Abstract
Function is a key central concept to the practice of biomimicry. Many published models of the biomimicry process include steps to identify, understand, and translate function of biological systems. Examples include functional modeling, decomposition, or abstraction with tools specifically designed to facilitate such steps. A functional approach to biomimicry yields a semantic bridge between biology and engineering, enabling practitioners from a variety of backgrounds to more easily communicate and collaborate in a biomimicry design process. Although analysis of function is likely a necessary part of biomimicry design, recent work suggests it is not sufficient without a more systematic understanding of the complex biological context in which a function exists (e.g., scale and trade-offs). Consequently, emerging tools such as ontologies are being developed that attempt to capture the intricacies of biological systems (including functions), such as their complex environmental and behavioral interactions. However, due to the complexity of such tools, they may be under-utilized. Here, we propose a solution through a computer-aided user interface tool which integrates a biomimetic ontology with a thesaurus-based functional approach to biomimicry. Through a proof of concept illustrative case study, we demonstrate how merging existing tools can facilitate the biomimicry process in a systematic and collaborative way, broadening solution discovery. This work offers an approach to making existing tools, specifically the BioMimetic Ontology, more accessible and encompassing of different perspectives via semantic translation and interface design. This provides the user with the opportunity to interface and extract information from both the Engineering-to-Biology Thesaurus and the BioMimetic Ontology in a way that was not possible before. The proposed E2BMO tool not only increases the accessibility of the BioMimetic Ontology, which ultimately aims to streamline engineers’ interaction with the bio-inspired design process, but also provides an option for practitioners to traverse biological knowledge along the way, encouraging greater interdisciplinary collaboration and consideration when conducting biomimicry research.
1. Introduction
Biomimicry and biomimetics used interchangeably in this work, can be defined as innovation inspired by the emulation of biological forms, processes, patterns, and systems [1]. Although an emerging field, biomimicry has led to innovations across many disciplines including materials science, chemistry, engineering, waste management, information technology, and organizational design [2,3,4,5,6]. In a recent publication, the Fermanian Business & Economic Institute projected that biomimicry as an innovation process is to account for $1.6 trillion global output by 2030 [7]. However, despite the innovative potential of biomimicry, challenges still exist with practice. One such challenge is the interdisciplinary nature of biomimicry; it encourages practitioners (designers, engineers, biologists, etc.) to engage in a collaborative process that transcends disciplinary boundaries, while also requiring knowledge transfer and communication across such disciplines. To add to this challenge, the tools created to facilitate biomimicry practice tend to be defined by one field of knowledge, primarily that of the tool developer [8]. For example, engineers tend to take a functional approach to problem decomposition while biologists more commonly tend to decompose the complex interactions within biological systems. To date, there are copious tools available to facilitate biomimicry practice, many supporting an engineering type functional approach, and recently, a surge of more complex systematic and computational tools that capture the intricacies of biological systems. As tool development often reflects the discipline of its creator, it is required that most practitioners, within the interdisciplinary field of biomimicry, learn a new nomenclature specific to that tool. This is a hurdle to the adoption of biomimicry and the advancement of the field as a whole, especially within an industrial context, as practitioners within a corporate innovation setting rarely have the interest or time to engage at this level, thus requiring the constant assistance of an outside consultant to use the tools. An intuitive user interface may be half the challenge.
In general, the practice of biomimicry can be approached from two perspectives: (1) solution-driven, where the biological model of inspiration is the driving factor for innovation or (2) problem-driven, where the technical challenge of interest is the driving factor [9,10,11]. As the problem-driven process aims to solve a technical challenge, it is more often utilized within industry or engineering design and is thus the approach of interest in this paper (Figure 1).
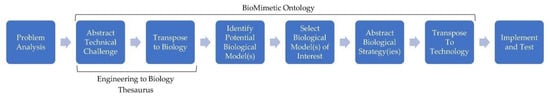
Figure 1.
Illustration of the traditional problem-driven biomimicry design process and steps that selected tools facilitate. E2B Thesaurus (Step 2 and 3) and BioMimetic Ontology (Step 2 to 7) [12].
The functional approach has long been a key central concept to the practice of biomimicry and has been identified as a solution to overcome the difficulties of its interdisciplinary nature. A recent review of biomimetic tools and processes identified several models of the problem-driven approach and correlated them with the general problem-solving process of Massay & Wallace 1956 [13], commonly used in engineering and design [12]. Of the 12 biomimetic processes identified, half clearly state a functional step referred to as either function identification and definition [14,15,16], functional keyword searches [17,18] or functional modeling [19]. Function offers a semantic bridge between disciplines, specifically biology and engineering, enabling practitioners from biological and other backgrounds to more easily communicate and collaborate in a biomimicry design process. This semantic bridge assists in reframing biological terminology in an engineering context or vice versa, reframing a technical challenge in terms of function that can better be identified in biological text. This increases the accessibility of biological information to practitioners with varying biological knowledge, but also allows for more efficient communication with a biologist, which we believe is a necessary endeavor to understand the complexity of biological systems. A recent review [8] identified 43 tools that facilitate the biomimicry process, including literature navigation tools, natural language tools, and interdisciplinary semantic translation tools that have been developed to facilitate this functional semantic bridge to biomimicry practice.
Recent work has suggested that functional analysis alone is not sufficient for biomimicry practice without a more systemic understanding of the complex biological context (e.g., scale and trade-offs) in which a function exists [20]. The concept of function as a cross-domain semantic bridge is deemed effective only if the definition of that particular function is unified across both domains, which with biology and engineering is rarely the case. Functional decomposition of a technical system is a logical approach from an engineering perspective as each component of that system has a specific role. In contrast, biological systems always have multi-functional components that contribute to several roles that can differ across scales and context. Take, for example, the adhesive properties of gecko toe pads. Gecko toe pads are composed of arrays of hierarchical fibers that make close contact with a surface such that enough van der Waals intermolecular forces can be generated to support the gecko’s mass [21,22,23,24,25]. The gecko adhesive system is incredibly multifunctional with properties including reusability, ease of release, underwater adhesion, and self-cleaning [21,26,27,28]. Materials scientists and biologists alike have been researching this system with a goal of designing and fabricating gecko-like synthetic adhesives that function in a variety of contexts. Even so, few synthetic adhesives have been able to fully replicate the multifunctional aspects of the gecko adhesive system and this may be a by-product of not fully understanding how the integrated system operates, as a whole, under a variety of natural conditions [27]. Thus, during functional analysis, it is necessary to consider the particular function of interest within a more systematic context to ensure neither misinterpretation or oversimplification of the abstracted biological principle. This is particularly important during the search for relevant biological information within biological texts, the most comprehensive and effective source of information [29], as such texts rarely define a function in a way that is recognizable to an engineer, making the search for relevant biological information difficult and time consuming for the practitioner. Consequently, tools such as ontologies are being developed that attempt to capture the intricacies of biological systems to enable practitioners, especially those outside of a biological discipline, to facilitate deeper understanding of the biological system.
Nevertheless, modeling such complex biological systems is not without its challenges. Practitioners, not familiar with complex system mapping models and software, often do not understand the objective or complexity of such tools and struggle interacting with them. The effort required to learn and become effective in the use of such tools, and most tools that support the bio-inspired design process, can be time consuming and intimidating, often leaving these tools and approaches underutilized [30]. Our goal is not to suggest that ‘function’ is not an effective approach to biomimicry practice or to deter practitioners from exploring complex systematic tools. We would simply like to draw attention to the limitations and consequences of each approach. Furthermore, it has been shown that an approach that elaborates on function to include complex environmental and behavioral interactions increases the precision of retrieved results compared to analysis through function alone [29]. This integration of approaches to biomimicry practice could be facilitated through an integration of tools which would aid in overcoming another hurdle to biomimicry practice, a fluid advancement through the process. Currently, the difficult task of advancing through the biomimicry process step-by-step is left to the practitioner to navigate, a challenge that, if overcome, could aid in the advancement of biomimicry practice.
The aim of this paper is to draw attention to the copious tools available that facilitate the biomimicry process and to demonstrate how merging existing tools through a user interface design could potentially facilitate practitioners’ navigation through the process in a more systematic and collaborative way, broadening solution discovery. Through this paper, we propose a computer-aided user interface that elaborates on a thesaurus-based functional approach to biomimicry to include complex environmental and behavioral interactions through integration with the BioMimetic Ontology (BMO) [31]. The focus of this work is to provide a portal for users to access information from within the BMO; we do not aim to make this information more interpretable to the user, we simply provide the opportunity to access it. For practitioners, this integration of tools captures the beneficial factors of both approaches to biomimicry practice, and the user interface creates an access point to important biological information for practitioners not necessarily familiar with biological research. This is not to say that this is the only approach to bio inspired design, but it is an approach that the authors deem to be of value, specifically to practitioners who wish to interact with the BMO. Thus, the integration of tools may assist in overcoming significant hurdles to the design process which could aid in the expanded application of biomimicry.
2. Materials and Methods
2.1. Selection of Tools
This paper is the result of a collaborative course that took place at the University of Akron and was composed of Integrated Bioscience Ph.D. students with diverse backgrounds including computer science, biology, physics, design, engineering, and architecture (four class members were Biomimicry Fellows, students actively working to integrate biomimicry practice into industry). Initially, we reviewed the biomimicry process and tools that support each step and identified a number of key points: (1) there are many tools available within the field of biomimicry, (2) these tools rarely cover more than one or two steps of the process, (3) the transition through the process or between tools is not facilitated, and (4) the more complex systematic tools, although useful, are too complex and largely inaccessible to potential users. Thus, two tools were selected that the team agreed effectively facilitated several steps, but if integrated, could further streamline the biomimicry process (Figure 1). Upon interviewing innovation consultants actively practicing biomimicry within an industrial context, the Engineering-to-Biology (E2B) Thesaurus [19] was selected, as it fulfilled our specific criteria for usability, accessibility, and cross-domain functionality. Our second tool of choice, the BioMimetic Ontology (BMO) [31], was selected as ontologies, due to their ability to organize complex collections of data, have proven to be successful across a variety of fields, including bioinformatics and medicine [32]. Therefore, an ontology that captures the intricacies of natural systems became the obvious tool of choice. The integration of such tools could bridge the gap between steps, therefore creating a more systematic process with direct access to identified and vetted biological strategies, opening up greater potential for biomimicry practice.
2.1.1. Engineering-to-Biology Thesaurus (E2B Thesaurus)
The E2B Thesaurus is a bio-inspired design tool that hierarchically maps biological terms to engineering terms of the Functional Basis (qualitative modeling lexicon) [19]. The Functional Basis is a standardized set of function-related terminology used to represent a formal function needed to support functional modeling, a process aimed at easing communication, modeling, and computation difficulties within design [33]. The E2B Thesaurus has two sets of terms that were derived from the Functional Basis: flows and function. The flow set is categorized into the flows of material, energy, and signal, and the function set is categorized into eight classes: branch, channel, connect, control magnitude, convert, provision, signal, and support. These terms are further categorized into secondary and tertiary engineering-related terminology, then mapped to biological flow and function correspondents (Figure 2). The E2B Thesaurus was developed to encourage collaboration between biologists and engineers for discovery and creation of biologically-inspired engineering solutions. It allows one to make associations between engineering and biology, thus strengthening the designer’s ability to utilize biological information for solving technical challenges.
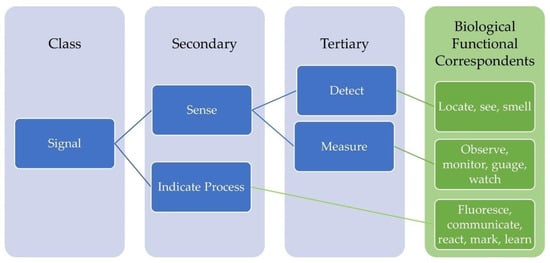
Figure 2.
Abstracts from the Engineering-to-Biology Thesaurus [18] representing both the biological Function and Flow correspondents. The hierarchical layout of the tool begins with classes derived from the Functional Basis, which is further categorized into secondary and tertiary terms, then correlated with biological correspondents.
Therefore, the E2B Thesaurus facilitates Step 2 and 3 of the biomimicry process (Figure 1). During Step 2, ‘Abstract Technical Challenge’, users are encouraged to define their technical challenge as a function to be attained and select such functions from the engineering-related functional terms of the E2B Thesaurus. During Step 3, ‘Transpose to Biology’, the user utilizes the term identified in Step 2 and is directed to several biological functional terms that directly correspond to the engineering-related term. This enables the user to better communicate with a biologist or access biological information that eases the transition to Step 4 ‘Identify Potential Biological Model(s)’. For instance, perhaps a team of engineers from an industrial R&D team would like to design a sorting mechanism to identify ripeness of fruit, for example bananas or plums, through sensor-activated image recognition while being transported on a conveyor belt. If they are interested in finding biological models to inspire this product, they may choose the engineering term ‘signal’ from the thesaurus (Figure 2), as to ideate around a signaling mechanism between the sensor and the conveyor belt. ‘Signal’ could further be characterized by the secondary and tertiary engineering related functional terms that are most relevant to their application, either ‘sense’ related to ‘detect’ and ‘measure’ or ‘indicate process’. After translating ‘signal’ from Engineering-to-Biology via the E2B thesaurus, they would find that ‘signal’ can refer to biological functions such as ‘locate’, ‘observe’, ‘fluoresce’, ‘learn’, etc. The team, using these terms, could then peruse scientific literature, other biomimetic tools, or collaborate with a biologist to locate potential biological models for their particular application. The necessity of collaborating with a biologist is important as to ensure neither a misinterpretation nor oversimplification of the biological strategy identified and provide an accurate transposition of this strategy back to technology, Step 7 of the biomimicry process (Figure 1).
2.1.2. BioMimetic Ontology (BMO)
An ontology is an assemblage of data in which information is organized hierarchically and relationships made specific. In this way, information becomes knowledge. The BioMimetic Ontology (BMO) aims to bridge the gap between engineering and biology by using trade-offs, making biological strategies more accessible to engineering practitioners [31]. TRIZ (acronym for ‘Teorija Reshenija Izobretateliskih Zadatch’, which translates to ‘Theory of Solving Problems Inventively’), developed in the second half of the last century by Genrich Altshuller [34], is a method for solving technical challenges. After the review of several thousand patents, Altshuller created a list of 39 engineering parameters that were characteristic factors that could affect the performance of an engineering machine, material or structure. These 39 engineering parameters were used to create the ‘contradiction matrix’, trading off one against the other. A user defines the problem being investigated by selecting two engineering parameters. For example, the problem of a bridge needing to take more traffic where it is not possible to increase the size of the foundation could be defined as a conflict between engineering parameters ‘resisting greater force’ and ‘changing supporting shape’. Defining a problem in a neutral and objective manner generalizes the problem so that solutions to problems, that can be defined in the same or similar way, can offer insight. This list of potential solutions or design insights are known as the inventive principles (IPs) and are numbered 1 to 40 [31]. In short, through the ‘contradiction matrix’ of TRIZ, conflicts help to break down problems and find innovative solutions, called the inventive principles (IPs). Ontologies can also be merged to further expand their content making them even more attractive for capturing complex natural systems. Merging is facilitated by a basic framework such as that of the basic formal ontology (BFO). The BioMimetic Ontology (BMO) follows the BFO framework, thus can merge with parts of other ontologies, increasing its range and usefulness.
One of the authors (J.F.V.V.), due to his specialization in biology and understanding of both biology and engineering, has translated a number of biological strategies from scientific papers, expressing them as trade-offs and IPs, and integrating them into the BMO. This is a difficult task as IPs, due to their engineering foundations, can be somewhat limited in capturing the intricate complexities of natural systems, hence this task of translation was undertaken by J.F.V.V., a subject matter expert. Also due to the vast quantity of biological information that is currently available and continues to be discovered, the BMO is not an exhaustive database of biological strategies and a process to automate this work to more efficiently capture these strategies has become a very active area of research, although with limited success to date. For more information, refer to Section 4. While implicit in TRIZ [31], trade-offs become the core concept of the BMO. BMO selects which TRIZ IP’s recommend possible solutions or insights to a specific trade-off, based on biological literature describing how living role models resolved such trade-off. Therefore, within the BMO, trade-offs are the factors whose resolution the biological organism is addressing; IPs interpret the biological strategies the relevant organism used to resolve a trade-off. Therefore, the BMO facilitates the biomimicry process (Figure 1) across several steps. Step 2, ‘Abstract Technical Challenge’, is facilitated through the selection of a trade-off. Step 3, ‘Transpose to Biology’, is bypassed as the need to interact with biological information directly is negated through the use of trade-offs. Step 4 and 6 ‘Identify Potential Biological Model(s)’ and ‘Abstract Biological Strategy(ies)’, respectively, are automated internally through the BMO. Step 5, ‘Select Biological Model of Interest’, is completely removed as the need to understand the complexity of each biological model has been completed through the work of one of the authors (J.F.V.V.). Step 7, ‘Transpose to Technology’, is also removed, as the problem was never transposed to biology to begin with. Overall, this accelerates the design process as the user begins by defining their technical challenge as a trade-off and then, through the BMO, are given direct access to potential solutions of biological strategies already transposed as IPs. The success of this approach depends on the universality of trade-offs.
Although very useful tools, ontologies in general have been considered difficult to navigate through the current available platforms, and the complex hierarchical relationships difficult to comprehend. This may be a potential limitation on the widespread use of the BMO as practitioners find it difficult to integrate it into the design process. To illustrate the complexity of this tool, Figure 3 illustrates the BMO within Protégé, a free open-source ontology editor, which is the form in which the BMO is currently available to practitioners [35]. The current available version of the BMO is a research tool with finite data that can be greatly shortened without losing its essential functionality, which other versions of the BMO may adopt. The example below is a continuation of the R&D challenge to design a sorting mechanism to identify ripeness of fruit through sensor activated image recognition. In the case given, a compromise needs to be made between the speed of the conveyor belt and the accuracy of the images captured by the sensors. The technical challenge abstracted in the form of a trade-off would be speed vs. accuracy. The results of querying the BMO for speed-accuracy trade-off through Protégé can be seen in Figure 3.
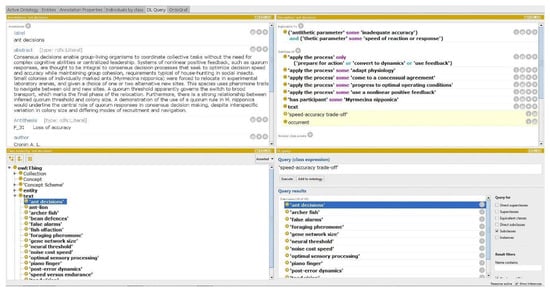
Figure 3.
Results of querying the BMO for speed/accuracy trade-off within Protégé. Bottom right image shows ‘DL query’ tab with ‘Query results’, a list of Biological Models as potential solutions. ‘Ant decision’ is selected, thus information is provided in the description tab which outlines IPs attached to that specific biology strategy. The top left tab, ‘Annotations’, provides an abstract of the biological text containing the specific biological strategy identified. Note: to ensure this is the information retrieved from the query the user must know how to retrieve these specific tabs.
2.2. Merging of Tools
2.2.1. Simplified Knowledge Organization System (SKOS)
To combine the E2B Thesaurus and the BMO, the E2B Thesaurus needed to be a format that could interact and communicate with the BMO. This was accomplished by expressing the E2B Thesaurus as a Simplified Knowledge Organization System (SKOS) [36], a method for organizing information from knowledge organization systems (e.g., thesauri, classification schemes, and taxonomies). All terms included in a SKOS are categorized into one of two categories: concepts or concept schemes. Concepts are the main elements of a SKOS; they are the specific terms from the original knowledge organization system (in this case, terms from the E2B Thesaurus). Associated concepts can then be organized into concept schemes. Once concepts and concept schemes are generated, various semantic relationships can be implemented between concepts to create a cohesive classification system [36,37]. SKOS was chosen as it allows terms to be loosely connected in an organized framework that could be inserted into an ontology directly. SKOS enables the selection of any chosen description system for such loose connections, giving the opportunity of introducing simpler or cross-disciplinary semantics that any user can quickly understand, despite the complexity of the ontology.
To begin, the E2B Thesaurus needed to be expressed as a SKOS. This was accomplished by first generating concept schemes using the ‘class’ engineering terms from the thesaurus (e.g., branch, channel, connect, etc.). Only the functional terms from the E2B Thesaurus were transferred to a SKOS (i.e., verbs or words that could be easily translated to verb form), as we believe that these terms are most likely to be utilized by practitioners during the biomimicry design process. Following the generation of concept schemes, the remainder of the functional terms from the thesaurus were integrated as concepts. To replicate the hierarchical relationships observed in the E2B Thesaurus, we connected engineering terms to biological function correspondents by adding the semantic relationships ‘has narrower’ and ‘has broader’ to the concepts. To illustrate this, consider the R&D design challenge example previously stated in Section 2.1.1. ‘Signal’ is considered a class term in the E2B Thesaurus and is connected to the tertiary engineering term ‘measure’ via the secondary term ‘sense’. ‘Measure’ is then related to the following biological function correspondents: ‘observe’, ‘monitor’, ‘gauge’, and ‘watch’. The following connections between concepts were made to capture these semantic relationships: (1) signal is a concept scheme and has top conceptsense, (2) senseis top concept in schemesignal and has narrower term measure, (3) measurehas broader term sense and has narrower terms observe, monitor, gauge, and watch. Here, thesaurus engineering terms are in red, thesaurus biological function correspondents are in blue, and SKOS vocabulary are in purple. This classification system enables the preservation of the hierarchical relationships in the thesaurus, so users could navigate from engineering to biological domains and back.
Once the E2B Thesaurus was expressed as a SKOS, biological function correspondents were connected to the BMO via IPs. E2B biological functional correspondents, were tagged with IPs rather than the biological text data from the BMO as the biological data within the BMO will be continually updated by J.F.V.V., but the IPs (which have already been tagged with words from the E2B Thesaurus) will remain intact. Therefore, the connection between the E2B Thesaurus and the BMO will be automatically updated as the BMO is expanded. Relevant biological function correspondents for a given IP were selected during several meetings by four of the authors (S.J.M., A.M.G., C.K.U., and A.R.) whose expertise included education, ecology, engineering, and zoology. A biological functional correspondent was connected to an IP only if all four authors agreed that the given biological functional correspondent was related to a particular IP. If one or more of the authors listed above disagreed with the particular connection being discussed, the biological functional correspondent was not tagged to that particular IP. The subjective biological function correspondents to IP connections are listed in Table 1.

Table 1.
List of all subjective biological function correspondents to IP connections.
2.2.2. Creating a Combined User Interface
E2BMO is formed using SKOS as a communication tool connecting the E2B to the BMO within Protégé. Protégé provides a graphic user interface to define ontologies using the Web Ontology Language (OWL). It also enables the ontology to be exported into RDF_XML format. RDF, Resource Description Framework, is a format used in semantic web applications. Unlike the original web that is concerned with exchange of documents, the semantic web is concerned with creating a common format for data integration and sharing. RDF uses triples to represent information; subject, predicate, and object, where each subject (trade-off) and its objects (IPs) are given a code number (BMO_xxxx). These relations are then saved in an XML (Extensible Markup Language) format, (Figure S1) which uses a set of rules to encode information in a human and machine-readable format.
SPARQL, a query language that uses a series of protocols in semantic web to access data, was used to access information from the RDF_XML format. SPARQL queries were written in Python, (Figure S2 shows one example of SPARQL query searching for IPs associated with biological models of a given trade-off). Other queries find E2B text incorporated into IPs. These are then connected to biological models using same IPs.
We used two exported BMO files, one integrated with E2B and the other without. Using IPs, we were able to connect these and generate 25 text files, (Figure S3 containing information on all currently available query words. Then, we used JavaScript, (Figure S4), in HTML (HyperText Markup Language) to develop the E2BMO website that takes users input, finds related datafile and finally creates an appropriate table and/or graph. A descriptive illustration of the operation of the interface design is presented in Figure 4.
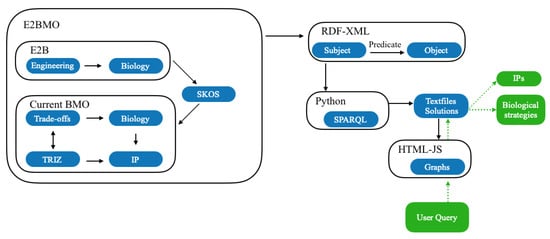
Figure 4.
Green boxes present the user interaction with E2BMO and blue boxes are the underlying building blocks of E2BMO.
3. Results
3.1. E2BMO Website
The homepage of the ‘E2BMO’ website (www.uakron.edu/BRIC/E2BMO; Figure 5) follows a simple layout featuring a search bar and two main access points: (1) using BMO trade-off concepts and (2) using E2B functional words. Through both approaches, the user is free to choose the type of solution discovery approach they wish to pursue. Those who wish to take a more technical engineering approach to solution discovery may pursue IPs and sub-IPs. Users interested in a more interdisciplinary innovation process may choose to pursue biological literature. Users wishing to embark on broader ideation and exploratory types of the creative process may pursue biological functional correspondent terms. Thus, the aim of this interface is to create a platform for a variety of users to interact with the biomimicry design process through an ontology.
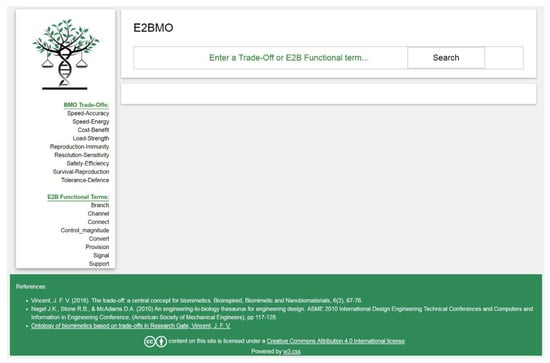
Figure 5.
Homepage of E2BMO Website.
3.1.1. BMO Trade-Off Concepts
Here the user can choose one of the nine trade-offs currently suggested, under the ‘BMO Trade-Offs’ column on the left-hand side and type the one selected into the query box. The search engine then identifies IPs, biological strategies previously abstracted from the BMO and tagged as potential solutions for this specific trade-off. The results appear as a table (Figure 6). The main table presents, in the first and second column, the nature of the trade-off being solved. The third column provides the IPs and sub-IPs derived from biology as potential solutions. The second table contains references to the biological models that these strategies were abstracted from, if the user wishes to explore the biological content. This second table is greyed out to avoid confusion between the engineering related results and the biological models, but by simply moving the mouse over the cell, the content will become clearly visible (Figure 6). Figure 6 shows a subset of the definitions to the ‘speed-accuracy’ trade-off query; 11 examples are currently available in the BMO. The results appear on the same page and a user can scroll down to access the table for more examples.

Figure 6.
E2BMO query result for BMO Trade-Off Speed–Accuracy.
3.1.2. E2B Functional Words
Through this option, the user can select one of the eight terms available under the ‘E2B Functional Words’ column on the left-hand side and type the one selected into the query box. Figure 7 and Figure 8 show the hierarchical graphs and results table that generate from querying the functional term ‘signal’. Figure 7 illustrates the class, secondary, and tertiary engineering related functional terms from the E2B Thesaurus in black. The green and orange terms represent the functional biological correspondents of the E2B thesaurus that are connected to IPs of the BMO (in red). Figure 8 illustrates sub-IPs in purple, which are connected to biological models presented in blue, from which both IPs and sub-IPs were abstracted from. On the interface, below this hierarchical graph is a trade-off table (Figure 8). This table represents how the E2B functional term can result in trade-offs connected to IPs and sub-IPs, and if the desired, the biological models these IPs were generated from. This approach allows a user who has not identified a trade-off they wish to solve for, to explore trade-offs and IPs through a functional term search. These IPs and trade-offs could inform a new way of thinking about a technical challenge and spark ideation. The user is free to navigate through the hierarchical tree and the results table to identify an output that is most relevant to their intended purpose: functional biological correspondent terms, IPs, and sub-IPs, or biological literature.
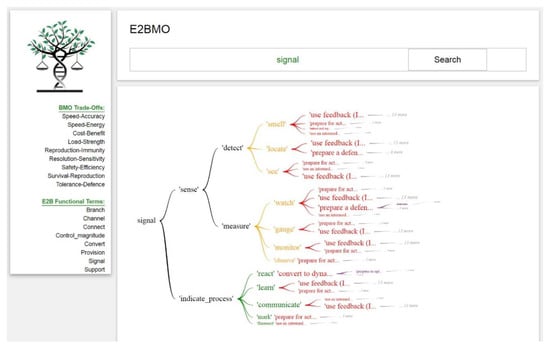
Figure 7.
E2BMO query result for E2B functional word ‘signal’ to IPs in red.
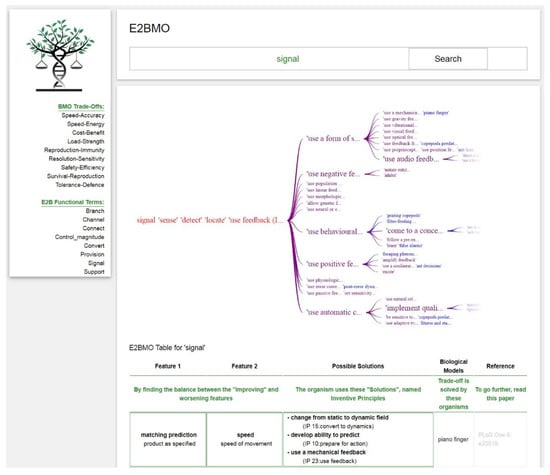
Figure 8.
E2BMO query result for E2B functional word ‘signal’ from IPs to biological texts in blue through sub-IPs in purple, results table included on the same page.
3.2. Case Study
The following proof of concept case study illustrates how the proposed E2BMO website could be used to navigate through the biomimicry design process, as outlined in Figure 1, leading to a simplified model for both approaches (Figure 9). The hypothetical technical challenge faced in this case study is the same challenge described throughout this paper, a sorting mechanism to identify ripeness of fruit, for example bananas or plums, through sensor activated image recognition as transported on a conveyor belt. We will explore the two options: (1) BMO Trade-Offs and (2) E2B Functional Words, available through this website.
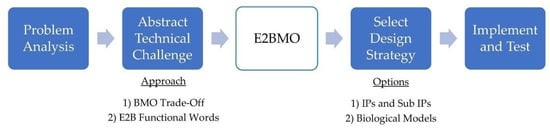
Figure 9.
Illustration of the simplified biomimicry process through use of the E2BMO website.
3.2.1. BMO Trade-Offs
● Step 1: Problem Analysis
The user must define the components contributing to their technical challenge. In this case study, the contributing factors identified are the speed of the conveyor belt and the accuracy of the sensor activated image recognition system.
● Step 2: Abstract Technical Challenge
These contributing components then need to be abstracted in terms of a trade-off in relation to the BMO Trade-Offs available. In the given case, a compromise needs to be made between the speed of the conveyor belt and the accuracy of the images captured. Although a higher belt speed would lead to more fruit processed, if the accuracy of the image capture system is not correlated this could lead to a higher failure rate and a manufacturing time and monetary loss. The possible technical challenge abstraction in the form of a trade-off would be speed versus accuracy.
● Step 3: Transpose to Biology
This step is bypassed as the trade-offs have been previously tagged to biological texts through IPs, thus bypassing the need to transpose engineering terminology to biological.
● Step 4: Identify Potential Biological Models
This fourth step is completed by the software which outputs biological information in a table format for the user to explore, if they wish to do so. This step is not required to advance through the process. In this case study, 11 biological models within the speed and accuracy trade-off were identified (Figure 6).
● Step 5: Select Biological Model(s) of Interest
This step traditionally is to narrow the selection of biological models to a reasonable subset that can be explored and understood in a more plausible amount of time. This complex step to understand biological text has been previously accomplished through the BMO, thus the user can choose to completely skip this step if they wish. If the user is involved in an interdisciplinary design team and participants wish to explore the biological information, they may choose to scroll through the main table, specifically the first and second column (Figure 6). Then, they can select which context of the speed-accuracy trade-off is most relevant to their context and explore the biological information made available in the corresponding biological reference table.
● Step 6: Abstract Biological Strategies
This step is completed by the software as the BMO has previously abstracted the biological strategies and tagged them with IPs. Thus, the IPs and sub-IPs generated in the results table are the biological strategies in engineering terminology. These IPs and sub IPs describe technical strategies used to solve for the functional challenge queried. Thus, saving the user the laborious task of reading and comprehending biological text to best abstract biological strategies. Of the 11 biological models identified, they included 9 unique IPs and 28 unique sub-IPs. Two recurring IPs are: (1) integrate several objects and systems, and (2) change response threshold (Figure 6).
● Step 7: Transpose to Technology
This step is bypassed as the technical challenge was never transposed from technology to biology in the first place, thus a re-transposition to technology is not necessary.
● Step 8: Implement and Test
The IP chosen from Step 6: integrate several objects or systems, could translate to a solution which integrates triggers along the conveyor belt. These triggers reduce the speed of a passing fruit creating a localized opportunity providing the sensor with sufficient time to capture a more accurate image. Testing several iterations of this technical solution is necessary to optimize the design.
3.2.2. E2B Functional Words
● Step 1: Problem Analysis
Same as outlined through the BMO Trade-Off approach, refer to Step 1 in Section 3.2.1.
● Step 2: Abstract Technical Challenge
Same as outlined through the BMO Trade-Off approach, refer to Step 2 in Section 3.2.1. The possible technical challenge abstraction would be to use a signaling mechanism between the sensors and conveyor belt. Hence, in the form of an E2B Functional Word the abstracted technical challenge would be ‘Signal’.
● Step 3. Transpose to Biology
The query of ‘Signal’ will result in a tree diagram (Figure 7 and Figure 8) that enables the user to navigate through the hierarchical structure of the Engineering-to-Biology Thesaurus selecting engineering related terms, in black, that are most suitable to their specific technical challenge. These terms are directly connected to corresponding biological functional words. For example, with our case study, the user may choose to query the engineering class ‘signal’ then select ‘sense’ as a more applicable engineering related term compared to ‘indicate_process’. ‘Sense’ is then further categorized to tertiary engineering terms, such as ‘detect’ and ‘measure’. These terms are directly correlated with biological functional correspondents, in green and orange (Figure 7), such as ‘see’ and ‘locate’.
● Step 4: Identify Potential Biological Models
Same as outlined through the BMO Trade-Off approach, refer to Step 4 in Section 3.2.1. In this case study, 27 biological models within the ‘Signal’ query were identified (Figure 8), 11 of which are the same as those identified through the BMO Trade-Off approach. Biological models, in blue, may also be identified if the user continues to navigate through the hierarchical structure of the tree diagram as IPs and sub-IPs are tagged to the biological models they originated from (Figure 8).
● Step 5: Select Biological Model(s) of Interest
Same as outlined through the BMO Trade-Off approach, refer to Step 5 in Section 3.2.1 and Figure 8.
● Step 6: Abstract Biological Strategies
Same as outlined through the BMO Trade-Off approach, refer to Step 6 in Section 3.2.1. In this case study as shown in Figure 7 and Figure 8, several IPs are repeated such as: ‘Use Feedback’, Prepare for Action’ and ‘Convert to Dynamics’. Thus, the user may correlate these to create a set of potential solutions or select one to explore further. For example, ‘Use Feedback’ is heavily repeated, and thus is a visibly relevant IP for this technical challenge.
● Step 7: Transpose to Technology
This step is bypassed as previously accomplished within Step 6.
● Step 8: Implement and Test
The IP chosen from Step 6 ‘Use Feedback’, could translate to a solution which integrates triggers along the conveyor belt to provide feedback for image capture. These triggers reduce the speed of a passing fruit creating a localized feedback for the sensor to activate image capture. Testing several iterations of this technical solution is necessary to optimize the design.
4. Related Work
Ontologies and other related computer-aided biomimetic tools have and are being developed by practitioners. Kruiper et al. shared a design process for computer-aided biomimetics where trade-offs are used to bridge the biology and engineering disciplines to make biological information more accessible to engineers (2018). The trade-off based BMO is only one of a few ontologies created for biomimicry. Also developed are the Biomimetics Ontology Database, a database with an embedded keyword exploration tool to find and extract information through an interface [38], and the Ontology for Bio-inspired Design, which centers around functional, architectural, and behavioral links between biology, engineering, and bio-inspired design solutions based on classification hierarchies built on description logic [39]. As noted, Kozaki and Mizoguchi (2014) have also approached the development of an ontology interface tool to interact with the Biomimetics Ontology Database. Their tool is grounded in association-based inference of the database, and exploits not only function, but other kinds of analogical inference surrounding such things as property, structure, and living environment [38].
The Function–Behavior–Structure (FBS) Ontology and other related FBS models could also be useful for automatically searching and extracting information from literature to populate an ontology for biomimicry, or potentially aiding in front-end querying, for example navigating the E2B thesaurus hierarchy to narrow down functional search terms [40]. This aspect of keyword retrieval work is close-ended enabling the retrieval of a pre-convinced set output, such as a keyword. This is common with many other biomimetic tools but is only partially relevant to the E2BMO, since the E2B is essentially a keyword system. With the entire E2BMO, the retrieval is for a biomimetic database—in essence a compendium of biomimetic effects and solutions which have already been solved. The ambition of the E2BMO is greater in that it is designed to capture and support biomimetic novelty—i.e., discovering new biomimetic solutions and concepts, making it an open-ended tool. Related work includes that of Li Shu who uses an open-ended prompt system to search cases within a digitized biology textbook [41].
Extracting information to populate databases and ontologies used for biomimicry is a factor limiting the comprehensiveness of these tools. Instances, such as Julian Vincent’s efforts to abstract and translate knowledge from biological texts to populate the BMO, or the Biomimicry Institute’s work curating and translating data for AskNature.org, are arduous manual tasks that rely on extensive time and intellectual commitment. Data extraction methods are being developed to help gather data to automatically populate databases and ontologies. For example, using Artificial Intelligence to mine relevant information from biological texts could potentially be used to populate the BMO in the future. A current area of research in Scotland is using Natural Language Processing, Naive Bayes, and neural nets, combined with manual annotation to develop a faster technique for extracting trade-offs and their resolution from biological and agricultural research papers. Although attempts at automation have historically had variable success, this is an active, up-and-coming research area with high potential.
Other research that could be deemed as ‘related work’ is that of bioTRIZ, although there are a number of distinct differences to the work presented here and specifically that of the BMO. bioTRIZ is the result of work done by J.F.V.V. and associates [42] and is not traditional TRIZ. In this paper, they stated that the bioTRIZ philosophy is, relating inventive principles of TRIZ into a new order that more closely reflects the biological route to the resolution of conflicts, with conflicts being among aspects of energy, space, time, substance, structure, and information. ‘These six operational fields re-organize and condense the TRIZ classification both of the features used to generate the conflict statements and the inventive principles’ (p. 476). This classification was performed by close interpretation of Altshuller’s definitions of the IPs as recorded in numerous Russian texts. bioTRIZ does not use a thesaurus, and the IPs are lumped into larger groupings in contrast to the BMO, which has introduced much more detail into the IPs specifically to make it easier to relate them to biological phenomena. In addition, bioTRIZ was not created within an ontology. The BMO has very specifically defined trade-offs identified by biologists in their literature work whereas bioTRIZ retained the original TRIZ contradiction matrix approach whereby each pair of engineering parameters was deemed to represent a conflict.
5. Discussion
The biomimicry design process, as outlined in the literature, traditionally follows 8 steps, as seen in Figure 1, or a variation of it. Through the illustrative case study described in Section 3.2 it is clear to see that the proposed E2BMO tool does not conform to these 8 steps (Figure 9). Both approaches have steps that are either bypassed, skipped entirely, or are completed by the software. Step 7, ‘Transpose to Technology’, is bypassed by both approaches. Traditionally, this step requires a complex understanding of the biological strategy and system of interest in sufficient detail to comprehensively translate this information into technological terminology that can be communicated effectively to an interdisciplinary group. This complicated transition from biological to engineering terminology is identified as one of the most difficult steps of the biomimicry process. However, through the E2BMO website, this step has been accomplished within the BMO, through tagging IPs with the biological strategies identified within the literature. Because the BMO Trade-Off approach begins by specifically abstracting the technical challenge as a trade-off, a concept more familiar to practitioners from technical backgrounds, Step 3 is also bypassed. This task of tagging biological strategies with IPs requires a complex understanding of both biological and technical terminology. A current project with the BMO is the development of an automated process that analyzes biological information to populate the ontology, while monitored by subject matter experts to ensure validity. Although Step 3 is not bypassed through the E2B Functional Word approach, it is heavily facilitated. Users can navigate through an interactive hierarchical tree representing the Engineering-to-Biology Thesaurus by advancing through each level of the hierarchy by simply selecting one of the available terms that they believe is most applicable to their technical challenge. They have the option to advance through IPs that have been drawn from biological texts that are attached for reference at the end. By-passing these steps has a number of disadvantages; loss of interaction and learning of biological content with the potential loss of addition inspiration or features for an engineering design, and advantages; saving time and effort spent comprehending the biological information and due to this possible increasing the adoption of biomimicry. To attempt to mitigate these advantages and disadvantages we have left it to the user to decide whether or not to interact with the biological content, by making it readily available on the same pane of view. It is important to note that the bypass and simplification of translation Steps 3 and 7 not only eases the communication and collaboration of interdisciplinary teams, it also calls into question the need for a semantic bridge if the biological information does not need identification, exploration, or comprehension.
The E2BMO software completes Steps 4 and 6 for both approaches and the nature of the results generated leaves Step 5 as an optional step for the user to explore if they so wish. This reduces the overall length of the process, another promising factor to increase user interest and the further adoption of biomimicry by other disciplines, especially for industrial use where strict deadlines and severe time constraints are common problems. Step 4 and 6, ‘Identify Potential Biological Models’ and ‘Abstract Biological Strategies’, are completed by the software which outputs biological strategies in a table format, as seen in Figure 6 and Figure 8, for the user to explore. This also reduces the time spent searching for credible sources of biological information that may be relevant to the technical challenge of interest, a difficult task for a user not familiar with biological journals or publications. This, again, reduces the users’ need to transcend disciplinary boundaries to advance through the biomimicry process, increasing user interest. This results table serves a double purpose: (1) a cohesive and simplified representation of complex information; and (2) direct access to a plethora of potential solutions: IPs and sub-IPs.
The illustrative case study was written to demonstrate that both approaches can result in similar solution discovery through two identified IPs. With that said, upon selecting the IP of interest, the interpretation and application of such is dependent upon the users’ understanding of both the strategy explained and the technical challenge of application. Thus, even with only one relevant IP as a potential design strategy, several solutions can be generated based on the composition of the design team tasked with applying it. Upon exploration of either approach within the E2BMO website, several IPs and several more sub IPs are presented that can be either used individually, similar to the case study provided, or compounded to create a more complex design strategy, broadening solution discovery.
This current iteration of the BMO is a research tool with finite data, but it will continue to be populated by J.F.V.V., increasing its capabilities for the discovery of solutions to real-world problems. This paper has made significant advances in the development of a simple front-end for the BMO and in its current iteration could be useful as an ideation platform for real-world problems as demonstrated through the case study (Section 3.2). The E2BMO was created using this current iteration of the BMO and allows the user direct access to IPs without the need to explore biological text, a difficult and time-consuming task for anyone, especially those from outside a biological discipline. Also, having these biological texts tagged with IPs by a subject matter expert reduces the risk of a user misunderstanding or misinterpreting complex biological strategies. However, further investigation on the potential misuse of biological information by non-experts during the biomimetic process is needed. The option to explore biological text is available for the user to make their own judgement on the biological strategy if they wish to do so, but with the E2BMO tool this step is not required to advance through the biomimicry process.
Our efforts to link these tools aim to bring awareness to the copious tools available, and provide practitioners with a possible approach to more easily navigate through the biomimicry process and two selected tools. Therefore, the E2BMO could enable greater potential for future biomimicry practice. We demonstrate that the developed E2BMO website enables practitioners to both interface and extract information from the E2B Thesaurus and the BMO, especially in ways that were not possible before. We anticipate that this user-interface will give the average practitioner a more intuitive interaction with these tools, possibly increasing the usability of the biomimicry process, functional analysis, and an ontology. Users are encouraged to delve into each of the options available within the website and to freely navigate through the software to explore the innovative potential that each of these approaches offer. Through use of the E2BMO, practitioners will be able to better interact with complex biological knowledge without a heavy investment of time and energy, encouraging a more widespread implementation of biomimicry.
The E2BMO Website is in a beta-testing phase, thus future work is necessary, which may include:
- Expanding E2B thesaurus words through SKOS to include additional biological functions such as ‘pollinate, harvest, mate, etc.’.
- Translating the E2B Thesaurus Flow Terms to SKOS terms and tagging to inventive principles.
- Creating direct links to images or diagrams within the biological text data that may explain the design principle and help navigate the biological text in an interdisciplinary project.
- Analyzing the tool within an industrial context solving for real world technical challenges.
- Future work could potentially include incorporating data contributions from biologists into the E2BMO thus acting as an information sharing platform to a more diverse audience, possibly leading to potential collaboration and funding.
Supplementary Materials
The following is available online at http://www.mdpi.com/2411-9660/2/4/53/s1, Figure S1: This is the XML-RDF format of the OWL exported file from Protégé. <owl:Class> and </owl:Class> show the start and end of a subject, in here ‘speed-accuracy’ class, Figure S2: Red section of the code is SPARQL query. This query looks for objects connected to a class/subject using their properties. ‘query_with_placeholder’ passes the query string to ‘g.query’ from ‘rdflib’ library. “$ENTITY_CODE” is a placeholder for the name of class provided by the user or other queries, Figure S3: This is an example of contents in final text file for ‘speed-accuracy trade-off’, each variable is separated by ‘::’ which helps when extracting data from the file. It starts by name of biological model, its TRIZ thesis and antithesis, abstract of the paper, followed by IPs and Sub IPs and any other information needed, Figure S4: This is a portion of JavaScript code in HTML where E2BMO graph is created, we are using google Word-Tree visualization to develop the hierarchical graph shown in Figure 7.
Author Contributions
Conceptualization, S.J.M., B.K., A.M.G., T.H., A.R., C.KU., N.W., and P.H.N.; Methodology, S.J.M., B.K., A.M.G., T.H., A.R., C.K.U., N.W., J.F.V.V., J.K.N., and P.H.N.; Software, B.K.; Investigation, S.J.M., B.K., A.M.G., T.H., A.R., C.K.U., N.W., and P.H.N.; Resources, J.F.V.V., J.K.N., and P.H.N.; Data curation, S.J.M., B.K., A.M.G., A.R., and C.K.U.; Writing—original draft preparation, S.J.M., B.K., A.M.G., and T.H.; Writing—review and editing, S.J.M., B.K., A.M.G., T.H., A.R., C.K.U., N.W., J.F.V.V., J.K.N., and P.H.N.; Visualization, T.H.; Supervision, S.J.M.; Project administration, P.H.N.
Funding
This research received no external funding.
Acknowledgments
This work was conducted using the Protégé resources, which is supported by grant GM10331601 from the National Institute of General Medical Sciences of the United States National Institutes of Health.
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
| E2BMO | Engineering-to-BioMimetic Ontology |
| E2B | Engineering-to-Biology Thesaurus |
| BMO | BioMimetic Ontology |
| TRIZ | Teorija Reshenija Izobretateliskih Zadatch, which translates to Theory of Solving Problems Inventively |
| BFO | Basic Formal Ontology |
| IPs | Inventive Principles |
| SKOS | Simplified Knowledge Organization System |
| OWL | Web Ontology Language |
| RDF | Resource Description Framework |
| XML | Extensible Markup Language |
| SPARQL | a RDF query language |
| HTML | Hypertext markup language |
| JS | JavaScript |
| CSS | Cascading Style Sheets |
References
- Benyus, J.M. Spreading the Meme: A Biomimicry Primer. In Biomimicry Resource Handbook: A Seed Bank of Best Practice; Biomimicry 3.8: Missoula, MT, USA, 2013; pp. 7–19. ISBN 9781505634648. [Google Scholar]
- Biomimicry Global Design Challenge Toolbox: Case Studies. Available online: https://toolbox.biomimicry.org/references/case-studies/ (accessed on 12 October 2018).
- Mann, E.E.; Manna, D.; Mettetal, M.R.; May, R.M.; Dannemiller, E.M.; Chung, K.K.; Brennan, A.B.; Reddy, S.T. Surface micropattern limits bacterial contamination. Antimicrob. Resist. Infect. Control 2014, 3, 28. [Google Scholar] [CrossRef] [PubMed]
- Faure, E.; Lecomte, P.; Lenoir, S.; Vreuls, C.; Van De Weerdt, C.; Archambeau, C.; Martial, J.; Jérôme, C.; Duwez, A.-S.; Detrembleur, C. Sustainable and bio-inspired chemistry for robust antibacterial activity of stainless steel. J. Mater. Chem. 2011, 21, 7901–7904. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, Z.; Zhang, Y.; Wei, D.; Deng, Y. Route selection for emergency logistics management: A bio-inspired algorithm. Saf. Sci. 2013, 54, 87–91. [Google Scholar] [CrossRef]
- Kerbel, M.; Hoeller, N.; McKeag, T. REGEN Energy: The Power of Ants and Bees. Zygote Quarterly. 2012. Available online: https://zqjournal.org/editions/zq01.html (accessed on 12 October 2018).
- Fermanian Business & Economic Institute. Bioinspiration: An Economic Progress Report [Internet]; Point Loma Nazarene University: San Diego, CA, USA, 2013; Available online: http://www.magnefico.com/fileadmin/user_upload/Dokumente/PLNU_Bioinspiration_Da_Vinci_Index_A_Progress_Report_November_2013_Final.pdf (accessed on 28 March 2018).
- Wanieck, K.; Fayemi, P.-E.; Maranzana, N.; Zollfrank, C.; Jacobs, S. Biomimetics and its tools. Bioinspired Biomim. Nanobiomater. 2017, 6, 53–66. [Google Scholar] [CrossRef]
- Vattam, S.; Helms, M.; Goel, A.K. Biologically-Inspired Innovation in Engineering Design: A Cognitive Study; Georgia Institute of Technology: Atlanta, Georgia, 2007; p. 41. [Google Scholar]
- Helms, M.E.; Vattam, S.S.; Goel, A.K.; Yen, J.; Weissburg, M. Problem-Driven and Solution-Based Design. In Proceedings of the 28th Annual Conference of the Association for Computer Aided Design in Architecture (ACADIA), Minneapolis, MN, USA, 16–19 October 2008; pp. 94–101. [Google Scholar]
- Helms, M.; Vattam, S.S.; Goel, A.K. Biologically inspired design: Process and products. Des. Stud. 2009, 30, 606–622. [Google Scholar] [CrossRef]
- Fayemi, P.-E.; Wanieck, K.; Zollfrank, C.; Maranzana, N.; Aoussat, A. Biomimetics: Process, tools and practice. Bioinspir. Biomim. 2017, 12, 011002. [Google Scholar] [CrossRef] [PubMed]
- Massey, A.P.; Wallace, W.A. Understanding and facilitating group problem structuring and formulation: Mental representations, interaction, and representation aids. Decis. Support Syst. 1996, 17, 253–274. [Google Scholar] [CrossRef]
- Sartori, J.; Pal, U.; Chakrabarti, A. A methodology for supporting “transfer” in biomimetic design. Artif. Intell. Eng. Des. Anal. Manuf. 2010, 24, 483–506. [Google Scholar] [CrossRef]
- Baumeister, D. Biomimicry Resource Handbook: A Seed Bank of Best Practice; Biomimicry 3.8: Missoula, MT, USA, 2013; ISBN 9781505634648. [Google Scholar]
- Bogatyrev, N.R. Microfluidic Actuation in Living Organisms: A Biomimetic Catalogue. In Proceedings of the First European Conference on Microfluidics, Bologna, Italy, 10–12 December 2008; p. 175. [Google Scholar]
- Lenau, T. Biomimetics as a Design Methodology—Possibilities and Challenges. In Proceedings of the 17th International Conference on Engineering Design (ICED 09), Palo Alto, CA, USA, 24–27 August 2009. Volume 5 Design Methods and Tools (pt. 1). [Google Scholar]
- Cheong, H.; Chiu, I.; Shu, L.H.; Stone, R.B.; McAdams, D.A. Biologically Meaningful Keywords for Functional Terms of the Functional Basis. J. Mech. Des. 2011, 133, 021007. [Google Scholar] [CrossRef]
- Nagel, J.K.S.; Stone, R.B.; McAdams, D.A. An Engineering-to-Biology Thesaurus for Engineering Design. In Volume 5: 22nd International Conference on Design Theory and Methodology; Special Conference on Mechanical Vibration and Noise, Montreal, QC, Canada, 15–18 August 2010; ASME: New York, NY, USA, 2010; pp. 117–128. [Google Scholar]
- Kruiper, R.; Vincent, J.; Abraham, E.; Soar, R.; Konstas, I.; Chen-Burger, J.; Desmulliez, M. Towards a Design Process for Computer-Aided Biomimetics. Biomimetics 2018, 3, 14. [Google Scholar] [CrossRef]
- Autumn, K. How Gecko Toes Stick. Am. Sci. 2006, 94, 123–132. Available online: https://webdisk.lclark.edu/xythoswfs/webui/_xy-1594799_1-t_d9VVAITO (accessed on 12 October 2018). [CrossRef]
- Autumn, K.; Sitti, M.; Liang, Y.A.; Peattie, A.M.; Hansen, W.R.; Sponberg, S.; Kenny, T.W.; Fearing, R.; Israelachvili, J.N.; Full, R.J. Evidence for van der Waals adhesion in gecko setae. Proc. Natl. Acad. Sci. USA 2002, 99, 12252–12256. [Google Scholar] [CrossRef] [PubMed]
- Hiller, U. Untersuchungen zum Feinbau und zur Funktion der Haftborsten von Reptilien. Z. Morphol. Tiere 1968, 62, 307–362. [Google Scholar] [CrossRef]
- Maderson, P.F.A. Keratinized Epidermal Derivatives as an Aid to Climbing in Gekkonid Lizards. Nature 1964, 203, 780–781. [Google Scholar] [CrossRef]
- Ruibal, R.; Ernst, V. The structure of the digital setae of lizards. J. Morphol. 1965, 117, 271–293. [Google Scholar] [CrossRef] [PubMed]
- Hansen, W.R.; Autumn, K. Evidence for self-cleaning in gecko setae. Proc. Natl. Acad. Sci. USA 2005, 102, 385–389. [Google Scholar] [CrossRef] [PubMed]
- Niewiarowski, P.H.; Stark, A.Y.; Dhinojwala, A. Sticking to the story: Outstanding challenges in gecko-inspired adhesives. J. Exp. Biol. 2016, 219, 912–919. [Google Scholar] [CrossRef] [PubMed]
- Stark, A.Y.; Badge, I.; Wucinich, N.A.; Sullivan, T.W.; Niewiarowski, P.H.; Dhinojwala, A. Surface wettability plays a significant role in gecko adhesion underwater. Proc. Natl. Acad. Sci. USA 2013, 110, 6340–6345. [Google Scholar] [CrossRef] [PubMed]
- Kaiser, M.K.; Hashemi Farzaneh, H.; Lindemann, U. An approach to support searching for biomimetic solutions based on system characteristics and its environmental interactions. In DS 70: Proceedings of DESIGN 2012, the 12th International Design Conference, Dubrovnik, Croatia, 21–24 May 2012; pp. 969–978.
- Jacobs, S.R.; Nichol, E.C.; Helms, M.E. “Where Are We Now and Where Are We Going?” The BioM Innovation Database. J. Mech. Des. 2014, 136, 111101. [Google Scholar] [CrossRef]
- Vincent, J.F.V. The trade-off: A central concept for biomimetics. Bioinspired Biomim. Nanobiomater. 2017, 6, 67–76. [Google Scholar] [CrossRef]
- The OBO Foundry. Available online: http://www.obofoundry.org/ (accessed on 12 October 2018).
- Hirtz, J.; Stone, R.B.; McAdams, D.A.; Szykman, S.; Wood, K.L. A functional basis for engineering design: Reconciling and evolving previous efforts. Res. Eng. Des. 2002, 13, 65–82. [Google Scholar] [CrossRef]
- Altshuller, G.S. Creativity as an Exact Science: The Theory of the Solution of Inventive Problems; CRC Press LLC: Boca Raton, FL, USA, 1984; ISBN 9780677212302. [Google Scholar]
- Musen, M.A. The Protégé Project: A Look Back and a Look Forward. AI Matters 2015, 1, 4–12. [Google Scholar] [CrossRef] [PubMed]
- Miles, A.; Bechhofer, S.; SKOS Simple Knowledge Organization System Reference [Internet]. W3C Recommendation c2009. Available online: https://www.w3.org/TR/2009/REC-skos-reference-20090818/ (accessed on 28 March 2018).
- Baker, T.; Bechhofer, S.; Isaac, A.; Miles, A.; Schreiber, G.; Summers, E. Key choices in the design of Simple Knowledge Organization System (SKOS). Web Semant. Sci. Serv. Agents World Wide Web 2013, 20, 35–49. [Google Scholar] [CrossRef]
- Kozaki, K.; Mizoguchi, R. A Keyword Exploration for Retrieval from Biomimetics Databases. In Semantic Technology; Supnithi, T., Yamaguchi, T., Pan, J.Z., Wuwongse, V., Buranarach, M., Eds.; Springer International Publishing: Cham, Switzerland, 2015; Volume 8943, pp. 361–377. ISBN 978-3-319-15614-9. [Google Scholar]
- Yim, S.; Wilson, J.O. Development of an Ontology for Bio-Inspired Design using Description Logics. In Proceedings of the International Conference on Product Lifecycle Management, Seoul University, South Korea; July 2008. [Google Scholar]
- Gero, J.S.; Kannengiesser, U. A function-behavior-structure ontology of processes. AI Edam 2007, 21, 379–391. [Google Scholar] [CrossRef]
- Shu, L.H. A natural-language approach to biomimetic design. Artif. Intell. Eng. Des. Anal. Manuf. 2010, 24, 507–519. [Google Scholar] [CrossRef]
- Vincent, J.F.V.; Bogatyreva, O.A.; Bogatyrev, N.R.; Bowyer, A.; Pahl, A.-K. Biomimetics: Its practice and theory. J. R. Soc. Interface 2006, 3, 471–482. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).