A Genotype-Independent, Simple, Effective and Efficient in Planta Agrobacterium-Mediated Genetic Transformation Protocol
Abstract
:1. Introduction
2. General Procedure of the Protocol
2.1. Plant Material
2.2. Vector for Transformation
2.3. Preparation of Agrobacterium Culture for the Transformation
2.3.1. Inoculum Preparation from Stock Culture (Day 1) • Timing 24 h
- Streak the stock culture initially on LB agar plates containing the appropriate antibiotic(s).
- Incubate the plates overnight at 28 °C(?) (TROUBLESHOOTING, Table 1).
2.3.2. Inoculum Culture for Further Studies (Day 2) • Timing 24 h
- Prepare the inoculum culture for further studies in LB broth wherein a single colony from the LB agar plate is inoculated in 10 mL LB broth containing the appropriate antibiotic(s).
- Incubate LB broth with the Agrobacterium inoculum at 28 °C and 120 rpm in an orbital shaking incubator overnight.
- Record O.D. of the culture at a wavelength of 600 nm(?) (TROUBLESHOOTING).
2.4. Agrobacterium-Mediated Transformation (Day 3)
- Surface sterilize the seeds with 0.1% HgCl2 solution for 10 min. Wash the seeds with sterilized distilled water multiple times to eliminate any traces of mercuric chloride. • Timing: 10–15 min.
- After sterilization and washing, incubate the seeds in the Agrobacterium culture (O.D. = 0.6) overnight in an orbital shaker at 100 rpm. • Timing: 24 h.
- Wash the incubated seeds with distilled water and transfer to potted soil(?) (TROUBLESHOOTING).
2.5. Screening of Putative Transgenic Plants for Presence of the Transgene
Isolation of Genomic DNA from Putative Transgenic Plants (2 Days)
2.6. Screening of Putative Transgenic Plants for Presence of the Transgene
2.6.1. Screening by PCR
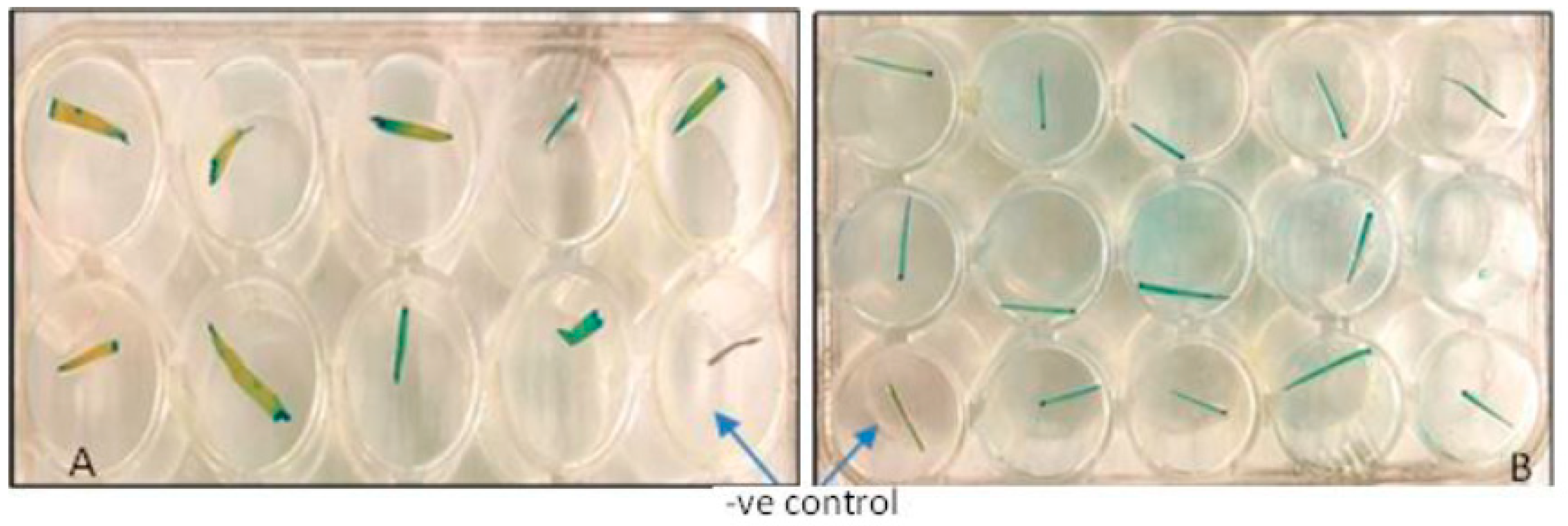
2.6.2. Screening of Putative Transgenic Plants by Expression of the Reporter Gene
2.7. Detection of Integration and Copy Number of the Transgene in Transgenic Plants (3 Days)
2.7.1. Perform Southern Hybridization for Detection of Integration and Copy Number of the Transgene in the T0 Transgenic Plants Using the Approach Given in [16]
2.7.2. Determination of Transgene Copy Number Using Real-Time PCR
2.8. Evaluation of Transgene Efficacy in Transgenic Plants
3. Results
3.1. Plant Material
3.2. Vector for Transformation
3.3. Preparation of Agrobacterium Culture for the Transformation
3.3.1. Inoculum Preparation from Stock Culture
3.3.2. Inoculum Culture for Further Studies
3.4. Agrobacterium-Mediated Transformation
3.5. Screening of Putative Transgenic Plants for Presence of the Transgene
3.6. Determination of Copy Number by Southern Hybridization and Real-Time PCR
3.7. Inheritance Pattern of Transgene in T1 Generation
3.8. Evaluation of Transgene Efficacy in Transgenic Plants
3.8.1. Evaluation of Transgene Efficacy for Insect Resistance in Transgenic Pigeon Pea
3.8.2. Evaluation of Transgene Efficacy for Insect Resistance in Transgenic Chick Pea Lines
3.8.3. Evaluation of Transgene Efficacy for Salt Stress Tolerance in Transgenic Plants
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Alves, A.C.; Quecini, V.M.; Vieira, M.L.C. Transformação de plantas: Avanços e perspectivas. Sci. Agric. 1999, 56, 1–8. [Google Scholar]
- Matsumoto, T.; Gonsalves, D.K. Biolistic and other non-Agrobacterium technologies of plant transformation. In Plant Biotechnology and Agriculture, 1st ed.; Altman, A., Hasegawa, P.W., Eds.; Academic Press: Cambridge, MA, USA, 2012; pp. 117–129. [Google Scholar]
- Jan, S.A.; Shinwari, Z.K.; Shah, S.H.; Shahzad, A.; Zia, M.A.; Ahmad, N. In-planta transformation: Recent advances. Rom. Biotechnol. Lett. 2016, 21, 11085–11091. [Google Scholar]
- Feldmann, K.A.; Marks, M.D. Agrobacterium-mediated transformation of germinating seeds of Arabidopsis thaliana: A non-tissue culture approach. Mol. Gen. Genet. MGG 1987, 208, 1–9. [Google Scholar]
- Chang, S.S.; Park, S.K.; Kim, B.C.; Kang, B.J.; Kim, D.U.; Nam, H.G. Stable genetic transformation of Arabidopsis thaliana by Agrobacterium inoculation in planta. Plant J. 1994, 5, 551–558. [Google Scholar]
- Ali, A.; Bang, S.W.; Chung, S.M.; Staub, J.E. Plant transformation via pollen tube-mediated gene transfer. Plant Mol. Biol. Rep. 2015, 33, 742–747. [Google Scholar]
- Bechtold, N.; Bouchez, D. In planta Agrobacterium-mediated transformation of adult Arabidopsis thaliana plants by vacuum infiltration. In Gene Transfer to Plants; Springer: Berlin/Heidelberg, Germany, 1995; pp. 19–23. [Google Scholar]
- Kesiraju, K.; Sreevathsa, R. Apical meristem-targeted in planta transformation strategy: An overview on its utility in crop improvement. Agric. Res. Technol. Open Access J. 2017, 8, 555734. [Google Scholar]
- Niazian, M.; Noori, S.S.; Galuszka, P.; Mortazavian, S.M.M. Tissue culture-based Agrobacterium-mediated and in planta transformation methods. Soil Water Res. 2017, 53, 133–143. [Google Scholar]
- Shou, H.; Palmer, R.G.; Wang, K. Irreproducibility of the soybean pollen-tube pathway transformation procedure. Plant Mol. Biol. Rep. 2002, 20, 325–334. [Google Scholar]
- Surekha, C.H.; Beena, M.R.; Arundhati, A.; Singh, P.K.; Tuli, R.; Dutta-Gupta, A.; Kirti, P.B. Agrobacterium-mediated genetic transformation of pigeon pea (Cajanus cajan (L.) Millsp.) using embryonal segments and development of transgenic plants for resistance against Spodoptera. Plant Sci. 2005, 169, 1074–1080. [Google Scholar]
- Sharma, K.K.; Sreelatha, G.; Dayal, S. Pigeonpea (Cajanus cajn (L.) Millsp.). Methods Mol. Biol. 2006, 343, 359–367. [Google Scholar] [CrossRef]
- Sawardekar, S.V.; Mhatre, N.K.; Sawant, S.S.; Bhave, S.G.; Gokhale, N.B.; Narangalkar, A.L.J. Agrobacterium-mediated genetic transformation of pigeon pea [Cajanus cajan (L.) Millisp] for pod borer resistance: Optimization of Protocol. Indian J. Genet. Plant Bread 2012, 72, 380–383. [Google Scholar]
- Parekh, M.J.; Mahatma, M.K.; Kansara, R.V.; Patel, D.H.; Jha, S.; Chauhan, D.A. Agrobacterium Mediated Genetic Transformation of Pigeon Pea (Cajanus cajan L. Millsp.) using Embryonic Axes for Resistance to Lepidopteron Insect. Indian J. Agric. Biochem. 2014, 27, 176–179. [Google Scholar]
- Jefferson, A.R. Assaying chimeric genes in plants: The GUS gene fusion system. Plant Mol. Biol. Rep. 1987, 5, 387–405. [Google Scholar]
- Khatodia, S.; Kharb, P.; Batra, P.; Chowdhury, V.K. Real time PCR based detection of transgene copy number in transgenic chickpea lines expressing Cry1Aa3 and Cry1 Ac. Int. J. Pure App. Biosci. 2014, 2, 100–105. [Google Scholar]
- Khatodia, S.; Kharb, P.; Batra, P.; Kumar, P.A.; Chowdhury, V.K. Molecular characterization of Bt chickpea (Cicer arietinum L.) plants carrying cry1Aa3 gene. Int. J. Curr. Microbiol. App. Sci. 2014, 3, 632–642. [Google Scholar]
- Khatodia, S.; Kharb, P.; Batra, P.; Chowdhury, V.K. Development and characterization of transgenic chickpea (Cicer arietinum L.) plants with cry1Ac gene using tissue culture independent protocol. Int. J. Adv. Res. 2014, 2, 323–331. [Google Scholar]
- Jain, M.; Khatodia, S.; Kharb, P.; Batra, P.; Chowdhury, V.K. Determination of Cry1Ac copy number in transgenic pigeonpea plants using quantitative real time PCR. Legume Res. Int. J. 2017, 40, 643–648. [Google Scholar]
- Singh, R.; Sharma, S.; Kharb, P.; Saifi, S.; Tuteja, N. OsRuvB transgene induces salt tolerance in pigeon pea. J. Plant Interact. 2020, 15, 17–26. [Google Scholar] [CrossRef]
- Pratibha. Development of Transgenic Pigeon Pea (Cajanus cajan L.) Plants Containing Lectin Receptor Like Kinase (Lec-RLK) Gene for Improving Salt Tolerance. Ph.D. Thesis, CCS Haryana Agricultural University, Hisar, Haryana, India, 2019.
- Kharb, P.N. Psp68, A Dead Box Helicase Confers Salinity Tolerance in Transgenic PigeonPea. Int. J. Curr. Microbiol. Appl. Sci. 2019, 8, 309–324. [Google Scholar]
- Preeti, P.K. Engineering chickpea variety HC-1 with OsRuvB gene for salt stress tolerance. Legume Res.-Int. J. 2020, 1, 6. [Google Scholar]
- Singh, G. Development and Characterization of Transgenic Chickpea (Cicer arietinum L.) Plants with OsLec-RLK Gene for Salt Stress Tolerance. Ph.D. Thesis, CCS Haryana Agricultural University, Hisar, Haryana, India, 2017. [Google Scholar]
- Khasa, R. Development and Characterization of Transgenic Wheat (Triticum aestivum L.) with OsRuvB Gene for Salinity Tolerance. Ph.D. Thesis, CCS Haryana Agricultural University, Hisar, Haryana, India, 2022. [Google Scholar]
- Kharb, P.; Singh, R.; Kumar, U. An Efficient Method for Transformation in Monocots. Indian Patent Application No. 201811012100, 4 October 2019. [Google Scholar]
- Kharb, P.; Batra, P.; Chowdhury, V.K. A Novel Process of Genetic Transformation in Chickpea Using Agrobacterium. Patent No. 252590, 2005. [Google Scholar]
- Kaur, R.P.; Devi, S. In planta transformation in plants: A review. Agric. Rev. 2019, 40, 159–174. [Google Scholar]
- Saifi, S.K.; Passricha, N.; Tuteja, R.; Pushpa, K.; Narendra, T. In planta transformation: A smart way of crop improvement. In Advancement in Crop Improvement Techniques; Tuteja, N., Tuteja, R., Passricha, N., Saifi, S.K., Eds.; Woodhead Publishing: Sawston, UK, 2020; pp. 351–362. ISBN 9780128185810. [Google Scholar] [CrossRef]
- Wang, K.; Shi, L.; Liang, X.; Zhao, P.; Wang, W.; Liu, J.; Chang, Y.; Hiei, Y.; Yanagihara, C.; Du, L.; et al. The gene TaWOX5 overcomes genotype dependency in wheat genetic transformation. Nat. Plants 2022, 8, 110–117. [Google Scholar] [PubMed]
- Saifi, S.K.; Passricha, N.; Tuteja, R.; Nath, M.; Gill, S.S.; Tuteja, N. OsRuvBL1a DNA Helicase Boost Salinity and Drought Tolerance in Transgenic Indica Rice Raised by In-Planta Transformation. Res. Sq. 2021. [Google Scholar] [CrossRef]
- Kesiraju, K.; Mishra, P.; Bajpai, A.; Sharma, M.; Rao, U.; Sreevathsa, R. Agrobacterium tumefaciens-mediated in planta transformation strategy for development of transgenics in cotton (Gossypium hirsutum L.) with GFP as a visual marker. Physiol. Mol. Biol. Plants 2020, 26, 2319–2327. [Google Scholar] [CrossRef]
- Tang, W.; Newton, R.J.; Weidner, D.A. Genetic transformation and gene silencing mediated by multiple copies of a transgene in eastern white pine. J. Exp. Bot. 2007, 58, 545–554. [Google Scholar] [CrossRef]
- Shou, H.; Frame, B.R.; Whitham, S.A.; Wang, K. Assessment of transgenic maize events produced by particle bombardment or Agrobacterium-mediated transformation. Mol. Breed. 2004, 13, 201–208. [Google Scholar]
- Travella, S.; Ross, S.M.; Harden, J.; Everett, C.; Snape, J.W.; Harwood, W.A. A comparison of transgenic barley lines produced by particle bombardment and Agrobacterium-mediated techniques. Plant Cell Rep. 2005, 23, 780–789. [Google Scholar] [PubMed]
- Mark, A.J.; David, J.A.; Robert, G.B. Comparison of Agrobacterium and particle bombardment using whole plasmid or minimal cassette for production of high-expressing, low-copy transgenic plants. Transgenic Res. 2013, 22, 143–151. [Google Scholar] [CrossRef]
- Li, S.; Cong, Y.; Liu, Y.; Wang, T.; Shuai, Q.; Chen, N.; Gai, J.; Li, Y. Optimization of Agrobacterium-Mediated Transformation in Soybean. Front. Plant Sci. 2017, 8, 246. [Google Scholar] [CrossRef]



| S. No. | Step | Problem | Possible Reason | Solution |
|---|---|---|---|---|
| 1 | Inoculum preparation from stock culture | The culture shows no growth | The culture is not viable. | Start a new culture from a freshly plated colony. |
| 2 | Inoculum culture for further studies | Thread-like growth in the culture | The culture is contaminated. | Start the culture again and properly sterilize the inoculating needle. |
| 3 | Transformation with the gene of interest | Seeds do not germinate | Over-treatment with HgCl2. | Strictly follow the time and amount mentioned. |
| 4 | Isolation of genomic DNA of putative transgenic plants | Sharp and clear DNA bands are not visible | The DNA might be sheared. | Follow the given protocol without any modifications. |
| 5 | Screening of putative transgenic plants carrying the transgene | No bands or smeared patterns or multiple amplification products | The amplification did not take place or there is some problem with the primer binding. | Check for the annealing temperature based on primers and adjust it accordingly. Do not go too low to avoid non-specific binding of the primer. |
| Sr. No. | Agrobacterium tumefaciens Strain | Crop Modified | Trait | Transformation Efficiency (%) | Reference |
|---|---|---|---|---|---|
| 1 | EHA105 harboring pBinAR-35Scry1Aa3 | Chickpea cv. C-235 | Insect resistance | 18.1 (6/33) | [17] |
| 2 | LBA4404 harboring pBinAR-35Scry1Ac | Chickpea cv. C-235 | Insect resistance | 14.3 (25/174) | [18] |
| 3 | LBA4404 harboring pBinAR-35Scry1Ac | Chickpea cv. HC-1 | Insect resistance | 40.9 (18/44) | [18] |
| 4 | LBA4404 harboring pBinAR-35Scry1Ac | Pigeon pea cv. Manak | Insect resistance | 45.0 (9/20) | [19] |
| 5 | LBA4404 harboring pCAMBIA 1301-OsRuvB gene | Pigeon pea cv. Manak | Salt tolerance | 35.7 (25/70) | [20] |
| 6 | LBA4404 harboring pCAMBIA 1301- OsLec-RLK | Pigeon pea cv. Manak | Salt tolerance | 18.6 (16/86) | [21] |
| 7 | LBA4404 containing pCAMBIA1300 harboring Psp68 | Pigeon pea cv. Manak | Salt tolerance | 16.0 (16/100) | [22] |
| 8 | LBA4404 harboring pCAMBIA 1301-OsRuvB gene | Chickpea cv. HC-1 | Salt tolerance | 17.0 (17/100) | [23] |
| 9 | LBA4404 harboring pCAMBIA 1301-OsLec-RLK | Chickpea cv. HC-1 | Salt tolerance | 17.8 (18/101) | [24] |
| 10 | LBA4404 harboring pCAMBIA 1301-OsRuvB gene | Wheat var. WH1184 | Salt tolerance | 27.0 (26/96) | [25] |
| 11 | LBA4404 harboring pCAMBIA 1301-OsRuvB gene | Wheat var. 1105 | -- | 58.9 (33/56) | [26] |
| 12 | LBA4404 harboring pCAMBIA 1301-OsRuvB gene | Indica rice | -- | 93.8 (45/49) | [26] |
| S. No. | Agrobacterium tumefaciens Strain | Crop Modified | Generation Analyzed | Restriction Endonuclease Used for Digestion of Genomic DNA | Probe Used for Detection of the Transgene Integration | Panel in Figure 2 | Reference |
|---|---|---|---|---|---|---|---|
| 1 | LBA4404 harboring pBinAR-35Scry1Ac | Chickpea cv. HC-1 | T2 | Hind III | PCR-amplified fragment of cry1Ac gene | Figure 2F | [18] |
| 2 | LBA4404 harboring pBinAR-35Scry1Ac | Pigeon pea cv. Manak | T1 | Hind III | PCR-amplified fragment of cry1Ac gene | Figure 2E | [19] |
| 3 | LBA4404 harboring pCAMBIA 1301-OsRuvB gene | Pigeon pea cv. Manak | T0 | EcoRI | PCR-amplified fragment of OsRuvB gene | Figure 2B | [20] |
| 4 | LBA4404 harboring pCAMBIA 1301- OsLec-RLK | Pigeon pea cv. Manak | T0 | Kpn1 | PCR-amplified fragment of OsLec-RLK gene | Figure 2G | [21] |
| 5 | LBA4404 containing pCAMBIA1300 harboring Psp68 | Pigeon pea cv. Manak | T0 | EcoRI | Figure 2A | [22] | |
| 6 | LBA4404 harboring pCAMBIA 1301-OsRuvB gene | Chickpea cv. HC-1 | T0 | EcoRI | PCR-amplified fragment of OsRuvB gene | Figure 2D | [23] |
| 7 | LBA4404 harboring pCAMBIA 1301- OsLec-RLK | Chickpea cv. HC-1 | T1 | Kpn1 | Figure 2C | [24] | |
| 8 | LBA4404 harboring pCAMBIA 1301-OsRuvB gene | Wheat var. WH1184 | T0 | EcoRI | PCR-amplified fragment of OsRuvB gene | Figure 2H | [25] |
| S. No. | Line | No. of T1 Plants Screened | OsRuvB +ve | OsRuvB −ve | Observed Ratio | χ2-Value | p-Value |
|---|---|---|---|---|---|---|---|
| 1 | L-10 | 43 | 31 | 12 | 2.6:1 | 0.21 | 0.64 |
| 2 | L-17 | 42 | 32 | 10 | 3.2:1 | 0.12 | 0.72 |
| 3 | L-32 | 47 | 34 | 13 | 2.6:1 | 0.10 | 0.75 |
| 4 | L-37 | 50 | 36 | 14 | 2.6:1 | 0.09 | 0.76 |
| 5 | L-66 | 45 | 33 | 12 | 2.8:1 | 0.11 | 0.74 |
| 6 | L-107 | 36 | 33 | 3 | 11:1 | 0.52 | 0.47 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kharb, P.; Chaudhary, R.; Tuteja, N.; Kaushik, P. A Genotype-Independent, Simple, Effective and Efficient in Planta Agrobacterium-Mediated Genetic Transformation Protocol. Methods Protoc. 2022, 5, 69. https://doi.org/10.3390/mps5050069
Kharb P, Chaudhary R, Tuteja N, Kaushik P. A Genotype-Independent, Simple, Effective and Efficient in Planta Agrobacterium-Mediated Genetic Transformation Protocol. Methods and Protocols. 2022; 5(5):69. https://doi.org/10.3390/mps5050069
Chicago/Turabian StyleKharb, Pushpa, Rinku Chaudhary, Narendra Tuteja, and Prashant Kaushik. 2022. "A Genotype-Independent, Simple, Effective and Efficient in Planta Agrobacterium-Mediated Genetic Transformation Protocol" Methods and Protocols 5, no. 5: 69. https://doi.org/10.3390/mps5050069
APA StyleKharb, P., Chaudhary, R., Tuteja, N., & Kaushik, P. (2022). A Genotype-Independent, Simple, Effective and Efficient in Planta Agrobacterium-Mediated Genetic Transformation Protocol. Methods and Protocols, 5(5), 69. https://doi.org/10.3390/mps5050069







