Recent Advances in miRNA Delivery Systems
Abstract
1. Introduction
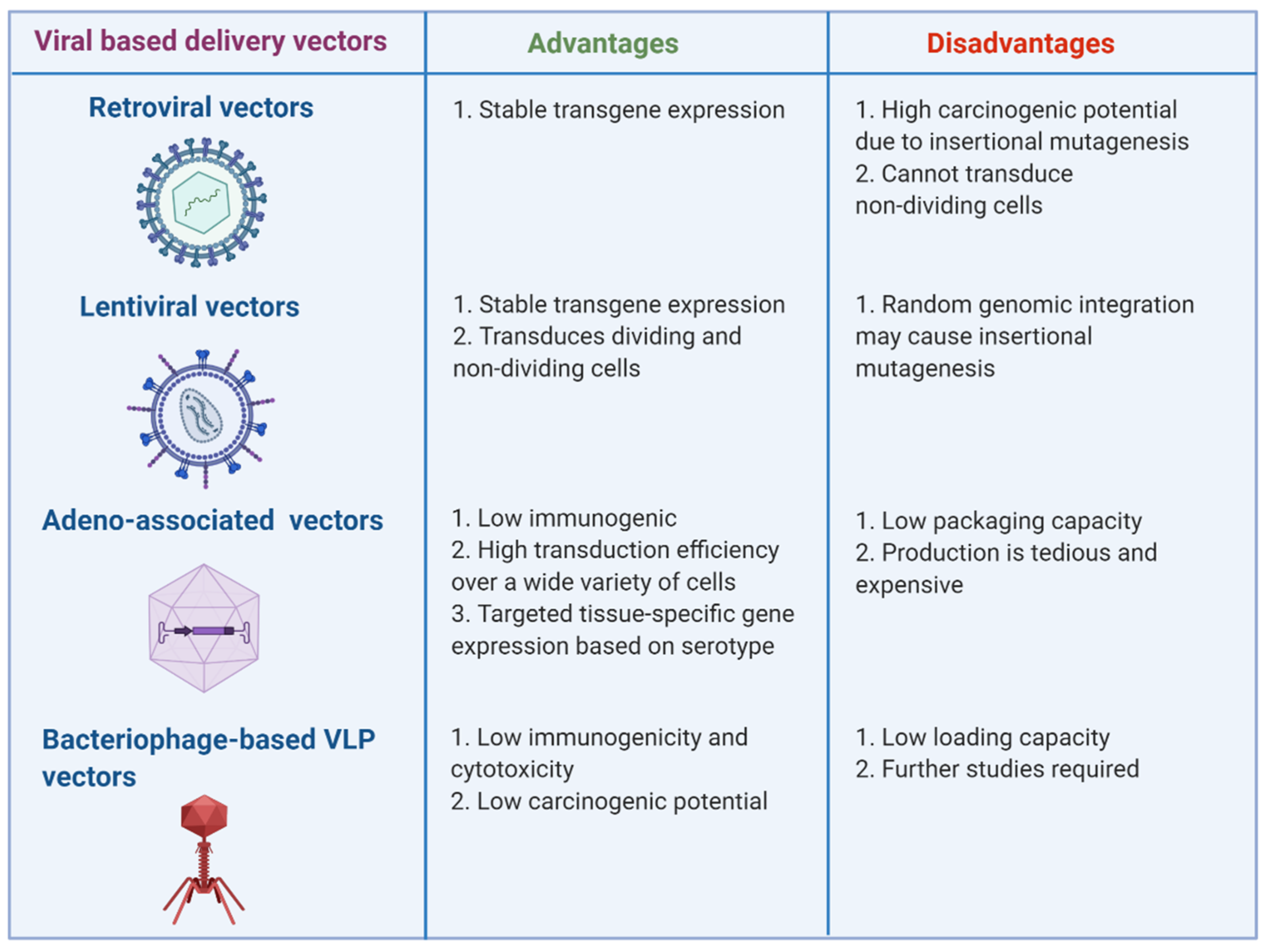
2. Virus-Based miRNA and Anti-miRNA Oligonucleotide Delivery Systems
2.1. Retroviral and Lentiviral Vectors
2.2. Adenovirus and Adeno-Associated Virus Vectors
2.3. Bacteriophage-Based VLP Vectors
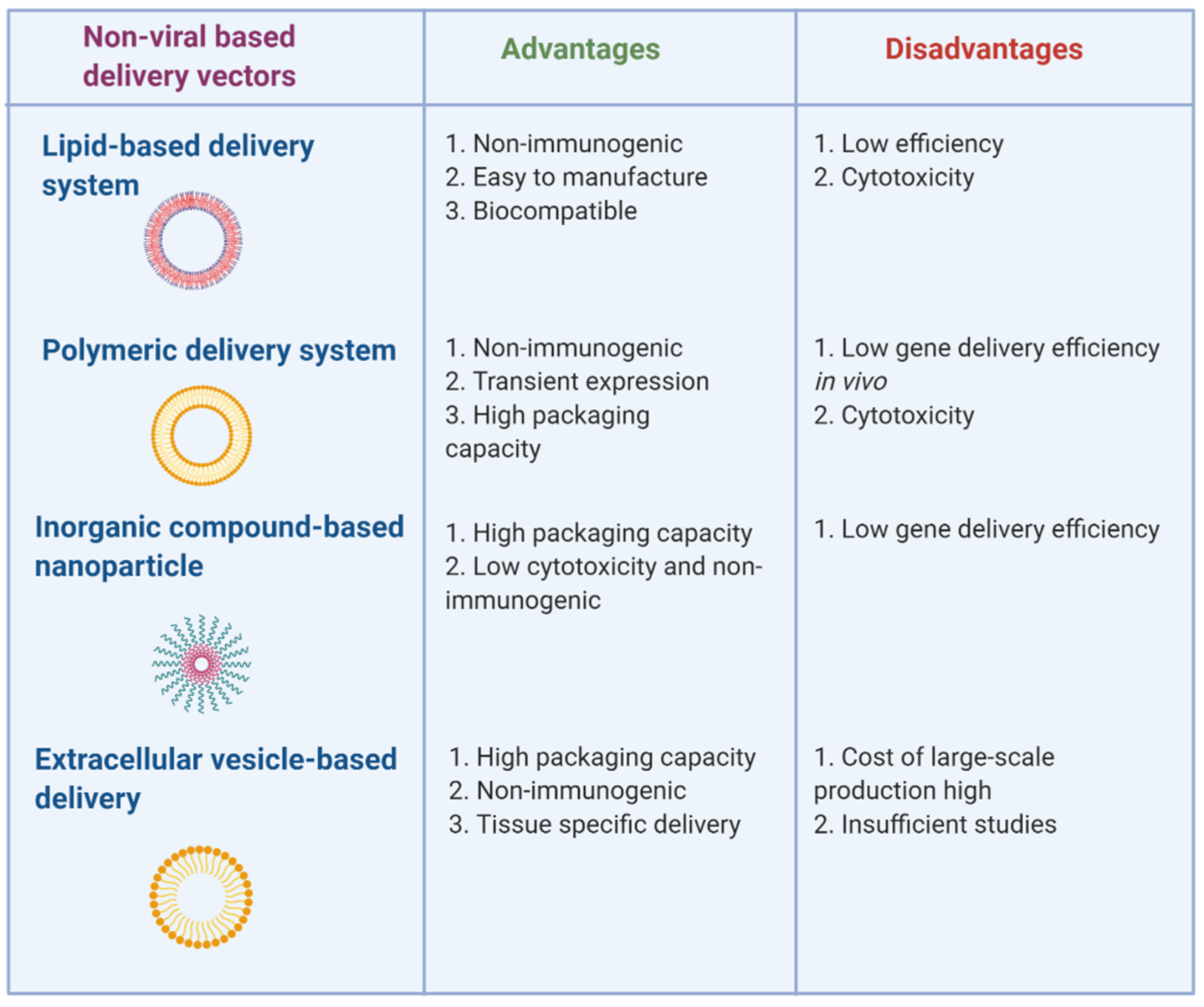
3. Non-Viral-Based miRNA and Anti-miRNA Oligonucleotide Delivery Systems
3.1. Lipid-Based Delivery Systems
3.2. Polymeric Delivery Systems
3.3. Inorganic Compound-Based Delivery Systems
3.4. Extracellular Vesicle-Based Delivery Systems
3.5. Emerging Methods of miRNA Delivery Systems
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- He, L.; Hannon, G.J. MicroRNAs: Small RNAs with a big role in gene regulation. Nat. Rev. Genet. 2004, 5, 522–531. [Google Scholar] [CrossRef] [PubMed]
- Lai, E.C. Micro RNAs are complementary to 3′ UTR sequence motifs that mediate negative post-transcriptional regulation. Nat. Genet. 2002, 30, 363–364. [Google Scholar] [CrossRef] [PubMed]
- Lim, L.P.; Lau, N.C.; Garrett-Engele, P.; Grimson, A.; Schelter, J.M.; Castle, J.; Bartel, D.P.; Linsley, P.S.; Johnson, J.M. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 2005, 433, 769–773. [Google Scholar] [CrossRef] [PubMed]
- Olsen, P.H.; Ambros, V. The lin-4 regulatory RNA controls developmental timing in Caenorhabditis elegans by blocking LIN-14 protein synthesis after the initiation of translation. Dev. Biol. 1999, 216, 671–680. [Google Scholar] [CrossRef]
- Hammond, S.M. An overview of microRNAs. Adv. Drug Deliv. Rev. 2015, 87, 3–14. [Google Scholar] [CrossRef]
- Kozomara, A.; Birgaoanu, M.; Griffiths-Jones, S. miRBase: From microRNA sequences to function. Nucleic Acids Res. 2019, 47, D155–D162. [Google Scholar] [CrossRef]
- Labatut, A.E.; Mattheolabakis, G. Non-viral based miR delivery and recent developments. Eur. J. Pharm. Biopharm. 2018, 128, 82–90. [Google Scholar] [CrossRef]
- Wilson, R.C.; Doudna, J.A. Molecular mechanisms of RNA interference. Annu. Rev. Biophys. 2013, 42, 217–239. [Google Scholar] [CrossRef]
- Landgraf, P.; Rusu, M.; Sheridan, R.; Sewer, A.; Iovino, N.; Aravin, A.; Pfeffer, S.; Rice, A.; Kamphorst, A.O.; Landthaler, M.; et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 2007, 129, 1401–1414. [Google Scholar] [CrossRef]
- Ludwig, N.; Leidinger, P.; Becker, K.; Backes, C.; Fehlmann, T.; Pallasch, C.; Rheinheimer, S.; Meder, B.; Stahler, C.; Meese, E.; et al. Distribution of miRNA expression across human tissues. Nucleic Acids Res. 2016, 44, 3865–3877. [Google Scholar] [CrossRef]
- Drury, R.E.; O’Connor, D.; Pollard, A.J. The Clinical Application of MicroRNAs in Infectious Disease. Front. Immunol. 2017, 8, 1182. [Google Scholar] [CrossRef] [PubMed]
- Finotti, A.; Fabbri, E.; Lampronti, I.; Gasparello, J.; Borgatti, M.; Gambari, R. MicroRNAs and Long Non-coding RNAs in Genetic Diseases. Mol. Diagn. Ther. 2019, 23, 155–171. [Google Scholar] [CrossRef] [PubMed]
- Markopoulos, G.S.; Roupakia, E.; Tokamani, M.; Alabasi, G.; Sandaltzopoulos, R.; Marcu, K.B.; Kolettas, E. Roles of NF-kappaB Signaling in the Regulation of miRNAs Impacting on Inflammation in Cancer. Biomedicines 2018, 6, 40. [Google Scholar] [CrossRef] [PubMed]
- McCoy, C.E. miR-155 Dysregulation and Therapeutic Intervention in Multiple Sclerosis. Adv. Exp. Med. Biol. 2017, 1024, 111–131. [Google Scholar] [CrossRef]
- Miya Shaik, M.; Tamargo, I.A.; Abubakar, M.B.; Kamal, M.A.; Greig, N.H.; Gan, S.H. The Role of microRNAs in Alzheimer’s Disease and Their Therapeutic Potentials. Genes 2018, 9, 174. [Google Scholar] [CrossRef]
- Strumidlo, A.; Skiba, S.; Scott, R.J.; Lubinski, J. The potential role of miRNAs in therapy of breast and ovarian cancers associated with BRCA1 mutation. Hered Cancer Clin. Pract. 2017, 15, 15. [Google Scholar] [CrossRef]
- Wojciechowska, A.; Braniewska, A.; Kozar-Kaminska, K. MicroRNA in cardiovascular biology and disease. Adv. Clin. Exp. Med. 2017, 26, 865–874. [Google Scholar] [CrossRef]
- Esau, C.C.; Monia, B.P. Therapeutic potential for microRNAs. Adv. Drug Deliv. Rev. 2007, 59, 101–114. [Google Scholar] [CrossRef]
- Li, Z.; Rana, T.M. Therapeutic targeting of microRNAs: Current status and future challenges. Nat. Rev. Drug Discov. 2014, 13, 622–638. [Google Scholar] [CrossRef]
- Bader, A.G.; Brown, D.; Winkler, M. The promise of microRNA replacement therapy. Cancer Res. 2010, 70, 7027–7030. [Google Scholar] [CrossRef]
- Banales, J.M.; Saez, E.; Uriz, M.; Sarvide, S.; Urribarri, A.D.; Splinter, P.; Tietz Bogert, P.S.; Bujanda, L.; Prieto, J.; Medina, J.F.; et al. Up-regulation of microRNA 506 leads to decreased Cl-/HCO3- anion exchanger 2 expression in biliary epithelium of patients with primary biliary cirrhosis. Hepatology 2012, 56, 687–697. [Google Scholar] [CrossRef] [PubMed]
- Gambari, R.; Brognara, E.; Spandidos, D.A.; Fabbri, E. Targeting oncomiRNAs and mimicking tumor suppressor miRNAs: New trends in the development of miRNA therapeutic strategies in oncology. Int. J. Oncol. 2016, 49, 5–32. [Google Scholar] [CrossRef] [PubMed]
- Cheng, C.J.; Bahal, R.; Babar, I.A.; Pincus, Z.; Barrera, F.; Liu, C.; Svoronos, A.; Braddock, D.T.; Glazer, P.M.; Engelman, D.M.; et al. MicroRNA silencing for cancer therapy targeted to the tumour microenvironment. Nature 2015, 518, 107–110. [Google Scholar] [CrossRef] [PubMed]
- Tang, L.; Chen, H.Y.; Hao, N.B.; Tang, B.; Guo, H.; Yong, X.; Dong, H.; Yang, S.M. microRNA inhibitors: Natural and artificial sequestration of microRNA. Cancer Lett. 2017, 407, 139–147. [Google Scholar] [CrossRef]
- Bravo, V.; Rosero, S.; Ricordi, C.; Pastori, R.L. Instability of miRNA and cDNAs derivatives in RNA preparations. Biochem. Biophys. Res. Commun. 2007, 353, 1052–1055. [Google Scholar] [CrossRef]
- Cho, W.C. Role of miRNAs in lung cancer. Expert Rev. Mol. Diagn. 2009, 9, 773–776. [Google Scholar] [CrossRef]
- Grimm, D.; Streetz, K.L.; Jopling, C.L.; Storm, T.A.; Pandey, K.; Davis, C.R.; Marion, P.; Salazar, F.; Kay, M.A. Fatality in mice due to oversaturation of cellular microRNA/short hairpin RNA pathways. Nature 2006, 441, 537–541. [Google Scholar] [CrossRef]
- Pecot, C.V.; Calin, G.A.; Coleman, R.L.; Lopez-Berestein, G.; Sood, A.K. RNA interference in the clinic: Challenges and future directions. Nat. Rev. Cancer 2011, 11, 59–67. [Google Scholar] [CrossRef]
- Wang, H.; Jiang, Y.; Peng, H.; Chen, Y.; Zhu, P.; Huang, Y. Recent progress in microRNA delivery for cancer therapy by non-viral synthetic vectors. Adv. Drug Deliv. Rev. 2015, 81, 142–160. [Google Scholar] [CrossRef]
- Crooke, S.T.; Graham, M.J.; Zuckerman, J.E.; Brooks, D.; Conklin, B.S.; Cummins, L.L.; Greig, M.J.; Guinosso, C.J.; Kornbrust, D.; Manoharan, M.; et al. Pharmacokinetic properties of several novel oligonucleotide analogs in mice. J. Pharmacol. Exp. Ther. 1996, 277, 923–937. [Google Scholar]
- Van Rooij, E.; Kauppinen, S. Development of microRNA therapeutics is coming of age. EMBO Mol. Med. 2014, 6, 851–864. [Google Scholar] [CrossRef] [PubMed]
- Wahlestedt, C.; Salmi, P.; Good, L.; Kela, J.; Johnsson, T.; Hokfelt, T.; Broberger, C.; Porreca, F.; Lai, J.; Ren, K.; et al. Potent and nontoxic antisense oligonucleotides containing locked nucleic acids. Proc. Natl. Acad. Sci. USA 2000, 97, 5633–5638. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Guo, Q.; Wei, W.; Robertson, S.; Yuan, Y.; Luo, X. Current Progress on MicroRNA-Based Gene Delivery in the Treatment of Osteoporosis and Osteoporotic Fracture. Int. J. Endocrinol. 2019, 2019, 6782653. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Gao, D.Y.; Huang, L. In vivo delivery of miRNAs for cancer therapy: Challenges and strategies. Adv. Drug. Deliv. Rev. 2015, 81, 128–141. [Google Scholar] [CrossRef] [PubMed]
- Forterre, A.; Komuro, H.; Aminova, S.; Harada, M. A Comprehensive Review of Cancer MicroRNA Therapeutic Delivery Strategies. Cancers 2020, 12, 1852. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.G.; Song, J.; Zhang, Y.Q.; Wang, P.C. MicroRNA-193b is a regulator of amyloid precursor protein in the blood and cerebrospinal fluid derived exosomal microRNA-193b is a biomarker of Alzheimer’s disease. Mol. Med. Rep. 2014, 10, 2395–2400. [Google Scholar] [CrossRef]
- Theis, T.; Yoo, M.; Park, C.S.; Chen, J.; Kugler, S.; Gibbs, K.M.; Schachner, M. Lentiviral delivery of miR-133b improves functional recovery after spinal cord injury in mice. Mol. Neurobiol. 2017, 54, 4659–4671. [Google Scholar] [CrossRef]
- Trang, P.; Medina, P.P.; Wiggins, J.F.; Ruffino, L.; Kelnar, K.; Omotola, M.; Homer, R.; Brown, D.; Bader, A.G.; Weidhaas, J.B.; et al. Regression of murine lung tumors by the let-7 microRNA. Oncogene 2010, 29, 1580–1587. [Google Scholar] [CrossRef]
- Yin, L.; Keeler, G.D.; Zhang, Y.; Hoffman, B.E.; Ling, C.; Qing, K.; Srivastava, A. AAV3-miRNA vectors for growth suppression of human hepatocellular carcinoma cells in vitro and human liver tumors in a murine xenograft model in vivo. Gene. Ther. 2020. [Google Scholar] [CrossRef]
- Pourshafie, N.; Lee, P.R.; Chen, K.L.; Harmison, G.G.; Bott, L.C.; Fischbeck, K.H.; Rinaldi, C. Systemic delivery of microRNA using recombinant adeno-associated virus serotype 9 to treat neuromuscular diseases in rodents. J. Vis. Exp. 2018. [Google Scholar] [CrossRef]
- Martier, R.; Sogorb-Gonzalez, M.; Stricker-Shaver, J.; Hubener-Schmid, J.; Keskin, S.; Klima, J.; Toonen, L.J.; Juhas, S.; Juhasova, J.; Ellederova, Z.; et al. Development of an AAV-Based microRNA gene therapy to treat Machado-Joseph Disease. Mol. Ther. Methods Clin. Dev. 2019, 15, 343–358. [Google Scholar] [CrossRef] [PubMed]
- Gokita, K.; Inoue, J.; Ishihara, H.; Kojima, K.; Inazawa, J. Therapeutic potential of LNP-mediated delivery of miR-634 for cancer therapy. Mol. Ther. Nucleic Acids 2020, 19, 330–338. [Google Scholar] [CrossRef] [PubMed]
- Misso, G.; Di Martino, M.T.; De Rosa, G.; Farooqi, A.A.; Lombardi, A.; Campani, V.; Zarone, M.R.; Gulla, A.; Tagliaferri, P.; Tassone, P.; et al. Mir-34: A new weapon against cancer? Mol. Ther. Nucleic Acids 2014, 3, e194. [Google Scholar] [CrossRef] [PubMed]
- Kheirolomoom, A.; Kim, C.W.; Seo, J.W.; Kumar, S.; Son, D.J.; Gagnon, M.K.; Ingham, E.S.; Ferrara, K.W.; Jo, H. Multifunctional nanoparticles facilitate molecular targeting and miRNA delivery to inhibit atherosclerosis in ApoE(-/-) mice. ACS Nano 2015, 9, 8885–8897. [Google Scholar] [CrossRef] [PubMed]
- Rai, K.; Takigawa, N.; Ito, S.; Kashihara, H.; Ichihara, E.; Yasuda, T.; Shimizu, K.; Tanimoto, M.; Kiura, K. Liposomal delivery of microRNA-7-expressing plasmid overcomes epidermal growth factor receptor tyrosine kinase inhibitor-resistance in lung cancer cells. Mol. Cancer Ther. 2011, 10, 1720–1727. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Crawford, M.; Mao, Y.; Lee, R.J.; Davis, I.C.; Elton, T.S.; Lee, L.J.; Nana-Sinkam, S.P. Therapeutic Delivery of MicroRNA-29b by Cationic Lipoplexes for Lung Cancer. Mol. Ther. Nucleic Acids 2013, 2, e84. [Google Scholar] [CrossRef] [PubMed]
- Cortez, M.A.; Valdecanas, D.; Zhang, X.; Zhan, Y.; Bhardwaj, V.; Calin, G.A.; Komaki, R.; Giri, D.K.; Quini, C.C.; Wolfe, T.; et al. Therapeutic delivery of miR-200c enhances radiosensitivity in lung cancer. Mol. Ther. 2014, 22, 1494–1503. [Google Scholar] [CrossRef] [PubMed]
- De Antonellis, P.; Liguori, L.; Falanga, A.; Carotenuto, M.; Ferrucci, V.; Andolfo, I.; Marinaro, F.; Scognamiglio, I.; Virgilio, A.; De Rosa, G.; et al. MicroRNA 199b-5p delivery through stable nucleic acid lipid particles (SNALPs) in tumorigenic cell lines. Naunyn Schmiedebergs Arch. Pharmacol. 2013, 386, 287–302. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, A.F.; Weirauch, U.; Thomas, M.; Grunweller, A.; Hartmann, R.K.; Aigner, A. MicroRNA replacement therapy for miR-145 and miR-33a is efficacious in a model of colon carcinoma. Cancer Res. 2011, 71, 5214–5224. [Google Scholar] [CrossRef] [PubMed]
- Hu, Q.; Wang, K.; Sun, X.; Li, Y.; Fu, Q.; Liang, T.; Tang, G. A redox-sensitive, oligopeptide-guided, self-assembling, and efficiency-enhanced (ROSE) system for functional delivery of microRNA therapeutics for treatment of hepatocellular carcinoma. Biomaterials 2016, 104, 192–200. [Google Scholar] [CrossRef] [PubMed]
- Seo, Y.E.; Suh, H.W.; Bahal, R.; Josowitz, A.; Zhang, J.; Song, E.; Cui, J.; Noorbakhsh, S.; Jackson, C.; Bu, T.; et al. Nanoparticle-mediated intratumoral inhibition of miR-21 for improved survival in glioblastoma. Biomaterials 2019, 201, 87–98. [Google Scholar] [CrossRef] [PubMed]
- Qian, X.; Long, L.; Shi, Z.; Liu, C.; Qiu, M.; Sheng, J.; Pu, P.; Yuan, X.; Ren, Y.; Kang, C. Star-branched amphiphilic PLA-b-PDMAEMA copolymers for co-delivery of miR-21 inhibitor and doxorubicin to treat glioma. Biomaterials 2014, 35, 2322–2335. [Google Scholar] [CrossRef] [PubMed]
- Hiraki, M.; Nishimura, J.; Takahashi, H.; Wu, X.; Takahashi, Y.; Miyo, M.; Nishida, N.; Uemura, M.; Hata, T.; Takemasa, I.; et al. Concurrent targeting of KRAS and AKT by miR-4689 is a novel treatment against mutant KRAS colorectal cancer. Mol. Ther. Nucleic Acids 2015, 4, e231. [Google Scholar] [CrossRef] [PubMed]
- Inoue, A.; Mizushima, T.; Wu, X.; Okuzaki, D.; Kambara, N.; Ishikawa, S.; Wang, J.; Qian, Y.; Hirose, H.; Yokoyama, Y.; et al. A miR-29b byproduct sequence exhibits potent tumor-suppressive activities via inhibition of NF-kappaB signaling in KRAS-mutant colon cancer cells. Mol. Cancer Ther. 2018, 17, 977–987. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, M.; Sawada, K.; Miyamoto, M.; Shimizu, A.; Yamamoto, M.; Kinose, Y.; Nakamura, K.; Kawano, M.; Kodama, M.; Hashimoto, K.; et al. Exploring the potential of engineered exosomes as delivery systems for tumor-suppressor microRNA replacement therapy in ovarian cancer. Biochem. Biophys. Res. Commun. 2020, 527, 153–161. [Google Scholar] [CrossRef] [PubMed]
- Ohno, S.; Takanashi, M.; Sudo, K.; Ueda, S.; Ishikawa, A.; Matsuyama, N.; Fujita, K.; Mizutani, T.; Ohgi, T.; Ochiya, T.; et al. Systemically injected exosomes targeted to EGFR deliver antitumor microRNA to breast cancer cells. Mol. Ther. 2013, 21, 185–191. [Google Scholar] [CrossRef] [PubMed]
- Lou, G.; Song, X.; Yang, F.; Wu, S.; Wang, J.; Chen, Z.; Liu, Y. Exosomes derived from miR-122-modified adipose tissue-derived MSCs increase chemosensitivity of hepatocellular carcinoma. J. Hematol. Oncol. 2015, 8, 122. [Google Scholar] [CrossRef] [PubMed]
- Vazquez-Rios, A.J.; Molina-Crespo, A.; Bouzo, B.L.; Lopez-Lopez, R.; Moreno-Bueno, G.; de la Fuente, M. Exosome-mimetic nanoplatforms for targeted cancer drug delivery. J. Nanobiotechnol. 2019, 17, 85. [Google Scholar] [CrossRef]
- Fu, Y.; Chen, J.; Huang, Z. Recent progress in microRNA-based delivery systems for the treatment of human disease. ExRNA 2019, 1, 24. [Google Scholar] [CrossRef]
- Yang, N. An overview of viral and nonviral delivery systems for microRNA. Int. J. Pharm. Investig. 2015, 5, 179–181. [Google Scholar] [CrossRef]
- Pan, Y.; Zhang, Y.; Jia, T.; Zhang, K.; Li, J.; Wang, L. Development of a microRNA delivery system based on bacteriophage MS2 virus-like particles. FEBS J. 2012, 279, 1198–1208. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Sun, Y.; Zhao, R. Establishment of microRNA delivery system by PP7 bacteriophage-like particles carrying cell-penetrating peptide. J. Biosci. Bioeng. 2017, 124, 242–249. [Google Scholar] [CrossRef] [PubMed]
- Pages, J.C.; Bru, T. Toolbox for retrovectorologists. J. Gene Med. 2004, 6 (Suppl. 1), S67–S82. [Google Scholar] [CrossRef] [PubMed]
- Cavazzana-Calvo, M.; Hacein-Bey, S.; de Saint Basile, G.; Gross, F.; Yvon, E.; Nusbaum, P.; Selz, F.; Hue, C.; Certain, S.; Casanova, J.L.; et al. Gene therapy of human severe combined immunodeficiency (SCID)-X1 disease. Science 2000, 288, 669–672. [Google Scholar] [CrossRef] [PubMed]
- Hacein-Bey-Abina, S.; Von Kalle, C.; Schmidt, M.; McCormack, M.P.; Wulffraat, N.; Leboulch, P.; Lim, A.; Osborne, C.S.; Pawliuk, R.; Morillon, E.; et al. LMO2-associated clonal T cell proliferation in two patients after gene therapy for SCID-X1. Science 2003, 302, 415–419. [Google Scholar] [CrossRef]
- Ye, D.; Wang, G.; Liu, Y.; Huang, W.; Wu, M.; Zhu, S.; Jia, W.; Deng, A.M.; Liu, H.; Kang, J. MiR-138 promotes induced pluripotent stem cell generation through the regulation of the p53 signaling. Stem Cells 2012, 30, 1645–1654. [Google Scholar] [CrossRef]
- Berkowitz, R.; Ilves, H.; Lin, W.Y.; Eckert, K.; Coward, A.; Tamaki, S.; Veres, G.; Plavec, I. Construction and molecular analysis of gene transfer systems derived from bovine immunodeficiency virus. J. Virol. 2001, 75, 3371–3382. [Google Scholar] [CrossRef]
- Leroux, C.; Cadore, J.L.; Montelaro, R.C. Equine Infectious Anemia Virus (EIAV): What has HIV’s country cousin got to tell us? Vet. Res. 2004, 35, 485–512. [Google Scholar] [CrossRef]
- Mangeot, P.E.; Negre, D.; Dubois, B.; Winter, A.J.; Leissner, P.; Mehtali, M.; Kaiserlian, D.; Cosset, F.L.; Darlix, J.L. Development of minimal lentivirus vectors derived from simian immunodeficiency virus (SIVmac251) and their use for gene transfer into human dendritic cells. J. Virol. 2000, 74, 8307–8315. [Google Scholar] [CrossRef]
- Poeschla, E.; Gilbert, J.; Li, X.; Huang, S.; Ho, A.; Wong-Staal, F. Identification of a human immunodeficiency virus type 2 (HIV-2) encapsidation determinant and transduction of nondividing human cells by HIV-2-based lentivirus vectors. J. Virol. 1998, 72, 6527–6536. [Google Scholar] [CrossRef]
- Poeschla, E.M.; Wong-Staal, F.; Looney, D.J. Efficient transduction of nondividing human cells by feline immunodeficiency virus lentiviral vectors. Nat. Med. 1998, 4, 354–357. [Google Scholar] [CrossRef] [PubMed]
- Milone, M.C.; O’Doherty, U. Clinical use of lentiviral vectors. Leukemia 2018, 32, 1529–1541. [Google Scholar] [CrossRef] [PubMed]
- Baum, C.; Dullmann, J.; Li, Z.; Fehse, B.; Meyer, J.; Williams, D.A.; von Kalle, C. Side effects of retroviral gene transfer into hematopoietic stem cells. Blood 2003, 101, 2099–2114. [Google Scholar] [CrossRef] [PubMed]
- Von Kalle, C.; Fehse, B.; Layh-Schmitt, G.; Schmidt, M.; Kelly, P.; Baum, C. Stem cell clonality and genotoxicity in hematopoietic cells: Gene activation side effects should be avoidable. Semin. Hematol. 2004, 41, 303–318. [Google Scholar] [CrossRef] [PubMed]
- Laufs, S.; Guenechea, G.; Gonzalez-Murillo, A.; Zsuzsanna Nagy, K.; Luz Lozano, M.; del Val, C.; Jonnakuty, S.; Hotz-Wagenblatt, A.; Jens Zeller, W.; Bueren, J.A.; et al. Lentiviral vector integration sites in human NOD/SCID repopulating cells. J. Gene Med. 2006, 8, 1197–1207. [Google Scholar] [CrossRef]
- Montini, E.; Cesana, D.; Schmidt, M.; Sanvito, F.; Bartholomae, C.C.; Ranzani, M.; Benedicenti, F.; Sergi, L.S.; Ambrosi, A.; Ponzoni, M.; et al. The genotoxic potential of retroviral vectors is strongly modulated by vector design and integration site selection in a mouse model of HSC gene therapy. J. Clin. Investig. 2009, 119, 964–975. [Google Scholar] [CrossRef] [PubMed]
- Montini, E.; Cesana, D.; Schmidt, M.; Sanvito, F.; Ponzoni, M.; Bartholomae, C.; Sergi Sergi, L.; Benedicenti, F.; Ambrosi, A.; Di Serio, C.; et al. Hematopoietic stem cell gene transfer in a tumor-prone mouse model uncovers low genotoxicity of lentiviral vector integration. Nat. Biotechnol. 2006, 24, 687–696. [Google Scholar] [CrossRef]
- Kasar, S.; Salerno, E.; Yuan, Y.; Underbayev, C.; Vollenweider, D.; Laurindo, M.F.; Fernandes, H.; Bonci, D.; Addario, A.; Mazzella, F.; et al. Systemic in vivo lentiviral delivery of miR-15a/16 reduces malignancy in the NZB de novo mouse model of chronic lymphocytic leukemia. Genes Immun. 2012, 13, 109–119. [Google Scholar] [CrossRef]
- Liu, Y.; Lai, L.; Chen, Q.; Song, Y.; Xu, S.; Ma, F.; Wang, X.; Wang, J.; Yu, H.; Cao, X.; et al. MicroRNA-494 is required for the accumulation and functions of tumor-expanded myeloid-derived suppressor cells via targeting of PTEN. J. Immunol. 2012, 188, 5500–5510. [Google Scholar] [CrossRef]
- Daya, S.; Berns, K.I. Gene therapy using adeno-associated virus vectors. Clin. Microbiol. Rev. 2008, 21, 583–593. [Google Scholar] [CrossRef]
- Schultz, B.R.; Chamberlain, J.S. Recombinant adeno-associated virus transduction and integration. Mol. Ther. 2008, 16, 1189–1199. [Google Scholar] [CrossRef] [PubMed]
- Miyazaki, Y.; Adachi, H.; Katsuno, M.; Minamiyama, M.; Jiang, Y.M.; Huang, Z.; Doi, H.; Matsumoto, S.; Kondo, N.; Iida, M.; et al. Viral delivery of miR-196a ameliorates the SBMA phenotype via the silencing of CELF2. Nat. Med. 2012, 18, 1136–1141. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.; Zhang, H.; Song, Y.; Zhou, D.; Wang, J. Hemagglutinin-targeting artificial microRNAs expressed by adenovirus protect mice from different clades of H5N1 infection. Mol. Ther. Nucleic Acids 2016, 5, e311. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Vandenberghe, L.H.; Wilson, J.M. AAV as an immunogen. Curr. Gene Ther. 2007, 7, 325–333. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Li, J.; Fu, F.H.; Chen, C.; Zhu, X.; Zhou, L.; Jiang, X.; Xiao, X. Construction and analysis of compact muscle-specific promoters for AAV vectors. Gene Ther. 2008, 15, 1489–1499. [Google Scholar] [CrossRef]
- Qiao, C.; Yuan, Z.; Li, J.; He, B.; Zheng, H.; Mayer, C.; Li, J.; Xiao, X. Liver-specific microRNA-122 target sequences incorporated in AAV vectors efficiently inhibits transgene expression in the liver. Gene Ther. 2011, 18, 403–410. [Google Scholar] [CrossRef]
- Tripp, R.A.; Mark Tompkins, S. Antiviral effects of inhibiting host gene expression. Curr. Top. Microbiol. Immunol. 2015, 386, 459–477. [Google Scholar] [CrossRef]
- Cooray, S.; Howe, S.J.; Thrasher, A.J. Retrovirus and lentivirus vector design and methods of cell conditioning. Methods Enzymol. 2012, 507, 29–57. [Google Scholar] [CrossRef]
- Vannucci, L.; Lai, M.; Chiuppesi, F.; Ceccherini-Nelli, L.; Pistello, M. Viral vectors: A look back and ahead on gene transfer technology. New Microbiol. 2013, 36, 1–22. [Google Scholar]
- Pan, Y.; Jia, T.; Zhang, Y.; Zhang, K.; Zhang, R.; Li, J.; Wang, L. MS2 VLP-based delivery of microRNA-146a inhibits autoantibody production in lupus-prone mice. Int. J. Nanomed. 2012, 7, 5957–5967. [Google Scholar] [CrossRef]
- Wang, G.; Jia, T.; Xu, X.; Chang, L.; Zhang, R.; Fu, Y.; Li, Y.; Yang, X.; Zhang, K.; Lin, G.; et al. Novel miR-122 delivery system based on MS2 virus like particle surface displaying cell-penetrating peptide TAT for hepatocellular carcinoma. Oncotarget 2016, 7, 59402–59416. [Google Scholar] [CrossRef] [PubMed]
- Yao, Y.; Jia, T.; Pan, Y.; Gou, H.; Li, Y.; Sun, Y.; Zhang, R.; Zhang, K.; Lin, G.; Xie, J.; et al. Using a novel microRNA delivery system to inhibit osteoclastogenesis. Int. J. Mol. Sci. 2015, 16, 8337–8350. [Google Scholar] [CrossRef] [PubMed]
- Ye, X.; Liu, Z.; Hemida, M.G.; Yang, D. Targeted delivery of mutant tolerant anti-coxsackievirus artificial microRNAs using folate conjugated bacteriophage Phi29 pRNA. PLoS ONE 2011, 6, e21215. [Google Scholar] [CrossRef] [PubMed]
- Guo, P.X.; Erickson, S.; Anderson, D. A small viral RNA is required for in vitro packaging of bacteriophage phi 29 DNA. Science 1987, 236, 690–694. [Google Scholar] [CrossRef]
- Guo, P. Bacterial virus phi29 DNA-packaging motor and its potential applications in gene therapy and nanotechnology. Methods Mol. Biol. 2005, 300, 285–324. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Wang, Z.; Gemeinhart, R.A. Progress in microRNA delivery. J. Control. Release 2013, 172, 962–974. [Google Scholar] [CrossRef] [PubMed]
- Hsu, S.H.; Yu, B.; Wang, X.; Lu, Y.; Schmidt, C.R.; Lee, R.J.; Lee, L.J.; Jacob, S.T.; Ghoshal, K. Cationic lipid nanoparticles for therapeutic delivery of siRNA and miRNA to murine liver tumor. Nanomedicine 2013, 9, 1169–1180. [Google Scholar] [CrossRef] [PubMed]
- Pedroso de Lima, M.C.; Simões, S.; Pires, P.; Faneca, H.; Düzgüneş, N. Cationic lipid–DNA complexes in gene delivery: From biophysics to biological applications. Adv. Drug Deliv. Rev. 2001, 47, 277–294. [Google Scholar] [CrossRef]
- Hara, E.S.; Ono, M.; Eguchi, T.; Kubota, S.; Pham, H.T.; Sonoyama, W.; Tajima, S.; Takigawa, M.; Calderwood, S.K.; Kuboki, T. miRNA-720 controls stem cell phenotype, proliferation and differentiation of human dental pulp cells. PLoS ONE 2013, 8, e83545. [Google Scholar] [CrossRef] [PubMed]
- Sluijter, J.P.; van Mil, A.; van Vliet, P.; Metz, C.H.; Liu, J.; Doevendans, P.A.; Goumans, M.J. MicroRNA-1 and -499 regulate differentiation and proliferation in human-derived cardiomyocyte progenitor cells. Arterioscler. Thromb. Vasc. Biol. 2010, 30, 859–868. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Sun, G.; Li, S.; Shi, Y. A feedback regulatory loop involving microRNA-9 and nuclear receptor TLX in neural stem cell fate determination. Nat. Struct. Mol. Biol. 2009, 16, 365–371. [Google Scholar] [CrossRef] [PubMed]
- Sundaram, G.M.; Common, J.E.; Gopal, F.E.; Srikanta, S.; Lakshman, K.; Lunny, D.P.; Lim, T.C.; Tanavde, V.; Lane, E.B.; Sampath, P. ‘See-saw’ expression of microRNA-198 and FSTL1 from a single transcript in wound healing. Nature 2013, 495, 103–106. [Google Scholar] [CrossRef] [PubMed]
- Silva, B.F.; Majzoub, R.N.; Chan, C.L.; Li, Y.; Olsson, U.; Safinya, C.R. PEGylated cationic liposome-DNA complexation in brine is pathway-dependent. Biochim. Biophys. Acta 2014, 1838, 398–412. [Google Scholar] [CrossRef] [PubMed]
- Endo-Takahashi, Y.; Negishi, Y.; Nakamura, A.; Ukai, S.; Ooaku, K.; Oda, Y.; Sugimoto, K.; Moriyasu, F.; Takagi, N.; Suzuki, R.; et al. Systemic delivery of miR-126 by miRNA-loaded bubble liposomes for the treatment of hindlimb ischemia. Sci. Rep. 2014, 4, 3883. [Google Scholar] [CrossRef] [PubMed]
- Shi, S.; Han, L.; Gong, T.; Zhang, Z.; Sun, X. Systemic delivery of microRNA-34a for cancer stem cell therapy. Angew. Chem. Int. Ed. Engl. 2013, 52, 3901–3905. [Google Scholar] [CrossRef] [PubMed]
- Piao, L.; Zhang, M.; Datta, J.; Xie, X.; Su, T.; Li, H.; Teknos, T.N.; Pan, Q. Lipid-based nanoparticle delivery of pre-miR-107 inhibits the tumorigenicity of head and neck squamous cell carcinoma. Mol. Ther. 2012, 20, 1261–1269. [Google Scholar] [CrossRef]
- Xu, F.; Liao, J.Z.; Xiang, G.Y.; Zhao, P.X.; Ye, F.; Zhao, Q.; He, X.X. MiR-101 and doxorubicin codelivered by liposomes suppressing malignant properties of hepatocellular carcinoma. Cancer Med. 2017, 6, 651–661. [Google Scholar] [CrossRef]
- Reddy, T.L.; Garikapati, K.R.; Reddy, S.G.; Reddy, B.V.; Yadav, J.S.; Bhadra, U.; Bhadra, M.P. Simultaneous delivery of Paclitaxel and Bcl-2 siRNA via pH-Sensitive liposomal nanocarrier for the synergistic treatment of melanoma. Sci. Rep. 2016, 6, 35223. [Google Scholar] [CrossRef]
- Nishimura, M.; Jung, E.J.; Shah, M.Y.; Lu, C.; Spizzo, R.; Shimizu, M.; Han, H.D.; Ivan, C.; Rossi, S.; Zhang, X.; et al. Therapeutic synergy between microRNA and siRNA in ovarian cancer treatment. Cancer Discov. 2013, 3, 1302–1315. [Google Scholar] [CrossRef]
- Yang, D.; Sun, Y.; Hu, L.; Zheng, H.; Ji, P.; Pecot, C.V.; Zhao, Y.; Reynolds, S.; Cheng, H.; Rupaimoole, R.; et al. Integrated analyses identify a master microRNA regulatory network for the mesenchymal subtype in serous ovarian cancer. Cancer Cell 2013, 23, 186–199. [Google Scholar] [CrossRef]
- Pecot, C.V.; Rupaimoole, R.; Yang, D.; Akbani, R.; Ivan, C.; Lu, C.; Wu, S.; Han, H.D.; Shah, M.Y.; Rodriguez-Aguayo, C.; et al. Tumour angiogenesis regulation by the miR-200 family. Nat. Commun. 2013, 4, 2427. [Google Scholar] [CrossRef] [PubMed]
- Zylberberg, C.; Matosevic, S. Pharmaceutical liposomal drug delivery: A review of new delivery systems and a look at the regulatory landscape. Drug Deliv. 2016, 23, 3319–3329. [Google Scholar] [CrossRef] [PubMed]
- Wilner, S.E.; Levy, M. Synthesis and Characterization of Aptamer-Targeted SNALPs for the Delivery of siRNA. Methods Mol. Biol. 2016, 1380, 211–224. [Google Scholar] [CrossRef] [PubMed]
- Hobel, S.; Aigner, A. Polyethylenimines for siRNA and miRNA delivery in vivo. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2013, 5, 484–501. [Google Scholar] [CrossRef]
- Jager, M.; Schubert, S.; Ochrimenko, S.; Fischer, D.; Schubert, U.S. Branched and linear poly(ethylene imine)-based conjugates: Synthetic modification, characterization, and application. Chem. Soc. Rev. 2012, 41, 4755–4767. [Google Scholar] [CrossRef]
- Gao, S.; Tian, H.; Guo, Y.; Li, Y.; Guo, Z.; Zhu, X.; Chen, X. miRNA oligonucleotide and sponge for miRNA-21 inhibition mediated by PEI-PLL in breast cancer therapy. Acta Biomater. 2015, 25, 184–193. [Google Scholar] [CrossRef]
- Biray Avci, C.; Ozcan, I.; Balci, T.; Ozer, O.; Gunduz, C. Design of polyethylene glycol-polyethylenimine nanocomplexes as non-viral carriers: Mir-150 delivery to chronic myeloid leukemia cells. Cell Biol. Int. 2013, 37, 1205–1214. [Google Scholar] [CrossRef]
- Cheng, C.J.; Saltzman, W.M. Polymer nanoparticle-mediated delivery of microRNA inhibition and alternative splicing. Mol. Pharm. 2012, 9, 1481–1488. [Google Scholar] [CrossRef]
- Wang, F.; Zhang, B.; Zhou, L.; Shi, Y.; Li, Z.; Xia, Y.; Tian, J. Imaging dendrimer-grafted graphene oxide mediated anti-miR-21 delivery with an activatable luciferase reporter. ACS Appl. Mater. Interfaces 2016, 8, 9014–9021. [Google Scholar] [CrossRef]
- Salzano, G.; Costa, D.F.; Sarisozen, C.; Luther, E.; Mattheolabakis, G.; Dhargalkar, P.P.; Torchilin, V.P. Mixed nanosized polymeric micelles as promoter of doxorubicin and miRNA-34a co-delivery triggered by dual stimuli in tumor tissue. Small 2016, 12, 4837–4848. [Google Scholar] [CrossRef]
- Suh, J.S.; Lee, J.Y.; Choi, Y.S.; Chung, C.P.; Park, Y.J. Peptide-mediated intracellular delivery of miRNA-29b for osteogenic stem cell differentiation. Biomaterials 2013, 34, 4347–4359. [Google Scholar] [CrossRef] [PubMed]
- Gao, S.; Chen, J.; Dong, L.; Ding, Z.; Yang, Y.H.; Zhang, J. Targeting delivery of oligonucleotide and plasmid DNA to hepatocyte via galactosylated chitosan vector. Eur. J. Pharm. Biopharm. 2005, 60, 327–334. [Google Scholar] [CrossRef]
- Gao, S.; Chen, J.; Xu, X.; Ding, Z.; Yang, Y.H.; Hua, Z.; Zhang, J. Galactosylated low molecular weight chitosan as DNA carrier for hepatocyte-targeting. Int. J. Pharm. 2003, 255, 57–68. [Google Scholar] [CrossRef]
- Martirosyan, A.; Olesen, M.J.; Howard, K.A. Chitosan-based nanoparticles for mucosal delivery of RNAi therapeutics. Adv. Genet. 2014, 88, 325–352. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Xianyu, Y.; Jiang, X. Surface modification of gold nanoparticles with small molecules for biochemical analysis. Acc. Chem. Res. 2017, 50, 310–319. [Google Scholar] [CrossRef] [PubMed]
- Schade, A.; Delyagina, E.; Scharfenberg, D.; Skorska, A.; Lux, C.; David, R.; Steinhoff, G. Innovative strategy for microRNA delivery in human mesenchymal stem cells via magnetic nanoparticles. Int. J. Mol. Sci. 2013, 14, 10710–10726. [Google Scholar] [CrossRef] [PubMed]
- Bitar, A.; Ahmad, N.M.; Fessi, H.; Elaissari, A. Silica-based nanoparticles for biomedical applications. Drug Discov. Today 2012, 17, 1147–1154. [Google Scholar] [CrossRef] [PubMed]
- Jia, C.; Chen, H.; Wei, M.; Chen, X.; Zhang, Y.; Cao, L.; Yuan, P.; Wang, F.; Yang, G.; Ma, J. Gold nanoparticle-based miR155 antagonist macrophage delivery restores the cardiac function in ovariectomized diabetic mouse model. Int. J. Nanomed. 2017, 12, 4963–4979. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, R.; Singh, L.C.; Shohet, J.M.; Gunaratne, P.H. A gold nanoparticle platform for the delivery of functional microRNAs into cancer cells. Biomaterials 2013, 34, 807–816. [Google Scholar] [CrossRef]
- Li, Y.; Duo, Y.; Bi, J.; Zeng, X.; Mei, L.; Bao, S.; He, L.; Shan, A.; Zhang, Y.; Yu, X. Targeted delivery of anti-miR-155 by functionalized mesoporous silica nanoparticles for colorectal cancer therapy. Int. J. Nanomed. 2018, 13, 1241–1256. [Google Scholar] [CrossRef]
- Mamaeva, V.; Sahlgren, C.; Linden, M. Mesoporous silica nanoparticles in medicine-recent advances. Adv. Drug Deliv. Rev. 2013, 65, 689–702. [Google Scholar] [CrossRef] [PubMed]
- Sun, S.; Wang, Y.; Zhou, R.; Deng, Z.; Han, Y.; Han, X.; Tao, W.; Yang, Z.; Shi, C.; Hong, D.; et al. Targeting and regulating of an oncogene via nanovector delivery of microRNA using patient-derived xenografts. Theranostics 2017, 7, 677–693. [Google Scholar] [CrossRef] [PubMed]
- Andaloussi, S.E.; Mager, I.; Breakefield, X.O.; Wood, M.J. Extracellular vesicles: Biology and emerging therapeutic opportunities. Nat. Rev. Drug Discov. 2013, 12, 347–357. [Google Scholar] [CrossRef] [PubMed]
- De Jong, O.G.; Kooijmans, S.A.A.; Murphy, D.E.; Jiang, L.; Evers, M.J.W.; Sluijter, J.P.G.; Vader, P.; Schiffelers, R.M. Drug Delivery with Extracellular Vesicles: From Imagination to Innovation. Acc. Chem. Res. 2019, 52, 1761–1770. [Google Scholar] [CrossRef] [PubMed]
- Ha, D.; Yang, N.; Nadithe, V. Exosomes as therapeutic drug carriers and delivery vehicles across biological membranes: Current perspectives and future challenges. Acta Pharm. Sin. B 2016, 6, 287–296. [Google Scholar] [CrossRef]
- Simons, M.; Raposo, G. Exosomes-vesicular carriers for intercellular communication. Curr. Opin. Cell Biol. 2009, 21, 575–581. [Google Scholar] [CrossRef] [PubMed]
- Munir, J.; Yoon, J.K.; Ryu, S. Therapeutic miRNA-Enriched Extracellular Vesicles: Current Approaches and Future Prospects. Cells 2020, 9, 2271. [Google Scholar] [CrossRef]
- Jiang, M.; Wang, H.; Jin, M.; Yang, X.; Ji, H.; Jiang, Y.; Zhang, H.; Wu, F.; Wu, G.; Lai, X.; et al. Exosomes from MiR-30d-5p-ADSCs Reverse Acute Ischemic Stroke-Induced, Autophagy-Mediated Brain Injury by Promoting M2 Microglial/Macrophage Polarization. Cell Physiol. Biochem. 2018, 47, 864–878. [Google Scholar] [CrossRef]
- Li, D.; Huang, S.; Yin, Z.; Zhu, J.; Ge, X.; Han, Z.; Tan, J.; Zhang, S.; Zhao, J.; Chen, F.; et al. Increases in miR-124-3p in Microglial Exosomes Confer Neuroprotective Effects by Targeting FIP200-Mediated Neuronal Autophagy Following Traumatic Brain Injury. Neurochem. Res. 2019, 44, 1903–1923. [Google Scholar] [CrossRef]
- Luarte, A.; Henzi, R.; Fernandez, A.; Gaete, D.; Cisternas, P.; Pizarro, M.; Batiz, L.F.; Villalobos, I.; Masalleras, M.; Vergara, R.; et al. Astrocyte-Derived Small Extracellular Vesicles Regulate Dendritic Complexity through miR-26a-5p Activity. Cells 2020, 9, 930. [Google Scholar] [CrossRef]
- Xin, H.; Li, Y.; Buller, B.; Katakowski, M.; Zhang, Y.; Wang, X.; Shang, X.; Zhang, Z.G.; Chopp, M. Exosome-mediated transfer of miR-133b from multipotent mesenchymal stromal cells to neural cells contributes to neurite outgrowth. Stem Cells 2012, 30, 1556–1564. [Google Scholar] [CrossRef] [PubMed]
- Luo, Q.; Guo, D.; Liu, G.; Chen, G.; Hang, M.; Jin, M. Exosomes from MiR-126-Overexpressing Adscs Are Therapeutic in Relieving Acute Myocardial Ischaemic Injury. Cell Physiol. Biochem. 2017, 44, 2105–2116. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.; Zhang, C.; Zhang, J.; Jiao, Z.; Dong, N.; Wang, G.; Wang, Z.; Wang, L. Localized injection of miRNA-21-enriched extracellular vesicles effectively restores cardiac function after myocardial infarction. Theranostics 2019, 9, 2346–2360. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Chen, C.; Yang, D.; Liao, Q.; Luo, H.; Wang, X.; Zhou, F.; Yang, X.; Yang, J.; Zeng, C.; et al. Mesenchymal stem cells-derived extracellular vesicles, via miR-210, improve infarcted cardiac function by promotion of angiogenesis. Biochim. Biophys. Acta Mol. Basis Dis. 2017, 1863, 2085–2092. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Tang, Y.; Liu, Y.; Zhang, P.; Lv, L.; Zhang, X.; Jia, L.; Zhou, Y. Exosomes derived from miR-375-overexpressing human adipose mesenchymal stem cells promote bone regeneration. Cell Prolif. 2019, 52, e12669. [Google Scholar] [CrossRef]
- Li, X.; Liu, L.L.; Yao, J.L.; Wang, K.; Ai, H. Human Umbilical Cord Mesenchymal Stem Cell-Derived Extracellular Vesicles Inhibit Endometrial Cancer Cell Proliferation and Migration through Delivery of Exogenous miR-302a. Stem Cells Int. 2019, 2019, 8108576. [Google Scholar] [CrossRef]
- Wang, F.; Li, L.; Piontek, K.; Sakaguchi, M.; Selaru, F.M. Exosome miR-335 as a novel therapeutic strategy in hepatocellular carcinoma. Hepatology 2018, 67, 940–954. [Google Scholar] [CrossRef]
- Morel, L.; Regan, M.; Higashimori, H.; Ng, S.K.; Esau, C.; Vidensky, S.; Rothstein, J.; Yang, Y. Neuronal exosomal miRNA-dependent translational regulation of astroglial glutamate transporter GLT1. J. Biol. Chem. 2013, 288, 7105–7116. [Google Scholar] [CrossRef]
- Zhou, Y.; Li, P.; Goodwin, A.J.; Cook, J.A.; Halushka, P.V.; Chang, E.; Fan, H. Exosomes from Endothelial Progenitor Cells Improve the Outcome of a Murine Model of Sepsis. Mol. Ther. 2018, 26, 1375–1384. [Google Scholar] [CrossRef]
- Wen, D.; Peng, Y.; Liu, D.; Weizmann, Y.; Mahato, R.I. Mesenchymal stem cell and derived exosome as small RNA carrier and Immunomodulator to improve islet transplantation. J. Control. Release 2016, 238, 166–175. [Google Scholar] [CrossRef]
- Ortega, A.; Martinez-Arroyo, O.; Forner, M.J.; Cortes, R. Exosomes as Drug Delivery Systems: Endogenous Nanovehicles for Treatment of Systemic Lupus Erythematosus. Pharmaceutics 2020, 13, 3. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Jia, S.; Xu, H. Potential therapeutic applications of exosomes in different autoimmune diseases. Clin. Immunol. 2019, 205, 116–124. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.Y.; Park, J.K.; Lee, E.Y.; Lee, E.B.; Song, Y.W. Correction to: Circulating exosomes from patients with systemic lupus erythematosus induce an proinflammatory immune response. Arthritis Res. Ther. 2020, 22, 109. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Bai, M.; Deng, T.; Liu, R.; Wang, X.; Qu, Y.; Duan, J.; Zhang, L.; Ning, T.; Ge, S.; et al. Cell-derived microvesicles mediate the delivery of miR-29a/c to suppress angiogenesis in gastric carcinoma. Cancer Lett. 2016, 375, 331–339. [Google Scholar] [CrossRef] [PubMed]
- Zernecke, A.; Bidzhekov, K.; Noels, H.; Shagdarsuren, E.; Gan, L.; Denecke, B.; Hristov, M.; Koppel, T.; Jahantigh, M.N.; Lutgens, E.; et al. Delivery of microRNA-126 by apoptotic bodies induces CXCL12-dependent vascular protection. Sci. Signal 2009, 2, ra81. [Google Scholar] [CrossRef] [PubMed]
- Gasparello, J.; Lomazzi, M.; Papi, C.; D’Aversa, E.; Sansone, F.; Casnati, A.; Donofrio, G.; Gambari, R.; Finotti, A. Efficient delivery of microRNA and antimiRNA molecules using an argininocalix[4]arene macrocycle. Mol. Ther. Nucleic Acids 2019, 18, 748–763. [Google Scholar] [CrossRef]
- Gasparello, J.; Manicardi, A.; Casnati, A.; Corradini, R.; Gambari, R.; Finotti, A.; Sansone, F. Efficient cell penetration and delivery of peptide nucleic acids by an argininocalix[4]arene. Sci. Rep. 2019, 9, 3036. [Google Scholar] [CrossRef]
- Soto-Sanchez, C.; Martinez-Navarrete, G.; Humphreys, L.; Puras, G.; Zarate, J.; Pedraz, J.L.; Fernandez, E. Enduring high-efficiency in vivo transfection of neurons with non-viral magnetoparticles in the rat visual cortex for optogenetic applications. Nanomedicine 2015, 11, 835–843. [Google Scholar] [CrossRef]
- Titze de Almeida, S.S.; Horst, C.H.; Soto-Sanchez, C.; Fernandez, E.; Titze de Almeida, R. Delivery of miRNA-targeted oligonucleotides in the rat striatum by magnetofection with neuromag((R)). Molecules 2018, 23, 1825. [Google Scholar] [CrossRef]
- Zhang, Q.; Ding, F.; Liu, X.; Shen, J.; Su, Y.; Qian, J.; Zhu, X.; Zhang, C. Nanobody-guided targeted delivery of microRNA via nucleic acid nanogel to inhibit the tumor growth. J. Control. Release 2020, 328, 425–434. [Google Scholar] [CrossRef]
- Qian, H.; Tay, C.Y.; Setyawati, M.I.; Chia, S.L.; Lee, D.S.; Leong, D.T. Protecting microRNAs from RNase degradation with steric DNA nanostructures. Chem. Sci. 2017, 8, 1062–1067. [Google Scholar] [CrossRef] [PubMed]
- Nahar, S.; Nayak, A.K.; Ghosh, A.; Subudhi, U.; Maiti, S. Enhanced and synergistic downregulation of oncogenic miRNAs by self-assembled branched DNA. Nanoscale 2017, 10, 195–202. [Google Scholar] [CrossRef] [PubMed]
- Ban, E.; Kwon, T.H.; Kim, A. Delivery of therapeutic miRNA using polymer-based formulation. Drug Deliv. Transl. Res. 2019, 9, 1043–1056. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, C.; Sharma, A.R.; Sharma, G.; Doss, C.G.P.; Lee, S.S. Therapeutic miRNA and siRNA: Moving from bench to clinic as next generation medicine. Mol. Ther. Nucleic Acids 2017, 8, 132–143. [Google Scholar] [CrossRef] [PubMed]
- Bikram, M.; Lee, M.; Chang, C.W.; Janat-Amsbury, M.M.; Kern, S.E.; Kim, S.W. Long-circulating DNA-complexed biodegradable multiblock copolymers for gene delivery: Degradation profiles and evidence of dysopsonization. J. Control. Release 2005, 103, 221–233. [Google Scholar] [CrossRef] [PubMed]
- Bertucci, A.; Prasetyanto, E.A.; Septiadi, D.; Manicardi, A.; Brognara, E.; Gambari, R.; Corradini, R.; De Cola, L. Combined delivery of Temozolomide and anti-miR221 PNA using mesoporous silica nanoparticles induces apoptosis in resistant glioma cells. Small 2015, 11, 5687–5695. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Wu, W.; Yang, W.; Qing, X.; Shao, Z. Surface engineering of nanoparticles with ligands for targeted delivery to osteosarcoma. Colloids Surf. B Biointerfaces 2020, 190, 110891. [Google Scholar] [CrossRef]
- Li, M.; Zhang, W.; Wang, B.; Gao, Y.; Song, Z.; Zheng, Q.C. Ligand-based targeted therapy: A novel strategy for hepatocellular carcinoma. Int. J. Nanomed. 2016, 11, 5645–5669. [Google Scholar] [CrossRef]
- Lee, T.J.; Yoo, J.Y.; Shu, D.; Li, H.; Zhang, J.; Yu, J.G.; Jaime-Ramirez, A.C.; Acunzo, M.; Romano, G.; Cui, R.; et al. RNA nanoparticle-based targeted therapy for glioblastoma through inhibition of oncogenic miR-21. Mol. Ther. 2017, 25, 1544–1555. [Google Scholar] [CrossRef]
- Zhang, T.; Xue, X.; He, D.; Hsieh, J.T. A prostate cancer-targeted polyarginine-disulfide linked PEI nanocarrier for delivery of microRNA. Cancer Lett. 2015, 365, 156–165. [Google Scholar] [CrossRef]
- Chen, H.; Li, M.; Wan, T.; Zheng, Q.; Cheng, M.; Huang, S.; Wang, Y. Design and synthesis of dual-ligand modified chitosan as a liver targeting vector. J. Mater. Sci. Mater. Med. 2012, 23, 431–441. [Google Scholar] [CrossRef] [PubMed]
- MacDiarmid, J.A.; Mugridge, N.B.; Weiss, J.C.; Phillips, L.; Burn, A.L.; Paulin, R.P.; Haasdyk, J.E.; Dickson, K.A.; Brahmbhatt, V.N.; Pattison, S.T.; et al. Bacterially derived 400 nm particles for encapsulation and cancer cell targeting of chemotherapeutics. Cancer Cell 2007, 11, 431–445. [Google Scholar] [CrossRef] [PubMed]
- Glover, A.R.; Zhao, J.T.; Gill, A.J.; Weiss, J.; Mugridge, N.; Kim, E.; Feeney, A.L.; Ip, J.C.; Reid, G.; Clarke, S.; et al. MicroRNA-7 as a tumor suppressor and novel therapeutic for adrenocortical carcinoma. Oncotarget 2015, 6, 36675–36688. [Google Scholar] [CrossRef] [PubMed]
- Williams, M.; Kirschner, M.B.; Cheng, Y.Y.; Hanh, J.; Weiss, J.; Mugridge, N.; Wright, C.M.; Linton, A.; Kao, S.C.; Edelman, J.J.; et al. miR-193a-3p is a potential tumor suppressor in malignant pleural mesothelioma. Oncotarget 2015, 6, 23480–23495. [Google Scholar] [CrossRef] [PubMed]
- Hirschi, K.D.; Pruss, G.J.; Vance, V. Dietary delivery: A new avenue for microRNA therapeutics? Trends Biotechnol. 2015, 33, 431–432. [Google Scholar] [CrossRef] [PubMed]
| Delivery System | miRNA | Target Disease | Cellular Targets | Reference |
|---|---|---|---|---|
| Viral Systems | ||||
| Lentiviral-Based Delivery Systems | ||||
| Lentiviral | miR-133b | Spinal cord regeneration | RhoA, Xylt1, Epha7, P2X, P2RX4 | [37] |
| Lentiviral | let-7 | Non-small-cell lung cancer (NSCLC) | RAS, MYC, HMGA2, CDC25A, CDK6, cyclin-D2 | [38] |
| Adeno-associated virus (AAV) serotype 3 | miR-26a miR-122 | Liver tumor | PIK3C2α/Akt/HIF-1α/VEGFA Bcl-2, Bcl-w, Bcl-xl, and Mcl-1 | [39] |
| AAV serotype 9 | miR-298 | Spinal and bulbar muscular atrophy | Androgen receptor | [40] |
| AAV serotype 5 | miATXN3 | Spinocerebellar ataxia type 3 (SCA3) | ATXN3 | [41] |
| Non-Viral Systems | ||||
| Lipid-Based Delivery Systems | ||||
| Lipid nanoparticle | ds-miR-634 | Pancreatic cancer | OPA1, TFAM, APIP, XIAP, and BIRC5, NRF2, LAMP2 | [42] |
| Neutral liposome | miR-34a | Lung cancer | BCL-2, c-Met, KRAS | [43] |
| Cationic liposome | anti-miR-712 | Atherosclerosis | TIMP3, MMPs, ADAMs | [44] |
| Cationic liposome | miR-143 miR-145 | Colorectal carcinoma | ERK5, K-ras, CHEK2 MYCN, FOS, YES, FLI, cyclin D2, cyclin CDK3, MAP3K3, MAPK4K4 | [44] |
| Cationic liposome | miR-7 | Lung cancer | IRS-1, RAF-1, EGFR | [45] |
| Cationic liposome | miR-29b | Lung cancer | CDK6, DNMT3B, MCL1 | [46] |
| Ionizable liposome | miR-200c | Lung cancer | PRDX2, SESN1, GAPB/Nrf2 | [47] |
| Ionizable cationic lipid nanoparticles | miR-199b-5p | Colon, breast, prostate, glioblastoma, medulloblastoma cancer | Hes-1 | [48] |
| Polymeric Delivery Systems | ||||
| Polyethyleneimines (PEI) | miR-145 miR-33a | Colon carcinoma | c-Myc, ERK5 | [49] |
| PEI-PEG | miR-34a | Hepatocellular carcinoma | SNAI1 | [50] |
| PACE polymer | anti-miR-21 | Glioblastoma | PTEN | [51] |
| Polymer micelle | anti-miR-21 | Glioma | PTEN | [52] |
| Inorganic Compound-Based Delivery Systems | ||||
| Carbonate apatite | miR-29b miR-4689 | Colorectal cancers | BCL-2, MCL1 KRAS, AKT1 | [53,54] |
| Exosome-Based Delivery Systems | ||||
| Exosomes | miR-199a-3p | Ovarian cancer | c-Met, mTOR, IKKβ, and CD44 | [55] |
| Exosome-GE11 peptide | let-7 | Breast cancer | HMGA2 | [56] |
| Exosome | miR-122 | Hepatocellular carcinoma | ADAM10, IGF1R, CCNG1 | [57] |
| Exosome | miR-145 | Lung cancer | CDH2 | [58] |
| Developmental Drug | miRNA | Disease | Phase | Agency/Company |
|---|---|---|---|---|
| Miravirsen | anti-miR-122 | Hepatitis C virus infection | II | Santaris Pharma |
| RG-101 | anti-miR-122 | Hepatitis C virus infection | II | Regulus Therapeutics |
| MRX34 | miR-34 | Cancer treatment | I | Mirna Therapeutics |
| RG-012 | anti-miR-21 | Alport syndrome | I | Regulus Therapeutics |
| MesomiR-1 | miR-16 | Malignant pleural mesothelioma or NSCLC | I | EnGeneIC/Asbestos Diseases Research Institute |
| MRG-201 | miR-29 | Scleroderma | I | miRagen Therapeutics |
| MRG-106 | anti-miR-155 | Cutaneous T cell lymphoma | I | miRagen Therapeutics |
| RG-125 | anti-miR-103/107 | Non-alcoholic steatohepatitis | I | Regulus Therapeutics |
| RG-125 (AZD4076) | anti-miR-103/107 | Type 2 diabetes | I | AstraZeneca |
| RGLS4326 | anti-miR-17 | Polycystic kidney disease (PKD) | I | Regulus Therapeutics |
| Cobomarsen (MRG-106) | anti-miR-155 | Cutaneous T-cell lymphoma (CTCL) | I | miRagen therapeutics |
| MRG-110 | anti-miR-92a | Ischemia | I | miRagen therapeutics |
| TargomiRs | miR-16 | Malignant pleural mesothelioma | I | Asbestos Diseases Research Foundation |
| MRG-106 | anti-miR-155 | Cutaneous T cell lymphoma and mycosis fungoides | I | miRagen Therapeutics |
| MRG-107 | anti-miR-155 | Amyotrophic lateral sclerosis (ALS) | Entering clinical trial | miRagen therapeutics |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dasgupta, I.; Chatterjee, A. Recent Advances in miRNA Delivery Systems. Methods Protoc. 2021, 4, 10. https://doi.org/10.3390/mps4010010
Dasgupta I, Chatterjee A. Recent Advances in miRNA Delivery Systems. Methods and Protocols. 2021; 4(1):10. https://doi.org/10.3390/mps4010010
Chicago/Turabian StyleDasgupta, Ishani, and Anushila Chatterjee. 2021. "Recent Advances in miRNA Delivery Systems" Methods and Protocols 4, no. 1: 10. https://doi.org/10.3390/mps4010010
APA StyleDasgupta, I., & Chatterjee, A. (2021). Recent Advances in miRNA Delivery Systems. Methods and Protocols, 4(1), 10. https://doi.org/10.3390/mps4010010





