Translational Venomics: Third-Generation Antivenomics of Anti-Siamese Russell’s Viper, Daboia siamensis, Antivenom Manufactured in Taiwan CDC’s Vaccine Center
Abstract
1. Introduction
2. Materials and Methods
2.1. Venom and Antivenom
2.2. Determination of Venom LD50 and Antivenom ED50
2.3. Isolation and Initial Characterization of D. siamensis Venom Proteins
2.4. Venomics Characterization and Quantification of the Venom Proteome of Taiwanese Russell’s Viper
2.5. Two-Dimensional (IEF/SDS-PAGE) Gel Electrophoresis (2-DE)
2.6. Third-Generation Antivenomics
3. Results and Discussion
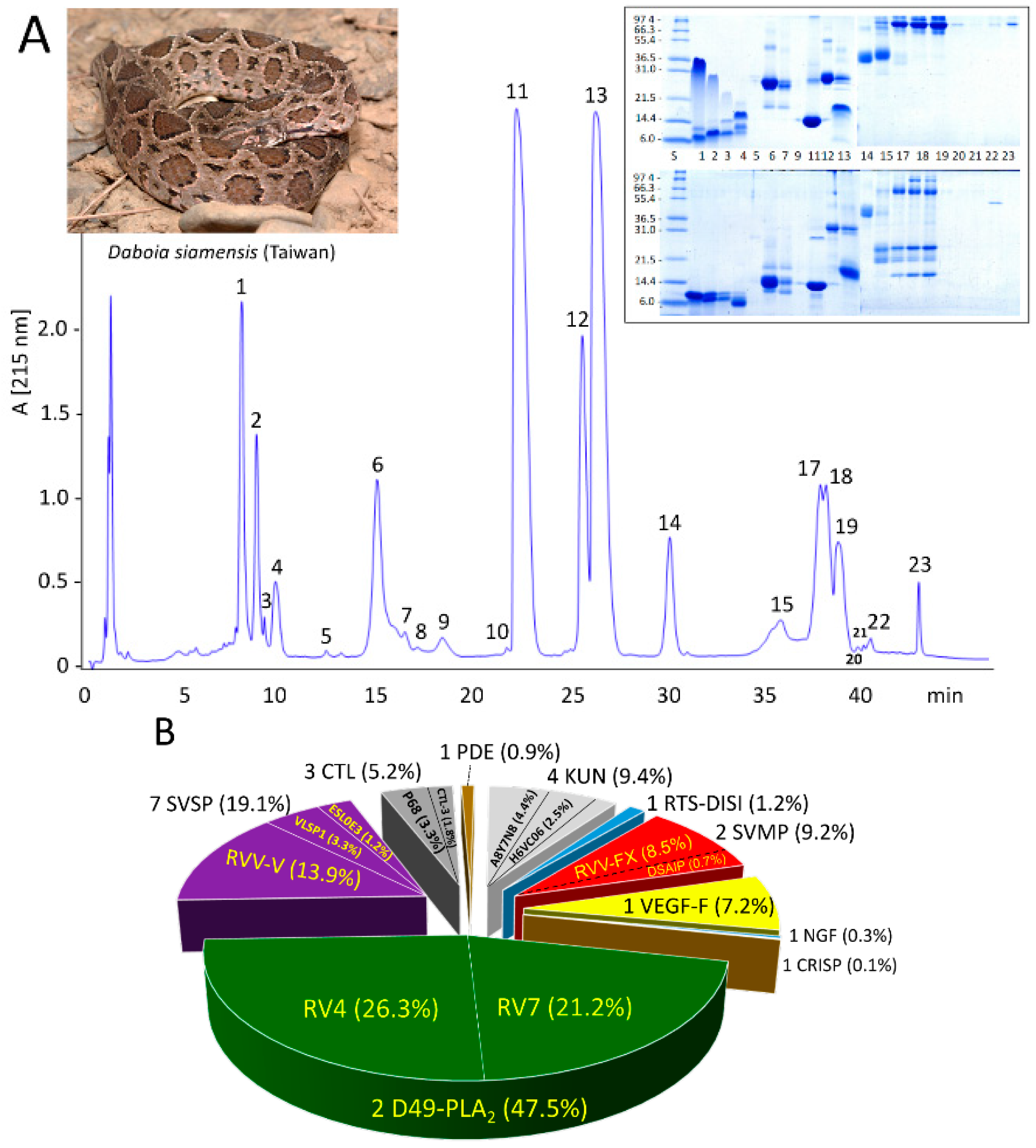
3.1. The Venom Proteome of Siamese Russell’s Viper from Taiwan
3.2. Comparison of the Venom Proteomes of D. siamensis from Taiwan and Myanmar
3.3. Functional Correlations
3.4. In Vivo and In Vitro Analysis of the Preclinical Efficacy of Anti-Daboia siamensis (Taiwan) Antivenom
3.4.1. Venom Lethality and Antivenom Neutralization of the Venom’s Lethal Effect
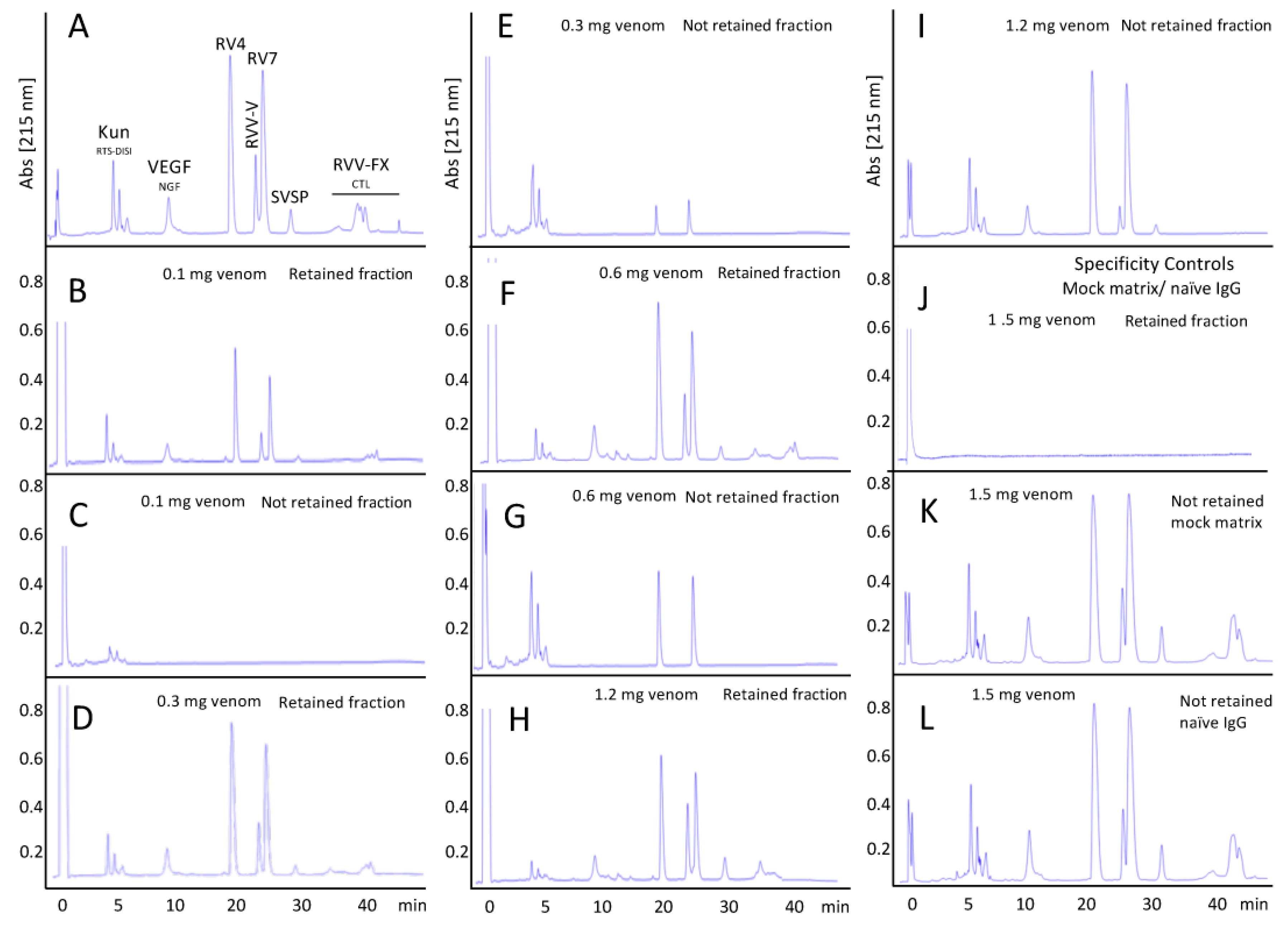
3.4.2. Assessment of the Immunorecognition Landscape of Taiwanese Anti-D. siamensis Antivenom through Third-Generation (3G) Antivenomics
3.4.3. Functional Comparison of the CDC Antivenom with Other Antivenoms against Russell’s Viper Venoms
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Chen, C.-K.; Lin, C.-C.; Shih, F.-Y.; Chaou, C.-H.; Lin, J.C.-C.; Lai, T.-I.; Tseng, C.-Y.; Fang, C.-C. Population-based study of venomous snakebite in Taiwan. J. Acute Med. 2015, 5, 38–42. [Google Scholar] [CrossRef]
- Hung, D.-Z. Taiwan’s venomous snakebite: Epidemiological evolution and geographic differences. Trans. R. Soc. Trop. Med. Hyg. 2004, 98, 96–101. [Google Scholar] [CrossRef]
- Chieh-Fan, C.; Tzeng-Jih, L.; Wen-Chi, H.; Hua-Wei, Y. Appropriate antivenom doses for six types of envenomations caused by snakes in Taiwan. J. Venom. Anim. Toxins incl. Trop. Dis. 2009, 15, 479–490. [Google Scholar] [CrossRef]
- Sawai, Y. Snakebites on Taiwan. Snake 1969, 1, 9–18. [Google Scholar]
- Kuo, T.O.; Wu, C.S. Clinico-pathological studies on snakebites in Taiwan. J. Formos. Med. Assoc. 1972, 71, 447–466. [Google Scholar]
- Alirol, E.; Sharma, S.K.; Bawaskar, H.S.; Kuch, U.; Chappuis, F. Snakebite in South Asia: A review. PLoS Negl. Trop. Dis. 2010, 4, e603. [Google Scholar]
- Thorpe, R.S.; Pook, C.E.; Malhotra, A. Phylogeography of the Russell’s viper (Daboia russelii) complex in relation to variation in the colour pattern and symptoms of envenoming. Herpetol. J. 2007, 17, 209–218. [Google Scholar]
- Uetz, P.; Etzold, T. The EMBL/EBI Reptile Database. Herpetol. Rev. 1996, 27, 174–175. [Google Scholar]
- Theakston, R.D.; Reid, H.A. Development of simple standard assay procedures for the characterization of snake venom. Bull. World Health Organ. 1983, 61, 949–956. [Google Scholar] [PubMed]
- Mallow, D.; Ludwig, D.; Nilson, G. True Vipers: Natural History and Toxinology of Old World Vipers; Krieger Publishing Company: Malabar, FL, USA, 2013; p. 359. ISBN 0-89464-877-2. [Google Scholar]
- Myint-Lwin; Warrell, D.A.; Phillips, R.E.; Tin-Nu-Swe; Tun-Pe; Maung-Maung-Lay. Bites by Russell’s viper (Vipera russellii siamensis) in Burma: Haemostatic, vascular, and renal disturbances and response to treatment. Lancet 1985, 2, 1259–1264. [Google Scholar] [CrossRef]
- Warrell, D.A. Snake venoms in science and clinical medicine. 1. Russell’s viper: Biology, venom and treatment of bites. Trans. R. Soc. Trop. Med. Hyg. 1989, 83, 732–740. [Google Scholar] [CrossRef]
- Tun-Pe; Phillips, R.E.; Warrell, D.A.; Moore, R.A.; Tin-Nu-Swe; Myint-Lwin; Burke, C.W. Acute and chronic pituitary failure resembling Sheehan’s syndrome following bites by Russell’s viper in Burma. Lancet 1987, 2, 763–767. [Google Scholar] [CrossRef]
- Tun-Pe; Ba-Aye; Aye-Aye-Myint; Tin-Nu-Swe; Warrell, D.A. A. Bites by Russell’s vipers (Daboia russelii siamensis) in Myanmar: Effect of the snake’s length and recent feeding on venom antigenaemia and severity of envenoming. Trans. R. Soc. Trop. Med. Hyg. 1991, 85, 804–808. [Google Scholar] [CrossRef]
- Tun-Pe; Aye-Aye-Myint; Kyi-May-Htwe; Khin-Aung-Cho; Theingi. Geographical Variation of Biological Properties of Russell’s Viper (Daboia russelii siamensis) Venom. Seminar on Management of Snakebite and Research, WHO Regional Office for South-East Asia: New Delhi, India, 2002; 65–72.
- Belt, P.; Malhotra, A.; Thorpe, R.S.; Warrell, D.A.; Wüster, W. Russell’s viper in Indonesia: Snakebite and systematics. In Venomous Snakes. Ecology, Evolution and Snakebite; Thorpe, R.S., Wüster, W., Malhotra, A., Eds.; Clarendon Press: Oxford, UK, 1987; pp. 207–217. [Google Scholar]
- Mitrakul, C.; Juzi, U.; Pongrujikorn, W. Antivenom therapy in Russell’s viper bite. Am. J. Clin. Pathol. 1991, 95, 412–417. [Google Scholar] [CrossRef] [PubMed]
- Hung, D.-Z.; Wu, M.L.; Deng, J.F.; Yang, D.Y.; Lin-Shiau, S.Y. Multiple thrombotic occlusions of vessels after Russell’s viper envenoming. Pharmacol. Toxicol. 2002, 91, 106–110. [Google Scholar] [CrossRef] [PubMed]
- Hung, D.-Z.; Wu, M.L.; Deng, J.F.; Lin-Shiau, S.Y. Russell’s viper snakebite in Taiwan: Differences from other Asian countries. Toxicon 2002, 40, 1291–1298. [Google Scholar] [CrossRef]
- Hung, D.-Z.; Yu, Y.-J.; Hsu, C.-L.; Lin, T.-J. Antivenom treatment and renal dysfunction in Russell’s viper snakebite in Taiwan: A case series. Trans. R. Soc. Trop. Med. Hyg. 2006, 100, 489–494. [Google Scholar] [CrossRef] [PubMed]
- Warrell, D.A. Researching nature’s venoms and poisons. Trans. R. Soc. Trop. Med. Hyg. 2009, 103, 860–866. [Google Scholar] [CrossRef] [PubMed]
- Warrell, D.A. Geographical and intraspecies variation in the clinical manifestations of envenoming by snakes. In Venomous Snakes. Ecology, Evolution and Snakebite; Thorpe, R.S., Wüster, W., Malhotra, A., Eds.; Clarendon Press: Oxford, UK, 1997; pp. 189–203. [Google Scholar]
- Risch, M.; Georgieva, D.; von Bergen, M.; Jehmlich, N.; Genov, N.; Arni, R.K.; Betzel, C. Snake venomics of the Siamese Russell’s viper (Daboia russellii siamensis)—relation to pharmacological activities. J. Proteomics 2009, 72, 256–269. [Google Scholar] [CrossRef] [PubMed]
- Maung-Maung-Thwin; Gopalakrishnakone, P.; Yuen, R.; Tan, C.H. A major lethal factor of the venom of Burmese Russell’s viper (Daboia russellii siamensis): Isolation, N-terminal sequencing and biological activities of daboiatoxin. Toxicon 1995, 33, 63–76. [Google Scholar] [CrossRef]
- Liau, M.Y.; Huang, R.J. Toxoids and antivenoms of venomous snakes in Taiwan. Toxin Rev. 1997, 16, 163–175. [Google Scholar]
- Lin, C.-C.; Chaou, C.-H.; Tseng, C.-Y. An investigation of snakebite antivenom usage in Taiwan. J. Formos. Med. Assoc. 2016, 115, 672–677. [Google Scholar] [CrossRef] [PubMed]
- Hsu, Y.-L.; Chia-Jung, W.; Tsu-Chen, C.; Wen-Chin, H.; Cheng, Y.-F.; Chiang, J.-R. Retrospection and prospection for manufacturing of snake antivenins in Taiwan. Taiwan Epidemiol. Bull. 2013, 29, 50–56. [Google Scholar]
- Calvete, J.J. Venomics: Integrative venom proteomics and beyond. Biochem. J. 2017, 474, 611–634. [Google Scholar] [CrossRef] [PubMed]
- Pla, D.; Rodríguez, Y.; Calvete, J.J. Third generation antivenomics: pushing the limits of the in vitro preclinical assessment of antivenoms. Toxins 2017, 9, 158. [Google Scholar]
- Finney, D.J. Probit Analysis; Cambridge University Press: Cambridge, UK, 1971; 333p, ISBN 052108041X. [Google Scholar]
- Eichberg, S.; Sanz, L.; Calvete, J.J.; Pla, D. Constructing comprehensive venom proteome reference maps for integrative venomics. Expert Rev. Proteomics 2015, 12, 557–573. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Calvete, J.J. Next-generation snake venomics: Protein-locus resolution through venom proteome decomplexation. Expert Rev. Proteomics 2014, 11, 315–329. [Google Scholar] [CrossRef] [PubMed]
- Calderón-Celis, F.; Cid-Barrio, L.; Ruiz Encinar, J.; Sanz-Medel, A.; Calvete, J.J. Absolute venomics: Absolute quantification of intact venom proteins through elemental mass spectrometry. J. Proteomics 2017, 164, 33–42. [Google Scholar] [CrossRef] [PubMed]
- Howard, G.C.; Kaser, M.R. Making and Using Antibodies: A Practical Handbook, 2nd ed.; CRC Press: Boca Raton, FL, USA, 2014; p. 458. ISBN 978-1-43-986908-6. [Google Scholar]
- Perbandt, M.; Tsai, I.H.; Fuchs, A.; Banumathi, S.; Rajashankar, K.R.; Georgieva, D.; Kalkura, N.; Singh, T.P.; Genov, N.; Betzel, C. Structure of the heterodimeric neurotoxic complex viperotoxin F (RV-4/RV-7) from the venom of Vipera russelliformosensis at 1.9 A resolution. Acta Crystallogr. D Biol. Crystallogr. 2003, 59, 1679–1687. [Google Scholar] [CrossRef] [PubMed]
- Takeda, S.; Igarashi, T.; Mori, H. Crystal structure of RVV-X: An example of evolutionary gain of specificity by ADAM proteinases. FEBS Lett. 2007, 581, 5859–5864. [Google Scholar] [CrossRef] [PubMed]
- Than-Than; Hutton, R.A.; Myint-Lwin; Khin-Ei-Han; Soe-Soe; Tin-Nu-Swe; Phillips, R.E.; Warrell, D.A. Haemostatic disturbances in patients bitten by Russell’s viper (Vipera russellii siamensis) in Burma. Br. J. Haematol. 1988, 69, 513–520. [Google Scholar] [CrossRef] [PubMed]
- Aung-Khin, M.; Ma-Ma, K.; Zin, T. Effects of Russell’s viper venom on blood coagulation, platelets and the fibrinolytic enzyme system. Jpn. J. Med. Sci. Biol. 1977, 30, 101–108. [Google Scholar] [CrossRef] [PubMed]
- Phillips, R.E.; Theakston, R.D.G.; Warrell, D.A.; Galigedara, Y.; Abeysekera, D.T.; Dissanayaka, P.; Hutton, R.A.; Aloysius, D.J. Paralysis, rhabdomyolysis and hemolysis cause by bites of Russell’s viper (Vipera russellipulchella) in Sri Lanka: Failure of Indian (Haffkine) antivenin. Q. J. Med. 1988, 68, 691–716. [Google Scholar] [PubMed]
- Than-Than; Francis, N.; Tin-Nu-Swe; Myint-Lwin; Tun-Pe; Soe-Soe; Maung-Maung-Oo; Phillips, R.E.; Warrell, D.A. Contribution of focal hemorrhage and microvascular fibrin deposition to fatal envenoming by Russell’s viper (Vipera russellii siamensis) in Burma. Acta Trop. 1989, 46, 23–38. [Google Scholar] [CrossRef]
- Tin-Nu-Swe; Tin-Tun; Myint-Lwin; Thein-Than; Tun-Pe; Robertson, J.I.; Leckie, B.J.; Phillips, R.E.; Warrell, D.A. Renal ischemia, transient glomerular leak and acute renal tubular damage in patients envenomed by Russell’s vipers (Daboia russellii siamensis) in Myanmar. Trans. R. Soc. Trop. Med. Hyg. 1993, 87, 678–681. [Google Scholar] [CrossRef]
- Morita, T. Proteases which activate factor X. In Enzymes from Snake Venom; Bailey, G.S., Ed.; Alaken: Fort Collins, CO, USA, 1998; pp. 179–208. [Google Scholar]
- Tans, G.; Rosing, J. Snake venom activators of factor X: An overview. Haemostasis 2001, 31, 225–233. [Google Scholar] [CrossRef] [PubMed]
- Nicolaes, G.A.F.; Rosing, J. Activation of factor V by venom proteases. J. Toxicol. Toxin Rev. 2006, 25, 217–234. [Google Scholar] [CrossRef]
- Nakayama, D.; Ben Ammar, Y.; Miyata, T.; Takeda, S. Structural basis of coagulation factor V recognition for cleavage by RVV-V. FEBS Lett. 2011, 585, 3020–3025. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.Y. Toxicological studies on the venom of Vipera russelliformosensis maki. Part 1. Toxicity and pharmacological properties. J. Formos. Med. Assoc. 1948, 47, 65–98. [Google Scholar]
- Jayanthi, J.P.; Gowda, T.V. Geographical variation in India in the composition and lethal potency of Russell’s viper (Vipera russelli) venom. Toxicon 1988, 26, 257–264. [Google Scholar] [CrossRef]
- Silva, A.; Maduwage, K.; Sedgwick, M.; Pilapitiya, S.; Weerawansa, P.; Dahanayaka, N.J.; Buckley, N.A.; Siribaddana, S.; Isbister, G.K. Neurotoxicity in Russell’s viper (Daboia russelii) envenoming in Sri Lanka: A clinical and neurophysiological study. Clin. Toxicol. 2016, 54, 411–419. [Google Scholar] [CrossRef] [PubMed]
- Silva, A.; Kuruppu, S.; Othman, I.; Goode, R.J.; Hodgson, W.C.; Isbister, G.K. Neurotoxicity in Sri Lankan Russell’s viper (Daboia russelii) envenoming is primarily due to U1-viperitoxin-Dr1a, a pre-synaptic neurotoxin. Neurotox. Res. 2017, 31, 11–19. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.M.; Lu, P.J.; Ho, C.L.; Tsai, I.H. Characterization and molecular cloning of neurotoxic phospholipase A2 from Taiwan viper (Vipera russelliformosensis). Eur. J. Biochem. 1992, 209, 635–641. [Google Scholar] [CrossRef] [PubMed]
- Calvete, J.J.; Sanz, L.; Cid, P.; de la Torre, P.; Flores-Díaz, M.; Dos Santos, M.C.; Borges, A.; Bremo, A.; Angulo, Y.; Lomonte, B.; et al. Snake venomics of the Central American rattlesnake Crotalus simus and the South American Crotalus durissus complex points to neurotoxicity as an adaptive paedomorphic trend along Crotalus dispersal in South America. J. Proteome Res. 2010, 9, 528–544. [Google Scholar] [CrossRef] [PubMed]
- Slotta, K.H.; Fraenkel-Conrat, H. Estudos químicos sobre os venenos ofídicos. 4-Purificacão e cristalizacão do veneno da cobra cascavel. Mem. Inst. Butantan 1938, 12, 505–512. [Google Scholar]
- Gutiérrez, J.M.; Dos Santos, M.C.; Furtado, M.F.; Rojas, G. Biochemical and pharmacological similarities between the venoms of newborn Crotalus durissusdurissus and adult Crotalus durissus terrificus rattlesnakes. Toxicon 1991, 29, 1273–1277. [Google Scholar] [CrossRef]
- Bon, C. Multicomponent neurotoxic phospholipases A2. In Venom Phospholipase A2. In Venom Phospholipase A2 Enzymes: Structure, Function and Mechanism; Kini, R.M., Ed.; Wiley: Chichester, UK, 1997; pp. 269–285. [Google Scholar]
- Santoro, M.L.; Sousa-e-Silva, M.C.; Gonçalves, L.R.; Almeida-Santos, S.M.; Cardoso, D.F.; Laporta-Ferreira, I.L.; Saiki, M.; Peres, C.A.; Sano-Martins, I.S. Comparison of the biological activities in venoms from three subspecies of the South American rattlesnake (Crotalus durissus terrificus, C. durissus cascavella and C. durissus collilineatus). Comp. Biochem. Physiol. C Pharmacol. Toxicol. Endocrinol. 1999, 122, 61–73. [Google Scholar] [CrossRef]
- Faure, G.; Xu, H.; Saul, F.A. Crystal structure of crotoxin reveals key residues involved in the stability and toxicity of this potent heterodimeric β-neurotoxin. J. Mol. Biol. 2011, 412, 176–191. [Google Scholar] [CrossRef] [PubMed]
- Faure, G.; Saul, F. Crystallographic characterization of functional sites of crotoxin and ammodytoxin, potent β-neurotoxins from Viperidae venom. Toxicon 2012, 60, 531–535. [Google Scholar] [CrossRef] [PubMed]
- Bolaños, R.; Marín, O.; Mora-Medina, E.; Alfaro, E.A. El accidente ofídico por cascabela (Crotalus durissus durissus) en Costa Rica. Acta Med. Costarric. 1981, 24, 211–214. [Google Scholar]
- Saravia, P.; Rojas, E.; Arce, V.; Guevara, C.; López, J.C.; Chaves, E.; Velásquez, R.; Rojas, G.; Gutiérrez, J.M. Geographic and ontogenic variability in the venom of the neotropical rattlesnake Crotalus durissus: Pathophysiological and therapeutic implications. Rev. Biol. Trop. 2002, 50, 337–346. [Google Scholar] [PubMed]
- Gutiérrez, J.M. Snakebite envenomation in Central America. In Handbook of Venoms and Toxins of Reptiles; Mackessy, S.P., Ed.; CRC Press: Boca Raton, FL, USA, 2009; pp. 491–507. [Google Scholar]
- Azevedo-Marques, M.M.; Cupo, P.; Coimbra, T.M.; Hering, S.E.; Rossi, M.E.; Laure, C.J. Myonecrosis, myoglobinuria and acute renal failure induced by South American rattlesnake (Crotalus durissus terrificus) envenomation in Brazil. Toxicon 1985, 23, 631–636. [Google Scholar] [CrossRef]
- Fan, H.W.; Cardoso, J.L. Clinical toxicology of snake bites South America. In Handbook of Clinical Toxicology of Animal Venoms and Poisons; Meier, J., White, J., Eds.; CRC Press: Boca Raton, FL, USA, 1995; pp. 667–688. [Google Scholar]
- Aye-Kyaw; San-Aye; Tin-Win; Hla-Pe; Mya-Maung. Purification, characterization and biological activities of phospholipase A from Russell’s viper (Vipera russelli) venom. Int. J. Biochem. 1994, 26, 79–83. [Google Scholar] [CrossRef]
- Napathorn, S.; Tejachokviwat, M.; Maneesri, S.; Kasantikul, V.; Stitprija, V. Effects of Russell’s viper venom on human erythrocytes in vitro. J. Nat. Toxins 1998, 7, 73–85. [Google Scholar] [PubMed]
- Mukherjee, A.K.; Mackessy, S.P. Pharmacological properties and pathophysiological significance of a Kunitz-type protease inhibitor (rusvikunin-II) and its protein complex (rusvikunin complex) purified from Daboia russelii russelii venom. Toxicon 2014, 89, 55–66. [Google Scholar] [CrossRef] [PubMed]
- Cheng, A.C.; Tsai, I.H. Functional characterization of a slow and tight-binding inhibitor of plasmin isolated from Russell’s viper venom. Biochim. Biophys. Acta 2014, 1840, 153–159. [Google Scholar] [CrossRef] [PubMed]
- Cheng, A.C.; Wu, H.L.; Shi, G.Y.; Tsai, I.H. A novel heparin-dependent inhibitor of activated protein C that potentiates consumptive coagulopathy in Russell’s viper envenomation. J. Biol. Chem. 2012, 287, 15739–15748. [Google Scholar] [CrossRef] [PubMed]
- Clemetson, K.J. Snaclecs (snake C-type lectins) that inhibit or activate platelets by binding to receptors. Toxicon 2010, 56, 1236–1246. [Google Scholar] [CrossRef] [PubMed]
- Arlinghaus, F.T.; Eble, J.A. C-type lectin-like proteins from snake venoms. Toxicon 2012, 60, 512–519. [Google Scholar] [CrossRef] [PubMed]
- Zhong, S.R.; Jin, Y.; Wu, J.B.; Chen, R.Q.; Jia, Y.H.; Wang, W.Y.; Xiong, Y.L.; Zhang, Y. Characterization and molecular cloning of dabocetin, a potent antiplatelet C-type lectin-like protein from Daboia russellii siamensis venom. Toxicon 2006, 47, 104–112. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, A.K.; Dutta, S.; Mackessy, S.P. A new C-type lectin (RVsnaclec) purified from venom of Daboia russelii russelii shows anticoagulant activity via inhibition of FXa and concentration-dependent differential response to platelets in a Ca2+-independent manner. Thromb. Res. 2014, 134, 1150–1156. [Google Scholar] [CrossRef] [PubMed]
- Mitra, J.; Bhattacharyya, D. Phosphodiesterase from Daboia russelli russelli venom: Purification, partial characterization and inhibition of platelet aggregation. Toxicon 2014, 88, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Aird, S.D. Ophidian envenomation strategies and the role of purines. Toxicon 2002, 40, 335–393. [Google Scholar] [CrossRef]
- Yamazaki, Y.; Nakano, Y.; Imamura, T.; Morita, T. Augmentation of vascular permeability of VEGF is enhanced by KDR-binding proteins. Biochem. Biophys. Res. Commun. 2007, 355, 693–699. [Google Scholar] [CrossRef] [PubMed]
- Calvete, J.J. unpublished data.
- Sapsutthipas, S.; Leong, P.K.; Akesowan, S.; Pratanaphon, R.; Tan, N.H.; Ratanabanangkoon, K. Effective equine immunization protocol for production of potent poly-specific antisera against Calloselasma rhodostoma, Cryptelytrops albolabris and Daboia siamensis. PLoS Negl. Trop. Dis. 2015, 9, e0003609. [Google Scholar] [CrossRef] [PubMed]
- Leong, P.K.; Tan, C.H.; Sim, S.M.; Fung, S.Y.; Sumana, K.; Sitprija, V.; Tan, N.H. Cross-neutralization of common Southeast Asian viperid venoms by a Thai polyvalent snake antivenom (Hemato Polyvalent Snake Antivenom). Acta Trop. 2014, 132, 7–14. [Google Scholar] [CrossRef] [PubMed]
- Kanjanabuch, T.; Sitprija, V. Snakebite nephrotoxicity in Asia. Semin. Nephrol. 2008, 28, 363–372. [Google Scholar] [CrossRef] [PubMed]
- Merchant, M.R.; Khanna, U.B.; Almeida, A.F.; Acharya, V.N.; Mittal, B.V. Clinicopathological study of acute renal failure following viperine snake bite. J. Assoc. Physicians India 1989, 37, 430–443. [Google Scholar] [PubMed]
- Acharya, V.N.; Khanna, U.B.; Almeida, A.F.; Merchant, M.R. Acute renal failure due to viperine snake bite as seen in tropical western India. Ren. Fail. 1989, 11, 33–35. [Google Scholar] [CrossRef] [PubMed]

| Protein | Species | |||
|---|---|---|---|---|
| P18964 | SVSP | RVV-V alpha | FV activator | D. siamensis |
| P18965 | SVSP | RVV-V gamma | D. siamensis | |
| P86530 | SVSP | RVV-V homolog-1 | D. russelii | |
| P86531/2 | SVSP | RVV-V homolog-4 | D. russelii | |
| E0Y418 | SVSP | VLSP-1 | M. lebetina | |
| E0Y419 | SVSP | VLBF | β-fibrinogenase | M. lebetina |
| ESL0E3 | SVSP | VLAF | α-fibrinogenase | D. siamensis |
| A8Y7N8 | KUN | KUN-5 | D. siamensis | |
| H6VC06 | KUN | DrK-In-II | D. russelii | |
| AFE83617 | KUN | D. siamensis | ||
| A8Y7N6/7 | KUN | KUN-3/4 | D. siamensis | |
| ADK22825 | CTL | P68 a-subunit | Light chains of PIV-SVMP RVV-FX (αβ)n, n ≥ 2 | D. siamensis |
| Q4PRD0 | CTL | C-type lectin-3 | D. siamensis | |
| Q4PRC9 | CTL | C-type lectin-4 | D. siamensis | |
| Q4PRD1 | CTL | RVV-FX Light Chain-1 | PIV-SVMP FX activator | D. siamensis |
| ADJ67473 | CTL | RVV-FX Light Chain-2 | D. siamensis | |
| Q7LZ61 | SVMP | RVV-FX Heavy Chain | D. siamensis | |
| AUF41660 | SVMP | DSAIP | PIII-SVMP Daborhagin-K | D. siamensis |
| Q02471 | PLA2 | Basic subunit RV4 | Viperotoxin-F | D. siamensis |
| P31100 | PLA2 | Acidic subunit RV7 | D. siamensis | |
| AUF41658 | DISI | RTS-disintegrin | Russelistatin | D. siamensis |
| P0DL42 | VEGF | VEGF-F | D. siamensis | |
| P30894 | NGF | D. russelii | ||
| ACE73567 | CRISP | D. russelii | ||
| AHJ80885 | PDE | M. lebetina | ||
| RP-HPLC Fraction | Daboia siamensis (Taiwan) Total Venom Proteins (mg) | Major Toxins in RP-HPLC Fraction | ||||||
|---|---|---|---|---|---|---|---|---|
| 100 | 300 | 600 | 900 | 1200 | 1500 | |||
| 1 | mg TOTAL | 5.62 | 16.86 | 33.72 | 50.58 | 67.44 | 94.40 | KUN-5 [A8Y7N8], RTS-DISI [AUF41658] |
| mg RET | 4.17 | 7.06 | 9.33 | 8.61 | 8.81 | 8.00 | ||
| 2 | mg TOTAL | 3.08 | 9.24 | 18.48 | 27.72 | 36.96 | 46.20 | KUN DrKIn-II [H6VC06], KUN-1/5 AFE83617] |
| mg RET | 1.85 | 3.83 | 5.06 | 3.82 | 4.06 | 4.21 | ||
| 3 | mg TOTAL | 0.23 | 0.69 | 1.38 | 2.07 | 2.76 | 3.45 | |
| mg RET | 0.20 | 0.60 | 1.11 | 0.89 | 0.84 | 0.90 | ||
| 4 | mg TOTAL | 1.67 | 5.01 | 10.02 | 15.03 | 20.04 | 28.53 | KUN DrKIn-II [H6VC06], [A8Y7N6/7] |
| mg RET | 1.27 | 2.91 | 4.39 | 1.81 | 3.14 | 0.00 | ||
| 6 | mg TOTAL | 7.02 | 21.06 | 42.12 | 63.18 | 84.24 | 28.53 | VEGF-F [P0DL42] |
| mg RET | 7.02 | 21.06 | 42.12 | 39.85 | 34.84 | 33.16 | ||
| 11 | mg TOTAL | 26.21 | 78.63 | 157.26 | 235.89 | 314.52 | 393.15 | PLA2 RV4 [Q02471] |
| mg RET | 27.70 | 77.23 | 122.74 | 105.19 | 117.07 | 119.20 | ||
| 12 | mg TOTAL | 8.07 | 24.21 | 48.42 | 72.63 | 96.84 | 121.05 | SVSP RVV-V [P18964] |
| mg RET | 8.07 | 24.21 | 48.42 | 72.63 | 78.92 | 73.75 | ||
| 13 | mg TOTAL | 26.87 | 80.61 | 161.22 | 241.83 | 322.44 | 403.05 | PLA2 RV7 [P31100] |
| mg RET | 27.73 | 74.73 | 116.50 | 114.58 | 107.27 | 108.89 | ||
| 14 | mg TOTAL | 3.32 | 9.96 | 19.92 | 29.88 | 39.84 | 49.80 | SVSP VLSP-1 |
| mg RET | 3.07 | 9.21 | 18.41 | 27.62 | 32.07 | 31.76 | ||
| 15 | mg TOTAL | 2.45 | 7.35 | 14.70 | 22.05 | 29.40 | 36.75 | PIV-SVMP RVV-FX |
| mg RET | 1.55 | 4.64 | 9.28 | 13.91 | 18.55 | 23.19 | ||
| 17 | mg TOTAL | 4.44 | 13.32 | 26.64 | 39.96 | 53.28 | 66.60 | CTL-4 [Q4PRC9], P68 [ADK22825] |
| mg RET | 4.44 | 13.32 | 26.64 | 39.96 | 53.28 | 66.60 | ||
| 18 | mg TOTAL | 4.39 | 13.17 | 26.34 | 39.51 | 52.68 | 65.85 | CTL-4, CTL-3 [4QPRD0], PIV-SVMP RVV-FX |
| mg RET | 4.39 | 13.17 | 26.34 | 39.51 | 52.68 | 65.85 | ||
| 19 | mg TOTAL | 3.99 | 11.97 | 23.94 | 35.91 | 47.88 | 59.85 | CTL-4, CTL-3, PIV-SVMP RVV-FX |
| mg RET | 3.99 | 11.97 | 23.94 | 35.91 | 47.88 | 59.85 | ||
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sanz, L.; Quesada-Bernat, S.; Chen, P.Y.; Lee, C.D.; Chiang, J.R.; Calvete, J.J. Translational Venomics: Third-Generation Antivenomics of Anti-Siamese Russell’s Viper, Daboia siamensis, Antivenom Manufactured in Taiwan CDC’s Vaccine Center. Trop. Med. Infect. Dis. 2018, 3, 66. https://doi.org/10.3390/tropicalmed3020066
Sanz L, Quesada-Bernat S, Chen PY, Lee CD, Chiang JR, Calvete JJ. Translational Venomics: Third-Generation Antivenomics of Anti-Siamese Russell’s Viper, Daboia siamensis, Antivenom Manufactured in Taiwan CDC’s Vaccine Center. Tropical Medicine and Infectious Disease. 2018; 3(2):66. https://doi.org/10.3390/tropicalmed3020066
Chicago/Turabian StyleSanz, Libia, Sarai Quesada-Bernat, Pei Yu Chen, Cheng Dow Lee, Jen Ron Chiang, and Juan J. Calvete. 2018. "Translational Venomics: Third-Generation Antivenomics of Anti-Siamese Russell’s Viper, Daboia siamensis, Antivenom Manufactured in Taiwan CDC’s Vaccine Center" Tropical Medicine and Infectious Disease 3, no. 2: 66. https://doi.org/10.3390/tropicalmed3020066
APA StyleSanz, L., Quesada-Bernat, S., Chen, P. Y., Lee, C. D., Chiang, J. R., & Calvete, J. J. (2018). Translational Venomics: Third-Generation Antivenomics of Anti-Siamese Russell’s Viper, Daboia siamensis, Antivenom Manufactured in Taiwan CDC’s Vaccine Center. Tropical Medicine and Infectious Disease, 3(2), 66. https://doi.org/10.3390/tropicalmed3020066






