Rapid Electrophoretic Staining and Destaining of Polyacrylamide Gels
Abstract
:1. Introduction
2. Experimental Design
2.1 Materials and Methods
2.1.1. Gel Electrophoresis
2.1.2. Conventional CBB Staining and Destaining
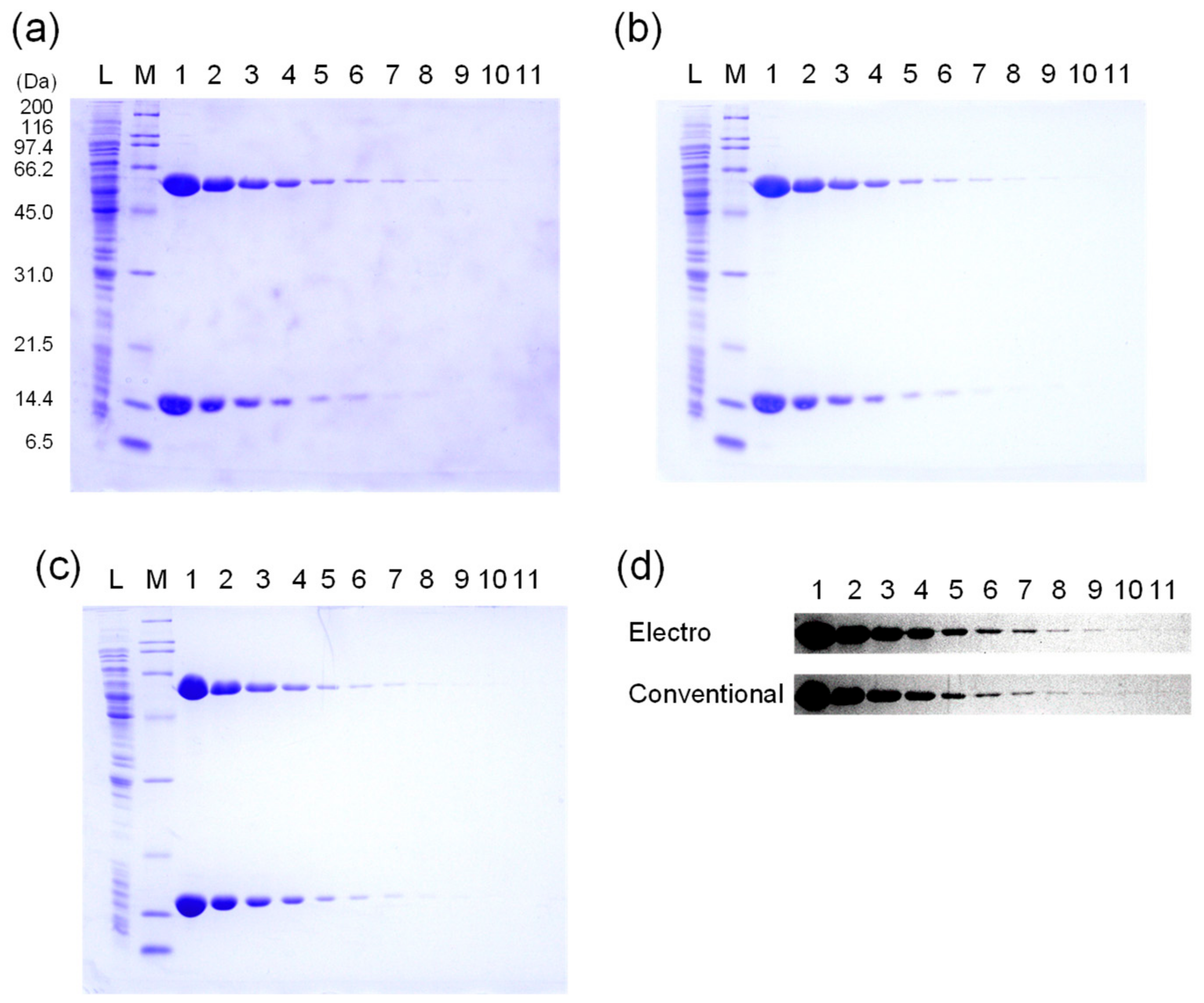
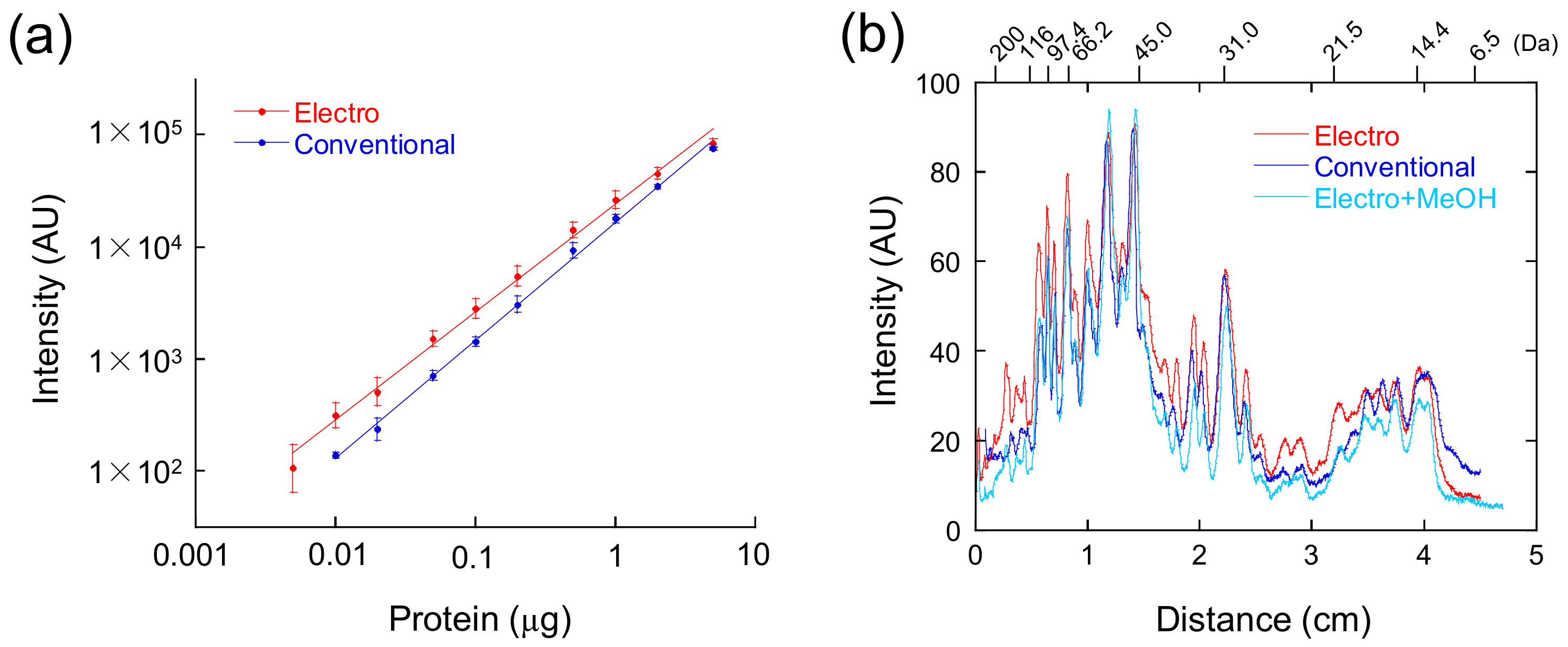
2.1.3. Quantification of CBB-Stained Bands
2.2 Experimental Setup
- A semi-dry transfer unit: The semi-dry transfer unit used in this study had an anode and cathode made of stainless and platinum-coated titanium, respectively (Bio craft, Tokyo, Japan, Cat. No.: BE-320). The carbon electrode may absorb CBB and affect the general use for western blotting.
- High current power supply: MP-3AP (Major science, Saratoga, CA, USA) was used in this study.
- Filter paper was cut to the size of a PAGE gel. In this study, 0.9 mm thick filter paper (ATTO, Tokyo, Japan, Cat. No.: CB-09A) was used.
- The cathode staining solution: 20% (v/v) ethanol, 10% (v/v) acetic acid, 0.1 M glycine, and 0.08% (w/v) CBB-R 250.
- The anode solution: 20% (v/v) ethanol, 10% (v/v) acetic acid, and 0.1 M glycine.
- The destaining solution: 20% (v/v) ethanol and 5% (v/v) acetic acid.
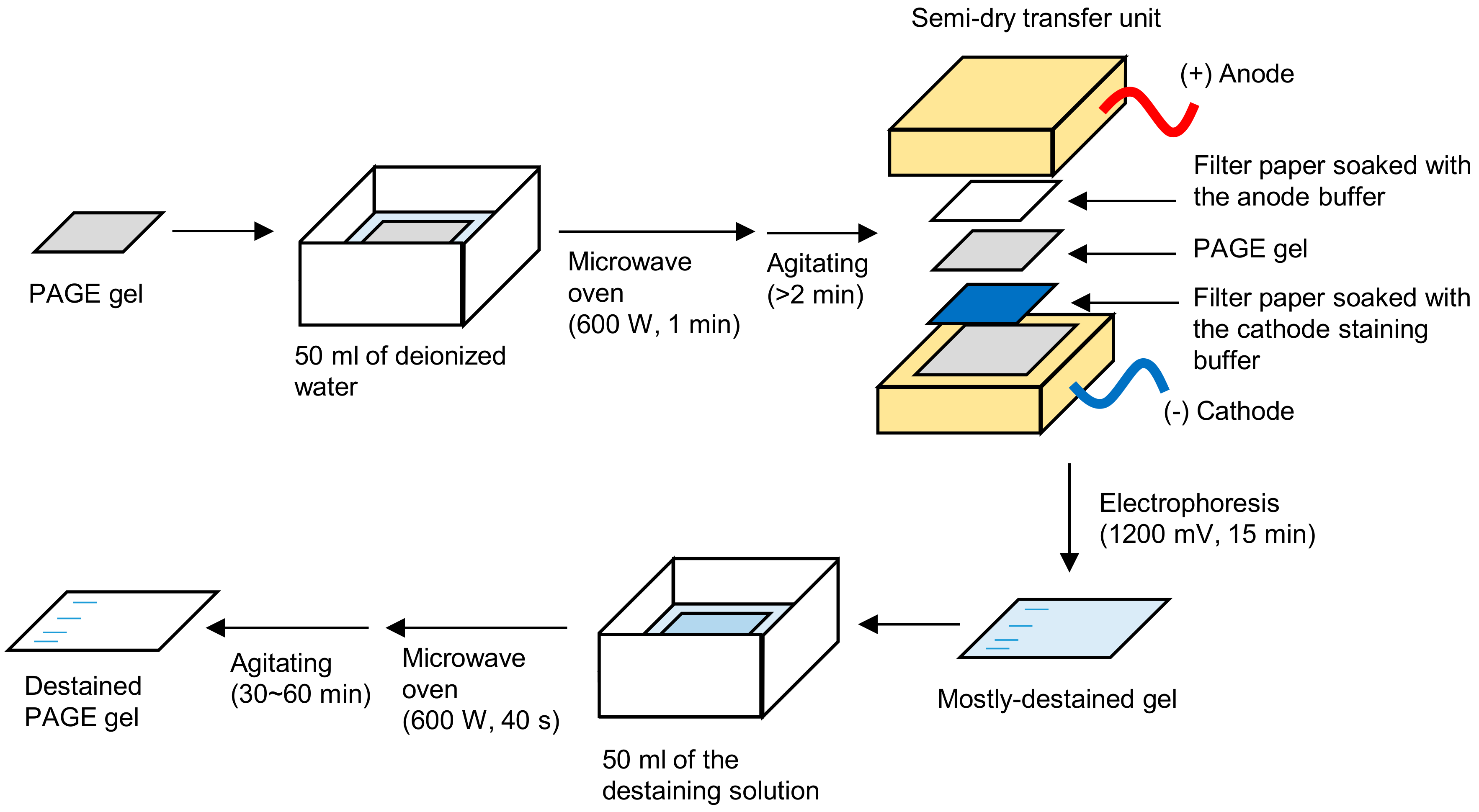
3. Procedure
3.1. Fixing Protein Bands by Boiling. Time for Completion: ~5 min
- Immerse a PAGE gel into 50 mL of deionized water in a plastic container, such as a ZipLoc® container (S. C. Johnson, Racine, WI, USA, Cat. No.: small square, 156 × 156 × 57 mm).
- The lid of the container should be opened to release the pressure from the boiled water.
- Heat the solution in a microwave oven at 600 W for 60 s to boil the solution and fix the proteins. Extend the heating time if the solution did not boil.
- Agitate it for more than 2 min to remove SDS and cool by adding water.
3.2. Electrophoretic-Staining/Destaining. Time for Completion: ~20 min
- Soak the filter paper in the cathode staining solution and spread the solution on the cathode of the semi-dry transfer unit with the filter paper, avoiding air bubbles at the interface.
- Place the soaked filter paper on the cathode.
- Immerse the fixed PAGE gel in the anode solution for a few seconds and place it on the filter paper. Avoid air bubbles between the filter paper and the gel.
- Soak a filter paper in the anode solution and place it on the PAGE gel.
- Spread the anode solution on the anode to avoid air bubbles.
- Place the anode on the filter paper.
- Place a weight of ~1 kg on the semi-dry transfer unit for close contact between the filter paper and electrodes.
- Run at a constant current of 1200 mA for 15 min by using a high current power supply. A higher current generates heat, resulting in bending of the gel and uneven destaining.
- If the CBB remained in the gel, change the arrangement of gel and the filter paper and run for an extra ~5 min.
- Wash the semi-dry transfer unit and cool down the electrode before the next staining.
3.3. Secondary Destaining. Time for Completion: 30–60 min
- Heat the gel in the destaining solution in a microwave oven for 40 s and subsequently agitate at room temperature (~25 °C) for 30–60 min for complete destaining. It takes 1–2 h without heating.
- OPTIONAL STEP: If the background level of CBB in the gel is high, replace the destaining solution containing 0.0001% (w/v) CBB R-250 to avoid excess destaining.
4. Expected Results and Discussion
5. Conclusions
6. Reagents Setup
Acknowledgments
Conflicts of Interest
References
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef] [PubMed]
- Meyer, T.S.; Lamberts, B.L. Use of coomassie brilliant blue R250 for the electrophoresis of microgram quantities of parotid saliva proteins on acrylamide-gel strips. Biochim. Biophys. Acta 1965, 107, 144–145. [Google Scholar] [CrossRef]
- Merril, C.R.; Goldman, D.; Sedman, S.A.; Ebert, M.H. Ultrasensitive stain for proteins in polyacrylamide gels shows regional variation in cerebrospinal fluid proteins. Science 1981, 211, 1437–1438. [Google Scholar] [CrossRef] [PubMed]
- Weber, K.; Osborn, M. The reliability of molecular weight determinations sulfate-polyacrylamide gel electrophoresis. J. Biol. Chem. 1969, 244, 4406–4412. [Google Scholar] [PubMed]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning: A Laboratory Manual, 2nd ed.; CSH Laboratory Press: Cold Spring Harbor, NY, USA, 1989. [Google Scholar]
- Hervieu, G. A quick and safe method for destaining Coomassie-blue-stained protein gels. Tech. Tips Online 1997, 2, 94–96. [Google Scholar] [CrossRef]
- Kurien, B.T.; Scofield, R.H. Heat mediated quick Coomassie blue protein staining and destaining of SDS-PAGE gels. Indian J. Biochem. Biophys. 1998, 35, 385–389. [Google Scholar] [PubMed]
- Kurien, B.T.; Scofield, R.H. Accelerated coomassie blue staining and destaining of SDS-PAGE gels with application of heat. Methods Mol. Biol. 2012, 869, 471–479. [Google Scholar] [CrossRef] [PubMed]
- Fairbanks, G.; Steck, T.L.; Wallach, D.F. Electrophoretic analysis of the major polypeptides of the human erythrocyte membrane. Biochemistry 1971, 10, 2606–2617. [Google Scholar] [CrossRef] [PubMed]
- Wong, C.; Sridhara, S.; Bardwell, J.C.; Jakob, U. Heating greatly speeds Coomassie blue staining and destaining. Biotechniques 2000, 28, 426–432. [Google Scholar] [PubMed]
- Dong, W.H.; Wang, T.Y.; Wang, F.; Zhang, J.H. Simple, time-saving dye staining of proteins for sodium dodecyl sulfate-polyacrylamide gel electrophoresis using coomassie blue. PLoS ONE 2011, 6, 6–8. [Google Scholar] [CrossRef] [PubMed]
- Lawrence, A.-M.; Besir, H.U.S. Staining of proteins in gels with Coomassie G-250 without organic solvent and acetic acid. J. Vis. Exp. 2009, 2–4. [Google Scholar] [CrossRef] [PubMed]
- Kang, D.; Gho, Y.S.; Suh, M.; Kang, C. Highly sensitive and fast protein detection with Coomassie brilliant blue in sodium dodecyl sulfate-polyacrylamide gel electrophoresis. Bull. Korean Chem. Soc. 2002, 23, 1511–1512. [Google Scholar] [CrossRef]
- Dyballa, N.; Metzger, S. Fast and sensitive colloidal coomassie G-250 staining for proteins in polyacrylamide gels. J. Vis. Exp. 2009, 2–5. [Google Scholar] [CrossRef] [PubMed]
- Wenk, M.R.; Fernandis, A.Z. A Manual for Biochemistry Protocols; World Scientific: Singapore, 2007; Volume 3. [Google Scholar]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef] [PubMed]
- Chial, H.J.; Thompson, H.B.; Splittgerber, A.G. A spectral study of the charge forms of Coomassie blue G. Anal. Biochem. 1993, 209, 258–266. [Google Scholar] [CrossRef] [PubMed]
- Motojima, F.; Makio, T.; Aoki, K.; Makino, Y.; Kuwajima, K.; Yoshida, M. Hydrophilic residues at the apical domain of GroEL contribute to GroES binding but attenuate polypeptide binding. Biochem. Biophys. Res. Commun. 2000, 267, 842–849. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Motojima, F. Rapid Electrophoretic Staining and Destaining of Polyacrylamide Gels. Methods Protoc. 2018, 1, 13. https://doi.org/10.3390/mps1020013
Motojima F. Rapid Electrophoretic Staining and Destaining of Polyacrylamide Gels. Methods and Protocols. 2018; 1(2):13. https://doi.org/10.3390/mps1020013
Chicago/Turabian StyleMotojima, Fumihiro. 2018. "Rapid Electrophoretic Staining and Destaining of Polyacrylamide Gels" Methods and Protocols 1, no. 2: 13. https://doi.org/10.3390/mps1020013
APA StyleMotojima, F. (2018). Rapid Electrophoretic Staining and Destaining of Polyacrylamide Gels. Methods and Protocols, 1(2), 13. https://doi.org/10.3390/mps1020013





