Newborn Screening for Severe Combined Immunodeficiency in Taiwan
Abstract
:1. Introduction
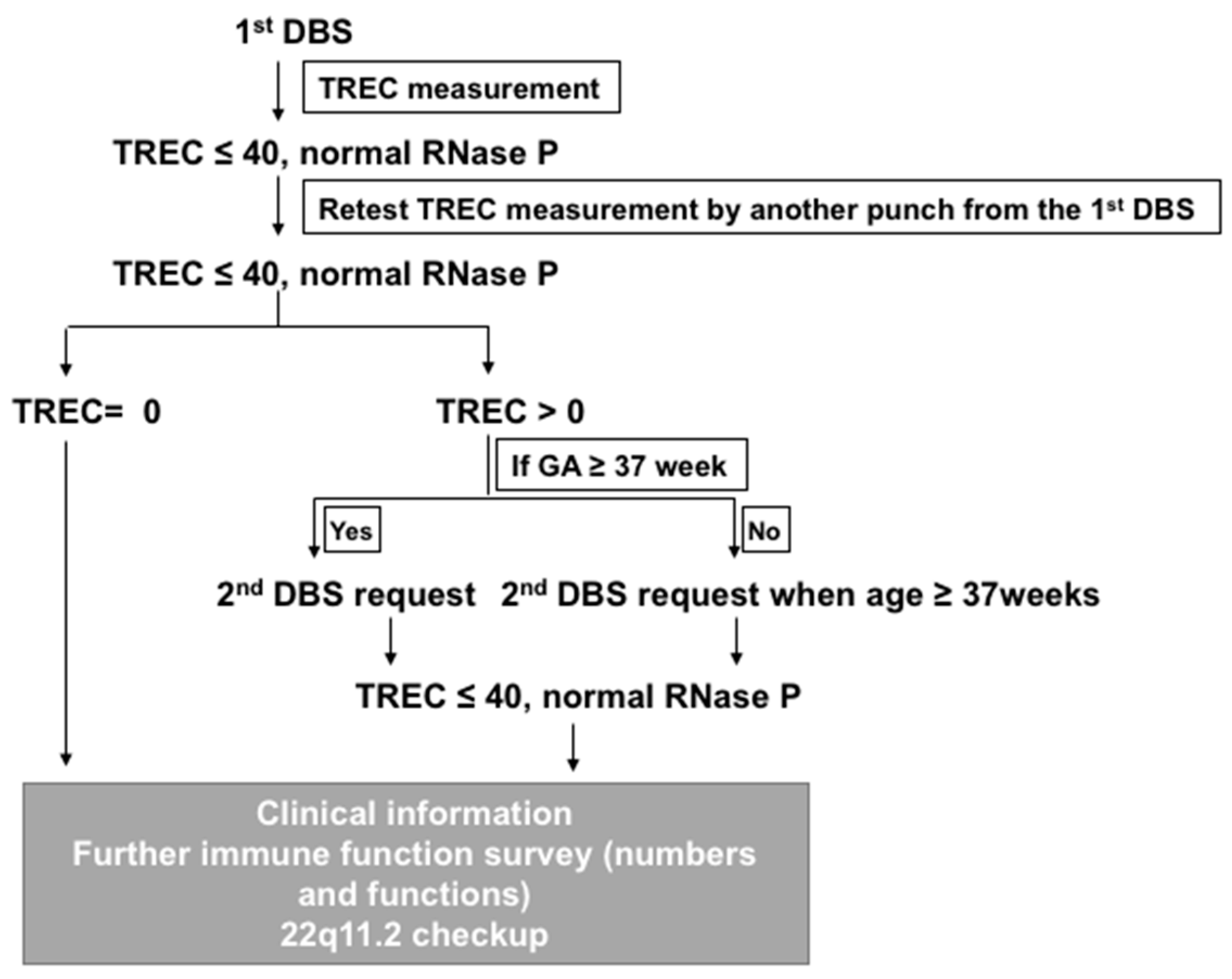
2. Method Development in NTUH
3. Updated Results of Screening
4. 22q11.2 Deletion Syndrome
5. Secondary Etiologies for T-Lymphopenia
6. Rare Conditions
7. Differences in the Methods Used and Their Rates of Detection
8. BCG Policy and Impact
9. Under Evaluation
ADA Deficiency
10. Kappa-Deleting Element Recombination Circle (KREC) Screening
11. Summary
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Kwan, A.; Puck, J.M. History and current status of newborn screening for severe combined immunodeficiency. Semin. Perinatol. 2015, 39, 194–205. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.L.; Lu, M.Y.; Jou, S.T.; Lin, K.H.; Lin, D.T. Hematopoietic stem cell transplantation in Taiwanese children with primary immunodeficiency. J. Formos. Med. Assoc. 2005, 104, 101–106. [Google Scholar] [PubMed]
- Abd Hamid, I.J.; Slatter, M.A.; McKendrick, F.; Pearce, M.S.; Gennery, A.R. Long-term outcome of hematopoietic stem cell transplantation for IL2RG/JAK3 SCID: A cohort report. Blood 2017, 129, 2198–2201. [Google Scholar] [CrossRef] [PubMed]
- Myers, L.A.; Patel, D.D.; Puck, J.M.; Buckley, R.H. Hematopoietic stem cell transplantation for severe combined immunodeficiency in the neonatal period leads to superior thymic output and improved survival. Blood 2002, 99, 872–878. [Google Scholar] [CrossRef] [PubMed]
- Shaw, K.L.; Garabedian, E.; Mishra, S.; Barman, P.; Davila, A.; Carbonaro, D.; Shupien, S.; Silvin, C.; Geiger, S.; Nowicki, B.; et al. Clinical efficacy of gene-modified stem cells in adenosine deaminase-deficient immunodeficiency. J. Clin. Investig. 2017, 127, 1689–1699. [Google Scholar] [CrossRef] [PubMed]
- Hacein-Bey-Abina, S.; Pai, S.Y.; Gaspar, H.B.; Armant, M.; Berry, C.C.; Blanche, S.; Bleesing, J.; Blondeau, J.; de Boer, H.; Buckland, K.F.; et al. A modified gamma-retrovirus vector for X-linked severe combined immunodeficiency. N. Engl. J. Med. 2014, 371, 1407–1417. [Google Scholar] [CrossRef] [PubMed]
- De Ravin, S.S.; Wu, X.; Moir, S.; Anaya-O’Brien, S.; Kwatemaa, N.; Littel, P.; Theobald, N.; Choi, U.; Su, L.; Marquesen, M.; et al. Lentiviral hematopoietic stem cell gene therapy for X-linked severe combined immunodeficiency. Sci. Transl. Med. 2016, 8, 335ra357. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.S.; Shyur, S.D.; Lin, H.Y. Severe combined immunodeficiency with B-lymphocytes (T-B+SCID): Report of two cases. Zhonghua Min Guo Xiao Er Ke Yi Xue Hui Za Zhi 1998, 39, 406–411. [Google Scholar] [PubMed]
- Lee, W.I.; Huang, J.L.; Jaing, T.H.; Shyur, S.D.; Yang, K.D.; Chien, Y.H.; Chiang, B.L.; Soong, W.J.; Chiou, S.S.; Shieh, C.C.; et al. Distribution, clinical features and treatment in Taiwanese patients with symptomatic primary immunodeficiency diseases (PIDs) in a nationwide population-based study during 1985–2010. Immunobiology 2011, 216, 1286–1294. [Google Scholar] [CrossRef] [PubMed]
- Morio, T.; Atsuta, Y.; Tomizawa, D.; Nagamura-Inoue, T.; Kato, K.; Ariga, T.; Kawa, K.; Koike, K.; Tauchi, H.; Kajiwara, M.; et al. Outcome of unrelated umbilical cord blood transplantation in 88 patients with primary immunodeficiency in Japan. Br. J. Haematol. 2011, 154, 363–372. [Google Scholar] [CrossRef] [PubMed]
- Chan, P.C.; Huang, L.M.; Kuo, H.S. Is neonatal bacillus calmette-guerin vaccination protective in Taiwan? J. Formos. Med. Assoc. 2008, 107, 195–197. [Google Scholar] [CrossRef]
- Huang, L.H.; Shyur, S.D.; Weng, J.D.; Shin, C.; Huang, F.Y.; Tzen, C.Y. Disseminated cutaneous bacille calmette-guerin infection identified by polymerase chain reaction in a patient with X-linked severe combined immunodeficiency. Pediatr. Dermatol. 2006, 23, 560–563. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.I.; Huang, J.L.; Yeh, K.W.; Jaing, T.H.; Lin, T.Y.; Huang, Y.C.; Chiu, C.H. Immune defects in active mycobacterial diseases in patients with primary immunodeficiency diseases (PIDs). J. Formos. Med. Assoc. 2011, 110, 750–758. [Google Scholar] [CrossRef] [PubMed]
- Marciano, B.E.; Huang, C.Y.; Joshi, G.; Rezaei, N.; Carvalho, B.C.; Allwood, Z.; Ikinciogullari, A.; Reda, S.M.; Gennery, A.; Thon, V.; et al. BCG vaccination in patients with severe combined immunodeficiency: Complications, risks, and vaccination policies. J. Allergy Clin. Immunol. 2014, 133, 1134–1141. [Google Scholar] [CrossRef] [PubMed]
- Chien, Y.H.; Chiang, S.C.; Chang, K.L.; Yu, H.H.; Lee, W.I.; Tsai, L.P.; Hsu, L.W.; Hu, M.H.; Hwu, W.L. Incidence of severe combined immunodeficiency through newborn screening in a Chinese population. J. Formos. Med. Assoc. 2015, 114, 12–16. [Google Scholar] [CrossRef] [PubMed]
- Chang, T.Y.; Jaing, T.H.; Lee, W.I.; Chen, S.H.; Yang, C.P.; Hung, I.J. Single-institution experience of unrelated cord blood transplantation for primary immunodeficiency. J. Pediatr. Hematol. Oncol. 2015, 37, e191–e193. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.Y.; Chang, Y.L. Adverse events induced by BCG immunization and immunization schedule in Taiwan and in Japan. Taiwan Epidemiol. Bull. 2015, 31, 631–633. [Google Scholar] [CrossRef]
- Baker, M.W.; Grossman, W.J.; Laessig, R.H.; Hoffman, G.L.; Brokopp, C.D.; Kurtycz, D.F.; Cogley, M.F.; Litsheim, T.J.; Katcher, M.L.; Routes, J.M. Development of a routine newborn screening protocol for severe combined immunodeficiency. J. Allergy Clin. Immunol. 2009, 124, 522–527. [Google Scholar] [CrossRef] [PubMed]
- Chan, K.; Puck, J.M. Development of population-based newborn screening for severe combined immunodeficiency. J. Allergy Clin. Immunol. 2005, 115, 391–398. [Google Scholar] [CrossRef] [PubMed]
- Hazenberg, M.D.; Otto, S.A.; Cohen Stuart, J.W.; Verschuren, M.C.; Borleffs, J.C.; Boucher, C.A.; Coutinho, R.A.; Lange, J.M.; Rinke de Wit, T.F.; Tsegaye, A.; et al. Increased cell division but not thymic dysfunction rapidly affects the T-cell receptor excision circle content of the naive T cell population in HIV-1 infection. Nat. Med. 2000, 6, 1036–1042. [Google Scholar] [CrossRef] [PubMed]
- Yokota, M.; Tatsumi, N.; Nathalang, O.; Yamada, T.; Tsuda, I. Effects of heparin on polymerase chain reaction for blood white cells. J. Clin. Lab. Anal. 1999, 13, 133–140. [Google Scholar] [CrossRef]
- Al-Soud, W.A.; Radstrom, P. Purification and characterization of PCR-inhibitory components in blood cells. J. Clin. Microbiol. 2001, 39, 485–493. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.; Huang, C.H.; Cheong, M.L.; Hung, K.L.; Lin, L.H.; Yu, Y.S.; Chien, C.C.; Huang, H.C.; Chen, C.W.; Huang, C.J. Unambiguous molecular detections with multiple genetic approach for the complicated chromosome 22q11 deletion syndrome. BMC Med. Genet. 2009, 10, 16. [Google Scholar] [CrossRef] [PubMed]
- McHugh, D.; Cameron, C.A.; Abdenur, J.E.; Abdulrahman, M.; Adair, O.; Al Nuaimi, S.A.; Ahlman, H.; Allen, J.J.; Antonozzi, I.; Archer, S.; et al. Clinical validation of cutoff target ranges in newborn screening of metabolic disorders by tandem mass spectrometry: A worldwide collaborative project. Genet. Med. 2011, 13, 230–254. [Google Scholar] [CrossRef] [PubMed]
- Kwan, A.; Abraham, R.S.; Currier, R.; Brower, A.; Andruszewski, K.; Abbott, J.K.; Baker, M.; Ballow, M.; Bartoshesky, L.E.; Bonilla, F.A.; et al. Newborn screening for severe combined immunodeficiency in 11 screening programs in the United States. JAMA 2014, 312, 729–738. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.I.; Jaing, T.H.; Hsieh, M.Y.; Kuo, M.L.; Lin, S.J.; Huang, J.L. Distribution, infections, treatments and molecular analysis in a large cohort of patients with primary immunodeficiency diseases (PIDs) in Taiwan. J. Clin. Immunol. 2006, 26, 274–283. [Google Scholar] [CrossRef] [PubMed]
- Shearer, W.T.; Rosenblatt, H.M.; Gelman, R.S.; Oyomopito, R.; Plaeger, S.; Stiehm, E.R.; Wara, D.W.; Douglas, S.D.; Luzuriaga, K.; McFarland, E.J.; et al. Lymphocyte subsets in healthy children from birth through 18 years of age: The pediatric AIDS clinical trials group P1009 study. J. Allergy Clin. Immunol. 2003, 112, 973–980. [Google Scholar] [CrossRef] [PubMed]
- Lee, N.C.; Huang, C.H.; Hwu, W.L.; Chien, Y.H.; Chang, Y.Y.; Chen, C.H.; Ko, T.M. Pseudogene-derived IKBKG gene mutations in incontinentia pigmenti. Clin. Genet. 2009, 76, 417–419. [Google Scholar] [CrossRef] [PubMed]
- Taylor, J.L.; Lee, F.K.; Yazdanpanah, G.K.; Staropoli, J.F.; Liu, M.; Carulli, J.P.; Sun, C.; Dobrowolski, S.F.; Hannon, W.H.; Vogt, R.F. Newborn blood spot screening test using multiplexed real-time PCR to simultaneously screen for spinal muscular atrophy and severe combined immunodeficiency. Clin. Chem 2015, 61, 412–419. [Google Scholar] [CrossRef] [PubMed]
- Borte, S.; Wang, N.; Oskarsdottir, S.; von Dobeln, U.; Hammarstrom, L. Newborn screening for primary immunodeficiencies: Beyond SCID and XLA. Ann. N. Y. Acad. Sci. 2011, 1246, 118–130. [Google Scholar] [CrossRef] [PubMed]
- Chiu, N.C.; Lin, M.C.; Lin, W.L.; Wang, S.Y.; Chi, H.; Huang, L.M.; Tang, R.B.; Huang, Y.C.; Liu, C.C.; Huang, F.Y.; et al. Mycobacterium bovis BCG-associated osteomyelitis/osteitis, Taiwan. Emerg. Infect. Dis. 2015, 21, 539–540. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.I.; Liang, F.C.; Huang, J.L.; Jaing, T.H.; Wang, C.H.; Lin, T.Y.; Huang, Y.C.; Huang, W.L.; Jou, R.; Hsieh, M.Y.; et al. Immunologic analysis of HIV-uninfected Taiwanese children with BCG-induced disease. J. Clin. Immunol. 2009, 29, 319–329. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.Y.; Chang, Y.L. Evolution and determinants of BCG immunization policy: Canada’s experience. Taiwan Epidemiol. Bull. 2016, 32, 351–362. [Google Scholar] [CrossRef]
- La Marca, G.; Canessa, C.; Giocaliere, E.; Romano, F.; Duse, M.; Malvagia, S.; Lippi, F.; Funghini, S.; Bianchi, L.; Della Bona, M.L.; et al. Tandem mass spectrometry, but not T-cell receptor excision circle analysis, identifies newborns with late-onset adenosine deaminase deficiency. J. Allergy Clin. Immunol. 2013, 131, 1604–1610. [Google Scholar] [CrossRef] [PubMed]
- Van Zelm, M.C.; Szczepanski, T.; van der Burg, M.; van Dongen, J.J. Replication history of B lymphocytes reveals homeostatic proliferation and extensive antigen-induced B cell expansion. J. Exp. Med. 2007, 204, 645–655. [Google Scholar] [CrossRef] [PubMed]
- Speckmann, C.; Neumann, C.; Borte, S.; la Marca, G.; Sass, J.O.; Wiech, E.; Fisch, P.; Schwarz, K.; Buchholz, B.; Schlesier, M.; et al. Delayed-onset adenosine deaminase deficiency: Strategies for an early diagnosis. J. Allergy Clin. Immunol. 2012, 130, 991–994. [Google Scholar] [CrossRef] [PubMed]
- Chan, H.Y.; Yang, Y.H.; Yu, H.H.; Chien, Y.H.; Chiang, L.L.; Chiang, B.L. Clinical characteristics and outcomes of primary antibody deficiency: A 20-year follow-up study. J. Formos Med. Assoc. 2014, 113, 340–348. [Google Scholar] [CrossRef] [PubMed]
- Lin, M.T.; Chien, Y.H.; Shyur, S.D.; Huang, L.H.; Chiang, Y.C.; Wen, D.C.; Liang, P.H.; Yang, H.C. De novo mutation in the BTK gene of atypical X-linked agammaglobulinemia in a patient with recurrent pyoderma. Ann. Allergy Asthma Immunol. 2006, 96, 744–748. [Google Scholar] [CrossRef]
- Jou, S.T.; Chien, Y.H.; Yang, Y.H.; Wang, T.C.; Shyur, S.D.; Chou, C.C.; Chang, M.L.; Lin, D.T.; Lin, K.H.; Chiang, B.L. Identification of variations in the human phosphoinositide 3-kinase p110delta gene in children with primary B-cell immunodeficiency of unknown aetiology. Int. J. Immunogenet. 2006, 33, 361–369. [Google Scholar] [CrossRef] [PubMed]
- Sottini, A.; Ghidini, C.; Zanotti, C.; Chiarini, M.; Caimi, L.; Lanfranchi, A.; Moratto, D.; Porta, F.; Imberti, L. Simultaneous quantification of recent thymic T-cell and bone marrow B-cell emigrants in patients with primary immunodeficiency undergone to stem cell transplantation. Clin. Immunol. 2010, 136, 217–227. [Google Scholar] [CrossRef] [PubMed]
- Tsai, H.Y.; Yu, H.H.; Chien, Y.H.; Chu, K.H.; Lau, Y.L.; Lee, J.H.; Wang, L.C.; Chiang, B.L.; Yang, Y.H. X-linked hyper-IgM syndrome with CD40LG mutation: Two case reports and literature review in Taiwanese patients. J. Microbiol. Immunol. Infect. 2015, 48, 113–118. [Google Scholar] [CrossRef] [PubMed]
- Barbaro, M.; Ohlsson, A.; Borte, S.; Jonsson, S.; Zetterstrom, R.H.; King, J.; Winiarski, J.; von Dobeln, U.; Hammarstrom, L. Newborn screening for severe primary immunodeficiency diseases in Sweden-a 2-year pilot TREC and KREC screening study. J. Clin. Immunol. 2017, 37, 51–60. [Google Scholar] [CrossRef] [PubMed]
| Institution | NTUH | CFOH | TIP | Total |
|---|---|---|---|---|
| Duration of study | 78 m | 51 m | 47 m | |
| No. newborns screened | 439,289 | 267,945 | 213,164 | 920,398 |
| No. referred for additional testing (% of total screened) | 105 (0.02%) | 60 (0.02%) | 10 (0.00%) | 175 (0.02%) |
| No. with T lymphopenia (incidence) | 84 (19.12) | 48 (17.91) | 4 (1.88) | 136 (14.78) |
| No. with SCID (incidence) | 5 (1.14) | 2 (0.75) | 0 | 7 (0.76) |
| No. with variant SCID (incidence) | 6 (1.37) | 2 (0.75) | 0 | 8 (0.87) |
| No. with 22q11.2 deletion (incidence) | 18 (4.10) | 1 (0.37) | 1 (0.47) | 20 (2.17) |
| No. of T cell loss | 21 | 1 | 2 | 24 |
| No. of premature infants | 23 | 36 | 0 | 59 |
| No. with other conditions | 10 | 3 | 1 | 14 |
| No | BBW (gm) | GA (wk) | Sex | 1st TREC (Copies/μL) | Age of Confirm (Days) | WBC (/μL) | ALC (/μL) | T Cells (%) | B Cells (%) | NK Cells (%) | T Cells (/μL) | B Cells (/μL) | NK Cells (/μL) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 2890 | 38 | M | 15 | 19 | - | - | - | - | - | - | - | - |
| 2 | 3000 | 40 | M | 15 | 39 | 12,700 | 3556 | 33 | 8 | 55 | 1173 | 284 | 1956 |
| 3 | 1286 | 31 | F | 34 | 32 | 10,050 | 3146 | 57 | 25 | 11 | 1806 | 783 | 359 |
| 4 | 3000 | 37 | M | 17 | 23 | 7520 | 3309 | 75 | 15 | 7 | 2478 | 506 | 245 |
| 5 | 2850 | 35 | M | 31 | 24 | - | - | - | - | - | - | - | - |
| 6 * | 2710 | 38 | F | 13 | 67 | 17,170 | 3795 | 75 | 12 | 5 | 2846 | 444 | 187 |
| 7 | 2300 | 39 | M | 16 | 40 | 13,170 | 5663 | 23 | 46 | 29 | 1314 | 2582 | 1658 |
| 8 | 3200 | 38 | M | 23 | 18 | 7000 | 3003 | 50 | 25 | 20 | 1515 | 742 | 601 |
| 9 | 3350 | 40 | M | 21 | 20 | 11,970 | 3878 | 49 | 27 | 13 | 1903 | 1032 | 491 |
| 10 | 3650 | 38 | M | 28 | 15 | 8140 | 2035 | 44 | 37 | 13 | 898 | 751 | 264 |
| 11 | 1620 | 33 | M | 15 | 43 | 11,110 | 5666 | 27 | 37 | 30 | 1519 | 2094 | 1710 |
| 12 | 2745 | 36 | M | 6 | 19 | 6780 | 1627 | 65 | 27 | 6 | 1059 | 438 | 98 |
| 13 | 3260 | 38 | M | 7 | 22 | - | - | 46 | 20 | 31 | - | - | - |
| 14 | 2294 | 39 | M | 15 | 46 | 6080 | 2402 | 33 | 56 | 8 | 793 | 1345 | 192 |
| 15 | 3420 | 37 | M | 14 | 25 | 6450 | 2838 | 49 | 29 | 15 | 1391 | 823 | 426 |
| 16 * | 2710 | 38 | M | 17 | 79 | 9230 | 2446 | 49 | 17 | 30 | 1199 | 416 | 734 |
| 17 | 2870 | 40 | F | 8 | 29 | 6570 | 3002 | 32 | 46 | 11 | 961 | 1381 | 330 |
| 18 | 2880 | 38 | M | 11 | 23 | 4850 | 2149 | 53 | 16 | 21 | 1139 | 344 | 451 |
| Reference median (10th–90th percentile) [27] | 10,600 (7200–18,000) | 5400 (3400–7600) | 73 (53–84) | 15 (6–32) | 8 (4–18) | 3680 (2500–5500) | 730 (300–2000) | 420 (170–1100) | |||||
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chien, Y.-H.; Yu, H.-H.; Lee, N.-C.; Ho, H.-C.; Kao, S.-M.; Lu, M.-Y.; Jaing, T.-H.; Lee, W.-I.; Chang, K.-W.; Shieh, C.-C.; et al. Newborn Screening for Severe Combined Immunodeficiency in Taiwan. Int. J. Neonatal Screen. 2017, 3, 16. https://doi.org/10.3390/ijns3030016
Chien Y-H, Yu H-H, Lee N-C, Ho H-C, Kao S-M, Lu M-Y, Jaing T-H, Lee W-I, Chang K-W, Shieh C-C, et al. Newborn Screening for Severe Combined Immunodeficiency in Taiwan. International Journal of Neonatal Screening. 2017; 3(3):16. https://doi.org/10.3390/ijns3030016
Chicago/Turabian StyleChien, Yin-Hsiu, Hsin-Hui Yu, Ni-Chung Lee, Hui-Chen Ho, Shu-Min Kao, Meng-Yao Lu, Tang-Her Jaing, Wen-I Lee, Kuei-Wen Chang, Chi-Chang Shieh, and et al. 2017. "Newborn Screening for Severe Combined Immunodeficiency in Taiwan" International Journal of Neonatal Screening 3, no. 3: 16. https://doi.org/10.3390/ijns3030016





