Abstract
The analytical microdosimetric function (AMF) implemented in the Monte Carlo code PHITS is a unique tool that bridges the gap between macro- and microscopic scales of radiation interactions, enabling accurate microdosimetric calculations over macroscopic bodies. The original AMF was published in 2006, based on the results of track structure calculations. Recently, a newer version of the AMF was proposed, incorporating an improved description of the energy loss at the microscopic scale. This study compares the older and the newer AMFs in computing microdosimetric probability distributions, mean values, and the relative biological effectiveness (RBE). To this end, 16000 microdosimetric lineal energy probability density distributions were simulated with PHITS for ions from 1H to 238U over a broad energy range (1–1000 MeV/n). The newer AMF was found to offer superior performance, particularly for very heavy ions, producing results that align more closely with published in vitro clonogenic survival experiments. These findings suggest that the updated AMF provides a more reliable tool for microdosimetric calculations and RBE modeling, essential for ion radiation therapy and space radiation protection.
1. Introduction
Due to the markedly different microscopic pattern of energy deposition, the biological effects of ions are markedly different from those of photons [1,2]. Macroscopic quantities such as the linear energy transfer (LET, defined as the mean energy lost by a charged particle per unit of path [3]) are unsuitable to univocally describe biological effects of ions on cells. As an example, different ions with the same LET can lead to significantly different biological effects, due to differences in their track-structure [4,5].
Microdosimetry aims to understand the correlation between ionizing radiation and biological effects by analyzing the stochastic nature of energy deposition at a biologically relevant scale (~µm) [6,7,8]. The variation in the microscopic pattern of energy deposition is quantified by means of probability distributions of microdosimetric quantities such as the lineal energy. The lineal energy () is the quotient between the energy imparted to a microscopic volume by a single energy-deposition event () and the mean chord length of that volume () [3,8]. The probability distribution of the lineal energy depends on the shape and dimensions of the microscopic volume, the type of radiation (i.e., photons or ions), and the kinetic energy of the particle [6,7,8,9,10].
The relative biological effectiveness (RBE), defined as the ratio of the dose of conventional photons to the dose of ions required to produce the same biological effect [11], is a central concept used for treatment planning in ion radiotherapy [12,13,14] and space radiation protection [15]. Microdosimetry-based RBE calculations [16,17] require three fundamental components: the microdosimetric distributions for the radiation types under study (i.e., ions and the reference photons), a biophysical RBE model, and cell- and endpoint-specific model parameters.
Monte Carlo track structure radiation transport codes [18,19,20,21,22,23] are commonly used to calculate micro- and nano- dosimetric probability distributions in microscopic targets. Due to the very long computational time, track structure calculations cannot be practically employed to compute microdosimetric probability distributions over macroscopic geometries such as a patient or an astronaut body. The Monte Carlo radiation transport code PHITS [24] includes a unique analytical approach for the computation of microdosimetric distributions, named the analytical microdosimetric function (AMF) [25]. The AMF serves as a bridge between largely different scales of radiation interactions, thus allowing the user to accurately compute microdosimetric calculations (~µm) over macroscopic bodies (~cm) in a reasonable amount of time [26].
The AMF [25] was successfully employed to model the particle-specific radiation-induced luminescence response of different dosimeters such as LiF:Mg,Ti and LiF:Mg,Cu,P thermoluminescent detectors [27,28,29,30], Al2O3:C optically stimulated luminescent detectors [31], and BaFBr:Eu optically stimulated luminescent detectors [32]. Furthermore, microdosimetric calculations with the AMF were instrumental for the development and benchmark of biophysical models describing and predicting in vitro clonogenic cell survival [33,34,35,36,37,38,39] and in vivo skin reactions [40].
Recently, an updated version of the AMF has been proposed based on the results of more recent track structure calculations [41]. This article presents a systematic comparison between the older [25] and the newer AMFs [41] in the computation of microdosimetric probability distributions, their mean values, and the RBE for ions from 1H to 238U.
2. Methodology
2.1. RBE
The endpoint chosen for the RBE calculation was 10% clonogenic survival (RBE10%). This quantity was defined as the ratio (Dref/Dion) of the absorbed doses needed to obtain a 10% cell surviving fraction when irradiating the cells with reference photons (Dref) and the ion beam under study (Dion).
The cell surviving fraction (S) as a function of the absorbed dose (D) was described by means of the linear quadratic model (LQM, Equation (1) [42,43]), where α and β are exposure- and cell-specific fitting parameters.
Subsequently, knowing the LQM terms for both the reference radiation (αref and βref) and the radiation under investigation (α and β), the RBE10% was calculated with Equation (2) [37] in the case of S = 10%.
2.2. Experimental RBE Data
Published in vitro results were extracted from the Particle Irradiation Data Ensemble (PIDE) version 3.4 [2,44] in the form of LQM terms for the ion exposure (α and β) and for the reference photons (αref and βref). The data filtering process follows that of previous studies [17,38,39]; thus, only a summary is given here. At first, we excluded experiments in which the cells were irradiated with ion energies lower than 1 MeV/n. In the case of hydrogen ions (1H and 2H), we included the results of exposures along monoenergetic and spread out Bragg peaks (SOBP), as the LET-dependence of the RBE was shown not to markedly depend on the irradiation scenario [45,46]. For heavier ions, we selected only results for monoenergetic beams since the LET-dependence of the RBE appears to differ between monoenergetic and SOBP exposures [45]. The V79 cell line was chosen as it is the most used mammalian cell line in the PIDE [2,44]. We selected the results only for the ten ions with most entries, as listed in Table 1.

Table 1.
Overview of the in vitro data for the V79 cell line: ions, number of survival curves for each ion, mean value of the α for the reference photon exposures (), and the mean value of the ratio of α and β for the reference photon exposures .
2.3. Computer Simulations
The PHITS code v3.32 [24] was used for all microdosimetric calculations. The simulations were carried out over an infinitesimal layer of water (track segment calculations) for clinically and space-relevant 1H, 4He, 12C, 16O, 20Ne, 40Ar, 56Fe, 84Kr, 132Xe, and 238U with 400 logarithmically spaced kinetic energy values between 1 and 1000 MeV/n.
The dose probability density distributions of the lineal energy d(y) were computed for spherical water targets of diameters equal to 0.3 µm and 0.5 µm using the older [25] and the newer AMFs [41] implemented in the [T-SED] tally of PHITS. The dose probability density distribution of the lineal energy d(y) represents the probability that one event deposits a fraction of the total absorbed dose in the lineal energy range between y and y + dy [3,6,8].
The mathematical formalism of the two AMFs, based on the work of Olko and Booz [9], was presented in detail in previous articles [25,26,41]. The simulated lineal energy distributions were scored using 500 logarithmically spaced bins between 10−3 and 107 keV/µm (50 bins per decade). The production of secondary particles was deactivated by setting cmin = 1010 MeV in the PHITS input file. As recommended by the International Commission on Radiation Units and Measurements (ICRU), the mean excitation energy of water was set to 78 eV [47].
The non-stochastic expectation value of the dose density distribution of the lineal energy is the dose-mean lineal energy and it is calculated as in Equation (3) [3,6,8].
The probability density distribution of the unrestricted LET in water was scored using the [T-LET] tally of PHITS, based on the ATIMA stopping power model (https://web-docs.gsi.de/~weick/atima/, accessed on 28 June 2024). As the simulated source was purely monoenergetic and the calculations were carried out in track segment conditions (no slowing down, no nuclear reaction) a single LET value was determined for each ion-energy combination.
For comparison purposes, the LET was also assessed using the Stopping and Range of Ions in Matter (SRIM) v2013 [48].
2.4. RBE Modeling
The Mayo Clinic Florida microdosimetric kinetic model (MCF MKM) [38,39] is a biophysical model that describes the change in the RBE of different particles based on their microscopic pattern of energy deposition. The MCF MKM is based on the theoretical framework of the MKM [49,50,51], grounded in the theory of dual radiation action [52]. In the MKM and its subsequent iterations, the cell nucleus is conceptualized as comprising subnuclear radiation structures, called domains, where the accumulation of lethal and sublethal radiation damage culminates in the loss of the cell’s clonogenic capability. In the MCF MKM, the domains are supposed to represent giant loops of chromatin, subnuclear structures containing ~2 Mbp of DNA [53,54,55].
The MCF MKM [38,39] was developed to address certain drawbacks of previous MKMs [17] such as decreased accuracy at high LET and low surviving fractions. Furthermore, the MCF MKM introduced novel strategies to overcome the need for ion-irradiated in vitro data to determine model parameters [38,39].
For each ion-energy combination included in this study, the LQM terms of clonogenic survival were predicted with the MCF MKM using Equations (4) and (5) (paragraph SM1 of Supplementary Materials)
where y is the lineal energy, d(y) is the dose probability density of the lineal energy, αref is the linear term of the LQM for the reference photons, βref is the quadratic term of the LQM for the reference photons, is the dose-mean lineal energy for the reference photons, ρ is the density (=1 g/cm3), rd is the mean radius of the subnuclear domains, and is the mean radius of the cell nucleus.
The ion-specific numerical values of αref and αref/βref used in the MCF MKM calculations match those of the in vitro experiments and are listed in Table 1. A representative, average value of = 4 keV/µm was used in all calculations, as the cell irradiations were carried out with different photon radiation types, the details of which are generally underreported. Furthermore, previous results suggest that the choice of has a relatively minor effect on the calculated RBE, especially for ions heavier than protons [40,46]. The cell-specific values of rd and for the V79 cell line were previously determined from measurable cell characteristics and equal to 0.27 µm and 4.0 µm, respectively [38,39].
2.5. Structure of the Study
The article comprises four main parts. At first, we present a comparison between the dose probability density distributions d(y) of the lineal energy assessed with the older [25] and the newer AMFs [41] for selected ions (1H, 12C, 56Fe, and 238U) and energies (1 MeV/n and 100 MeV/n) in the case of spherical water targets with radii of 0.3 µm and 0.5 µm. The spherical targets with a 0.3 µm radius were chosen due to their relevance for the RBE calculations, as they represent the mean calculated dimension of giant chromatin loops [17,38,39]. On the other hand, a spherical volume with a radius equal to 0.5 µm represents the most common target size for experimental measurements with microdosimetric gas detectors [7,8].
Secondly, we analyzed the dose-mean lineal energy as a function of the kinetic energy for all ions included in this study (1H, 4He, 12C, 16O, 20Ne, 40Ar, 56Fe, 84Kr, 132Xe, and 238U), the two AMFs, and the two microdosimetric targets (water spheres with radii equal to 0.3 µm and 0.5 µm).
Thirdly, we evaluated the RBE10% of the V79 cell line using the MCF MKM [38,39] in combination with the microdosimetric distributions calculated with the two AMFs [25,41] for the reference microdosimetric targets (water spheres with radii equal to 0.3 µm). Corresponding in vitro values from the PIDE [2,44] for 1H, 4He, 12C, 16O, 20Ne, 40Ar, 56Fe, 84Kr, 132Xe, and 238U ions were used as validation.
Finally, using the newer AMF [41], we investigated the extent to which the simulated RBE10% of the V79 cell line is affected by the target size used in the microdosimetric calculations (spheres with radii equal to 0.3 µm or 0.5 µm, i.e., reference targets vs. the most used target size).
3. Results and Discussion
3.1. LET
As a first validation of our calculations, Figure 1 compares the unrestricted LET in water computed using PHITS and SRIM (1H, 4He, 12C, 16O, 20Ne, 40Ar, 56Fe, 84Kr, 132Xe, and 238U ions) with published data from the Errata and Addenda for ICRU Report 73 (C, O, Ne, and Ar ions) [56], the ICRU Report 90 (H, He, and C ions) [47], and calculations with the track structure code KURBUC (H, He, C, O, Ne, and Fe ions) [19].
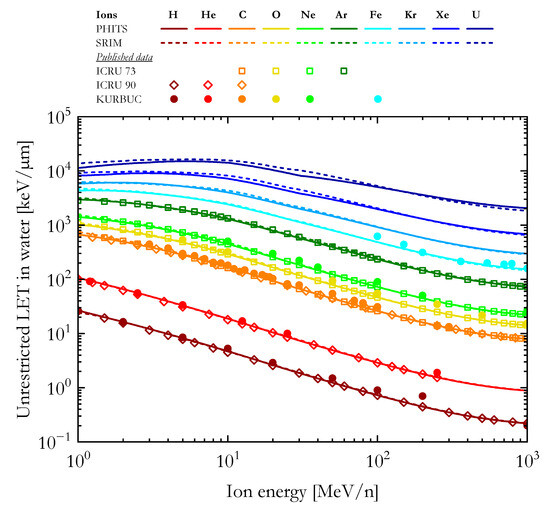
Figure 1.
The unrestricted LET in water as a function of the ion energy calculated with PHITS and SRIM in comparison with published data from the Errata and Addenda for ICRU Report 73 [56], ICRU Report 90 [47], and KURBUC [19] simulations.
Figure 1.
The unrestricted LET in water as a function of the ion energy calculated with PHITS and SRIM in comparison with published data from the Errata and Addenda for ICRU Report 73 [56], ICRU Report 90 [47], and KURBUC [19] simulations.
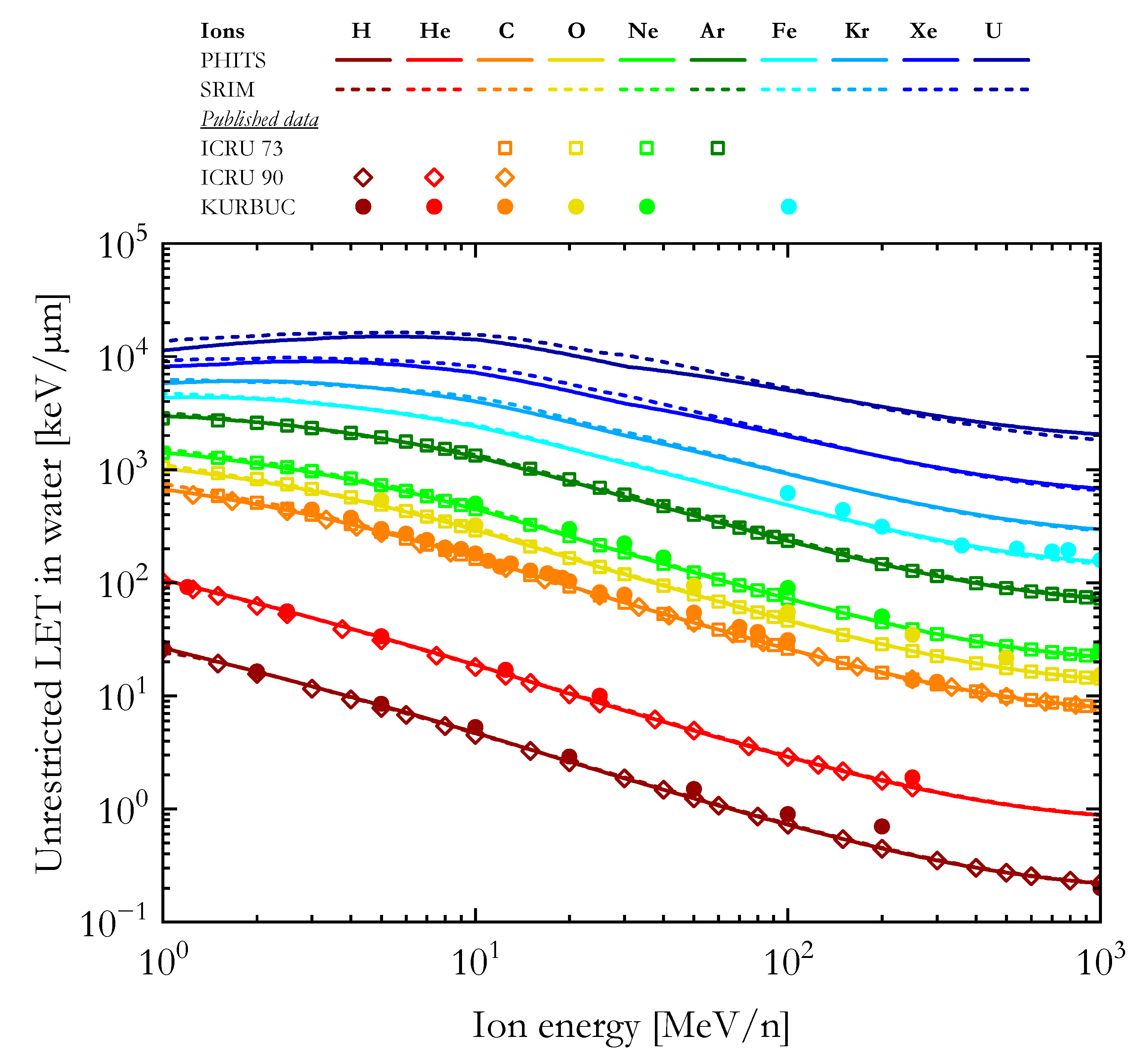
For ions from 1H to 40Ar, the LET values computed with PHITS and SRIM are very similar and agree well with the data from ICRU Reports 73 and 90 [47,56]. By contrast, the results of the KURBUC calculations [19] are generally higher than the other datasets, especially for ions with energy >100 MeV/n. The PHITS and SRIM LET values for 56Fe ions agree well with each other and reasonably well with the high energy KURBUC calculations [19]. No ICRU data was available for ions heavier than argon. For the three heaviest ions included in this article (Kr, Xe, and U), the LET values calculated with PHITS are lower than the SRIM results for ions with energies higher than 10 MeV/n, consistent with what was reported in a previous study [57].
3.2. Lineal Energy Distributions
Figure 2 compares the dose distribution of the lineal energy for 1H and 12C ions calculated with the two AMFs [25,41] for spherical water targets with radii of 0.3 µm and 0.5 µm. Two energies, 1 and 100 MeV/n, were simulated for each ion. In panels A, B, E, and F, the distributions are plotted in the standard semilogarithmic yd(y) vs. y representation where the area under the curve between two energy lineal points is proportional to the absorbed dose in the considered interval [6]. In contrast, the results in panels C, D, G, and H are plotted on a bilogarithmic scale to highlight the differences between the distributions at low lineal energy values (y < 1 keV/µm).
For 100 MeV protons (Figure 2A), the distributions are composed of two peaks corresponding to the proton events (at lower y) and the secondary electrons (extending up to 20–40 keV/µm). The edge of the electron peak is located at higher lineal energy values for the simulations with the smaller spherical targets. It is worth noting that the maximum of the proton peak calculated with the newer AMF is located at higher lineal energy values than the other distributions (~0.8 keV/µm instead of ~0.6 keV/µm). The four dose distributions of the lineal energy for 1 MeV protons (Figure 2B) are similar, featuring a single peak with a maximum between 30–40 keV/µm. For both simulated target sizes, the results of the newer AMF span over higher lineal energy values. Furthermore, the dose contribution of low lineal energy events was found to be larger in the calculations using the older AMF, as shown in Figure 2D.
The lineal energy distributions for 100 MeV/n carbon ions (Figure 2E) include a clear contribution of secondary electrons up to ~10 keV/µm. However, the electron edge is not visible, as it overlaps with the peak from carbon ion events. As for the low energy protons (Figure 2B), the peak of 1 MeV/n and 100 MeV/n carbon ion events extend to higher lineal energy values in the case of calculations with the newer AMF, as shown in Figure 2E,F. Similarly, a higher dose contribution of low lineal energy events was observed in the results of the older AMF (Figure 2G,H). These differences are attributed to several assumptions introduced in the track-structure simulation for developing the older AMF, including the core and penumbra concept [58] and the free-gas-based secondary electron production model [59].
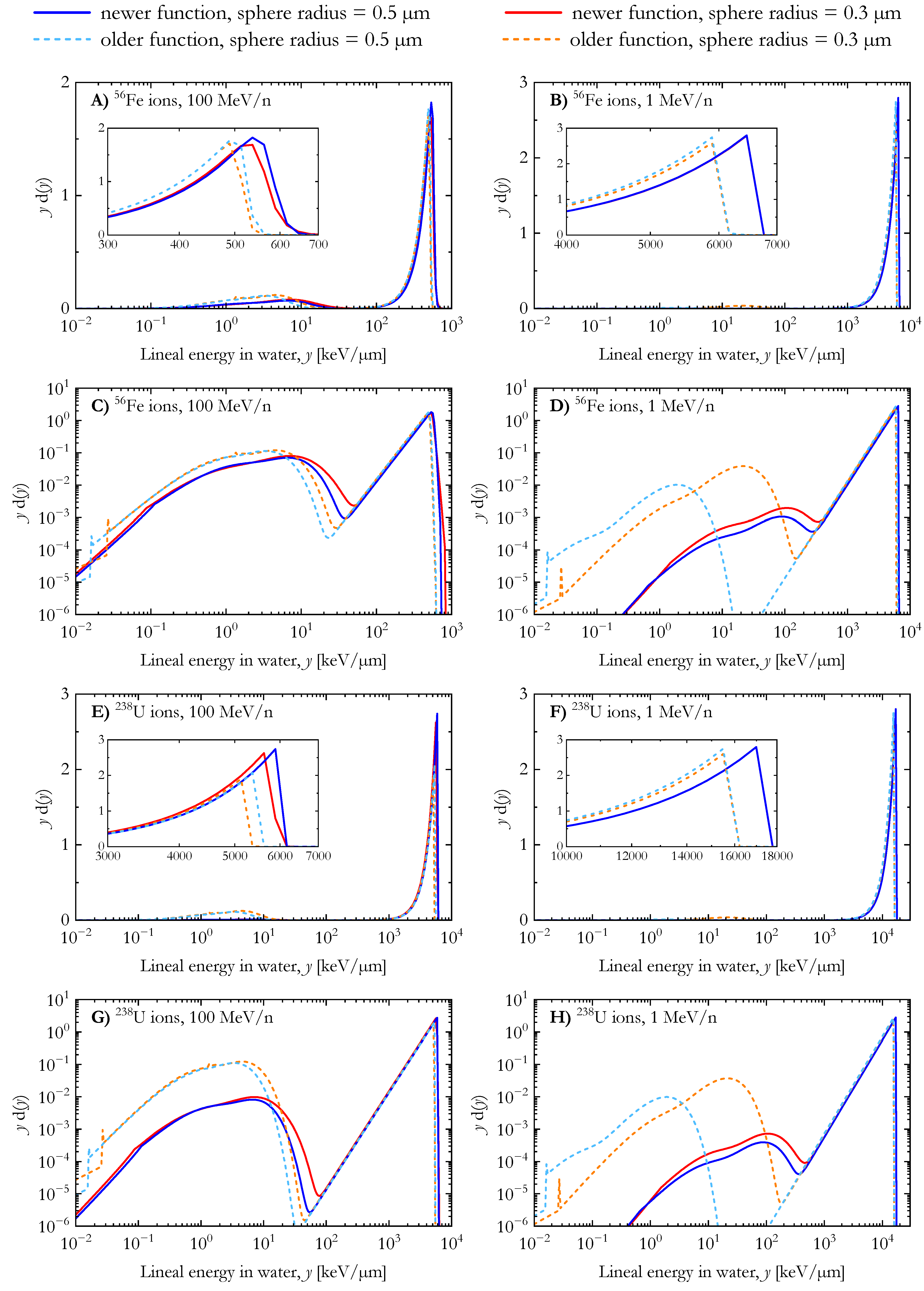
For 56Fe and 238U ions (Figure 3), the differences in the results of the two AMFs follow the same pattern as those for the carbon ions of Figure 2, namely, (a) the relative dose contribution from low lineal energy events (secondary electrons) is higher in the case of calculations using the older AMF and (b) the peak in the distribution due to ion events spans over higher lineal energy values for the calculations with the newer AMF.
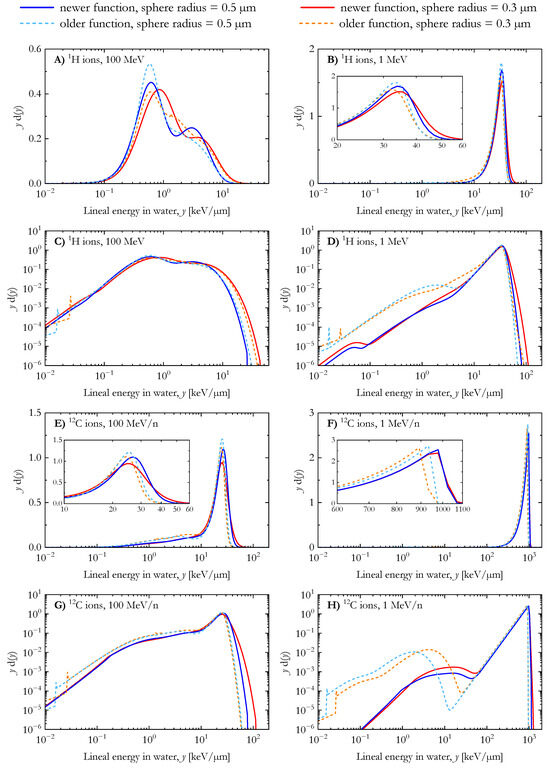
Figure 2.
Dose distribution of the lineal energy for 1H and 12C ions (energy: 1 MeV/n and 100 MeV/n) calculated with the newer and the older analytical microdosimetric functions in the case of spherical water targets with radii of 0.3 µm and 0.5 µm.
Figure 2.
Dose distribution of the lineal energy for 1H and 12C ions (energy: 1 MeV/n and 100 MeV/n) calculated with the newer and the older analytical microdosimetric functions in the case of spherical water targets with radii of 0.3 µm and 0.5 µm.
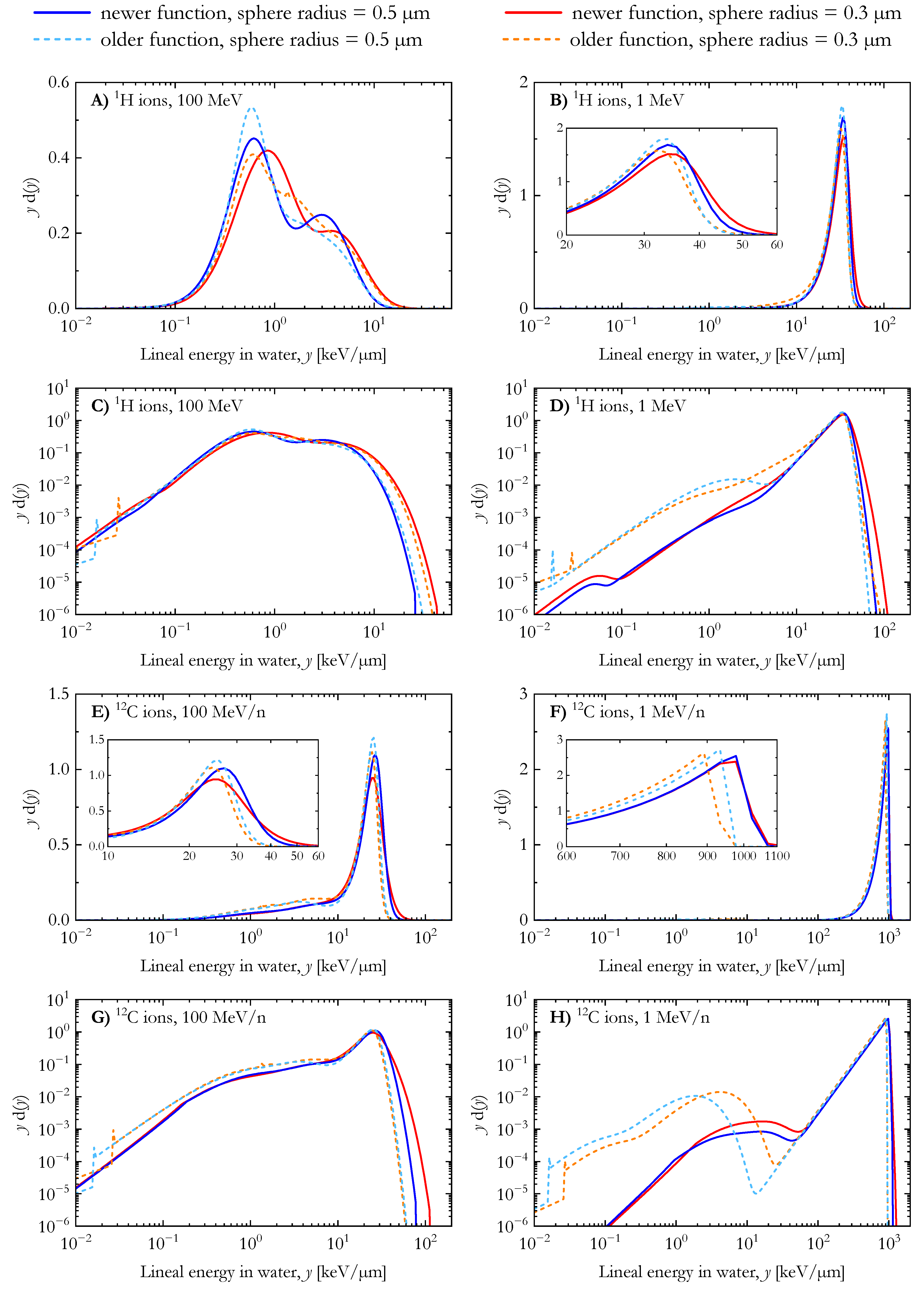
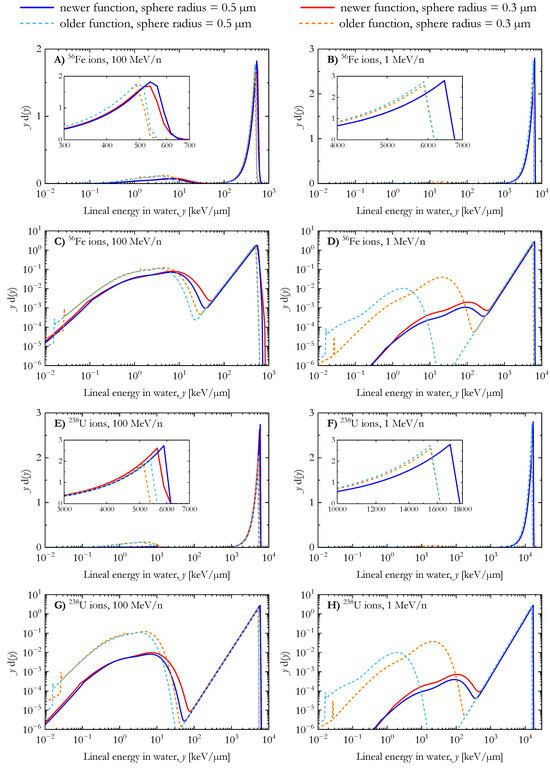
Figure 3.
Dose distribution of the lineal energy for 56Fe and 238U ions (energy: 1 MeV/n and 100 MeV/n) calculated with the newer and the older analytical microdosimetric functions in the case of spherical water targets with radii of 0.3 µm and 0.5 µm.
Figure 3.
Dose distribution of the lineal energy for 56Fe and 238U ions (energy: 1 MeV/n and 100 MeV/n) calculated with the newer and the older analytical microdosimetric functions in the case of spherical water targets with radii of 0.3 µm and 0.5 µm.
3.3. Dose-Mean Lineal Energy
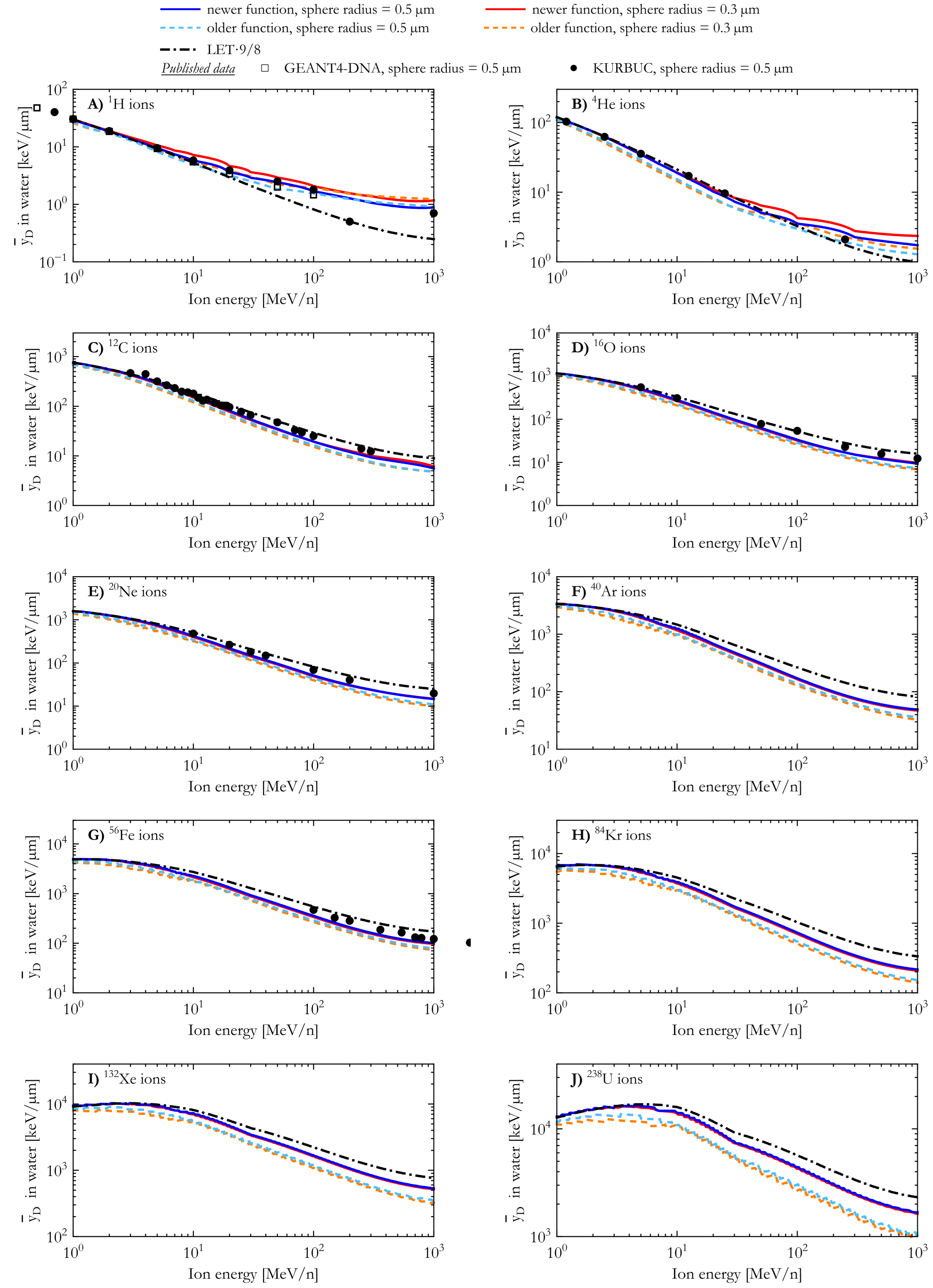
The simulated dose-mean lineal energy () of 1H, 4He, 12C, 16O, 20Ne, 40Ar, 56Fe, 84Kr, 132Xe, and 238U ions is plotted in Figure 4 as a function of the ion energy. The results of both AMFs and both spherical target sizes (sphere radius = 0.3 µm and 0.5 µm) were compared with published results [19,60] obtained with the track structure codes GEANT4-DNA and KURBUC for spherical targets of radius 0.5 µm.
Furthermore, was also calculated by multiplying the simulated LET of Figure 1 by a fixed factor of 9/8. This factor can be theoretically derived from the ratio between the second and first moment of the chord length distribution for a spherical target [61,62] and represents an estimate of /LET at relatively low energies. This correlation does not hold at energies above a few MeV because of changes in the energy-, ion-, and target- dependent contribution of secondary electrons.
For 1H and 4He ions and both AMFs (Figure 4A,B), the for the spherical targets with a 0.3 µm radius was found to be generally higher than the corresponding calculations with the larger spheres (radius = 0.5 µm). For a chosen AMF, the differences between the results of the two target sizes become smaller at low energies. For a chosen target size, the values calculated using the newer AMF were generally higher than those for the older AMF.
For proton energies > 10 MeV/n, the spread in the literature results (sphere radius = 0.5 µm) of the KURBUC and GEANT4-DNA codes [19,60] prevents determining which AMF aligns better with the existing data. At lower proton energies, the results of the newer AMF (sphere radius = 0.5 µm) are in good agreement with the literature results and the LET-based calculations. By contrast, the older AMF (sphere radius = 0.5 µm) appears to underestimate the other datasets. In the case of 4He ions and spheres with a 0.5 µm radius, the published assessed with the KURBUC [19] are best described by the corresponding results of the newer AMF.
For 12C and heavier ions, the values of a given AMF appear to be minorly affected by the choice of the target size (sphere radius = 0.3 µm and 0.5 µm). The results of the newer AMF were systematically higher than those of the older AMF and closer to the corresponding KURBUC data [19] and the theoretical LET-based calculations of . It should be noted that the values calculated with the newer AMF were lower than the KURBUC results in a similar manner to how our LET calculations, which are in good agreement with the ICRU LET data [47,56], were lower than the LET values estimated with KURBUC (see Figure 1).
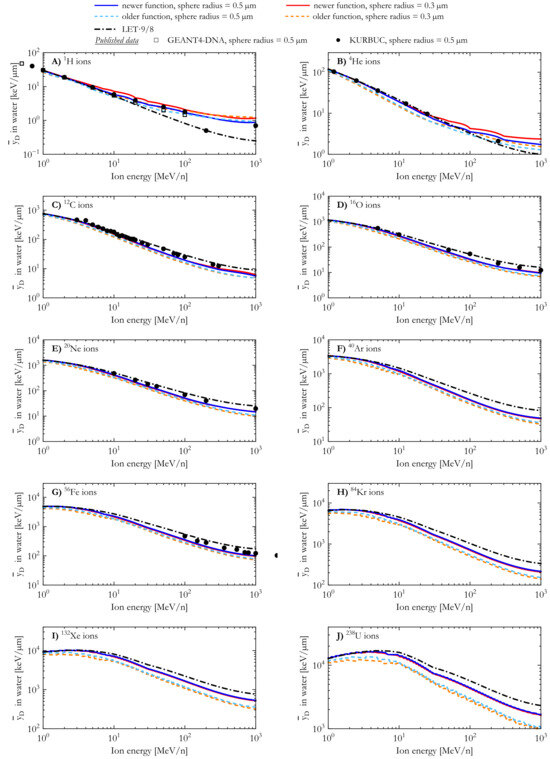
Figure 4.
Dose-mean lineal energy for 1H, 4He, 12C, 16O, 20Ne, 40Ar, 56Fe, 84Kr, 132Xe, and 238U ions calculated with the newer and the older analytical microdosimetric functions in the case of spherical water targets with radii of 0.3 µm and 0.5 µm. The results are compared with published results from the GEANT4-DNA (1H ions [60]) and KURBUC (1H, 4He, 12C, 16O, 20Ne, and 56Fe ions [19]) track structure codes, and corresponding theoretically derived LET-based estimates.
Figure 4.
Dose-mean lineal energy for 1H, 4He, 12C, 16O, 20Ne, 40Ar, 56Fe, 84Kr, 132Xe, and 238U ions calculated with the newer and the older analytical microdosimetric functions in the case of spherical water targets with radii of 0.3 µm and 0.5 µm. The results are compared with published results from the GEANT4-DNA (1H ions [60]) and KURBUC (1H, 4He, 12C, 16O, 20Ne, and 56Fe ions [19]) track structure codes, and corresponding theoretically derived LET-based estimates.
3.4. RBE
At first, we investigated the differences in the RBE10% of the V79 cell line calculated with the MCF MKM in conjunction with microdosimetric distributions simulated in the reference microdosimetric targets (spheres with a radius of 0.3 µm) using the older and the newer AMFs. The results are shown in Figure 5 as a function of the LET for the ten ions included in this study. Corresponding in vitro data from the PIDE [2,44] are also plotted in Figure 5.
For 1H and 4He, the RBE10% values calculated using the newer AMF were systematically higher than those obtained with the older AMF (Figure 5A,B). For ions from 12C to 132Xe (Figure 5C–I), the newer AMF initially yielded lower RBE10% values compared to the older AMF. Taking the RBE10% computed with the newer AMF as a reference, an increase in the ion LET led to a relative increase in the results of the older AMF. At LET > 1000 keV/µm, the RBE10% values obtained with the newer AMF were systematically lower than those of the older AMF.
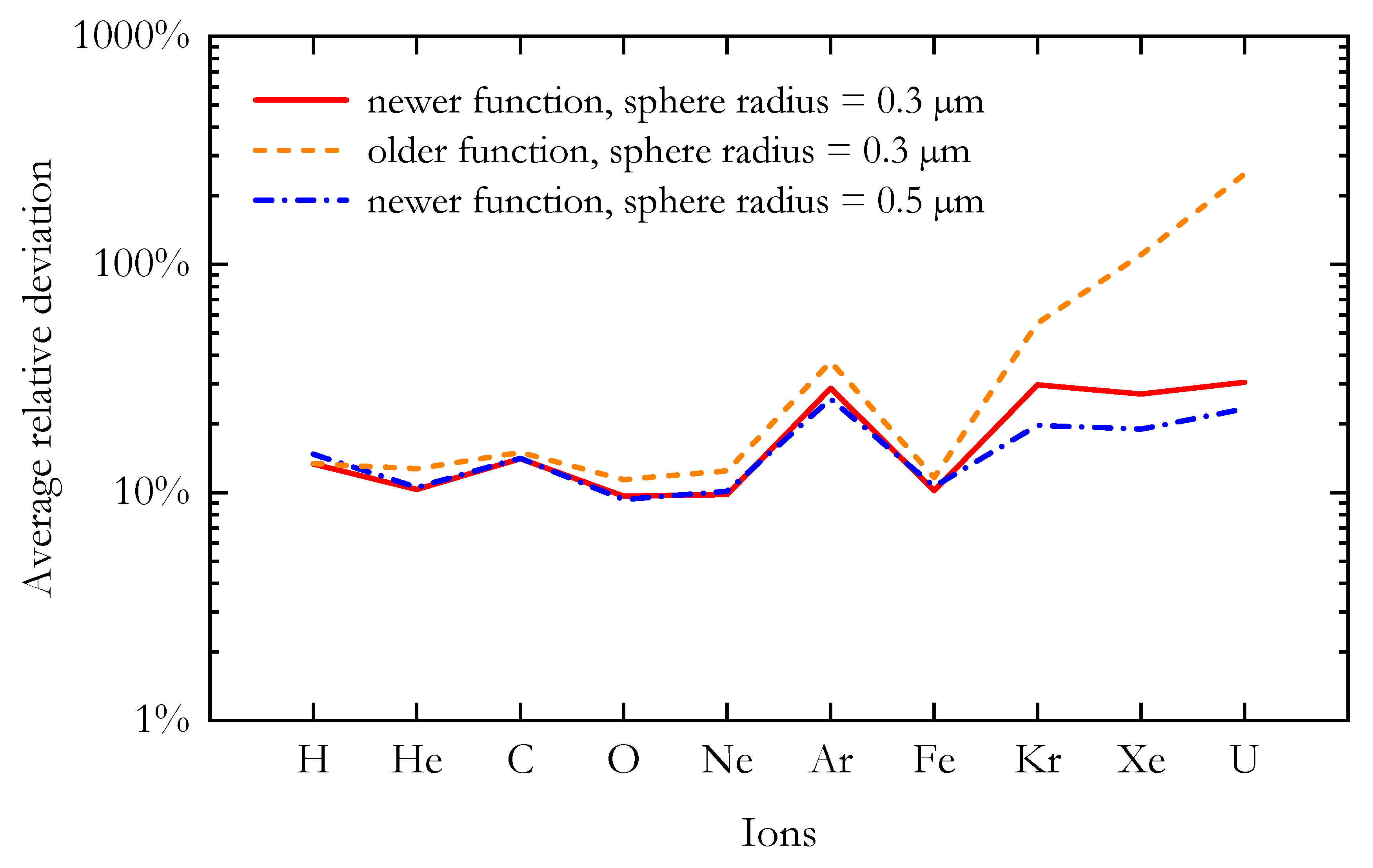
A comparison between the predicted RBE10% values and the corresponding in vitro data suggests that the newer AMF appears to better reproduce the experimental results, especially at high LET. In order to better visualize the agreement between the modelled and the corresponding n experimental data for a chosen ion, Figure 6 shows the average relative deviation () for the ten different ions included in this study. The average relative deviation between in silico and in vitro RBE data was lower for the newer AMF across all tested ions.
Secondly, we investigated the effect of carrying out the microdosimetric calculations with the newer AMF in two different target sizes: the reference water spheres with a radius of 0.3 µm (representative of the dimensions of the subnuclear domains in the MCF MKM) and spheres with a radius of 0.5 µm (representative of the most common target size employed in experimental microdosimetric measurements with gas detectors). As can be seen from Figure 5 and Figure 6, the differences between the two datasets were relatively minor, especially for ions heavier than protons. For very heavy ions (84Kr, 132Xe, and 238U), the calculations for a 0.5 µm target radius were closer to the experimental results. However, most of those results were extracted from a relatively old publication [63] whose LET calculations were reported to differ from the PHITS-based LET results [57], thus introducing additional uncertainties which are difficult to quantify.
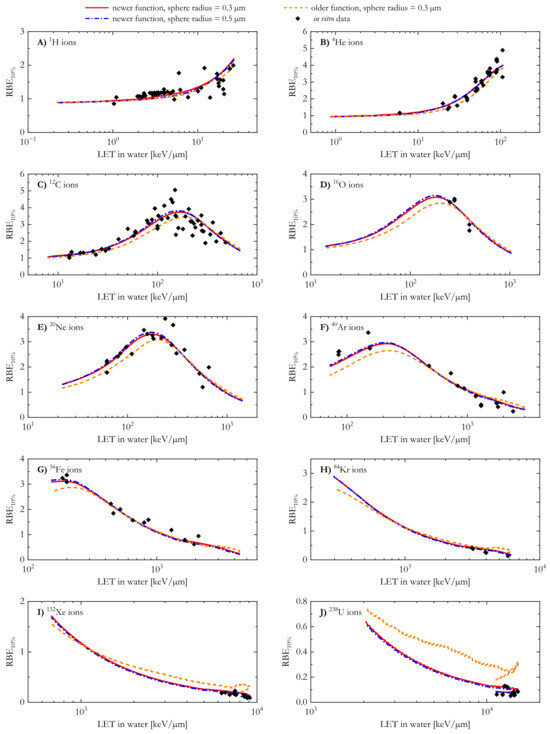
Figure 5.
Clonogenic survival RBE10% calculated with the newer and the older analytical microdosimetric functions for Chinese Hamster lung fibroblasts (V79 cell line) in comparison with corresponding in vitro results from the PIDE [2,44].
Figure 5.
Clonogenic survival RBE10% calculated with the newer and the older analytical microdosimetric functions for Chinese Hamster lung fibroblasts (V79 cell line) in comparison with corresponding in vitro results from the PIDE [2,44].
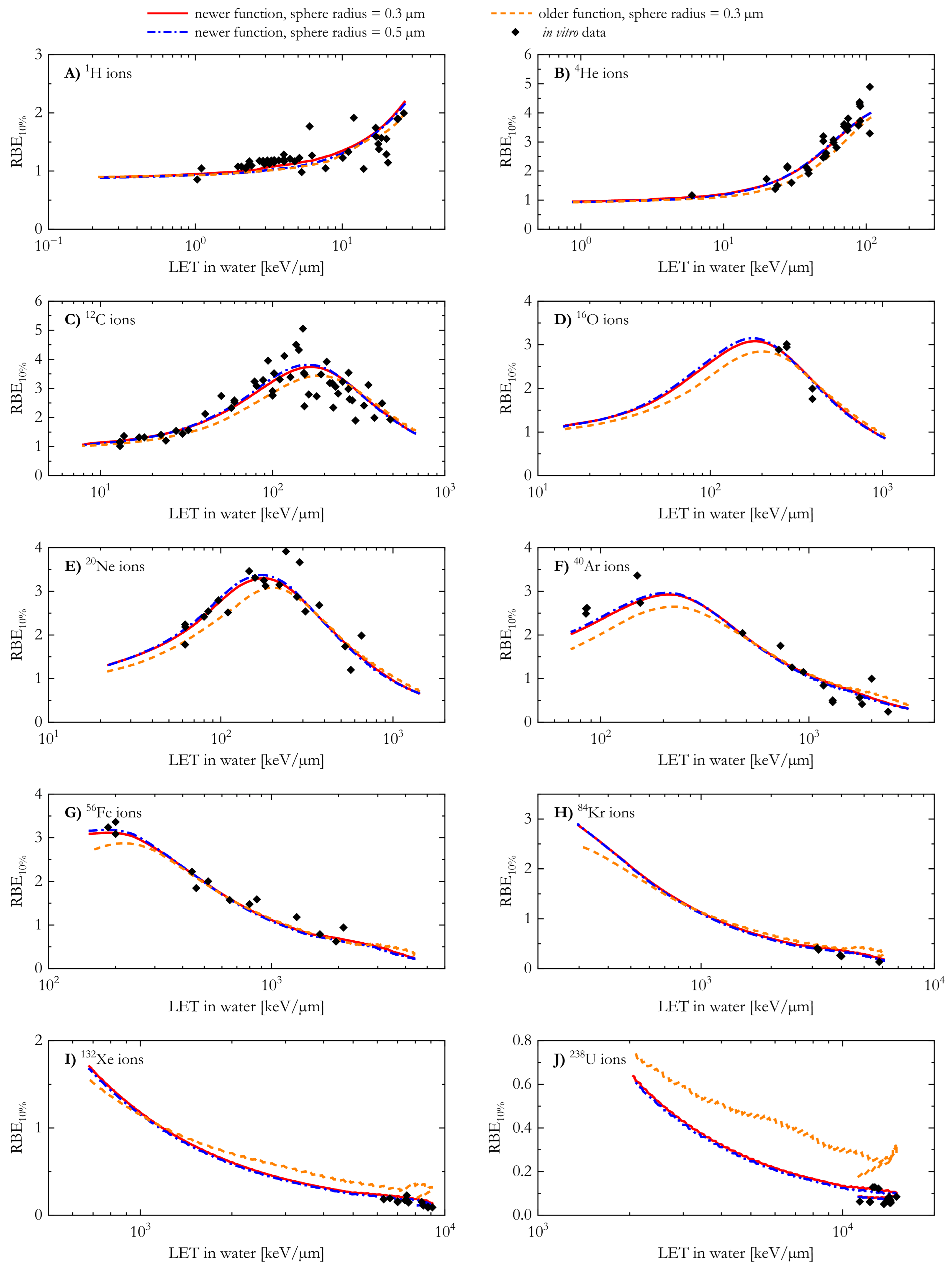
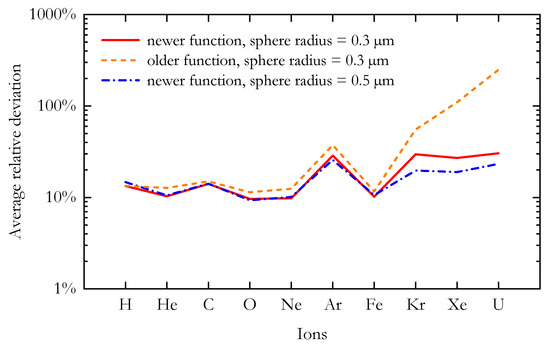
Figure 6.
The average relative deviation between the RBE10% calculated with the MCF MKM and the corresponding in vitro results for the ten ions included in this study. The microdosimetric distributions needed for the RBE calculations were simulated using the older and the newer analytical microdosimetric functions implemented in PHITS.
Figure 6.
The average relative deviation between the RBE10% calculated with the MCF MKM and the corresponding in vitro results for the ten ions included in this study. The microdosimetric distributions needed for the RBE calculations were simulated using the older and the newer analytical microdosimetric functions implemented in PHITS.
4. Conclusions
A comprehensive comparison of two analytical microdosimetric functions (AMFs) [25,41] was carried out in 4000 different scenarios (400 energy points, 10 ions from 1H to 238U) for water spherical targets with radii of 0.3 µm (the reference scale for RBE calculations) and 0.5 µm (the most common target size used in the literature). The two AMFs were compared with regard to the shape of the simulated probability distributions of the lineal energy, their mean values, and the RBE10% computed in combination with the MCF MKM for the most used mammalian cell line (Chinese hamster lung fibroblasts, V79 cell line). Published data from in vitro RBE experiments and track structure simulations were used to benchmark our results.
The results of this work indicate that the newer AMF offers superior performance in calculating microdosimetric quantities and RBE modeling, especially in the case of very heavy ions.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/qubs8030018/s1.
Author Contributions
Software: A.P. and T.S.; formal analysis: A.P.; visualization: A.P.; writing—original draft: A.P.; writing—review and editing: A.P., K.M.F., T.S. and C.J.B.; funding acquisition: K.M.F. and C.J.B. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The data presented in this study are available on request from the corresponding author.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Scholz, M. Effects of Ion Radiation on Cells and Tissues. In Radiation Effects on Polymers for Biological Use; Springer: Berlin/Heidelberg, Germany, 2003; Volume 162, pp. 95–155. [Google Scholar]
- Friedrich, T.; Scholz, U.; Elsasser, T.; Durante, M.; Scholz, M. Systematic analysis of RBE and related quantities using a database of cell survival experiments with ion beam irradiation. J. Radiat. Res. 2013, 54, 494–514. [Google Scholar] [CrossRef] [PubMed]
- ICRU. ICRU Report 85: Fundamental Quantities and Units for Ionizing Radiation; Technical Report for International Commission on Radiation Units & Measurements: Bethesda, MD, USA, 2011. [Google Scholar]
- Furusawa, Y.; Fukutsu, K.; Aoki, M.; Itsukaichi, H.; Eguchi-Kasai, K.; Ohara, H.; Yatagai, F.; Kanai, T.; Ando, K. Inactivation of Aerobic and Hypoxic Cells from Three Different Cell Lines by Accelerated3He-,12C- and20Ne-Ion Beams. Radiat. Res. 2000, 154, 485–496. [Google Scholar] [CrossRef] [PubMed]
- Friedrich, T.; Durante, M.; Scholz, M. Particle species dependence of cell survival RBE: Evident and not negligible. Acta Oncol. 2013, 52, 589–603. [Google Scholar] [CrossRef] [PubMed]
- ICRU. ICRU Report 36: Microdosimetry; Technical Report for International Commission on Radiation Units & Measurements: Bethesda, MD, USA, 1983. [Google Scholar]
- Rossi, H.H.; Zaider, M. Microdosimetry and Its Applications; Springer: Berlin/Heidelberg, Germany, 1996. [Google Scholar] [CrossRef]
- ICRU. ICRU Report 98: Stochastic Nature of Radiation Interactions: Microdosimetry. J. ICRU 2023, 23, 1–168. [Google Scholar] [CrossRef]
- Olko, P.; Booz, J. Energy deposition by protons and alpha particles in spherical sites of nanometer to micrometer diameter. Radiat. Environ. Biophys. 1990, 29, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Parisi, A.; Olko, P.; Swakon, J.; Horwacik, T.; Jablonski, H.; Malinowski, L.; Nowak, T.; Struelens, L.; Vanhavere, F. Microdosimetric characterization of a clinical proton therapy beam: Comparison between simulated lineal energy distributions in spherical water targets and experimental measurements with a silicon detector. Phys. Med. Biol. 2022, 67, 015006. [Google Scholar] [CrossRef]
- IAEA. TRS 461: Relative Biological Effectiveness in Ion Beam Therapy; Technical Report Series; IAEA: Vienna, Austria, 2008. [Google Scholar]
- Kanai, T.; Endo, M.; Minohara, S.; Miyahara, N.; Koyama-ito, H.; Tomura, H.; Matsufuji, N.; Futami, Y.; Fukumura, A.; Hiraoka, T.; et al. Biophysical characteristics of HIMAC clinical irradiation system for heavy-ion radiation therapy. Int. J. Radiat. Oncol. Biol. Phys. 1999, 44, 201–210. [Google Scholar] [CrossRef] [PubMed]
- Kramer, M.; Scholz, M. Rapid calculation of biological effects in ion radiotherapy. Phys. Med. Biol. 2006, 51, 1959–1970. [Google Scholar] [CrossRef] [PubMed]
- Inaniwa, T.; Kanematsu, N.; Matsufuji, N.; Kanai, T.; Shirai, T.; Noda, K.; Tsuji, H.; Kamada, T.; Tsujii, H. Reformulation of a clinical-dose system for carbon-ion radiotherapy treatment planning at the National Institute of Radiological Sciences, Japan. Phys. Med. Biol. 2015, 60, 3271–3286. [Google Scholar] [CrossRef] [PubMed]
- ICRP. ICRP, 123: Assessment of radiation exposure of astronauts in space. ICRP Publication 123. Ann. ICRP 2013, 42, 1–339. [Google Scholar] [CrossRef] [PubMed]
- Bellinzona, V.E.; Cordoni, F.; Missiaggia, M.; Tommasino, F.; Scifoni, E.; La Tessa, C.; Attili, A. Linking Microdosimetric Measurements to Biological Effectiveness in Ion Beam Therapy: A Review of Theoretical Aspects of MKM and Other Models. Front. Phys. 2021, 8, 578492. [Google Scholar] [CrossRef]
- Parisi, A.; Furutani, K.M.; Beltran, C.J. On the calculation of the relative biological effectiveness of ion radiation therapy using a biological weighting function, the microdosimetric kinetic model (MKM) and subsequent corrections (non-Poisson MKM and modified MKM). Phys. Med. Biol. 2022, 67, 095014. [Google Scholar] [CrossRef] [PubMed]
- Friedland, W.; Dingfelder, M.; Kundrat, P.; Jacob, P. Track structures, DNA targets and radiation effects in the biophysical Monte Carlo simulation code PARTRAC. Mutat. Res. 2011, 711, 28–40. [Google Scholar] [CrossRef] [PubMed]
- Nikjoo, H.; Emfietzoglou, D.; Liamsuwan, T.; Taleei, R.; Liljequist, D.; Uehara, S. Radiation track, DNA damage and response-a review. Rep. Prog. Phys. 2016, 79, 116601. [Google Scholar] [CrossRef] [PubMed]
- Incerti, S.; Kyriakou, I.; Bernal, M.A.; Bordage, M.C.; Francis, Z.; Guatelli, S.; Ivanchenko, V.; Karamitros, M.; Lampe, N.; Lee, S.B.; et al. Geant4-DNA example applications for track structure simulations in liquid water: A report from the Geant4-DNA Project. Med. Phys. 2018, 45, e722–e739. [Google Scholar] [CrossRef] [PubMed]
- Schuemann, J.; McNamara, A.L.; Ramos-Mendez, J.; Perl, J.; Held, K.D.; Paganetti, H.; Incerti, S.; Faddegon, B. TOPAS-nBio: An Extension to the TOPAS Simulation Toolkit for Cellular and Sub-cellular Radiobiology. Radiat. Res. 2019, 191, 125–138. [Google Scholar] [CrossRef] [PubMed]
- Plante, I.; Poignant, F.; Slaba, T. Track Structure Components: Characterizing Energy Deposited in Spherical Cells from Direct and Peripheral HZE Ion Hits. Life 2021, 11, 1112. [Google Scholar] [CrossRef]
- Ogawa, T.; Hirata, Y.; Matsuya, Y.; Kai, T. Development and validation of proton track-structure model applicable to arbitrary materials. Sci. Rep. 2021, 11, 24401. [Google Scholar] [CrossRef] [PubMed]
- Sato, T.; Iwamoto, Y.; Hashimoto, S.; Ogawa, T.; Furuta, T.; Abe, S.-I.; Kai, T.; Matsuya, Y.; Matsuda, N.; Hirata, Y.; et al. Recent improvements of the particle and heavy ion transport code system—PHITS version 3.33. J. Nucl. Sci. Technol. 2023, 61, 127–135. [Google Scholar] [CrossRef]
- Sato, T.; Watanabe, R.; Niita, K. Development of a calculation method for estimating specific energy distribution in complex radiation fields. Radiat. Prot. Dosim. 2006, 122, 41–45. [Google Scholar] [CrossRef] [PubMed]
- Sato, T.; Watanabe, R.; Sihver, L.; Niita, K. Applications of the microdosimetric function implemented in the macroscopic particle transport simulation code PHITS. Int. J. Radiat. Biol. 2012, 88, 143–150. [Google Scholar] [CrossRef] [PubMed]
- Parisi, A.; Van Hoey, O.; Mégret, P.; Vanhavere, F. Microdosimetric specific energy probability distribution in nanometric targets and its correlation with the efficiency of thermoluminescent detectors exposed to charged particles. Radiat. Meas. 2019, 123, 1–12. [Google Scholar] [CrossRef]
- Parisi, A.; Dabin, J.; Schoonjans, W.; Van Hoey, O.; Mégret, P.; Vanhavere, F. Photon energy response of LiF:Mg,Ti (MTS) and LiF:Mg,Cu,P (MCP) thermoluminescent detectors: Experimental measurements and microdosimetric modeling. Radiat. Phys. Chem. 2019, 163, 67–73. [Google Scholar] [CrossRef]
- Parisi, A.; Struelens, L.; Vanhavere, F. Nanoscale calculation of the relative efficiency of 7LiF: Mg, Ti (MTS-7) and 7LiF: Mg, Cu, P (MCP-7) thermoluminescent detectors for measuring electrons and positrons. J. Phys. Conf. Ser. 2020, 1662, 012025. [Google Scholar] [CrossRef]
- Parisi, A.; Olko, P.; Swakon, J.; Horwacik, T.; Jablonski, H.; Malinowski, L.; Nowak, T.; Struelens, L.; Vanhavere, F. Modeling the radiation-induced cell death in a therapeutic proton beam using thermoluminescent detectors and radiation transport simulations. Phys. Med. Biol. 2020, 65, 015008. [Google Scholar] [CrossRef] [PubMed]
- Parisi, A.; Sawakuchi, G.; Granville, D.; Yukihara, E.G. Microdosimetric modeling of the relative efficiency of Al2O3:C (Luxel, blue emission) optically stimulated luminescent detectors exposed to ions from 1H to 132Xe. Radiat. Meas. 2022, 150, 106678. [Google Scholar] [CrossRef]
- Hirata, Y.; Sato, T.; Watanabe, K.; Ogawa, T.; Parisi, A.; Uritani, A. Theoretical and experimental estimation of the relative optically stimulated luminescence efficiency of an optical-fiber-based BaFBr:Eu detector for swift ions. J. Nucl. Sci. Technol. 2022, 59, 915–924. [Google Scholar] [CrossRef]
- Sato, T.; Watanabe, R.; Kase, Y.; Tsuruoka, C.; Suzuki, M.; Furusawa, Y.; Niita, K. Analysis of cell-survival fractions for heavy-ion irradiations based on microdosimetric kinetic model implemented in the particle and heavy ion transport code system. Radiat. Prot. Dosim. 2011, 143, 491–496. [Google Scholar] [CrossRef] [PubMed]
- Sato, T.; Furusawa, Y. Cell survival fraction estimation based on the probability densities of domain and cell nucleus specific energies using improved microdosimetric kinetic models. Radiat. Res. 2012, 178, 341–356. [Google Scholar] [CrossRef] [PubMed]
- Matsuya, Y.; Sasaki, K.; Yoshii, Y.; Okuyama, G.; Date, H. Integrated Modelling of Cell Responses after Irradiation for DNA-Targeted Effects and Non-Targeted Effects. Sci. Rep. 2018, 8, 4849. [Google Scholar] [CrossRef]
- Matsuya, Y.; McMahon, S.J.; Ghita, M.; Yoshii, Y.; Sato, T.; Date, H.; Prise, K.M. Intensity Modulated Radiation Fields Induce Protective Effects and Reduce Importance of Dose-Rate Effects. Sci. Rep. 2019, 9, 9483. [Google Scholar] [CrossRef] [PubMed]
- Parisi, A.; Sato, T.; Matsuya, Y.; Kase, Y.; Magrin, G.; Verona, C.; Tran, L.; Rosenfeld, A.; Bianchi, A.; Olko, P.; et al. Development of a new microdosimetric biological weighting function for the RBE(10) assessment in case of the V79 cell line exposed to ions from (1)H to (238)U. Phys. Med. Biol. 2020, 65, 235010. [Google Scholar] [CrossRef] [PubMed]
- Parisi, A.; Beltran, C.J.; Furutani, K.M. The Mayo Clinic Florida microdosimetric kinetic model of clonogenic survival: Formalism and first benchmark against in vitro and in silico data. Phys. Med. Biol. 2022, 67, 185013. [Google Scholar] [CrossRef] [PubMed]
- Parisi, A.; Beltran, C.J.; Furutani, K.M. The Mayo Clinic Florida Microdosimetric Kinetic Model of Clonogenic Survival: Application to Various Repair-Competent Rodent and Human Cell Lines. Int. J. Mol. Sci. 2022, 23, 2491. [Google Scholar] [CrossRef] [PubMed]
- Sato, T.; Matsuya, Y.; Hamada, N. Microdosimetric Modeling of Relative Biological Effectiveness for Skin Reactions: Possible Linkage Between In Vitro and In Vivo Data. Int. J. Radiat. Oncol. Biol. Phys. 2022, 114, 153–162. [Google Scholar] [CrossRef] [PubMed]
- Sato, T.; Matsuya, Y.; Ogawa, T.; Kai, T.; Hirata, Y.; Tsuda, S.; Parisi, A. Improvement of the hybrid approach between Monte Carlo simulation and analytical function for calculating microdosimetric probability densities in macroscopic matter. Phys. Med. Biol. 2023, 68, 155005. [Google Scholar] [CrossRef]
- Fowler, J.F. The linear-quadratic formula and progress in fractionated radiotherapy. Br. J. Radiol. 1989, 62, 679–694. [Google Scholar] [CrossRef]
- McMahon, S.J. The linear quadratic model: Usage, interpretation and challenges. Phys. Med. Biol. 2018, 64, 01TR01. [Google Scholar] [CrossRef] [PubMed]
- Friedrich, T.; Pfuhl, T.; Scholz, M. Update of the particle irradiation data ensemble (PIDE) for cell survival. J. Radiat. Res. 2021, 62, 645–655. [Google Scholar] [CrossRef] [PubMed]
- Grun, R.; Friedrich, T.; Traneus, E.; Scholz, M. Is the dose-averaged LET a reliable predictor for the relative biological effectiveness? Med. Phys. 2019, 46, 1064–1074. [Google Scholar] [CrossRef] [PubMed]
- Parisi, A.; Beltran, C.J.; Furutani, K.M. Variable RBE in proton radiotherapy: A comparative study with the predictive Mayo Clinic Florida microdosimetric kinetic model and phenomenological models of cell survival. Phys. Med. Biol. 2023, 68, 185020. [Google Scholar] [CrossRef] [PubMed]
- ICRU. ICRU Report 90: Key Data For Ionizing-Radiation Dosimetry; Technical Report for International Commission on Radiation Units & Measurements: Bethesda, MD, USA, 2016. [Google Scholar]
- Ziegler, J.F.; Ziegler, M.D.; Biersack, J.P. SRIM—The stopping and range of ions in matter (2010). Nucl. Instrum. Methods Phys. Res. Sect. B Beam Interact. Mater. At. 2010, 268, 1818–1823. [Google Scholar] [CrossRef]
- Hawkins, R.B. A statistical theory of cell killing by radiation of varying linear energy transfer. Radiat. Res. 1994, 140, 366–374. [Google Scholar] [CrossRef] [PubMed]
- Hawkins, R.B. A microdosimetric-kinetic theory of the dependence of the RBE for cell death on LET. Med. Phys. 1998, 25, 1157–1170. [Google Scholar] [CrossRef] [PubMed]
- Hawkins, R.B. A microdosimetric-kinetic model for the effect of non-Poisson distribution of lethal lesions on the variation of RBE with LET. Radiat. Res. 2003, 160, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Kellerer, A.M.; Rossi, H.H. The theory of dual radiation action. Curr. Top. Radiat. Res. Quart. 1972, 8, 85–158. [Google Scholar]
- Yokota, H.; Vandenengh, G.; Hearst, J.E.; Sachs, R.K.; Trask, B.J. Evidence for the Organization of Chromatin in Megabase Pair-Sized Loops Arranged Along a Random-Walk Path in the Human G0/G1 Interphase Nucleus. J. Cell Biol. 1995, 130, 1239–1249. [Google Scholar] [CrossRef] [PubMed]
- Rogakou, E.P.; Pilch, D.R.; Orr, A.H.; Ivanova, V.S.; Bonner, W.M. DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139. J. Biol. Chem. 1998, 273, 5858–5868. [Google Scholar] [CrossRef] [PubMed]
- Friedrich, T.; Durante, M.; Scholz, M. Modeling cell survival after photon irradiation based on double-strand break clustering in megabase pair chromatin loops. Radiat. Res. 2012, 178, 385–394. [Google Scholar] [CrossRef]
- Sigmund, P.; Schinner, A.; Paul, H. Errata and Addenda for ICRU Report 73. J. ICRU 2009, 5, 1–10. [Google Scholar]
- Parisi, A.; Struelens, L.; Vanhavere, F. Comparison between the results of a recently-developed biological weighting function (V79-RBE(10)BWF) and thein vitroclonogenic survival RBE(10) of other repair-competent asynchronized normoxic mammalian cell lines and ions not used for the development of the model. Phys. Med. Biol. 2021, 66, 235006. [Google Scholar] [CrossRef]
- Chatterjee, A.; Schaefer, H.J. Microdosimetric structure of heavy ion tracks in tissue. Radiat. Environ. Biophys. 1976, 13, 215–227. [Google Scholar] [CrossRef] [PubMed]
- Butts, J.J.; Katz, R. Theory of RBE for heavy ion bombardment of dry enzymes and viruses. Radiat. Res. 1967, 30, 855–871. [Google Scholar] [CrossRef] [PubMed]
- Vassiliev, O.N.; Peterson, C.B.; Cao, W.; Grosshans, D.R.; Mohan, R. Systematic microdosimetric data for protons of therapeutic energies calculated with Geant4-DNA. Phys. Med. Biol. 2019, 64, 215018. [Google Scholar] [CrossRef] [PubMed]
- ICRU. ICRU Report 40: The Quality Factor in Radiation Protection; Technical Report for International Commission on Radiation Units & Measurements: Bethesda, MD, USA, 1986. [Google Scholar]
- Magrin, G. A method to convert spectra from slab microdosimeters in therapeutic ion-beams to the spectra referring to microdosimeters of different shapes and material. Phys. Med. Biol. 2018, 63, 215021. [Google Scholar] [CrossRef]
- Wulf, H.; Kraft-Weyrather, W.; Miltenburger, H.G.; Blakely, E.A.; Tobias, C.A.; Kraft, G. Heavy-ion effects on mammalian cells: Inactivation measurements with different cell lines. Radiat. Res. Suppl. 1985, 8, S122–S134. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
