A Machine Learning Perspective on Personalized Medicine: An Automized, Comprehensive Knowledge Base with Ontology for Pattern Recognition
Abstract
1. Introduction
Personalized medicine is a broad and rapidly advancing field of health care that is informed by each person’s unique clinical, genetic, genomic, and environmental information.
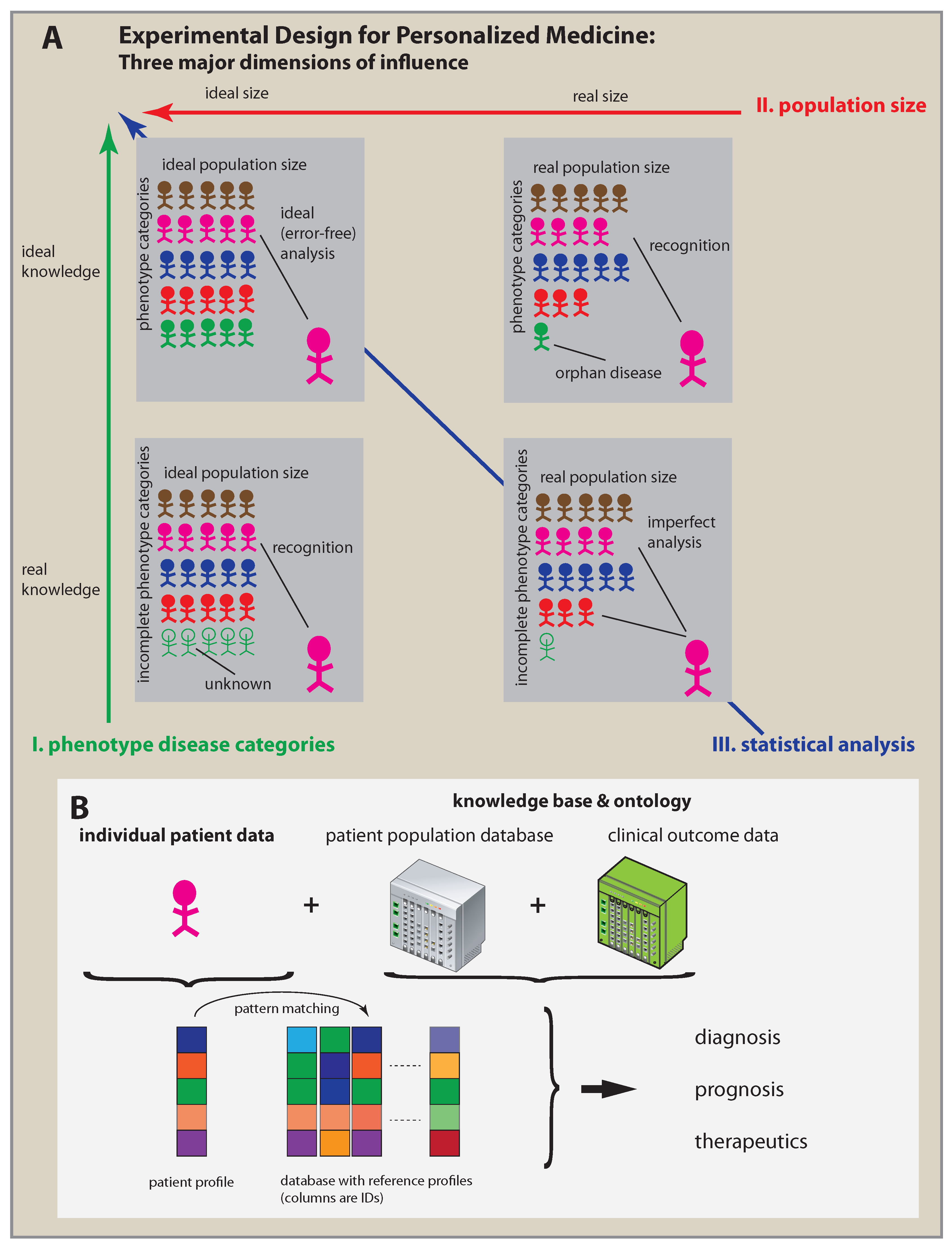
2. Three Key Factors of Personalized Medicine
3. Advances Required to Implement Personalized Medicine
4. A Machine Learning Perspective

5. Practical Personalized Medicine

- knowledge base;
- ontology;
- pattern recognition;
- patient profiles.
6. Closing the Loop
7. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Katsnelson, A. Momentum grows to make ‘personalized’ medicine more ‘precise’. Nat. Med. 2013, 19, 249. [Google Scholar] [CrossRef] [PubMed]
- Auffray, C.; Chen, Z.; Hood, L. Systems medicine: The future of medical genomics and healthcare. Genome Med. 2009, 1, 2. [Google Scholar] [CrossRef] [PubMed]
- Chin, L.; Andersen, J.N.; Futreal, P.A. Cancer genomics: From discovery science to personalized medicine. Nat. Med. 2011, 17, 297–303. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.; Snyder, M. Promise of personalized omics to precision medicine. Wiley Interdiscip. Rev. 2013, 5, 73–82. [Google Scholar] [CrossRef] [PubMed]
- Seo, D.; Ginsburg, G.S. Genomic medicine: Bringing biomarkers to clinical medicine. Curr. Opin. Chem. Biol. 2005, 9, 381–386. [Google Scholar] [CrossRef] [PubMed]
- Tian, Q.; Price, N.D.; Hood, L. Systems cancer medicine: Towards realization of predictive, preventive, personalized and participatory (P4) medicine. J. Intern. Med. 2012, 271, 111–121. [Google Scholar] [CrossRef] [PubMed]
- International Human Genome Sequencing Consortium. Finishing the euchromatic sequence of the human genome. Nature 2004, 431, 931–945. [Google Scholar] [CrossRef] [PubMed]
- Lander, E.S.; Linton, L.M.; Birren, B.; Nusbaum, C.; Zody, M.C.; Baldwin, J.; Devon, K.; Dewar, K.; Doyle, M.; FitzHugh, W.; et al. Initial sequencing and analysis of the human genome. Nature 2001, 409, 860–921. [Google Scholar] [PubMed]
- Venter, J.C.; Adams, M.D.; Myers, E.W.; Li, P.W.; Mural, R.J.; Sutton, G.G.; Smith, H.O.; Yandell, M.; Evans, C.A.; Holt, R.A.; et al. The Sequence of the Human Genome. Science 2001, 291, 1304–1351. [Google Scholar] [CrossRef] [PubMed]
- Ginsburg, G.S.; Willard, H.F. Genomic and personalized medicine: Foundations and applications. Transl. Res. 2009, 154, 277–287. [Google Scholar] [CrossRef] [PubMed]
- Emmert-Streib, F.; Tuomisto, L.; Yli-Harja, O. The Need for Formally Defining ‘Modern Medicine’ by Means of Experimental Design. Front. Genet. 2016, 7, 60. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Angulo, A.M.; Hennessy, B.T.; Mills, G.B. Future of Personalized Medicine in Oncology: A Systems Biology Approach. J. Clin. Oncol. 2010, 28, 2777–2783. [Google Scholar] [CrossRef] [PubMed]
- Welch, B.M.; Kawamoto, K. Clinical decision support for genetically guided personalized medicine: A systematic review. J. Am. Med. Inform. Assoc. 2012, 20, 388–400. [Google Scholar] [CrossRef] [PubMed]
- Lesko, L.J. Personalized medicine: Elusive dream or imminent reality? Clin. Pharmacol. Ther. 2007, 81, 807–816. [Google Scholar] [CrossRef] [PubMed]
- Sorlie, T.; Tibshirani, R.; Parker, J.; Hastie, T.; Marron, J.S.; Nobel, A.; Deng, S.; Johnsen, H.; Pesich, R.; Geisler, S.; et al. Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc. Natl. Acad. Sci. USA 2003, 100, 8418–8423. [Google Scholar] [CrossRef] [PubMed]
- Perou, C.M.; Sorlie, T.; Eisen, M.B.; Van De Rijn, M.; Jeffrey, S.S.; Rees, C.A.; Pollack, J.R.; Ross, D.T.; Johnsen, H.; Akslen, L.A.; et al. Molecular portraits of human breast tumours. Nature 2000, 406, 747–752. [Google Scholar] [CrossRef] [PubMed]
- Curtis, C.; Shah, S.P.; Chin, S.F.; Turashvili, G.; Rueda, O.M.; Dunning, M.J.; Speed, D.; Lynch, A.G.; Samarajiwa, S.; Yuan, Y.; et al. The genomic and transcriptomic architecture of 2000 breast tumours reveals novel subgroups. Nature 2012, 486, 346. [Google Scholar] [CrossRef] [PubMed]
- Prat, A.; Parker, J.; Karginova, O.; Fan, C.; Livasy, C.; Herschkowitz, J.; He, X.; Perou, C. Phenotypic and molecular characterization of the claudin-low intrinsic subtype of breast cancer. Breast Cancer Res. 2010, 12, R68. [Google Scholar] [CrossRef] [PubMed]
- The Cancer Genome Atlas Research Network. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 2008, 455, 1061–1068. [Google Scholar] [CrossRef] [PubMed]
- Huck, J.H.; Verhoeven, N.M.; Struys, E.A.; Salomons, G.S.; Jakobs, C.; van der Knaap, M.S. Ribose-5-phosphate isomerase deficiency: New inborn error in the pentose phosphate pathway associated with a slowly progressive leukoencephalopathy. Am. J. Hum. Genet. 2004, 74, 745–751. [Google Scholar] [CrossRef] [PubMed]
- Wamelink, M.M.; Grüning, N.M.; Jansen, E.E.; Bluemlein, K.; Lehrach, H.; Jakobs, C.; Ralser, M. The difference between rare and exceptionally rare: Molecular characterization of ribose 5-phosphate isomerase deficiency. J. Mol. Med. 2010, 88, 931–939. [Google Scholar] [CrossRef] [PubMed]
- Schieppati, A.; Henter, J.I.; Daina, E.; Aperia, A. Why rare diseases are an important medical and social issue. Lancet 2008, 371, 2039–2041. [Google Scholar] [CrossRef]
- Lehman, E. Testing Statistical Hypotheses; Springer: Berlin, Germany, 2005. [Google Scholar]
- Mankoff, S.P.; Brander, C.; Ferrone, S.; Marincola, F.M. Lost in translation: Obstacles to translational medicine. J. Transl. Med. 2004, 2, 14. [Google Scholar] [CrossRef] [PubMed]
- R Development Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2008; ISBN 3-900051-07-0. [Google Scholar]
- Gentleman, R.; Carey, V.; Bates, D.E.A. Bioconductor: Open software development for computational biology and bioinformatics. Genome Biol. 2004, 5, R80. [Google Scholar] [CrossRef] [PubMed]
- Diggle, P.J.; Zeger, S.L. Embracing the concept of reproducible research. Biostatistics 2010, 11, 375. [Google Scholar] [CrossRef] [PubMed]
- Peng, R.D. Reproducible Research in Computational Science. Science 2011, 334, 1226–1227. [Google Scholar] [CrossRef] [PubMed]
- Casscells, W.; Schoenberger, A.; Graboys, T.B. Interpretation by Physicians of Clinical Laboratory Results. N. Engl. J. Med. 1978, 299, 999–1001. [Google Scholar] [CrossRef] [PubMed]
- Manrai, A.; Bhatia, G.; Strymish, J.; Kohane, I.; Jain, S. Medicine’s uncomfortable relationship with math: Calculating positive predictive value. JAMA Intern. Med. 2014, 174, 991–993. [Google Scholar] [CrossRef] [PubMed]
- Metzker, M. Sequencing technologies-the next generation. Nat. Rev. Genet. 2009, 11, 31–46. [Google Scholar] [CrossRef] [PubMed]
- Stupnikov, A.; Tripathi, S.; de Matos Simoes, R.; McArt, D.; Salto-Tellez, M.; Glazko, G.; Emmert-Streib, F. samExploreR: Exploring reproducibility and robustness of RNA-seq results based on SAM files. Bioinformatics 2016, 32, 3345–3347. [Google Scholar] [CrossRef] [PubMed]
- Quackenbush, J. Microarray analysis and tumor classification. N. Engl. J. Med. 2006, 345, 2463–2472. [Google Scholar] [CrossRef] [PubMed]
- Emmert-Streib, F.; de Matos Simoes, R.; Glazko, G.; McDade, S.; Haibe-Kains, B.; Holzinger, A.; Dehmer, M.; Campbell, F. Functional and genetic analysis of the colon cancer network. BMC Bioinform. 2014, 15, 6. [Google Scholar] [CrossRef] [PubMed]
- Chandrasekaran, B.; Josephson, J.R.; Benjamins, V.R. What are ontologies, and why do we need them? IEEE Intell. Syst. Their Appl. 1999, 14, 20–26. [Google Scholar] [CrossRef]
- Fonseca, F. The double role of ontologies in information science research. J. Am. Soc. Inform. Sci. Technol. 2007, 58, 786–793. [Google Scholar] [CrossRef]
- Kieseberg, P.; Malle, B.; Frühwirt, P.; Weippl, E.; Holzinger, A. A tamper-proof audit and control system for the doctor in the loop. Brain Inform. 2016, 3, 269–279. [Google Scholar] [CrossRef] [PubMed]
- Holzinger, A.; Dehmer, M.; Jurisica, I. Knowledge discovery and interactive data mining in bioinformatics-state-of-the-art, future challenges and research directions. BMC Bioinform. 2014, 15, I1. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Emmert-Streib, F.; Dehmer, M. A Machine Learning Perspective on Personalized Medicine: An Automized, Comprehensive Knowledge Base with Ontology for Pattern Recognition. Mach. Learn. Knowl. Extr. 2019, 1, 149-156. https://doi.org/10.3390/make1010009
Emmert-Streib F, Dehmer M. A Machine Learning Perspective on Personalized Medicine: An Automized, Comprehensive Knowledge Base with Ontology for Pattern Recognition. Machine Learning and Knowledge Extraction. 2019; 1(1):149-156. https://doi.org/10.3390/make1010009
Chicago/Turabian StyleEmmert-Streib, Frank, and Matthias Dehmer. 2019. "A Machine Learning Perspective on Personalized Medicine: An Automized, Comprehensive Knowledge Base with Ontology for Pattern Recognition" Machine Learning and Knowledge Extraction 1, no. 1: 149-156. https://doi.org/10.3390/make1010009
APA StyleEmmert-Streib, F., & Dehmer, M. (2019). A Machine Learning Perspective on Personalized Medicine: An Automized, Comprehensive Knowledge Base with Ontology for Pattern Recognition. Machine Learning and Knowledge Extraction, 1(1), 149-156. https://doi.org/10.3390/make1010009





