Investigation of the Prevalence of High-Risk Human Papillomavirus, Human Herpesvirus-8, and Herpes Simplex Virus-2 in Cervical Biopsy Samples Using the Real-Time PCR Method
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Design and Sample Collection
2.2. Ethical Consideration
2.3. Histopathological Evaluation
2.4. DNA Extraction and Sample Preparation
2.5. Detection of HR-HPV, HHV-8, and HSV-1/2 DNA
2.6. Statistical Analysis
3. Results
3.1. Patient Demographics and HPV Prevalence
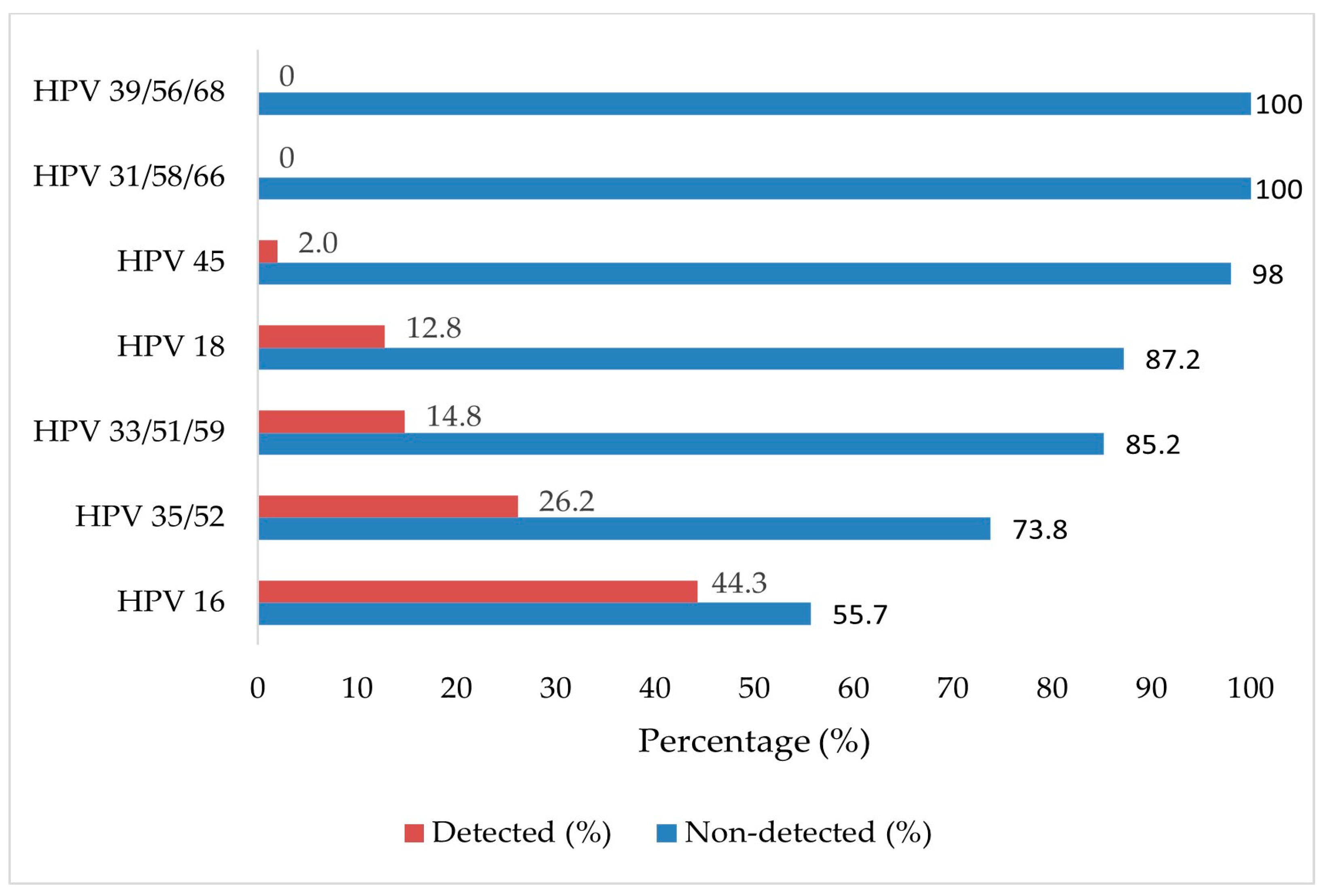
3.2. HPV Genotype Distribution and Infection Patterns
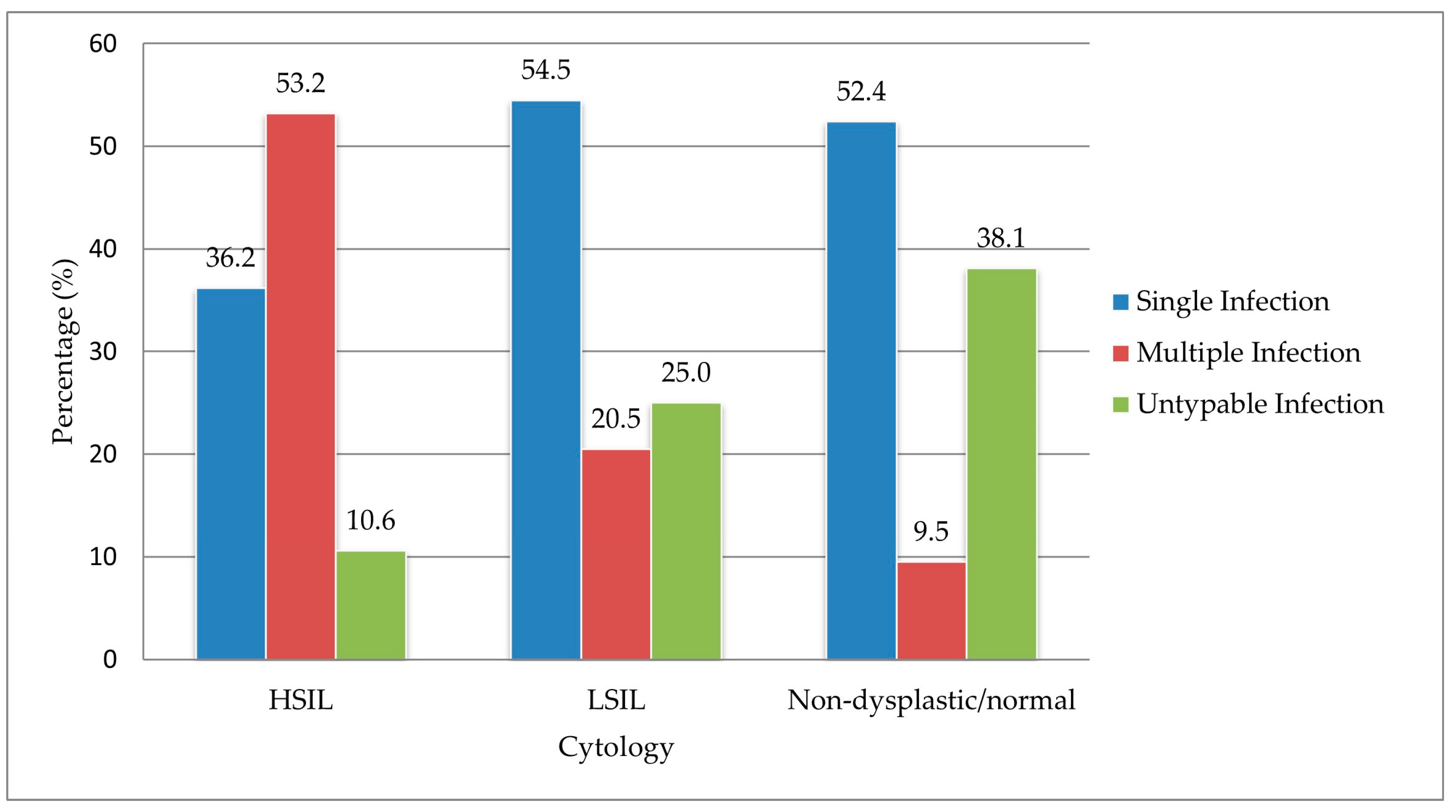
3.3. Histopathological Correlations
3.4. Association Between HR-HPV Status, Age, Infection Pattern, and Cervical Lesion Severity
3.5. Herpesvirus Detection
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| HPV | Human papillomavirus |
| HR-HPV | High-risk human papillomavirus |
| HSV-1 | Herpes simplex virus 1 |
| HSV-2 | Herpes simplex virus 2 |
| HHV-8 | Human herpes virus-8 |
| FFPE | Formalin-fixed paraffin-embedded |
| LSIL | Low-grade squamous intraepithelial lesion |
| HSIL | High-grade squamous intraepithelial lesion |
References
- Wang, J.; Tian, Z.; Wang, J. Risk Factors for Persistent Infection of High-Risk HPV in Patients with Cervical Intraepithelial Neoplasia. Am. J. Transl. Res. 2025, 17, 2992–3000. [Google Scholar] [CrossRef] [PubMed]
- Pimple, S.A.; Mishra, G.A. Global strategies for cervical cancer prevention and screening. Minerva Ginecol. 2019, 71, 313–320. [Google Scholar] [CrossRef] [PubMed]
- Habtemariam, L.W.; Zewde, E.T.; Simegn, G.L. Cervix Type and Cervical Cancer Classification System Using Deep Learning Techniques. Med. Devices 2022, 15, 163–176. [Google Scholar] [CrossRef] [PubMed]
- Tawe, L.; Grover, S.; Narasimhamurthy, M.; Moyo, S.; Gaseitsiwe, S.; Kasvosve, I.; Paganotti, G.M. Molecular Detection of Human Papillomavirus (HPV) in Highly Fragmented DNA from Cervical Cancer Biopsies using Double-Nested PCR. MethodsX 2018, 5, 569–578. [Google Scholar] [CrossRef] [PubMed]
- Staykova, J.; Belovska, T.; Murad, A.; Kakid, S.; Nacheva, A.; Shikova, E. Cervical Viral Infections Among Asymptomatic Bulgarian Women. Cent. Eur. J. Public Health 2016, 24, 176–179. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Cai, S.; Xia, Y.; Lin, Y.; Zhou, G.; Yu, Y.; Feng, M. Association between Human Herpesvirus Infection and Cervical Carcinoma: A Systematic Review and Meta-Analysis. Virol. J. 2023, 20, 288. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, M.D.; de Melo, Y.L. Association between Human Papillomavirus and Epstein–Barr Virus Infections and Cancer of the Uterine Cervix. Crit. Rev. Oncog. 2019, 24, 379–383. [Google Scholar] [CrossRef] [PubMed]
- Blanco, R.; Carrillo-Beltrán, D.; Osorio, J.C.; Calaf, G.M.; Aguayo, F. Role of Epstein–Barr virus and Human Papillomavirus Coinfection in Cervical Cancer: Epidemiology, Mechanisms and Perspectives. Pathogens 2020, 9, 685. [Google Scholar] [CrossRef] [PubMed]
- Abebe, M.; Eshetie, S.; Tessema, B. Prevalence of Sexually Transmitted Infections Among Cervical Cancer Suspected Women at University of Gondar Comprehensive Specialized Hospital, Northwest Ethiopia. BMC Infect. Dis. 2021, 21, 378. [Google Scholar] [CrossRef] [PubMed]
- Ferber, M.J.; Montoya, D.P.; Yu, C.; Aderca, I.; McGee, A.; Thorland, E.C.; Nagorney, D.M.; Gostout, B.S.; Burgart, L.J.; Boix, L.; et al. Integrations of the Hepatitis B Virus (HBV) and Human Papillomavirus (HPV) into the Human Telomerase Reverse Transcriptase (Htert) Gene in Liver and Cervical Cancers. Oncogene 2003, 22, 3813–3820. [Google Scholar] [CrossRef] [PubMed]
- Luo, C.; Yu, S.; Zhang, J.; Wu, X.; Dou, Z.; Li, Z.; Yang, E.; Zhang, L. Hepatitis B or C Viral Infection and the Risk of Cervical Cancer. Infect. Agents Cancer 2022, 17, 54. [Google Scholar] [CrossRef] [PubMed]
- Mertelsmann, A.M.; Mukerebe, C.; Miyaye, D.; Shigella, P.; Mhango, L.; Lutonja, P.; Corstjens, P.L.A.M.; de Dood, C.; van Dam, G.J.; Colombe, S.; et al. Clinical and Demographic Factors Associated with Kaposi Sarcoma-Associated Herpesvirus Shedding in Saliva or Cervical Secretions in a Cohort of Tanzanian Women. Open Forum Infect. Dis. 2024, 11, ofae161. [Google Scholar] [CrossRef] [PubMed]
- Chavoshpour-Mamaghani, S.; Shoja, Z.; Mollaei-Kandelous, Y.; Sharifian, K.; Jalilvand, S. The Prevalence of Human herpesvirus 8 in Normal, Premalignant, and Malignant Cervical Samples of Iranian Women. Virol. J. 2021, 18, 144. [Google Scholar] [CrossRef] [PubMed]
- Moharreri, M.; Sohrabi, A. Characteristics of HSV-2, M. genitalium and C. trachomatis in HPV Genotypes Associated with Cervical Intraepithelial Neoplasia and Genital Infections. Infect. Disord. Drug Targets 2021, 21, 112–118. [Google Scholar] [PubMed]
- Vitali, D.; Bagri, P.; Wessels, J.M.; Arora, M.; Ganugula, R.; Parikh, A.; Mandur, T.; Felker, A.; Garg, S.; Kumar, M.N.V.R.; et al. Curcumin Can Decrease Tissue Inflammation and the Severity of HSV-2 Infection in the Female Reproductive Mucosa. Int. J. Mol. Sci. 2020, 21, 337. [Google Scholar] [CrossRef] [PubMed]
- DiPaolo, M.; Woodworth, C.D.; Popescu, N.C.; Koval, D.L.; Lopez, J.V.; Doniger, J. HSV-2-Induced Tumorigenicity in HPV16-Immortalized Human Genital Keratinocytes. Virology 1990, 177, 777–779. [Google Scholar] [CrossRef] [PubMed]
- Hara, Y.; Kimoto, T.; Okuno, Y.; Minekawa, Y. Effect of Herpes Simplex Virus on the DNA of Human Papillomavirus 18. J. Med. Virol. 1997, 53, 4–12. [Google Scholar] [CrossRef]
- Li, S.; Wen, X. Seropositivity to Herpes Simplex Virus Type 2, but Not Type 1 is Associated with Cervical Cancer: NHANES (1999–2014). BMC Cancer 2017, 17, 726. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.S.; Herrero, R.; Bosetti, C.; Muñoz, N.; Bosch, F.X.; Eluf-Neto, J.; Castellsagué, X.; Meijer, C.J.; Van den Brule, A.J.; Franceschi, S.; et al. Herpes Simplex Virus-2 as a Human Papillomavirus Cofactor in the Etiology of Invasive Cervical Cancer. J. Natl. Cancer Inst. 2002, 94, 1604–1613. [Google Scholar] [CrossRef] [PubMed]
- Mogtomo, M.L.K.; Ngane, A.N.; Nganwa, G.D.; Wankam, M.; Epaka, C.B.; Zollo, P.H.A. Association of Cervical Inflammation and Cervical Abnormalities in Women Infected with Herpes Simplex Virus Type 2. Int. J. Trop. Med. Public Health 2014, 1, 1–4. [Google Scholar]
- Maza, M.; Gage, J.C. Considerations for HPV Primary Screening in Lower-Middle Income Countries. Prev. Med. 2017, 98, 39–41. [Google Scholar] [CrossRef] [PubMed]
- Steinau, M.; Patel, S.S.; Unger, E.R. Efficient DNA Extraction for HPV Genotyping in Formalin-Fixed, Paraffin-Embedded Tissues. J. Mol. Diagn. 2011, 13, 377–381. [Google Scholar] [CrossRef] [PubMed]
- Bruni, L.; Albero, G.; Serrano, B.; Mena, M.; Collado, J.J.; Gómez, D.; Muñoz, J.; Bosch, F.X.; de Sanjosé, S. Human Papillomavirus and Related Diseases in the World. ICO/IARC HPV Information Centre. Summary Report, 10 March 2023. Available online: https://hpvcentre.net/statistics/reports/XWX.pdf?t=1712869358562 (accessed on 12 April 2024).
- de Sanjosé, S.; Marshall, V.; Solà, J.; Palacio, V.; Almirall, R.; Goedert, J.J.; Bosch, F.X.; Whitby, D. Prevalence of Kaposi’s Sarcoma-Associated Herpesvirus Infection in Sex Workers and Women from the General Population in Spain. Int. J. Cancer 2002, 98, 155–158. [Google Scholar] [CrossRef] [PubMed]
- Brasil, C.M.; Ribeiro, C.M.; Leão, J.C. Oral and Genital Human Herpesvirus 8 and Human Papillomavirus in Heterosexual Partners. J. Oral. Pathol. Med. 2013, 42, 61–65. [Google Scholar] [CrossRef] [PubMed]
- Enbom, M.; Strand, A.; Falk, K.I.; Linde, A. Detection of Epstein–Barr Virus, but Not Human Herpesvirus 8, DNA in Cervical Secretions from Swedish Women by Real-Time Polymerase Chain Reaction. Sex. Transm. Dis. 2001, 28, 300–306. [Google Scholar] [CrossRef] [PubMed]
- Chan, P.K.; Li, W.H.; Chan, M.Y.; Cheng, A.F. Detection of Human Herpesvirus 8 in Cervical Cells of Chinese Women with Abnormal Papanicolaou Smears. Clin. Infect. Dis. 1999, 29, 1584–1585. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Cao, S.; Gan, Y.; Dong, X.; Lu, Z. Herpes Simplex Virus Type 2 and the Risk of Cervical Cancer: A Meta-Analysis of Observational Studies. Arch. Gynecol. Obstet. 2014, 290, 1059–1066. [Google Scholar] [CrossRef] [PubMed]
- Luo, Q.; Zeng, X.; Luo, H.; Pan, L.; Huang, Y.; Zhang, H.; Han, N. Epidemiologic Characteristics of High-Risk HPV and the Correlation between Multiple Infections and Cervical Lesions. BMC Infect. Dis. 2023, 23, 667. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Xia, X.; Zheng, W.; Dai, Y.; Zhuang, X. HPV Prevalence and Genotype Distribution Among Women in Eastern China During the COVID-19 Pandemic. Hum. Vaccin. Immunother. 2023, 19, 2212571. [Google Scholar] [CrossRef] [PubMed]
- Dickson, E.L.; Vogel, R.I.; Bliss, R.L.; Downs, L.S. Cervical Cytology and Multiple Type HPV Infection: A Study of 8182 Women Ages 31–65. Gynecol. Oncol. 2014, 133, 405–408. [Google Scholar] [CrossRef] [PubMed]
- Ahmadi, M.; Rasi, H.; Mostafazadeh, M.; Hajazimian, S.; Maroufi, N.F.; Nahaei, M.R.; Rahaee, S.; Isazadeh, A. Analysis of Cervical Lesions for Presence of HSV-2 and HPV-16 and HPV-18 in Iranian Patients by PCR. Horm. Mol. Biol. Clin. Investig. 2017, 31, 20170019. [Google Scholar] [CrossRef] [PubMed]
- Rocha, D.A.; Mariño, J.M.; Santos, C.M. Detection of Human Cytomegalovirus and Herpes Simplex Virus Type 2 in Cervical Sample. Rev. Bras. Ginecol. Obstet. 2012, 34, 499–504. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Joharinia, N.; Faghihinejad, S.; Seyedi, K.; Farhadi, A.; Hosseini, S.Y.; Safaei, A.; Bahrampour, H.; Sarvari, J. Co-Existing of HSV1/2 or EBV Infection with the Presence of High-Risk HPV DNA in Cervical Lesions in the Southwest of Iran. Asian Pac. J. Cancer Prev. 2020, 21, 1459–1464. [Google Scholar] [CrossRef] [PubMed]
- Pisani, S.; Imperi, M.; Seganti, L.; Superti, F.; Tinari, A.; Bucci, M.; Degener, A.M. Effect of HSV-2 Infection on the Expression of HPV 16 Genes in CaSki Cells. Int. J. Immunopathol. Pharmacol. 2004, 17, 65–70. [Google Scholar] [CrossRef] [PubMed]
- Lehtinen, M.; Koskela, P.; Jellum, E.; Bloigu, A.; Anttila, T.; Hallmans, G.; Luukkaala, T.; Thoresen, S.; Youngman, L.; Dillner, J.; et al. Herpes Simplex Virus and Risk of Cervical Cancer: A Longitudinal, Nested Case-Control Study in the Nordic Countries. Am. J. Epidemiol. 2002, 156, 687–692. [Google Scholar] [CrossRef] [PubMed]
- Okoye, J.O.; Ngokere, A.A.; Erinle, C.; Mbamalu, C. Co-Existence of Herpes Simplex Virus Type 2 and Two Other Oncoviruses is Associated with Cervical Lesions in Women Living with HIV in South-Western Nigeria. Afr. Health Sci. 2020, 20, 1015–1023. [Google Scholar] [CrossRef] [PubMed]
- Bahena-Román, M.; Sánchez-Alemán, M.A.; Contreras-Ochoa, C.O.; Lagunas-Martínez, A.; Olamendi-Portugal, M.; López-Estrada, G.; Delgado-Romero, K.; Guzmán-Olea, E.; Madrid-Marina, V.; Torres-Poveda, K. Prevalence of Active Infection by Herpes Simplex Virus Type 2 in Patients with High-Risk Human Papillomavirus Infection. J. Med. Virol. 2020, 92, 1246–1252. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Cao, X.; Zheng, Y.; Tang, J.; Cai, W.; Wang, H.; Gao, Y.; Wang, Y. Relationship between Cervical Disease and Infection with HPV Types 16 and 18, and HSV-1 and HSV-2. J. Med. Virol. 2012, 84, 1920–1927. [Google Scholar] [CrossRef] [PubMed]
- de Abreu, A.L.; Malaguti, N.; Souza, R.P.; Uchimura, N.S.; Ferreira, É.C.; Pereira, M.W.; Carvalho, M.D.; Pelloso, S.M.; Bonini, M.G.; Gimenes, F.; et al. Association of HPV, Neisseria gonorrhoeae and Chlamydia trachomatis Co-Infections on the Risk of High-Grade Squamous Intraepithelial Cervical Lesion. Am. J. Cancer Res. 2016, 6, 1371–1383. [Google Scholar] [PubMed]
- Pérez, L.O.; Barbisan, G.; Abba, M.C.; Laguens, R.M.; Dulout, F.N.; Golijow, C.D. Herpes Simplex Virus and HPV Infection in Cervical Disease in Argentine Women. Int. J. Gynecol. Pathol. 2006, 25, 42–47. [Google Scholar] [CrossRef] [PubMed]
- Bass, B.P.; Engel, K.B.; Greytak, S.R.; Moore, H.M. A Review of Preanalytical Factors Affecting Molecular, Protein, and Morphological Analysis of Formalin-Fixed, Paraffin-Embedded (FFPE) Tissue: How Well Do You Know Your FFPE Specimen? Arch. Pathol. Lab. Med. 2014, 138, 1520–1530. [Google Scholar] [CrossRef] [PubMed]
- Palmer, M.; Katanoda, K.; Saito, E.; Martellucci, C.A.; Tanaka, S.; Ikeda, S.; Sakamoto, H.; Machelek, D.; Brotherton, J.M.L.; Hocking, J.S. Genotype Prevalence and Age Distribution of Human Papillomavirus from Infection to Cervical Cancer in Japanese Women: A Systematic Review and Meta-Analysis. Vaccine 2022, 40, 5971–5996. [Google Scholar] [CrossRef] [PubMed]
- Kelly, H.; Mayaud, P.; Segondy, M.; Pai, N.P.; Peeling, R.W. A Systematic Review and Meta-Analysis of Studies Evaluating the Performance of Point-of-Care Tests for Human Papillomavirus Screening. Sex. Transm. Infect. 2017, 93, S36–S45. [Google Scholar] [CrossRef] [PubMed]
- Mai, Q.; Yang, X.; Cheng, H.; Wu, G.; Wu, Z. Prevalence and Genotype Distribution of Human Papillomavirus Among Women With Cervical Lesions in Shenzhen City, China. Hum. Vaccin. Immunother. 2021, 17, 965–971. [Google Scholar] [CrossRef] [PubMed]
- Yaman, M.; Özcan, H.E.A.; Bakir, A. High-Risk Human Papilloma Virus Genotype Distribution and Correlation with Cervical Cytomorphological Data in Turkish and Immigrant Women in Mersin Province. New Microbiol. 2024, 47, 88–97. [Google Scholar] [PubMed]
- Gunes, A.C.; Ozgul, N.; Turkyılmaz, M.; Kara, F.; Unlu, F.; Ayhan, A.; Gultekin, M. Evaluation of Colposcopy After the Addition of Human Papillomavirus Testing to the Turkish Cervical Cancer Screening Program. Cancer Med. 2023, 12, 21751–21760. [Google Scholar] [CrossRef] [PubMed]
- Akış, S.; Öztürk, U.K.; Keleş, E.; Alınca, C.M.; Kabaca, C.; Api, M. The Role of Multiple High-Risk Human Papillomavirus Infections for Cervical Biopsies and Findings in Colposcopic Procedures. J. Turk. Ger. Gynecol. Assoc. 2023, 24, 101–108. [Google Scholar] [CrossRef] [PubMed]
- Gecer, M. High-Risk Human Papillomavirus (hrHPV) Prevalence and Genotype Distribution Among Turkish Women. J. Cytol. 2023, 40, 42–48. [Google Scholar] [CrossRef] [PubMed]
- Agadayi, E.; Karademir, D.; Karahan, S. Knowledge, Attitudes and Behaviors of Women Who Have or Have Not Had Human Papillomavirus Vaccine in Turkey About the Virus and the Vaccine. J. Community Health 2022, 47, 650–657. [Google Scholar] [CrossRef] [PubMed]
- Hosseini, S.Z.; Makvandi, M.; Samarbafzade, A.; Timori, A.; Ranjbar, N.; Saki, N.; Nisi, N.; Shahani, T.; Varnaseri, M.; Angali Ahmadi, K. Frequency of Human Papillomavirus (HPV) 16 and 18 Detection in Paraffin-Embedded Laryngeal Carcinoma Tissue. Asian Pac. J. Cancer Prev. 2017, 18, 889–893. [Google Scholar] [PubMed]
- Cohen, J.I. Therapeutic Vaccines for Herpesviruses. J. Clin. Investig. 2024, 134, e179483. [Google Scholar] [CrossRef] [PubMed]

| Age Groups (Years) | p Value | |||||
|---|---|---|---|---|---|---|
| 30–39 | 40–49 | 50–59 | ≥60 | Total | ||
| HR-HPV genotype | n = 86 | n = 91 | n = 60 | n = 39 | n = 276 | |
| Negative | 42 (48.8) | 51 (56.0) | 39 (65.0) | 32 (82.1) | 164 (59.4) | 0.004 |
| Positive | 44 (51.2) | 40 (44.0) | 21 (35.0) | 7 (17.9) | 112 (40.6) | |
| Single infection | 18 (20.9) | 18 (19.8) | 12 (20.0) | 4 (10.3) | 52 (18.8) | |
| HPV 16 | 16 (18.6) | 17 (18.7) | 9 (15.0) | 4 (10.3) | 46 (16.7) | |
| HPV 18 | 2 (2.3) | 1 (1.1) | 3 (5.0) | 0 (0.0) | 6 (2.2) | |
| Multiple infection | 20 (23.3) | 12 (13.2) | 4 (6.7) | 0 (0.0) | 36 (13.0) | |
| HPV 35/52, HPV 33/51/59 | 4 (4.7) | 8 (8.8) | 0 (0.0) | 0 (0.0) | 12 (4.3) | |
| HPV 16, HPV 18 | 6 (7.0) | 1 (1.1) | 1 (1.7) | 0 (0.0) | 8 (2.9) | |
| HPV 16, HPV 35/ 52 | 4 (4.7) | 1 (1.1) | 2 (3.3) | 0 (0.0) | 7 (2.5) | |
| HPV 16, HPV 18, HPV 35/52 | 0 (0.0) | 1 (1.1) | 0 (0.0) | 0 (0.0) | 1 (0.4) | |
| HPV 18, HPV 35/52 | 3 (3.5) | 0 (0.0) | 1 (1.7) | 0 (0.0) | 4 (1.4) | |
| HPV 16, HPV 45 | 2 (2.3) | 1 (1.1) | 0 (0.0) | 0 (0.0) | 3 (1.1) | |
| HPV 16, HPV 33/51/59 | 1 (1.2) | 0 (0.0) | 0 (0.0) | 0 (0.0) | 1 (0.4) | |
| Untypable | 6 (7.0) | 10 (11.0) | 5 (8.3) | 3 (7.7) | 24 (8.7) | |
| HPV 35/52 | 4 (4.7) | 5 (5.5) | 5 (8.3) | 1 (2.6) | 15 (5.4) | |
| HPV 33/51/59 | 2 (2.3) | 5 (5.5) | 0 (0.0) | 2 (5.1) | 9 (3.3) | |
| HR-HPV DNA | Normal/ Non-Dysplastic | LSIL | HSIL | Total | p Value |
|---|---|---|---|---|---|
| n = 177 | n = 50 | n = 49 | n = 276 | ||
| Negative | 156 (88.1) | 6 (12.0) | 2 (4.1) | 164 (59.4) | <0.001 |
| Positive | 21 (11.9) | 44 (88.0) | 47 (95.9) | 112 (40.6) | |
| Single-type infection | 11 (6.2) | 24 (48.0) | 17 (34.7) | 52 (18.8) | |
| HPV 16 | 9 (5.1) | 21 (42.0) | 16 (32.7) | 46 (16.7) | |
| HPV 18 | 2 (1.1) | 3 (6.0) | 1 (2.0) | 6 (2.2) | |
| Multiple-type infection | 2 (1.1) | 9 (18.0) | 25 (51.0) | 36 (13.0) | |
| HPV 35/52, HPV 33/51/59 | 0 (0.0) | 0 (0.0) | 12 (24.5) | 12 (4.3) | |
| HPV 16, HPV 18 | 1 (0.6) | 6 (12.0) | 1 (2.0) | 8 (2.9) | |
| HPV 16, HPV 35/ 52 | 0 (0.0) | 1 (2.0) | 6 (12.2) | 7 (2.5) | |
| HPV 16, HPV 18, HPV 35/52 | 0 (0.0) | 0 (0.0) | 1 (2.0) | 1 (0.4) | |
| HPV 18, HPV 35/52 | 0 (0.0) | 2 (4.0) | 2 (4.1) | 4 (1.4) | |
| HPV 16, HPV 45 | 0 (0.0) | 0 (0.0) | 3 (6.1) | 3 (1.1) | |
| HPV 16, HPV 33/51/59 | 1 (0.6) | 0 (0.0) | 0 (0.0) | 1 (0.4) | |
| Untypable | 8 (4.5) | 11 (22.0) | 5 (10.2) | 24 (8.7) | |
| HPV 35/52 | 7 (4.0) | 6 (12.0) | 2 (4.1) | 15 (5.4) | |
| HPV 33/51/59 | 1 (0.6) | 5 (10.0) | 3 (6.1) | 9 (3.3) |
| Normal/Non Displastic | LSIL, HSIL | OR | 95% CI | p Value | |
|---|---|---|---|---|---|
| n | n | ||||
| HR-HPV status | |||||
| Negative | 156 | 8 | Reference | ||
| Positive | 21 | 91 | 84.5 | 35.96–198.57 | <0.001 |
| Genotype | |||||
| HPV-negative | 156 | 8 | Reference | ||
| HPV 16 | 9 | 37 | 80.17 | 28.98–221.78 | <0.001 |
| HPV 18 | 2 | 4 | 39.0 | 6.19–245.59 | <0.001 |
| HPV 16, HPV 18 | 1 | 7 | 136.5 | 14.94–1247.43 | <0.001 |
| HPV 35/52 | 7 | 8 | 19.5 | 5.5–69.15 | <0.001 |
| HPV 33/51/59 | 1 | 8 | 156.0 | 17.34–1403.63 | <0.001 |
| Age groups | |||||
| 30–39 | 6 | 38 | Reference | ||
| 40–49 | 8 | 32 | 1.06 | 0.26–4.31 | 0.928 |
| 50–59 | 6 | 15 | 1.36 | 0.23–8.17 | 0.734 |
| ≥60 | 1 | 6 | 1.12 | 0.10–12.15 | 0.928 |
| Infection pattern | |||||
| Single | 11 | 41 | Reference | ||
| Multiple | 2 | 34 | 4.56 | 0.95–22.0 | 0.059 |
| Untypable | 8 | 16 | 0.54 | 0.18–1.58 | 0.258 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bakır, A.; Yüzügüldü, B.; Tanık, E.B.; Kürkçü, M.F.; Korkut, G.; Duran, F.Ş. Investigation of the Prevalence of High-Risk Human Papillomavirus, Human Herpesvirus-8, and Herpes Simplex Virus-2 in Cervical Biopsy Samples Using the Real-Time PCR Method. Trop. Med. Infect. Dis. 2025, 10, 200. https://doi.org/10.3390/tropicalmed10070200
Bakır A, Yüzügüldü B, Tanık EB, Kürkçü MF, Korkut G, Duran FŞ. Investigation of the Prevalence of High-Risk Human Papillomavirus, Human Herpesvirus-8, and Herpes Simplex Virus-2 in Cervical Biopsy Samples Using the Real-Time PCR Method. Tropical Medicine and Infectious Disease. 2025; 10(7):200. https://doi.org/10.3390/tropicalmed10070200
Chicago/Turabian StyleBakır, Ayfer, Betül Yüzügüldü, Eylül Beren Tanık, Muhammed Furkan Kürkçü, Gizem Korkut, and Firdevs Şahin Duran. 2025. "Investigation of the Prevalence of High-Risk Human Papillomavirus, Human Herpesvirus-8, and Herpes Simplex Virus-2 in Cervical Biopsy Samples Using the Real-Time PCR Method" Tropical Medicine and Infectious Disease 10, no. 7: 200. https://doi.org/10.3390/tropicalmed10070200
APA StyleBakır, A., Yüzügüldü, B., Tanık, E. B., Kürkçü, M. F., Korkut, G., & Duran, F. Ş. (2025). Investigation of the Prevalence of High-Risk Human Papillomavirus, Human Herpesvirus-8, and Herpes Simplex Virus-2 in Cervical Biopsy Samples Using the Real-Time PCR Method. Tropical Medicine and Infectious Disease, 10(7), 200. https://doi.org/10.3390/tropicalmed10070200







