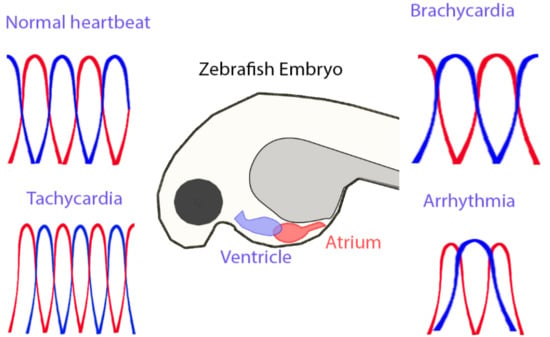
A Simple ImageJ-Based Method to Measure Cardiac Rhythm in Zebrafish Embryos
Abstract
:1. Introduction
2. Materials and Methods
2.1. Zebrafish Maintenance and Sample Preparation
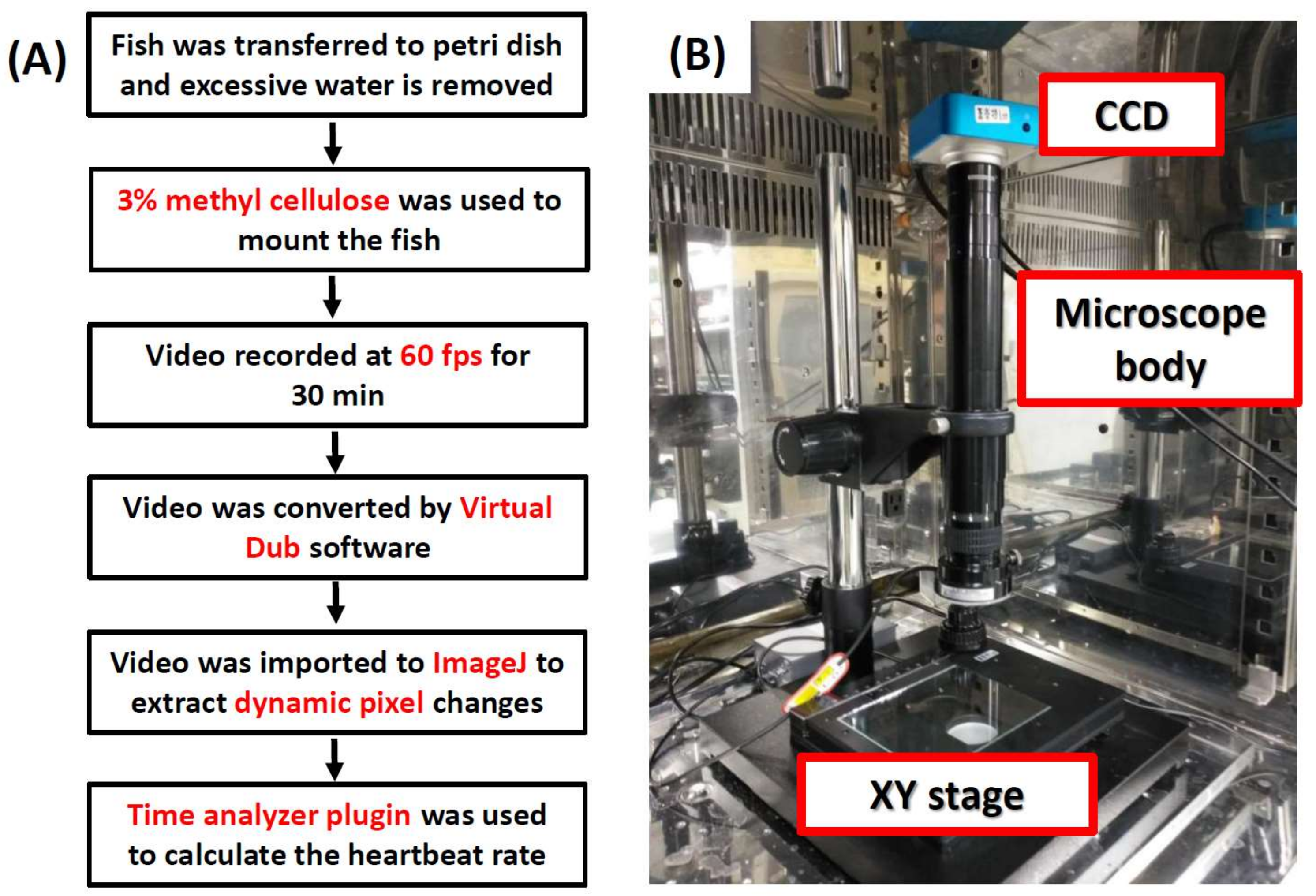
2.2. Video Recording
2.3. Video Processing
2.4. Time Interval and BPM Analysis
2.5. Drug Treatment
2.6. Statistical Analysis
3. Results
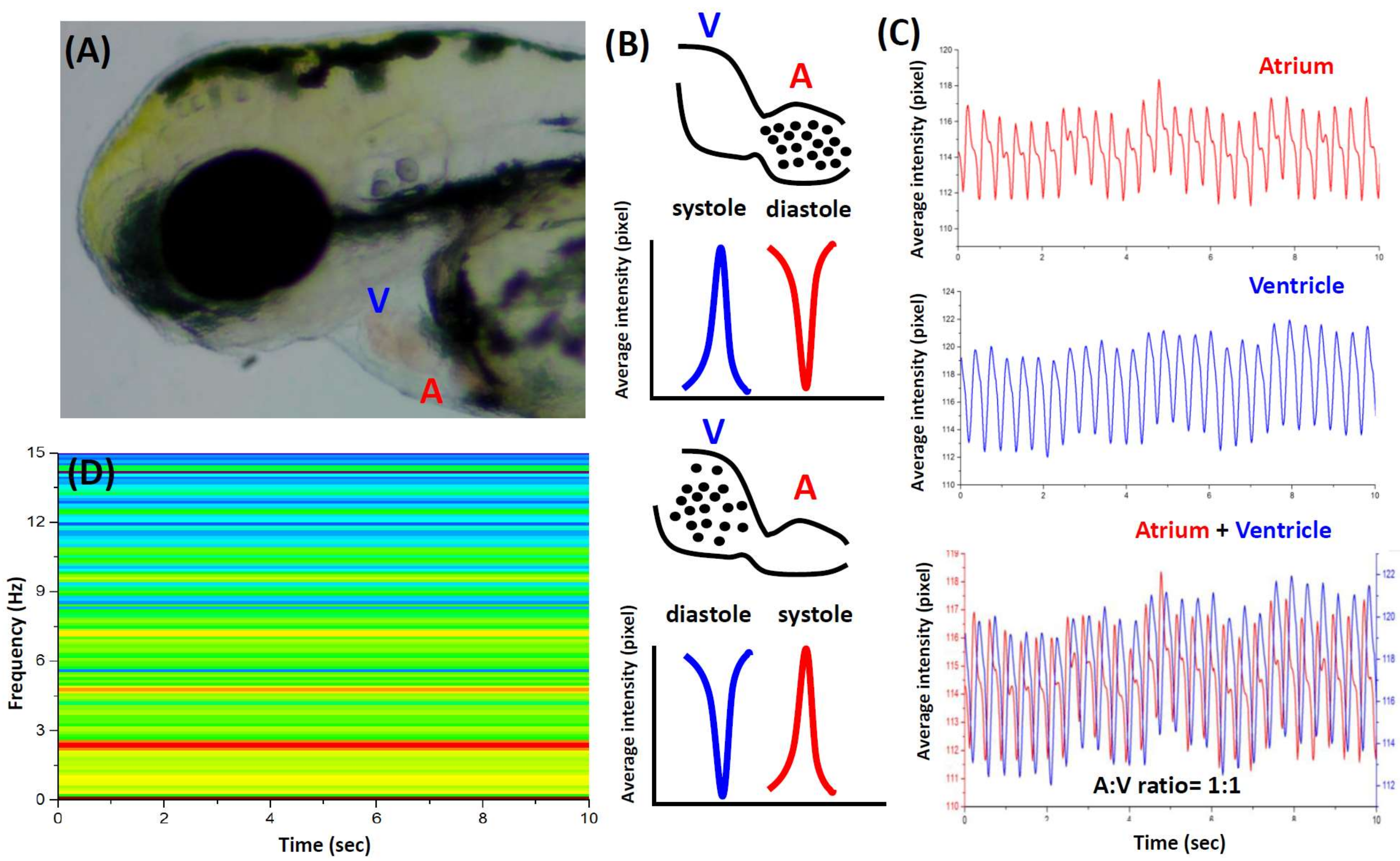
3.1. Cardiac Rhythm Can Be Detected by the Dynamic Pixel Method
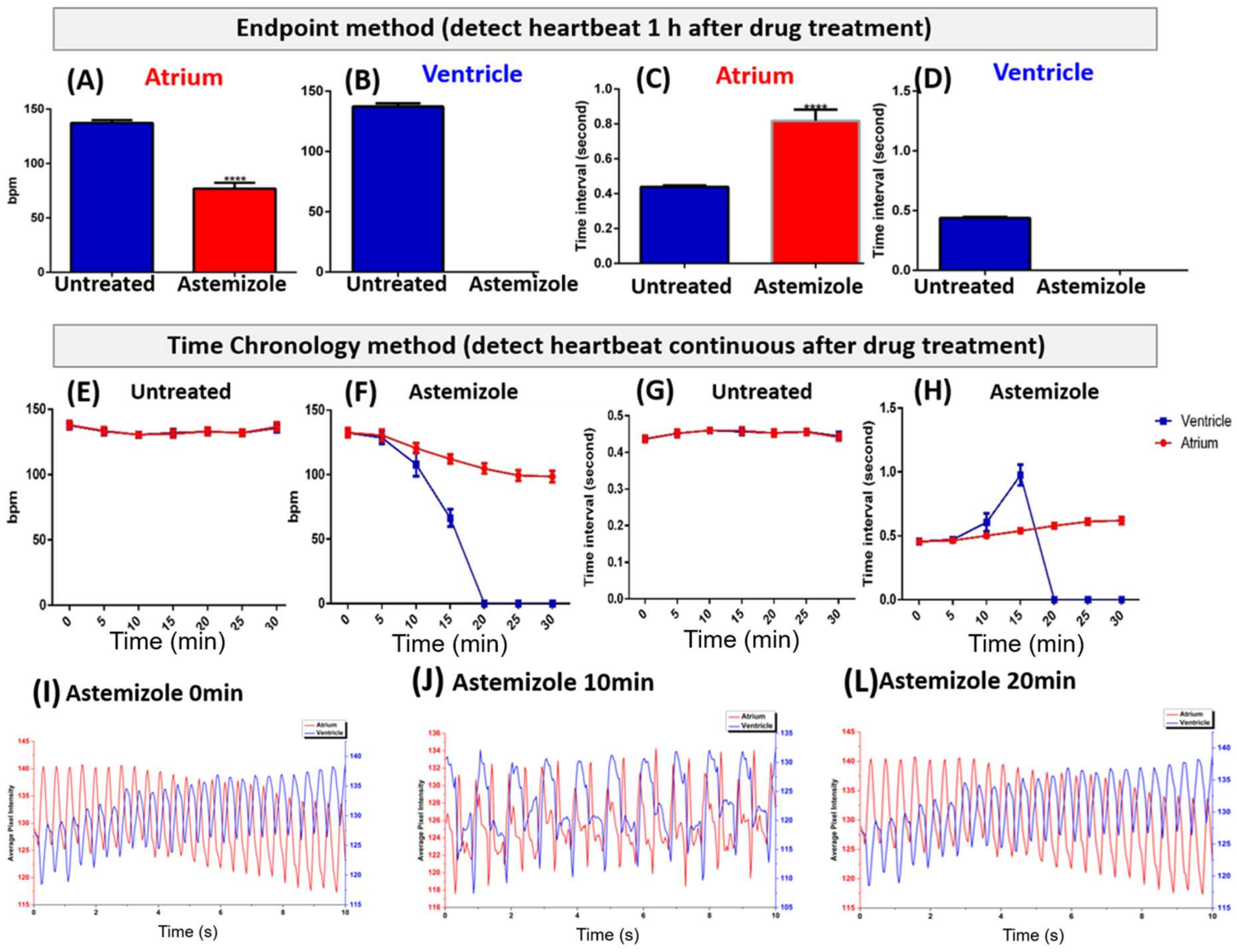
3.2. Endpoint Method of Astemizole Incubation
3.3. Time Chronology Method of Astemizole
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Barrionuevo, W.; Burggren, W. O2 consumption and heart rate in developing zebrafish (Danio rerio): Influence of temperature and ambient O2. Am. J. Physiol. Regul. Integr. Comp. Physiol. 1999, 276, R505–R513. [Google Scholar] [CrossRef]
- Fishman, M.C.; Stainier, D.Y.; Breitbart, R.E.; Westerfield, M. Zebrafish: Genetic and embryological methods in a transparent vertebrate embryo. Methods Cell Boil. 1997, 52, 67–82. [Google Scholar]
- Stainier, D.Y. Zebrafish genetics and vertebrate heart formation. Nat. Rev. Genet. 2001, 2, 39–48. [Google Scholar] [CrossRef] [PubMed]
- Nemtsas, P.; Wettwer, E.; Christ, T.; Weidinger, G.; Ravens, U. Adult zebrafish heart as a model for human heart? An electrophysiological study. J. Mol. Cell. Cardiol. 2010, 48, 161–171. [Google Scholar] [CrossRef] [PubMed]
- Staudt, D.; Stainier, D. Uncovering the molecular and cellular mechanisms of heart development using the zebrafish. Ann. Rev. Genet. 2012, 46, 397–418. [Google Scholar] [CrossRef] [PubMed]
- Milan, D.J.; Peterson, T.A.; Ruskin, J.N.; Peterson, R.T.; MacRae, C.A. Drugs that induce repolarization abnormalities cause bradycardia in zebrafish. Circulation 2003, 107, 1355–1358. [Google Scholar] [CrossRef] [PubMed]
- Burns, C.G. High-throughput assay for small molecules that modulate zebrafish embryonic heart rate. Nat. Chem. Boil. 2005, 1, 263–264. [Google Scholar] [CrossRef] [PubMed]
- De Luca, E. ZebraBeat: A flexible platform for the analysis of the cardiac rate in zebrafish embryos. Sci. Rep. 2014, 4, 4898. [Google Scholar] [CrossRef]
- Lai, Y.-C.; Chang, W.-T.; Lin, K.-Y.; Liau, I. Optical assessment of the cardiac rhythm of contracting cardiomyocytes in vitro and a pulsating heart in vivo for pharmacological screening. Biomed. Opt. Express 2014, 5, 1616–1625. [Google Scholar] [CrossRef] [PubMed]
- Chan, P.K.; Lin, C.C.; Cheng, S.H. Noninvasive technique for measurement of heartbeat regularity in zebrafish (Danio rerio) embryos. BMC Biotechnol. 2009, 9, 11. [Google Scholar] [CrossRef] [PubMed]
- Skinner, J.R.; Sharland, G. Detection and management of life threatening arrhythmias in the perinatal period. Early Hum. Dev. 2008, 84, 161–172. [Google Scholar] [CrossRef] [PubMed]
- Langheinrich, U.; Vacun, G.; Wagner, T. Zebrafish embryos express an orthologue of HERG and are sensitive toward a range of QT-prolonging drugs inducing severe arrhythmia. Toxicol. Appl. Pharmacol. 2003, 193, 370–382. [Google Scholar] [CrossRef] [PubMed]
- McGrath, P.; Li, C.-Q. Zebrafish: A predictive model for assessing drug-induced toxicity. Drug Discov. Today 2008, 13, 394–401. [Google Scholar] [CrossRef] [PubMed]
- Avdesh, A. Regular care and maintenance of a zebrafish (Danio rerio) laboratory: An introduction. J. Vis. Exp. 2012, 69, e4196. [Google Scholar] [CrossRef] [PubMed]
- Denvir, M.A.; Tucker, C.S.; Mullins, J.J. Systolic and diastolic ventricular function in zebrafish embryos: Influence of norepenephrine, MS-222 and temperature. BMC Biotechnol. 2008, 8, 21. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.-C. Combined use of MS-222 (tricaine) and isoflurane extends anesthesia time and minimizes cardiac rhythm side effects in adult zebrafish. Zebrafish 2010, 7, 297–304. [Google Scholar] [CrossRef] [PubMed]
- Romdhani, S.; Vetter, T. Estimating 3D shape and texture using pixel intensity, edges, specular highlights, texture constraints and a prior. In Proceedings of the CVPR 2005 IEEE Computer Society Conference on Computer Vision and Pattern Recognition, San Diego, CA, USA, 20–25 June 2005; pp. 986–993. [Google Scholar]
- Pylatiuk, C. Automatic zebrafish heartbeat detection and analysis for zebrafish embryos. Zebrafish 2014, 11, 379–383. [Google Scholar] [CrossRef] [PubMed]
- Müller, I.I. Functional modeling in zebrafish demonstrates that the atrial-fibrillation-associated gene GREM2 regulates cardiac laterality, cardiomyocyte differentiation and atrial rhythm. Dis. Models Mech. 2013, 6, 332–341. [Google Scholar] [CrossRef] [PubMed]
- Van Cauwenberge, P. Consensus statement on the treatment of allergic rhinitis. Allergy 2000, 55, 116–134. [Google Scholar] [CrossRef] [PubMed]
- Fung, M. Evaluation of the characteristics of safety withdrawal of prescription drugs from worldwide pharmaceutical markets—1960 to 1999. Drug Inf. J. 2001, 35, 293–317. [Google Scholar] [CrossRef]
- Olasińska-Wiśniewska, A.; Olasiński, J.; Grajek, S. Cardiovascular safety of antihistamines. Adv. Dermatol. Allergol. Postęp. Dermatol. I Alergol. 2014, 31, 182. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Vorperian, V.R.; Gong, Q.; Zhang, S.; January, C.T. Block of HERG potassium channels by the antihistamine astemizole and its metabolites desmethylastemizole and norastemizole. J. Cardiovasc. Electrophysiol. 1999, 10, 836–843. [Google Scholar] [CrossRef] [PubMed]
- Craft, T.M. Torsade de pointes after astemizole overdose. Br. Med. J. 1986, 292, 660. [Google Scholar] [CrossRef]
- Yozzo, K.L.; Isales, G.M.; Raftery, T.D.; Volz, D.C. High-content screening assay for identification of chemicals impacting cardiovascular function in zebrafish embryos. Environ. Sci. Technol. 2013, 47, 11302–11310. [Google Scholar] [CrossRef] [PubMed]
- Fink, M. A new method for detection and quantification of heartbeat parameters in Drosophila, zebrafish, and embryonic mouse hearts. Biotechniques 2009, 46, 101–103. [Google Scholar] [CrossRef] [PubMed]
- Lu, F.; Langenbacher, A.; Chen, J.-N. Transcriptional Regulation of Heart Development in Zebrafish. J. Cardiovasc. Dev. Dis. 2016, 3, 14. [Google Scholar] [CrossRef] [PubMed]
- Nerbonne, J.M.; Kass, R.S. Molecular physiology of cardiac repolarization. Physiol. Rev. 2005, 85, 1205–1253. [Google Scholar] [CrossRef] [PubMed]
- Milan, D.J.; Jones, I.L.; Ellinor, P.T.; MacRae, C.A. In vivo recording of adult zebrafish electrocardiogram and assessment of drug-induced QT prolongation. Am. J. Physiol. Heart Circ. Physiol. 2006, 291, H269–H273. [Google Scholar] [CrossRef] [PubMed]
- Roden, D.M. Drug-Induced Prolongation of the QT Interval. New Engl. J. Med. 2004, 350, 1013–1022. [Google Scholar] [CrossRef] [PubMed]
- Devarasetty, M. Optical Tracking and Digital Quantification of Beating Behavior in Bioengineered Human Cardiac Organoids. Biosensors 2017, 7, 24. [Google Scholar] [CrossRef] [PubMed]
- Roden, D.M.; Viswanathan, P.C. Genetics of acquired long QT syndrome. J. Clin. Investig. 2005, 115, 2025–2032. [Google Scholar] [CrossRef] [PubMed]
- Chiu, P.J. Validation of a [3H] astemizole binding assay in HEK293 cells expressing HERG K+ channels. J. Pharmacol. Sci. 2004, 95, 311–319. [Google Scholar] [CrossRef] [PubMed]
- Sallam, K.; Li, Y.; Sager, P.T.; Houser, S.R.; Wu, J.C. Finding the Rhythm of Sudden Cardiac Death. New Oppor. Using Induc. Pluripotent Stem Cell Deriv. Cardiomyocytes 2015, 116, 1989–2004. [Google Scholar] [CrossRef]
- Bass, A.S.; Darpo, B.; Valentin, J.P.; Sager, P.; Thomas, K. Moving towards better predictors of drug-induced torsades de pointes. Br. J. Pharmacol. 2008, 154, 1550–1553. [Google Scholar] [CrossRef] [PubMed]
| Parameter | This Study | De Luca et al. [8] | Lai et al. [9] | Yozzo et al. [25] | Fink et al. [26] | Chan et al. [10] |
|---|---|---|---|---|---|---|
| Require special transgenic line? | No | Yes | No | Yes | No | No |
| Require special script to run software? | No | Yes | Yes | Yes | Yes | Yes |
| Major platform to calculate heartbeat regularity | ImageJ | MATLAB | Not reported in original article | MetaXpress 4.0.0.24 software (commercial) | MATLAB | A custom-made program which developed in C# (C sharp) language was used for digital motion analysis |
| Require expensive setup to capture heartbeat video? | No | Yes | Yes | Yes | Yes | No |
| Major facility to capture heartbeat images | CCD mount onto dissecting microscope | Leica TCS SP5X II confocal laser-scanning inverted microscope equipped with a tandem scanning system | Dual-beam optical reflectometry set up in inverted microscope | ImageXpress Micro (IXM) Widefield High-Content Screening | Hamamatsu EM-CCD digital camera mounted on Leica DM-LFSA microscope | CCD mount onto dissecting microscope |
| Position to capture heartbeat information | Heart | Heart | Heart | Heart | Heart | Caudal blood vessel |
| What kind of message can be obtained? | Atrium and ventricle rhythm, whole heartbeat frequency | Atrium and ventricle rhythm, whole heartbeat frequency | Atrium and ventricle rhythm, whole heartbeat frequency | Whole heartbeat frequency | Atrium and ventricle rhythm, whole heartbeat frequency | Whole heartbeat frequency |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sampurna, B.P.; Audira, G.; Juniardi, S.; Lai, Y.-H.; Hsiao, C.-D. A Simple ImageJ-Based Method to Measure Cardiac Rhythm in Zebrafish Embryos. Inventions 2018, 3, 21. https://doi.org/10.3390/inventions3020021
Sampurna BP, Audira G, Juniardi S, Lai Y-H, Hsiao C-D. A Simple ImageJ-Based Method to Measure Cardiac Rhythm in Zebrafish Embryos. Inventions. 2018; 3(2):21. https://doi.org/10.3390/inventions3020021
Chicago/Turabian StyleSampurna, Bonifasius Putera, Gilbert Audira, Stevhen Juniardi, Yu-Heng Lai, and Chung-Der Hsiao. 2018. "A Simple ImageJ-Based Method to Measure Cardiac Rhythm in Zebrafish Embryos" Inventions 3, no. 2: 21. https://doi.org/10.3390/inventions3020021
APA StyleSampurna, B. P., Audira, G., Juniardi, S., Lai, Y.-H., & Hsiao, C.-D. (2018). A Simple ImageJ-Based Method to Measure Cardiac Rhythm in Zebrafish Embryos. Inventions, 3(2), 21. https://doi.org/10.3390/inventions3020021