A Method for the Isolation and Characterization of Mycosporine-Like Amino Acids from Cyanobacteria
Abstract
1. Introduction
2. Experimental Design
2.1. Materials
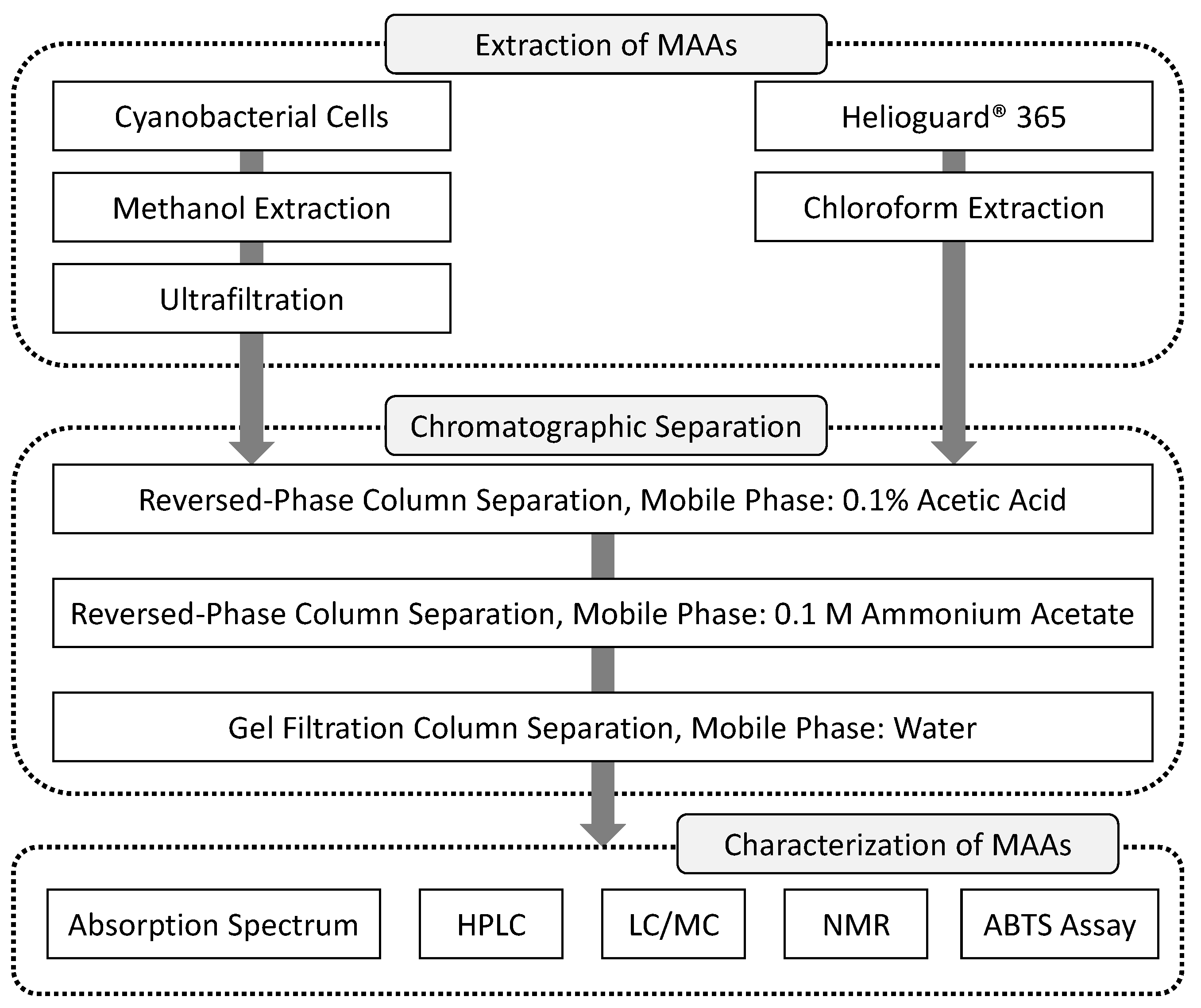
2.1.1. Extraction of MAAs
- Cyanobacterial cells (cyanobacterial cells were available from culture collections, such as the American Type Culture Collection (ATCC), Manassas, VA, USA).
- Helioguard®365 (Mibelle Biochemistry, Buchs, Switzerland)
- HPLC grade methanol (Wako Pure Chemicals Industries, Osaka, Japan; Cat. no. 132-06471)
- Chloroform (Kanto Chemical, Tokyo, Japan; Cat. no. 07278-00)
- Amicon Ultra-4 Ultracel-3K (Merck Millipore, Darmstadt, Germany; Cat. no. UFC800396)
2.1.2. Purification of MAAs
- Acetic acid (Kanto Chemical, Tokyo, Japan; Cat. no. 01021-00)
- Ammonium acetate (Sigma-Aldrich, Tokyo, Japan; Cat. no. 01-4390-5)
- Ethanol (Sigma-Aldrich, Tokyo, Japan; Cat. no. 09-0770-4)
- Nanosep Membrane Filter, 0.45 µm (Pall Life Sciences, MI, USA; Cat. no. ODGHPC34)
- Cosmosil 40C18-PREP (Nacalai Tesque, Kyoto, Japan; Cat. no. 37932-86)
- Sephadex G-10 (GE Healthcare, Uppsala, Sweden; Cat. no. 51185200-EG)
- Polycarbonate chromatography column (18 × 300 mm) (Eyela, Tokyo, Japan; Cat. no. 166130)
- Polycarbonate chromatography column (14 × 500 mm) (Eyela, Tokyo, Japan; Cat. no. 166090)
2.1.3. Characterization of MAAs
LC/MS Analysis
- Methanol for LC/MS (Kanto Chemical, Tokyo, Japan; Cat. no. 25185-79)
- APCI Positive Calibration Solution for the AB Sciex Triple TOF system (AB SCIEX, Framingham, MA, USA; Cat. no. 4460131)
- Triart C18 column (2.1 × 33 mm) (YMC, Kyoto, Japan; Cat. no. TA12S03-H3Q1PTH)
NMR Analysis
- Methanol-d4, for NMR (Acros Organics, NJ, USA; Cat. no. 351460075)
ABTS Assay
- ABTS (Tokyo Chemical Industry, Tokyo, Japan; Cat. no. A2166)
- Potassium peroxodisulfate (Sigma-Aldrich, Tokyo, Japan; Cat. no. 24-5220-2)
- 6-Hydroxy-1.5.7.8-tetramethyl-chroman-2-carboxylic acid (Trolox) (Tokyo Chemical Industry, Tokyo, Japan; Cat. no. H0726)
2.2. Equipment
2.2.1. Extraction of MAAs
- Sonicator VP-5s (Taitec, Tokyo, Japan)
- Rotary evaporator VC-15s (Taitec, Tokyo, Japan)
2.2.2. Purification of MAAs
- Low pressure liquid chromatography system AKTA prime (Amersham Pharmacia Biotech, NJ, USA)
- Freeze dryer FDS-1000 (Eyela, Tokyo, Japan)
- Photospectrometer BioSpec-nano (Shimadzu, Kyoto, Japan)
2.2.3. Characterization of MAAs
Absorption Spectra Analysis
- Photospectrometer BioSpec-nano (Shimadzu, Kyoto, Japan)
Analytical HPLC Analysis
- Analytical HPLC L-2000 system (Hitachi High Technologies, Tokyo, Japan)
LC/MS Analysis
- LC/MS Triple TOF 6600 system (AB SCIEX, Framingham, MA, USA) along with the Nexera XR system (Shimadzu, Kyoto, Japan)
NMR Analysis
- AVANCE III HD 600 NMR spectrometer equipped with a CryoProbe Prodigy (Bruker, Rheinstetten, Germany)
- NMR test tube HG-type (Wako Pure Chemical Industries, Osaka, Japan)
ABTS Assay
- Photospectrometer BioSpec-nano (Shimadzu, Kyoto, Japan)
3. Procedure
3.1. Extraction of MAAs (Time of Completion: 5–6 h for M2G, 2–3 h for SHI and P334)
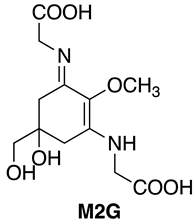
3.1.1. Extraction of M2G from A. halophytica Cells
- Collect A. halophytica cells from liquid cultures. For the induction of M2G bioproduction in A. halophytica, use a BG-11 liquid medium plus Turk Island salt solution, which contains 2.5 M NaCl. Typically, ~3 g fresh weight cells are obtained from a 500 mL culture from cells in the stationary growth phase.
- Add methanol and resuspend the cells. The volume of methanol was determined with the following formula: methanol volume [mL] = cell fresh weight [g] × 8.
- Disrupt the cells with a sonicator. The suspension should be cooled on ice during sonication. Typical parameters for disrupting ~3 g fresh weight cells using the VP-5s instrument comprise the following: output (7), on time (30 s), off time (30 s), and total on time (60 s).
CRITICAL STEP The cells should be completely disrupted.
- Allow the suspension to settle at room temperature for 15 min.
PAUSE STEP The suspension can be stored at 4 °C overnight.
- Centrifuge the suspension at 15,000 × g for 10 min at 25 °C.
- Transfer the supernatant to a new tube.
- Desiccate the supernatant with a rotary evaporator at room temperature. This step takes approximately 2 h.
- Dissolve the dried material in 4.0 mL of water.
- Remove the undissolved compounds by centrifugation at 15,000 × g for 10 min at 25 °C.
- Add 80 µL of chloroform to the supernatant, and shake the tube. In this step, the pigments move to the chloroform phase. With the addition of this amount of chloroform, the color of the solution turns from deep green to clear.
- Centrifuge the samples at 15,000 × g for 10 min at 25 °C.
- Transfer the supernatant (aqueous phase) to an Amicon Ultra-4 Ultracel-3K Centrifugal Filter.
- Centrifuge the filter at 15,000 × g at 25 °C until most of the solution is passed through the membrane. Collect the flow through fraction. This step can remove high molecular weight compounds and takes approximately 1–2 h. The samples should be stored at 4 °C until they are subjected to the separation step.
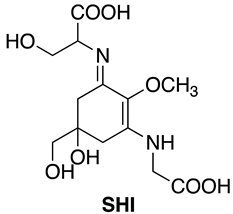
3.1.2. Extraction of SHI and P334 from Helioguard®365
- Concentrate 2.0 mL of Helioguard®365 to a volume of ~200 µL with a rotary evaporator. This step takes approximately 1–2 h.
- Dissolve the concentrated sample in 1.0 mL of water.
- Remove the undissolved compounds by centrifugation at 15,000 × g for 10 min at 25 °C.
- Add 1.0 mL of chloroform to the supernatant, and mix vigorously. Chloroform can disrupt the liposomal forms of SHI and P334. In this step, the MAAs move to the equal amount of the aqueous phase.
- Centrifuge the samples at 15,000 × g for 10 min at 25 °C.
- Transfer the upper phase (aqueous phase) into a new tube.
3.2. Preparation of Columns (Time of Completion: 2–3 h)
3.2.1. Preparation of Reversed-Phase Column
- Open the cap of the empty polycarbonate chromatography column (18 mm × 300 mm) and place it in a vertical position.
- Pour Cosmosil 40C18-PREP resin into the column until it is completely filled with the resin.
- Close the cap, and connect the column to a low-pressure liquid chromatography system.
- Apply at least three column volumes of ethanol to the column.
3.2.2. Preparation of Gel Filtration Column
- Open the cap of the empty polycarbonate chromatography column (14 mm × 500 mm) and place it in a vertical position.
- Soak Sephadex G-10 resin in water for at least 30 min.
- Pour the resin into the column and allow it to settle using gravity. Repeat this step until the column is completely filled with the resin.
- Close the cap, and connect the column to a low-pressure liquid chromatography system.
- Apply at least two column volumes of water to the column.
3.3. Purification of MAAs with a Three-Step Chromatographic Separation (Time of Completion: 3–4 Days)
3.3.1. Separation by Reversed-Phase Chromatography Using 1% Acetic Acid (v/v) as a Mobile Phase
Preparation of Samples
- Before applying the samples to the column, add acetic acid to the MAA samples to set the final concentration to 1% (v/v).
CRITICAL STEP Pass the sample through a filter (pore size: 0.45 µm) if the solution contains insoluble compounds.
Separation
- Before separation, apply at least one column volume (~80 mL) of 1% acetic acid in water to the column.
- After the equilibration of the column, inject the sample (~4 mL for M2G extracted from A. halophytica cells; ~1 mL for SHI and P334 extracted from Helioguard®365) into the column.
- Apply 1% acetic acid to the column at a flow rate of 3.0 mL/min, and collect 2.0 mL fractions.
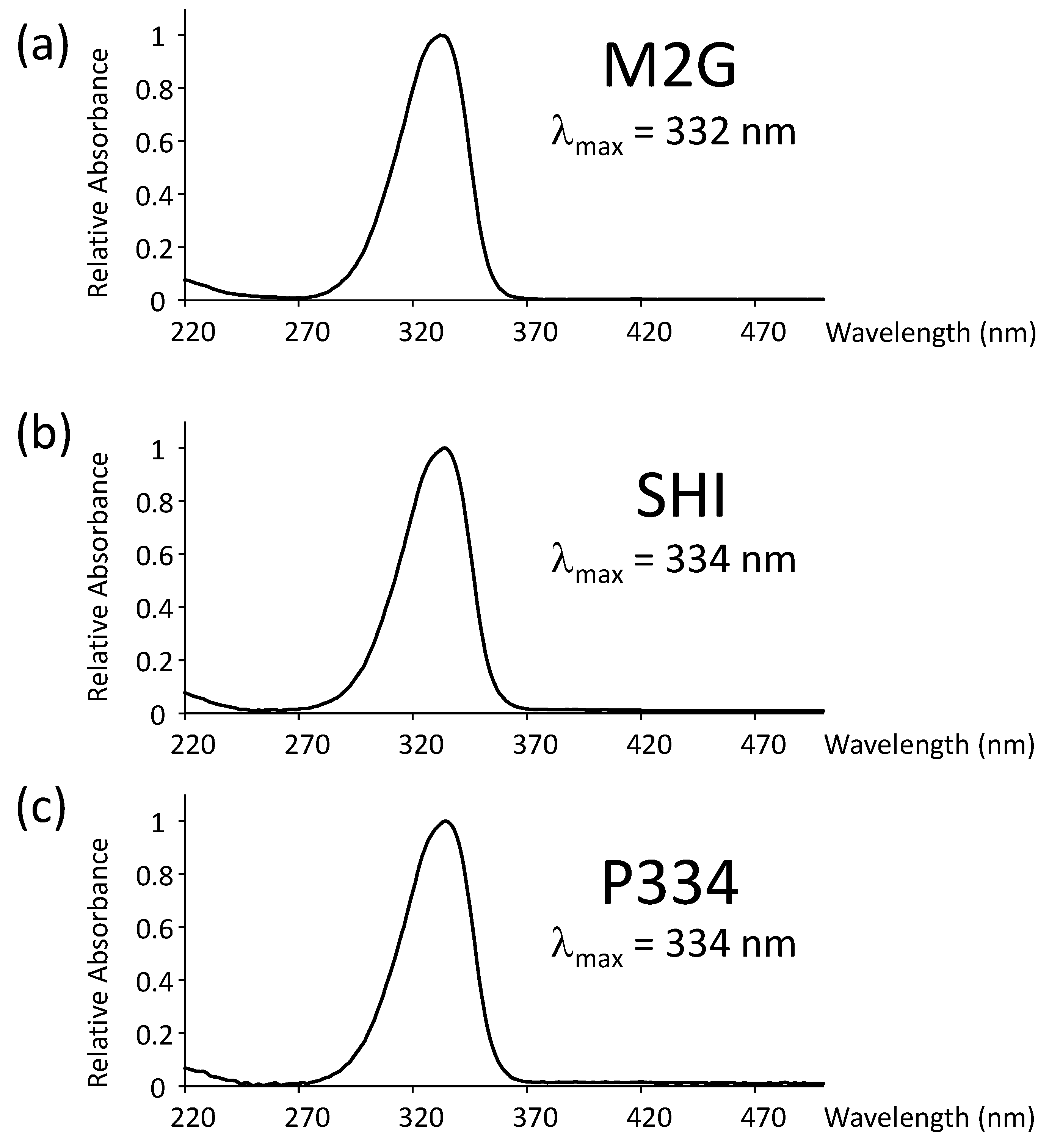
- Measure the absorption of the collected fractions at 330 nm with a photospectrometer. The eluted peaks of M2G, SHI, and P334 were typically found in fractions #38, #35, and #50, respectively.
- After separation, wash the column with 25 mL of 96% ethanol.
CRITICAL STEP Do not use methanol or other organic solvents if a polycarbonate column was used.
- Combine the fractions containing each MAA, and lyophilize them.
3.3.2. Separation by Reversed-Phase Chromatography Using 0.1 M Ammonium Acetate as a Mobile Phase
Preparation of Samples
- Dissolve the lyophilized samples in 4.0 mL of 0.1 M ammonium acetate.
CRITICAL STEP Pass the sample through a filter (pore size: 0.45 µm) if the solution contains insoluble compounds.
Separation
- Before separation, apply at least one column volume (~80 mL) of 0.1 M ammonium acetate to the column.
- After the equilibration of the column, inject the sample (~4 mL) into the column.
- Apply 0.1 M ammonium acetate to the column at a flow rate of 3.0 mL/min and collect 2.0 mL fractions.
- Measure the absorption of the collected fractions at 330 nm with a photospectrometer. The eluted peaks of M2G, SHI, and P334 are typically found in fractions #36, #33, and #50, respectively.
- After separation, wash the column with 25 mL of 96% ethanol.
CRITICAL STEP Do not use methanol or other organic solvents if a polycarbonate column was used.
- Combine the fractions containing each MAA, and lyophilize them.
3.3.3. Separation by Gel Filtration Chromatography
Preparation of Samples
 CRITICAL STEP Pass the sample through a filter (pore size: 0.45 µm) if the solution contains insoluble compounds.
CRITICAL STEP Pass the sample through a filter (pore size: 0.45 µm) if the solution contains insoluble compounds.Separation
- Before separation, apply at least one column volume (~80 mL) of water to the column.
- After the equilibration of the column, inject the sample (~4 mL) into the column.
- Apply water to the column at a flow rate of 2.0 mL/min, and collect 2.0 mL fractions.
- Measure the absorption of the collected fractions at 330 nm with a photospectrometer. The eluted peaks of M2G, SHI, and P334 were typically found in fractions #18–19.
- After separation, wash the column with 80 mL of water.
- Combine the fractions containing each MAA, and lyophilize them.
- The quantitation of purified MAAs were determined using the following molar extinction coefficients: SHI (44,668 M−1 cm−1) and P334 (42,300 M−1 cm−1). The molar extinction coefficient of M2G was assumed to be identical to that of the structurally similar compound SHI, as reported previously [13].
3.4. Characterization of MAAs (Time of Completion: 1–2 Days)
3.4.1. Absorption Spectra Analysis
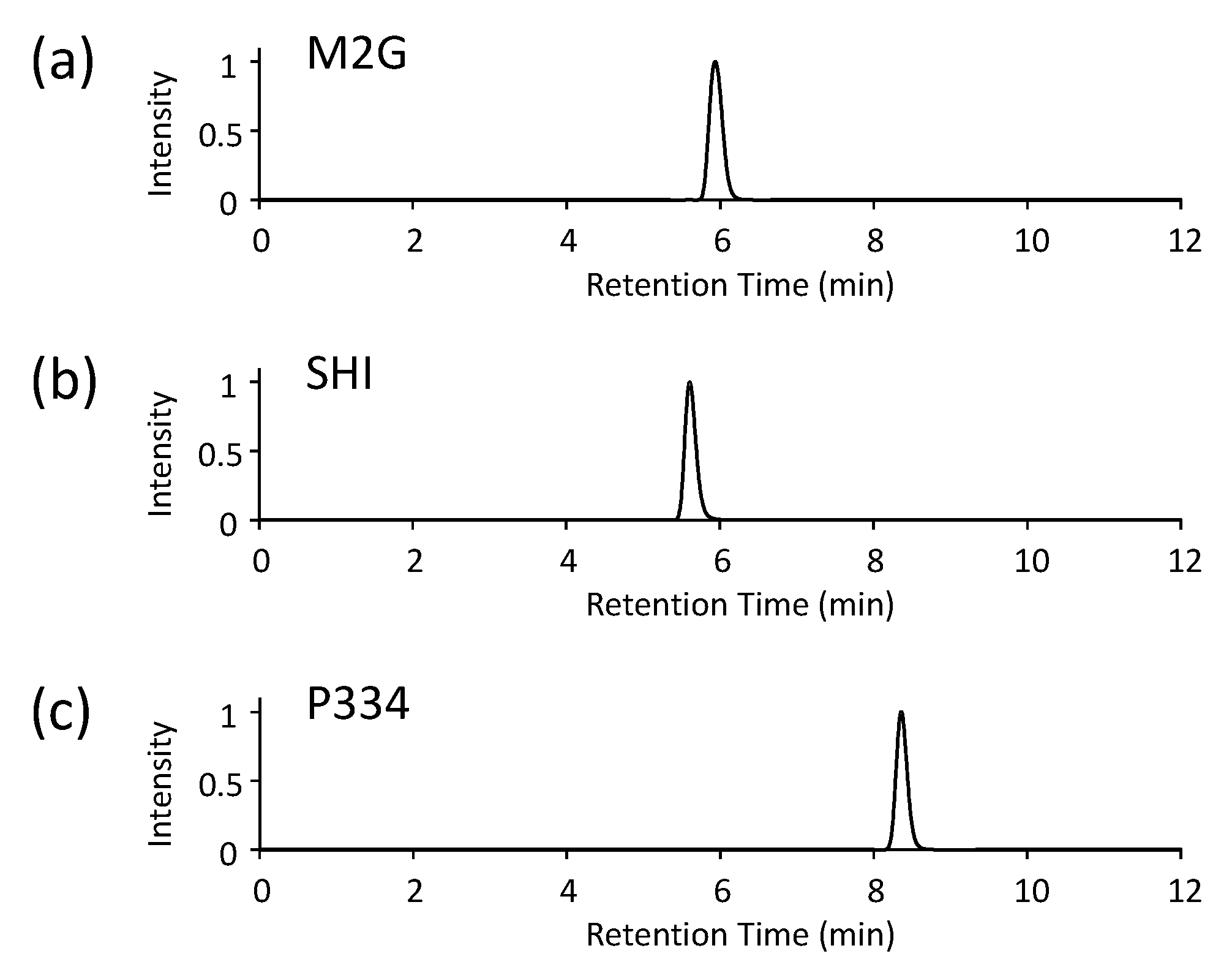
3.4.2. Analytical HPLC
- Analyze the MAAs using an analytical HPLC instrument. All of the MAAs examined in this study can be separated with a typical octadecylsilyl (ODS) column. The separation conditions in this study comprised the following:
- Columns: Shim-pack FC-ODS reversed-phase column (3 µm; 150 × 4.6 mm; Shimadzu, Kyoto Japan) connected to a guard column (30 × 4.6 mm) that contained the same packing material as the main column.
- Injection volume: 10 µL
- Mobile phase: 1% acetic acid (v/v) in water
- Flow rate: 0.4 mL/min
- Separation temperature: 35 °C
- Detection: 330 nm using a UV-visible detector
- If the authentic standards of MAAs are utilized, compare the retention times of the target MAAs to the standards.
3.4.3. LC/MS
- Prepare a 200 nM solution of MAAs by dissolving them in 10% methanol (v/v) in water.
- Mass calibration was conducted using the APCI Positive Calibration Solution (AB Sciex). The conditions for both HPLC and MS comprised the following:
- [HPLC] Nexera XR system (Shimadzu)
- Columns: Triart C18 column (3 µm; 33 × 2.1 mm) (YMC, Kyoto, Japan)
- Injection volume: 2 µL
- Mobile phase: 10% methanol in water (0 min) to 70% methanol in water (3 min) to 100% methanol (7 min); linear gradient
- Flow rate: 0.3 mL/min
- Separation temperature: 35 °C
- [MS] Triple TOF 6600 system
- Source housing: DuoSpray ion source
- Ionization: ESI positive ion mode
- Source temperature: 500 °C
- Experimental type: TOF MS
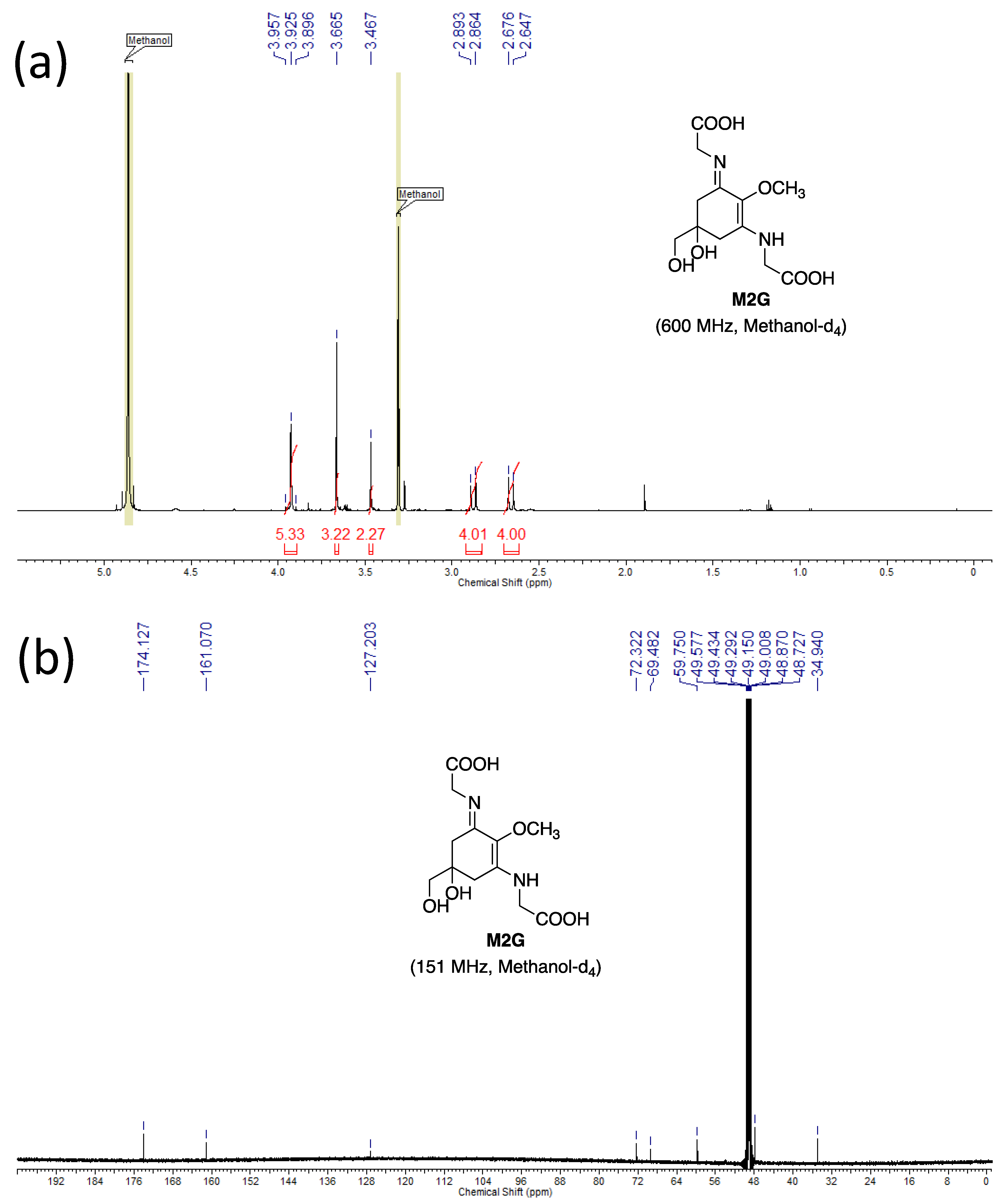
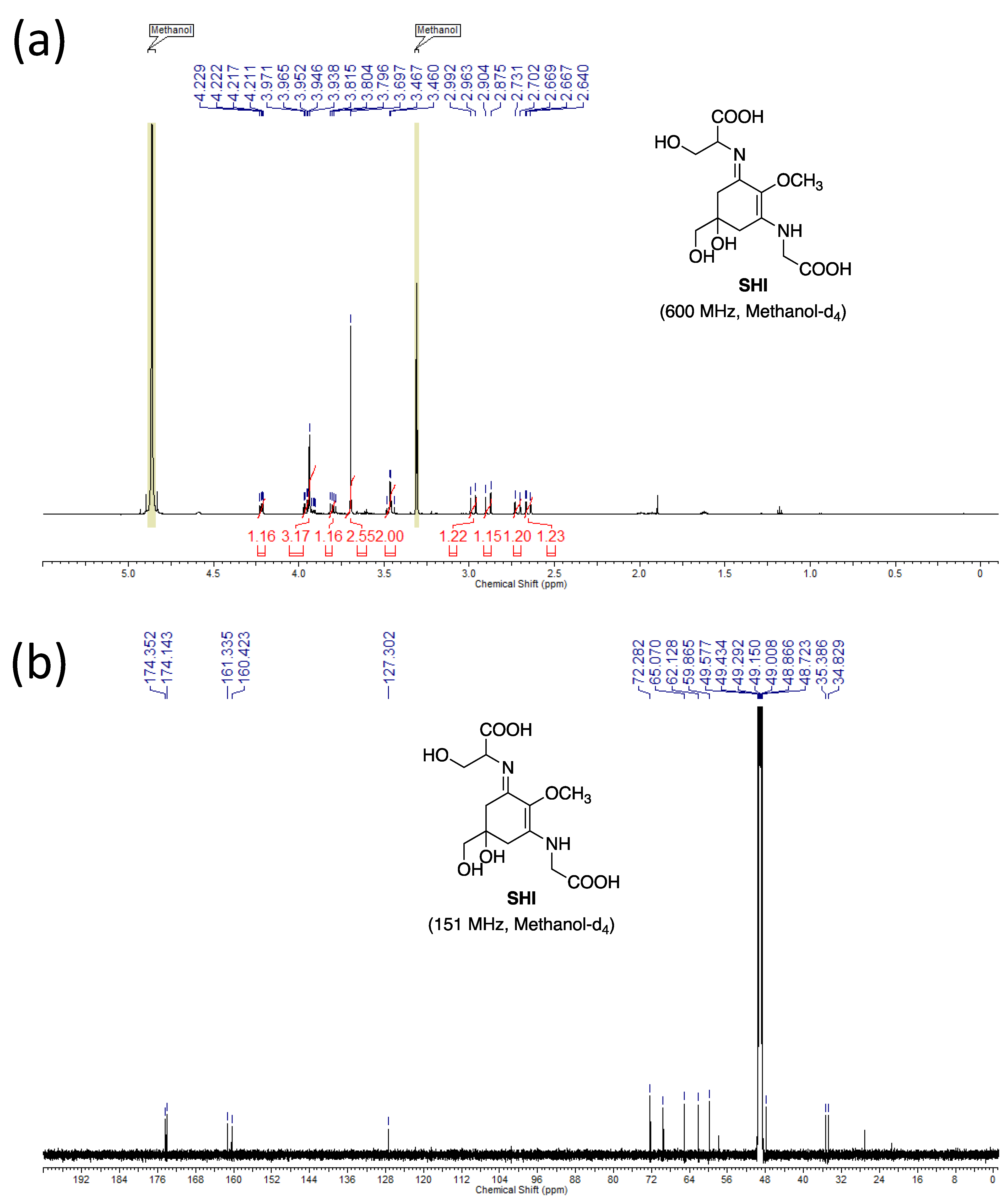
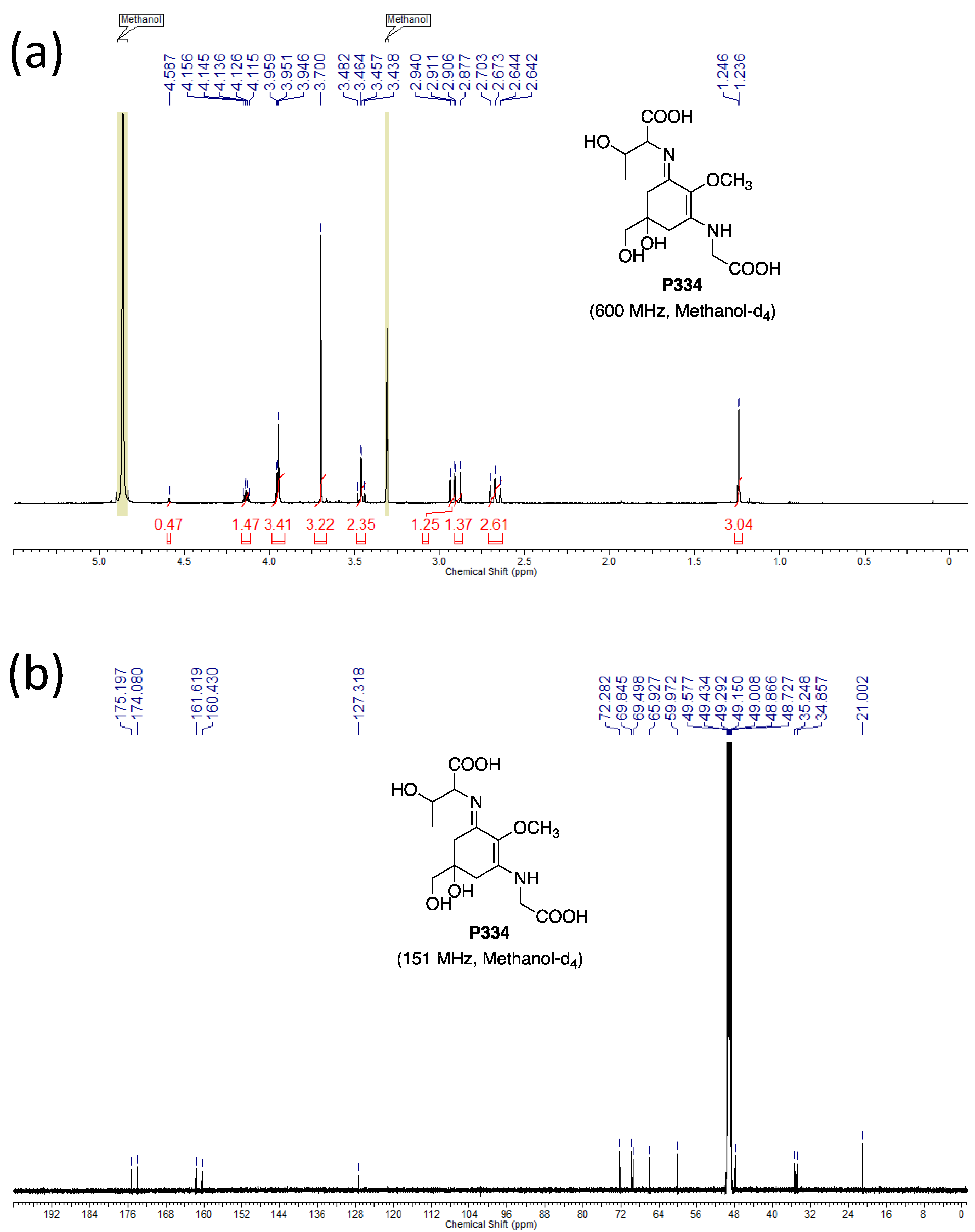
3.4.4. NMR
- Dissolve the samples (~600 µg) in 0.6 mL of methanol-d4.
- Transfer the resulting solution to an NMR test tube.
- Measure the NMR with an AVANCE III HD 600 spectrometer.
- Chemical shift values are reported using residual CD2HOD (δH 3.31) and CD3OD (δC 49.15) for the references. The 1H-NMR spectra are reported as follows: δ (number of protons, multiplicity, coupling constant J Hz). The multiplicities are indicated by s (singlet), d (doublet), and ABq (AB quartet).
3.4.5. ABTS Assay
- Prepare 7 mM ABTS and 2.45 mM potassium persulfate by dissolving in water.
- Mix the 7 mM ABTS solution with the 2.45 mM potassium persulfate solution (1/1, v/v), and allow the mixture to settle for 12–16 h at 25 °C in the dark. During this incubation, the ABTS radical cation (ABTS•+) is produced.
- Dilute the ABTS•+ solution with ethanol to an absorption of approximately 0.70 at 413 nm.
- Mix 18 µL of the diluted ABTS•+ solution with 2 µL of the test samples. In this study, Trolox was used as a positive control.
- Incubate the mixtures at 25 °C for 15 min and measure the absorption at 413 nm.
- Determine the % inhibition of the ABTS radical using the following equation: inhibition % = ((Ab − At)/Ab) × 100, where Ab and At are the absorbances of the blank and tested samples, respectively.
4. Expected Results

Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Wada, N.; Sakamoto, T.; Matsugo, S. Mycosporine-like amino acids and their derivatives as natural antioxidants. Antioxidant 2015, 4, 603–646. [Google Scholar] [CrossRef] [PubMed]
- Kageyama, H.; Waditee-Sirisattha, R. Mycosporine-like amino acids as multifunctional secondary metabolites in cyanobacteria: From biochemical to application aspects. Stud. Nat. Prod. Chem. 2018, 59, 153–194. [Google Scholar]
- Chrapusta, E.; Kaminski, A.; Duchnik, K.; Bober, B.; Adamski, M.; Bialczyk, J. Mycosporine-like amino acids: Potential health and beauty ingredients. Mar. Drugs 2017, 15, 326. [Google Scholar] [CrossRef] [PubMed]
- Cheewinthamrongrod, V.; Kageyama, H.; Palaga, T.; Takabe, T.; Waditee-Sirisattha, R. DNA damage protecting and free radical scavenging properties of mycosporine-2-glycine from the Dead Sea cyanobacterium in A375 human melanoma cell lines. J. Photochem. Photobiol. B 2016, 164, 289–295. [Google Scholar] [CrossRef] [PubMed]
- Patipong, T.; Hibino, T.; Waditee-Sirisattha, R.; Kageyama, H. Efficient bioproduction of mycosporine-2-glycine, which functions as potential osmoprotectant, using Escherichia coli cells. Nat. Prod. Commun. 2017, 12, 1593–1594. [Google Scholar]
- Hartmann, A.; Gostner, J.; Fuchs, J.E.; Chaita, E.; Aligiannis, N.; Skaltsounis, L.; Ganzera, M. Inhibition of collagenase by mycosporine-like amino acids from marine sources. Planta Med. 2015, 81, 813–820. [Google Scholar] [CrossRef] [PubMed]
- Tarasuntisk, S.; Patipong, T.; Hibino, Y.; Waditee-Sirisattha, R.; Kageyama, H. Inhibitory effects of mycosporine-2-glycineisolated from a halotolerant cyanobacteriaumon protein glycation and collagenase activity. Lett. Appl. Microbiol. 2018, 67, 314–320. [Google Scholar] [CrossRef] [PubMed]
- Volkmann, M.; Gorbushina, A.A. A broadly applicable method for extraction and characterization of mycosporines and mycosporine-like amino acids of terrestrial, marine and freshwater origin. FEMS Microbiol. Lett. 2006, 255, 286–295. [Google Scholar] [CrossRef] [PubMed]
- Matsui, K.; Nazifi, E.; Kunita, S.; Wada, N.; Matsugo, S.; Sakamoto, T. Novel glycosylated mycosporine-like amino acids with radical scavenging activity from the cyanobacterium Nostoc commune. J. Photochem. Photobiol. B 2011, 105, 81–89. [Google Scholar] [CrossRef]
- Waditee-Sirisattha, R.; Kageyama, H.; Sopun, W.; Tanaka, Y.; Takabe, T. Identification and upregulation of biosynthetic genes required for accumulation of mycosporine-2-glycine under salt stress conditions in the halotolerant cyanobacterium Aphanothece halophytica. Appl. Environ. Microbiol. 2014, 80, 1763–1769. [Google Scholar] [CrossRef]
- Hu, C.; Völler, G.; Süßmuth, R.; Dittmann, E.; Kehr, J.C. Functional assessment of mycosporine-like amino acids in Microcystis aeruginosa strain PCC 7806. Environ. Microbiol. 2015, 17, 1548–1559. [Google Scholar] [CrossRef] [PubMed]
- Shukri, D.; Darwis, D.; Santoni, A. Preparative HPLC for the Purification of major Anthocyanins from Ficus padana burm. L. Res. J. Chem. Sci. 2013, 3, 60–64. [Google Scholar]
- Shick, J.M.; Dunlap, W.C.; Chalker, B.E.; Banaszak, A.T.; Rosenzweig, T.K. Survey of ultraviolet radiation-absorbing mycosporine-like amino acids in organs of coral reef holothuroids. Mar. Ecol. Prog. Ser. 1992, 90, 139–148. [Google Scholar] [CrossRef]
- Carreto, J.I.; Carignan, M.O. Mycosporine-like amino acids: Relevant secondary metabolites. Chemical and ecological aspects. Mar. Drugs 2011, 9, 387–446. [Google Scholar] [CrossRef] [PubMed]
- Mushir, S.; Fatma, T. Ultraviolet radiation-absorbing mycosporine-like amino acids in cyanobacterium Aulosira fertilissima: Environmental perspective and characterization. Curr. Res. J. Biol. Sci. 2011, 3, 165–171. [Google Scholar]
- Balskus, E.P.; Walsh, C.Y. The genetic and molecular basis for sunscreen biosynthesis in cyanobacteria. Science 2010, 329, 1653–1656. [Google Scholar] [CrossRef] [PubMed]
- de la Coba, F.; Aguilera, J.; Figueroa, F.L.; de Gálvez, M.V.; Herrera, E. Antioxidant activity of mycosporine-like amino acids isolated from three red macroalgae and one marine lichen. J. Appl. Phycol. 2009, 21, 161–169. [Google Scholar] [CrossRef]
| Purification Step | M2G 1 | SHI 2 | P334 2 | |||
|---|---|---|---|---|---|---|
| Amount (µg) | Yield (%) | Amount (µg) | Yield (%) | Amount (µg) | Yield (%) | |
| Methanol extract or Chloroform extract | 781 | 100 | 496 | 100 | 900 | 100 |
| C18 (1% acetic acid) | 498 | 63.8 | 427 | 86.1 | 748 | 83.1 |
| C18 (0.1 M AcONH4) | 357 | 45.7 | 260 | 52.4 | 358 | 39.8 |
| SephadexG-10 | 288 | 36.9 | 227 | 45.8 | 322 | 35.8 |
| Compounds | IC50 1 (µM) |
|---|---|
| M2G | 40 |
| SHI | 94 |
| P334 | 133 |
| Trolox 2 | 10 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ngoennet, S.; Nishikawa, Y.; Hibino, T.; Waditee-Sirisattha, R.; Kageyama, H. A Method for the Isolation and Characterization of Mycosporine-Like Amino Acids from Cyanobacteria. Methods Protoc. 2018, 1, 46. https://doi.org/10.3390/mps1040046
Ngoennet S, Nishikawa Y, Hibino T, Waditee-Sirisattha R, Kageyama H. A Method for the Isolation and Characterization of Mycosporine-Like Amino Acids from Cyanobacteria. Methods and Protocols. 2018; 1(4):46. https://doi.org/10.3390/mps1040046
Chicago/Turabian StyleNgoennet, Siripat, Yasuhiro Nishikawa, Takashi Hibino, Rungaroon Waditee-Sirisattha, and Hakuto Kageyama. 2018. "A Method for the Isolation and Characterization of Mycosporine-Like Amino Acids from Cyanobacteria" Methods and Protocols 1, no. 4: 46. https://doi.org/10.3390/mps1040046
APA StyleNgoennet, S., Nishikawa, Y., Hibino, T., Waditee-Sirisattha, R., & Kageyama, H. (2018). A Method for the Isolation and Characterization of Mycosporine-Like Amino Acids from Cyanobacteria. Methods and Protocols, 1(4), 46. https://doi.org/10.3390/mps1040046







