tan, Z.; Qian, X.; Jiang, R.; Liu, Q.; Wang, Y.; Chen, C.; Wang, X.; Ryffel, B.; Sun, B. IL-17A plays a critical role in the pathogenesis of liver fibrosis through hepatic stellate cell activation. J. Immunol. 2013, 191, 1835–1844. [Google Scholar] [CrossRef]
- Fabre, T.; Kared, H.; Friedman, S.L.; Shoukry, N.H. IL-17A enhances the expression of profibrotic genes through upregulation of the TGF-β receptor on hepatic stellate cells in a JNK-dependent manner. J. Immunol. 2014, 193, 3925–3933. [Google Scholar] [CrossRef]
- Paratore, M.; Santopaolo, F.; Cammarota, G.; Pompili, M.; Gasbarrini, A.; Ponziani, F.R. Fecal Microbiota Transplantation in Patients with HBV Infection or Other Chronic Liver Diseases: Update on Current Knowledge and Future Perspectives. J. Clin. Med. 2021, 10, 2605. [Google Scholar] [CrossRef]
- Sehgal, R.; Bedi, O.; Trehanpati, N. Role of Microbiota in Pathogenesis and Management of Viral Hepatitis. Front. Cell Infect. Microbiol. 2020, 10, 341. [Google Scholar] [CrossRef]
- Wang, J.; Zhou, X.; Li, X.; Guo, W.; Zhu, Q.; Zhu, B.; Lu, Y.; Zheng, X.; Yang, D.; Wang, B. Fecal Microbiota Transplantation Alters the Outcome of Hepatitis B Virus Infection in Mice. Front. Cell Infect. Microbiol. 2022, 12, 844132. [Google Scholar] [CrossRef]
- Ren, Y.D.; Ye, Z.S.; Yang, L.Z.; Jin, L.X.; Wei, W.J.; Deng, Y.Y.; Chen, X.X.; Xiao, C.X.; Yu, X.F.; Xu, H.Z.; et al. Fecal microbiota transplantation induces hepatitis B virus e-antigen (HBeAg) clearance in patients with positive HBeAg after long-term antiviral therapy. Hepatology 2017, 65, 1765–1768. [Google Scholar] [CrossRef]
- Chauhan, A.; Kumar, R.; Sharma, S.; Mahanta, M.; Vayuuru, S.K.; Nayak, B.; Kumar, S.; Shalimar. Fecal Microbiota Transplantation in Hepatitis B e Antigen-Positive Chronic Hepatitis B Patients: A Pilot Study. Dig. Dis. Sci. 2021, 66, 873–880. [Google Scholar] [CrossRef]
- Mukherjee, A.; Lordan, C.; Ross, R.P.; Cotter, P.D. Gut microbes from the phylogenetically diverse genus Eubacterium and their various contributions to gut health. Gut Microbes 2020, 12, 1802866. [Google Scholar] [CrossRef]
- Xia, X.; Chen, J.; Xia, J.; Wang, B.; Liu, H.; Yang, L.; Wang, Y.; Ling, Z. Role of probiotics in the treatment of minimal hepatic encephalopathy in patients with HBV-induced liver cirrhosis. J. Int. Med. Res. 2018, 46, 3596–3604. [Google Scholar] [CrossRef]
- Lu, H.; Zhu, X.; Wu, L.; Lou, X.; Pan, X.; Liu, B.; Zhang, H.; Zhu, L.; Li, L.; Wu, Z. Alterations in the intestinal microbiome and metabolic profile of patients with cirrhosis supplemented with lactulose, Clostridium butyricum, and Bifidobacterium longum infantis: A randomized placebo-controlled trial. Front. Microbiol. 2023, 14, 1169811. [Google Scholar] [CrossRef]
- Polaris Observatory HCV Collaborators. Global change in hepatitis C virus prevalence and cascade of care between 2015 and 2020: A modelling study. Lancet Gastroenterol. Hepatol. 2022, 7, 396–415. [Google Scholar] [CrossRef]
- Aly, A.M.; Adel, A.; El-Gendy, A.O.; Essam, T.M.; Aziz, R.K. Gut microbiome alterations in patients with stage 4 hepatitis C. Gut Pathog. 2016, 8, 42. [Google Scholar] [CrossRef]
- Heidrich, B.; Vital, M.; Plumeier, I.; Döscher, N.; Kahl, S.; Kirschner, J.; Ziegert, S.; Solbach, P.; Lenzen, H.; Potthoff, A.; et al. Intestinal microbiota in patients with chronic hepatitis C with and without cirrhosis compared with healthy controls. Liver Int. 2018, 38, 50–58. [Google Scholar] [CrossRef]
- Mizutani, T.; Ishizaka, A.; Koga, M.; Tsutsumi, T.; Yotsuyanagi, H. Role of Microbiota in Viral Infections and Pathological Progression. Viruses 2022, 14, 950. [Google Scholar] [CrossRef]
- Preveden, T.; Scarpellini, E.; Milić, N.; Luzza, F.; Abenavoli, L. Gut microbiota changes and chronic hepatitis C virus infection. Expert. Rev. Gastroenterol. Hepatol. 2017, 11, 813–819. [Google Scholar] [CrossRef]
- Dolganiuc, A.; Norkina, O.; Kodys, K.; Catalano, D.; Bakis, G.; Marshall, C.; Mandrekar, P.; Szabo, G. Viral and host factors induce macrophage activation and loss of toll-like receptor tolerance in chronic HCV infection. Gastroenterology 2007, 133, 1627–1636. [Google Scholar] [CrossRef]
- Inoue, T.; Funatsu, Y.; Ohnishi, M.; Isogawa, M.; Kawashima, K.; Tanaka, M.; Moriya, K.; Kawaratani, H.; Momoda, R.; Iio, E.; et al. Bile acid dysmetabolism in the gut-microbiota-liver axis under hepatitis C virus infection. Liver Int. 2022, 42, 124–134. [Google Scholar] [CrossRef]
- Ponziani, F.R.; Putignani, L.; Paroni Sterbini, F.; Petito, V.; Picca, A.; Del Chierico, F.; Reddel, S.; Calvani, R.; Marzetti, E.; Sanguinetti, M.; et al. Influence of hepatitis C virus eradication with direct-acting antivirals on the gut microbiota in patients with cirrhosis. Aliment. Pharmacol. Ther. 2018, 48, 1301–1311. [Google Scholar] [CrossRef] [PubMed]
- Iwata, R.; Stieger, B.; Mertens, J.C.; Müller, T.; Baur, K.; Frei, P.; Braun, J.; Vergopoulos, A.; Martin, I.V.; Schmitt, J.; et al. The role of bile acid retention and a common polymorphism in the ABCB11 gene as host factors affecting antiviral treatment response in chronic hepatitis C. J. Viral Hepat. 2011, 18, 768–778. [Google Scholar] [CrossRef]
- Inoue, T.; Nakayama, J.; Moriya, K.; Kawaratani, H.; Momoda, R.; Ito, K.; Iio, E.; Nojiri, S.; Fujiwara, K.; Yoneda, M.; et al. Gut Dysbiosis Associated With Hepatitis C Virus Infection. Clin. Infect. Dis. 2018, 67, 869–877. [Google Scholar] [CrossRef] [PubMed]
- Atarashi, K.; Tanoue, T.; Shima, T.; Imaoka, A.; Kuwahara, T.; Momose, Y.; Cheng, G.; Yamasaki, S.; Saito, T.; Ohba, Y.; et al. Induction of colonic regulatory T cells by indigenous Clostridium species. Science 2011, 331, 337–341. [Google Scholar] [CrossRef] [PubMed]
- Furusawa, Y.; Obata, Y.; Fukuda, S.; Endo, T.A.; Nakato, G.; Takahashi, D.; Nakanishi, Y.; Uetake, C.; Kato, K.; Kato, T.; et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 2013, 504, 446–450. [Google Scholar] [CrossRef]
- Ullah, N.; Kakakhel, M.A.; Khan, I.; Gul Hilal, M.; Lajia, Z.; Bai, Y.; Sajjad, W.; Yuxi, L.; Ullah, H.; Almohaimeed, H.M.; et al. Structural and compositional segregation of the gut microbiota in HCV and liver cirrhotic patients: A clinical pilot study. Microb. Pathog. 2022, 171, 105739. [Google Scholar] [CrossRef] [PubMed]
- Ashour, Z.; Shahin, R.; Ali-Eldin, Z.; El-Shayeb, M.; El-Tayeb, T.; Bakr, S. Potential impact of gut Lactobacillus acidophilus and Bifidobacterium bifidum on hepatic histopathological changes in non-cirrhotic hepatitis C virus patients with different viral load. Gut Pathog. 2022, 14, 25. [Google Scholar] [CrossRef]
- Moon, M.S.; Quinn, G.; Townsend, E.C.; Ali, R.O.; Zhang, G.Y.; Bradshaw, A.; Hill, K.; Guan, H.; Hamilton, D.; Kleiner, D.E.; et al. Bacterial Translocation and Host Immune Activation in Chronic Hepatitis C Infection. Open Forum Infect. Dis. 2019, 6, ofz255. [Google Scholar] [CrossRef]
- Sultan, S.; El-Mowafy, M.; Elgaml, A.; El-Mesery, M.; El Shabrawi, A.; Elegezy, M.; Hammami, R.; Mottawea, W. Alterations of the Treatment-Naive Gut Microbiome in Newly Diagnosed Hepatitis C Virus Infection. ACS Infect. Dis. 2021, 7, 1059–1068. [Google Scholar] [CrossRef]
- Wellhöner, F.; Döscher, N.; Tergast, T.L.; Vital, M.; Plumeier, I.; Kahl, S.; Potthoff, A.; Manns, M.P.; Maasoumy, B.; Wedemeyer, H.; et al. The impact of proton pump inhibitors on the intestinal microbiota in chronic hepatitis C patients. Scand. J. Gastroenterol. 2019, 54, 1033–1041. [Google Scholar] [CrossRef]
- El-Mowafy, M.; Elgaml, A.; El-Mesery, M.; Sultan, S.; Ahmed, T.A.E.; Gomaa, A.I.; Aly, M.; Mottawea, W. Changes of Gut-Microbiota-Liver Axis in Hepatitis C Virus Infection. Biology 2021, 10, 55. [Google Scholar] [CrossRef]
- Neag, M.A.; Mitre, A.O.; Catinean, A.; Buzoianu, A.D. Overview of the microbiota in the gut-liver axis in viral B and C hepatitis. World J. Gastroenterol. 2021, 27, 7446–7461. [Google Scholar] [CrossRef]
- Bajaj, J.S.; Sterling, R.K.; Betrapally, N.S.; Nixon, D.E.; Fuchs, M.; Daita, K.; Heuman, D.M.; Sikaroodi, M.; Hylemon, P.B.; White, M.B.; et al. HCV eradication does not impact gut dysbiosis or systemic inflammation in cirrhotic patients. Aliment. Pharmacol. Ther. 2016, 44, 638–643. [Google Scholar] [CrossRef]
- Pérez-Matute, P.; Íñiguez, M.; Villanueva-Millán, M.J.; Recio-Fernández, E.; Vázquez, A.M.; Sánchez, S.C.; Morano, L.E.; Oteo, J.A. Short-term effects of direct-acting antiviral agents on inflammation and gut microbiota in hepatitis C-infected patients. Eur. J. Intern. Med. 2019, 67, 47–58. [Google Scholar] [CrossRef]
- Wellhöner, F.; Döscher, N.; Woelfl, F.; Vital, M.; Plumeier, I.; Kahl, S.; Potthoff, A.; Manns, M.P.; Pieper, D.H.; Cornberg, M.; et al. Eradication of Chronic HCV Infection: Improvement of Dysbiosis Only in Patients Without Liver Cirrhosis. Hepatology 2021, 74, 72–82. [Google Scholar] [CrossRef]
- Yilmaz, B.; Ruckstuhl, L.; Müllhaupt, B.; Magenta, L.; Kuster, M.H.; Clerc, O.; Torgler, R.; Semmo, N. Pilot Sub-Study of the Effect of Hepatitis C Cure by Glecaprevir/Pibrentasvir on the Gut Microbiome of Patients with Chronic Hepatitis C Genotypes 1 to 6 in the Mythen Study. Pharmaceuticals 2021, 14, 931. [Google Scholar] [CrossRef]
- Huang, P.Y.; Chen, C.H.; Tsai, M.J.; Yao, C.C.; Wang, H.M.; Kuo, Y.H.; Chang, K.C.; Hung, C.H.; Chuah, S.K.; Tsai, M.C. Effects of direct anti-viral agents on the gut microbiota in patients with chronic hepatitis C. J. Formos. Med. Assoc. 2023, 122, 157–163. [Google Scholar] [CrossRef]
- Hsu, Y.C.; Chen, C.C.; Lee, W.H.; Chang, C.Y.; Lee, F.J.; Tseng, C.H.; Chen, T.H.; Ho, H.J.; Lin, J.T.; Wu, C.Y. Compositions of gut microbiota before and shortly after hepatitis C viral eradication by direct antiviral agents. Sci. Rep. 2022, 12, 5481. [Google Scholar] [CrossRef]
- Honda, T.; Ishigami, M.; Yamamoto, K.; Takeyama, T.; Ito, T.; Ishizu, Y.; Kuzuya, T.; Nakamura, M.; Kawashima, H.; Miyahara, R.; et al. Changes in the gut microbiota after hepatitis C virus eradication. Sci. Rep. 2021, 11, 23568. [Google Scholar] [CrossRef]
- Chuaypen, N.; Jinato, T.; Avihingsanon, A.; Nookaew, I.; Tanaka, Y.; Tangkijvanich, P. Long-term benefit of DAAs on gut dysbiosis and microbial translocation in HCV-infected patients with and without HIV coinfection. Sci. Rep. 2023, 13, 14413. [Google Scholar] [CrossRef]
- Lattanzi, B.; Baroncelli, S.; De Santis, A.; Galluzzo, C.M.; Mennini, G.; Michelini, Z.; Lupo, M.; Ginanni Corradini, S.; Rossi, M.; Palmisano, L.; et al. Microbial translocation and T cell activation are modified by direct-acting antiviral therapy in HCV-infected patients. Aliment. Pharmacol. Ther. 2018, 48, 1146–1155. [Google Scholar] [CrossRef]
- Torres-Barceló, C. The disparate effects of bacteriophages on antibiotic-resistant bacteria. Emerg. Microbes Infect. 2018, 7, 168. [Google Scholar] [CrossRef] [PubMed]
- Stern, J.; Miller, G.; Li, X.; Saxena, D. Virome and bacteriome: Two sides of the same coin. Curr. Opin. Virol. 2019, 37, 37–43. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Bortolanza, M.; Zhai, G.; Shang, A.; Ling, Z.; Jiang, B.; Shen, X.; Yao, Y.; Yu, J.; Li, L.; et al. Gut microbiota dysbiosis associated with plasma levels of Interferon-γ and viral load in patients with acute hepatitis E infection. J. Med. Virol. 2022, 94, 692–702. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Huang, F.; Ling, Z.; Liu, S.; Liu, J.; Fan, J.; Yu, J.; Wang, W.; Jin, X.; Meng, Y.; et al. Altered faecal microbiota on the expression of Th cells responses in the exacerbation of patients with hepatitis E infection. J. Viral Hepat. 2020, 27, 1243–1252. [Google Scholar] [CrossRef] [PubMed]
- Kreuzer, S.; Machnowska, P.; Aßmus, J.; Sieber, M.; Pieper, R.; Schmidt, M.F.; Brockmann, G.A.; Scharek-Tedin, L.; Johne, R. Feeding of the probiotic bacterium Enterococcus faecium NCIMB 10415 differentially affects shedding of enteric viruses in pigs. Vet Res. 2012, 43, 58. [Google Scholar] [CrossRef]
- Ishizaka, A.; Koga, M.; Mizutani, T.; Lim, L.A.; Adachi, E.; Ikeuchi, K.; Ueda, R.; Aoyagi, H.; Tanaka, S.; Kiyono, H.; et al. Prolonged Gut Dysbiosis and Fecal Excretion of Hepatitis A Virus in Patients Infected with Human Immunodeficiency Virus. Viruses. 2021, 13, 2101. [Google Scholar] [CrossRef]
- Kefalakes, H.; Rehermann, B. Inflammation drives an altered phenotype of mucosal-associated invariant T cells in chronic hepatitis D virus infection. J. Hepatol. 2019, 71, 237–239. [Google Scholar] [CrossRef]
- Bhat, M.; Arendt, B.M.; Bhat, V.; Renner, E.L.; Humar, A.; Allard, J.P. Implication of the intestinal microbiome in complications of cirrhosis. World J. Hepatol. 2016, 8, 1128–1136. [Google Scholar] [CrossRef]
- Kakiyama, G.; Pandak, W.M.; Gillevet, P.M.; Hylemon, P.B.; Heuman, D.M.; Daita, K.; Takei, H.; Muto, A.; Nittono, H.; Ridlon, J.M.; et al. Modulation of the fecal bile acid profile by gut microbiota in cirrhosis. J. Hepatol. 2013, 58, 949–955. [Google Scholar] [CrossRef]
- Chen, Y.; Ji, F.; Guo, J.; Shi, D.; Fang, D.; Li, L. Dysbiosis of small intestinal microbiota in liver cirrhosis and its association with etiology. Sci. Rep. 2016, 6, 34055. [Google Scholar] [CrossRef]
- Bajaj, J.S.; Heuman, D.M.; Hylemon, P.B.; Sanyal, A.J.; White, M.B.; Monteith, P.; Noble, N.A.; Unser, A.B.; Daita, K.; Fisher, A.R.; et al. Altered profile of human gut microbiome is associated with cirrhosis and its complications. J. Hepatol. 2014, 60, 940–947. [Google Scholar] [CrossRef]
- Assimakopoulos, S.F.; Tsamandas, A.C.; Tsiaoussis, G.I.; Karatza, E.; Triantos, C.; Vagianos, C.E.; Spiliopoulou, I.; Kaltezioti, V.; Charonis, A.; Nikolopoulou, V.N.; et al. Altered intestinal tight junctions’ expression in patients with liver cirrhosis: A pathogenetic mechanism of intestinal hyperpermeability. Eur. J. Clin. Investig. 2012, 42, 439–446. [Google Scholar] [CrossRef]
- Teltschik, Z.; Wiest, R.; Beisner, J.; Nuding, S.; Hofmann, C.; Schoelmerich, J.; Bevins, C.L.; Stange, E.F.; Wehkamp, J. Intestinal bacterial translocation in rats with cirrhosis is related to compromised Paneth cell antimicrobial host defense. Hepatology 2012, 55, 1154–1163. [Google Scholar] [CrossRef]
- Simbrunner, B.; Trauner, M.; Reiberger, T. Review article: Therapeutic aspects of bile acid signalling in the gut-liver axis. Aliment. Pharmacol. Ther. 2021, 54, 1243–1262. [Google Scholar] [CrossRef]
- Shao, J.W.; Ge, T.T.; Chen, S.Z.; Wang, G.; Yang, Q.; Huang, C.H.; Xu, L.C.; Chen, Z. Role of bile acids in liver diseases mediated by the gut microbiome. World J. Gastroenterol. 2021, 27, 3010–3021. [Google Scholar] [CrossRef]
- Sauerbruch, T.; Hennenberg, M.; Trebicka, J.; Beuers, U. Bile Acids, Liver Cirrhosis, and Extrahepatic Vascular Dysfunction. Front. Physiol. 2021, 12, 718783. [Google Scholar] [CrossRef]
- Liu, Y.; Chen, K.; Li, F.; Gu, Z.; Liu, Q.; He, L.; Shao, T.; Song, Q.; Zhu, F.; Zhang, L.; et al. Probiotic Lactobacillus rhamnosus GG Prevents Liver Fibrosis Through Inhibiting Hepatic Bile Acid Synthesis and Enhancing Bile Acid Excretion in Mice. Hepatology 2020, 71, 2050–2066. [Google Scholar] [CrossRef]
- Albillos, A.; Martin-Mateos, R.; Van der Merwe, S.; Wiest, R.; Jalan, R.; Álvarez-Mon, M. Cirrhosis-associated immune dysfunction. Nat. Rev. Gastroenterol. Hepatol. 2022, 19, 112–134. [Google Scholar] [CrossRef]
- Kaliannan, K. Compromise of α-Defensin Function in Liver Cirrhosis Facilitates the Toxic Relationship Between Gut Permeability and Endotoxemia. Dig. Dis. Sci. 2018, 63, 2492–2494. [Google Scholar] [CrossRef]
- Hassan, M.; Moghadamrad, S.; Sorribas, M.; Muntet, S.G.; Kellmann, P.; Trentesaux, C.; Fraudeau, M.; Nanni, P.; Wolski, W.; Keller, I.; et al. Paneth cells promote angiogenesis and regulate portal hypertension in response to microbial signals. J. Hepatol. 2020, 73, 628–639. [Google Scholar] [CrossRef]
- Hrncir, T.; Hrncirova, L.; Kverka, M.; Hromadka, R.; Machova, V.; Trckova, E.; Kostovcikova, K.; Kralickova, P.; Krejsek, J.; Tlaskalova-Hogenova, H. Gut Microbiota and NAFLD: Pathogenetic Mechanisms, Microbiota Signatures, and Therapeutic Interventions. Microorganisms 2021, 9, 957. [Google Scholar] [CrossRef]
- Kassa, Y.; Million, Y.; Gedefie, A.; Moges, F. Alteration of Gut Microbiota and Its Impact on Immune Response in Patients with Chronic HBV Infection: A Review. Infect. Drug Resist. 2021, 14, 2571–2578. [Google Scholar] [CrossRef]
- Kisseleva, T.; Brenner, D. Molecular and cellular mechanisms of liver fibrosis and its regression. Nat. Rev. Gastroenterol. Hepatol. 2021, 18, 151–166. [Google Scholar] [CrossRef]
- Parola, M.; Pinzani, M. Liver fibrosis: Pathophysiology, pathogenetic targets and clinical issues. Mol. Aspects Med. 2019, 65, 37–55. [Google Scholar] [CrossRef]
- Strowig, T.; Henao-Mejia, J.; Elinav, E.; Flavell, R. Inflammasomes in health and disease. Nature 2012, 481, 278–286. [Google Scholar] [CrossRef]
- Schroder, K.; Tschopp, J. The inflammasomes. Cell 2010, 140, 821–832. [Google Scholar] [CrossRef]
- Watanabe, A.; Sohail, M.A.; Gomes, D.A.; Hashmi, A.; Nagata, J.; Sutterwala, F.S.; Mahmood, S.; Jhandier, M.N.; Shi, Y.; Flavell, R.A.; et al. Inflammasome-mediated regulation of hepatic stellate cells. Am. J. Physiol. Gastrointest. Liver Physiol. 2009, 296, G1248–G1257. [Google Scholar] [CrossRef]
- Boaru, S.G.; Borkham-Kamphorst, E.; Tihaa, L.; Haas, U.; Weiskirchen, R. Expression analysis of inflammasomes in experimental models of inflammatory and fibrotic liver disease. J. Inflamm. 2012, 9, 49. [Google Scholar] [CrossRef]
- Gurung, P.; Li, B.; Subbarao Malireddi, R.K.; Lamkanfi, M.; Geiger, T.L.; Kanneganti, T.D. Chronic TLR Stimulation Controls NLRP3 Inflammasome Activation through IL-10 Mediated Regulation of NLRP3 Expression and Caspase-8 Activation. Sci. Rep. 2015, 5, 14488. [Google Scholar] [CrossRef]
- Chuang, S.Y.; Yang, C.H.; Chou, C.C.; Chiang, Y.P.; Chuang, T.H.; Hsu, L.C. TLR-induced PAI-2 expression suppresses IL-1β processing via increasing autophagy and NLRP3 degradation. Proc. Natl. Acad. Sci. USA 2013, 110, 16079–16084. [Google Scholar] [CrossRef]
- Arab, J.P.; Martin-Mateos, R.M.; Shah, V.H. Gut-liver axis, cirrhosis and portal hypertension: The chicken and the egg. Hepatol. Int. 2018, 12, 24–33. [Google Scholar] [CrossRef]
- Johnson, K.V.; Foster, K.R. Why does the microbiome affect behaviour? Nat. Rev. Microbiol. 2018, 16, 647–655. [Google Scholar] [CrossRef]
- Ding, J.H.; Jin, Z.; Yang, X.X.; Lou, J.; Shan, W.X.; Hu, Y.X.; Du, Q.; Liao, Q.S.; Xie, R.; Xu, J.Y. Role of gut microbiota via the gut-liver-brain axis in digestive diseases. World J. Gastroenterol. 2020, 26, 6141–6162. [Google Scholar] [CrossRef]
- Smith, M.L.; Wade, J.B.; Wolstenholme, J.; Bajaj, J.S. Gut microbiome-brain-cirrhosis axis. Hepatology 2023. [Google Scholar] [CrossRef]
- Won, S.M.; Oh, K.K.; Gupta, H.; Ganesan, R.; Sharma, S.P.; Jeong, J.J.; Yoon, S.J.; Jeong, M.K.; Min, B.H.; Hyun, J.Y.; et al. The Link between Gut Microbiota and Hepatic Encephalopathy. Int. J. Mol. Sci. 2022, 23, 8999. [Google Scholar] [CrossRef]
- Bajaj, J.S.; Hylemon, P.B.; Ridlon, J.M.; Heuman, D.M.; Daita, K.; White, M.B.; Monteith, P.; Noble, N.A.; Sikaroodi, M.; Gillevet, P.M. Colonic mucosal microbiome differs from stool microbiome in cirrhosis and hepatic encephalopathy and is linked to cognition and inflammation. Am. J. Physiol. Gastrointest. Liver Physiol. 2012, 303, G675–G685. [Google Scholar] [CrossRef]
- Zhu, R.; Liu, L.; Zhang, G.; Dong, J.; Ren, Z.; Li, Z. The pathogenesis of gut microbiota in hepatic encephalopathy by the gut-liver-brain axis. Biosci. Rep. 2023, 43, BSR20222524. [Google Scholar] [CrossRef]
- Xie, G.; Wang, X.; Jiang, R.; Zhao, A.; Yan, J.; Zheng, X.; Huang, F.; Liu, X.; Panee, J.; Rajani, C.; et al. Dysregulated bile acid signaling contributes to the neurological impairment in murine models of acute and chronic liver failure. EBioMedicine 2018, 37, 294–306. [Google Scholar] [CrossRef]
- Ridlon, J.M.; Alves, J.M.; Hylemon, P.B.; Bajaj, J.S. Cirrhosis, bile acids and gut microbiota: Unraveling a complex relationship. Gut Microbes 2013, 4, 382–387. [Google Scholar] [CrossRef]
- Sung, C.M.; Lin, Y.F.; Chen, K.F.; Ke, H.M.; Huang, H.Y.; Gong, Y.N.; Tsai, W.S.; You, J.F.; Lu, M.J.; Cheng, H.T.; et al. Predicting Clinical Outcomes of Cirrhosis Patients With Hepatic Encephalopathy From the Fecal Microbiome. Cell Mol. Gastroenterol. Hepatol. 2019, 8, 301–318.e2. [Google Scholar] [CrossRef]
- Zhang, Z.; Zhai, H.; Geng, J.; Yu, R.; Ren, H.; Fan, H.; Shi, P. Large-scale survey of gut microbiota associated with MHE Via 16S rRNA-based pyrosequencing. Am. J. Gastroenterol. 2013, 108, 1601–1611. [Google Scholar] [CrossRef]
- Luo, M.; Hu, F.R.; Xin, R.J.; Yao, L.; Hu, S.J.; Bai, F.H. Altered gut microbiota is associated with sleep disturbances in patients with minimal hepatic encephalopathy caused by hepatitis B-related liver cirrhosis. Expert. Rev. Gastroenterol. Hepatol. 2022, 16, 797–807. [Google Scholar] [CrossRef]
- Bajaj, J.S.; Ridlon, J.M.; Hylemon, P.B.; Thacker, L.R.; Heuman, D.M.; Smith, S.; Sikaroodi, M.; Gillevet, P.M. Linkage of gut microbiome with cognition in hepatic encephalopathy. Am. J. Physiol. Gastrointest. Liver Physiol. 2012, 302, G168–G175. [Google Scholar] [CrossRef]
- Ahluwalia, V.; Betrapally, N.S.; Hylemon, P.B.; White, M.B.; Gillevet, P.M.; Unser, A.B.; Fagan, A.; Daita, K.; Heuman, D.M.; Zhou, H.; et al. Impaired Gut-Liver-Brain Axis in Patients with Cirrhosis. Sci. Rep. 2016, 6, 26800. [Google Scholar] [CrossRef]
- Yukawa-Muto, Y.; Kamiya, T.; Fujii, H.; Mori, H.; Toyoda, A.; Sato, I.; Konishi, Y.; Hirayama, A.; Hara, E.; Fukuda, S.; et al. Distinct responsiveness to rifaximin in patients with hepatic encephalopathy depends on functional gut microbial species. Hepatol. Commun. 2022, 6, 2090–2104. [Google Scholar] [CrossRef]
- Bajaj, J.S.; Shamsaddini, A.; Fagan, A.; McGeorge, S.; Gavis, E.; Sikaroodi, M.; Brenner, L.A.; Wade, J.B.; Gillevet, P.M. Distinct gut microbial compositional and functional changes associated with impaired inhibitory control in patients with cirrhosis. Gut Microbes 2021, 13, 1953247. [Google Scholar] [CrossRef]
- Bajaj, J.S.; Fagan, A.; White, M.B.; Wade, J.B.; Hylemon, P.B.; Heuman, D.M.; Fuchs, M.; John, B.V.; Acharya, C.; Sikaroodi, M.; et al. Specific Gut and Salivary Microbiota Patterns Are Linked With Different Cognitive Testing Strategies in Minimal Hepatic Encephalopathy. Am. J. Gastroenterol. 2019, 114, 1080–1090. [Google Scholar] [CrossRef]
- Luo, M.; Xin, R.J.; Hu, F.R.; Yao, L.; Hu, S.J.; Bai, F.H. Role of gut microbiota in the pathogenesis and therapeutics of minimal hepatic encephalopathy via the gut-liver-brain axis. World J. Gastroenterol. 2023, 29, 144–156. [Google Scholar] [CrossRef]
- Alberts, C.J.; Clifford, G.M.; Georges, D.; Negro, F.; Lesi, O.A.; Hutin, Y.J.; de Martel, C. Worldwide prevalence of hepatitis B virus and hepatitis C virus among patients with cirrhosis at country, region, and global levels: A systematic review. Lancet Gastroenterol. Hepatol. 2022, 7, 724–735. [Google Scholar] [CrossRef]
- Li, Y.G.; Yu, Z.J.; Li, A.; Ren, Z.G. Gut microbiota alteration and modulation in hepatitis B virus-related fibrosis and complications: Molecular mechanisms and therapeutic inventions. World J. Gastroenterol. 2022, 28, 3555–3572. [Google Scholar] [CrossRef]
- Xu, M.; Luo, K.; Li, J.; Li, Y.; Zhang, Y.; Yuan, Z.; Xu, Q.; Wu, X. Role of Intestinal Microbes in Chronic Liver Diseases. Int. J. Mol. Sci. 2022, 23, 12661. [Google Scholar] [CrossRef]
- Wu, Z.W.; Lu, H.F.; Wu, J.; Zuo, J.; Chen, P.; Sheng, J.F.; Zheng, S.S.; Li, L.J. Assessment of the fecal lactobacilli population in patients with hepatitis B virus-related decompensated cirrhosis and hepatitis B cirrhosis treated with liver transplant. Microb. Ecol. 2012, 63, 929–937. [Google Scholar] [CrossRef]
- Deng, Y.D.; Peng, X.B.; Zhao, R.R.; Ma, C.Q.; Li, J.N.; Yao, L.Q. The intestinal microbial community dissimilarity in hepatitis B virus-related liver cirrhosis patients with and without at alcohol consumption. Gut Pathog. 2019, 11, 58. [Google Scholar] [CrossRef]
- Cani, P.D.; Jordan, B.F. Gut microbiota-mediated inflammation in obesity: A link with gastrointestinal cancer. Nat. Rev. Gastroenterol. Hepatol. 2018, 15, 671–682. [Google Scholar] [CrossRef]
- Kajihara, M.; Koido, S.; Kanai, T.; Ito, Z.; Matsumoto, Y.; Takakura, K.; Saruta, M.; Kato, K.; Odamaki, T.; Xiao, J.Z.; et al. Characterisation of blood microbiota in patients with liver cirrhosis. Eur. J. Gastroenterol. Hepatol. 2019, 31, 1577–1583. [Google Scholar] [CrossRef]
- Zheng, R.; Wang, G.; Pang, Z.; Ran, N.; Gu, Y.; Guan, X.; Yuan, Y.; Zuo, X.; Pan, H.; Zheng, J.; et al. Liver cirrhosis contributes to the disorder of gut microbiota in patients with hepatocellular carcinoma. Cancer Med. 2020, 9, 4232–4250. [Google Scholar] [CrossRef]
- Sun, X.; Chi, X.; Zhao, Y.; Liu, S.; Xing, H. Characteristics and Clinical Significance of Intestinal Microbiota in Patients with Chronic Hepatitis B Cirrhosis and Type 2 Diabetes Mellitus. J. Diabetes Res. 2022, 2022, 1826181. [Google Scholar] [CrossRef]
- Fasano, A.; Not, T.; Wang, W.; Uzzau, S.; Berti, I.; Tommasini, A.; Goldblum, S.E. Zonulin, a newly discovered modulator of intestinal permeability, and its expression in coeliac disease. Lancet 2000, 355, 1518–1519. [Google Scholar] [CrossRef]
- Jiang, J.W.; Chen, X.H.; Ren, Z.; Zheng, S.S. Gut microbial dysbiosis associates hepatocellular carcinoma via the gut-liver axis. Hepatobiliary Pancreat. Dis. Int. 2019, 18, 19–27. [Google Scholar] [CrossRef]
- Behary, J.; Amorim, N.; Jiang, X.T.; Raposo, A.; Gong, L.; McGovern, E.; Ibrahim, R.; Chu, F.; Stephens, C.; Jebeili, H.; et al. Gut microbiota impact on the peripheral immune response in non-alcoholic fatty liver disease related hepatocellular carcinoma. Nat. Commun. 2021, 12, 187. [Google Scholar] [CrossRef]
- Ponziani, F.R.; Bhoori, S.; Castelli, C.; Putignani, L.; Rivoltini, L.; Del Chierico, F.; Sanguinetti, M.; Morelli, D.; Paroni Sterbini, F.; Petito, V.; et al. Hepatocellular Carcinoma Is Associated With Gut Microbiota Profile and Inflammation in Nonalcoholic Fatty Liver Disease. Hepatology 2019, 69, 107–120. [Google Scholar] [CrossRef]
- Khalyfa, A.A.; Punatar, S.; Yarbrough, A. Hepatocellular Carcinoma: Understanding the Inflammatory Implications of the Microbiome. Int. J. Mol. Sci. 2022, 23, 8164. [Google Scholar] [CrossRef]
- Rajapakse, J.; Khatiwada, S.; Akon, A.C.; Yu, K.L.; Shen, S.; Zekry, A. Unveiling the complex relationship between gut microbiota and liver cancer: Opportunities for novel therapeutic interventions. Gut Microbes 2023, 15, 2240031. [Google Scholar] [CrossRef]
- Liu, S.; Yang, X. Intestinal flora plays a role in the progression of hepatitis-cirrhosis-liver cancer. Front. Cell Infect. Microbiol. 2023, 13, 1140126. [Google Scholar] [CrossRef]
- Mohamadkhani, A. On the potential role of intestinal microbial community in hepatocarcinogenesis in chronic hepatitis B. Cancer Med. 2018, 7, 3095–3100. [Google Scholar] [CrossRef]
- Wu, L.; Feng, J.; Li, J.; Yu, Q.; Ji, J.; Wu, J.; Dai, W.; Guo, C. The gut microbiome-bile acid axis in hepatocarcinogenesis. Biomed. Pharmacother. 2021, 133, 111036. [Google Scholar] [CrossRef]
- Marascio, N.; De Caro, C.; Quirino, A.; Mazzitelli, M.; Russo, E.; Torti, C.; Matera, G. The Role of the Microbiota Gut-Liver Axis during HCV Chronic Infection: A Schematic Overview. J. Clin. Med. 2022, 11, 5936. [Google Scholar] [CrossRef]
- Virseda-Berdices, A.; Brochado-Kith, O.; Díez, C.; Hontañon, V.; Berenguer, J.; González-García, J.; Rojo, D.; Fernández-Rodríguez, A.; Ibañez-Samaniego, L.; Llop-Herrera, E.; et al. Blood microbiome is associated with changes in portal hypertension after successful direct-acting antiviral therapy in patients with HCV-related cirrhosis. J. Antimicrob. Chemother. 2022, 77, 719–726. [Google Scholar] [CrossRef]
- Bajaj, J.S.; Salzman, N.H.; Acharya, C.; Sterling, R.K.; White, M.B.; Gavis, E.A.; Fagan, A.; Hayward, M.; Holtz, M.L.; Matherly, S.; et al. Fecal Microbial Transplant Capsules Are Safe in Hepatic Encephalopathy: A Phase 1, Randomized, Placebo-Controlled Trial. Hepatology 2019, 70, 1690–1703. [Google Scholar] [CrossRef]
- Bloom, P.P.; Donlan, J.; Torres Soto, M.; Daidone, M.; Hohmann, E.; Chung, R.T. Fecal microbiota transplant improves cognition in hepatic encephalopathy and its effect varies by donor and recipient. Hepatol. Commun. 2022, 6, 2079–2089. [Google Scholar] [CrossRef]
- Sarangi, A.N.; Goel, A.; Singh, A.; Sasi, A.; Aggarwal, R. Faecal bacterial microbiota in patients with cirrhosis and the effect of lactulose administration. BMC Gastroenterol. 2017, 17, 125. [Google Scholar] [CrossRef] [PubMed]
- Moratalla, A.; Ampuero, J.; Bellot, P.; Gallego-Durán, R.; Zapater, P.; Roger, M.; Figueruela, B.; Martínez-Moreno, B.; González-Navajas, J.M.; Such, J.; et al. Lactulose reduces bacterial DNA translocation, which worsens neurocognitive shape in cirrhotic patients with minimal hepatic encephalopathy. Liver Int. 2017, 37, 212–223. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.Y.; Bajaj, J.S.; Wang, J.B.; Shang, J.; Zhou, X.M.; Guo, X.L.; Zhu, X.; Meng, L.N.; Jiang, H.X.; Mi, Y.Q.; et al. Lactulose improves cognition, quality of life, and gut microbiota in minimal hepatic encephalopathy: A multicenter, randomized controlled trial. J. Dig. Dis. 2019, 20, 547–556. [Google Scholar] [CrossRef] [PubMed]
- Bajaj, J.S.; Heuman, D.M.; Sanyal, A.J.; Hylemon, P.B.; Sterling, R.K.; Stravitz, R.T.; Fuchs, M.; Ridlon, J.M.; Daita, K.; Monteith, P.; et al. Modulation of the metabiome by rifaximin in patients with cirrhosis and minimal hepatic encephalopathy. PLoS ONE 2013, 8, e60042. [Google Scholar] [CrossRef]
- Bajaj, J.S.; Heuman, D.M.; Hylemon, P.B.; Sanyal, A.J.; Puri, P.; Sterling, R.K.; Luketic, V.; Stravitz, R.T.; Siddiqui, M.S.; Fuchs, M.; et al. Randomised clinical trial: Lactobacillus GG modulates gut microbiome, metabolome and endotoxemia in patients with cirrhosis. Aliment. Pharmacol. Ther. 2014, 39, 1113–1125. [Google Scholar] [CrossRef]
- Manzhalii, E.; Moyseyenko, V.; Kondratiuk, V.; Molochek, N.; Falalyeyeva, T.; Kobyliak, N. Effect of a specific Escherichia coli Nissle 1917 strain on minimal/mild hepatic encephalopathy treatment. World J. Hepatol. 2022, 14, 634–646. [Google Scholar] [CrossRef]
- Zuo, Z.; Fan, H.; Tang, X.D.; Chen, Y.M.; Xun, L.T.; Li, Y.; Song, Z.J.; Zhai, H.Q. Effect of different treatments and alcohol addiction on gut microbiota in minimal hepatic encephalopathy patients. Exp. Ther. Med. 2017, 14, 4887–4895. [Google Scholar] [CrossRef]
- Liu, Q.; Duan, Z.P.; Ha, D.K.; Bengmark, S.; Kurtovic, J.; Riordan, S.M. Synbiotic modulation of gut flora: Effect on minimal hepatic encephalopathy in patients with cirrhosis. Hepatology 2004, 39, 1441–1449. [Google Scholar] [CrossRef]
- Bajaj, J.S.; Kassam, Z.; Fagan, A.; Gavis, E.A.; Liu, E.; Cox, I.J.; Kheradman, R.; Heuman, D.; Wang, J.; Gurry, T.; et al. Fecal microbiota transplant from a rational stool donor improves hepatic encephalopathy: A randomized clinical trial. Hepatology 2017, 66, 1727–1738. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).