In Vitro Screening Method for Characterization of Macrophage Activation Responses
Abstract
:1. Introduction
2. Materials
3. Procedure
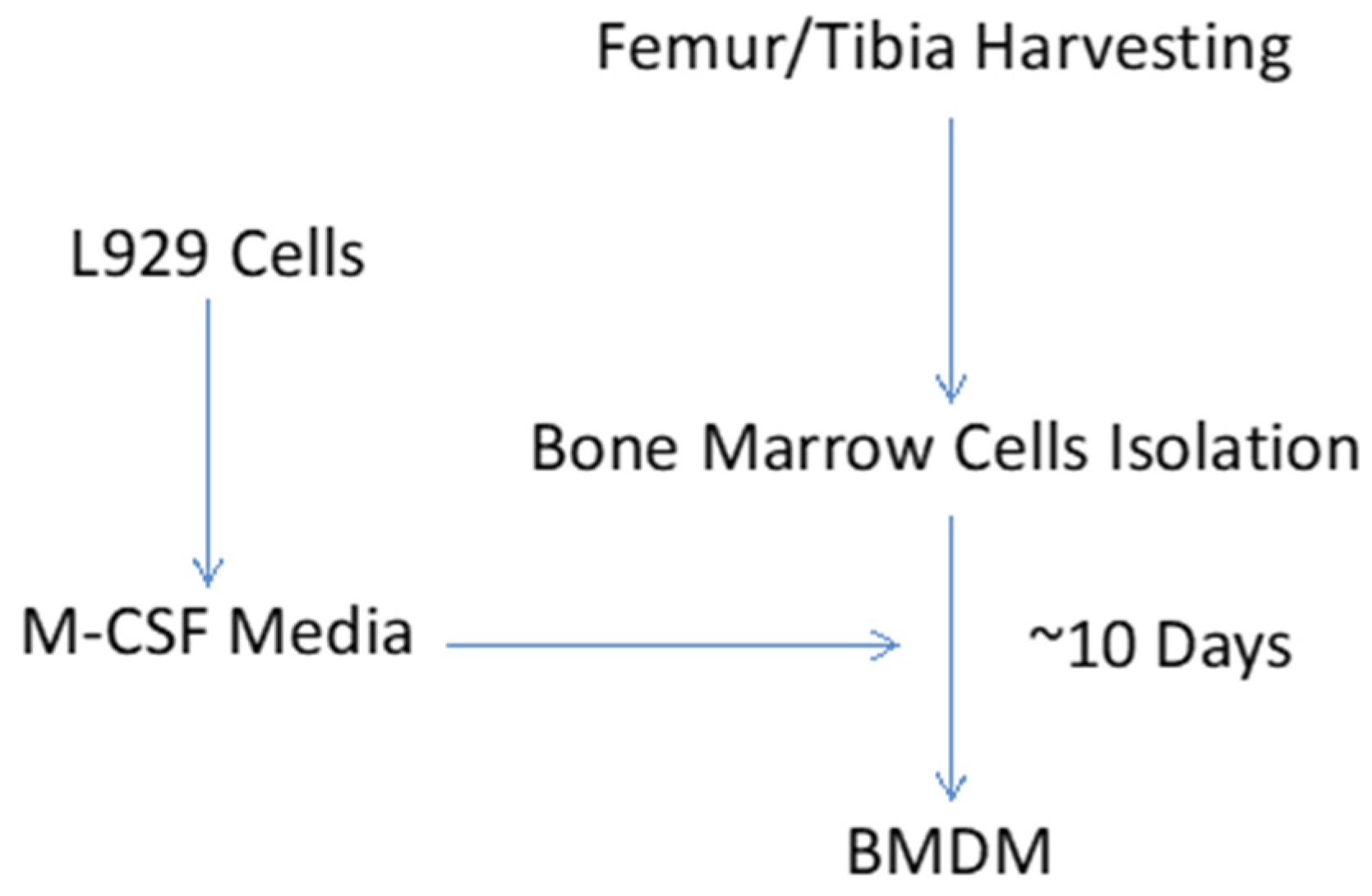
3.1. Generation of Macrophage-Colony Stimulating Factor (M-CSF) Containing L929 Medium
- (1)
- Plate 106 L929 cells (M-CSF producing cell line, CCL-1, was purchased from ATCC) in 20 mL of DMEM (Gibco, # 11995-065) supplemented with 10% heat inactivated fetal bovine serum (FBS), and 1% penicillin/streptomycin (Lonza, #17-602E) in a T75 cell culture flask.
- (2)
- Grow cells for one week (or until the flask is ~90% confluent) at 37°C, 5% CO2. It is important to keep the seeding density and harvesting schedule consistent to avoid variation in L929 concentration is the L929 medium.
- (3)
- Collect the medium and spin at 400× g for 5 min to remove any floating cells and cellular debris.
- (4)
- Filter the supernatant through a 0.22 μm filter. Store the filtered L-929 medium in 150 mL aliquots at −20 °C.
- (5)
- L929-conditioned macrophage media is then prepared by mixing L-929 medium (30%) and RPMI 1640 (Caisson Labs, #RPL09-500 mL) complete medium (70%).
3.2. Collection of Bone Marrow Cells
- (1)
- Euthanize six-week-old male mice on C57BL/6J background by CO2 inhalation and cervical dislocation. Disinfect euthanized mice with 70% ethanol and betadine (this step can be performed on bench-top).
- (2)
- Using sterile surgical instruments, isolate femur and tibial bones, and transfer to 100 mm cell culture dish containing 5 mL of RPMI 1640 complete media (10% FBS, and 1% pen/strep).
- (3)
- Transfer cell culture dish containing bones to cell culture hood for further processing. Using new set of surgical instruments, cut bones from both ends to expose the marrow cavity. Flush the bone marrow into new cell culture dish with 10 mL syringe (containing RPMI 1640 complete media) fitted with 22G needle.
- (4)
- Dissociate bone marrow with repeated passing through 25G needle. Strain the cell suspension through 70 μm cell strainer in to 50 mL sterilized conical tube.
- (5)
- Spin the cell suspension at 400× g for 5 min. Resuspend the cell pellet in 20 mL of L929-conditioned macrophage medium and plate in T25 cell culture flask for overnight incubation. Next morning (~12 h wait), replate non-adherent cells from T25 flask into 100 mm petri dishes (20 dishes per mouse) for differentiation of bone marrow cells to macrophages.
- (6)
- Add 3 mL fresh L929-conditioned macrophage medium every day. Switch L929-conditioned macrophage medium at 3-day intervals (7 mL per dish). Collect adherent cells (mature macrophages) on 10th day of culture and proceed to in vitro macrophage activation step.
3.3. BMDM Macrophage Activation
- (1)
- Seed BMDMs in designated wells (seeding density: 40,000 per well in 200 μL volume) of 96-well cell culture plate. Incubate the plate at 37 °C for 1 h.
- (2)
- To prevent the likely influence of serum constituents on the macrophage responsiveness at baseline, replace the media with serum-free RPMI 1640 complete medium, and incubate the plate for overnight serum starvation. Of note, in case of possible loss of viability of cells due to serum-free environment, RPMI 1640 complete medium with 1–2% serum could be used.
- (3)
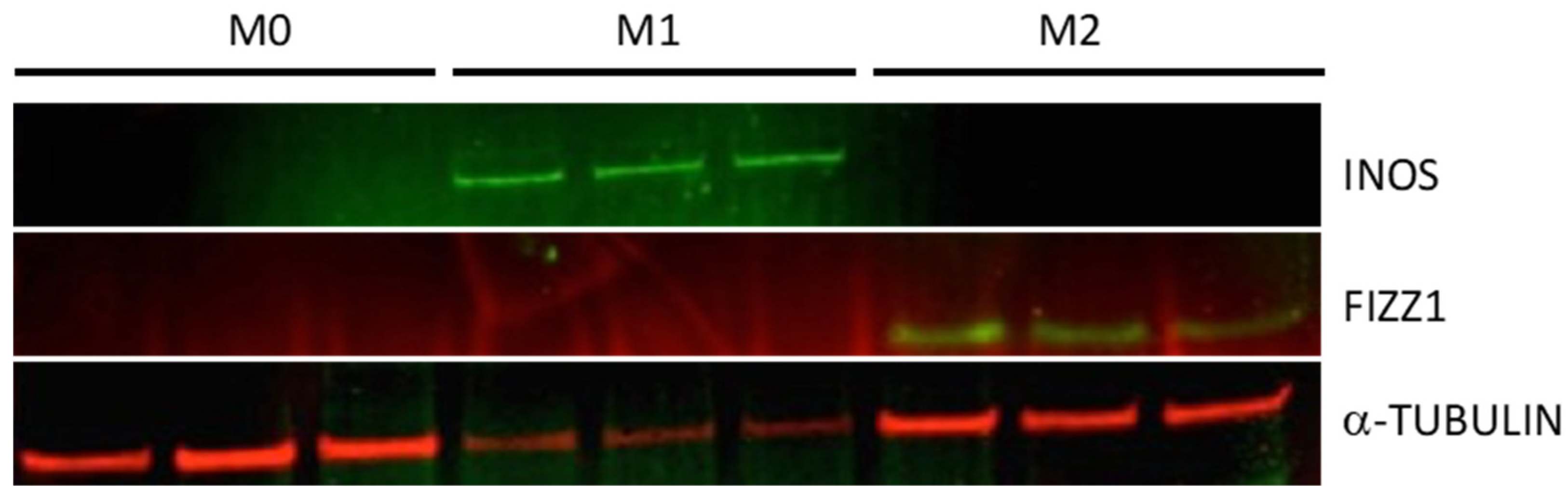
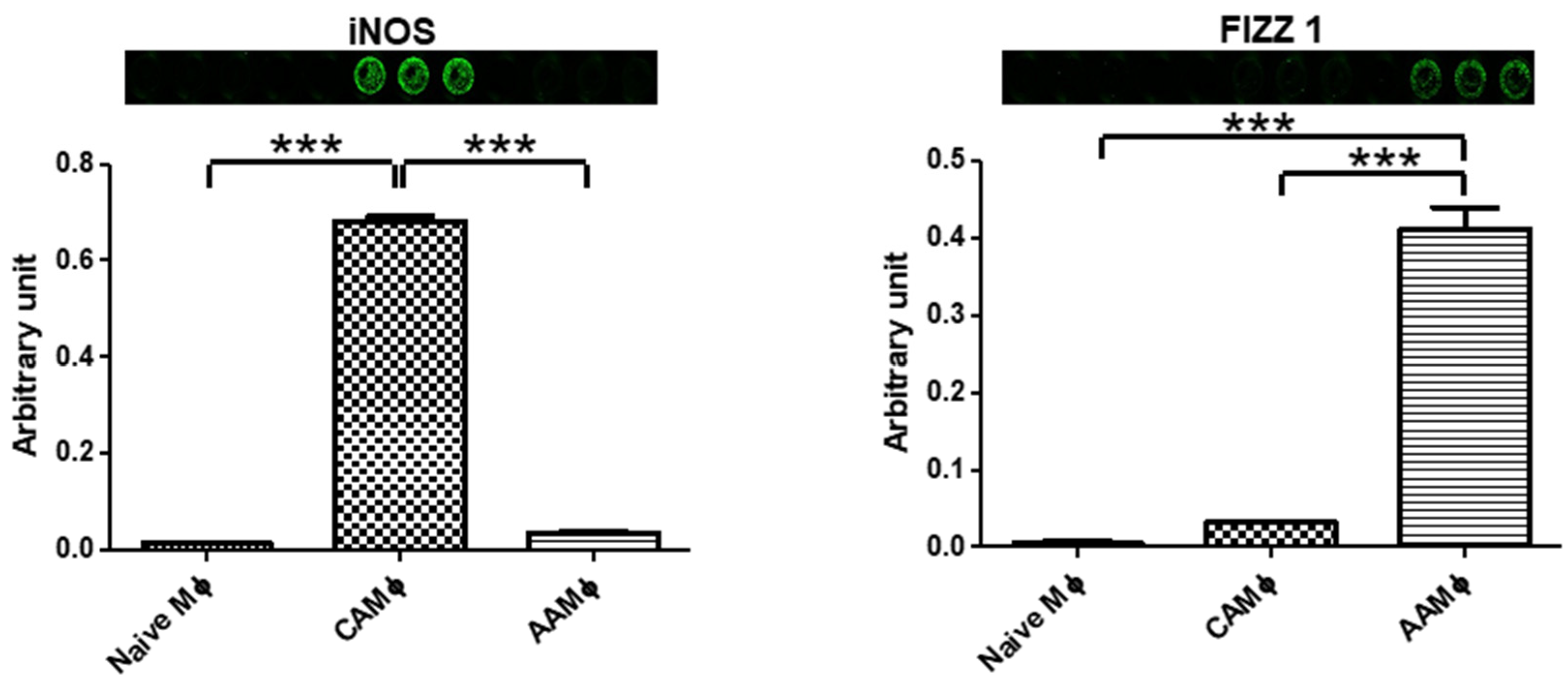
- Next day, label wells for M1, M2, and M0 (non-activated). Replace media (24 wells/treatment) with fresh serum-free RPMI 1640 complete medium containing IFN-γ (215 U/mL) + LPS (10 ng/mL; added 8 h after the start of IFN-γ treatment) for M1 activation or IL-4 (20 U/mL) for M2 activation. Wells assigned to M0 group will receive fresh serum- free RPMI1640 complete medium with no additives [8].
3.4. Screening for Macrophage Activation Markers
- (1)
- Wash once with 1× PBS buffer and fix with 100 μL of 10% neutral buffered formalin (Fisher Scientific) for 20 min, at room temperature. Use of multichannel pipette is highly recommended. To avoid dislodgment of cells, it is important to add buffers/solutions alongside the walls.
- (2)
- Centrifuge the 96 well plate at 500× g for 5 min. Aspirate the formalin fixative and wash cells once with 1× PBS.
- (3)
- Wash cells three times (5 min for each wash) with 0.1% Triton X-100 PBS (1× PBSTr) to permeabilize cells.
- (4)
- Wash cells once with 1× PBS. Add 200 µL of LI-COR Odyssey blocking buffer per well to block cells. Incubate on a rocker for 1 h at room temperature.
- (5)
- Add 50 μL of primary antibody solution (prepared in Odyssey Blocking Buffer) to each well and incubate overnight on rocker in a cold room.
- (6)
- Wash the plate three times with 0.1% Tween-20 PBS (1× PBSTw) for 5 min at room temperature.
- (7)
- Incubate with 50 μL of goat anti rabbit IRDYE 800CW secondary antibody solution (1:1000) for 1 h at room temperature. At this step, include DNA labeling dye (1:10,000 DRAQ5 stain) for normalizing cell numbers in each well.
- (8)
- Wash with 1× PBSTw for 5 min, three times.
- (9)
- Wash with 1× PBS for 5 min. Discard wash solution and dry the plate on paper towels.
- (10)
- Scan the plate in both the 700 nm and 800 nm detection channels using Odyssey Imager (Odyssey CLx Infrared Imaging System, Lincoln, NE). Normalize the primary antibody signal by dividing value obtained on 800 nm channel with values obtained on 700 nm channel for each well.
4. Results
5. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
References
- Hussell, T.; Bell, T.J. Alveolar macrophages: Plasticity in a tissue-specific context. Nat. Rev. Immunol. 2014, 14, 81–93. [Google Scholar] [CrossRef] [PubMed]
- Mosser, D.M.; Edwards, J.P. Exploring the full spectrum of macrophage activation. Nat. Rev. Immunol. 2008, 8, 958–969. [Google Scholar] [CrossRef] [PubMed]
- Glass, C.K.; Natoli, G. Molecular control of activation and priming in macrophages. Nat. Immunol. 2016, 17, 26–33. [Google Scholar] [CrossRef] [PubMed]
- Bain, C.C.; MacDonald, A.S. The impact of the lung environment on macrophage development, activation and function: Diversity in the face of adversity. Mucosal. Immunol. 2022, 15, 223–234. [Google Scholar] [CrossRef] [PubMed]
- Bleriot, C.; Chakarov, S.; Ginhoux, F. Determinants of Resident Tissue Macrophage Identity and Function. Immunity 2020, 52, 957–970. [Google Scholar] [CrossRef] [PubMed]
- Murray, P.J.; Wynn, T.A. Protective and pathogenic functions of macrophage subsets. Nat. Rev. Immunol. 2011, 11, 723–737. [Google Scholar] [CrossRef] [PubMed]
- Gordon, S.; Taylor, P.R. Monocyte and macrophage heterogeneity. Nat. Rev. Immunol. 2005, 5, 953–964. [Google Scholar] [CrossRef] [PubMed]
- Mosser, D.M.; Zhang, X. Activation of murine macrophages. Curr. Protoc. Immunol. 2008, 83, 14.2.1–14.2.8. [Google Scholar] [CrossRef] [PubMed]
- Edin, S.; Wikberg, M.L.; Dahlin, A.M.; Rutegard, J.; Oberg, A.; Oldenborg, P.A.; Palmqvist, R. The distribution of macrophages with a M1 or M2 phenotype in relation to prognosis and the molecular characteristics of colorectal cancer. PLoS ONE 2012, 7, e47045. [Google Scholar] [CrossRef]
- Abdelaziz, M.H.; Abdelwahab, S.F.; Wan, J.; Cai, W.; Huixuan, W.; Jianjun, C.; Kumar, K.D.; Vasudevan, A.; Sadek, A.; Su, Z.; et al. Alternatively activated macrophages; a double-edged sword in allergic asthma. J. Transl. Med. 2020, 18, 58. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yao, Y.; Xu, X.H.; Jin, L. Macrophage Polarization in Physiological and Pathological Pregnancy. Front. Immunol. 2019, 10, 792. [Google Scholar] [CrossRef] [PubMed]
- Raes, G.; Noel, W.; Beschin, A.; Brys, L.; de Baetselier, P.; Hassanzadeh, G.H. FIZZ1 and Ym as tools to discriminate between differentially activated macrophages. Dev. Immunol. 2002, 9, 151–159. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.X.; Zhang, S.X.; Wu, H.J.; Rong, X.L.; Guo, J. M2b macrophage polarization and its roles in diseases. J. Leukoc. Biol. 2019, 106, 345–358. [Google Scholar] [CrossRef] [PubMed]
- Arora, S.; Dev, K.; Agarwal, B.; Das, P.; Syed, M.A. Macrophages: Their role, activation and polarization in pulmonary diseases. Immunobiology 2018, 223, 383–396. [Google Scholar] [CrossRef] [PubMed]
- Saini, Y.; Dang, H.; Livraghi-Butrico, A.; Kelly, E.J.; Jones, L.C.; O’Neal, W.K.; Boucher, R.C. Gene expression in whole lung and pulmonary macrophages reflects the dynamic pathology associated with airway surface dehydration. BMC Genom. 2014, 15, 726. [Google Scholar] [CrossRef] [PubMed]
- Choudhary, I.; Vo, T.; Paudel, K.; Patial, S.; Saini, Y. Compartment-specific transcriptomics of ozone-exposed murine lungs reveals sex- and cell type-associated perturbations relevant to mucoinflammatory lung diseases. Am. J. Physiol. Lung Cell. Mol. Physiol. 2021, 320, L99–L125. [Google Scholar] [CrossRef] [PubMed]
- Kradin, R.L.; McCarthy, K.M.; Preffer, F.I.; Schneeberger, E.E. Flow-cytometric and ultrastructural analysis of alveolar macrophage maturation. J. Leukoc. Biol. 1986, 40, 407–417. [Google Scholar] [CrossRef] [PubMed]
- Choudhary, I.; Vo, T.; Paudel, K.; Wen, X.; Gupta, R.; Kesimer, M.; Patial, S.; Saini, Y. Vesicular and extravesicular protein analyses from the airspaces of ozone-exposed mice revealed signatures associated with mucoinflammatory lung disease. Sci. Rep. 2021, 11, 23203. [Google Scholar] [CrossRef] [PubMed]
- MacMicking, J.; Xie, Q.W.; Nathan, C. Nitric oxide and macrophage function. Annu. Rev. Immunol. 1997, 15, 323–350. [Google Scholar] [CrossRef] [PubMed]
- Raes, G.; De Baetselier, P.; Noel, W.; Beschin, A.; Brombacher, F.; Hassanzadeh Gh, G. Differential expression of FIZZ1 and Ym1 in alternatively versus classically activated macrophages. J. Leukoc. Biol. 2002, 71, 597–602. [Google Scholar] [PubMed]



| Reagent | Company/Institute | Cat. Number | Comments |
|---|---|---|---|
| C57BL/6J | Jackson Laboratory | 000664 | Mouse Strain |
| L-929 Cell line | ATCC | CCL-1 | Mouse connective tissue fibroblast cell line |
| Dulbecco’s Modified Eagle Medium (DMEM) | Gibco (Life Technologies) | 11995-065 | With Glucose, L-Glutamine and Sodium Pyruvate |
| RPMI 1640 | Gibco (Life Technologies) | 22400-089 | With L-Glutamine and HEPES |
| Fetal Bovine Serum | Atlanta Biologicals | S11550H | Heat Inactivated |
| Penicillin/Streptomycin (100X) | Sigma-Aldrich | P4333 | Working concentration (1X) |
| Phosphate Buffered Saline (PBS) | Sigma-Aldrich | P5368 | 0.01 M PBS; pH 7.5 |
| Interleukin 4 | EMD Millipore | IL016 | Recombinant (Murine) |
| Interferon Gamma (IFN-γ) | EMD Millipore | IF005 | Recombinant (Murine) |
| Lipopolysaccharide (LPS) | EMD Millipore | LPS25 | LPS, E. Coli O111:B4 |
| 96-well plates | Thermo Scientific | 165305 | Black with polymer base |
| Filtration Unit | Genesee | 25-227 | 0.22 μm |
| Betadine | Purdue Products | 67618-151-17 | Povidone-iodine, 7.5% |
| Cell strainer | Corning Life Sciences | 352350 | 70 μm |
| T25 culture flask | Genesee | 25-207 | |
| T75 culture flasks | Genesee | 25-209 | |
| 100 mm Petri-dishes | Thermo Scientific | 130182 | |
| 10% Buffered Formalin | Fisher Scientific | SF100-20 | pH 7.0 |
| Triton x-100 | Fisher Bioreagents | BP151-100 | Electrophoresis Grade |
| Tween-20 | Fisher Bioreagents | BP337-100 | Electrophoresis Grade |
| FIZZ1A | ABCAM | AB39626 | Rabbit Polyclonal |
| YM1/2 | National Institute of Health | Rabbit Polyclonal | A kind gift from Dr. Shioko Kimura |
| INOS | ABCAM | AB15326 | Rabbit Polyclonal |
| HIF2a | ABCAM | AB199 | Rabbit Polyclonal |
| COX 1 | Cell Signaling | 4841 | Rabbit Polyclonal |
| MMP12 | ABCAM | AB15326 | Rabbit Polyclonal |
| ARG1 | Santa-Cruz Biotechnology | SC-20150 | Rabbit Polyclonal |
| Odyssey Blocking Buffer | LI-COR | 927-40000 | PBS-based buffer |
| Odyssey Imager | LI-COR | Odyssey CLx | Infra-red Imager |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lewis, B.W.; Patial, S.; Saini, Y. In Vitro Screening Method for Characterization of Macrophage Activation Responses. Methods Protoc. 2022, 5, 68. https://doi.org/10.3390/mps5050068
Lewis BW, Patial S, Saini Y. In Vitro Screening Method for Characterization of Macrophage Activation Responses. Methods and Protocols. 2022; 5(5):68. https://doi.org/10.3390/mps5050068
Chicago/Turabian StyleLewis, Brandon W., Sonika Patial, and Yogesh Saini. 2022. "In Vitro Screening Method for Characterization of Macrophage Activation Responses" Methods and Protocols 5, no. 5: 68. https://doi.org/10.3390/mps5050068
APA StyleLewis, B. W., Patial, S., & Saini, Y. (2022). In Vitro Screening Method for Characterization of Macrophage Activation Responses. Methods and Protocols, 5(5), 68. https://doi.org/10.3390/mps5050068






