Overcoming Nonlinear Dynamics in Diabetic Retinopathy Classification: A Robust AI-Based Model with Chaotic Swarm Intelligence Optimization and Recurrent Long Short-Term Memory
Abstract
1. Introduction
- The presence of chaos in the images of each DR disease class along with the healthy class is revealed by fractal analysis.
- Feature groups are extracted for each family by applying two-dimensional stationary wavelet transform (2D-SWT) with biorthogonal, reverse biorthogonal, Daubechies, Coiflet, symlet, and Fejer–Korovkin wavelet families to the dataset consisting of each DR disease class together with the healthy class.
- The entropy- and statistical-based feature groups extracted for the 12 image matrices obtained as a result of three-level decomposition contain nonlinear dynamics representing the DR disease classes.
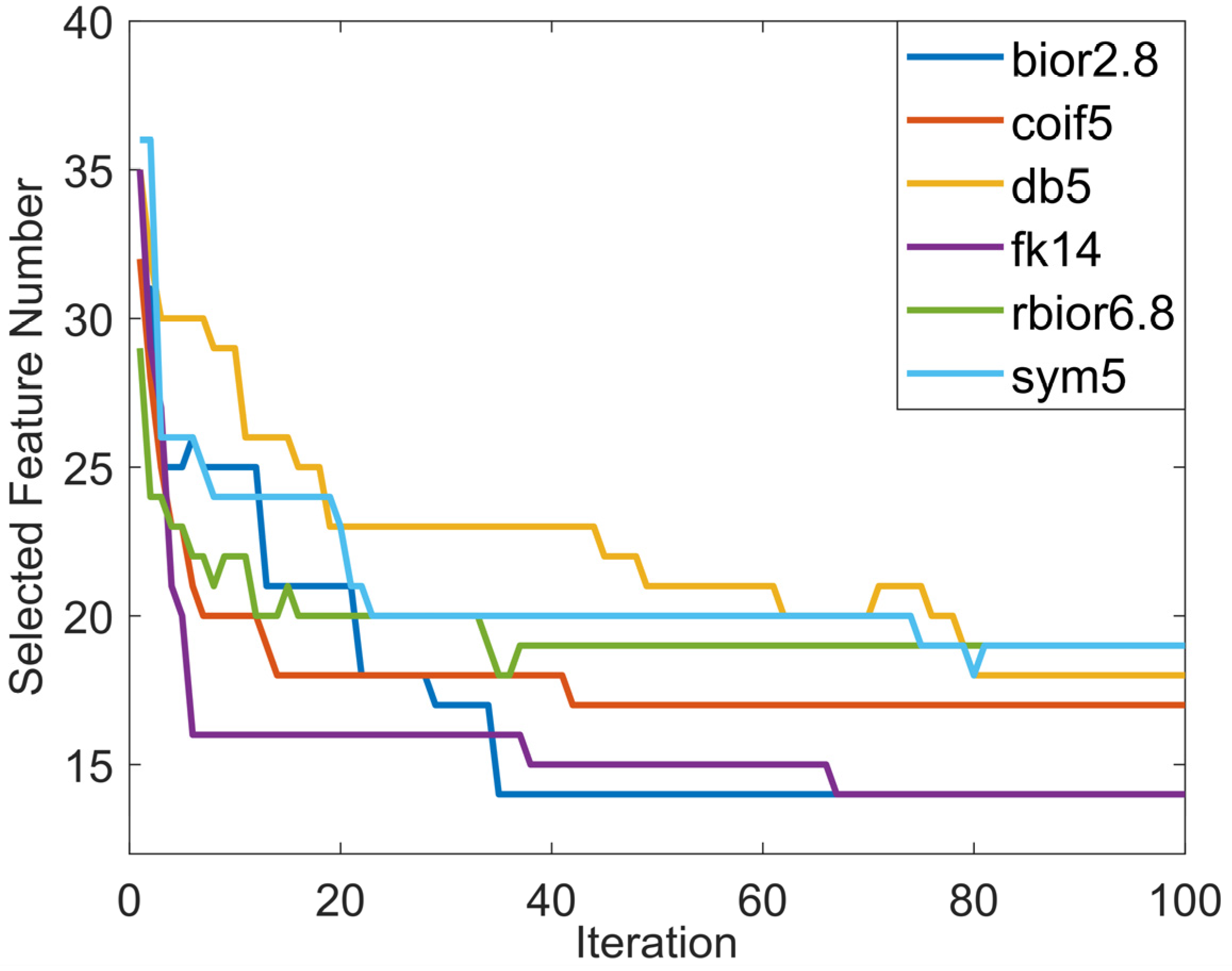
- The features that keep the model performance high are selected with a wrapper approach consisting of the chaotic particle swarm optimization (CPSO) and k-nearest neighbor (kNN) algorithms in order to keep the computational complexity of the model at a minimum and to cope with the chaoticity in the fundus images.
- The most suitable chaotic map, which improves the convergence speed and optimum solution of the optimization algorithm, is determined and included in the optimization process to obtain the highest classification accuracy with the least features.
- The effect of the features selected by the chaotic wrapper approach on the model performance is examined for each wavelet family.
- The selected optimum feature vectors are finally fed into the recurrent neural network-long short-term memory (RNN-LSTM) for classifying DR disease sub-types like PDR, mild NPDR, moderate NPDR, and severe NPDR.
- The model with the best performance is proposed, which includes the three-level 2D-SWT technique based on the ‘bior 2.8’ wavelet family, the wrapper approach consisting of logistic-chaotic-map-based CPSO and kNN, and the RNN-LSTM network for DR disease classification.
- It is shown by experimental results that the proposed model can cope with nonlinear dynamics, has low computational complexity, and can be used in real-time applications thanks to these features.
2. Background and Related Works
3. Framework of the Diagnosis and Classification Algorithm for Diabetic Retinopathy
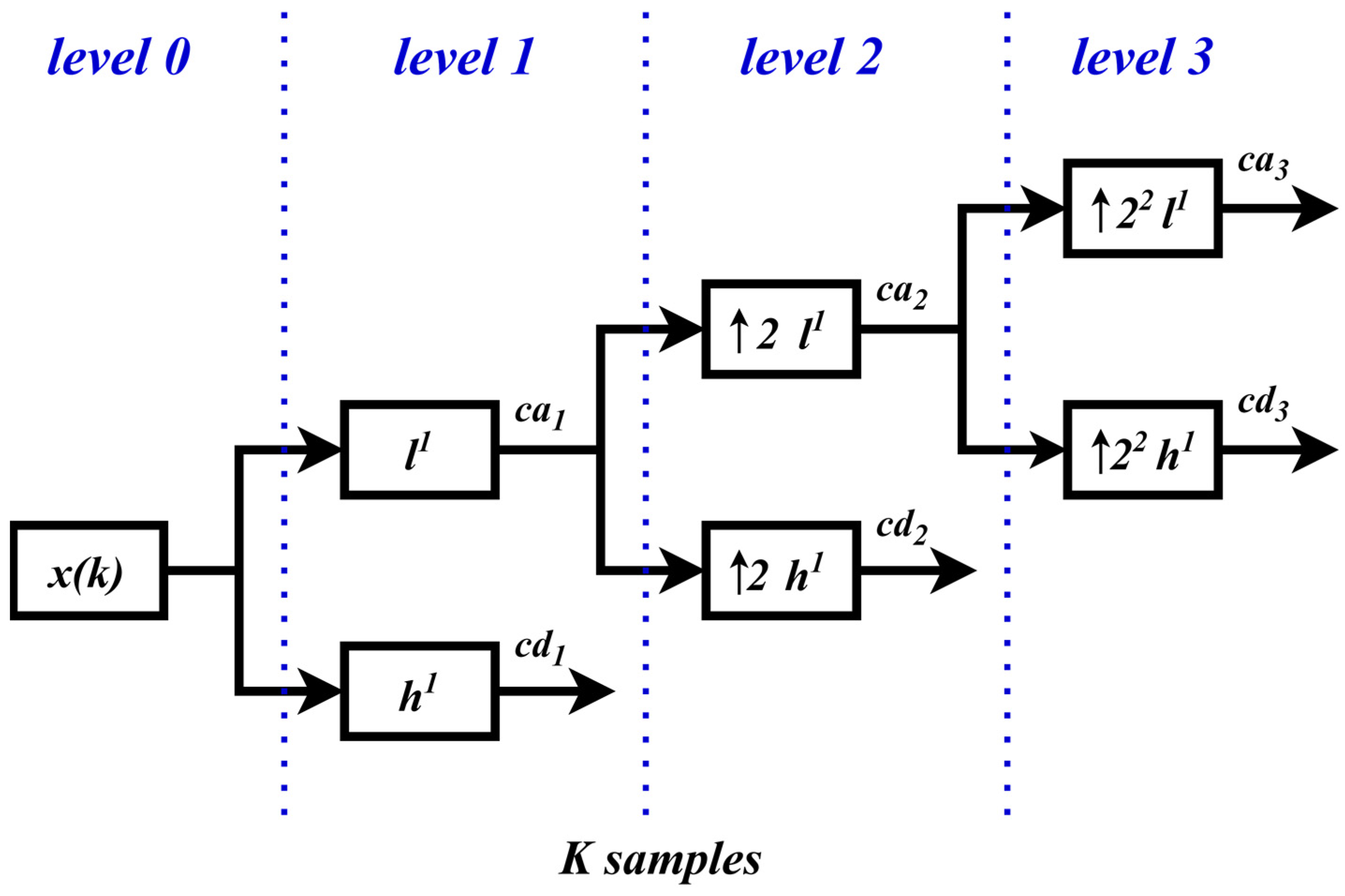
3.1. Stationary Wavelet Transform
3.2. Multiresolution Analysis
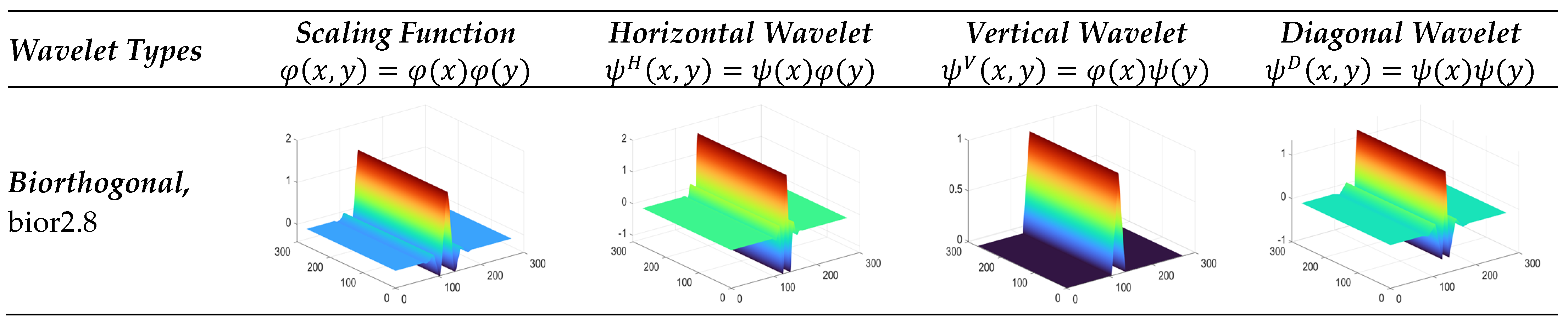
3.3. Wavelet Families
3.3.1. Biorthogonal Wavelet
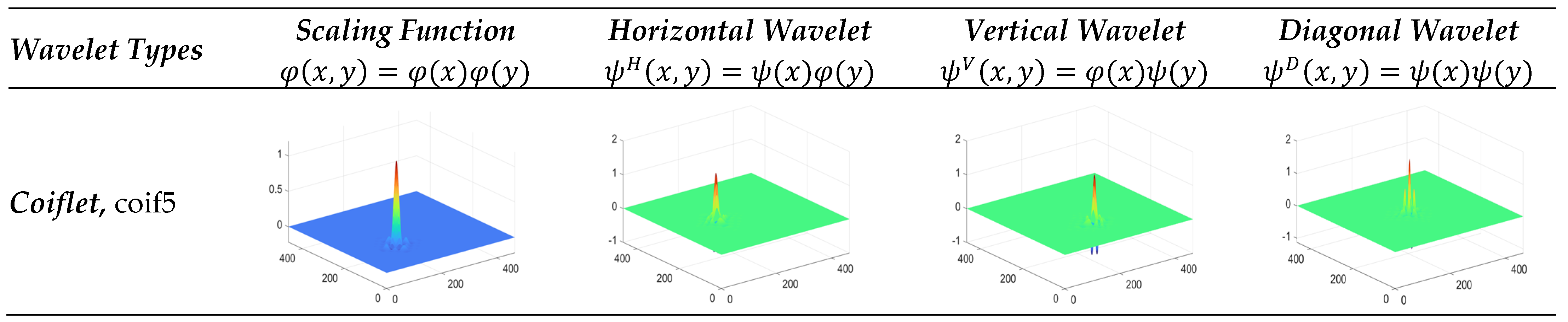
3.3.2. Coiflet Wavelet
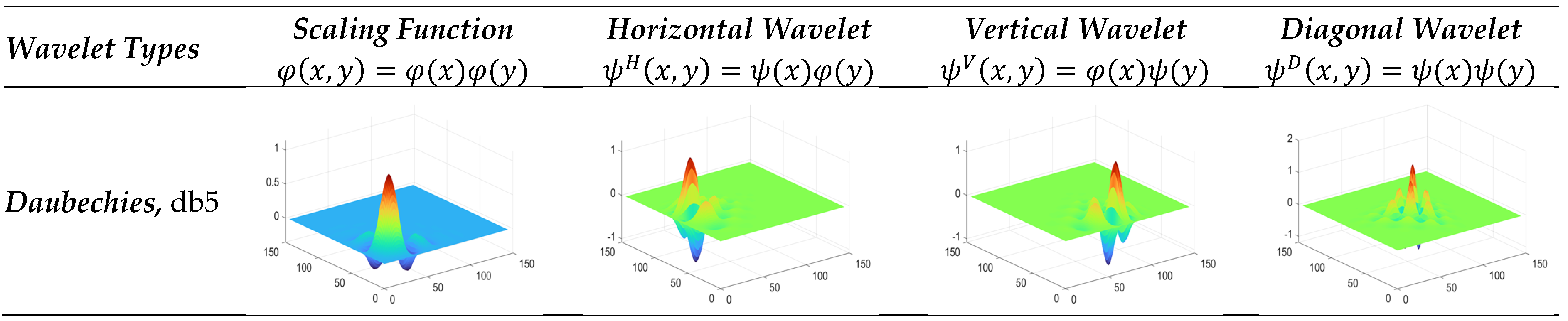
3.3.3. Daubechies Wavelet
3.3.4. Fejer–Korovkin Wavelet
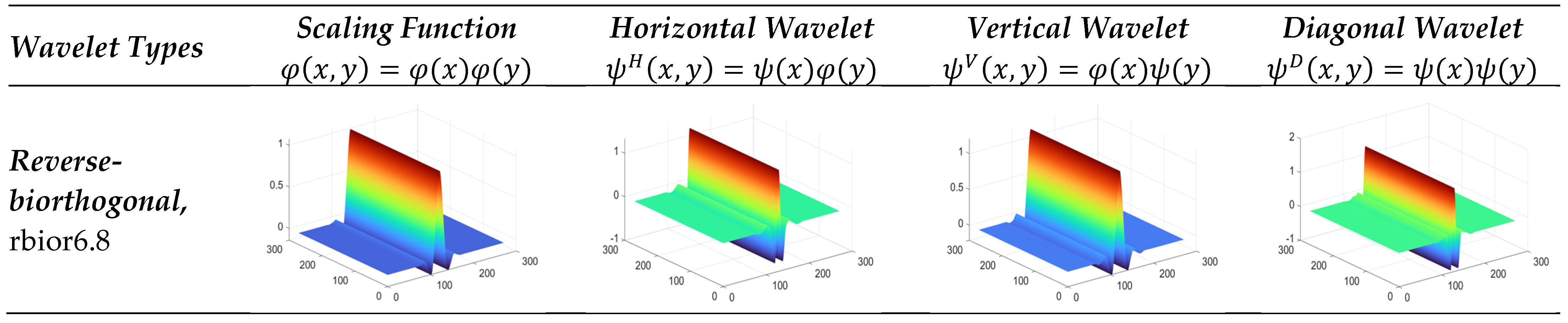
3.3.5. Reverse Biorthogonal Wavelet
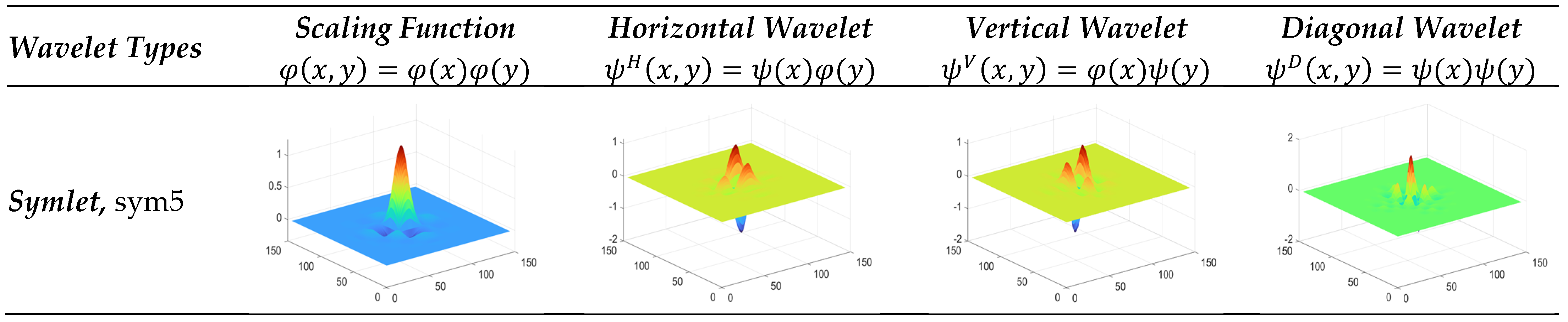
3.3.6. Symlet Wavelet
3.4. Two-Dimensional Stationary Wavelet Transform (2D-SWT)
3.5. Chaotic Particle Swarm Optimization
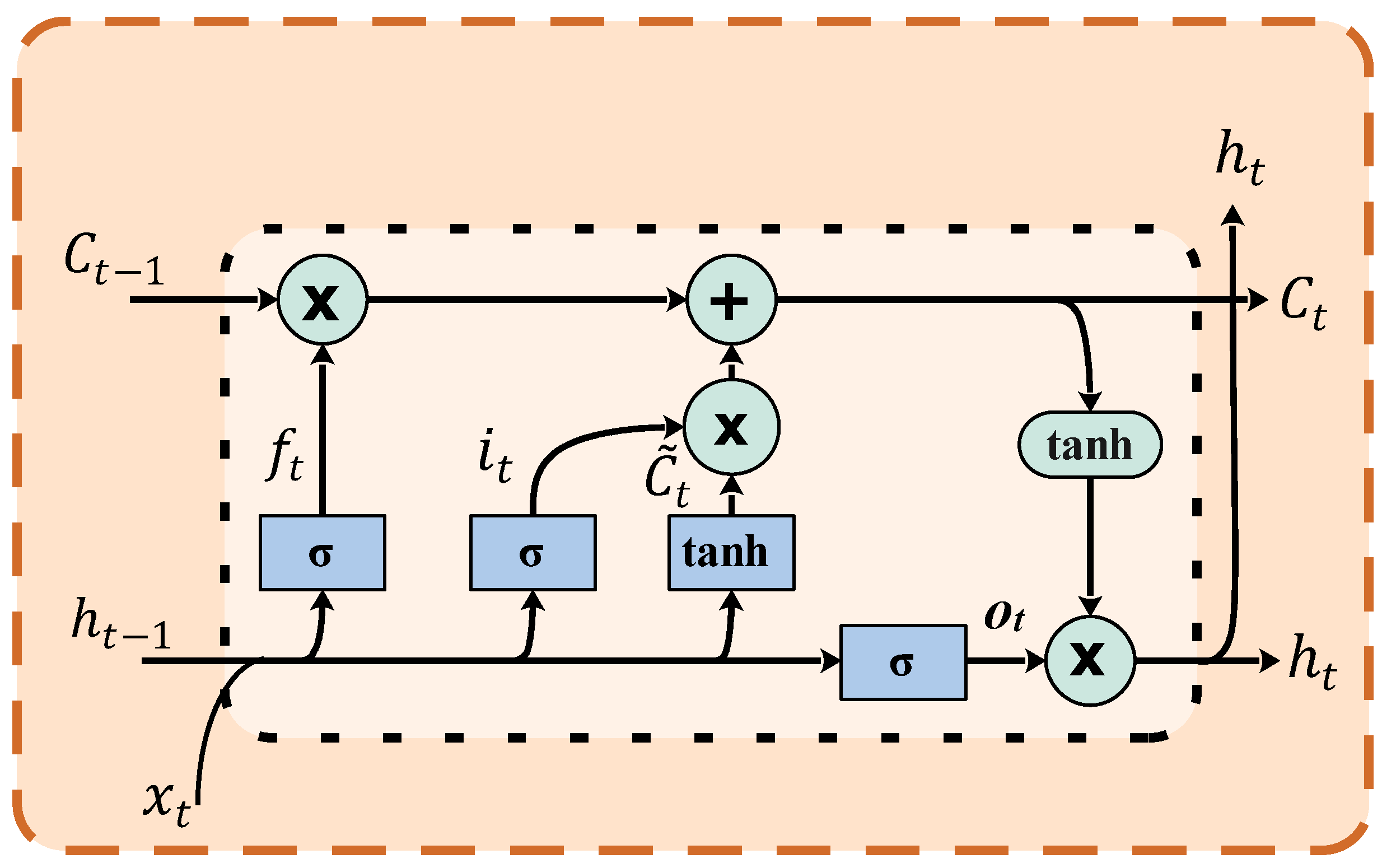
3.6. Classification Using Recurrent Neural Network-Long Short-Term Memory (RNN-LSTM)
- represents the input at time step .
- denotes the hidden state at time step .
- represents the cell state at time step .
- , , , and denote the weight matrices for the input at time step associated with the input gate, forget gate, output gate, and candidate cell state, respectively.
- , , , and represent the weight matrices for the hidden state at time step associated with the input gate, forget gate, output gate, and candidate cell state, respectively.
- , , , and denote the bias vectors for the input gate, forget gate, output gate, and candidate cell state, respectively.
- (i)
- Input gate ()
- (ii)
- Forget gate ()
- (iii)
- Output gate ()
- (iv)
- Candidate cell state ()
- (v)
- Cell state update ()
- (vi)
- Hidden state ()
3.7. Performance Metrics for Classification
3.8. Framework of the Proposed DR Disease Classification Model
4. Results and Discussion
4.1. Dataset
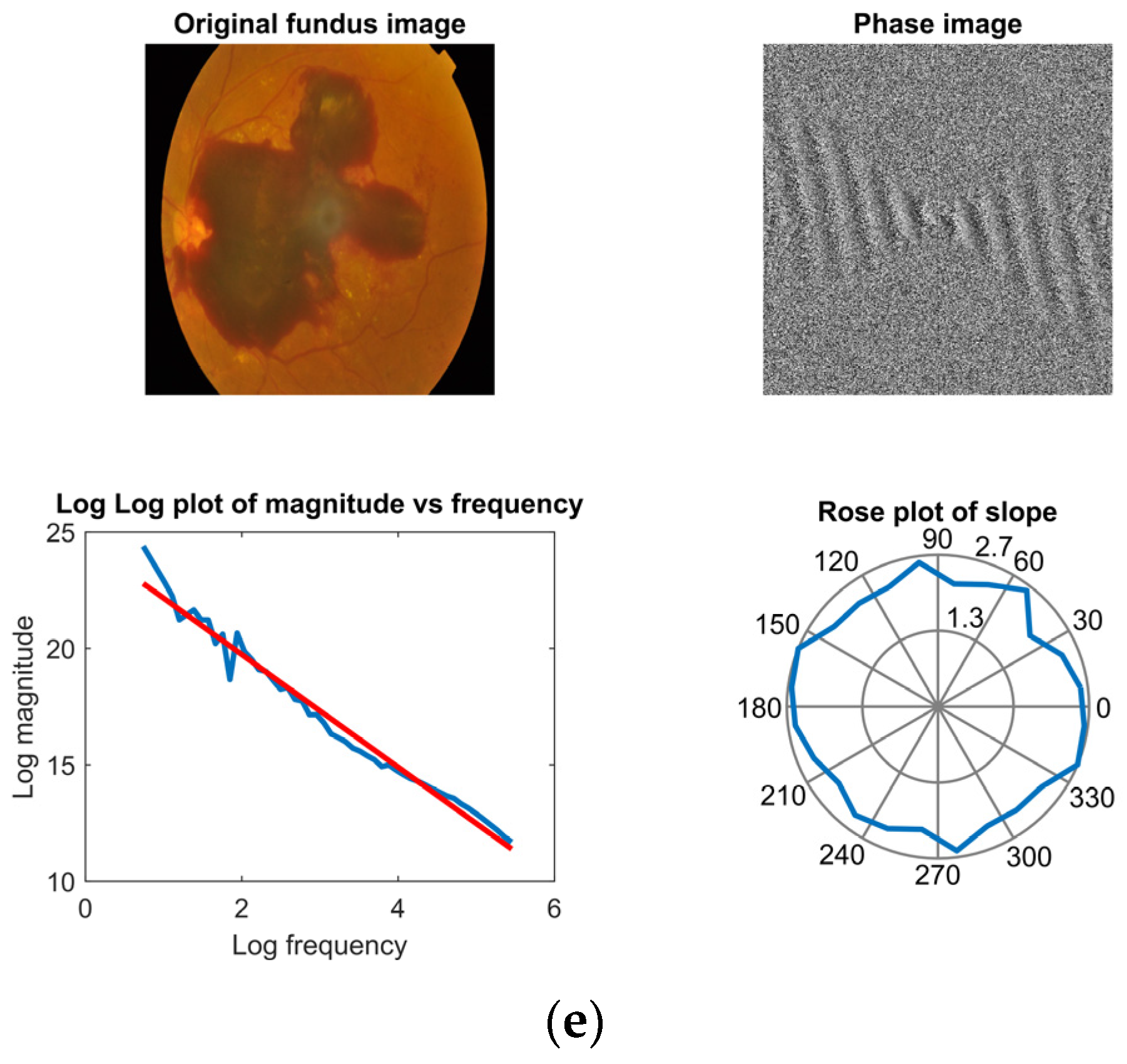
4.2. Fractal Dimension Analysis with Fourier Power Spectrum
4.3. Feature Extraction Applying 2D-SWT with Wavelet Families
4.4. Feature Selection with CPSO-kNN
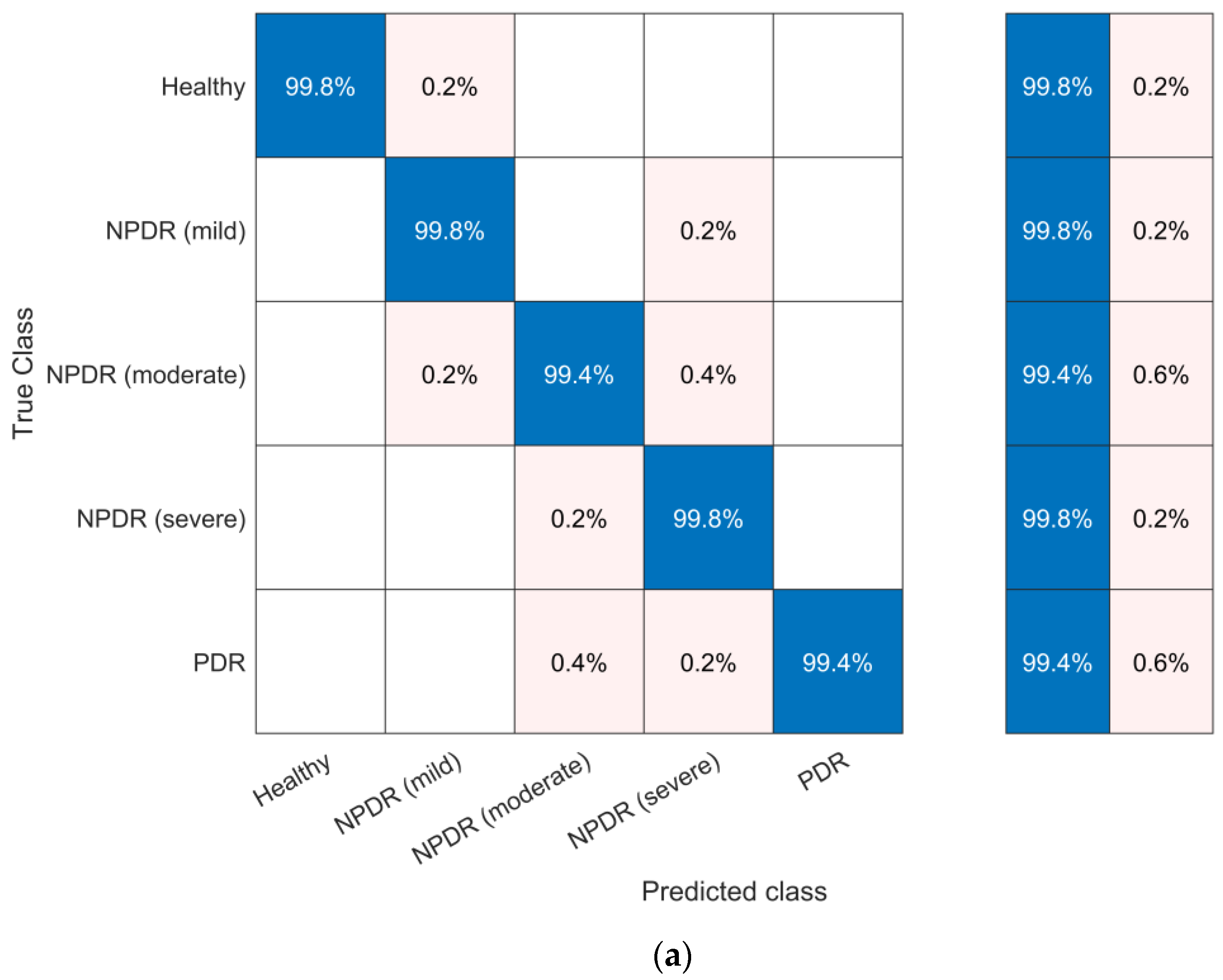
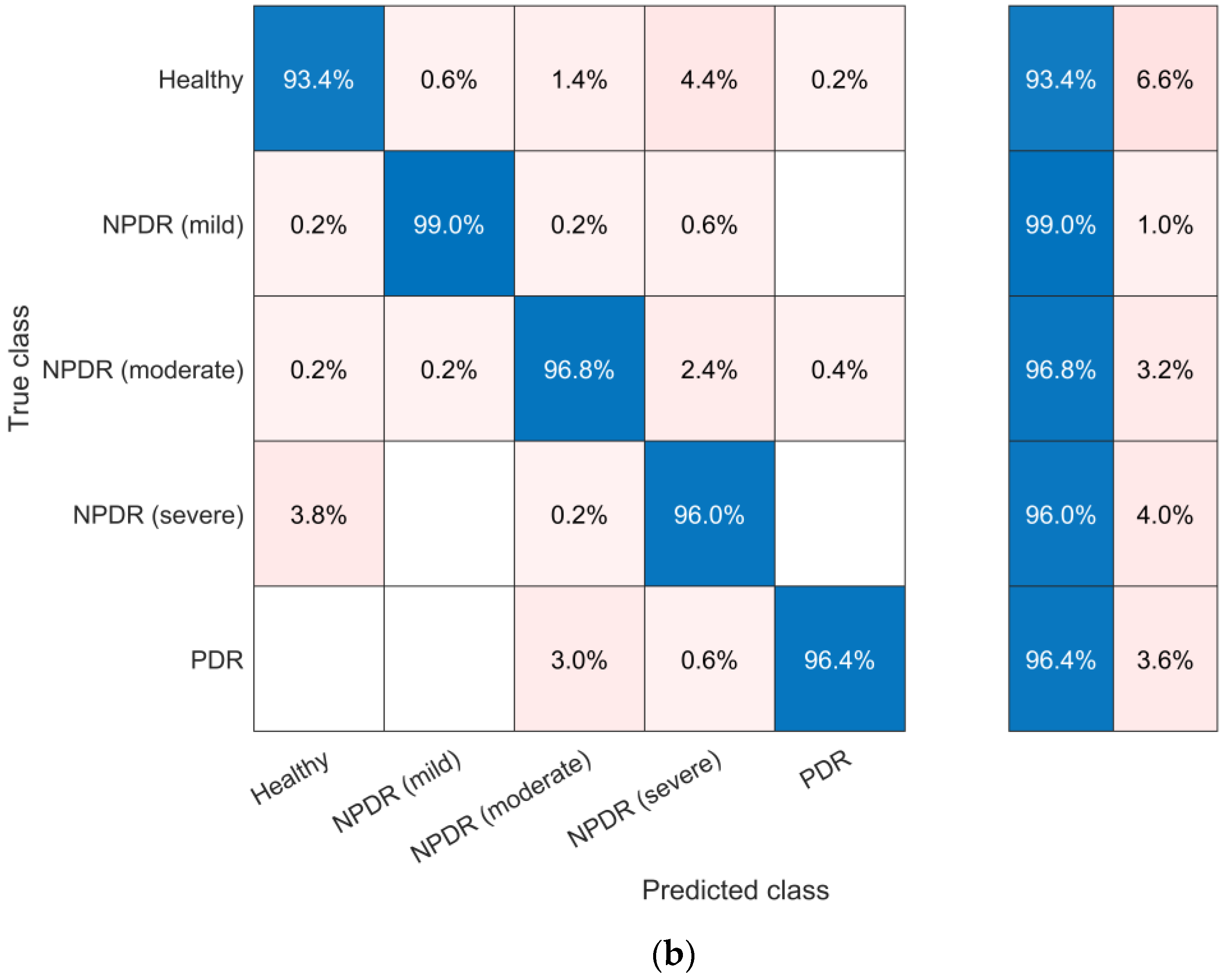
4.5. Evaluation and Discussion of Classification Models
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Saeedi, P.; Petersohn, I.; Salpea, P.; Malanda, B.; Karuranga, S.; Unwin, N.; Colagiuri, S.; Guariguata, L.; Motala, A.A.; Ogurtsova, K.; et al. Global and regional diabetes prevalence estimates for 2019 and projections for 2030 and 2045: Results from the International Diabetes Federation Diabetes Atlas. Diabetes Res. Clin. Pract. 2019, 157, 107843. [Google Scholar] [CrossRef]
- Wong, T.Y.; Cheung, C.M.G.; Larsen, M.; Sharma, S.; Simó, R. Diabetic retinopathy (Primer). Nat. Rev. Dis. Primers 2016, 2, 16012. [Google Scholar] [CrossRef] [PubMed]
- Zago, G.T.; Andreão, R.V.; Dorizzi, B.; Salles, E.O.T. Diabetic retinopathy detection using red lesion localization and convolutional neural networks. Comput. Biol. Med. 2020, 116, 103537. [Google Scholar] [CrossRef] [PubMed]
- Lee, R.; Wong, T.Y.; Sabanayagam, C. Epidemiology of diabetic retinopathy, diabetic macular edema and related vision loss. Eye Vis. 2015, 2, 1–25. [Google Scholar] [CrossRef]
- Kobrin Klein, B.E. Overview of epidemiologic studies of diabetic retinopathy. Ophthalmic Epidemiol. 2007, 14, 179–183. [Google Scholar] [CrossRef] [PubMed]
- Wan, S.; Liang, Y.; Zhang, Y. Deep convolutional neural networks for diabetic retinopathy detection by image classification. Comput. Electr. Eng. 2018, 72, 274–282. [Google Scholar] [CrossRef]
- Nazir, T.; Irtaza, A.; Shabbir, Z.; Javed, A.; Akram, U.; Mahmood, M.T. Diabetic retinopathy detection through novel tetragonal local octa patterns and extreme learning machines. Artif. Intell. Med. 2019, 99, 101695. [Google Scholar] [CrossRef]
- Aiello, L.M. Perspectives on diabetic retinopathy. Am. J. Ophthalmol. 2003, 136, 122–135. [Google Scholar] [CrossRef]
- Stratton, I.M.; Kohner, E.M.; Aldington, S.J.; Turner, R.C.; Holman, R.R.; Manley, S.E.; Matthews, D.R. for the UKPDS Group. UKPDS 50: Risk factors for incidence and progression of retinopathy in type II diabetes over 6 years from diagnosis. Diabetologia 2001, 44, 156–163. [Google Scholar] [CrossRef]
- Wu, B.; Zhu, W.; Shi, F.; Zhu, S.; Chen, X. Automatic detection of microaneurysms in retinal fundus images. Comput. Med. Imaging Graph. 2017, 55, 106–112. [Google Scholar] [CrossRef]
- García, M.; Sánchez, C.I.; López, M.I.; Abásolo, D.; Hornero, R. Neural network-based detection of hard exudates in retinal images. Comput. Methods Programs Biomed. 2009, 93, 9–19. [Google Scholar] [CrossRef]
- Faust, O.; Acharya U, R.; Ng, E.Y.K.; Ng, K.H.; Suri, J.S. Algorithms for the automated detection of diabetic retinopathy using digital fundus images: A review. J. Med. Syst. 2012, 36, 145–157. [Google Scholar] [CrossRef]
- Bilal, A.; Zhu, L.; Deng, A.; Lu, H.; Wu, N. AI-based automatic detection and classification of diabetic retinopathy using U-Net and deep learning. Symmetry 2022, 14, 1427. [Google Scholar] [CrossRef]
- Math, L.; Fatima, R. Adaptive machine learning classification for diabetic retinopathy. Multimed. Tools Appl. 2021, 80, 5173–5186. [Google Scholar] [CrossRef]
- Qiao, L.; Zhu, Y.; Zhou, H. Diabetic retinopathy detection using prognosis of microaneurysm and early diagnosis system for non-proliferative diabetic retinopathy based on deep learning algorithms. IEEE Access 2020, 8, 104292–104302. [Google Scholar] [CrossRef]
- Gadekallu, T.R.; Khare, N.; Bhattacharya, S.; Singh, S.; Maddikunta, P.K.R.; Ra, I.H.; Alazab, M. Early detection of diabetic retinopathy using PCA-firefly based deep learning model. Electronics 2020, 9, 274. [Google Scholar] [CrossRef]
- Kukkar, A.; Gupta, D.; Beram, S.M.; Soni, M.; Singh, N.K.; Sharma, A.; Neware, R.; Shabaz, M.; Rizwan, A. Optimizing deep learning model parameters using socially implemented IoMT systems for diabetic retinopathy classification problem. IEEE Trans. Comput. Soc. Syst. 2022. Early Access. [Google Scholar]
- Gundluru, N.; Rajput, D.S.; Lakshmanna, K.; Kaluri, R.; Shorfuzzaman, M.; Uddin, M.; Rahman Khan, M.A. Enhancement of detection of diabetic retinopathy using Harris hawks optimization with deep learning model. Comput. Intell. Neurosci. 2022, 2022, 8512469. [Google Scholar] [CrossRef] [PubMed]
- Uppamma, P.; Bhattacharya, S. Diabetic retinopathy detection: A blockchain and African vulture optimization algorithm-based deep learning framework. Electronics 2023, 12, 742. [Google Scholar] [CrossRef]
- Gupta, S.; Thakur, S.; Gupta, A. Optimized hybrid machine learning approach for smartphone based diabetic retinopathy detection. Multimed. Tools Appl. 2022, 81, 14475–14501. [Google Scholar] [CrossRef]
- Beevi, S.Z. Multi-Level severity classification for diabetic retinopathy based on hybrid optimization enabled deep learning. Biomed. Signal Process. Control 2023, 84, 104736. [Google Scholar] [CrossRef]
- Rachapudi, V.; Rao, K.S.; Rao, T.S.M.; Dileep, P.; Deepika Roy, T.L. Diabetic retinopathy detection by optimized deep learning model. Multimed. Tools Appl. 2023, 82, 27949–27971. [Google Scholar] [CrossRef]
- Williams, J.R.; Amaratunga, K. Introduction to wavelets in engineering. Int. J. Numer. Methods Eng. 1994, 37, 2365–2388. [Google Scholar] [CrossRef]
- Kim, C.H.; Aggarwal, R. Wavelet transforms in power systems. Part 1: General introduction to the wavelet transforms. Power Eng. J. 2000, 14, 81–87. [Google Scholar]
- Mallat, S. A Wavelet Tour of Signal Processing, 2nd ed.; Elsevier: Berkeley, CA, USA, 1999; ISBN 978-0-12-466606-1. [Google Scholar]
- Nason, G.P.; Silverman, B.W. The discrete wavelet transform in S. J. Comput. Graph. Stat. 1994, 3, 163–191. [Google Scholar]
- Vetterli, M.; Herley, C. Wavelets and filter banks: Theory and design. IEEE Trans. Signal Process. 1992, 40, 2207–2232. [Google Scholar] [CrossRef]
- Nason, G.P.; Silverman, B.W. The stationary wavelet transform and some statistical applications. In Wavelets and Statistics; Springer: New York, NY, USA, 1995; pp. 281–299. [Google Scholar]
- Pesquet, J.C.; Krim, H.; Carfantan, H. Time-invariant orthonormal wavelet representations. IEEE Trans. Signal Process. 1996, 44, 1964–1970. [Google Scholar] [CrossRef]
- Merah, M.; Abdelmalik, T.A.; Larbi, B.H. R-peaks detection based on stationary wavelet transform. Comput. Methods Programs Biomed. 2015, 121, 149–160. [Google Scholar] [CrossRef]
- Mallat, S.G. A theory for multiresolution signal decomposition: The wavelet representation. IEEE Trans. Pattern Anal. Mach. Intell. 1989, 11, 674–693. [Google Scholar] [CrossRef]
- Keinert, F. Biorthogonal wavelets for fast matrix computations. Appl. Comput. Harmon. Anal. 1994, 1, 147–156. [Google Scholar] [CrossRef]
- Monzón, L.; Beylkin, G.; Hereman, W. Compactly supported wavelets based on almost interpolating and nearly linear phase filters (coiflets). Appl. Comput. Harmon. Anal. 1999, 7, 184–210. [Google Scholar] [CrossRef]
- Daubechies, I. Ten Lectures on Wavelets; Society for Industrial and Applied Mathematics: Philadelphia, PA, USA, 1992; ISBN 0-89871-274-2. [Google Scholar]
- Daubechies, I. Orthonormal bases of compactly supported wavelets. Commun. Pure Appl. Math. 1988, 41, 909–996. [Google Scholar] [CrossRef]
- Daubechies, I. The wavelet transform, time-frequency localization and signal analysis. IEEE Trans. Inf. Theory 1990, 36, 961–1005. [Google Scholar] [CrossRef]
- Nielsen, M. On the construction and frequency localization of finite orthogonal quadrature filters. J. Approx. Theory 2001, 108, 36–52. [Google Scholar] [CrossRef]
- Szewczyk, R.; Grabowski, K.; Napieralska, M.; Sankowski, W.; Zubert, M.; Napieralski, A. A reliable iris recognition algorithm based on reverse biorthogonal wavelet transform. Pattern Recognit. Lett. 2012, 33, 1019–1026. [Google Scholar] [CrossRef]
- Yang, C.; Liu, P.; Yin, G.; Jiang, H.; Li, X. Defect detection in magnetic tile images based on stationary wavelet transform. NDT E Int. 2016, 83, 78–87. [Google Scholar] [CrossRef]
- Kennedy, J.; Eberhart, R. Particle swarm optimization. In Proceedings of the ICNN’95 International Conference on Neural Networks, Perth, Australia, 27 November–1 December 1995. [Google Scholar]
- dos Santos Coelho, L.; Herrera, B.M. Fuzzy identification based on a chaotic particle swarm optimization approach applied to a nonlinear yo-yo motion system. IEEE Trans. Ind. Electron. 2007, 54, 3234–3245. [Google Scholar] [CrossRef]
- Wei, D.; Wang, B.; Lin, G.; Liu, D.; Dong, Z.; Liu, H.; Liu, Y. Research on unstructured text data mining and fault classification based on RNN-LSTM with malfunction inspection report. Energies 2017, 10, 406. [Google Scholar] [CrossRef]
- Karasu, S.; Altan, A. Crude oil time series prediction model based on LSTM network with chaotic Henry gas solubility optimization. Energy 2022, 242, 122964. [Google Scholar] [CrossRef]
- Hochreiter, S.; Schmidhuber, J. Long short-term memory. Neural Comput. 1997, 9, 1735–1780. [Google Scholar] [CrossRef]
- Hua, Y.; Mou, L.; Zhu, X.X. Recurrently exploring class-wise attention in a hybrid convolutional and bidirectional LSTM network for multi-label aerial image classification. ISPRS J. Photogramm. Remote Sens. 2019, 149, 188–199. [Google Scholar] [CrossRef]
- Sokolova, M.; Lapalme, G. A systematic analysis of performance measures for classification tasks. Inf. Process. Manag. 2009, 45, 427–437. [Google Scholar] [CrossRef]
- APTOS 2019 Blindness Detection. Available online: https://www.kaggle.com/c/aptos2019-blindness-detection (accessed on 29 June 2023).
- Quevedo, R.; Carlos, L.G.; Aguilera, J.M.; Cadoche, L. Description of food surfaces and microstructural changes using fractal image texture analysis. J. Food Eng. 2022, 53, 361–371. [Google Scholar] [CrossRef]







| No | Name | Description | Range |
|---|---|---|---|
| 1 | Logistic | ||
| 2 | Chebyshev | ||
| 3 | Sine | ||
| 4 | Sinusoidal | ||
| 5 | Singer | ||
| 6 | Iterative | ||
| 7 | Circle | ||
| 8 | Tent | ||
| 9 | Gauss/mouse | ||
| 10 | Piecewise |
| Wavelet Family | Filter Length |
|---|---|
| Biorthogonal | (1.) 1, 3, 5, (2.) 2, 4, 6, 8, (3.) 1, 3, 5, 7, 9, (4.) 4, (5.) 5, (6.) 8 |
| Coiflet | 1, 2, 3, 4, 5 |
| Daubechies | 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 |
| Fejer–Korovkin | 4, 6, 8, 14, 18, 22 |
| Reverse biorthogonal | (1.) 1, 3, 5, (2.) 2, 4, 6, 8, (3.) 1, 3, 5, 7, 9, (4.) 4, (5.) 5, (6.) 8 |
| Symlet | 2, 3, 4, 5, 6, 7, 8 |
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | |
| 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | |
| 17 | 18 | 19 | 20 | 21 | 22 | 23 | 24 | |
| 25 | 26 | 27 | 28 | 29 | 30 | 31 | 32 | |
| 33 | 34 | 35 | 36 | 37 | 38 | 39 | 40 | |
| 41 | 42 | 43 | 44 | 45 | 46 | 47 | 48 | |
| 49 | 50 | 51 | 52 | 53 | 54 | 55 | 56 | |
| 57 | 58 | 59 | 60 | 61 | 62 | 63 | 64 | |
| 65 | 66 | 67 | 68 | 69 | 70 | 71 | 72 | |
| 73 | 74 | 75 | 76 | 77 | 78 | 79 | 80 | |
| 81 | 82 | 83 | 84 | 85 | 86 | 87 | 88 | |
| 89 | 90 | 91 | 92 | 93 | 94 | 95 | 96 |
| Label | Feature Name | Mathematical Representation |
|---|---|---|
| Arithmetic mean | ||
| Entropy | ||
| Standard deviation | ||
| Skewness | ||
| Kurtosis | ||
| Energy | ||
| Shannon entropy | ||
| Renyi entropy |
| Parameters | Value |
|---|---|
| total number of solutions | 200 |
| total number of features | 96 |
| total number of iterations | 100 |
| threshold | 0.5 |
| cognitive factor | 2 |
| social factor | 2 |
| inertia weight | 0.99 |
| fitness function | maximization of classifier performance and minimization of the number of selected features |
| Parameters | Value |
|---|---|
| number of hidden units | 100 |
| fully connected layer | 5 |
| output mode | last |
| state activation function | |
| gate activation function | hard-sigmoid |
| optimization algorithm | Adam |
| maximum number of epochs | 200 |
| minimum batch size | 32 |
| initial learning rate | 0.01 |
| gradient threshold | 1 |
| Maps | Wavelet Family | LSTM (%) | SVM (%) |
|---|---|---|---|
| Chebyshev | rbior2.6 | 99.32 ± 0.12 | 94.80 ± 0.40 |
| Circle | rbior2.2 | 99.44 ± 0.34 | 95.76 ± 0.36 |
| Gauss | coif1 | 99.44 ± 0.24 | 97.84 ± 0.04 |
| Iterative | bior2.4 | 99.48 ± 0.28 | 98.36 ± 0.36 |
| Logistic | bior2.8 | 99.64 ± 0.04 | 96.32 ± 0.32 |
| Piecewise | rbior6.8 | 99.36 ± 0.36 | 96.56 ± 0.16 |
| Sine | bior2.4 | 99.40 ± 0.20 | 98.04 ± 0.24 |
| Singer | db5 | 99.40 ± 0.20 | 96.12 ± 0.08 |
| Sinusoidal | db5 | 99.44 ± 0.04 | 96.84 ± 0.24 |
| Tent | coif4 | 99.56 ± 0.36 | 96.72 ± 0.52 |
| Wavelet Family | Number of Selected Features | Selected Features | Accuracy (%) | ||
|---|---|---|---|---|---|
| RNN-LSTM | SVM | ||||
| Biorthogonal | 1.1 | 13 | 12, 19, 46, 50, 52, 53, 58, 74, 84, 88, 92, 94, 96 | 98.88 ± 0.08 | 95.52 ± 0.28 |
| 1.3 | 17 | 3, 4, 5, 13, 17, 18, 31, 32, 44, 46, 52, 70, 78, 82, 84, 88, 92 | 98.24 ± 1.04 | 92.88 ± 0.32 | |
| 1.5 | 15 | 12, 24, 25, 42, 44, 46, 53, 60, 73, 79, 83, 84, 89, 92, 96 | 98.16 ± 0.16 | 94.88 ± 0.08 | |
| 2.2 | 16 | 4, 9, 12, 18, 32, 37, 47, 61, 66, 67, 70, 71, 74, 76, 81, 87 | 99.16 ± 0.16 | 95.60 ± 0.20 | |
| 2.4 | 18 | 3, 4, 9, 12, 14, 15, 18, 20, 27, 28, 32, 36, 45, 54, 70, 72, 92, 93 | 98.84 ± 0.24 | 95.56 ± 0.04 | |
| 2.6 | 14 | 4, 5, 11, 12, 18, 23, 51, 53, 55, 60, 68, 76, 83, 89 | 99.00 ± 0.02 | 96.28 ± 0.08 | |
| 2.8 | 14 | 1, 7, 9, 12, 15, 58, 61, 69, 71, 74, 76, 89, 92, 93 | 99.64 ± 0.04 | 96.32 ± 0.32 | |
| 3.1 | 14 | 3, 8, 11, 18, 20, 23, 28, 30, 33, 49, 52, 75, 81, 96 | 97.68 ± 0.12 | 91.12 ± 0.12 | |
| 3.3 | 16 | 7, 20, 21, 29, 44, 46, 48, 52, 60, 75, 79, 81, 85, 90, 91, 94 | 98.72 ± 0.12 | 95.68 ± 0.28 | |
| 3.5 | 17 | 2, 7, 9, 12, 16, 20, 27, 37, 44, 47, 52, 54, 60, 61, 74, 81, 90 | 98.88 ± 0.12 | 95.28 ± 0.08 | |
| 3.7 | 18 | 10, 18, 20, 21, 25, 28, 44, 52, 53, 57, 59, 62, 63, 71, 75, 76, 86, 93 | 98.64 ± 0.04 | 94.08 ± 0.08 | |
| 3.9 | 14 | 3, 4, 10, 20, 27, 28, 34, 37, 44, 52, 53, 78, 89, 92 | 98.72 ± 0.12 | 94.60 ± 0.01 | |
| 4.4 | 16 | 8, 14, 20, 23, 25, 36, 37, 40, 44, 45, 50, 60, 71, 76, 78, 85 | 99.20 ± 0.02 | 97.60 ± 0.02 | |
| 5.5 | 19 | 4, 9, 14, 18, 36, 38, 42, 49, 52, 57, 68, 72, 75, 76, 80, 81, 85, 92, 96 | 99.00 ± 0.20 | 94.20 ± 0.20 | |
| 6.8 | 13 | 2, 12, 26, 28, 32, 42, 44, 47, 53, 61, 62, 67, 82 | 99.12 ± 0.32 | 96.20 ± 0.40 | |
| Coiflet | 1 | 14 | 4, 5, 11, 12, 23, 35, 44, 58, 74, 79, 80, 92, 93, 96 | 99.32 ± 0.12 | 98.24 ± 0.24 |
| 2 | 18 | 4, 12, 20, 21, 25, 44, 47, 48, 58, 63, 64, 67, 68, 73, 75, 87, 93, 95 | 98.88 ± 0.08 | 93.68 ± 0.68 | |
| 3 | 14 | 2, 3, 12, 17, 30, 32, 36, 44, 55, 59, 76, 84, 92, 95 | 99.12 ± 0.72 | 95.48 ± 0.28 | |
| 4 | 13 | 4, 6, 9, 20, 44, 53, 58, 61, 64, 66, 85, 86, 92 | 98.76 ± 0.96 | 94.48 ± 0.08 | |
| 5 | 17 | 4, 7, 12, 18, 28, 29, 31, 51, 52, 54, 62, 73, 78, 91, 93, 94, 96 | 99.36 ± 0.16 | 95.48 ± 0.32 | |
| Daubechies | 1 | 13 | 16, 20, 27, 29, 30, 33, 34, 43, 44, 53, 60, 84, 87 | 99.04 ± 0.04 | 95.04 ± 0.04 |
| 2 | 16 | 4, 9, 10, 12, 22, 23, 25, 27, 36, 37, 44, 48, 50, 60, 84, 96 | 99.12 ± 0.12 | 96.68 ± 0.48 | |
| 3 | 13 | 7, 12, 20, 37, 56, 58, 60, 70, 71, 74, 76, 82, 84 | 98.32 ± 0.12 | 87.56 ± 0.56 | |
| 4 | 13 | 20, 28, 44, 52, 53, 58, 67, 69, 74, 76, 79, 80, 85 | 98.64 ± 0.64 | 92.20 ± 0.02 | |
| 5 | 18 | 4, 5, 10, 12, 14, 17, 20, 29, 50, 52, 55, 58, 60, 65, 66, 70, 85, 86 | 99.28 ± 0.48 | 96.32 ± 0.32 | |
| 6 | 13 | 4, 20, 28, 30, 42, 44, 52, 69, 80, 88, 89, 93, 96 | 99.00 ± 0.01 | 97.72 ± 0.12 | |
| 7 | 19 | 3, 12, 17, 19, 20, 25, 26, 32, 50, 52, 55, 58, 60, 61, 65, 72, 82, 84, 93 | 98.72 ± 0.97 | 95.52 ± 0.12 | |
| 8 | 13 | 4, 5, 8, 9, 12, 57, 59, 60, 68, 79, 84, 93, 96 | 98.32 ± 0.32 | 95.04 ± 0.24 | |
| 9 | 13 | 2, 4, 5, 19, 52, 53, 54, 57, 58, 60, 61, 69, 72 | 99.16 ± 0.36 | 97.40 ± 0.20 | |
| 10 | 13 | 6, 8, 10, 20, 27, 41, 44, 65, 68, 86, 92, 93, 95 | 97.48 ± 0.48 | 91.00 ± 0.20 | |
| Fejer–Korovkin | 4 | 19 | 3, 5, 7, 10, 15, 17, 18, 24, 38, 46, 52, 60, 67, 73, 77, 83, 84, 85, 88 | 98.40 ± 0.01 | 94.20 ± 0.01 |
| 6 | 13 | 2, 4, 8, 12, 22, 28, 37, 38, 44, 46, 52, 72, 79 | 98.56 ± 0.24 | 92.60 ± 0.40 | |
| 8 | 15 | 6, 7, 12, 16, 26, 33, 37, 51, 52, 58, 60, 68, 82, 87, 96 | 98.40 ± 0.60 | 94.00 ± 0.40 | |
| 14 | 14 | 2, 3, 20, 31, 38, 42, 44, 49, 50, 53, 61, 76, 80, 92 | 99.28 ± 0.08 | 96.68 ± 0.28 | |
| 18 | 14 | 3, 4, 5, 13, 20, 44, 48, 69, 74, 76, 84, 91, 92, 95 | 99.00 ± 0.20 | 96.20 ± 0.20 | |
| 22 | 13 | 4, 7, 11, 19, 28, 30, 44, 52, 59, 65, 81, 87, 96 | 98.44 ± 0.44 | 95.32 ± 0.28 | |
| Reverse Biorthogonal | 1.1 | 13 | 2, 3, 15, 52, 62, 65, 66, 69, 71, 76, 80, 84, 85 | 97.56 ± 0.36 | 95.20 ± 0.20 |
| 1.3 | 13 | 3, 12, 20, 38, 51, 52, 60, 73, 83, 84, 85, 86, 93 | 97.80 ± 0.01 | 92.60 ± 0.20 | |
| 1.5 | 17 | 2, 4, 10, 17, 20, 23, 25, 27, 33, 39, 43, 47, 52, 54, 69, 84, 92 | 98.32 ± 0.32 | 94.40 ± 0.20 | |
| 2.2 | 13 | 6, 12, 24, 32, 41, 47, 50, 58, 65, 68, 72, 84, 92 | 98.88 ± 0.28 | 94.28 ± 0.08 | |
| 2.4 | 17 | 4, 5, 6, 8, 9, 11, 15, 22, 27, 31, 35, 36, 52, 65, 76, 89, 95 | 98.96 ± 0.04 | 94.36 ± 0.36 | |
| 2.6 | 16 | 12, 15, 16, 21, 22, 29, 37, 44, 45, 68, 70, 72, 88, 91, 94, 95 | 98.96 ± 0.36 | 95.08 ± 0.52 | |
| 2.8 | 13 | 9, 12, 18, 21, 28, 32, 44, 52, 59, 65, 68, 88, 94 | 98.72 ± 0.72 | 93.32 ± 0.32 | |
| 3.1 | 13 | 5, 11, 13, 31, 35, 41, 48, 52, 57, 60, 71, 82, 84 | 98.08 ± 0.08 | 93.04 ± 0.04 | |
| 3.3 | 16 | 2, 4, 10, 23, 28, 31, 40, 41, 49, 55, 71, 76, 82, 84, 88, 93 | 98.32 ± 0.08 | 94.84 ± 0.04 | |
| 3.5 | 16 | 7, 16, 19, 31, 33, 37, 41, 44, 53, 60, 72, 74, 75, 76, 77, 88 | 97.68 ± 0.12 | 91.52 ± 0.12 | |
| 3.7 | 21 | 4, 7, 8, 10, 18, 20, 23, 28, 30, 32, 37, 38, 39, 44, 46, 48, 53, 75, 83, 85, 91 | 98.20 ± 0.01 | 93.76 ± 0.24 | |
| 3.9 | 15 | 1, 3, 4, 16, 19, 20, 26, 37, 48, 57, 60, 63, 69, 75, 88 | 98.56 ± 0.16 | 93.44 ± 0.44 | |
| 4.4 | 14 | 4, 12, 14, 22, 28, 45, 55, 60, 62, 69, 74, 76, 83, 86 | 99.20 ± 0.40 | 96.84 ± 0.44 | |
| 5.5 | 14 | 2, 4, 6, 12, 17, 25, 31, 35, 38, 41, 43, 68, 84, 88 | 98.64 ± 0.24 | 94.84 ± 0.04 | |
| 6.8 | 19 | 12, 22, 25, 26, 27, 28, 32, 36, 40, 52, 54, 62, 72, 75, 77, 78, 80, 89, 95 | 99.24 ± 0.24 | 96.00 ± 0.40 | |
| Symlet | 2 | 13 | 4, 16, 26, 36, 53, 56, 61, 65, 69, 76, 87, 92, 94 | 98.64 ± 0.64 | 96.96 ± 0.36 |
| 3 | 16 | 3, 19, 20, 29, 31, 53, 60, 69, 71, 73, 76, 77, 84, 85, 92, 95 | 98.20 ± 1.40 | 94.68 ± 0.28 | |
| 4 | 15 | 12, 20, 28, 32, 37, 40, 41, 44, 56, 57, 65, 70, 82, 85, 95 | 98.48 ± 0.32 | 93.40 ± 0.40 | |
| 5 | 19 | 4, 7, 8, 17, 19, 20, 34, 37, 39, 42, 46, 61, 64, 67, 68, 85, 88, 91, 92 | 98.96 ± 0.36 | 94.48 ± 0.12 | |
| 6 | 18 | 9, 12, 20, 22, 28, 32, 34, 37, 39, 44, 53, 55, 72, 74, 76, 85, 86, 94 | 98.68 ± 0.12 | 94.08 ± 0.08 | |
| 7 | 15 | 4, 13, 20, 23, 46, 48, 56, 57, 59, 60, 67, 74, 84, 85, 90 | 98.48 ± 0.32 | 92.32 ± 0.08 | |
| 8 | 15 | 4, 14, 20, 27, 28, 30, 36, 43, 44, 49, 52, 53, 61, 84, 96 | 98.56 ± 0.04 | 95.40 ± 0.40 | |
| Wavelet | Chaotic Map | Classifier | Accuracy (%) | Precision (%) | Recall (%) | F1-Score (%) |
|---|---|---|---|---|---|---|
| bior 2.8 | logistic | LSTM | 99.64 | 99.64 | 99.64 | 99.64 |
| SVM | 96.32 | 96.37 | 96.32 | 96.35 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Özçelik, Y.B.; Altan, A. Overcoming Nonlinear Dynamics in Diabetic Retinopathy Classification: A Robust AI-Based Model with Chaotic Swarm Intelligence Optimization and Recurrent Long Short-Term Memory. Fractal Fract. 2023, 7, 598. https://doi.org/10.3390/fractalfract7080598
Özçelik YB, Altan A. Overcoming Nonlinear Dynamics in Diabetic Retinopathy Classification: A Robust AI-Based Model with Chaotic Swarm Intelligence Optimization and Recurrent Long Short-Term Memory. Fractal and Fractional. 2023; 7(8):598. https://doi.org/10.3390/fractalfract7080598
Chicago/Turabian StyleÖzçelik, Yusuf Bahri, and Aytaç Altan. 2023. "Overcoming Nonlinear Dynamics in Diabetic Retinopathy Classification: A Robust AI-Based Model with Chaotic Swarm Intelligence Optimization and Recurrent Long Short-Term Memory" Fractal and Fractional 7, no. 8: 598. https://doi.org/10.3390/fractalfract7080598
APA StyleÖzçelik, Y. B., & Altan, A. (2023). Overcoming Nonlinear Dynamics in Diabetic Retinopathy Classification: A Robust AI-Based Model with Chaotic Swarm Intelligence Optimization and Recurrent Long Short-Term Memory. Fractal and Fractional, 7(8), 598. https://doi.org/10.3390/fractalfract7080598








