Montivipera bornmuelleri Venom: Inhibitory Effect on Staphylococcus epidermidis and Escherichia coli F1F0-ATPases and Cytotoxicity on HCT116 Cancer Cell Lines
Abstract
:1. Introduction
2. Materials and Methods
2.1. Reagents and Chemicals
2.2. Bacterial Cultures
2.3. Preparation of the Bacterial Sample
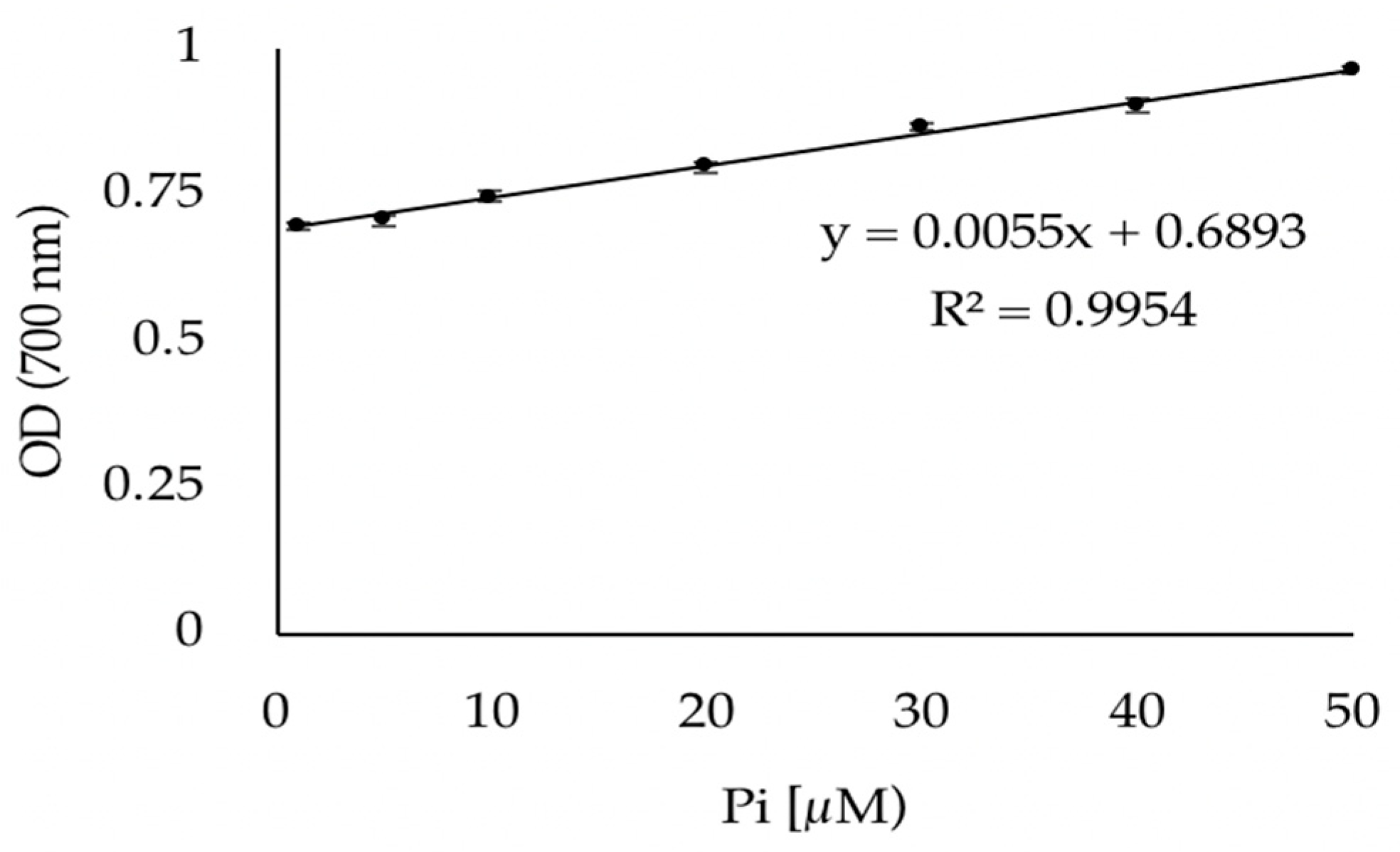
2.4. Phosphate Quantification Protocol
2.5. Study of the Inhibition of the Membrane-Bound S. epidermidis and E. coli ATP Synthases by Quercetin
2.6. Study of the Potential Inhibitory Effect of M. bornmuelleri Venom on the Membrane-Bound S. epidermidis and E. coli F1F0-ATPases
2.7. HCT116 Cell Culture
2.8. Cellular Viability Assay
2.9. Statistical Analysis
3. Results
3.1. Validation of the Phosphate Dosage Method
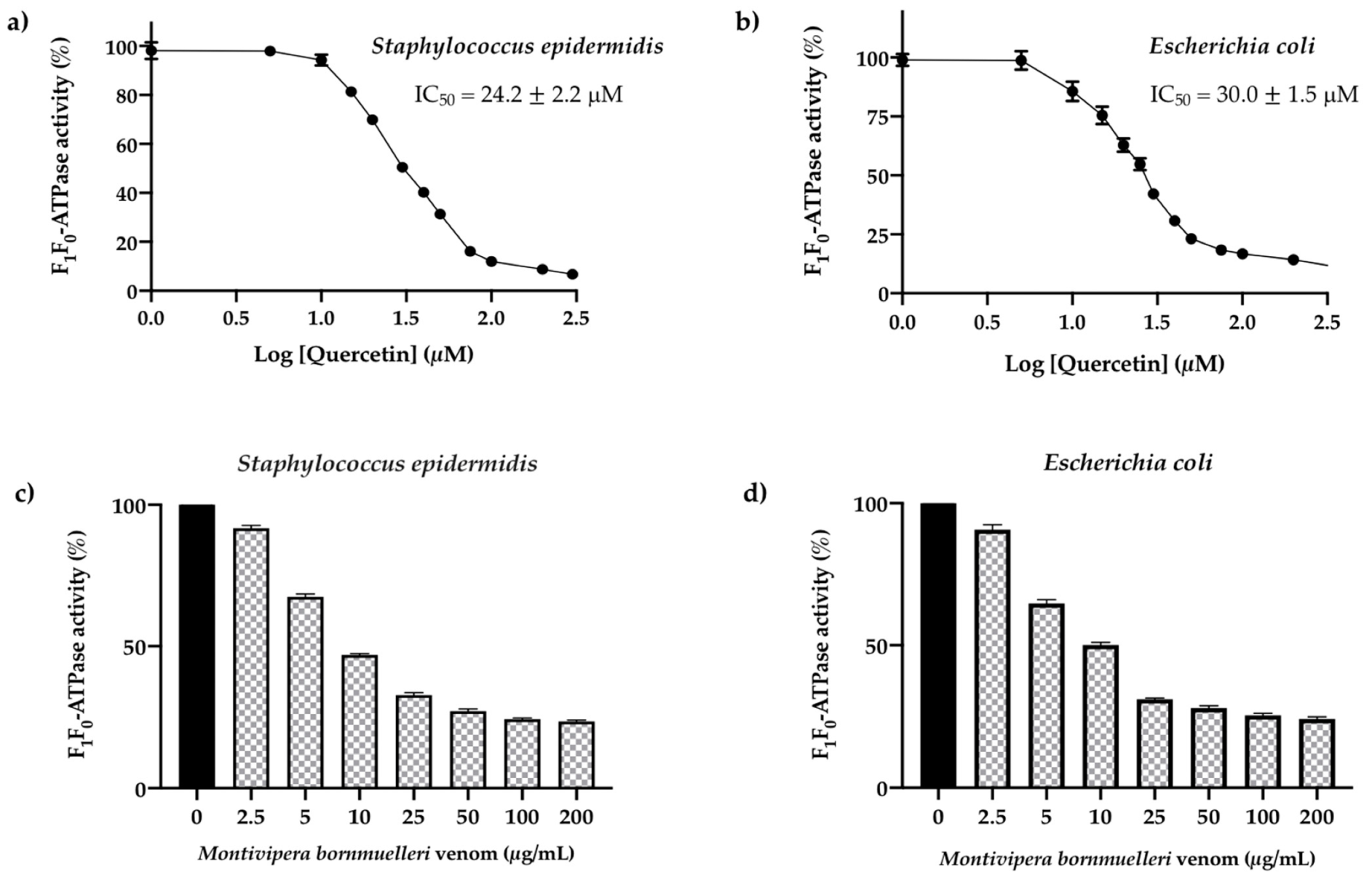
3.2. Screening of the Reference Inhibitor Quercetin on S. epidermidis and E. coli F1F0-ATPase Enzymes
3.3. Effect of the M. bornmuelleri Venom on the Membrane S. epidermidis and E. coli. F1F0-ATPase Enzymes
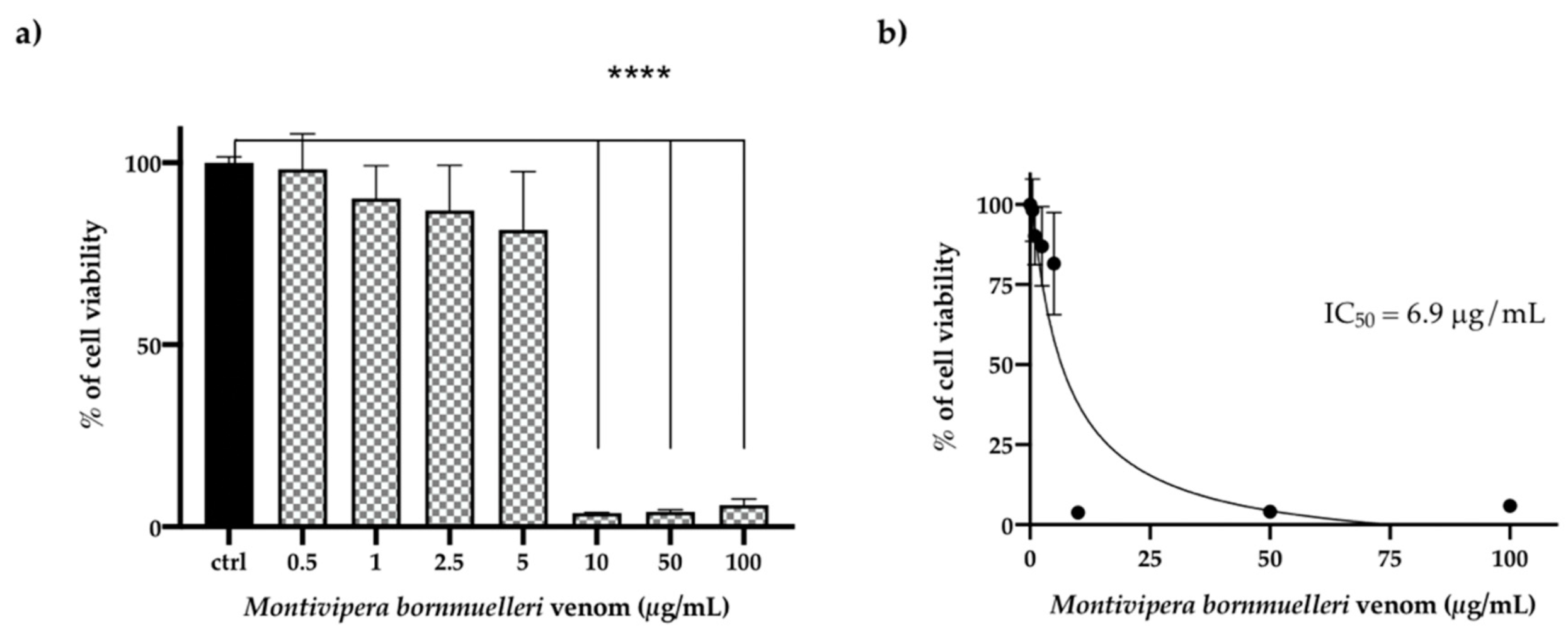
3.4. Effect of the M. bornmulleri Venom on HCT116 Cancer Cell Lines
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Rima, M.; Naini, S.M.A.; Karam, M.; Sadek, R.; Sabatier, J.-M.; Fajloun, Z. Vipers of the Middle East: A Rich Source of Bioactive Molecules. Molecules 2018, 23, 2721. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hraoui-bloquet, S.; Sadek, R.; Hleihel, W.; Fajloun, Z. An ecological study of the Lebanon mountain viper Montivipera bornmuelleri (werner, 1898) with a preliminary biochemical characterization of its venom. Lebanese Sci. J. 2012, 13, 89–101. [Google Scholar]
- Rima, M.; Accary, C.; Haddad, K.; Sadek, R.; Hraoui-Bloquet, S.; Desfontis, J.C.; Fajloun, Z. Identification of L-amino acid oxidase (Mb-LAAO) with antibacterial activity in the venom of Montivipera bornmuelleri, a viper from Lebanon. Infect. Disord. Drug Targets 2014, 13, 337–343. [Google Scholar] [CrossRef] [PubMed]
- Accary, C.; Hraoui-Bloquet, S.; Hamze, M.; Mallem, Y.; Omar, F.; Sabatier, J.-M.; Desfontis, J.-C.; Fajloun, Z. Protein Content Analysis and Antimicrobial Activity of the Crude Venom of Montivipera bornmuelleri; a Viper from Lebanon. Infect. Disord. Drug Targets 2014, 14, 49–55. [Google Scholar] [CrossRef] [PubMed]
- Sawan, S.; Yaacoub, T.; Hraoui-Bloquet, S.; Sadek, R.; Hleihel, W.; Fajloun, Z.; Karam, M. Montivipera bornmuelleri venom selectively exhibits high cytotoxic effects on keratinocytes cancer cell lines. Exp. Toxicol. Pathol. 2017, 69, 173–178. [Google Scholar] [CrossRef]
- Yacoub, T.; Rima, M.; Sadek, R.; Hleihel, W.; Fajloun, Z.; Karam, M. Montivipera bornmuelleri venom has immunomodulatory effects mainly up-regulating pro-inflammatory cytokines in the spleens of mice. Toxicol. Rep. 2018, 5, 318–323. [Google Scholar] [CrossRef] [PubMed]
- Accary, C.; Rima1, M.; Kouzayha, A.; Hleihel, W.; Sadek, S.; Desfontis, J.-C.; Fajloun, Z.; Hraoui-Bloquet, S. Effect of the Montivipera bornmuelleri snake venom on human blood: Coagulation disorders and hemolytic activities. Open J. Hematol. 2014, 5. [Google Scholar] [CrossRef] [Green Version]
- Accary, C.; Hraoui-Bloquet, S.; Sadek, R.; Alameddine, A.; Fajloun, Z.; Desfontis, J.-C.; Mallem, Y. The relaxant effect of the Montivipera bornmuelleri snake venom on vascular contractility. J. Venom Res. 2016, 7, 10–15. [Google Scholar]
- Accary, C.; Mantash, A.; Mallem, Y.; Fajloun, Z.; Elkak, A. Separation and Biological Activities of Phospholipase A2 (Mb-PLA2) from the Venom of Montivipera bornmuelleri, a Lebanese Viper. J. Liq. Chromatogr. Relat. Technol. 2015, 38, 833–839. [Google Scholar] [CrossRef]
- Aminov, R.I. A Brief History of the Antibiotic Era: Lessons Learned and Challenges for the Future. Front. Microbiol. 2010, 1, 134. [Google Scholar] [CrossRef] [Green Version]
- Alanis, A.J. Resistance to Antibiotics: Are We in the Post-Antibiotic Era? Arch. Med. Res. 2005, 36, 697–705. [Google Scholar] [CrossRef]
- Namvar, A.E.; Bastarahang, S.; Abbasi, N.; Ghehi, G.S.; Farhadbakhtiarian, S.; Arezi, P.; Hosseini, M.; Baravati, S.Z.; Jokar, Z.; Chermahin, S.G. Clinical characteristics of Staphylococcus epidermidis: A systematic review. GMS Hyg. Infect. Control 2014, 9, Doc23. [Google Scholar] [CrossRef] [PubMed]
- Penelitian, K. Strategi Implementasi., Baragina Widyaningrum, Program Pascasarjana. Nat. Rev. Microbiol. 2008, 7, 555–567. [Google Scholar] [CrossRef]
- Blount, Z.D. The unexhausted potential of E. coli. eLife 2015, 4, e05826. [Google Scholar] [CrossRef]
- Reinthaler, F.; Posch, J.; Feierl, G.; Wüst, G.; Haas, D.; Ruckenbauer, G.; Mascher, F.; Marth, E. Antibiotic resistance of E. coli in sewage and sludge. Water Res. 2003, 37, 1685–1690. [Google Scholar] [CrossRef]
- Hermolin, J.; Fillingame, R.H. H+-ATPase activity of Escherichia coli F1F0 is blocked after reaction of dicyclohexylcarbodiimide with a single proteolipid (subunit c) of the F0 complex. J. Biol. Chem. 1989, 264, 3896–3903. [Google Scholar] [CrossRef]
- Uribe-Alvarez, C.; Chiquete-Félix, N.; Contreras-Zentella, M.; Guerrero-Castillo, S.; Peña, A.; Uribe-Carvajal, S. Staphylococcus epidermidis: Metabolic adaptation and biofilm formation in response to different oxygen concentrations. Pathog. Dis. 2015, 74, ftv111. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hong, S.; Pedersen, P.L. ATP Synthase and the Actions of Inhibitors Utilized To Study Its Roles in Human Health, Disease, and Other Scientific Areas. Microbiol. Mol. Biol. Rev. 2008, 72, 590–641. [Google Scholar] [CrossRef] [Green Version]
- Devenish, R.J.; Prescott, M.; Rodgers, A.J. The Structure and Function of Mitochondrial F1F0-ATP Synthases. Int. Rev. Cell Mol. Biol. 2008, 267, 1–58. [Google Scholar] [CrossRef]
- Nesci, S.; Trombetti, F.; Algieri, C.; Pagliarani, A. A Therapeutic Role for the F1FO-ATP Synthase. SLAS Discov. Adv. Life Sci. R&D 2019, 24, 893–903. [Google Scholar] [CrossRef]
- Viegas, S.; Ladeira, C.; Costa-Veiga, A.; Perelman, J.; Gajski, G. Forgotten public health impacts of cancer—An overview. Arch. Ind. Hyg. Toxicol. 2017, 68, 287–297. [Google Scholar] [CrossRef] [Green Version]
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [Green Version]
- Shen, H.; Yang, J.; Huang, Q.; Jiang, M.-J.; Tan, Y.-N.; Fu, J.-F.; Zhu, L.-Z.; Fang, X.-F.; Yuan, Y. Different treatment strategies and molecular features between right-sided and left-sided colon cancers. World J. Gastroenterol. 2015, 21, 6470–6478. [Google Scholar] [CrossRef]
- Lindley, C.; McCune, J.S.; Thomason, T.E.; Lauder, D.; Sauls, A.; Adkins, S.; Msawyer, W.T.S. Perception of Chemotherapy Side Effects Cancer versus Noncancer Patients. Cancer Pract. 1999, 7, 59–65. [Google Scholar] [CrossRef]
- Monsuez, J.-J.; Charniot, J.-C.; Vignat, N.; Artigou, J.-Y. Cardiac side-effects of cancer chemotherapy. Int. J. Cardiol. 2010, 144, 3–15. [Google Scholar] [CrossRef]
- Issa, D.; Najjar, A.; Greige-Gerges, H.; Nehme, H.; Nehme, H. Screening of Some Essential Oil Constituents as Potential Inhibitors of the ATP Synthase of Escherichia coli. J. Food Sci. 2019, 84, 138–146. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- The Determination of Inorganic Phosphate in the Presence of Labile Phosphate Esters–PubMed. Available online: https://pubmed.ncbi.nlm.nih.gov/21018750/ (accessed on 30 May 2021).
- Mosmann, T. Rapid colorimetric assay for cellular growth and survival: Application to proliferation and cytotoxicity assays. J. Immunol. Methods 1983, 65, 55–63. [Google Scholar] [CrossRef]
- Nehme, H.; Ayde, H.; El Obeid, D.; Sabatier, J.M.; Fajloun, Z. Potential Inhibitory Effect of Apis mellifera’s Venom and of Its Two Main Components—Melittin and PLA2—On Escherichia coli F1F0-ATPase. Antibiotics 2020, 9, 824. [Google Scholar] [CrossRef] [PubMed]
- Dadi, P.K.; Ahmad, M.; Ahmad, Z. Inhibition of ATPase activity of Escherichia coli ATP synthase by polyphenols. Int. J. Biol. Macromol. 2009, 45, 72–79. [Google Scholar] [CrossRef]
- Breijyeh, Z.; Jubeh, B.; Karaman, R. Resistance of Gram-Negative Bacteria to Current Antibacterial Agents and Approaches to Resolve It. Molecules 2020, 25, 1340. [Google Scholar] [CrossRef] [Green Version]
- Courvalin, P. Transfer of Antibiotic Resistance Genes between Gram-Positive and Gram-Negative Bacteriat. Antimicrob. Agents Chemother. 1994, 38, 1447–1451. [Google Scholar] [CrossRef] [Green Version]
- Junior, N.G.D.O.; Cardoso, M.H.E.S.; Franco, O.L. Snake venoms: Attractive antimicrobial proteinaceous compounds for therapeutic purposes. Cell. Mol. Life Sci. 2013, 70, 4645–4658. [Google Scholar] [CrossRef] [PubMed]
- Samy, R.P.; Gopalakrishnakone, P.; Chow, V.; Ho, B. Viper metalloproteinase (Agkistrodon halys pallas) with antimicrobial activity against multi-drug resistant human pathogens. J. Cell. Physiol. 2008, 216, 54–68. [Google Scholar] [CrossRef]
- Alam, I.; Ojha, I.; Alam, A.; Quasimi, H.; Alam, O. Therapeutic potential of snake venoms as antimicrobial agents. Front. Drug Chem. Clin. Res. 2019, 2, 1–9. [Google Scholar] [CrossRef]
- Haddoub, C.; Rima, M.; Heurtebise, S.; Lawand, M.; Jundi, D.; Sadek, R.; Amigorena, S.; Fajloun, Z.; Karam, M.C. Cytotoxic effect of Montivipera bornmuelleri’s venom on cancer cell lines: In vitro and in vivo studies. PeerJ 2020, 8, e9909. [Google Scholar] [CrossRef]
- Park, M.H.; Jo, M.; Won, D.; Song, H.S.; Han, S.B.; Song, M.J.; Hong, J.T. Snake venom toxin from vipera lebetina turanicainduces apoptosis of colon cancer cells via upregulation of ROS- and JNK-mediated death receptor expression. BMC Cancer 2012, 12, 228. [Google Scholar] [CrossRef] [Green Version]
- Abidin, S.A.Z.; Rajadurai, P.; Chowdhury, E.H.; Othman, I.; Naidu, R. Cytotoxic, Anti-Proliferative and Apoptosis Activity of l-Amino Acid Oxidase from Malaysian Cryptelytrops purpureomaculatus (CP-LAAO) Venom on Human Colon Cancer Cells. Molecules 2018, 23, 1388. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tan, C.H.; Liew, J.L.; Navanesan, S.; Sim, K.S.; Tan, N.H.; Tan, K.Y. Cytotoxic and anticancer properties of the Malaysian mangrove pit viper (Trimeresurus purpureomaculatus) venom and its disintegrin (purpureomaculin). J. Venom. Anim. Toxins Incl. Trop. Dis. 2020, 26, e20200013. [Google Scholar] [CrossRef]
- Hiu, J.J.; Yap, M.K.K. Cytotoxicity of snake venom enzymatic toxins: Phospholipase A2 and l-amino acid oxidase. Biochem. Soc. Trans. 2020, 48, 719–731. [Google Scholar] [CrossRef] [Green Version]
- Galligan, L.; Longley, D.B.; McEwan, M.; Wilson, T.R.; McLaughlin, K.; Johnston, P.G. Chemotherapy and TRAIL-mediated colon cancer cell death: The roles of p53, TRAIL receptors, and c-FLIP. Mol. Cancer Ther. 2005, 4, 2026–2036. [Google Scholar] [CrossRef] [Green Version]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kfoury, M.; Mouawad, C.; Rifi, M.; Sadek, R.; Sabatier, J.-M.; Nehme, H.; Fajloun, Z. Montivipera bornmuelleri Venom: Inhibitory Effect on Staphylococcus epidermidis and Escherichia coli F1F0-ATPases and Cytotoxicity on HCT116 Cancer Cell Lines. Sci 2021, 3, 31. https://doi.org/10.3390/sci3030031
Kfoury M, Mouawad C, Rifi M, Sadek R, Sabatier J-M, Nehme H, Fajloun Z. Montivipera bornmuelleri Venom: Inhibitory Effect on Staphylococcus epidermidis and Escherichia coli F1F0-ATPases and Cytotoxicity on HCT116 Cancer Cell Lines. Sci. 2021; 3(3):31. https://doi.org/10.3390/sci3030031
Chicago/Turabian StyleKfoury, Milena, Charbel Mouawad, Mariam Rifi, Riyad Sadek, Jean-Marc Sabatier, Hala Nehme, and Ziad Fajloun. 2021. "Montivipera bornmuelleri Venom: Inhibitory Effect on Staphylococcus epidermidis and Escherichia coli F1F0-ATPases and Cytotoxicity on HCT116 Cancer Cell Lines" Sci 3, no. 3: 31. https://doi.org/10.3390/sci3030031
APA StyleKfoury, M., Mouawad, C., Rifi, M., Sadek, R., Sabatier, J.-M., Nehme, H., & Fajloun, Z. (2021). Montivipera bornmuelleri Venom: Inhibitory Effect on Staphylococcus epidermidis and Escherichia coli F1F0-ATPases and Cytotoxicity on HCT116 Cancer Cell Lines. Sci, 3(3), 31. https://doi.org/10.3390/sci3030031






